Abstract
Coral bleaching events from thermal stress are increasing globally in duration, frequency, and intensity. While bleaching can cause mortality, some corals survive, reacquire symbionts, and recover. We experimentally bleached Montipora capitata to examine molecular and physiological differences between corals that recover (resilient) and those that die (susceptible). Corals were collected and monitored for eight months post-bleaching to identify genets with long-term resilience. Using an integrated systems-biology approach that included quantitative proteomics, 16S rRNA sequencing to characterize the coral microbiome, total coral lipids, symbiont community composition and density, we explored molecular-level mechanisms of tolerance in corals pre- and post-bleaching. Prior to thermal stress, resilient corals have a more diverse microbiome and abundant proteins essential for carbon acquisition, symbiont retention, and pathogen resistance. Protein signatures of susceptible corals showed early symbiont rejection and utilized urea for carbon and nitrogen. Our results reveal molecular factors for surviving bleaching events and identify diagnostic protein biomarkers for reef management and restoration.
Similar content being viewed by others
Introduction
Coral reefs are one of the most diverse and structurally complex ecosystems on Earth, providing shelter and habitat for many organisms1. Tens of millions of people in more than 100 countries have coastlines adjacent to coral reefs and depend on them for their livelihoods2. Unfortunately, coral reefs are declining rapidly throughout the world due to pollution, coastal development, overexploitation3,4, and effects associated with climate change5,6. As global seawater surface temperatures increase and marine heatwaves sweep coastlines, large-scale thermal-induced coral bleaching events are becoming increasingly common worldwide7,8,9.
When water temperatures exceed local thermal thresholds for a given coral, the association between the coral host and its endosymbiotic algae, Symbiodiniaceae, breaks down. This breakdown is termed “coral bleaching” due to the loss of the pigmented symbionts3. In addition to the distinct colors, Symbiodiniaceae provide much of the energetic requirements for the host in the form of organic carbon and nitrogen photosynthetic byproducts10. The expulsion of Symbiodiniaceae during bleaching events metabolically compromises the host11, leading to reduced physiological performance, reduced reproductive capacity12,13,14, and can lead to widespread mortality15,16.
As a holobiont, corals host a diverse community of organisms, including algal endosymbionts, and a bacterial, archaeal, and eukaryotic microbiome that resides on and within the coral tissue, coral mucus, the coral gastric cavity and within endolithic skeleton The diversity of the coral microbiome frequently decreases with thermal-induced bleaching17,18. While some of these changes may be associated with the loss of Symbiodiniaceae, other changes in microbiome diversity appear to be directly driven by thermal stress19. If bleaching alters the microbiome in ways that disrupt beneficial bacteria or archaea that support coral metabolism or defend against pathogens, these changes may contribute to coral mortality after bleaching.
Coral microbiomes demonstrate host specificity, bolstering the hypothesis that bleaching-induced microbiome disruptions play an important role in host mortality20,21,22,23. Microbiome disruption can remove defensive benefits of normal coral microbiomes, which may include reducing pathogen infection success via multiple mechanisms (e.g., spatial occlusion and competition, direct predation, metabolic jamming, and quorum quenching)24,25,26,27. While gnotobiotic models are now being developed28, fully unraveling the complex relationships between corals and their microbiomes remains a challenging task. Moreover, although genomic evidence and correlations between taxonomic abundance and disease provide some hints, it is unclear which specific microbiome changes associated with bleaching or other stressors are helpful or harmful for the survival of the coral host. Indeed, while a rich literature documents alterations to microbiome structure and stability by many specific stressors and coral diseases—including changes to beta-diversity29,30, richness31, and the abundance of particular taxa—far fewer data are available on which of these diverse microbiome changes best predict the future survival of coral hosts. Thus, data linking microbiome changes with subsequent coral survival could be vital to interpreting the ecological consequences of shifts in bleaching-induced microbiome structure for corals.
Coral death due to the interconnected physiological and microbiological consequences of bleaching events can lead to total community collapse if mortality is widespread32,33. However, some coral species, and individuals within species, are resilient to the effects of bleaching and can recover. Corals resilient to thermal stress can reacquire symbionts, fully recover physiological performance and yield viable gametes34, although the timescale for post-bleaching recovery can vary from weeks to a year12,35,36,37. Post-bleaching recovery times may be shorter in corals that have previously been exposed to multiple annual bleaching events38, perhaps by priming the physiological responses necessary to survive bleaching.
There are several possible contributing factors that may provide greater coral holobiont tolerance and resilience. We define tolerance as the physiological traits that enable corals to avoid bleaching under stressful conditions. Resilience is the ability of a coral reef system to absorb or recover from disturbances like bleaching while maintaining its ecosystem functions39. Some of the factors included are species specific at the host and/or symbiont level40,41,42,43, or are dependent on host genotype44,45,46,47,48,49, or the microbial constituents of the holobiont’s microbiome50,51. At the center of this complex equation is the host’s metabolic capacity before, during, and after bleaching. Without adequate carbon and nitrogen, the coral cannot maintain systems of cellular and tissue repair, retain or reacquire symbionts, or fight off pathogens52,53. This suggests that the thermal threshold and physiological condition of the coral host may be the most important factors in determining resilience to bleaching stress and mortality.
The interactions between an organism and its environment are complex and depend upon ecological and evolutionary history. Corals in Kāneo’he Bay, O’ahu, Hawai’i have endured an increased frequency of bleaching events since 1996, similar to worldwide trends54,55,56. Results from bleaching surveys in the bay have shown a decrease in the proportion of corals bleached over time (62% in 1996 to 30% in 2015) but an increase in the number of corals dying (<1% in 1996 to 22% in 2015)57. Reefs in Kāneo’he Bay are dominated by the reef-building Scleractinian coral Montipora capitata58. These corals are typically found in tropical waters, living within 1–2 °C of their upper thermal limit59,60. This suggests that projected future water temperatures will specifically threaten this species. However, M. capitata has been shown to have higher thermal tolerance than other Hawaiian coral species61 and is thereby an ideal species to study the long-term effects of thermal acclimatization or adaptation to reveal the underlying molecular physiology that supports bleaching resilience.
In this study, we report the results of a joint coral physiology, proteomic, lipid, and microbiome analysis comparing M. capitata that recovered in situ from experimental bleaching against those that died. Visually healthy M. capitata from Kāneo’he Bay were exposed to an experimentally simulated thermal stress (30 °C for three weeks) coincident with the typical summer thermal maximum, to induce bleaching. Samples for proteomics, bacterial and archaeal microbiome diversity, total lipids, Symbiodiniaceae density and diversity were collected before thermal stress (T1) and four weeks after bleaching was initiated (T2) (Fig. 1). Paired controls collected from the same source coral did not undergo thermal-induced bleaching. Following the experimental treatment, corals were outplanted back to a natural reef and monitored for eight months to distinguish those that recovered from the thermal-induced bleaching event and those that died. These mortality outcomes were then retroactively used to label the previously collected samples as either resilient (R) or susceptible (S) corals. The proteomic, microbial, and physiological differences uncovered in this study describe intraspecific differences associated with bleaching resilience in the field. Because corals were collected from and outplanted to the same location at Moku O Lo’e island in Kāneo’he Bay, they experienced similar environmental conditions throughout the recovery period.
a, b Average total cell counts (x109) for Symbiodiniaceae per gdw−1 (n = 6 genets for each point, error bars represent standard deviations) illustrating the experimental design to track effects of thermally induced bleaching on 12 Montipora capitata coral genets that were monitored for 8 months. Paired subsamples of all 12 coral genets were either maintained at ambient temperatures (a) or exposed to thermal bleaching (b). Symbiodiniaceae density for the (a) control cohort maintained at ambient (25 °C) temperature and (b) the experimental cohort that underwent thermally induced bleaching (30 °C) for 4 weeks; light green: (n = 6 genets) resilient corals reacquired symbionts and recovered post-bleaching; light purple: (n = 6 genets) corals susceptible to thermally induced bleaching that did not recover and died. Coral sub-samples were collected before (T1) and after exposure to thermal stress (T2) to assess host performance and symbiont and microbial composition in corals. Note that resilient and susceptible corals were identified three months later (December); after this period, no additional mortality was observed. c Symbiont community composition in the resilient and susceptible genets prior to thermal stress (T1) for resilient corals (green: dark green Cladocopium, medium green Durusdinium, light green mixed community) and susceptible corals (purple: dark purple Cladocopium, medium purple Durusdinium, light purple mixed community) with reported percentages and number of samples used for community composition (n = 6 genets and n = 4 genets*, respectively). *The variation in the number of genets analyzed was due to limitations in the size of the archived sample collection.
Comparison of resilient vs. susceptible M. capitata using mass spectrometry based proteomics allowed for the generation of metabolic maps of abundant enzymes and outlined the coral’s energetic priorities that may confer resilience in a changing climate. Several proteins significantly differed between resilient and susceptible corals prior to experimental thermal stress. These resilience-associated proteins suggest nutritional and metabolic strategies may underpin the ability to survive bleaching. The proteome also revealed evidence for symbiont disassociation, antiviral activity, enhanced immune response to pathogens, and carbon and nitrogen pathways exclusive to resilient corals. Additional physiological and molecular-level metrics of the holobionts were monitored, including total lipids, bacterial and archaeal microbiome diversity, and symbiont density and genus, allowing us to identify whole-organism biomarkers of resilience. These molecular-level signatures could be used to predict coral resilience or susceptibility prior to a bleaching event. Last, we identify biomarkers in corals that have the potential to predict survivorship during future bleaching events, the linchpin to coral management, propagation efforts, and restoration success.
Results
In September 2017, seventy-four Montipora capitata were tagged and collected. Corals were cut in half, acclimated in tanks for two weeks, and sampled (T1). Then, each coral half was either maintained at ambient temperatures (27.7 °C ± 0.6, range = 25.8–28.8 °C) for the duration of the experiment (Fig. 1a) or gradually exposed to increasing water temperatures to reach 30 °C. Coral fragments in the high-temperature treatment were held at 30 °C ± 0.5 °C (range = 28.9–31 °C) for three weeks to simulate a thermal bleaching event (Fig. 1b). After the thermal stress, experimental tanks were returned to ambient temperatures over four days. After 24 h at the ambient temperature, corals were sampled again (T2), placed on racks in the natural environment, and monitored for long-term survival and recovery for eight months (Fig. 1a, b, and Fig. S1). After the first three months (December; Fig. 1b), 22 corals died; these coral genets are referred to as “susceptible” (i.e., S). Fifty-two corals recovered and reacquired symbionts; these coral genets are referred to as “resilient” (i.e., R). After December, no additional coral mortality was observed. By May, all the corals that survived reached pre-bleaching coloration (Fig. 1b). At T1 and T2, we obtained coral samples to examine physiological performance and recovery. Six of the 52 resilient corals and six of the 22 susceptible corals were randomly selected for this study and frozen sub-samples from both timepoints were used for mass spectrometry-based proteomics, total lipid content, microbial community composition, Symbiodiniaceae density, and dominance by Cladocopium vs. Durusdinium Symbiodiniaceae. Because our microbiome assays were based on 16S rRNA primers targeting bacteria and archaea, microbiome results describe the bacterial and archaeal portions of the microbiome (but not microbial eukaryotes).
Symbiodiniaceae community
At T1, no significant difference in total symbiont density was detected between the resilient and susceptible corals (p = 1; Fig. 1c, Supplementary Data 1a). During the thermal stress event, symbiont density decreased in both cohorts at the same rate; within three months after T2, the resilient cohort returned to pre-bleaching Symbiodiniaceae density. Corals were classified as being Cladocopium- or Durusdinium-dominated or having mixed communities (see Supplemental Methods; Fig. S2, Table S1). Notably, there were representatives of each community type in both the resilient and susceptible categories (Fig. 1c). A logistic regression analysis was conducted to assess whether the proportion of Durusdinium symbionts influenced survival rates. The analysis indicated a positive association between the proportion of Durusdinium symbionts and post-bleaching mortality (coefficient = 2.38); however, this relationship was not statistically significant (p = 0.342). Additionally, the model’s intercept was not statistically significant (p = 0.345), suggesting that baseline mortality was not strongly predicted in this dataset (Supplementary Data 1b).
Biological enrichment analysis of proteomic data
A total of 2193 coral proteins were identified at T1, 2161 coral proteins were identified at T2, and 1424 coral proteins were shared between those timepoints, indicating constitutive expression (Fig. S3, Supplementary Data 2a–g). Analysis of the resilient and susceptible corals independent of timepoint revealed 2276 proteins detected in the resilient cohort and 2066 proteins in susceptible corals.
Enrichment analysis of Gene Ontology (GO) terms was completed using MetaGOmics62, an unbiased method to identify significantly different metabolic processes represented in proteomes of the resilient vs. susceptible cohorts at both timepoints using log-fold change (base 2; LFC) of GO term assignments (Supplementary Data 3a, b). Resilient corals at T1 were characterized by multiple cellular responses to external signals, receptor activity, and monosaccharide binding. Sterol esterase activity was the most enriched term (LFC = 5.0; Fig. 2a). Six GO terms were significantly enriched in susceptible corals prior to bleaching (T1S) and included proteins involved in urea and amide catabolism, nickel binding, and the removal of superoxides (Fig. 2a). After thermal bleaching, 51 GO terms were enriched in resilient corals, 33 of which were unique (Fig. 2b). Proteomes of resilient corals post-bleaching (T2R) were enriched in regulation of phagocytosis and meiotic cell cycle, vesicle-mediated transport, hormone regulation, and cardiac muscle processes. Although some of the labels for these processes may not seem to apply to corals, proteins associated with the “cardiac muscle process” GO term are involved in sodium and calcium exchange, while “regulation of systemic arterial blood pressure” proteins include sodium-driven chloride bicarbonate exchange proteins involved in pH regulation. Many of the identified GO terms in susceptible corals post-bleaching involved the cellular processing of metabolites, including sterols, methionine, betaine, sarcosine, lipids, and glutamate. Additionally, several terms were related to DNA- or RNA-processing.
Molecular data on proteins (a–d) and lipids (Fig. 4a, b), completed on the same 12 coral genets used throughout the study (n = 6 susceptible, n = 6 resilient). a, b Volcano plots depicting -Log (p-value) vs the Laplace corrected Log Fold Change (LFC) for protein-associated Gene Ontology terms. Colored dots indicate GO terms that were significantly different between resilient and susceptible corals (Laplace corrected Log fold ≤−0.5 or ≥0.5 and p ≤ 0.01). a T1 with negative LFC indicating proteins more abundant in resilient corals (T1R) while positive values correspond to proteins that were at higher abundance in susceptible corals (T1S). b T2, where negative LFC indicates greater abundance in resilient (T2R) M. capitata while GO terms with positive values are higher in susceptible corals (T2S). c, d Volcano Plots of individual protein abundances. LFC ≤ −0.5 are proteins that were detected in significantly higher abundance in the resilient coral cohorts (green) at c T1 and d T2. LFC ≥ 0.5 are proteins that were detected in higher abundance in the susceptible coral cohorts (purple) before thermal-stress-induced bleaching (c: T1) and after (d: T2).
Immune system responses
To elucidate complete metabolic pathways being preferentially utilized by either the resilient or susceptible corals at the two timepoints, significant differential abundances of proteins were calculated (Fig. 2c, d; significance only reported when p ≤ 0.05; Supplementary Data 2g, h). Comparisons of resilient and susceptible coral proteomes before the simulated thermal bleaching event (T1), revealed that resilient corals had 39 proteins at significantly higher abundances than the susceptible cohort, whereas susceptible corals possessed 56 proteins at significantly higher abundances (Fig. 2c). In T1R corals, signal peptidase (LepB) yielded the highest differential abundance, followed by F-type H + -transporting ATPase subunit alpha (ATPeF1A), and a membrane attack complex component/perforin (MACPF) domain-containing protein known to lyse virus-infected and pathogenic bacterial cells. Susceptible corals before bleaching (T1) significantly increased the abundance of Fibropellin-1 (EGF1), a component of the apical lamina, the surface glycoprotein melanoma-associated antigen p97 (MFI2), an enzyme involved in glycosylating proteins, alpha 1,2-mannosyltransferase (KTR1_3), and the urea degrading enzyme urease (URE1).
After the corals were bleached (T2), the resilient cohort was characterized by 108 proteins that significantly increased in abundance between timepoints, while 63 proteins significantly increased in abundance in the susceptible cohort between T1 and T2 (Fig. 2d). A CyanoVirin-N domain-containing protein (CVNH; Fig. 2d) was identified to have the most consistent expression across the 12 corals tested (i.e., lowest p-value), a high LFC in the T1R proteome (compared to T1S), and the highest LFC in T2R (compared to T2S) (Fig. 2d). Further analysis of this protein sequence against the conserved domain database (Fig. S5) revealed that it contains four CyanoVirin-N conserved domains with viricidal activity that interact with the glycoproteins on the viral envelope63. Conversely, post-bleaching (T2S), chitinase (CHIC), an enzyme capable of degrading chitin, exhibited the highest LFC in the susceptible corals (Fig. 2d).
Cluster analysis of proteins that were identified across all experiments to be significantly abundant in at least one treatment (LFC ≥ | 1 | , p < 0.01) are represented in a heatmap that spans fifteen metabolic pathways (Fig. 3). Resilient and susceptible coral proteomes are most similar at T1 (Fig. 3: clusters 1–3, 9–12) compared to the proteomes at T2 (Fig. 3). Furthermore, resilient corals at T1 (prior to bleaching) exhibited higher abundances of several proteins involved in anti-viral activity, immune response, and symbiosome maintenance (Fig. 3: clusters 4–7) than susceptible corals. After thermal bleaching, the resilient corals maintain a significantly higher abundance of six enzymes, including CNVH (Fig. 3: cluster 1), compared to susceptible corals. The T2R cohort uniquely increased the abundance of 37 additional proteins (Fig. 3: cluster 3) involved in nitrogen metabolism, immune response, endosome/symbiosome activity, and DNA translation, among others. Pre-bleaching susceptible corals (T1S), despite exhibiting somewhat similar proteomic trends to T1R, revealed one unique cluster (Fig. 3: cluster 8) of 9 proteins that were significantly increased in abundance. These proteins play a role in structures and functions such as the extracellular matrix, and immune response, or are associated with lysosome/phagosome activity. Post-bleaching susceptible corals (T2S) had the most distinct proteomic response, with depletion of nearly all proteins represented by clusters 1–8 (Fig. 3) and enrichment in a unique suite of proteins in carbon, nitrogen and lipid metabolism, the biosynthesis of secondary metabolites, the extracellular matrix, and the immune system (Fig. 3: clusters 11–12).
Heatmap shades of blue indicate averaged normalized spectral abundance factor (NSAF) values for bioreplicates per condition, normalized by the row mean. Rows are clustered using a correlation algorithm and a dendrogram was set to cut 12 distinct clusters (indicated by #s 1–12 black). Right panel dot-matrix indicates metabolic categories identified through KEGG, UniProt and GO (D) viral defense or (R) reproduction, CO2 or as a substrate (CO2-) or product (CO2+), or Ca- binding domain (i.e., Ca-binding).
Resilient corals retain lipids through the thermal bleaching event
Previous investigations on recovery from thermal bleaching events revealed that M. capitata, unlike other corals, has the unique ability to replenish energy reserves within 1–2 months after the bleaching event64. Pre-bleaching, resilient corals had significantly greater abundances of enzymes involved in lipid degradation compared to the susceptible cohort (e.g., PSAP and LIP, Fig. 2a). Total lipids were measured to determine if pre-bleaching lipid biomass (i.e., T1) is a significant and predictable metric to identify M. capitata corals that will recover from thermal bleaching events. For lipid and microbiome analyses an additional control cohort that was not bleached was followed from T1 through to T2 (as outlined in the methods). T2NB refers to lipids at T2 from Non-Bleached, control corals. Using a two-way analysis of variance (ANOVA), no significant difference in coral lipid content between T1R and T1S corals was found before the simulated thermal event (T1R vs. T1S: p = 0.187; Supplementary Data 1d, e; Fig. 4a). Coral lipid content varied significantly by bleaching status at T2 (T2B vs. T2NB: p = 0.00079; Fig. 4b) and resilience, or recovery capacity after bleaching (T2R vs. T2S: p = 0.00095; Fig. 4a). Interaction of the two variables (i.e., bleaching status and timepoint) was also significant (p = 0.044): susceptible corals experienced a decrease in mean lipid content by 44% after exposure to thermal stress (T1S and T2S), while resilient corals decreased by only 16% (T1R and T2R).
Average lipid biomass (g/g dry weight) measurements and reported standard deviations (error bars) on all resilient (green n = 6 genets) and susceptible (purple n = 6 genets) samples from the different cohorts at a T1 before bleaching and b T2 after bleaching (gray bars with “B” labels) and non-bleached controls maintained at 25 °C (light green and purple bars with “NB” labels).
Resilient corals have more diverse bacterial communities
To understand the effects of bleaching on the coral-associated microbiome, phylum-level composition and alpha diversity were analyzed using 16S rRNA sequencing (Fig. 5). Alpha diversity of the microbiome based on the 16S rRNA V4 variable region was quantified using Faith’s phylogenetic diversity (Faith’s PD), a measure of microbiome richness that accounts for phylogeny. Faith’s PD significantly differed among groups defined by each combination of timepoint, bleaching resilience, and bleaching status (Kruskal Wallis p = 0.0045). Prior to bleaching (T1), resilient M. capitata had higher alpha diversity compared to susceptible corals (Kruskal Wallis p = 0.016; Fig. 5a). However, this difference could be attributable to multiple comparisons (false discovery rate (FDR) q = 0.061). After bleaching, previously low-diversity susceptible corals exhibited significantly increased alpha diversity (p = 0.0065, FDR q = 0.048), while previously high-diversity resilient corals did not (p = 0.52, FDR q = 0.6). Thus, resilient corals showed smaller microbiome changes during bleaching and greater stability in microbiome richness over time than susceptible corals.
a Boxplots display microbiome alpha diversity (as measured by Faith’s Phylogenetic diversity) for resilient (R, green) and susceptible (S, purple) coral hosts across two timepoints (T1 and T2) and bleaching conditions. Data points represent individual samples, with boxplot whiskers indicating variability outside the upper and lower quartiles. The y-axis categorizes corals based on their susceptibility to heat stress-induced bleaching and treatments. b T1 (T1R: dark green, T1S: dark purple), c T2 after thermal stress (T2R: gray with green outline, T1S: white with dark purple outline), and d T2 Non-Bleached control (NB) samples not exposed to thermal stress (NBT2R: light green, NBT2S: light purple). Coral genet ID numbers are listed on the x-axis. These same 12 coral IDs were used for all analyses presented. Size of the dot represents the log transformation of the phylum-level counts as indicated in legend presented in (a).
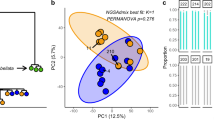
Beta diversity was also quantified to determine the similarities of the bacterial communities between cohorts. Across combinations of timepoint, resilience, and bleaching status, microbial community composition differed qualitatively (Unweighted UniFrac PERMANOVA; p = 0.002) and quantitatively (Weighted UniFrac PERMANOVA; p = 0.041). These differences were not attributable to differences in microbiome dispersion (Weighted and Unweighted UniFrac p > 0.05).
The main taxonomic drivers of community differences revealed that Gammaproteobacteria were well represented in the six resilient corals and nearly absent in susceptible corals (Fig. 5b–d), consistent with the overall community differences detected in beta-diversity analysis. At the family level, multiple microbial families showed striking differences between the resilient and susceptible cohorts. The clearest of these differences was seen in Moraxellaceae, a family in class Gammaproteobacteria that was only found in resilient corals at T1. Microbiome Multivariate Association with Linear Models65, a statistical analysis method that identifies multivariable association between microbial features and metadata (i.e., time, resilience, bleaching), confirmed that Moraxellaceae significantly correlated with both time and resilience (MaAsLin2TIME p = 0.0001; MaAsLin2RvS p = 0.0007; Table S2). Additionally, the Caulobacteraceae microbial family—in the phylum Proteobacteria—was present at elevated abundance in resilient corals, and lower abundance in susceptible ones, irrespective of timepoint or bleaching status (MaAsLin2RvS p = 0.0006).
PICRUSt2 analysis66 was performed on coral microbiomes from resilient (T1R) and susceptible (T1S) cohorts prior to bleaching to estimate their predicted functional capacities. While a comprehensive analysis was beyond the original scope of the study, several notable functional differences emerged (Fig. S6a), shedding light on the potential ecological dynamics of these microbiomes. Among the most significantly altered metabolic pathways as determined by MetaCyc (a database of metabolic pathways), was a lower predicted capacity for antibiotic production via the (5 R)-carbapenem carboxylate biosynthesis pathway in resilient corals compared to susceptible corals. The PICRUSt2 analysis also revealed a lower predicted capacity for aerobactin biosynthesis in the microbiomes of resilient corals at T1. Aerobactin, a siderophore utilized for iron scavenging by microbes, including some pathogens, may play a role in mediating microbial competition and nutrient acquisition. Additionally, resilient coral microbiomes exhibited a reduction in predicted androstenedione degradation relative to susceptible coral microbiomes at T1. This pathway has been recently implicated as a potential mechanism by which Endozoicomonas bacteria could modulate host hormone signaling, particularly under environmental stress67. Notably, while no consistent functional categories were enriched in the microbiomes of resilient corals at T1, many categories were depleted compared to susceptible corals.
Discussion
Our multidisciplinary approach provides unprecedented biochemical details that link metabolism to expressed physiology in resilient and susceptible Montipora capitata before and after thermal-induced bleaching. Although multiple coral species have had their proteomes analyzed (e.g., Eunicea calyculata68; Acropora microphthalma69; Acropora millepora52, Montipora capitata70,71), to date, no studies have exhaustively explored pre-bleaching protein-level physiology in combination with multiple other molecular-level factors to determine if there are traits predictive of resilience to bleaching events. Previous research has shown that bleaching susceptibility in M. capitata is strongly linked to symbiont community composition, with Durusdinium-dominated colonies being more resistant to bleaching than their Cladocopium-dominated conspecifics72. However, corals with Durusdinium- and Cladocopium-dominated symbiont communities were represented in both the resilient and susceptible cohorts in the present study and logistic regression showed no significant relationship between the proportion of Durusdinium pre-bleaching and post-bleaching mortality (p = 0.342). These findings indicate that symbiont community composition was not a primary driver of mortality following complete bleaching (Fig. 1c, Fig. S2a, b). Our primary hypothesis is that holobiont-specific molecular phenotypic differences will enhance the ability of some individual corals to mitigate the effects of bleaching. Before thermal stress occurred (T1), the only significant differences between R and S corals were protein abundance differences in specific pathways and differences in the microbiome diversity. Examination of the significantly changing metabolic pathways in the resilient and susceptible cohorts reveals how nutritional strategies, antiviral mechanisms, and microbiome diversity pre-and post-bleaching are associated with corals’ likelihood to survive a bleaching event.
Phenotypic advantages in resilient corals prior to bleaching
In general, more proteins were detected in all corals at T2 post-bleaching relative to T1 pre-bleaching, a common response to exogenous stressors73,74,75. Several proteins in metabolic pathways associated with maintenance of a functional symbiont-host relationship were present at increased abundance in the resilient coral proteome prior to the bleaching event. The primary active pathways that were enhanced in resilient corals pre-bleaching include sterol and lipid degradation, cellular respiration, oxidative phosphorylation, and carbon metabolism (Fig. 2a). Despite discovering significant enrichments in proteins associated with the Gene Ontology term lipid degradation, no significant difference in total lipid content between T1R and T1S cohorts was found (Fig. 4a).
Prior to bleaching, resilient corals, compared to the susceptible corals, have an increased relative abundance of enzymes that allow them to utilize heterotrophic feeding and symbiont photosynthate. The GO term sterol esterase was the most significantly increased term in resilient M. capitata at T1. The dominating protein contributors to the enrichment of this GO term included a gastric triacylglycerol lipase-like protein (LIP) (Fig. 3; cluster 4) and the lipid-specific degradation enzyme saposin (PSAP; Fig. 2c). These enzymes are typically involved in digestion and likely reside in the coral gastric cavity, suggesting that at T1, pre-bleaching, the resilient corals are using a heterotrophic feeding strategy in addition to photosynthate from Symbiodiniaceae. This was not unexpected as M. capitata has been previously observed to utilize heterotrophy when not bleached76. T1 resilient corals also possessed a higher abundance of early endosome antigen 1 (EEA1), an essential protein in symbiosis establishment77,78,79, providing resilient corals with an advantage for maintaining symbiont relationships compared to the susceptible corals. Utilization of diverse feeding strategies would provide a distinct advantage to the resilient corals as the excess carbon can be shuttled into lipid storage vesicles76,80.
Resilient corals present evidence of more active cellular respiration pathways at T1, prior to the bleaching event. Although many enzymes in the carbon metabolism pathway are constitutively expressed in resilient and susceptible corals, the increased abundance of isocitrate lyase (ICL; Fig. 2c) enables resilient corals to use the glyoxylate shunt, a TCA-cycle bypass that allows cells to complete anabolic reactions with 2-carbon units without losing carbon as CO2 (the opposite of what is observed in susceptible corals at T1; Fig. S4a). Further, increased abundances of glycine hydroxymethyltransferase (GLYA; Fig. 2c) provides single carbon (1 C) units to the cell, fueling the glyoxylate shunt for the generation of larger carbon-storage molecules (i.e., lipids) from the excess 1–2 carbon unit small molecules, again bypassing the generation/loss of CO281,82. Additional evidence, such as increased abundance of pyruvate dehydrogenase (PDHa; Fig. 2c) and multiple acetyl-CoA transferases, support a hypothesis of increased cellular respiration in resilient corals, which may contribute to both immediate physiological demands and the potential to store excess energy83.
Prior to bleaching, the resilient corals appeared to prime anti-viral activity and exhibited a significantly more diverse microbiome, which likely supported their immune response84,85,86. Previous demonstrations of “frontloading” immune response or pathogen-fighting enzymes have been shown to increase survival in corals87. Cyanovirin-like protein (CVNH) was one of the most differentially abundant proteins that was elevated in resilient corals at T1 (Fig. 2c). CVNH is an evolutionarily conserved protein that binds to viruses and blocks entry into the cell88. CVNH was also detected at higher abundances in the resilient cohort as it progressed through T2, post-bleaching (Figs. 2d and 3, cluster 1). To determine if there were identifiable viral proteins in the mass spectrometry data from the whole-coral protein extractions, the data were re-analyzed using a larger database that included 5 coral-associated viral proteomes (Table S3). The number of confident peptides associated with the viral proteins detected were not statistically different between resilient and susceptible corals at T1, yet their presence in whole-holobiont protein extract corroborates the need for corals to produce antiviral proteins. Further studies should explore the viral influence on downstream susceptibility, including methods such as tracking viral load with transmission electron microscopy to quantify and locate viral particles (i.e., in the endosymbionts or corals). Additionally, the T1R corals hosted significantly more diverse microbiomes (Fig. 5a). The predicted functional capacity of the resilient microbiome (i.e., PICRUSt2; Fig. S6) suggested the possibility that microbial antibiotic production in susceptible corals may contribute to the reduced microbiome richness observed in T1S cohorts, potentially through mechanisms such as microbe-microbe competition. Higher microbiome diversity has previously been reported in more bleaching-resistant species, though this association does not always extend to bleaching susceptible or resilient conspecifics67,89. The resilient microbiome exclusively included the Moraxellaceae bacterial family. Moraxellaceae are associated with local wastewater and they are recognized to have high numbers of antibiotic resistance genes (ARGs)90,91. As the coral host’s immune system is activated against pathogenic bacteria and releases antimicrobial defenses, the Moraxellaceae bacterial family’s high number of ARGs may provide them with an advantage for long-term residence on the host. As Moraxellaceae has been found to be a common component of many shallow water coral microbiomes, these bacteria may be important in shaping a healthy coral holobiont92. The functional role of this bacterial family’s unique genome in conferring resilience against bleaching is unknown, but the close association of Moraxellaceae with resilient M. capitata corals merits further research. Additionally, the PICRUSt2 analysis of coral microbiomes identified notable functional differences in resilient corals prior to bleaching. These results may reflect lower microbiome richness in susceptible corals prior to bleaching and point to higher functional variance in the microbiome of resilient corals, which could confer greater adaptability and an ecological advantage in the face of environmental stressors.
Molecular signs of stress before thermal induced bleaching in susceptible corals
A detailed proteomics analysis revealed the metabolic processes identified in susceptible corals prior to bleaching, including urea and amide catabolism, nickel binding, and urease activity (Fig. 2a). Redirected nitrogen and carbon uptake pathways in susceptible corals suggest a decrease in symbiont-derived photosynthate at T1. Previous molecular-level investigations of symbiont-host relationships have demonstrated that the majority of nitrogen assimilation occurs via the symbiont-directed GS/GOGAT (glutamine synthase/glutamine oxoglutarate aminotransferase) activity or the coral host-directed glutamine synthetase or glutamate dehydrogenase activity93. Prior literature suggests that the majority of nitrogen uptake is from symbiont-transferred metabolites resulting from their utilization of free ammonia in the water column94, although it has been suggested that the assimilation of nitrogen by the host itself is underestimated95. Here, there is evidence that the susceptible corals utilized urea as their primary nitrogen source at T1 (Fig. S4).
We propose that high levels of the urease enzyme may be a biomarker of a dysfunctional metabolic relationship between the coral host and its algal symbiont. Urea, a soluble nitrogen-rich molecule, is degraded intracellularly to yield ammonia and carbon dioxide (CO2) via urease enzyme (URE1, Figs. 2a, c and S3). At T1 in susceptible corals, GO terms for nickel-binding activity and carbonic anhydrase activity are enriched (Fig. 2a). Proteins associated with these functions provide the required nickel cofactor for urease96 and increased abundances of carbonic anhydrase enzyme (CA) rapidly convert CO2, byproducts of the reaction, into carbonic acid (or bicarbonate). It has been suggested that coral cells rely on this pathway to acquire additional nitrogen when stressed97,98. For example, in corals lacking Symbiodiniaceae, urease activity increased to compensate for the lack of nitrogen provided by Symbiodiniaceae98. Previous experiments on corals revealed that urea- and nickel-enrichments increased photosynthesis and calcification rates, suggesting that these molecules support coral growth in adverse environmental conditions96. Isocitrate dehydrogenase (ICDH1; FC: 0.57), an enzyme that increases under nitrogen starvation, is slightly increased in T1S (compared to T1R) corals. ICDH1 links the carbon metabolism (TCA cycle) and nitrogen cycle together to generate glutamate (via GS-GOGAT; Fig. S4a). The increased presence of enzymes involved in these alternate routes of nitrogen and carbon acquisition provides molecular evidence for their use as potential biomarkers of environmental stress and/or the beginnings of dysfunctional symbiosis.
Following thermal bleaching, the abundance of urease (URE1) in the susceptible corals continued to increase, resulting in a greater divergence in URE1 levels between susceptible and resilient corals at T2 compared to the pre-bleaching levels observed at T1timepoint (Fig. 3, cluster 9). Therefore, urea-dominated nitrogen acquisition strategy in the host increases as the host-symbiont relationship becomes compromised in susceptible corals responding to thermal stress. Although prokaryotic host-associated microbiome could provide bioavailable nitrogen via nitrogen fixation when the host is stressed, the 16S rRNA does not provide species-level resolution that would definitively reveal if any of the noted bacterial families in the T2S microbiome were nitrogen-fixers. No definitive evidence was revealed with the analysis of the 16S rRNA-predicted functional capacities of the microbiome (i.e., PICRUSt2 analysis; Fig. S6).
Importantly, the susceptible corals displayed early evidence for the rejection and degradation of symbiosomes before the onset of thermal stress. The coral’s symbionts reside in phagosomes called symbiosomes; corals therefore have specific enzymatic and signaling pathways to disrupt the standard phagosome recycling mechanisms, ensuring the symbiont’s residence. Typical phagosome recycling via hydrolytic enzymes is directed by Rab11 expression in coral hosts99. Established, healthy symbiotic relationships therefore inhibit the Rab11 pathway, resulting in a decrease in Rab11 abundance99. Susceptible corals at T1 displayed significantly increased abundance of Rab11 (Fig. 2c), an early indication of a dysfunctional symbiotic relationship and potential host-directed degradation of the symbiosome, or symbiophagy100. This host-symbiont disequilibrium hypothesis in T1S corals is further supported by increased abundance of alpha-tubulin (TUBA), a phagocytosis protein recognized to be active in symbiont degradation101, among other functions. Three enzymes detected at significantly increased abundances were involved in glycan degradation, in particular the mechanism involved in cleaving mannose-based oligosaccharides: alpha-L-fucosidase (FUCA1), mannose-receptor (MRC1), and mannosyl-oligosaccharide alpha-1,3-glucosidase (GANAB) (Fig. 3, cluster 8). As mannose is recognized by lectin proteins in corals to identify pathogens and symbionts, the degradation of these mannose-based oligosaccharides may weaken physical associations of the host with symbionts and its microbiome102,103. Further, the degradation of these oligosaccharides would specifically provide easily accessed glucose monomers for supporting the energy demands of the susceptible coral at T1. Increased presence of mucin proteins (i.e., MUC4, Fig. 2c), a noted deterrent to pathogen colonization104, and lectins, pathogen recognition proteins (e.g., TLEC2; Fig. 3, cluster 8) suggest that T1 susceptible corals may also be more challenged by pathogens, further weakening their immune system prior to the bleaching event. Further studies on how increased mucin expression influences pathogen response in corals should be completed to test this hypothesis. The T1S microbiome was less diverse than the T1R corals’ microbiomes and more variable across the entire susceptible cohort (i.e., each T1S coral had a different taxonomic composition). Quantification of the panel of proteins suggested here linked to symbiont rejection could be developed into a biomarker test providing coral managers with a rapid tool for identifying stressed corals that may not be suitable for propagation, even under optimal environmental conditions.
Divergent metabolic strategies in resilient and susceptible corals post-bleaching
Constitutive post-bleaching (T2) response across all M. capitata included many components of the phagocytic and endocytic pathways, indicating that active symbiotic expulsion105 during thermal stress-induced bleaching was occurring regardless of whether the corals were resilient or susceptible. In particular, Rab11, the inhibitor of symbiosome degradation observed in T1S, was detected at increased abundances in both coral groups at T2 relative to the T1 samples. Other constitutively expressed immune response proteins detected at higher abundances at T2 included NOD, MAPK, WNT, and TOLL-like receptors. All of these signaling pathways have been previously observed in corals106 and their detection suggests that during bleaching the innate immune system is activated. At T2 we detected increased abundances of the protein responsible for the irreversible step in gluconeogenesis in susceptible corals, and increased abundances of two irreversible steps in glycolysis in the resilient corals (Fig. S4b). This proteomic evidence suggests that after bleaching, the resilient corals have a more accessible glucose source, whereas the susceptible corals are catabolizing non-carbohydrate sources, such as lipids and proteins. These enzymatic pathway analyses also provide a molecular foundation for the observed 49% decrease in lipid biomass in the susceptible corals and the insignificant change in lipids in the resilient corals between pre- and post-bleaching (Fig. 4b). The finding that prior lipid stores (in support of the proteomic biosignatures noted) was not informative in predicting bleaching resilience was unexpected. This could be due to the relative similarity of lipid stores pre-bleaching in these particular corals, or that prior energy storage is unpredictive of coral survival, even though proteomic analyses suggest bleached corals rely on stored lipids for survival. In either case, further studies across different coral species and experimental conditions, particularly those with greater variation in pre-bleaching lipid stores, are necessary to determine whether this pattern is broadly applicable.
Resilient corals diversify metabolic pathway utilization to recover from thermal bleaching
After bleaching, several GO terms were enriched in resilient M. capitata proteome compared to T1: amino acid synthesis (methionine and proline), sulfur amino acid metabolic process, immune response, cell signaling/oxidoreductase, endoplasmic reticulum (ER) organization, oxoacid metabolism, and ribosome assembly (Fig. 2b). Sulfur amino acids, such as methionine, are antioxidants and therefore capable of providing oxidative protection to cells107. Increased levels of these amino acids in resilient corals could be indicative of an increased need for protection against oxidative stress resulting from the heat. Sulfurtransferase enzymes are present in both susceptible and resilient corals; however, they are significantly increased in resilient corals (Fig. 3, clusters 1 and 2). Heat stress has been found to also induce an increase in endoplasmic reticulum transcripts in Acropora hyacinthus108, mirroring our findings of increased abundance of ER proteins. Increases in cell signaling and ribosome assembly proteins are likely indicative of more normal cellular trafficking in healthy host tissue, enabling recovery from thermal bleaching. Several metabolic pathways are discussed below that support the resilient metabolism through thermal stress compared to the susceptible coral cohort.
Here, resilient corals activate endocytic uptake pathways and heterotrophic feeding after bleaching events to aid in nutritional recovery. Multiple enzymes involved in endocytosis are increased in the T2R cohort providing a heterotrophic avenue for carbon and nitrogen acquisition (Fig. 3, cluster 3). Increased abundance of two alpha-tubulin proteins (TUBA), vacuolar sorting endocytic protein (VPS4), dynamines and coatamers (COPG, COPB2) imply increased endocytic activity of particles, such as pathogens or food101,109. Significantly increased abundance of lysosome-associated membrane protein (LAMP) may indicate that symbiont engulfment and degradation is an additional potential source of nutrition for resilient corals post bleaching100. Significantly increased peptide degradation enzymes in the T2R cohort included glutamyl amino peptidase (ENPEP, (Fig. 3, cluster 3), which cleaves acidic amino acids from the N-terminus of peptides for subsequent degradation to enhance cellular growth. T2R also significantly increased a vitellogenic carboxypeptidase (CPVL), a protein involved in the degradation of yolk proteins. This could be a biomarker of decreased lipid provisioning to developing gametes in bleached resilient corals since this species only transfers autotrophic carbon to its gametes110,111,112. Abundant protease/peptidases (Fig. 3, cluster 3) and lipases (e.g., triacylglycerol lipase PNLIP; Fig. 2d) in resilient corals can break the bonds of macromolecular complexes to generate mobile small molecules that can be recycled or further degraded for energy. In support of these findings suggesting adequate nutritional resources were available in the resilient cohort post-bleaching, the increased abundance of transketolase (TKT) in T2R may indicate a higher abundance of thiamine (vitamin B1) compared to T2S113,114.
To further aid in recovery, resilient corals appear to utilize several new pathways to aid in cellular nitrogen and carbon demands post bleaching. Although the urea degrading enzyme URE1 is detected in both T2S and T2R corals, T2R displayed increased abundance of polyamine oxidase (MPAO; Fig. 3 cluster 3) which may allow resilient corals to access polyamine-nitrogen as needed and produce beta-alanine. Beta-alanine (aminopropagrainnoic acid) is a degradation product of the nucleotide uracil and is a precursor to acetyl-CoA. Notably, it has been found to increase cellular oxygen consumption and respiration rates115. Detecting multiple enzymes involved in these pathways to be at significantly higher abundance in the T2R corals provides a molecular explanation for hypothesized improved energy production compared to the T2S corals when symbiont derived photosynthate is diminished. Further, this energy may have provided resilient corals the ability to significantly increase multiple enzymes responsible for DNA transcription and translational processes (Fig. 3, cluster 3).
Proteomics analyses also suggest that T2R corals launched an antiviral campaign during thermal stress to assist the immune system. Cyanovirin protein (CVNH) was detected in resilient corals at T1 and T2 at three-fold higher abundance compared to susceptible corals (Fig. 3, cluster 1). Although we do not believe this is the only mode of protection for the resilient cohort, high production of this protein could increase the disease resilience of these corals after bleaching events when they are simultaneously coping with multiple stressors.
Catastrophic metabolic fate of T2S corals
GO term enrichment analysis revealed a greater abundance of proteins participating in peptide degradation and protein transport in susceptible M. capitata post-bleaching (Fig. 2b, d; Fig. 3 cluster 12). Since symbiont-derived photosynthate nutrition is absent in the bleached corals, susceptible corals may have increased mobilization and degradation of coral host proteins and peptides to provide the energy needed for cellular maintenance. Decreases in the free amino acids pool resulting from protein degradation in thermally bleached Acropora aspera suggests that these amino acids are being metabolically leveraged to provide energy during low photosynthate yield116.
Specifically, susceptible hosts post-bleaching express an abundance of enzymes that suggest host-directed catabolism of remaining symbionts. Several lysosomal-targeted peptidases/degradation enzymes were significantly increased in the susceptible corals in response to bleaching (e.g., cathepsin CATL, galactosidase GLB, and Niemann Pick C2 protein NPC2, Fig. 3, clusters 7, 12). The early signs of a weakened host-symbiont relationship for T1S corals (discussed earlier) appear to have progressed further in the susceptible cohort by T2. At T2, health-compromised/dead symbionts may be leaking organic substrates that are degraded by lysosomal and intracellular peptidases and hydrolase enzymes (Fig. 3, cluster 12). NPC2 enzymes are concentrated in the symbiosome and participate in the direct sterol transfer from symbionts117. Evidence of increased host-directed transfer of sterols from the symbiont combined with increased lysosomal-catabolic processes indicate that after bleaching, the symbiosome and its contents are targeted for rapid degradation in susceptible corals100.
Further, the decrease in symbiont-derived photosynthate in susceptible corals leads to activated gluconeogenesis and the degradation of glycine betaine. Glucose is one of the primary carbon molecules transferred to holobionts in cnidarian dinoflagellate symbiosis118,119. The increased abundance of the irreversible enzymes pyruvate carboxylase (PC) and phosphoenolpyruvate carboxykinase (PCK1) in T2 susceptible corals reveals that gluconeogenesis (i.e., the generation of glucose from pyruvate) is more active than glycolysis (i.e., the degradation of glucose; Fig. S4b). Increased abundance of these enzymes suggests that susceptible corals are more glucose-limited compared to the resilient corals after bleaching events. Gluconeogenesis depends on the catabolism of amino acids, glycine betaine, and lipids (Fig. 3, cluster 12). Lipid degradation is evidenced by the significant decrease in total lipid biomass between T1S and T2S corals (Fig. 4a, b). Only recently were glycine betaines recognized to be a major reservoir of nitrogen for corals and the near-complete glycine betaine catabolic and biosynthesis pathways have been uncovered in several genomes of cnidarians120. T2S corals increased abundance of the betaine-degrading enzymes betaine-homocysteine S-methyltransferase (BHMT), glycine N-methyltransferase (GNMT), and sarcosine dehydrogenase (SARDH). Ngugi et al. 120, suggest that glycine betaines are abundant nitrogen reservoirs that are easily degraded into other nitrogen compounds such as amino acids.
Susceptible coral proteomes post-bleaching also revealed a trend of potential decreased immune function at T2. Immune pathway results indicate that the NOD, MAPK, and TOLL-like signaling pathways are suppressed in T2S corals (p < 0.10; Supplementary Data 2h). The suppression or inactivation of these important immune pathways may make the corals vulnerable to disease and coral mortality. The suppressed beta diversity in T2S corals reveals a shift to a less diverse symbiotic bacterial community, which may be an indicator of the onset of infection121. Susceptible M. capitata corals also increased tyrosinase (TYR), an indicator of immune response to an infection122 or neutralization of reactive oxygen species123 demonstrating that T2S corals are being challenged.
Coral management and restoration applications
The impacts of climate change are accelerating, and there is a growing desire to harness mechanisms of thermal adaptation and acclimatization to increase the efficiency of reef restoration124. Biomarkers predictive of thermal resilience can provide another layer of information that practitioners can integrate into their restoration planning to help select corals for propagation and outplanting124. We identified two key biological factors that distinguish resilient from susceptible corals before the onset of thermal bleaching that forecast their long-term health: protein abundances (Fig. 2a, c) and microbiome diversity (Fig. 5a–d). Here, we focus on the proteomic results as we have no conclusive evidence that links a particular microbial taxa to bleaching tolerance, although recent research suggests that the microbiome may influence thermotolerance125. As a high-throughput technology, proteomic biomarkers can be tailored to provide informative results on the molecular-level health of M. capitata across a range of resolutions. We have identified seven proteins that could be quantified in corals before bleaching events as a resilience-based assay to select corals for propagation or other management approaches (Fig. 6). Proteins identified to be potential biomarkers for resilience include a skeletal precipitation-related protein126, ubiquinone oxidoreductase activity, a lipoxygenase protein involved in immune function and pest resistance127. Two subunits of V-type proton ATPase were identified as potential biomarkers and participate in the carbon-concentrating mechanism that regulates the photosynthetic rate of their symbiotic algae128. If using mass spectrometry, quantifying five peptides through the detection of five or more diagnostic fragment ions from each of these proteins would provide the user with high confidence on both positive and negative signals of pre-bleaching resilience. This short list could also be expanded to generate a 60-min assay with up to 250 peptides that are simultaneously monitored, providing further information on heterotrophic feeding/lipid degradation (i.e., LIP, PSAP), antiviral activity (i.e., CNVH), symbiophagy (i.e., RAB11, TUBA), pathogen recognition (i.e., TLEC), mucin proteins (i.e., MUC4), urea degradation (i.e.,URE1), and mannose degradation (i.e., FUCA1, MRC1, GANAB). Alternatively, selected proteins identified here as resilience biomarkers could be developed into a hand-held rapid antigen test with multiple test and control lines that could be assessed in the field on rice-grain size coral tissue samples. Future, more targeted, analyses need to be conducted on a larger coral cohort to verify the applicability of these biomarkers.
Intensity of pink color depicts (Normalized Spectral Abundance Factor) NSAF values for each of the 12 coral genets analyzed, normalized by row mean. Clustered dendrograms were completed with the correlation algorithm on the x and y axis to generate groups for significantly increased or decreased abundances in the resilient vs. susceptible cohorts.
Concluding Remarks
Using mass spectrometry-based proteomics, 16S rRNA sequencing of the microbiome, total lipids, and symbiont density and community composition, we identify intrinsic differences predictive of and likely involved in recovery and survival of corals to thermal bleaching events. Resilient corals have a significantly higher abundance of antiviral proteins and express multiple enzymes involved in a diverse range of carbon and nitrogen acquisition pathways such as lipid degradation, heterotrophic feeding, and respiration. Conversely, corals that did not survive thermal bleaching had elevated abundances of pre-bleaching molecular markers that play an active role in symbiont rejection, pathogen recognition, and mannose and urea degradation. These differences were evident despite identical environmental histories, as experimental coral were sourced from the same reef site, suggesting that these functional differences likely reflect fixed genetic effects. In addition to characterizing the physiological mechanisms underpinning resilience we have identified a set of candidate proteomic biomarkers which can confidently identify resilient and susceptible coral prior to stress events. The proteins represented in each of these pathways and cellular mechanisms can be fully developed into rapid molecular assays to help assess corals and guide the development of mitigation strategies for reef management.
To enhance the robustness of future studies and better account for biological variation, we recommend collecting additional Montipora capitata coral samples in situ prior to the next natural heat-induced thermal stress event. Increasing the sample size and including more sites with a broader range of M. capitata genotypes would help control for biological variability and increase confidence in identifying biosignatures. These efforts should incorporate proteomics, lipidomics, symbiont density measurements, microbiome analyses, and pathogen susceptibility assays, along with metabolomics across different coral sites and species, to provide a comprehensive understanding of coral physiological states. Additionally, monitoring coral resilience or susceptibility after thermal bleaching is essential for validating proposed biosignatures. While we acknowledge the logistical challenges of such studies, advancing these efforts is crucial for identifying reliable resilience markers and informing effective coral management strategies.
Methods
Coral collection and experiment
Seventy-four Montipora capitata (~30 cm in diameter) were collected from at 1.5–2 m depth Moku O Lo’e island (patch reef) located in Kāne’ohe Bay, O’ahu, Hawai’i (21.428°N, 157.792°W) in August 2017. Corals were brought to shore and acclimated in flow-through outdoor tanks at the Hawaii Institute of Marine Biology (HIMB) for two weeks at ambient conditions (27 ± 0.4 °C, max = 28.1, min = 25.7). At the time of collection, corals were divided into two pieces to compare physiological performance for the same genets with/without exposure to thermal stress (Fig. 1a, b). In September, one half of each coral was exposed to warmer water temperatures to simulate a natural thermal bleaching event70. To reach the 30 °C temperature target for the bleaching treatment, experimental tank temperatures were increased by ~0.5 °C per day for four days using a 600 W Titanium Aquarium Heater (Bulk Reef Supply, United States). Throughout the bleaching experiment, the control tanks (n = 3) had a temperature of 28 ± 0.6 °C (range: 26.5–30.7 °C), and the high-temperature tanks (n = 3) had a temperature of 30 ± 0.5 °C (range: 28.9–31 °C)(Fig. S1a).
Corals were rotated once a week among the same treatment tanks to minimize tank effects. For the bleaching experiment, M. capitata were kept at this elevated temperature for three weeks to induce bleaching in all corals. After three weeks, the tank temperature was lowered, following the previously described rate, back to ambient temperature (27–28 °C) and another set of tissue subsamples were taken (temperature data: Supplementary Data 1b). The control experiment coral halves (T2NB) not exposed to thermal stress remained at 28 °C and were rotated each week among tanks of the same treatment to minimize tank effects52 and collected at the same time as the bleaching experiment (details below). Mean daytime photosynthetic active radiation (PAR) levels in the tanks were 584 µmol photons m−2 s−1, and mean PAR at 12:00 was 1249 µmol photons m−2 s−1, measured using a waterproof PAR logger (Odyssey, Dataflow Systems Limited, NZ; Fig. S1b). These irradiance values were very similar to values in the field where the coral colonies were located129. For more information on the experimental setup see refs. 70,130. After thermal exposure, all corals (bleached and not bleached) were placed on racks at 1.5–2 m depth off HIMB (where they were originally collected from) to monitor survival and physiological recovery in situ for eight months.
Bleaching assessments were conducted on all corals just prior to the start of the elevated temperature conditions, then every week for the duration of the bleaching experiment using the Coral Watch Card (The University of Queensland, Australia), along with assessments of mortality (see Fig. S1c). Corals that bleached and recovered were deemed part of the resilient cohort, while corals that bleached and died were deemed susceptible to bleaching. All resulting proteomic search results, protein accession numbers and annotation files, lipid data, symbiont density and clade data, chlorophyll data and R code for plot generation and data analysis have been deposited in GitHub (https://github.com/Nunn-Lab/Publication-coral-resilience).
Branches from twelve (six resilient and six susceptible) M. capitata coral genets were collected at two timepoints: (1) on August 30, 2017, after temperature acclimation in the tanks but before corals were bleached (T1) and (2) on October 1, 2017, 24 h after bleached corals were gradually returned to ambient temperature (T2). All coral samples were collected 1 cm from the tip of a branch and snap-frozen immediately in liquid nitrogen. Frozen samples were shipped to the University of Washington on dry ice and stored at −80 °C. Samples for protein extraction consisted of 2 mm thin cross-sections of the branches, encompassing both tissue and skeletal matrix.
Symbiont and chlorophyll analyses
Chlorophyll a concentrations and dinoflagellate symbiont (Symbiodiniaceae) densities from each of the coral genets were investigated, both using the methods and standardizations reported in Rodrigues and Grottoli64 (Supplementary Data 1a). Briefly, chlorophyll a was extracted with 100% acetone, and its absorbance was measured using a light spectrophotometer set to 663 nm (Table S1). Triplicate symbiont samples were taken from a single sample of ground coral tissue and homogenized prior to being counted using a hemocytometer. Chlorophyll a and symbiont densities were standardized to grams of ash-free dry weight (gdw) of coral tissue (Supplementary Data 1a, c). M. capitata in Kāne’ohe Bay forms symbioses with Symbiodiniaceae in the genera Durusdinium and Cladocopium110,131, and the identity of the dominant symbiont type can influence bleaching susceptibility72. To assess relative abundances of Cladocopium and Durusdinium in experimental corals, total DNA was extracted from a 4 mm piece of frozen M. capitata using a CTAB-chloroform protocol (https://doi.org/10.17504/protocols.io.dyq7vv). Relative abundances of Durusdinium and Cladocopium were quantified via qPCR using actin assays72,132 on an Agilent AriaMX system with two technical replicates and reference dye corrections run for 40 cycles. CT values were corrected for fluorescence and copy number, and Cladocopium:Durusdinium spp ratios were calculated following Innis et al. 133. CT values greater than 37 were taken to indicate that presence of the amplified sequence was below detection level for these assays134. Symbiont community composition was classified as Durusdinium-dominated or Cladocopium-dominated if only one clade was present, or as a mixed community if both Cladocopium and Durusdinium spp. were present in any proportion, based on the symbiont community composition in samples collected at T1 or from the non-bleached controls at T2. Fisher’s exact test was used to confirm that symbiont community type and resilience/susceptibility were independent. A logistic regression analysis was performed to evaluate the relationship between the proportion of Durusdinium symbionts pre-bleaching and coral mortality post-bleaching (Supplementary Data 1b). Further details found in SI Methods.
Lipid analyses
Total lipids were analyzed on each sample following the methods of Rodrigues and Grottoli64. Briefly, whole fragments (tissue plus skeleton) were crushed and digested in a 2:1 chloroform:methanol solution, sequentially washed in a 0.88% KCl solution, dried under grade 5.0 N2 gas to a constant weight and standardized to ash-free gdw of coral tissue (Supplementary Data 1d, e). A two-way ANOVA was performed to explore the single and interactive effects of bleaching status and tolerance to bleaching to better understand if total lipid content influenced coral mortality post-bleaching.
Proteomics
Six coral genets from the resilient cohort and six corals from the susceptible cohort, were randomly selected as bioreplicates to track phenotypic differences in protein abundance through time (i.e., T1 and T2). Details can be found in SI Methods. Briefly, proteins were extracted from whole coral fragments (4 mm diameter × 1 mm thick, tissue plus skeletal matrix) and resulting protein concentrations were determined with bicinchoninic acid (BCA) protein microplate assay. Protein lysates (50 µg per coral sample) were reduced, alkylated, and digested with Trypsin (modified porcine sequencing grade trypsin; Promega; 1:20 enzyme:coral protein). Each digested peptide sample was amended with Peptide Retention Time Calibration Mixture (PRTC; Pierce) such that 50 fmol of PRTC was analyzed with 1 µg of coral peptides for each mass spectrometry experiment.
M. capitata samples were analyzed using liquid chromatography coupled to tandem mass spectrometry (LC‒MS/MS) on a Q‒Exactive‒HF (Thermo Scientific) in Data Dependent Acquisition (DDA) Top 20 mode. Samples were separated using a heated (50 oC) 40 cm long analytical column packed with C18 beads (Dr. Maisch HPLC, Germany, 0.3 µm, 120 Å). Peptides were chromatographically separated on a Waters nanoAcquity UPLC using an acidified (0.01% formic acid) acetonitrile:water gradient of 2–45% over 120 min. Internal and external standards were monitored to ensure peptide peak area correlation variances were <10% through the duration of the analyses. Data were searched against a translated M. capitata transcriptome135 GSE97888_Montiporacapitata_transcriptome.fasta; Supplementary Data 6). Protein identifications from the whole-cell lysates are reported if two or more peptides were identified, at least one terminus was tryptic, and the false discovery rate <0.01 (Supplementary Data 2a–e). Differential relative protein abundances for resilient vs. susceptible corals and corresponding p-values were determined for each cohort (T1 and T2) using the QPROT-QSPEC package136(Supplementary Data 2g, h). Differential abundances of proteins are reported with the following p-value cutoff rules: (1) p < 0.10 if several proteins within a pathway, (2) p < 0.05 if significance of an individual protein, or (3) p < 0.01 if identifying a potential biomarker. A second analysis of the mass spectrometry data was performed following the same search parameters as a test to determine if known coral viral proteins could be detected from the whole-coral mass spectrometry analyses (details in Supplementary Information). The database searched consisted of the same translated M. capitata transcriptome135, common contaminants, and 5 viral proteomes (Supplementary Data 7).
MetaGOmics biological enrichment analysis
To determine if categories of proteins were enriched in the resilient vs. susceptible coral cohorts at the two timepoints, a biological enrichment strategy that analyzes Gene Ontology (GO) categorical terms was used to compare sets of detected proteins62. Top results are reported with a cutoff E-value < 1E-10. A fasta file of all M. capitata protein sequences confidently identified in these experiments (Supplementary Data 8) was analyzed with MetaGOmics v.0.1.1. Although MetaGOmics was designed to analyze microbiomes, the use of the software was modified to work with a single organism by ignoring the taxonomic enrichment analysis to instead examine functions that are significantly enriched or depleted in pairwise comparisons of coral cohorts. Additional details found in SI Methods.
Microbiome 16S rRNA Sequencing
Total DNA was extracted from the same subset of flash frozen coral fragment selected for the proteomic study (n = 12) using the Qiagen DNA extraction kit. Sample purity was measured using a NanoDrop ND1000 spectrophotometer (NanoDrop Technologies, Willmington DE). Samples showing good quality and concentrations were sent to Molecular Research LP (Shallowater, TX) for library prep and 16S rRNA amplicon sequencing. Library prep and sequencing was performed following details outlined in Brown et al.137. at Mr. DNA Molecular Research. Briefly, 16S rRNA amplicons were amplified using Caporaso et al.138. 515 f (5′-GTGCCAGCMGCCGCGGTAA-3′) 806r (5′-GGACTACHVGGGTWTCTAAT-3′) primers. Amplification conditions were a single step 30-cycle PCR using HotStarTaq plus Master Mix (Qiagen) at the following temperatures: 94 °C for 30 s, 53 °C for 40 s, and 72 °C for 1 min, followed by a final incubation step of 72 °C for 1 min after all cycles finished. Sequencing was performed on an Ion Torrent PGM. All 16S rRNA gene amplicon sequence data, processing steps and code for quality control on the microbiome data and analysis are available on GitHub (https://github.com/tanyabrown9/Resilient_vs_Susceptible_Mcapitata). Sequences are deposited in NCBI as bioproject PRJNA933787. Initial sequencing resulted in a collection of 2,472,819 total reads, with an average read depth of 68,689 (±28,107 SD) sequences per sample (Supplementary Data 4a, b).
Microbiome 16S rRNA data preparation and quality control
16S rRNA gene amplicon sequence data from Molecular Research LP were processed using the QIIME2 software package139. Forward and reverse reads from Molecular Research LP were imported into QIIME2 and demultiplexed (using the QIIME paired-end sequencing method). Sequences were then denoised using DADA2 and chimaera-checked (using the QIIME dada2 denoise-pyro method) to generate amplicon sequence variants (ASVs)140. Care was taken to identify mitochondria and chloroplasts in silico prior to downstream analysis as previously described137. Once annotated, chloroplast and mitochondria sequences were removed. Samples were rarefied to 10,000 sequences per sample to protect against false-positive results attributable to unequal sequencing depth between samples141.
Microbiome alpha and beta diversity analysis
Alpha and beta139 diversities were assessed on rarefied samples after the quality control steps. To conduct phylogenetic alpha and beta diversity metrics, a fragment insertion approach was used to insert short microbial 16S rRNA reads into a reference phylogeny. Sequences were placed on the QIIME2-compatible SILVA 128 tree included with the SEPP software using the QIIME fragment-insertion SEPP method142,143.
Alpha diversity was assessed on samples rarefied to 1000 sequences per sample, using multiple metrics: the number of unique observed ASVs in rarefied samples (observed features), Faith’s Phylogenetic Diversity, Simpson’s Evenness, and Shannon’s Diversity Indexes. Overall differences in alpha diversity across susceptibility status and timepoints were tested using Kruskal-Wallis tests. False Discovery Rate (FDR) for pairwise tests between susceptibility and time was controlled using FDR q-values reported by QIIME2. Beta diversity between samples was assessed using Weighted UniFrac distances and Bray-Curtis dissimilarities. The significance of differences in beta-diversity between susceptibility vs. resilient corals and across timepoints was tested using PERMANOVA144.
Microbiome differential abundance analysis
Following methods outlined in Anderson et al.144, the top 10 bacterial families in each sample type were selected for taxonomic analysis. Multivariate Association with Linear Models were performed on the 16S datasets using the R package MaAsLin249. Fixed effects for the analysis included bleaching status, resilience, timepoint, and whether the colony survived or succumbed to thermal stress (bleaching) by T2. Coral colony term was set as a random effect and the analysis method was “LM”. Results were considered significant if the q-value was <0.0565.
Microbial sequence data quality analysis and processing
All other statistical analyses of microbial taxonomy were conducted using R 4.0.3. The top 10 bacterial families in each sample type were selected for taxonomic analysis. Significant differences between bacterial families, susceptibility, and timepoints were carried out using a nested ANOVA. The Benjamini-Hochberg FDR was used to control the false discovery rate. Microbiome Multivariate Association with Linear Models was performed on the 16S rRNA sequencing data using the R package MaAsLin265. Fixed effects for the analysis included bleaching status, resilience, timepoint, and whether the colony survived or succumbed to thermal stress (bleaching) by T2. Coral colony term was set as a random effect and the analysis method was “LM”. Results were considered significant if the q-value was <0.05.
Microbiome functional prediction analysis
Predicted functional profiles for MetaCyc categories were generated from sequencing data using PICRUSt266. This method estimates gene content or other functional parameters based on phylogenetic modeling of 16S rRNA sequences relative to sequenced reference genomes. SETé enabled phylogenetic placement (SEPP145) was used for phylogenetic placement (--p-placement_tool sepp), while phylogenetic independent contrasts (PIC) was used for phylogenetic inference (--p-hsp-method pic). Predicted profiles of MetaCyc functional annotations at T1 were compared with ANCOM-BC146 to identify differentially abundant functional categories (Fig. S6, Supplementary Data 5).
Data availability
All raw MS proteomic data and protein FASTA files used for searching can be accessed at PRIDE accession PXD021262, a public proteomic data repository. All resulting proteomic search results, protein accession numbers and annotation files, lipid data, symbiont density and clade data, chlorophyll data can be found in GitHub https://github.com/Nunn-Lab/Publication-coral-resilience(https://github.com/Nunn-Lab/Publication-coral-resilience). All study data are included in the article and/or as Supplementary Data. All transcriptome and protein sequences used in this study are also publicly available through NCBI GenBank and ProteomeXchange PRIDE. Protein/gene accession numbers and annotations are provided as supplementary information (Supplementary Data 2) and all supplemental data can be found in Figshare project 237383. All 16S rRNA gene amplicon sequence data and PICRUSt2 functional predictions presented from the microbiome data and analysis are available on GitHub (https://github.com/tanyabrown9/Resilient_vs_Susceptible_Mcapitata).
Code availability
All R code required with file inputs for proteomic, lipid, and symbiont plot generation and data analysis have been deposited in GitHub https://github.com/Nunn-Lab/Publication-coral-resilience(https://github.com/Nunn-Lab/Publication-coral-resilience). No custom code was generated for this manuscript or analysis. The transcriptome was translated using Transdecoder v 2.0.1. and Comet 2022.01 was used to search the DDA mass spectrometry files. MetaGOmics software (version 0.2.0; https://www.yeastrc.org/metagomics) was used to identify the lowest common ancestor on each peptide’s functional Gene Ontology (GO) assignment. All code related to the analysis of the coral microbiome 16S rRNA gene amplicon sequence data and PICRUSt2 functional predictions are available on GitHub (https://github.com/tanyabrown9/Resilient_vs_Susceptible_Mcapitata).
References
Mumby, P. J. et al. Coral reef habitats as surrogates of species, ecological functions, and ecosystem services. Conserv. Biol. 22, 941–951 (2008).
Salvat, B. Coral reefs—a challenging ecosystem for human societies. Glob. Environ. change 2, 12–18 (1992).
Hughes, T. P. et al. Climate change, human impacts, and the resilience of coral reefs. Science 301, 929–933 (2003).
Pandolfi, J. M. et al. Global trajectories of the long-term decline of coral reef ecosystems. Science 301, 955–958 (2003).
Hoegh-Guldberg, O. et al. Coral reefs under rapid climate change and ocean acidification. Science 318, 1737–1742 (2007).
De’ath, G., Lough, J. M. & Fabricius, K. E. Declining coral calcification on the Great Barrier Reef. Science 323, 116–119 (2009).
Hughes, T. P. et al. Global warming and recurrent mass bleaching of corals. Nature 543, 373–377 (2017).
Reaser, J. K., Pomerance, R. & Thomas, P. O. Coral bleaching and global climate change: scientific findings and policy recommendations. Conserv. Biol. 14, 1500–1511 (2000).
Wellington, G. M. et al. Crisis on coral reefs linked to climate change. Eos Trans. Am. Geophys. Union 82, 1–5 (2001).
Muscatine, L., Falkowski, P., Porter, J. & Dubinsky, Z. Fate of photosynthetic fixed carbon in light-and shade-adapted colonies of the symbiotic coral Stylophora pistillata. Proc. R. Soc. Lond. Ser. B. Biol. Sci. 222, 181–202 (1984).
Perez, S. F., Cook, C. B. & Brooks, W. R. The role of symbiotic dinoflagellates in the temperature-induced bleaching response of the subtropical sea anemone Aiptasia pallida. J. Exp. Mar. Biol. Ecol. 256, 1–14 (2001).
Szmant, A. & Gassman, N. The effects of prolonged “bleaching” on the tissue biomass and reproduction of the reef coral Montastrea annularis. Coral Reefs 8, 217–224 (1990).
Baird, A. & Marshall, P. Mortality, growth and reproduction in scleractinian corals following bleaching on the Great Barrier Reef. Mar. Ecol. Prog. Ser. 237, 133–141 (2002).
Hoegh-Guldberg, O. Climate change, coral bleaching and the future of the world’s coral reefs. Mar. Freshwater Res.50, 839–866 (1999).
Helgoe, J., Davy, S. K., Weis, V. M. & Rodriguez-Lanetty, M. Triggers, cascades, and endpoints: connecting the dots of coral bleaching mechanisms. Biol. Rev. 99, 715–752 (2024).
Boilard, A. et al. Defining coral bleaching as a microbial dysbiosis within the coral holobiont. Microorganisms 8, 1682 (2020).
Grottoli, A. G. et al. Coral physiology and microbiome dynamics under combined warming and ocean acidification. PloS ONE 13, e0191156 (2018).
Bourne, D., Iida, Y., Uthicke, S. & Smith-Keune, C. Changes in coral-associated microbial communities during a bleaching event. ISME J. 2, 350–363 (2008).
Lima, L. F. et al. Modeling of the coral microbiome: the influence of temperature and microbial network. Mbio 11, e02691–02619 (2020).
Dunphy, C. M., Gouhier, T. C., Chu, N. D. & Vollmer, S. V. Structure and stability of the coral microbiome in space and time. Sci. Rep.-Uk 9, 1–13 (2019).
Chu, N. D. & Vollmer, S. V. Caribbean corals house shared and host-specific microbial symbionts over time and space. Environ. Microbiol. Rep. 8, 493–500 (2016).
Hester, E. R., Barott, K. L., Nulton, J., Vermeij, M. J. & Rohwer, F. L. Stable and sporadic symbiotic communities of coral and algal holobionts. ISME J. 10, 1157–1169 (2016).
Huggett, M. J. & Apprill, A. Coral microbiome database: integration of sequences reveals high diversity and relatedness of coral-associated microbes. Environ. Microbiol. Rep. 11, 372–385 (2019).
Peixoto, R. S., Rosado, P. M., Leite, D. Cd. A., Rosado, A. S. & Bourne, D. G. Beneficial microorganisms for corals (BMC): proposed mechanisms for coral health and resilience. Front. Microbiol. 8, 341 (2017).
Krediet, C. J., Ritchie, K. B., Paul, V. J. & Teplitski, M. Coral-associated micro-organisms and their roles in promoting coral health and thwarting diseases. Proc. R. Soc. B: Biol. Sci. 280, 20122328 (2013).
van de Water, J. A., Allemand, D. & Ferrier-Pagès, C. Host-microbe interactions in octocoral holobionts-recent advances and perspectives. Microbiome 6, 1–28 (2018).
Beatty, D. S. et al. Variable effects of local management on coral defenses against a thermally regulated bleaching pathogen. Sci. Adv. 5, eaay1048 (2019).
Costa, R. M. et al. Surface topography, bacterial carrying capacity, and the prospect of microbiome manipulation in the sea anemone coral model Aiptasia. Front. Microbiol. 12, 637834 (2021).
Zaneveld, J. R. et al. Overfishing and nutrient pollution interact with temperature to disrupt coral reefs down to microbial scales. Nat. Commun. 7, 1–12 (2016).
Mayer, B., Rixen, T. & Pohlmann, T. The spatial and temporal variability of air-sea CO2 fluxes and the effect of net coral reef calcification in the Indonesian Seas: A numerical sensitivity study. Front. Mar. Sci. 5, 116 (2018).
Morrow, K., Muller, E. & Lesser, M. Coral bleaching 153-188 (Springer, 2018).
Baker, A. C., Glynn, P. W. & Riegl, B. Climate change and coral reef bleaching: an ecological assessment of long-term impacts, recovery trends and future outlook. Estuar. Coast. shelf Sci. 80, 435–471 (2008).
Weis, V. M. Cellular mechanisms of Cnidarian bleaching: stress causes the collapse of symbiosis. J. Exp. Biol. 211, 3059–3066 (2008).
Thornhill, D. J., LaJeunesse, T. C., Kemp, D. W., Fitt, W. K. & Schmidt, G. W. Multi-year, seasonal genotypic surveys of coral-algal symbioses reveal prevalent stability or post-bleaching reversion. Mar. Biol. 148, 711–722 (2006).
Toller, W. W., Rowan, R. & Knowlton, N. Repopulation of zooxanthellae in the Caribbean corals Montastraea annularis and M. faveolata following experimental and disease-associated bleaching. Biol. Bull. 201, 360–373 (2001).
Jones, R. J. & Yellowlees, D. Regulation and control of intracellular algae (= zooxanthellae) in hard corals. Philos. Trans. R. Soc. Lond. Ser. B: Biol. Sci. 352, 457–468 (1997).
Jokiel, P. L. & Coles, S. Effects of temperature on the mortality and growth of Hawaiian reef corals. Mar. Biol. 43, 201–208 (1977).
Grottoli, A. G. et al. The cumulative impact of annual coral bleaching can turn some coral species winners into losers. Glob. Change Biol. 20, 3823–3833 (2014).
Salm, R. V. Coral reef resilience and resistance to bleaching(IUCN, 2005).
Berkelmans, R. & Van Oppen, M. J. The role of zooxanthellae in the thermal tolerance of corals: a ‘nugget of hope’for coral reefs in an era of climate change. Proc. R. Soc. B: Biol. Sci. 273, 2305–2312 (2006).
Silverstein, R. N., Cunning, R. & Baker, A. C. Change in algal symbiont communities after bleaching, not prior heat exposure, increases heat tolerance of reef corals. Glob. Change Biol. 21, 236–249 (2015).
Baker, A. C. Coral health and disease 177-194 (Springer, 2004).
Abrego, D., Ulstrup, K. E., Willis, B. L. & van Oppen, M. J. Species–specific interactions between algal endosymbionts and coral hosts define their bleaching response to heat and light stress. Proc. R. Soc. B: Biol. Sci. 275, 2273–2282 (2008).
Baird, A. H., Bhagooli, R., Ralph, P. J. & Takahashi, S. Coral bleaching: the role of the host. Trends Ecol. Evol. 24, 16–20 (2009).
Henley, E. M. et al. Growth and survival among Hawaiian corals outplanted from tanks to an ocean nursery are driven by individual genotype and species differences rather than preconditioning to thermal stress. PeerJ 10, e13112 (2022).
Drury, C. Resilience in reef-building corals: the ecological and evolutionary importance of the host response to thermal stress. Mol. Ecol. 29, 448–465 (2020).
Barshis, D. J. et al. Genomic basis for coral resilience to climate change. Proc. Natl. Acad. Sci. USA 110, 1387–1392 (2013).
Bay, R. A. & Palumbi, S. R. Multilocus adaptation associated with heat resistance in reef-building corals. Curr. Biol. 24, 2952–2956 (2014).
Weis, V. M. The susceptibility and resilience of corals to thermal stress: adaptation, acclimatization or both? Mol. Ecol. 19, 1515–1517 (2010).
Pootakham, W. et al. Heat-induced shift in coral microbiome reveals several members of the Rhodobacteraceae family as indicator species for thermal stress in Porites lutea. Microbiologyopen 8, e935 (2019).
Ziegler, M., Seneca, F. O., Yum, L. K., Palumbi, S. R. & Voolstra, C. R. Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat. Commun. 8, 14213 (2017).
Petrou, K., Nunn, B., Padula, M., Miller, D. & Nielsen, D. Broad scale proteomic analysis of heat-destabilised symbiosis in the hard coral Acropora millepora. Sci. Rep.11, 1–16 (2021).
Morris, L. A., Voolstra, C. R., Quigley, K. M., Bourne, D. G. & Bay, L. K. Nutrient availability and metabolism affect the stability of coral–Symbiodiniaceae symbioses. Trends Microbiol. 27, 678–689 (2019).
Bahr, K. D., Jokiel, P. L. & Toonen, R. J. The unnatural history of Kāne ‘ohe Bay: coral reef resilience in the face of centuries of anthropogenic impacts. PeerJ 3, e950 (2015).
Eakin, C. M., Sweatman, H. P. & Brainard, R. E. The 2014–2017 global-scale coral bleaching event: insights and impacts. Coral Reefs 38, 539–545 (2019).
Ritson-Williams, R. & Gates, R. D. Coral community resilience to successive years of bleaching in Kāne ‘ohe Bay, Hawai ‘i. Coral Reefs 39, 757–769 (2020).
Bahr, K. D., Rodgers, Ku. S. & Jokiel, P. L. Impact of three bleaching events on the reef resiliency of Kāne ‘ohe Bay, Hawai ‘i. Front. Mar. Sci. 4, 398 (2017).
Hughes, T. P. et al. Global warming transforms coral reef assemblages. Nature 556, 492–496 (2018).
Coles, S. L., Jokiel, P. L. & Lewis, C. R. Thermal tolerance in tropical versus subtropical Pacific reef corals. Pac. Sci. 30, 159–166 (1976).
Jokiel, P. L. & Brown, E. K. Global warming, regional trends and inshore environmental conditions influence coral bleaching in Hawaii. Glob. Change Biol. 10, 1627–1641 (2004).
Gibbin, E. M., Putnam, H. M., Gates, R. D., Nitschke, M. R. & Davy, S. K. Species-specific differences in thermal tolerance may define susceptibility to intracellular acidosis in reef corals. Mar. Biol. 162, 717–723 (2015).
Riffle, M. et al. MetaGOmics: A Web-Based Tool for Peptide-Centric Functional and Taxonomic Analysis of Metaproteomics Data. Proteomes 6, https://doi.org/10.3390/proteomes6010002 (2017).
Botos, I. & Wlodawer, A. Cyanovirin-N: a sugar-binding antiviral protein with a new twist. Cell. Mol. Life Sci. CMLS 60, 277–287 (2003).
Rodrigues, L. J. & Grottoli, A. G. Energy reserves and metabolism as indicators of coral recovery from bleaching. Limnol. Oceanogr. Lett. 52, 1874–1882 (2007).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
Ochsenkühn, M. A. et al. Endozoicomonas provides corals with steroid hormones during thermal stress. bioRxiv, 2023.2009. 2019.558257 (2023).
Ricci, C. A. et al. Proteomic investigation of a diseased gorgonian coral indicates disruption of essential cell function and investment in inflammatory and other immune processes. Int. Comp. Bio. 59, 830–844 (2019).
Weston, A. J. et al. Proteomics links the redox state to calcium signaling during bleaching of the scleractinian coral acropora microphthalma on exposure to high solar irradiance and thermal stress [S]. Mol. Cell. Proteom. 14, 585–595 (2015).
Axworthy, J. B. et al. Shotgun proteomics identifies active metabolic pathways in bleached coral tissue and intraskeletal compartments. Front. Mar. Sci. 9, 50 (2022).
Tisthammer, K. H., Timmins-Schiffman, E., Seneca, F. O., Nunn, B. L. & Richmond, R. H. Physiological and molecular responses of lobe coral indicate nearshore adaptations to anthropogenic stressors. Sci. Rep. 11, 3423 (2021).
Cunning, R., Ritson-Williams, R. & Gates, R. D. Patterns of bleaching and recovery of Montipora capitata in Kāne ‘ohe Bay, Hawai ‘i, USA. Mar. Ecol. Prog. Ser. 551, 131–139 (2016).
Nunn, B. L. et al. Diatom proteomics reveals unique acclimation strategies to mitigate Fe limitation. PLoS ONE 8, e75653 (2013).
Aertsen, A. & Michiels, C. W. Diversify or die: generation of diversity in response to stress. Crit. Rev. Microbiol. 31, 69–78 (2005).
Yan, S.-P., Zhang, Q.-Y., Tang, Z.-C., Su, W.-A. & Sun, W.-N. Comparative proteomic analysis provides new insights into chilling stress responses in rice. Mol. Cell. Proteom. 5, 484–496 (2006).
Palardy, J. E., Rodrigues, L. J. & Grottoli, A. G. The importance of zooplankton to the daily metabolic carbon requirements of healthy and bleached corals at two depths. J. Exp. Mar. Biol. Ecol. 367, 180–188 (2008).
Lawe, D. C., Patki, V., Heller-Harrison, R., Lambright, D. & Corvera, S. The FYVE domain of early endosome antigen 1 is required for both phosphatidylinositol 3-phosphate and Rab5 binding: critical role of this dual interaction for endosomal localization. J. Biol. Chem. 275, 3699–3705 (2000).
Mohamed, A. R. et al. Deciphering the nature of the coral–Chromera association. ISME J. 12, 776–790 (2018).
Koul, A., Herget, T., Klebl, B. & Ullrich, A. Interplay between mycobacteria and host signalling pathways. Nat. Rev. Microbiol. 2, 189–202 (2004).
Grottoli, A. G., Rodrigues, L. J. & Palardy, J. E. Heterotrophic plasticity and resilience in bleached corals. Nature 440, 1186–1189 (2006).
Swart, P. K. Carbon and oxygen isotope fractionation in scleractinian corals: a review. Earth-Sci. Rev. 19, 51–80 (1983).
Yano, T., Yoshida, N., Yu, F., Wakamatsu, M. & Takagi, H. The glyoxylate shunt is essential for CO2-requiring oligotrophic growth of Rhodococcus erythropolis N9T-4. Appl. Microbiol. Biotechnol. 99, 5627–5637 (2015).
Seibel, B. A. & Drazen, J. C. The rate of metabolism in marine animals: environmental constraints, ecological demands and energetic opportunities. Philos. Trans. R. Soc. B: Biol. Sci. 362, 2061–2078 (2007).
van Oppen, M. J. & Blackall, L. L. Coral microbiome dynamics, functions and design in a changing world. Nat. Rev. Microbiol. 17, 557–567 (2019).
Mohamed, A. R., Ochsenkühn, M. A., Kazlak, A. M., Moustafa, A. & Amin, S. A. The coral microbiome: towards an understanding of the molecular mechanisms of coral–microbiota interactions. FEMS Microbiol. Rev. 47, fuad005 (2023).
Arriaga-Piñón, Z. P. et al. Microbiomes of three coral species in the Mexican Caribbean and their shifts associated with the Stony Coral Tissue Loss Disease. Plos ONE 19, e0304925 (2024).
Voolstra, C. R. et al. Contrasting heat stress response patterns of coral holobionts across the Red Sea suggest distinct mechanisms of thermal tolerance. Mol. Ecol. 30, 4466–4480 (2021).
Percudani, R., Montanini, B. & Ottonello, S. The anti-HIV cyanovirin-N domain is evolutionarily conserved and occurs as a protein module in eukaryotes. Proteins: Struct. Funct. Bioinforma. 60, 670–678 (2005).
Gardner, S. G. et al. Coral microbiome diversity reflects mass coral bleaching susceptibility during the 2016 El Niño heat wave. Ecol. Evol. 9, 938–956 (2019).
Ziegler, M. et al. Coral bacterial community structure responds to environmental change in a host-specific manner. Nat. Commun. 10, 3092 (2019).
Stalder, T., Press, M. O., Sullivan, S., Liachko, I. & Top, E. M. Linking the resistome and plasmidome to the microbiome. ISME J. 13, 2437–2446 (2019).
Kellogg, C. A. Microbiomes of stony and soft deep-sea corals share rare core bacteria. Microbiome 7, 1–13 (2019).
Mashini, A. G. et al. The influence of symbiosis on the proteome of the Exaiptasia Endosymbiont Breviolum minutum. Microorganisms 11, 292 (2023).
Pernice, M. et al. A single-cell view of ammonium assimilation in coral–dinoflagellate symbiosis. ISME J. 6, 1314–1324 (2012).
Roberts, J. M., Fixter, L. & Davies, P. Ammonium metabolism in the symbiotic sea anemone Anemonia viridis. Hydrobiologia 461, 25–35 (2001).
Biscéré, T. et al. Enhancement of coral calcification via the interplay of nickel and urease. Aquat. Toxicol. 200, 247–256 (2018).
Grover, R., Maguer, J.-F., Allemand, D. & Ferrier-Pagès, C. Urea uptake by the scleractinian coral Stylophora pistillata. J. Exp. Mar. Biol. Ecol. 332, 216–225 (2006).
Barnes, D. & Crossland, C. Urease activity in the staghorn coral, Acropora acuminata. Comp. Biochem. Physiol. Part B: Comp. Biochem. 55, 371–376 (1976).
Chen, M.-C. et al. ApRab11, a cnidarian homologue of the recycling regulatory protein Rab11, is involved in the establishment and maintenance of the Aiptasia–Symbiodinium endosymbiosis. Biochem. Biophys. Res. Commun. 338, 1607–1616 (2005).
Downs, C. A. et al. Symbiophagy as a cellular mechanism for coral bleaching. Autophagy 5, 211–216 (2009).
Araki, N. Role of microtubules and myosins in Fc gamma receptor-mediated phagocytosis. Front. Biosci. 11, 1479–1490 (2006).
Kvennefors, E. C. E., Leggat, W., Hoegh-Guldberg, O., Degnan, B. M. & Barnes, A. C. An ancient and variable mannose-binding lectin from the coral Acropora millepora binds both pathogens and symbionts. Dev. Comp. Immunol. 32, 1582–1592 (2008).
Fujita, T., Matsushita, M. & Endo, Y. The lectin-complement pathway–its role in innate immunity and evolution. Immunol. Rev. 198, 185–202 (2004).
Moncada, D. M., Kammanadiminti, S. J. & Chadee, K. Mucin and Toll-like receptors in host defense against intestinal parasites. Trends Parasitol. 19, 305–311 (2003).
Rivera, H. & Davies, S. Symbiosis maintenance in the facultative coral, Oculina arbuscula, relies on nitrogen cycling, cell cycle modulation, and immunity. Sci. Rep.11, 21226 (2021).
Mydlarz, L. D., McGinty, E. S. & Harvell, C. D. What are the physiological and immunological responses of coral to climate warming and disease? J. Exp. Biol. 213, 934–945 (2010).
Bin, P., Huang, R. & Zhou, X. Oxidation resistance of the sulfur amino acids: methionine and cysteine. BioMed. Res. Int. 2017, 9584932 (2017).
Ruiz-Jones, L. J. & Palumbi, S. R. Tidal heat pulses on a reef trigger a fine-tuned transcriptional response in corals to maintain homeostasis. Sci. Adv. 3, e1601298 (2017).
Sundborger, A. C. & Hinshaw, J. E. Regulating dynamin dynamics during endocytosis. F1000prime Rep. 6, 85 (2014).
Padilla-Gamiño, J. L., Pochon, X., Bird, C., Concepcion, G. T. & Gates, R. D. From parent to gamete: vertical transmission of Symbiodinium (Dinophyceae) ITS2 sequence assemblages in the reef building coral Montipora capitata. PLoS ONE 7, e38440 (2012).
Rodrigues, L. J. & Padilla-Gamiño, J. L. Trophic provisioning and parental trade-offs lead to successful reproductive performance in corals after a bleaching event. Sci. Rep.12, 18702 (2022).
Jaffe, M. D., Padilla-Gamiño, J. L., Nunn, B. L. & Rodrigues, L. J. Coral trophic pathways impact the allocation of carbon and nitrogen for egg development after bleaching. Front. Ecol. Evol. 11, 1251220 (2023).
Lehninger, A. L., Nelson, D. L., Cox, M. M. & Cox, M. M. Lehninger principles of biochemistry (Macmillan, 2005).
Lindqvist, Y., Schneider, G., Ermler, U. & Sundström, M. Three-dimensional structure of transketolase, a thiamine diphosphate dependent enzyme, at 2.5 A resolution. EMBO J. 11, 2373–2379 (1992).
Schnuck, J. K., Sunderland, K. L., Kuennen, M. R. & Vaughan, R. A. Characterization of the metabolic effect of β-alanine on markers of oxidative metabolism and mitochondrial biogenesis in skeletal muscle. J. Exerc. Nutr. Biochem. 20, 34 (2016).
Hillyer, K. E., Tumanov, S., Villas-Bôas, S. & Davy, S. K. Metabolite profiling of symbiont and host during thermal stress and bleaching in a model cnidarian–dinoflagellate symbiosis. J. Exp. Biol. 219, 516–527 (2016).
Hambleton, E. A. et al. Sterol transfer by atypical cholesterol-binding NPC2 proteins in coral-algal symbiosis. Elife 8, e43923 (2019).
Tortorelli, G. et al. Cell surface carbohydrates of symbiotic dinoflagellates and their role in the establishment of cnidarian–dinoflagellate symbiosis. ISME J. 16, 190–199 (2022).
Kopp, C. et al. Subcellular investigation of photosynthesis-driven carbon assimilation in the symbiotic reef coral Pocillopora damicornis. Mbio 6, e02299–02214 (2015).
Ngugi, D. K., Ziegler, M., Duarte, C. M. & Voolstra, C. R. Genomic blueprint of glycine betaine metabolism in coral metaorganisms and their contribution to reef nitrogen budgets. Iscience 23, 101120 (2020).
O’Brien, P. A. et al. Diverse coral reef invertebrates exhibit patterns of phylosymbiosis. ISME J. 14, 2211–2222 (2020).
Palmer, C. & Traylor-Knowles, N. Towards an integrated network of coral immune mechanisms. Proc. R. Soc. B: Biol. Sci. 279, 4106–4114 (2012).
Matthews, J. L. et al. Optimal nutrient exchange and immune responses operate in partner specificity in the cnidarian-dinoflagellate symbiosis. Proc. Natl. Acad. Sci. USA 114, 13194–13199 (2017).
Parkinson, J. E. et al. Molecular tools for coral reef restoration: Beyond biomarker discovery. Conserv. Lett. 13, e12687 (2020).
Baquiran, J. I. P. et al. Microbiome stability is linked to coral thermotolerance. bioRxiv, 2024.2010. 2003.616497 (2024).
Ramos-Silva, P. et al. The skeletal proteome of the coral Acropora millepora: the evolution of calcification by co-option and domain shuffling. Mol. Biol. Evol. 30, 2099–2112 (2013).
Vick, B. A. & Zimmerman, D. C. Lipids: structure and function (Elsevier, 1987).
Barott, K. L., Thies, A. B. & Tresguerres, M. V-type H+-ATPase in the symbiosome membrane is a conserved mechanism for host control of photosynthesis in anthozoan photosymbioses. R. Soc. Open Sci. 9, 211449 (2022).
Padilla-Gamino, J. L. et al. Are all eggs created equal? A case study from the Hawaiian reef-building coral Montipora capitata. Coral Reefs 32, 137–152 (2013).
Timmins-Schiffman, E. et al. Reproductive resilience: pathways to gametogenic success in Montipora capitata after bleaching. Sci. Rep.14, 27765 (2024).
Stat, M. et al. Variation in Symbiodinium ITS2 sequence assemblages among coral colonies. PloS ONE 6, e15854 (2011).
Cunning, R. & Baker, A. C. Excess algal symbionts increase the susceptibility of reef corals to bleaching. Nat. Clim. Change 3, 259–262 (2013).
Innis, T., Cunning, R., Ritson-Williams, R., Wall, C. & Gates, R. Coral color and depth drive symbiosis ecology of Montipora capitata in Kāne ‘ohe Bay, O ‘ahu, Hawai ‘i. Coral Reefs 37, 423–430 (2018).
Matz, M. V., Wright, R. M. & Scott, J. G. No control genes required: Bayesian analysis of qRT-PCR data. PloS one 8, e71448 (2013).
Frazier, M., Helmkampf, M., Bellinger, M. R., Geib, S. M. & Takabayashi, M. De novo metatranscriptome assembly and coral gene expression profile of Montipora capitata with growth anomaly. BMC Genom. 18, 1–11 (2017).
Choi, H., Kim, S., Fermin, D., Tsou, C. C. & Nesvizhskii, A. I. QPROT: statistical method for testing differential expression using protein-level intensity data in label-free quantitative proteomics. J. Proteom. 129, 121–126 (2015).
Brown, T., Sonett, D., Zaneveld, J. R. & Padilla-Gamiño, J. L. Characterization of the microbiome and immune response in corals with chronic Montipora white syndrome. Mol. Ecol. 30, 2591–2606 (2021).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl. Acad. Sci. USA 108, 4516–4522 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Weiss, S. et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome 5, 27 (2017).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Yilmaz, P. et al. The SILVA and “All-species Living Tree Project (LTP)” taxonomic frameworks. Nucleic Acids Res. 42, D643–D648 (2014).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 26, 32–46 (2001).
Mirarab, S., Nguyen, N. & Warnow, T. Biocomputing. 2012 247-258 (World Scientific, 2012).
Lin, H. & Peddada, S. D. Analysis of compositions of microbiomes with bias correction. Nat. Commun. 11, 3514 (2020).
Acknowledgements
We would like to thank the Gates Coral Lab for hosting and supporting us during this experiment at the Hawaiʻi Institute of Marine Biology. We thank Brenner Wakayama, Gavin Kreitman, Melissa Jaffe, and Sean Frangos for their help collecting and culturing the experimental corals. Work was supported in part by the University of Washington’s Proteomics Resource (UWPR95794), by NSF IOS-IEP 1655682 awarded to B.L.N. and J.L.P.G., NSF IOS-IEP 1655888 to L.J.R., NSF-IOS 2041497 to B.L.N., NSF GFRP awarded to J.B.A., NSF CAREER 2044840 awarded to J.L.P.G., NIH-F31 awarded to M.C.M. and Alfred P. Sloan Research Fellowship to J.L.P.G. and by CCN (private funds to support environmentally relevant research aimed at making a difference) to B.L.N.
Author information
Authors and Affiliations
Contributions
Brook L. Nunn designed the proteomic study of resilient vs susceptible corals, performed all mass spectrometry, data analysis and synthesis and was the primary author of the manuscript. Tanya Brown was responsible for all field experiments, and performed microbiome analyses and interpretations with Jesse Zaneveld. Brook L. Nunn, Emma Timmins-Schiffman, and Tanya Brown performed the protein extractions, data analysis and interpretation. Tanya Brown and Jeremy B. Axworthy assisted with field work and symbiont counts. Miranda C. Mudge and Emma Timmins-Schiffman assisted in proteomic data interpretations and visualizations. Michael Riffle assisted in statistical proteomic analyses and MetaGOmics analyses. Jenna Dilworth and Carly D. Kenkel performed the qPCR of the symbiont genera and wrote relevant portions of the manuscript. Lisa J. Rodrigues assisted in the study design, field experiments, and performed the lipid analyses. Jacqueline L. Padilla-Gamiño designed the experimental time-series study design, led the fieldwork. Brook L. Nunn, Jacqueline L. Padilla-Gamiño, and Lisa J. Rodrigues collaborated to acquire funding. All co-authors contributed to the writing and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Earth & Environment thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Christopher Cornwall and Alice Drinkwater. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nunn, B.L., Brown, T., Timmins-Schiffman, E. et al. Protein signatures predict coral resilience and survival to thermal bleaching events. Commun Earth Environ 6, 191 (2025). https://doi.org/10.1038/s43247-025-02167-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43247-025-02167-7
This article is cited by
-
Symbiotic Symbiodiniaceae mediate coral-associated bacterial communities along a natural thermal gradient
Environmental Microbiome (2025)
-
Species-specific retention vs. recovery of coral thermal tolerance following nursery propagation
Communications Biology (2025)
-
Ten years of coral reef benthic changes in the Central Mexican Pacific
Discover Ecology (2025)
-
Responses of the Coral Symbiont Cladocopium goreaui to Extreme Temperature Stress in Relatively High-Latitude Reefs, South China Sea
Microbial Ecology (2025)