Abstract
Despite the promise of immune checkpoint blockade (ICB) in head and neck squamous cell carcinoma (HNSCC), mediators of response are poorly understood. To address this, here we analyzed oropharyngeal HNSCCs treated with neoadjuvant durvalumab (anti-PDL1) alone or in combination with tremelimumab (anti-CTLA4) from the CIAO clinical trial (NCT03144778). We found that only the total abundance of intratumoral bacteria predicted ICB response, which was validated in multiple independent cohorts. High intratumoral bacteria abundance was associated with an immunosuppressive tumor microenvironment, characterized by an accumulation of neutrophils coupled with depletion of T cells and other adaptive immune cells. Experimental elevation or reduction in intratumoral bacteria abundance in orthotopic models of HNSCC in female mice recapitulated immunological associations observed in participant tumors. Increasing intratumoral bacteria abundance was sufficient to induce resistance to anti-PDL1 ICB, irrespective of bacterial species tested. Together, these findings demonstrate that high intratumoral bacteria abundance is a key suppressor of antitumor immunity and promotes immunotherapy resistance.
Similar content being viewed by others
Main
Immune checkpoint blockade therapy (ICB) has improved clinical outcomes for a subset of persons with head and neck squamous cell carcinoma (HNSCC)1,2. The utility of clinical biomarkers to identify which persons with HNSCC may benefit from ICB remains unclear, including both PDL1 expression3,4 and tumor mutational burden (TMB)3,4. The complexity of processes underlying ICB response highlights the need for deeper mechanistic studies to enhance stratification and optimize immunotherapy treatment strategies. To that end, a growing number of studies have emphasized the importance of the role of the gut microbiome in shaping immunotherapy outcomes5. The relative abundance of specific gut bacteria has been shown to promote antitumor immunity, proliferation of tumor specific T cells and response to immunotherapy in persons with cancer6,7,8. Beyond the gut, bacteria have also emerged as an inherent element of the tumor microenvironment in some cancers9. Studies analyzing the relative microbial composition within tumors provided preclinical and clinical evidence supporting the relevance of the intratumoral microbiome in tumor biology and therapeutic outcomes10,11,12,13,14. However, in contrast to the gut, which harbors 109–1012 bacteria per milligram15, bacteria are typically excluded from the deep tissue parenchyma. While the relative abundance of bacterial species within the microbially rich gut can drive numerous host phenotypes16, in tumors, where total microbial biomass is orders of magnitude lower than in the gut10, the total abundance of ectopic bacteria may exert a larger influence on biology than relative composition. The normal oral and pharyngeal microbiome is microbially rich, comprising over 700 known species17, and pan-cancer analyses have similarly shown that HNSCC tumors harbor a rich microbial community9. Although many studies have focused on the influence of Fusobacterium nucleatum and other specific opportunistic pathogenic bacteria18,19,20,21 on carcinogenesis and progression of malignant disease, the role for total intratumoral bacterial abundance is underexplored.
Here, we examined how the total abundance of intratumoral bacteria corresponds with intratumoral microbial composition, tumor signaling, remodeling of the immune microenvironment and response to ICB. Integrating the analysis of prospective clinical trials and retrospective validation cohorts with experimental model systems, we provide evidence that the total amount of intratumoral bacteria acts as a primary signal to remodel the tumor immune microenvironment and suppress the response to ICB treatment.
Results
Intratumoral bacterial load predicts response to ICB
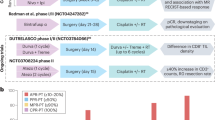
To study the drivers of response to ICB in HNSCC, we analyzed samples from the CIAO clinical trial, which evaluated neoadjuvant durvalumab (anti-PDL1) with or without tremelimumab (anti-CTLA4) in persons with resectable oropharyngeal HNSCC22. We previously reported that response rate was equivalent between single and combination therapy and that CD8 T cells were not associated with response at baseline but an increase in CD8 T cells following treatment was associated with response22. In addition to these observations, we found no differences in treatment response based on tumor stage, human papillomavirus (HPV) status or smoking status among the 28 participants treated (Fig. 1a). Molecular analyses revelated that neither PDL1 combined positivity score nor TMB was significantly associated with ICB response (Fig. 1b,c). Further analysis of the tumor immune microenvironment revealed no immune cell populations significantly associated with ICB response (Fig. 1d). As oropharyngeal tumors are often driven by HPV, which can result in the expression of cancer-specific antigens23,24, we next evaluated whether viral HPV load was associated with response to ICB in the CIAO cohort. We found that there was no significant difference in viral load in responders compared to nonresponders (Fig. 1e). As viral load showed no association with ICB response, we next evaluated whether intratumoral bacterial burden might better predict therapeutic outcome. We found tumor bacteria burden (TBB), defined as the number of bacterial reads per million human reads mapped, was significantly lower in ICB responders than nonresponders (Fig. 1f) and significantly lower for participants that had major or partial pathological response (Fig. 1g). Diversity, as measured by Shannon index, did not significantly differ between responders and nonresponders (Fig. 1h). In a recent study, benefit from immunotherapy in non-small cell lung cancer (NSCLC) was shown to be associated with relative fraction of Fusobacterium25. In samples from the CIAO clinical trial, we found that the relative fraction of Fusobacterium was positively correlated with TBB (Fig. 1i). However, Fusobacterium relative fraction did not predict ICB response in univariate regression analysis (Fig. 1j). In multivariate regression analysis, TBB remained a significant predictor of response after controlling for Fusobacterium (Fig. 1k).
a, Overview of study cohort and association of clinicopathological variables with response to neoadjuvant immunotherapy. P values were determined using a two-tailed Fisher’s exact test, except for stage, which was analyzed by a chi-squared test (n = 28). b, Comparison of PDL1 combined positivity score between responders (PR) and nonresponders (SD and PD). Data are shown as the mean and s.d. Statistical analysis was conducted using a two-tailed Welch’s t-test (responder, n = 15; nonresponder, n = 12). c, Comparison of TMB, defined as mutations per megabase of sequenced DNA, between responders (PR) and nonresponders (SD and PD). Data are shown as the median and interquartile range. Statistical analysis was conducted using a two-tailed rank-sum test (responder, n = 16; nonresponder, n = 12). d, Comparison of immune cell populations from RNA sequencing deconvolution using ssGSEA with signatures from Bindea et al.63,64 between responders (PR) and nonresponders (SD and PD). Statistical analysis was conducted using a two-tailed Welch’s t-test with false discovery rate (FDR) determined by Benjamini–Hochberg procedure (responder, n = 16; nonresponder, n = 12). e, Comparison of HPV viral burden, defined as the number of HPV reads per million human reads, between responders (PR) and nonresponders (SD and PD). Data are shown as the mean and s.d. Statistical analysis was conducted using a two-tailed Welch’s t-test (responder, n = 16; nonresponder, n = 12). f, Comparison of TBB between responders (PR) and nonresponders (SD and PD). Data are shown as the mean and s.d. Statistical analysis was conducted using a two-tailed Welch’s t-test (responder, n = 16; nonresponder, n = 12). g, Comparison of TBB between pathological responders and nonresponders. Data are shown as the mean and s.d. Statistical analysis was conducted using a two-tailed Welch’s t-test (responder, n = 14; nonresponder, n = 11). h, Comparison of intratumoral microbiome diversity, quantified by Shannon diversity index, between responders (PR) and nonresponders (SD and PD). Data are shown as the mean and s.d. Statistical analysis was conducted using a two-tailed Welch’s t-test (responder, n = 16; nonresponder, n = 12). i, Correlation of TBB with relative Fusobacterium. Statistical analysis was conducted using a two-tailed Spearman correlation coefficient (n = 28). j, Univariable regression assessing response to ICB as a function of TBB and relative Fusobacterium. Error bars indicate the 95% confidence interval (responder, n = 16; nonresponder, n = 12). k, Multivariable regression assessing response to ICB as a function of TBB and relative Fusobacterium. Error bars indicate the 95% confidence interval (responder, n = 16; nonresponder, n = 12). TEM cells, effector memory T cells.
Because of recent concerns regarding analysis of microbial content from The Cancer Genome Atlas (TCGA) bulk sequencing data26, we benchmarked multiple approaches to ensure our analysis from bulk sequencing data accurately recapitulated the expected microbiota in HNSCC. When comparing the relative bacterial abundances in oral cavity samples quantified from TCGA whole-genome sequencing (WGS) data using PathSeq to those from our prior study of oral cavity tumors analyzed using 16S ribosomal RNA (rRNA) sequencing12, we observed a high degree of concordance with a correlation coefficient of 0.73 (Extended Data Fig. 1a). In contrast, the method used in the now retracted work by Poore et al.27 shows no relationship with our prior 16S rRNA sequencing data (R = −0.03; Extended Data Fig. 1b). To further improve our analysis, we compared 16S rRNA sequencing of tumors to 16S rRNA sequencing of sham controls and identified numerous known contaminants such as Burkholderia, Propionibacterium and Rhizobium28,29 (Extended Data Fig. 1c). All further analysis was performed using a custom PathSeq index constructed from relevant reference genomes followed by subtraction of identified contaminants (Supplementary Tables 1 and 2). Next, we compared the relative abundances of bacteria recovered when analyzing matched tumors by 16S rRNA sequencing compared to whole-exome sequencing (WES). Overall, we found a strong correlation in average relative abundances, as well as per-sample correlations, at both the phylum and the genus level (Extended Data Fig. 1d–g). Comparing TBB in these samples to total bacterial load determined by qPCR also showed a strong correlation (Extended Data Fig. 1h), which was comparable to the correlation coefficient achieved by comparing 16S rRNA qPCR to 16S rRNA in situ staining (Extended Data Fig. 1i) and approached the correlation coefficient obtained by duplicate 16S rRNA in situ staining (Extended Data Fig. 1j).
To validate the association of TBB with ICB outcomes observed in the CIAO study analysis, we next examined an internal cohort of persons with HNSCC treated with ICB and analyzed by WES. We also found that high TBB was associated with poor outcomes following immunotherapy (Fig. 2a) but did not find an association between Fusobacteria and outcomes (Fig. 2b). To establish the generalizability of this observation, we analyzed the Hartwig NSCLC cohort, where high relative Fusobacterium was shown to predict resistance to ICB25. For these analyses, we compared the ability of total intratumoral bacteria, as quantified by TBB, to predict ICB resistance to both relative Fusobacterium and total Escherichia. In contrast to TBB and Fusobacterium, elevated Escherichia was recently shown to be associated with improved ICB response in NSCLC by Elkrief et al.30. We found that both TBB and relative Fusobacterium were significantly associated with resistance to ICB in NSCLC but did not detect an association between Escherichia and improved ICB response (Fig. 2c). Comparison of the three variables using a receiver operating characteristic curve suggested that TBB provides better predictive performance, with an area under the curve (AUC) of 0.74 compared to an AUC of 0.58 for relative Fusobacterium and 0.40 for total Escherichia (Fig. 2d). Additionally, stratifying tumors by Escherichia positivity to mirror the original comparison performed in Elkrief et al.30 also demonstrated a nonsignificant trend toward lower response in Escherichia-positive tumors, in contrast to the previously reported association with improved response30 (Fig. 2e). Performing multivariate regression analysis for ICB response rate as a function of TBB, relative Fusobacterium and total Escherichia in a combined cohort of NSCLC, HNSCC, urothelial carcinoma, mesothelioma and small intestine or colorectal cancer samples from the Hartwig cohort25, we found that TBB maintained predictive value but Fusobacterium was no longer significant after controlling for TBB (Fig. 2f). Repeating this analysis in tumors treated with either chemotherapy or targeted therapy indicated that neither TBB nor Fusobacterium was associated with response (Fig. 2g), suggesting that this effect is specific to immunotherapy.
a, Progression-free survival in an internal cohort of participants with HNSCC treated with immunotherapy stratified by median TBB determined from WES. Statistical analysis was conducted using a two-tailed log-rank test (n = 19). HR, hazard ratio. b, Progression-free survival in an internal cohort of participants with HNSCC treated with immunotherapy stratified median Fusobacteria abundance from using WES. Statistical analysis was conducted using a two-tailed log-rank test (n = 19). c, Regression coefficient for TBB, relative Fusobacterium or total Escherichia to predict response (CR or PR) to ICB in Hartwig NSCLC samples. Values were log-transformed and z-normalized before regression. Cancer type and biopsy site were taken as fixed effects and the sequencing platform was taken as a random effect. Error bars indicate the 95% confidence interval (n = 72). d, Receiver operating characteristic curve for ability of TBB, relative Fusobacterium or total Escherichia to predict response (CR or PR) to ICB in Hartwig NSCLC samples (responder, n = 12; nonresponder, n = 60). e, Percentage of responders (CR or PR) to ICB in Hartwig NSCLC samples stratified by positivity for Escherichia. Inset numbers indicate the number of responders among total participants. Statistical analysis was conducted using a two-tailed Fisher’s exact test. f, Multivariable regression for ability of TBB, relative Fusobacterium and total Escherichia to predict response to ICB in Hartwig NSCLC, HNSCC, urothelial carcinoma, mesothelioma and small intestine or colorectal cancer samples. Values were log-transformed and z-normalized before regression. Cancer type and biopsy site were taken as fixed effects and the sequencing platform was taken as a random effect. Error bars indicate the 95% confidence interval (responder, n = 23; nonresponder, n = 122). g, Multivariable regression for ability of TBB, relative Fusobacterium and total Escherichia to predict response (CR or PR) to chemotherapy or targeted therapies in Hartwig NSCLC, HNSCC, urothelial carcinoma, mesothelioma and small intestine or colorectal cancer samples. Values were log-transformed and z-normalized before regression. Cancer type and biopsy site were taken as fixed effects and the sequencing platform was taken as a random effect. Error bars indicate the 95% confidence interval (responder, n = 69; nonresponder, n = 273).
Landscape of intratumoral bacteria burden
To further understand the overall landscape of total intratumoral bacteria burden and associated molecular phenotypes, we analyzed microbiome composition and abundance extracted from next-generation sequencing data of HNSCC from TCGA9. Given that TCGA samples were collected and sequenced across multiple centers, we evaluated whether this could introduce technical artifacts. We first compared TBB determined from WGS and WES data generated independently from the same tumors at different sequencing sites, observing an extremely robust correlation (R = 0.88; Extended Data Fig. 2a). TBB values determined by WES were appreciably lower than those determined by WGS, consistent with WES enriching for human reads while retaining ~20–40% in nonenriched regions from imperfect capture31. A strong correlation between WES-based and WGS-based microbial analysis was also observed when comparing relative abundances of specific taxa (Extended Data Fig. 2b,c). We next compared whether TBB differed significantly by cancer center of origin or sequencing site and found no significant alterations using WGS data (Extended Data Fig. 2d,e). Likewise, no variation in TBB associated with cancer center of origin were noted for WES samples, which were all sequenced at a single site (Extended Data Fig. 2f). Together, these results indicate TBB was not impacted by technical or site variations.
Analysis of TCGA WGS data revealed a significant inverse relationship between bacterial diversity and TBB (Fig. 3a). Consistent with findings from the CIAO cohort, relative Fusobacterium was positively correlated with TBB in the full TCGA HNSCC dataset and when restricted to oropharynx cancer samples (Extended Data Fig. 3a,b). Beyond Fusobacterium, analysis of overall shifts in the microbiome landscape revealed that the relative abundances of additional tumor-associated bacteria (for example, Campylobacter and Treponema) were also positively correlated with TBB, whereas other bacteria (for example, Streptococcus, Rothia and Actinomyces) exhibited a negative correlation with TBB (Fig. 3b). When correlating TBB with the total abundance of intratumoral bacteria, we again observed Fusobacterium to be the most strongly correlated. However, in contrast to relative abundance, the total abundance of 19 of 31 genera was significantly positively correlated with TBB, in contrast to only one genus significantly negatively correlated with TBB (Fig. 3c). Notably, TBB was positively correlated with both the absolute level and the relative fraction of F. nucleatum across multiple cancer sites (Fig. 3d). Stratifying bacteria detected on the basis of aerophilicity, we found that the majority of bacteria were anaerobic (~77%; Extended Data Fig. 3c). TBB was positively correlated with the total abundance of both anaerobic and aerobic bacteria (Extended Data Fig. 3d,e), as well as the relative abundance of anaerobic bacteria (Extended Data Fig. 3f,g). The correlation of TBB with over half of the bacterial genera, including both aerobes and anaerobes, and not only highly studied taxa such as Fusobacterium further supports the hypothesis that intratumoral bacterial abundance is a primary effector within the tumor microenvironment.
a, Correlation of WGS TBB with intratumoral microbiome diversity. Statistical analysis was conducted using a two-tailed Spearman correlation coefficient (n = 130). b, Correlation of WGS TBB with relative fractions of indicated genera, quantified as fraction of reads for each genera among all mapped bacterial reads. Statistical analysis was conducted using a two-tailed Spearman correlation coefficient. Highlighted genera indicate an FDR < 0.05 (n = 130). c, Correlation of WGS TBB with absolute abundances of indicated genera, quantified as the number of reads per million human reads. Statistical analysis was conducted using a two-tailed Spearman correlation coefficient. Highlighted genera indicate an FDR < 0.05 (n = 130). d, Correlation of WGS TBB with relative fraction of F. nucleatum across cancers (HNSCC, n = 157; stomach adenocarcinoma (STAD), n = 128; esophageal carcinoma (ESCA), n = 62; colorectal adenocarcinoma and rectal adenocarcinoma (COADREAD), n = 170). Statistical analysis was conducted using a two-tailed Spearman correlation coefficient. e–g, WGS TBB in tumors from participants with HNSCC analyzed on the basis of pathological T stage (e; T1, n = 15; T2, n = 41; T3, n = 31; T4, n = 49), tumor location (f; oral cavity, n = 106; larynx, n = 24; oropharynx, n = 27) or tumor HPV status (g; HPV-negative, n = 110; HPV-positive, n = 38). The center white point is the median and the box is the interquartile range. Statistical analysis was conducted using a Kruskal–Wallis (e,f) or two-tailed rank-sum test (g). h, Difference in WES TBB in HNSCC tumors based on mutations in specific genes relevant to HNSCC. Statistical analysis was conducted using a two-tailed Welch’s t-test. Highlighted values indicate P < 0.05 (n = 507). i, Correlation between WES TBB and protein expression from reverse-phase protein array targeted proteomics in HNSCC tumors. Highlighted values indicate P values < 0.05 (n = 349). Statistical analysis was conducted using a two-tailed Spearman correlation coefficient. j, GSEA of genes correlated with WES TBB using Hallmark gene sets. Highlighted values indicate an FDR < 0.05 (n = 503).
Assessment of TBB across HNSCC tumors found no differences associated with tumor size based on pathological T stage (Fig. 3e), hypoxia score (Extended Data Fig. 4a), pathological and clinical stage (Extended Data Fig. 4b,c), tumor histologic grade (Extended Data Fig. 4d,e) or demographics (Extended Data Fig. 4f–h). When comparing tumors on the basis of head and neck subsite, we found that TBB was highest in oral cavity tumors and lowest in oropharynx tumors, with intermediate values for larynx tumors (Fig. 3f). No differences were observed between subsites within the oral cavity (Extended Data Fig. 4i). HPV-positive tumors exhibited significantly lower TBB compared to all HPV-negative HNSCCs, as well as when the HPV status was restricted to the oropharyngeal subsite (Fig. 3g and Extended Data Fig. 4j). No significant associations were observed between TBB and additional risk factors, including alcohol usage and smoking, according to univariate analysis (Extended Data Fig. 4k,l). However, multivariate analysis of HPV and smoking status indicated that current smokers had elevated TBB compared to never smokers (Extended Data Fig. 4m).
To improve statistical power in assessing molecular alterations associated with TBB in HNSCC, we used TBB values derived from WES for subsequent analyses, given its strong correlation with WGS-derived TBB in overall bacteria burden measurements (Extended Data Fig. 2a). Assessing common HNSCC mutations, we found that TBB was higher in tumors harboring NOTCH1 and CDKN2A mutations and lower in tumors harboring mutations in NFE2L2 (NRF2) and CTNNB1 (Fig. 3h). To control for potential confounding variables from the differential mutation landscape between HPV-positive and HPV-negative HNSCCs, we repeated this analysis with only HPV-negative tumors or as a multivariate analysis controlling for HPV status and found largely concordant results (Extended Data Fig. 5). The association of high TBB in NOTCH-active tumors was also observed at the protein level, where TBB was positively correlated with cleaved NOTCH1 (Fig. 3i). TBB was also positively correlated with immunomodulatory proteins cyclooxygenase 2 and macrophage inhibitory factor and the epithelial cell pattern recognition receptor EphA2. Additional gene expression analysis revealed further evidence of TBB activating inflammatory signaling, with gene set enrichment analysis (GSEA) demonstrating that high-TBB tumors were associated with upregulation of pathways associated with inflammatory signaling by both tumor necrosis factor and interferon-α (Fig. 3j).
High intratumoral bacteria abundance corresponds with an immunosuppressive tumor microenvironment
On the basis of the observed associations of TBB with ICB sensitivity and inflammatory signaling, we evaluated the relationship between TBB and immune cells within the tumor immune microenvironment. TBB from WGS in HNSCC tumors was strongly associated with an increase in neutrophil abundance and reduction in adaptive antitumor immune cells, including both T cells and B cells (Fig. 4a). We validated this result using independent HNSCC tumors from TCGA profiled by only WES not included in the WGS cohort (for example, nonoverlapping samples), finding highly concordant changes in the tumor microenvironment associated with high TBB (Fig. 4b). This result was further validated in samples from the CIAO clinical trial (Fig. 4c). As TCGA WGS and WES sequencing were performed independently and at different research centers, the highly concordant observation in the primary TCGA WGS analysis, secondary TCGA analysis of independent samples only profiled by WES and final validation in the independent CIAO cohort strongly suggests the immune correlations are nonspurious and unlikely to be because of research site-specific contamination. To summarize the observed immune microenvironment remodeling, we determined the intratumoral neutrophil-to-lymphocyte ratio (tNLR), defined as the inferred ratio of neutrophils to T cells and B cells. TBB was significantly positively correlated with tNLR in CIAO tumor samples (Fig. 4d) and tNLR was associated with worse response to ICB in the CIAO cohort (Fig. 4e). The positive correlation between tNLR and TBB was validated in TCGA HNSCC samples and was generalizable to multiple other cancer types (Fig. 4f). To confirm concordance of tumor immune cell infiltration with TBB using an orthogonal approach, we stained a validation cohort of oral cavity HNSCC tumors for T cells (CD3), neutrophils (CD66b) and bacterial (16S rRNA) fluorescence in situ hybridization (FISH). Results in our validation imaging-based cohort mirrored sequencing-based analyses, with high abundance of intratumoral bacteria being associated with low T cell infiltration (Fig. 4g), high neutrophil infiltration (Fig. 4h) and high ratio of neutrophils to T cells (Fig. 4i).
a, Correlation coefficients of TBB determined from WGS with immune cell populations in HNSCC tumors. Statistical analysis was conducted using a two-tailed Spearman correlation. Highlighted values indicate an FDR < 0.05 (n = 157). TFH cells, T follicular helper cells; TCM cells, central memory T cells. b, Correlation coefficients of TBB determined from WES in independent samples not analyzed by WGS (excluding samples from a) with immune cell populations in HNSCC tumors. Statistical analysis was conducted using a two-tailed Spearman correlation. Highlighted values indicate an FDR < 0.05 (n = 348). c, Correlation coefficients of TBB with immune cell populations in tumors from CIAO trial. Statistical analysis was conducted using a two-tailed Spearman correlation. Highlighted values indicate P < 0.05 (n = 28). d, Correlation of TBB with log-transformed tNLR in samples from CIAO trial. Statistical analysis was conducted using a two-tailed Spearman correlation (n = 28). e, Comparison of log-transformed tNLR between responders (PR) and nonresponders (SD and PD) from CIAO trial. Data are shown as the mean and 95% confidence interval. Statistical analysis was conducted using a two-tailed Welch’s t-test (nonresponder, n = 16; responder, n = 12). f, Spearman correlation coefficients of TBB with tNLR across cancer types for HNSCC by WGS (n = 157) and WES (n = 503), STAD by WGS (n = 114) and WES (n = 413), ESCA by WGS (n = 62) and WES (n = 183) and COADREAD by WGS (n = 155) and WES (n = 582). Statistical analysis was conducted using a two-tailed Spearman correlation. g–i, Correlation of imaging-based quantification of bacteria by 16S rRNA in situ hybridization with T cells detected by CD3 (g), neutrophils detected by CD66b (h) and difference between neutrophils and T cells (i). Inset values indicate the two-tailed Spearman correlation coefficient (n = 17). j, Changes induced by in vitro exposure of HNSCC cell lines to either Fusobacterium or Prevotella, with each dot representing an individual gene. Four separate cell lines were exposed to bacteria and the average fold change is shown. The inset value indicates the two-tailed Pearson correlation coefficient. Values represent the average of n = 4 cell lines, each analyzed in duplicate. k, Comparison correlation coefficients between individual genes and TBB in participant tumors (n = 503) with average change induced by in vitro exposure of HNSCC cell lines to Fusobacterium and Prevotella (n = 4 cell lines). In a–c, immune populations were determined by ssGSEA using signatures from Bindea et al.63,64.
To better understand the mechanisms underlying immunological remodeling associated with high abundance of intratumoral bacteria, we infected a panel of four HNSCC cell lines with two highly prevalent bacteria within the HNSCC tumor microbiome, F. nucleatum and Prevotella scopos. We found that bacteria-induced gene expression changes were conserved between the two different bacteria (Fig. 4j and Extended Data Fig. 6a–d). Leading-edge analysis of GSEA from Fig. 3j indicated multiple neutrophil and myeloid chemoattractants as primary drivers of the observed inflammatory signaling, including CSF2/3, CXCL1/2/3, CXCL8, IL1A/B and TNF. Heat-killing Fusobacterium before infecting HNSCC cells greatly blunted the induction of these genes compared to live bacteria (average of 85% reduction; Extended Data Fig. 6e–j), suggesting that signaling is not purely related to the pathogen-associated molecular pattern recognition of lipopolysaccharide or other biomolecules. These neutrophil and myeloid chemoattractants were also among the most highly induced genes following in vitro bacterial exposure and were generally moderately32 positively correlated with TBB in participant tumors (Fig. 4k). These data suggest that bacteria-induced chemoattractants are responsible for the accumulation of neutrophils in high-TBB tumors.
In vivo modulation of intratumoral bacteria burden alters immunosuppression and immunotherapy sensitivity
To mechanistically study the role of intratumoral bacteria burden in vivo, we leveraged murine models of head and neck cancer implanted either orthotopically into the tongue or subcutaneously in the flank (Fig. 5a) under specific-pathogen-free housing conditions. In these conditions, intratumoral bacteria accumulated in orthotopic tongue tumors but were absent in the subcutaneous tumors (Fig. 5b,c). Broad-spectrum antibiotics administered through drinking water, beginning 1 week before tumor implantation, effectively depleted bacterial load in orthotopic tumors (Fig. 5b,c). Antibiotic treatment of mice implanted with MOC2 tumors significantly attenuated tumor growth in the bacteria-rich orthotopic site (Fig. 5d) but had no effect at the low-intratumoral-bacteria subcutaneous site (Fig. 5e). Similar results were observed with MOC1 tumors (Fig. 5f,g). Notably, we did not detect Fusobacterium in murine orthotopic tumors but instead found ~95% Streptococcus, which was associated with low-bacteria-burden tumors when analyzing relative abundance (Fig. 3b and Extended Data Fig. 7), suggesting that the observed phenotypes are not driven by the presence of a specific opportunistic pathogen.
a, Experimental schematic. Syngeneic oral cancer cell lines were implanted either subcutaneously or in the orthotopic (tongue) location. For antibiotic studies, antibiotic treatment (ABX) was initiated 1 week before tumor implantation. b, Representative immunofluorescence staining detecting bacteria in tumors for the treatment groups (green, bacterial 16S rRNA); cell nuclei were stained with DAPI (blue). Scale bar, 100 μm. c, Quantification of relative bacteria in the ABX versus control treatment groups in the flank versus orthotopic location (n = 5). Data are shown as the mean and s.d. d–g, Comparison of tumor volume growth curves with and without ABX bacterial depletion for control (n = 10) and ABX (n = 8) MOC2 orthotopic tumors (d), control (n = 10) and ABX (n = 10) MOC2 subcutaneous tumors (e), control (n = 9) and ABX (n = 10) MOC1 orthotopic tumors (f) and control (n = 6) and ABX (n = 6) MOC1 subcutaneous tumors (g). All tumor volumes are presented as the mean and s.e.m. Statistical analysis was conducted using an unpaired two-tailed t-test. NS, not significant. h, Immune deconvolution of mRNA expression profiling using ssGSEA with signatures from Bindea et al.63,64 from orthotopic tumors in mice with or without ABX demonstrating increased neutrophils and decreased CD8 T cells (n = 5). Statistical analysis was conducted using a two-tailed t-test. The box represents the median and interquartile range and whiskers represent the range. i, Comparison of changes in intratumoral immune cell populations quantified by immune deconvolution using ssGSEA with signatures from Bindea et al.63,64 induced by ABX in mice bearing either orthotopic tumors (x axis) or subcutaneous tumors (y axis). Blue-highlighted populations indicate populations significantly changed in orthotopic tumors. Inset values indicate the two-tailed Spearman correlation coefficient (n = 5). j, Changes in cytokine expression induced by ABX in orthotopic tumors or subcutaneous tumors, as well as the interaction effect between ABX and tumor site. The dot color indicates the direction and magnitude of change; nominally significant relationships (P < 0.05) are encircled in black. Significance was determined using a two-way ANOVA (interaction effect, ABX × site), followed by Šidák post hoc analysis to compare antibiotic-induced changes in the orthotopic and subcutaneous sites (n = 5). k,l, Comparison of tumor volumes after depletion of CD4-positive and CD8-positive cells (k; n = 6) or Ly6G-positive cells (l; n = 9) in orthotopic MOC1 tumors. All tumor volumes are presented as the mean and s.e.m. Statistical analysis was conducted using an unpaired two-tailed Welch’s t-test.
Antibiotic treatment of orthotopic tumors resulted in a decrease in neutrophils and increase in CD8 T cells, implicating intratumoral bacteria in remodeling the tumor immune microenvironment (Fig. 5h). While previous studies demonstrated that antibiotic-mediated disruption of the gut microbiome can suppress antitumor immunity and reduce ICB efficacy in melanoma and other low-bioburden tumors33,34, its impact in HNSCC is more uncertain. We found that antibiotics failed to promote immunogenicity in subcutaneous tumors that harbor minimal intratumoral bacteria, exhibiting near-opposite changes to those observed in the orthotopic site (Fig. 5i). Likewise, analysis of cytokine gene expression following treatment with antibiotics revealed opposing changes between orthotopic and subcutaneous tumors, with numerous chemokines exhibiting a significant interaction effect between tumor site and antibiotic treatment (Fig. 5j). These data suggest that, in the high-bacterial-burden oral cavity, antibiotic-mediated depletion of intratumoral bacteria may counteract the potentially deleterious effects of gut microbiome disruption. On the other hand, in low-bacteria-burden tumors such as those at nonmucosal sites outside of the digestive tract, the negative impact on the gut microbiome may predominate. Consistent with the hypothesis that antibiotics may be beneficial in bacterially rich HNSCC, analysis of clinical benefit from anti-PD(L)1 treatment in participants with HNSCC revealed that those treated with concurrent antibiotics exhibited significantly improved responses to immunotherapy35 (Extended Data Fig. 8). Together, these findings suggest that eliminating intratumoral bacteria can enhance antitumor immunity.
We next evaluated how key immune cells contribute to the suppression of tumor growth by antibiotics. We hypothesized that the influx of T cells following antibiotic treatment may drive the observed suppression of tumor growth. We evaluated this hypothesis through systemic depletion of CD4+ and CD8+ T cells (Extended Data Fig. 9a–c) in mice bearing orthotopic MOC1 tumors. In contrast to our hypothesis, depletion of CD4+ and CD8+ cells failed to mitigate the phenotype, resulting in an even more pronounced growth phenotype (Fig. 5k), indicating CD4+ and CD8+ T cells were not the primary driver of reduced tumor growth with antibiotic treatment. To evaluate whether neutrophils may be the primary driver of the growth phenotype, we repeated the experiment with the depletion of Ly6G+ cells (Extended Data Fig. 9d,e). Depletion of Ly6G+ cells completely abrogated the effect of antibiotics on tumor growth (Fig. 5l), indicating that intratumoral bacteria can enhance tumor growth through the recruitment of tumor-promoting neutrophils. Proneutrophilic chemokines were induced by the infection of MOC1 cells with either Fusobacterium or Prevotella (Extended Data Fig. 6k), strongly mirroring the infection of human cell lines (Extended Data Fig. 6l). In vivo, antibiotics also suppressed these neutrophil chemokines in murine tumors (Extended Data Fig. 6m). These results demonstrate that the reduction in intratumoral bacteria abundance by antibiotic treatment promotes antitumor immunity by altering the intratumoral immune landscape.
To experimentally validate the role of intratumoral bacteria in ICB response, we sought to increase total intratumoral bacteria abundance in the ICB-sensitive MOC1 tumor model. To do so, we randomized mice to either control arms or oral administration of tumor-associated bacteria Fusobacterium, Prevotella or Campylobacter for 5 days and then implanted all study arms with MOC1 tumor cells (Fig. 6a). We found that all three bacteria increased total intratumoral bacteria (Fig. 6b,c). Increased bacteria within orthotopic MOC1 tumors led to increased neutrophil infiltration, reduced CD3+ T cells and a higher ratio of neutrophils to T cells (Fig. 6b,d–f). We hypothesized that the observed decrease in T cells may result in reduced efficacy of ICB and evaluated this hypothesis using anti-PDL1 treatment. Control orthotopic MOC1 tumors robustly responded to anti-PDL1 (Fig. 6g). Increasing intratumoral bacteria through the oral administration of Fusobacterium, Prevotella or Campylobacter was sufficient to abrogate the response of orthotopic MOC1 tumors to anti-PDL1 treatment (Fig. 6h–j). To validate that resistance to anti-PDL1 treatment following administration of bacteria was because of intratumoral accumulation and not influenced by leakage of bacteria into the gut as reported in colorectal cancer36, we repeated the anti-PDL1 treatment experiment in mice bearing subcutaneous MOC1 tumors. We found that oral administration of Fusobacterium had no impact on the sensitivity of subcutaneous MOC1 tumors to anti-PDL1 treatment, indicating that the effect of Fusobacterium on ICB therapy depends on the intratumoral delivery of bacteria and not simply bacterial administration into the alimentary tract (Fig. 6k,l). By using multiple different bacterial taxa, including both those that increase in relative abundance with increasing TBB and those that do not, our findings strongly support the hypothesis that intratumoral bacteria suppress antitumor immunity regardless of bacterial taxa.
a, Schematic of experimental bacterial inoculation and immunotherapy treatment. b, Representative images of control tumors or tumors inoculated with indicated microbes stained for CD3 (T cells, cyan), Ly6G (neutrophils, red), 16S rRNA (bacteria, green) and nuclear counterstain (DAPI, white). Scale bar, 100 μm. c–f, Quantifications of images in b for 16S rRNA staining (c), neutrophils (d), T cells (e) and log2 ratio of neutrophil to T cell density (f) (control, n = 8; otherwise, n = 7). Statistical analysis was conducted using a one-way ANOVA with Benjamini–Hochberg correction for multiple comparisons. The box center indicates the median, box edges indicate the interquartile range and whiskers indicate the minimum and maximum. g–j, Tumor volumes in orthotopic MOC1 tumors with and without anti-PDL1 treatment or IgG control for control mice (g) and mice orally inoculated with F. nucleatum (h), P. scopos (i) or C. rectus (j) (n = 8). Tumor volumes are presented as the mean and s.e.m. Statistical analysis was conducted using an unpaired two-tailed Welch’s t-test. k,l, Tumor volumes in subcutaneous MOC1 tumors with anti-PDL1 treatment or IgG control for control mice (k) and mice orally inoculated with F. nucleatum (l) (n = 8). Tumor volumes are presented as the mean and s.e.m. Statistical analysis was conducted using a unpaired two-tailed Welch’s t-test. Panel a created with BioRender.com.
In summary, our results indicate that high intratumoral bacteria burden promotes an immunosuppressive tumor microenvironment, exacerbating tumor growth and driving resistance to ICB.
Discussion
The immunosuppressive tumor microenvironment in HNSCC limits response to both traditional and immune-based therapies37. Despite the promise of ICBs, the majority of persons with HNSCC do not demonstrate clinical benefit1,38,39. Although some biomarkers may have predictive value for ICB response (for example, PDL1 staining) for persons with HNSCC3,4, improved biomarkers are needed because of the complexity of the tumor microenvironment. Here, we find that only TBB was predictive of benefit from ICB in the CIAO clinical trial and further validate TBB as a biomarker for ICB in independent cohorts. This observation is further supported in our companion paper analyzing the placebo-controlled phase 3 JAVELIN100 trial40. Moreover, experimental modulation of intratumoral bacteria burden is sufficient to remodel the immune microenvironment and modulate sensitivity to ICB. Together, our current data suggest that the intratumoral bacterial microbiome may be critical in shaping the tumor microenvironment and modulating response to immunotherapy.
Many important studies have demonstrated that the composition of the gut microbiome has profound implications regarding T cell-mediated ICB responses in persons with cancer, presenting prognostic and therapeutic opportunities6,7,8. Fewer studies have focused on the role of intratumoral bacterial landscape in shaping tumor immunogenicity. In lung cancer, airway dysbiosis driven by increased relative abundance of Veillonella parvula was associated with interleukin-17-dependent tumor progression14. Multiple studies have demonstrated that a high relative abundance of opportunistic pathogens such as F. nucleatum can have a negative influence on the tumor immune microenvironment and clinical outcomes in several cancer types, including HNSCC12,13,20,25,41,42,43,44. We found that the relative fraction of the genus Fusobacterium is strongly correlated with total tumor bacteria abundance within HNSCC tumors, suggesting that prior studies analyzing tumors with high relative Fusobacterium may also be analyzing high-TBB tumors.
In both preclinical models and participant tumor samples, we demonstrated that total abundance of bacteria is a primary driver of tumor microenvironment remodeling by promoting an immunosuppressive microenvironment, characterized by an accumulation of neutrophils and reduction in T cells. Critically, the ability to remodel the immune microenvironment through experimental modulation of intratumoral bacteria burden supports the hypothesis that bacteria are driving the immune changes, as opposed to a specific immune landscape permitting the persistence of bacteria. It is important to note that the murine orthotopic oral tumors are free of Fusobacterium and other known opportunistic pathogenic bacteria, instead being primarily colonized with commensal Streptococcus (~95%). Streptococcus is typically associated with noncancerous tissues, where it exhibits a higher relative abundance compared to HNSCC tumors12. Beyond specific taxa, prior work in the gut also suggested that microbiome diversity can impact immune activity7. However, the homogeneity of the untreated murine orthotopic tumor microbiome makes antibiotic-driven alterations in intratumoral microbial diversity an unlikely driver of TME changes, supported by a lack of association between diversity and treatment response in CIAO samples.
Expression of neutrophil chemoattractants associated with high intratumoral bacteria abundance in both preclinical mouse models and participant tumors was induced by multiple bacteria in HNSCC cell lines in culture, independent of bacteria type. These findings suggest that immune modulation may be primarily driven by the overall intratumoral bacteria abundance and not by the presence of a specific pathogenic bacterium. This is also mirrored by prior work showing that bacteria-rich regions of tumors are enriched in neutrophils11. Acute stimulation of neutrophil migration and activation may be beneficial in preventing infection or even tumor spread and can also have an essential role in maintaining oral health and preventing oral bacterial dysbiosis45. Likewise, recruitment of neutrophils after initiation of ICB was shown to be critical for tumor clearance46,47. However, dysregulation of neutrophils in the tumor microenvironment can lead to chronic immunosuppression. Our results may provide a mechanistic rationale for the accumulation of neutrophils and polymorphonuclear myeloid-derived suppressor cells (PMN-MDSCs) within certain tumors.
We observed divergent effects of bacterial depletion by antibiotics in the high-bacterial-load orthotopic site and low-bacterial-load subcutaneous site, likely because of signals from intratumoral bacteria driving effects in the orthotopic site versus signals from gut bacteria driving effects in the subcutaneous site. Antibiotics were shown to promote T cell accumulation and suppress tumor growth in a T cell-dependent manner in preclinical pancreatic cancer models; however, the relative impact from depletion of gut versus intratumoral microbiome was not assessed48. In contrast to these findings, we found that depletion of CD4+ and CD8+ T cells in mice bearing orthotopic tumors enhanced the antitumor effect of bacterial ablation. However, depletion of Ly6G+ cells abrogated the effect of bacterial ablation, indicating that this phenomenon is not dependent on adaptive T cell immunity but driven by Ly6G+ neutrophils and PMN-MDSCs. These results are mirrored by studies in preclinical lung cancer models, where a reduction in tumor bacteria with antibiotics reduced the proliferation of γδ T cells to suppress neutrophil recruitment and tumor growth49.
There is conflicting evidence regarding the use of antibiotics in the setting of solid tumors and ICB treatment. Multiple correlative studies focused on tumor types with low levels of intratumoral bacteria indicated that there is a reduced response to ICB when persons receive antibiotics50,51,52, whereas, in persons with HNSCC, we observed improved clinical benefit in those receiving antibiotics. There are only three published studies exploring the use of antibiotics in HNSCC and they showed conflicting results with limited generalizability. Specifically, the first study relied on overall survival and did not take relevant confounding variables or comorbidities into account53. In the second study, the results were skewed by the treatment line, where persons who received antibiotics were more likely receive ICB in the second or third line, whereas those who did not receive antibiotics were more likely to receive ICB as first-line treatment where response rates are higher54. In the third study, no difference in objective response was found and differences in progression-free and overall survival were only significant in persons over 70 years of age55. The HNSCC cohort in our analysis, where antibiotics were associated with improved ICB, near-universally received checkpoint blockade in the second or third line. We posit that the effect of antibiotics on persons with cancer receiving immunotherapy is highly dependent on the overall biomass of the cancer under investigation and the type and timing of antibiotics administered.
Intratumoral bacterial load may serve as a biomarker for ICB response in persons with HNSCC, as it was the only significant predictor of immunotherapy response in the CIAO clinical trial. For clinical translation, technical variation across TBB quantification assays must be explored, as variation across assays is known to impact the predictive reliability of clinically approved ICI biomarkers microsatellite instability status55, PDL1 staining56 and TMB57. The predictive accuracy of intratumoral bacteria abundance, quantified as TBB, was validated in additional retrospective cohorts of ICB-treated participants and was further verified in our companion paper analyzing the JAVELIN100 multicenter randomized phase 3 placebo-controlled clinical trial40. TBB could potentially be used to stratify persons more likely to benefit from neoadjuvant ICB treatment, a particularly timely finding given the recent positive findings reported from a completed neoadjuvant immunotherapy clinical trial58. Alternatively, abundance of intratumoral bacteria may be used to identify persons who may benefit from antibiotic use in combination with immunotherapy. Restricting antibiotic administration to persons with high-bacterial-load tumors may optimize the therapeutic balance by preserving the immunostimulatory benefits of intratumoral bacterial depletion while minimizing the potential adverse effects of antibiotics on the gut–immune axis. This approach could be further refined through the use of targeted antibiotics or alternative bacteria-directed strategies that selectively eliminate intratumoral bacteria while preserving commensal gut microbiota. Ongoing clinical trials (NCT06627270) are assessing the potential of antibiotics to reduce intratumoral microbial load and can inform the design of future studies.
In conclusion, our study demonstrates that the accumulation of intratumoral bacteria can drive immunosuppression and induce resistance to ICB. These findings have broad implications for basic tumor biology and the clinical treatment of cancer.
Methods
Ethical approval
Our research complied with all relevant ethical guidelines. Research was overseen by the Institutional Animal Care and Use Committee of Cleveland Clinic Foundation (protocol 2774) and Institutional Review Boards of the MD Anderson Cancer Center (CIAO trial) and Cleveland Clinic Foundation (all other participant samples).
Mammalian cell lines and culture
All cells were cultured in a humidified incubator at 37 °C and 5% CO2 and verified by Mycoplasma and short tandem repeat testing by the Cleveland Clinic Cell Services’ Cell and Media Production Core (Media Core). MOC1 and MOC2 cells (gifted by R. Uppaluri, Dana Farber Cancer Institute) were cultured in IMDM (Cytiva) and Ham’s Nutrient Mixture F12 (Cytiva) at a 2:1 mixture with 5% FBS (Cytiva), 5 ng ml−1 mEGF (Life Technologies), 400 ng ml−1 hydrocortisone (Sigma-Aldrich) and 5 mg ml−1 insulin (Sigma-Aldrich). FaDu (American Type Culture Collection (ATCC), HTB-43) cells were cultured in EMEM (Media Core) supplemented with 10% FBS (Gibco). OQ01 (provided by L.-J. Chang, University of Florida) and SCC-25 (ATCC, CRL-1628) cells were cultured in DMEM/F12 (Media Core) supplemented with 10% FBS (Gibco). PCI-15B (provided by R. Ferris, University of North Carolina) cells were cultured in DMEM (Media Core) supplemented with 10% FBS (Gibco).
Bacterial strains and culture
F. nucleatum subsp. nucleatum (ATCC, 23726) and P. scopos (German Collection of Microorganisms and Cell Cultures, 22613) were grown under anaerobic conditions in TSABYEP consisting of tryptic soy broth (Millipore) supplemented with 5 g L−1 yeast extract (BioBasic) and 10 g L−1 Bacto peptone (BioWorld). Both strains were streaked on TSABYEP agar plates prepared using the same medium with the addition of agar (Himedia) at 15 g L−1. Campylobacter rectus (ATCC, 33238) was grown with TSABYEP + SF2 broth (TSABYEP + 20 mM sodium formate and 20 mM sodium fumarate) and was streaked on TSABYEP + SF2 agar plates using the same medium with the addition of agar at 15 g L−1. The bacterial strain identified was confirmed by Sanger sequencing.
Tumor models and treatments
For all experiments, 8–12-week-old C57BL/6J female mice were obtained from Jackson Laboratory (stock 000664). Mice from Jackson Laboratories have been shown to be very homogeneous and stable in terms of gut microbiota across not only breeding pairs but also location and time59. Mice were housed under specific-pathogen-free conditions at the Cleveland Clinic Biological Research Unit in ventilated cages kept at constant temperature (20–26 °C), humidity (30–70%) and a 14-h light–dark cycle. Before initiating experiments, mice were allowed to equilibrate for 3–4 weeks. Mice were fed Envigo Diet 2918 (Envigo) ad libitum and DietGel 76A-72-07-5022 (ClearH2O) was supplemented to the diet 6–8 days after tumor implantation.
For flank models, 2 × 106 MOC1 or 1 × 105 MOC2 cells in sterile PBS were injected subcutaneously in the left flank. Injection site was shaved and sterilized with antimicrobial wipes before injection. For orthotopic implantation, 3 × 105 MOC1 or 3 × 104 MOC2 cells in sterile PBS were injected into the tongue. Mice were monitored for tumor growth and body condition weekly. Tumor size was recorded as the greatest longitudinal diameter (length) and the greatest transverse diameter (width) measured with an external caliper. Tumor volumes were calculated using the following formula: tumor volume = length × width2 × 0.5. Mice were excluded from analysis in the case of deaths that could not be associated with tumor growth (for example, no loss of bodyweight or other signs of poor condition and tumor burden far below maximum allowable volume). The maximum allowable tumor volume for subcutaneous flank models was 2 cm3. For the orthotopic tongue model, the tumor location prevented a prespecified size humane endpoint; instead, the mouse condition was considered, with endpoints of 15% weight loss, signs of respiratory distress, dehydration, overly hunched posture, tumor causing tongue protrusion or an ensemble body condition score.
Antibiotic treatment
Mice were treated with antibiotics in their drinking water 4 days before implantation until endpoint with the following cocktail: 1 g L−1 ampicillin sodium salt (Sigma-Aldrich, A9518), 1 g L−1 neomycin sulfate (GoldBio, N-620-25), 1 g L−1 metronidazole (GoldBio, M-295-100) and 0.5 g L−1 vancomycin hydrochloride (Sigma-Aldrich, V8138). Antibiotics were prepared fresh weekly. Depletion of oral microbiota was verified using 16S rRNA FISH in the tumor (Fig. 6b,c), aerobic and anaerobic cultures from oral swabs (Extended Data Fig. 10a) and 16S rRNA qPCR of oral swabs (Extended Data Fig. 10b).
Bacterial inoculations
Mice were orally inoculated five times per week with ~50 μl of F. nucleatum, P. scopus or C. rectus at 1 × 109 colony-forming units of bacteria per ml in 2% (w/v) low-viscosity carboxymethylcellulose (Sigma-Aldrich) dissolved in sterile PBS. For inoculation, scruffed mice were held ventral side up. The oral cavity was filled with inoculum and the mouse was held for 10 s before release back into the cage.
Immune cell depletion
For depletion antibody treatments, 500 μg of anti-CD4 (GK1.5, BioXcell) and anti-CD8α (2.43, BioXcell) antibodies were administered through two intraperitoneal injections on consecutive days 1 week before tumor implantation, followed by readministration every 14 days60. Anti-Ly6G (1A8, BioXcell) was administered through intraperitoneal injections of 350 μg twice weekly starting 1 week before tumor implantation until the experimental endpoint61.
ICB treatment
After the development of palpable tumors, mice were randomized to receive either anti-PDL1 treatment (clone 10F.9G2, BioXcell) or IgG2b isotype control (LTF-2, BioXcell). Antibodies were administered through an initial intraperitoneal injection of 400 μg, followed by twice-weekly intraperitoneal injections of 200 μg until the endpoint.
Flow cytometry
Fresh whole blood was obtained through retro-orbital bleeding of anesthetized mice. Blood was collected in tubes containing 1:6 v/v ACD anticoagulant. Samples were stained for 20 min at room temperature with fluorescently labeled antibodies to CD45, CD4 and CD8 or CD45, Ly6G and CD11b (Supplementary Table 5). Red blood cells (RBCs) were lysed with RBC lysis buffer (BioLegend) after staining. Samples were washed twice, resuspended in the cell staining buffer (Biolegend) and acquired using a Sony ID7000 spectral cell analyzer (Sony Biotechnology). Data were analyzed using FlowJo (version 10.8.0; BD Biosciences). Gating strategies used for the identification of different cell types are shown in Extended Data Fig. 10.
Bacterial coculture and RNA sequencing analysis
FaDu, OQ01, PCI-15B, SCC-25 and MOC1 cells were seeded 1 day before infection. Before infection, the medium was replaced with fresh prereduced medium on all cells. For bacterial infection, bacteria (F. nucleatum or P. scopos) were resuspended in prereduced mammalian cell culture medium and added at a multiplicity of infection (MOI) of 200 to desired wells. Cells were returned to a standard mammalian cell culture conditions for 4 h, consistent with similar prior studies11. RNA was isolated with a Qiagen RNeasy mini kit (Qiagen) using on-column DNase treatment (Qiagen). RNA sequencing was performed by the Cleveland Clinic Genomics Core using a stranded mRNA prep and ligation kit (Illumina) for library preparation followed by sequencing using the NovaSeq next-generation sequencing platform. Resulting reads were quantified as transcripts per million (TPM) using kallisto (version 0.44.0)62 aligned to either GRCh38 (FaDu, OQ01, PCI-15B and SCC-25) or GRCm39 (MOC1). The change following infection was taken as the average log2-transformed difference for bacterial-infected cells compared to mock-treated control cells for each cell line and then reported as the average fold change across all four human cell lines (Fig. 4j) or as individual cell lines (Extended Data Fig. 6a–d,k). To compare overall changes induced by bacterial infection (Fig. 4k and Extended Data Fig. 6l,m), gene expression changes induced by F. nucleatum and P. scopos were averaged.
For qPCR studies with heat-killed bacteria, cells were infected at an MOI of 200 with live F. nucleatum or dead heat-killed F. nucleatum or mock-infected as a control. RNA was isolated as above and then complementary DNA was synthesized using iScript reverse transcription supermix (Bio-Rad). qPCR was performed using a QuantStudio3 (Thermo Fisher Scientific) and SsoAdvanced Universal SYBR green supermix (Bio-Rad) with primers for desired gene targets and β-actin internal control (Supplementary Table 3). The relative levels of RNA expression in treated samples in comparison to mock-treated controls were quantified using the 2−∆∆Ct method.
Murine in vivo tumor RNA expression profiling
RNA from tumors of control or antibiotic-treated mice RNA was isolated using the Qiagen RNeasy mini kit (Qiagen) with on-column DNase treatment (Qiagen). The nCounter PanCancer immune profiling panel (XT-CSO-MIP1-12) was used to analyze gene expression through the NanoString Technology platform. Data were normalized by the implication of positive and negative control probes and housekeeping genes. Immune cell deconvolution was performed using ssGSEA with signatures from Bindea et al.63,64. Analysis of the interaction effect between tumor implantation site and antibiotic treatment was assessed by a two-way analysis of variance (ANOVA). Raw data are available in Supplementary Table 4.
Mouse tumor 16S rRNA sequencing and analysis
DNA from frozen tumors was isolated using the ZymoBIOMICS DNA miniprep kit (Zymo Research, D4300) as per the manufacturer’s protocols and used for 16S rRNA sequencing of the V4 region. Single-end sequences were imported into QIIME 2 (version 2018.8) using the Casava 1.8 single-end demultiplexed fastq format. The divisive amplicon denoising algorithm 2 (DADA2) pipeline65 was used to filter and trim sequences (tuncLen = 150, maxEE = 2, tuncQ = 2). Taxonomy was assigned to reads within the pipeline using the silva database (version 138.1). The output of the DADA2 pipeline (feature table of amplicon sequence variants) was processed for alpha and beta diversity analysis using base R. Taxa were filtered using PCR negatives and those found to be at higher relative quantities in the PCR negatives compared to samples were removed before analysis; these included Catenibacterium, Cellulomonas, Phascolarctobacterium and Ralstonia. Additional environmental controls were included to improve filtering for potential contaminants (Burkholderiaceae, Caballeronia, Paraburkholderia and Bradyrhizobiaceae). Microbial taxa abundance was estimated by the summed count of all reads in each sample, binned to each unique taxa level.
PathSeq analysis and validation approaches
Because of recent concerns about analysis of microbial content from bulk sequencing data, we took several steps to ensure analytical robustness. Preliminary analysis found that PathSeq-based approaches outperformed those used in the retracted work by Poore et al.27 (Extended Data Fig. 1a,b); thus, we selected PathSeq for further use.
Identification of contaminants from 16S rRNA sham sequencing controls
First, we 16S rRNA sequenced HNSCC formalin-fixed paraffin-embedded (FFPE) tumors and sham controls with Zymo Research using V4 sequencing primers and quantified using DADA2 by Zymo Research. Comparing sham controls to tumors, we identified numerous known contaminant taxa (Extended Data Fig. 1c). Taxa from this experiment, in conjunction with prior literature on known contaminants28,29 and manual curation of known oral microbes66, were subtracted from subsequent analyses (Supplementary Table 1).
Comparison of 16S rRNA sequencing with PathSeq
Using taxa identified in this analysis, along with manual curation of prior literature, we generated a custom index of relevant bacterial taxa to prevent erroneous mapping to nonsensical bacteria such as deep-sea hydrothermal vent extremophiles, as previously reported26. All microbial genomes used are given in Supplementary Table 2. For the human reference genome, we used the full telomere to telomere genome T2T-CHM1 (ref. 67). We next performed paired 16S rRNA sequencing (Zymo Research) and WES on a set of HNSCC tumors performed by the Broad Institute using their Research Human Exome pipeline. WES was analyzed with PathSeq and, after subtracting identified contaminants, we compared relative bacterial abundances at the phylum and genus levels, finding a high degree of concordance (Extended Data Fig. 1d–g). To quantify total intratumoral bacteria abundance, we defined TBB as the number of bacterial reads per million human reads. All analysis was performed using GATK (version 4.5.0.0)68.
Additional validations
Comparing matched 16S rRNA qPCR, as described below, to TBB determined from WES also gave a high degree of concordance (Extended Data Fig. 1h). Additional comparison of microbial content profiled by both WES and WGS, as described below, also showed a high degree of concordance (Extended Data Fig. 2a–c).
CIAO cohort
Cohort overview
The CIAO cohort (NCT03144778)22 consisted of participants with resectable oropharyngeal squamous cell carcinoma that was either newly diagnosed stage II–IVA per AJCC7 (ref. 69) or locoregionally recurrent. Participants were randomized to two cycles of neoadjuvant durvalumab (anti-PDL1) or durvalumab + tremelimumab (anti-CTLA4) before surgery. Responders were considered participants with a partial response (PR); nonresponders were considered participants with stable disease (SD) or progressive disease (PD). Analysis conducted here includes prespecified exploratory analysis of tumor-based biomarkers. HPV status was determined by p16 immunostaining and RNAscope as previously described70. PDL1 immunostaining was performed using clone 22C3, with the combined proportion score calculated as described71. All sexes were considered for trial enrollment but participants were enrolled on the basis of the first available who agreed to participate. Sex and gender were self-reported. All participants provided written informed consent before enrollment. No compensation was provided.
RNA sequencing analysis and immune deconvolution
RNA sequencing was performed using QuantSeq using Baylor College of Medicine core facilities. Resulting reads were quantified as TPM using kallisto (version 0.44.0)62 aligned to GRCh38. Immune cell deconvolution was performed using ssGSEA using signatures from Bindea et al.63,64.
Mutation calling
WES was performed by LC Biosciences using the SureSelect human all exon V6 kit (Agilent Technologies) on both tumor and normal germline controls. Reads were aligned to GRCh38 using BWA-MEM. Somatic single-nucleotide variants and insertion–deletions were called using MuTect2 as implemented in GATK4 (refs. 68,72). Alignment artifacts were flagged using GATK4 FilterAlignmentArtifacts and all flagged variants were manually inspected in Integrated Genomics Viewer73. FFPE and oxoG artifacts were removed by read pair orientation bias filters, as described previously74. TMB was calculated as the number of mutations per megabase of DNA sequenced. Only nonsilent coding mutations with a variant allele frequency > 0.05 were included for TMB calculation.
Microbiome analysis
Analysis of intratumoral bacteria was performed as described above. We additionally included viral genomes for HPV16, HPV18, HPV33 and HPV35 on the basis of initial HPV serotyping. Similar to TBB, we also defined HPV viral burden as the number of HPV viral reads per million human reads.
Analysis of Cleveland Clinic HNSCC ICB cohort
Participants from Cleveland Clinic with HNSCC treated with ICB were identified from a retrospective chart review. WES was performed as part of clinical sequencing with Caris. Progression-free survival was defined as the time from start of anti-PD(L)1 treatment until participants either progressed or transitioned to another therapy. All participants provided written informed consent before enrollment. WES data were processed as described above.
Analysis of Hartwig samples
For analysis of NSCLC, HNSCC, urothelial carcinoma, mesothelioma and small intestine or colorectal cancer samples from the Hartwig cohort, data were obtained directly from Battaglia et al.25. Responders were considered participants with complete response (CR) or PR; nonresponders were considered participants with SD or PD. For associations of TBB or relative Fusobacterium with response, biopsy site and cancer type (where applicable) were taken as fixed effects, whereas sequencing platform was taken as a random effect, consistent with original analyses25.
Analysis of TCGA samples
All data were obtained from TCGA pan-cancer atlas release, including self-reported race, ethnicity and sex. Additional quantification of microbial content was obtained as described previously9, using PathSeq75 with subsequent robust contaminant subtraction9. Both WES (n = 511) and WGS (n = 157) were used for analysis. As described above, TBB was determined as the number of bacterial reads per million human reads mapped derived from either WGS or WES data. Annotations of aerobic and anaerobic bacteria were taken from Battaglia et al.25. To prevent variations in read counts impacting bacterial diversity analysis76, bacterial read counts from WGS data were scaled to a uniform 103 total reads and samples with under 103 total reads (27 of 157 samples) were excluded from further analysis. After scaling, any bacterial taxon with fewer than one read was set to 0 before calculation of Shannon diversity. For comparison of specific taxon-relative abundances in WGS versus WES, we limited analysis to samples profiled by both WES and WGS with at least 50 WES bacterial reads, resulting in 67 total samples. All samples were used for all other analyses. For mutation analysis, we focused on mutations contained within OncoKB77 and those defined as drivers for HNSCC in DriverDB4 (ref. 78). To analyze pathways enriched with increasing TBB, we performed GSEA79 on the basis of the Spearman correlation coefficient between TBB and gene expression levels using Hallmark gene sets80. Immune cell deconvolution was performed using ssGSEA with signatures from Bindea et al.63,64. Hypoxia score was determined as described by Battaglia et al.25.
Immunostaining and in situ hybridization
FFPE slides from participants with oral cavity squamous cell carcinoma were warmed at 60 °C for 30 min on a slide warmer and then dewaxed and rehydrated following standard techniques (subsequent xylene and ethanol washes). Following rehydration, slides were rinsed briefly in double-distilled H2O slides, fixed in 10% neutral buffered formalin for 20 min and then transferred to PBS-T (0.05% Triton X-100). Antigen retrieval was performed using the TintoRetriever pressure cooker (BioSB, BSB-7087) in citrate buffer (pH 6.0) for 15 min at 115 °C. Tissues were then either blocked in 2% BSA in PBS-T for antibody staining or moved directly to in situ hybridization. For antibody staining, following primary and secondary antibody incubations (Supplementary Table 5), tyramide signal amplification was performed with CF550R dye (Biotium, 96077) in amplification buffer (0.1 M borate pH 8.5 with 0.003% hydrogen peroxide). Bacterial FISH was performed as described previously81 using the pan-bacterial 16S rRNA probe EUB338 (5′-GCTGCCTCCCGTAGGAGT-3′) with a 5′ conjugation of ATTO 647N (Millipore Sigma) at 2 µM. FISH was performed in a hybridization oven at 46 °C for 2 h, with 2 µM probe in hybridization buffer (20 mM Tris pH 7.5, 900 mM NaCl, 0.01% SDS and 20% formamide), followed by washing in prewarmed wash buffer (200 mM NaCl, 20 mM Tris pH 7.5 and 5 mM EDTA). Slides were then mounted with Vectashield antifade mounting medium with DAPI (Vector Laboratories, H-1200). Slides were imaged on a Nikon Eclipse Ti2-e fluorescence inverted microscope with 6–12 images per tissue captured using Nikon Elements by a blinded individual randomly selecting tumor regions for imaging, using the histological appearance of the DAPI stain to identify tumor regions. Total tissue area was identified using a median-based threshold across intensity for all channels. Segmentation of immune cells and bacteria was performed on the basis of intensity relative to autofluorescence background levels. Immune cells are reported as the average number of cells per tissue area. Bacteria signal is reported as the integrated intensity of segmented regions normalized to the tissue area. Values were averaged across all images captured. Unstained tissues were used to determine detection limit and sensitivity of segmentation. All analysis was performed and quantified in MATLAB (R2020a, Mathworks).
Bacterial quantification by qPCR
DNA was extracted from FFPE rolls with a QIAamp DNA FFPE kit (Qiagen) and eluted in EB buffer (Qiagen). DNA was diluted fivefold with PCR-grade DNA-free water (Molzym) and quantified with Quant-iT PicoGreen double-stranded DNA kit (Invitrogen). qPCR was run with PowerTrack SYBR green mastermix (Applied Biosystems) on a QuantStudio3 (Thermo Fisher Scientific) thermocycler with 16s rRNA and human SMARCA4 primers (Supplementary Table 3). Tissues with no amplification of human housekeeping gene SMARCA4 were excluded from further analysis. Values were reported as 1/Ct. Background was determined on the basis of mock-extracted paraffin and DNA isolated from cultured human cell lines.
Antibiotic use and anti-PD(L)1 clinical benefit
Clinical benefit from anti-PD(L)1 in the presence or absence of antibiotics was assessed in participants with HNSCC of the oral cavity, oropharynx, larynx, hypopharynx and nasopharynx using data from Valero et al.35. Participants with autoimmune diseases were excluded from analysis.
Statistics
Statistical comparisons were made using GraphPad Prism 10 (GraphPad Software), R (version 4.2.0) and MATLAB R2020a (Mathworks). No statistical method was used to predetermine sample size. Unless otherwise noted, no specific blinding or randomization was performed. A comparison of two groups was made using Welch’s t-test (normally distributed) or rank-sum test (not normally distributed). Normality was assessed using quantile–quantile plots. A comparison of more than two groups was made using one-way or two-way ANOVA (normally distributed) or Kruskal–Wallis test (not normally distributed) with the appropriate post hoc test to compare groups if necessary. A comparison between two continuous variables was made using the Pearson or Spearman correlation coefficient. Two-tailed tests were used for all analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Data for the CIAO cohort including clinical characteristics, microbiome and immune deconvolution are contained within Supplementary Table 6, with sequencing data deposited to the European Genome-Phenome Archive (EGA) under accession number EGAS50000001174. Data for the Cleveland Clinic HNSCC cohort including clinical characteristics and microbiome analysis are contained in Supplementary Table 7, with sequencing data deposited to EGA under accession EGAS50000001175. Raw sequencing data from CIAO and Cleveland Clinic are both available under controlled access because of privacy restrictions. Requests for access can be made through the EGA portal, after which the requestor will receive a data access agreement to be completed and signed in agreement with the Cleveland Clinic. Data for cell line RNA sequencing were deposited to the National Center for Biotechnology Institute Gene Expression Omnibus under accession number GSE307471 and in vivo murine tumor NanoString transcriptional profiling are contained in Supplementary Table 4. Data from TCGA samples were downloaded from the Genomic Data Commons or from Dohlman et al.9, available through the Duke Research Data Repository. Data from Hartwig samples are available as Supplementary Tables from Battaglia et al.25. Contaminants subtracted and genomes used for microbial analysis are included in Supplementary Tables 1 and 2. The remaining data are available within the article and its Supplementary Information or from the corresponding author on request. Source data are provided with this paper.
Code availability
Code used for analysis is described with corresponding references in Methods including GATK68 for PathSeq and mutational analysis, kallisto (version 0.44.0)62 for RNA sequencing analysis, DADA2 (ref. 65) for 16S rRNA sequencing analysis, Nikon Elements and MATLAB R2020a for image capture and postprocessing and FlowJo (version 10.8.0) for flow analysis. Additional statistical analysis was performed in GraphPad PRISM 10, R (version 4.2.0) and MATLAB R2020a.
References
Ferris, R. L. et al. Nivolumab for recurrent squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 375, 1856–1867 (2016).
Burtness, B. et al. Pembrolizumab alone or with chemotherapy versus cetuximab with chemotherapy for recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-048): a randomised, open-label, phase 3 study. Lancet 394, 1915–1928 (2019).
Ferris, R. L. & Licitra, L. PD-1 immunotherapy for recurrent or metastatic HNSCC. Lancet 394, 1882–1884 (2019).
Wildsmith, S. et al. Tumor mutational burden as a predictor of survival with durvalumab and/or tremelimumab treatment in recurrent or metastatic head and neck squamous cell carcinoma. Clin. Cancer Res. 29, 2066–2074 (2023).
Villemin, C. et al. The heightened importance of the microbiome in cancer immunotherapy. Trends Immunol. 44, 44–59 (2023).
Matson, V. et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science 359, 104–108 (2018).
Gopalakrishnan, V. et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 359, 97–103 (2018).
Routy, B. et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 359, 91–97 (2018).
Dohlman, A. B. et al. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe 29, 281–298 (2021).
Geller, L. T. et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 357, 1156–1160 (2017).
Galeano Niño, J. L. et al. Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature 611, 810–817 (2022).
Michikawa, C. et al. Fusobacterium is enriched in oral cancer and promotes induction of programmed death-ligand 1 (PD-L1). Neoplasia 31, 100813 (2022).
Riquelme, E. et al. Tumor microbiome diversity and composition influence pancreatic cancer outcomes. Cell 178, 795–806(2019).
Tsay, J. J. et al. Lower airway dysbiosis affects lung cancer progression. Cancer Discov. 11, 293–307 (2021).
Quigley, E. M. Gut bacteria in health and disease. Gastroenterol. Hepatol. (N. Y.) 9, 560–569 (2013).
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012).
Yamashita, Y. & Takeshita, T. The oral microbiome and human health. J. Oral Sci. 59, 201–206 (2017).
Wang, N. & Fang, J. Y. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 31, 159–172 (2023).
Brennan, C. A. & Garrett, W. S. Fusobacterium nucleatum—symbiont, opportunist and oncobacterium. Nat. Rev. Microbiol. 17, 156–166 (2019).
Bullman, S. et al. Analysis of Fusobacterium persistence and antibiotic response in colorectal cancer. Science 358, 1443–1448 (2017).
Yu, T. et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell 170, 548–563 (2017).
Ferrarotto, R. et al. Impact of neoadjuvant durvalumab with or without tremelimumab on CD8+ tumor lymphocyte density, safety, and efficacy in patients with oropharynx cancer: CIAO trial results. Clin. Cancer Res. 26, 3211–3219 (2020).
Shamseddine, A. A., Burman, B., Lee, N. Y., Zamarin, D. & Riaz, N. Tumor immunity and immunotherapy for HPV-related cancers. Cancer Discov. 11, 1896–1912 (2021).
Peng, S. et al. HLA-DQB1*02-restricted HPV-16 E7 peptide-specific CD4+ T-cell immune responses correlate with regression of HPV-16-associated high-grade squamous intraepithelial lesions. Clin. Cancer Res. 13, 2479–2487 (2007).
Battaglia, T. W. et al. A pan-cancer analysis of the microbiome in metastatic cancer. Cell 187, 2324–2335(2024).
Gihawi, A. et al. Major data analysis errors invalidate cancer microbiome findings. mBio 14, e01607–e01623 (2023).
Poore, G. D. et al. Retraction Note: Microbiome analyses of blood and tissues suggest cancer diagnostic approach. Nature 631, 694–694 (2024).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12, 87 (2014).
Dyrhovden, R. et al. Managing contamination and diverse bacterial loads in 16S rRNA deep sequencing of clinical samples: implications of the law of small numbers. mBio 12, e0059821 (2021).
Elkrief, A. et al. Intratumoral Escherichia is associated with improved survival to single-agent immune checkpoint inhibition in patients with advanced non-small-cell lung cancer. J. Clin. Oncol. 23, 01488 (2024).
Seaby, E. G., Pengelly, R. J. & Ennis, S. Exome sequencing explained: a practical guide to its clinical application. Brief. Funct. Genomics 15, 374–384 (2016).
Cohen, J. Statistical Power Analysis for the Behavioral Sciences 2nd edn (Routledge or Lawrence Erlbaum Associates, 1988).
Poizeau, F. et al. The association between antibiotic use and outcome among metastatic melanoma patients receiving immunotherapy. J. Natl Cancer Inst. 114, 686–694 (2022).
Mohiuddin, J. J. et al. Association of antibiotic exposure with survival and toxicity in patients with melanoma receiving immunotherapy. J. Natl Cancer Inst. 113, 162–170 (2021).
Valero, C. et al. Clinical–genomic determinants of immune checkpoint blockade response in head and neck squamous cell carcinoma. J. Clin. Invest. 133, e169823 (2023).
Jiang, S. S. et al. Fusobacterium nucleatum-derived succinic acid induces tumor resistance to immunotherapy in colorectal cancer. Cell Host Microbe 31, 781–797(2023).
Davis, R. J., Van Waes, C. & Allen, C. T. Overcoming barriers to effective immunotherapy: MDSCs, TAMs, and Tregs as mediators of the immunosuppressive microenvironment in head and neck cancer. Oral Oncol. 58, 59–70 (2016).
Ferris, R. L. Immunology and immunotherapy of head and neck cancer. J. Clin. Oncol. 33, 3293–3304 (2015).
Harrington, K. J. et al. Nivolumab versus standard, single-agent therapy of investigator’s choice in recurrent or metastatic squamous cell carcinoma of the head and neck (CheckMate 141): health-related quality-of-life results from a randomised, phase 3 trial. Lancet Oncol. 18, 1104–1115 (2017).
Chan, T. A. et al. Tumor ecosystem and microbiome features associated with efficacy from avelumab-based multimodal therapy in a phase III randomized trial. Nat. Cancer (in the press).
Mima, K. et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut 65, 1973–1980 (2016).
Mima, K. et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol. 1, 653–661 (2015).
Fujiwara, N. et al. Involvement of Fusobacterium species in oral cancer progression: a literature review including other types of cancer. Int. J. Mol. Sci. 21, 6207 (2020).
Serna, G. et al. Fusobacterium nucleatum persistence and risk of recurrence after preoperative treatment in locally advanced rectal cancer. Ann. Oncol. 31, 1366–1375 (2020).
Mantovani, A., Cassatella, M. A., Costantini, C. & Jaillon, S. Neutrophils in the activation and regulation of innate and adaptive immunity. Nat. Rev. Immunol. 11, 519–531 (2011).
Gungabeesoon, J. et al. A neutrophil response linked to tumor control in immunotherapy. Cell 186, 1448–1464(2023).
Hirschhorn, D. et al. T cell immunotherapies engage neutrophils to eliminate tumor antigen escape variants. Cell 186, 1432–1447(2023).
Pushalkar, S. et al. The pancreatic cancer microbiome promotes oncogenesis by induction of innate and adaptive immune suppression. Cancer Discov. 8, 403–416 (2018).
Jin, C. et al. Commensal microbiota promote lung cancer development via γδ T cells. Cell 176, 998–1013 (2019).
Pinato, D. J. et al. Association of prior antibiotic treatment with survival and response to immune checkpoint inhibitor therapy in patients with cancer. JAMA Oncol. 5, 1774–1778 (2019).
Huang, X. Z. et al. Antibiotic use and the efficacy of immune checkpoint inhibitors in cancer patients: a pooled analysis of 2740 cancer patients. Oncoimmunology 8, e1665973 (2019).
Derosa, L. et al. Negative association of antibiotics on clinical activity of immune checkpoint inhibitors in patients with advanced renal cell and non-small-cell lung cancer. Ann. Oncol. 29, 1437–1444 (2018).
Preissner, S., Heiland, M., Preissner, R., Wirth, M. & Wollenberg, B. Antibiotics significantly decrease the survival of head and neck carcinoma patients with immunotherapy: a real-world analysis of more than 3000 cases. Cancers (Basel) 15, 2342 (2023).
Ueta, R. et al. Antibiotics may interfere with nivolumab efficacy in patients with head and neck squamous cell carcinoma. Oncology 102, 252–259 (2024).
Minohara, K. et al. Impact of medications on the efficacy of immune checkpoint inhibitors in patients with recurrent or metastatic head and neck cancer. Int. J. Clin. Oncol. 30, 1572–1581 (2025).
Doroshow, D. B. et al. PD-L1 as a biomarker of response to immune-checkpoint inhibitors. Nat. Rev. Clin. Oncol. 18, 345–362 (2021).
McGrail, D. J. et al. High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann. Oncol. 32, 661–672 (2021).
Uppaluri, R. et al. Neoadjuvant and adjuvant pembrolizumab in locally advanced head and neck cancer. N. Engl. J. Med. 393, 37–50 (2025).
Guo, J. et al. Characteristics of gut microbiota in representative mice strains: implications for biological research. Animal Model. Exp. Med. 5, 337–349 (2022).
Laky, K. & Kruisbeek, A. M. In vivo depletion of T lymphocytes. Curr. Protoc. Immunol. 113, 4.1.1–4.1.9 (2016).
Davis, R. W. T. et al. Luminol chemiluminescence reports photodynamic therapy-generated neutrophil activity in vivo and serves as a biomarker of therapeutic efficacy. Photochem. Photobiol. 95, 430–438 (2019).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Şenbabaoğlu, Y. et al. Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol. 17, 231 (2016).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Dewhirst, F. E. et al. The human oral microbiome. J. Bacteriol. 192, 5002–5017 (2010).
Nurk, S. et al. The complete sequence of a human genome. Science 376, 44–53 (2022).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Lydiatt, W., O’Sullivan, B. & Patel, S. Major changes in head and neck staging for 2018. Am. Soc. Clin. Oncol. Educ. Book 38, 505–514 (2018).
Mirghani, H. et al. Diagnosis of HPV driven oropharyngeal cancers: comparing p16 based algorithms with the RNAscope HPV-test. Oral Oncol. 62, 101–108 (2016).
Seiwert, T. Y. et al. Safety and clinical activity of pembrolizumab for treatment of recurrent or metastatic squamous cell carcinoma of the head and neck (KEYNOTE-012): an open-label, multicentre, phase 1b trial. Lancet Oncol. 17, 956–965 (2016).
Benjamin, D. et al. Calling somatic SNVs and indels with Mutect2. Preprint at bioRxiv https://doi.org/10.1101/861054 (2019).
Robinson, J. T., Thorvaldsdóttir, H., Wenger, A. M., Zehir, A. & Mesirov, J. P. Variant review with the Integrative Genomics Viewer. Cancer Res. 77, e31–e34 (2017).
Shih, D. J. H. et al. Genomic characterization of human brain metastases identifies drivers of metastatic lung adenocarcinoma. Nat. Genet. 52, 371–377 (2020).
Walker, M. A. et al. GATK PathSeq: a customizable computational tool for the discovery and identification of microbial sequences in libraries from eukaryotic hosts. Bioinformatics 34, 4287–4289 (2018).
Schloss, P. D. Rarefaction is currently the best approach to control for uneven sequencing effort in amplicon sequence analyses. mSphere 9, e0035423 (2024).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017).
Liu, S. H. et al. DriverDBv3: a multi-omics database for cancer driver gene research. Nucleic Acids Res. 48, D863–D870 (2020).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Drewes, J. L. et al. High-resolution bacterial 16S rRNA gene profile meta-analysis and biofilm status reveal common colorectal cancer consortia. Biofilms Microbiomes 3, 34 (2017).
Acknowledgements
This work was supported by National Cancer Institute (NCI) grant R00CA240689 to D.J.M. and National Institute of Dental and Craniofacial Research (NIDCR) grant K08DE029503, American Cancer Society RSG-24-1322538-01-IBCD and Velosano8 to N.L.S. Additional support was provided by NIDCR grant R00DE031372 to A. Stacy, Velosano8 to M.D., National Institutes of Health NCI 5U24 CA264006 to R.A. and Cancer Prevention and Research Institute of Texas fellow grant RP210037 to S.V.K. We are also grateful to L.-J. Chang (University of Florida) for OQ01 cells, R. Uppaluri (Dana Farber Cancer Institute) for MOC1 and MOC2 cells, R. Ferris (University of North Carolina) for PCI-15B cells and C. Lee-Poturalski for her input on the paper. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
N.L.S, J.D., T.D.K, J.A., R.G., H.S., A. Santos, M.D. and D.J.M. performed the experiments. N.L.S., J.D., T.D.K., L.N-R., V.M., D.J.H.S. and D.J.M. analyzed the data. R.F. oversaw the CIAO clinical trial with N.G. and A.G.S. J.D., T.D.K, J.A., R.G., H.S., T.A., L.N-R., V.M., D.J.H.S., S.V.K., A. Santos, R.A., A.B., M.D., N.G., A.G.S., E.J.S., A. Stacy, C.J., T.A.C. and R.F. reviewed and approved the final paper. N.L.S and D.J.M. conceptualized and oversaw the study. N.L.S and D.J.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
T.A.C. is a cofounder of Gritstone Oncology and holds equity. T.A.C. holds equity in An2H. T.A.C. acknowledges grant funding from Bristol-Myers Squibb, AstraZeneca, Illumina, Pfizer, An2H and Eisai. T.A.C. has served as an advisor for Bristol-Myers, MedImmune, Squibb, Illumina, Eisai, AstraZeneca and An2H. T.A.C. is an inventor on intellectual property held by Memorial Sloan Kettering Cancer Center on using TMB to predict immunotherapy response, with a pending patent, which has been licensed to PGDx. E.J.S. consults for Siren Biotechnology and has immunotherapy patent applications under option to license agreements with iOncologi. N.G. reports grants or contracts and personal or consulting fees from Regeneron Pharmaceuticals, personal or consulting fees from DragonFly Therapeutics, Intuitive Surgical, Merck, Replimune and Sanofi/Genzyme US Companies and support for other professional activities from PDS Biotechnology Corporation outside the submitted work. R.F. reports consulting or advisory roles for Regeneron, Sanofi, Ayala Pharmaceuticals, Prelude Therapeutics, Elevar Therapeutics, G1 Therapeutics, Guidepoint, Expert Connect, Remix, Eisai and Bioatlas in the past 24 months and research funds from Prelude, Ayala, Merck, Genentech, Pfizer, Rakuten, Nanobiotix, EMD Serono, ISA, Viracta and Gilead in the past 24 months. D.J.M. holds intellectual property for transcriptional signatures to predict response to ICB. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cancer thanks N. Gopalakrishna Iyer, Bertrand Routy, Daniel Spakowicz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quantification of tumor bacteria burden from bulk DNA sequencing.
(A) Comparison of average relative abundance of bacterial genera in oral cavity tumors quantified from TCGA WGS data in this study (n = 87) to results from 16S rRNA sequencing from Michikawa et al.12 (n = 33). Two-tailed Pearson correlation coefficient. (B) Comparison of average relative abundance of bacterial genera in oral cavity tumors quantified from TCGA WGS data from the retracted work by Poore et al.27 (n = 157) to results from 16S rRNA sequencing from Michikawa et al.12 (n = 33). Two-tailed Pearson correlation coefficient. (C) Comparison of relative bacteria abundance by 16S rRNA sequencing in oral cavity samples (n = 66) compared to sham controls (n = 22). Red highlighted taxa indicate a subset of contaminants identified. See Table S1. (D) Comparison of average relative bacterial abundances at the phylum level in a cohort of samples analyzed by both whole exome sequencing and 16S rRNA sequencing. Two-tailed Pearson correlation coefficient. n = 22 (E) Sample-wise two-tailed Pearson correlation coefficients of average relative bacterial abundance at the phylum level in samples profiled by both whole exome sequencing and 16S rRNA sequencing. Shaded area indicates 95% confidence interval of correlation coefficients for unmatched samples (for example random pairs). n = 22. (F) Comparison of average relative bacterial abundances at the genus level in a cohort of samples analyzed by both whole exome sequencing and 16S rRNA sequencing. Two-tailed Pearson correlation coefficient. n = 22. (G) Sample-wise two-tailed Pearson correlation coefficients of average relative bacterial abundance at the genus level in samples profiled by both whole exome sequencing and 16S rRNA sequencing. Shaded area indicates 95% confidence interval of correlation coefficients for unmatched samples (for example random pairs). n = 22. (H) Comparison of tumor bacteria burden determine by whole exome sequencing with 16S rRNA qPCR quantification of intratumoral bacteria. Two-tailed Pearson correlation coefficient. n = 22. (I) Pearson correlation coefficients (two-tailed) of TBB determined from imaging 16S rRNA in situ hybridization with TBB determined by qPCR for 16S rRNA. Shaded region indicates detection limit. n = 17. (J) Pearson correlation coefficients (two-tailed) of TBB determined from imaging 16S rRNA in situ hybridization on two slides of the same tumor. Shaded region indicates detection limit. n = 17.
Extended Data Fig. 2 Evaluation of tumor bacteria burden in TCGA samples.
(A) Comparison of TBB determined based on whole genome sequencing (WGS) and whole exome sequencing (WES) in samples with both approaches available. Two-tailed Spearman correlation coefficient. n = 155. (B) Correlation of average relative microbial abundances between WGS and WES in matched samples. Analysis was restricted to WES samples with ≥50 total microbial reads for calculation of relative abundance values. Two-tailed Pearson correlation coefficient. n = 67. (C) Sample-wise comparison of relative microbial abundances determined by WGS and WES in matched samples. Each dot represents the two-tailed Pearson correlation coefficient. correlation coefficient from an individual tumor sample. Analysis was restricted to WES samples with ≥50 total microbial reads for calculation of relative abundance values. Median with interquartile range. The shaded region represents the mean with 95% confidence interval for correlation of unmatched samples. n = 67. (D) Comparison of tumor bacteria burden determined by whole genome sequencing (WGS) based on tissue collection site in oral cavity HNSC samples. Linear regression shown +/- 95% confidence interval. Raw P-value and P-value adjusted for multiple comparisons by Benjamini-Hochberg procedure shown. (E) Comparison of tumor bacteria burden determined by whole genome sequencing based on DNA sequencing site in oral cavity HNSC samples. Each dot represents a single tumor; mean +/- stdev. One-way ANOVA. HMS n = 62; MDACC n = 20; BCM n = 16; BI n = 8. (F) Comparison of tumor bacteria burden determined by whole exome sequencing (WES) based on tissue collection site in oral cavity HNSC samples. Linear regression shown +/- 95% confidence interval. Raw P-value and P-value adjusted for multiple comparisons by Benjamini-Hochberg procedure shown.
Extended Data Fig. 3 Correlation of bacterial species with tumor bacteria burden.
(A) Correlation of relative fractions of Fusobacterium, quantified as fraction of reads out of all mapped bacterial reads with TBB, quantified as bacterial reads per million human reads, in all TCGA HNSCC samples. Two-tailed Spearman correlation. n = 130. (B) Correlation of relative fractions of Fusobacterium, quantified as fraction of reads out of all mapped bacterial reads, with tumor bacteria burden in only oropharynx HNSCC samples from TCGA. Two-tailed Spearman correlation. n = 50. (C) Breakdown of relative proportion of bacteria in TCGA WGS HNSCC samples into anaerobic, aerobic, or not classified (unknown or genera with mixture of both aerobic and anaerobic). Inset numbers indicate mean +/- std. n = 130. (D) Correlation of tumor bacteria burden, defined as bacteria reads per million human reads, with total anaerobic bacteria, defined as anaerobic bacteria reads per million human reads, in TCGA HNSCC samples profiled by WGS. Two-tailed Spearman correlation coefficient. n = 130. (E) Correlation of tumor bacteria burden, defined as bacteria reads per million human reads, with total aerobic bacteria, defined as aerobic bacteria reads per million human reads, in TCGA HNSCC samples profiled by WGS. Two-tailed Spearman correlation coefficient. n = 130. (F) Correlation of tumor bacteria burden, defined as bacteria reads per million human reads, with relative anaerobic bacteria, defined as anaerobic bacteria reads per total bacteria reads, in TCGA HNSCC samples profiled by WGS. Two-tailed Spearman correlation coefficient. n = 130. (G) Correlation of tumor bacteria burden, defined as bacteria reads per million human reads, with relative aerobic bacteria, defined as aerobic bacteria reads per total bacteria reads, in TCGA HNSCC samples profiled by WGS. Two-tailed Spearman correlation coefficient. n = 130.
Extended Data Fig. 4 Clinical correlates with tumor bacteria burden in HNSCC.
Comparison of TBB based on (A) Hypoxia score determined from RNAseq (n = 157), (B) pathological stage (Stage I n = 10; Stage II n = 25; Stage III n = 23; Stage IV n = 74), (C) clinical stage (Stage I-II n = 38; Stage III n = 31; Stage IV n = 87), (D) tumor grade (Grade 1-2 n = 108; Grade 3 n = 44), (E) tumor grade in only oral cavity tumors (Grade 1-2 n = 79; Grade 3 n = 26), (F) sex (Male n = 114, Female n = 43), (G) ethnicity (White n = 141, Black/Afr. American n = 12, Asian n = 2), (H) patient age (n = 157), (I) subsite within the oral cavity (oral tongue n = 55, not otherwise specified n = 25, floor of mouth n = 12, other = 8, alveolar ridge n = 6), (J) HPV status for only oropharynx tumors (HPV+ n = 23, HPV- n = 3), (K) alcohol history (high n = 50, no/low n = 21), or (L) smoking status (n = 155). (M) Multivariable regression of HPV status and smoking status (current versus never), error bars represent 95% confidence interval. Unless otherwise noted, comparisons between two groups were made with a rank-sum test, comparisons between more than two groups were made with a Kruskal-Wallis test, and comparisons between two continuous variables were made using a Spearman correlation coefficient. Center white point is median, box is interquartile range.
Extended Data Fig. 5 Association of tumor bacterial burden with mutations.
(A) Difference in WES tumor bacteria burden (TBB) in HPV-negative HNSCC tumors based on mutations in specific genes relevant to HNSCC. Two-tailed Welch’s t-test. Highlighted values indicate P-values<0.05. n = 415. (B) Association of WES TBB with mutations in specific genes relevant to HNSCC and HPV status assessed by multivariable regression of log-transformed TBB with mutation status + HPV status. Highlighted values indicate P-values<0.05. n = 467.
Extended Data Fig. 6 Gene expression changes in cell lines infected with tumor-relevant bacteria in vitro.
(A-D) Comparison of gene expression changes following infection with either Fusobacterium nucleatum or Prevotella scopos in human HNSCC cell lines OQ01 (A), SCC25 (B), FaDu (C), and PCI-15B (D). Pearson correlation coefficients. n = 2 technical replicates per cell line from one representative experiment. (E-J) Expression of indicated genes in OQ01 cells infected with either live or dead (heat-killed) Fusobacterium nucleatum compared to mock-infected control. n = 4. Mean with s.e.m. (K) Comparison of gene expression changes following infection with either Fusobacterium nucleatum or Prevotella scopos in mouse cell line MOC1. Pearson correlation coefficients. n = 3 technical replicates from one representative experiment. (L) Comparison of average gene expression alterations following exposure to Prevotella scopos and Fusobacterium nucleatum in mouse (n = 1 cell line in technical triplicate) and human (n = 4 cell lines in technical duplicate from one representative experiment per cell line) HNSCC cell lines. Inset value indicates Spearman correlation coefficient. (M) Comparison of average gene expression alterations following exposure to Prevotella scopos and Fusobacterium nucleatum in MOC1 mouse cancer cells in vitro (n = 1 cell line in technical triplicate from one representative experiment) and orthotopic MOC1 tumors treated with antibiotics in vivo (n = 5). Dotted line indicates 0 for change post-antibiotics.
Extended Data Fig. 7 Murine intratumoral microbiome.
(A) Relative abundance of indicated bacteria in MOC1 orthotopic tumors, as determined by 16S rRNA sequencing. Showing taxa detected in at least two tumors. n = 5. (B) Average relative abundance for all detected taxa in MOC1 orthotopic tumors, as determined by 16S rRNA sequencing. n = 5.
Extended Data Fig. 8 Effect of antibiotics on immune checkpoint blockade response in HNSCC.
Clinical benefit rate for patients with HNSCC receiving anti-PD(L)1 immune checkpoint blockade in the presence or absence of concurrent antibiotics. Inset fraction indicates number with clinical benefit over total sample size. n = 111 total. Two-tailed Chi-square test.
Extended Data Fig. 9 In vivo immune cell depletion.
C57BL/6 mice were treated with depletion antibodies and whole blood was collected week 3 for flow cytometry analysis. (A) Blood was stained with anti-CD45, anti-CD4 and anti-CD8, followed by flow cytometry sorting of CD45+CD4+ and CD45+CD8+ population. Flow Cytometry gating strategy to identify CD45+CD4+ and CD45+CD8+ cells shown. (B and C) Quantification of percentage of subpopulations of sorted CD4+ and CD8+ cells in the CD45+ population. Data are presented as mean ± SEM (n=3). P values by two-tailed unpaired t test. (D) Blood was stained with anti-CD45, anti-CD11b, anti-Ly6G followed by flow cytometry sorting of CD45+CD11b+Ly6G+ population with the gating strategy. (E) Quantification of percentage of subpopulations of sorted Ly6G+ cells in the CD45+CD11b+ population. Data are presented as mean ± SEM (n=3). P values by two-tailed unpaired t test.
Extended Data Fig. 10 Depletion of oral microbiome with antibiotics.
(A) Culture of bacterial swabs in either aerobic or anerobic conditions from control mice and mice treated with antibiotics (ABX) for four days prior to tumor injection. Three representative technical replicates from two biological replicates shown. (B) Quantitative PCR on bacterial swabs from control mice and mice treated with antibiotics (ABX) for four days prior to tumor injection. Each dot represents an individual mouse. Median +/- interquartile range. Rank-sum test. n = 5.
Supplementary information
Supplementary Tables 1–7 (download XLSX )
Supplementary Tables 1–7.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Silver, N.L., Dai, J., Kerr, T.D. et al. Intratumoral bacteria are immunosuppressive and promote immunotherapy resistance in head and neck squamous cell carcinoma. Nat Cancer 7, 80–97 (2026). https://doi.org/10.1038/s43018-025-01067-1
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s43018-025-01067-1