Abstract
Background
Yellow fever virus remains a major public health threat in Brazil, where recent resurgence risks affecting both forest and periurban populations. Understanding viral movement across ecological settings is critical to support early detection and prevent outbreaks.
Methods
We performed genomic surveillance in two Brazilian states, a northern Amazon region and a southeastern region, between 2023 and 2025. Human and non-human primate samples were collected across forest, rural, and urban environments. Viral genomes were generated and analyzed using phylogenetic, phylogeographic, and temporal approaches to reconstruct viral transmission patterns.
Results
Here, we show evidence of continued yellow fever virus circulation and diversification in distinct ecological settings. We generate 25 genomes from humans and non-human primates, including the first human-derived genomes from the Amazon region. All genomes fall within the South American lineage. We identify one cluster in the Amazon region consistent with undetected viral persistence and reemergence, and a second cluster in the southeast associated with reintroduction followed by sustained local transmission.
Conclusions
These findings demonstrate ongoing yellow fever virus activity in Brazil, with forest regions serving as reservoirs for reemergence and periurban areas supporting continued spread. Strengthened genomic and epizootic surveillance is required to detect viral expansion early and inform targeted prevention strategies across Brazil and the Americas.
Plain language summary
Yellow fever is a serious mosquito-borne disease that affects both people and animals in Brazil. Although vaccination has helped control past outbreaks, the virus continues to circulate in forested regions and sometimes spreads toward towns and cities, creating a risk for new epidemics. In this study, we monitored yellow fever activity in two parts of Brazil between 2023 and 2025: the Amazon region in the north and a southeastern region where past outbreaks occurred. We analyzed samples from people and monkeys to track how the virus is spreading over time and across different environments. We found ongoing virus circulation in both regions, showing that yellow fever can persist quietly and then reappear. Strengthening surveillance is essential to detect early warning signs and protect at-risk communities.
Similar content being viewed by others
Introduction
Yellow fever (YF) is a mosquito-borne disease caused by Yellow Fever virus (species Orthoflavivirus flavi, family Flaviviridae), which continues to pose a significant public health threat in endemic regions of Africa and the Americas1. The virus is maintained in nature through two primary transmission cycles: a sylvatic cycle involving nonhuman primates (NHPs) and forest-dwelling mosquitoes such as Haemagogus and Sabethes spp., and an urban cycle involving Aedes aegypti, which has not been documented in Brazil since 19422. While eradication is not feasible due to the presence of animal reservoirs, sustained high vaccination coverage remains the most effective strategy to prevent urban outbreaks. Despite the availability of an effective vaccine, Brazil experienced repeated episodes of yellow fever virus (YFV) re-emergence between 2016 and 20223,4,5. In recent years, genomic surveillance and case reporting have raised concerns about ongoing viral circulation and expansion into new areas. In early 2023, a confirmed YFV case in an NHP in São Paulo was shown to be linked to a previously undetected introduction from the Midwest6. Subsequently, the Pan American Health Organization (PAHO) issued alerts regarding increasing YFV activity throughout the Americas, signaling persistent viral circulation and the potential for renewed spread7. In this study, we investigate the resurgence of YFV in Brazil between 2023 and 2025, focusing on its spatiotemporal dynamics in both forested and peri-urban settings using integrated genomic, epidemiological, and geographic analyses. We generate 25 YFV genomes from human and non-human primate samples collected in Pará (Northern Amazon) and Minas Gerais (Southeast), including the first human-derived sequences ever reported from the Amazon region. Phylogenetic and phylogeographic analyses reveal two distinct regional lineages: one in Pará showing evidence of undetected viral persistence and re-emergence, and another in Minas Gerais likely resulting from a more recent reintroduction from the Midwest followed by sustained local transmission. Overall, our findings demonstrate ongoing viral diversification and persistence across ecologically distinct regions, underscoring the importance of continuous genomic and epizootic surveillance to detect early signals of YFV expansion and guide targeted prevention strategies in Brazil and across the Americas.
Methods
Sample collection
Human and nonhuman primate (NHP) samples were collected by local and regional public health teams as part of Brazil’s national yellow fever surveillance program, coordinated by the Brazilian Ministry of Health (BrMoH)8. Human samples were obtained from individuals presenting with clinical symptoms consistent with yellow fever virus (YFV) infection. NHP samples were collected from animals found dead or visibly ill, particularly in areas where epizootics had been reported, including urban and peri-urban environments. All samples were submitted for molecular screening to the Central Public Health Laboratories (LACENs) in the states of Pará (Northern Brazil) and Minas Gerais (Southeastern Brazil), in accordance with established national protocols for arbovirus surveillance.
Molecular screening and whole genome sequencing
RNA was extracted from serum and tissue samples using either the QIAamp Viral RNA Mini Kit™ (Qiagen, Hilden, Germany) or an automated protocol on the MagNA Pure 96 platform, employing the MagNA Pure 96 DNA and Viral NA Small Volume Kit (Roche, Basel, Switzerland), following the manufacturers’ instructions. Detection of YFV RNA was performed using the RT-qPCR protocol described by Domingo et al.9. Sequencing was attempted on 25 RT-qPCR-positive samples, regardless of cycle threshold (Ct) values or the availability of associated epidemiological data3,4,5. From the total number of confirmed positive samples, those prioritized for sequencing were selected based on the availability of associated metadata, RNA integrity, and Ct values below 35—criteria that ensure suitability for successful amplification and sequencing. Geographic representation across affected municipalities within each state was also considered to enhance spatial coverage and improve the interpretability of the resulting genomic data. All positive samples underwent complementary DNA (cDNA) synthesis using the ProtoScript II First Strand cDNA Synthesis Kit (NEB). Subsequently, a multiplex tiling PCR was performed following a previously published YFV primer scheme, using Q5 High-Fidelity DNA Polymerase (NEB) for 35 amplification cycles, as described by Quick et al.10. Amplicons were purified using AMPure XP beads (Beckman Coulter), and DNA concentrations were quantified with the Qubit dsDNA High Sensitivity Assay Kit on a Qubit 3.0 fluorometer (Thermo Fisher Scientific). DNA libraries were prepared for sequencing using the COVIDSeq Assay (Illumina MiSeq platform) and the Native Barcoding Kit 96 V14 (Oxford Nanopore MinION platform), following the manufacturers’ instructions. Details regarding sample metadata, including host species, geographic origin (municipality), and collection dates, are provided in Table S1.
Generation of consensus sequences
Raw sequencing data were processed using platform-specific pipelines to generate high-quality consensus sequences. For MinION data, basecalling was performed using Guppy v4.5.4, and barcode demultiplexing was carried out with Qcat. Consensus sequences were first generated through de novo assembly using Genome Detective (https://www.genomedetective.com/)11,12. Briefly, Genome Detective employs DIAMOND to identify and classify candidate viral reads based on the viral subset of the SwissProt UniRef90 protein database. These reads are then mapped to reference sequences using NCBI BLASTn and aligned using AGA (Annotated Genome Aligner) and MAFFT, producing final contigs and consensus sequences in FASTA format. In parallel, a complementary reference-based approach was implemented using a custom pipeline adapted for each sequencing platform. For Illumina data, the pipeline included FastQC (v0.12.1) for quality control, Trimmomatic (v0.36) for adapter and low-quality base trimming, Minimap2 (v2.28) for read alignment, Samtools (v1.20) for alignment processing, Bcftools (v1.20) for variant calling, iVar (v1.4.2) for consensus genome generation, and Pilon (v1.24) for final sequence polishing. For MinION data, raw fast5 files were basecalled and demultiplexed using Guppy (v6.0), assessed for quality using NanoPlot (v1.43.0), and trimmed with Chopper (v0.8.0). Subsequent steps—alignment, variant calling, consensus generation, and polishing—mirrored those of the Illumina pipeline, ensuring consistency in genome reconstruction across sequencing platforms.
Phylogenetic and phylogeographic reconstruction
Genotyping of yellow fever virus (YFV) sequences was performed using the Yellow Fever Typing Tool available at https://www.genomedetective.com/app/typingtool/yellowfever/. The genome sequences generated in this study were combined with a reference dataset of 184 previously published YFV genomes, including recently released sequences1. Multiple sequence alignment was conducted using MAFFT12 and manually curated in AliView13 to remove potential artefacts. Maximum likelihood (ML) phylogenetic trees were reconstructed using IQ-TREE214, employing the GTR substitution model, which was selected as the best-fitting model by ModelFinder15, as implemented in IQ-TREE2. Statistical support for tree topology was assessed through 1000 bootstrap replicates. To evaluate the temporal signal in the dataset, root-to-tip genetic distances from the ML tree were regressed against sampling dates using TempEst v1.5.116. For time-calibrated phylogeographic reconstruction, Bayesian inference was performed using BEAST v1.10.417, under the GTR + Γ4 substitution model, an uncorrelated relaxed molecular clock with a lognormal distribution, and the Bayesian Skyline coalescent tree prior, following previously established protocols5. Two independent Markov Chain Monte Carlo (MCMC) runs were conducted for 100 million steps, with sampling every 10,000 steps. Convergence of MCMC chains and sufficient effective sample sizes (ESS > 200) were confirmed using Tracer v1.7.118. The resulting trees were summarized into a maximum clade credibility (MCC) tree using TreeAnnotator, discarding the initial 10% as burn-in. We categorized sequence sampling locations into six distinct geographic regions based on available genome data, as illustrated in Fig. 1e: North Brazil (Roraima, Pará, Tocantins); Midwest Brazil (Goiás, Federal District); Northeast Brazil (Bahia); Southeast Brazil (Minas Gerais, Espírito Santo, Rio de Janeiro, São Paulo); South Brazil (Paraná, Santa Catarina, Rio Grande do Sul); and South America, which includes sequences from Colombia and Venezuela.
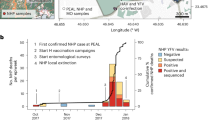
a Map of Brazil indicating the states of Pará (dark blue, North) and Minas Gerais (gold, Southeast) under investigation; b Proportion of YFV genomes by host (human: orange; NHP: green) and ecological classification (urban, rural/peri-urban, rural, peri-urban, forest/remote rural); c Sampling dates of sequenced genomes by state (PA and MG) and host; d Age distribution of human cases by state (Pará: n = 10 biologically independent samples; Minas Gerais: n = 4 biologically independent samples); e Time-scaled maximum likelihood phylogeny including genomes generated in this study and publicly available sequences. Tip colors indicate sampling location, defined as follows: North: Roraima (RR), Pará (PA), Tocantins (TO); Midwest: Goiás (GO), Federal District (DF); Northeast: Bahia (BA); Southeast: Minas Gerais (MG), Espírito Santo (ES), Rio de Janeiro (RJ), São Paulo (SP); South: Paraná (PR), Santa Catarina (SC), Rio Grande do Sul (RS); South America: Colombia (CO), Venezuela (VE). Tip fill denotes host (human: orange; NHP: green; mosquito: beige). Vertical bars to the right summarize host metadata. All sequences belong to the South American I lineage.
Ethical statement
This project was supported by the PAHO and the BrMoH as part of ongoing efforts to strengthen arboviral genomic surveillance. All procedures complied with Resolution 510/2016 of the Brazilian National Research Ethics Commission (CONEP—Comissão Nacional de Ética em Pesquisa, Ministério da Saúde). Under this resolution, the use of anonymized clinical samples collected during public health emergencies is permitted without informed consent, provided they are surplus to routine diagnostics and intended to support surveillance and outbreak response. The samples used in this study were obtained anonymously from material exceeding routine arbovirus diagnostics within Brazil’s public health laboratory network.
Results
Geographical spread of recent yellow fever activity
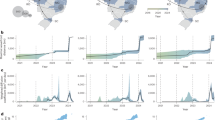
To characterize recent transmission dynamics, we first analyzed the geographic distribution of confirmed human cases between 2023 and 2025, which revealed a progressive eastward expansion of YFV circulation (Fig. 2). In 2023, the majority of cases remained concentrated in western Amazonian regions. However, in 2024 and early 2025, new clusters were identified in central and northern Brazil, with increasing detections in southeastern states, including previously unaffected areas of the states of São Paulo and Minas Gerais. This pattern is consistent with historical observations of YFV dissemination from enzootic Amazonian reservoirs into more populated transitional zones5. The country-level distribution of YFV cases demonstrates a marked increase in transmission across the region, with Colombia experiencing increases in 2024-2025 and Brazil experiencing the sharpest rise in 2025 (Fig. 2b). At the Brazilian state level, surveillance data highlight substantial increases in both case numbers and YFV-related mortality in Pará and Minas Gerais, which remain areas of concern due to their ecological connectivity and the presence of vulnerable populations with suboptimal vaccine coverage (Fig. 2c).
a Geographical distribution of confirmed human YFV cases overlaid on a land cover map of South America. Brazil is highlighted with a black border. Red circles indicate reported cases for each respective year (2023, 2024, and 2025); b Number of confirmed YFV cases reported per year by country (Bolivia, Brazil, Colombia, Guyana, and Peru). A marked increase in total cases is observed in 2025; c Number of confirmed cases (left) and deaths (right) reported annually by selected Brazilian states. Data highlight rising case numbers in Pará and Minas Gerais, with associated increases in YFV-related mortality.
Genome sequencing and sample characteristics
To investigate the emergence and spread of YFV in two affected Brazilian states - Pará (PA) and Minas Gerais (MG) - we implemented an integrated molecular and genomic surveillance approach. Clinical samples were processed at the state public health laboratories, where viral RNA was extracted and screened by RT-qPCR9. Positive samples underwent whole-genome sequencing on MiSeq (Illumina) and MinION (Oxford Nanopore) using the COVIDSeq Assay and Native Barcoding Kit 96 V14, respectively. Consensus genomes were aligned with complete YFV sequences from GenBank (as of March 24, 2025) using MAFFT12, and phylogenetic analyses were performed with IQ-TREE219 and BEAST v1.10.417. A total of 25 YFV genomes were generated from samples collected between 2023 and 2025, comprising 14 human and 11 NHP sequences. These genomes were obtained from different sample types, including serum (n = 14), liver (n = 9), spleen (n = 1), and lung (n = 1), and originated from forest, rural, peri-urban, and urban settings (Table S1). Samples were geographically distributed between Pará in the Northern region (n = 10) and Minas Gerais in the Southeastern region of Brazil (n = 15) (Fig. 1a).
Ecological Context and Epizootic Observations
Ecological classification of the sampling sites indicated that all human infections identified in Pará were associated with forested or remote rural environments. In contrast, viral genomes recovered from Minas Gerais were predominantly obtained from NHPs across a broader range of ecological contexts, including rural, peri-urban, and urban areas. These findings likely reflect regional differences in host surveillance and sampling coverage rather than true ecological distribution. Notably, the exclusive detection of human infections in Pará is of epidemiological significance given the limited documentation of YFV human cases in this northern region in recent years (Fig. 1b, Table S1). This is further supported by active surveillance near the municipality of Breves, where health authorities reported NHP carcasses and accounts from local residents of primate mortality occurring 2–3 months before human cases. Such observations underscore the role of NHP epizootic surveillance as a critical early warning tool for YFV circulation8. However, implementing this strategy in the Amazon remains challenging due to dense vegetation and limited access, which can hinder timely sample collection and transport.
Temporal dynamics and sequencing performance
Temporal analysis of the newly generated genomes indicated sustained YFV circulation in Minas Gerais from early 2023 to early 2025, whereas in Pará YFV genome detections were restricted to a shorter period, with all sequences clustering in 2025 (Fig. 1c). Demographic data from confirmed human infections (n = 14) showed that all cases occurred in males aged between 17 and 65 years. Comparative analysis revealed a higher median age among infected individuals in Minas Gerais (~52 years) relative to those in Pará (~39 years), although this difference was not statistically significant (t-test p = 0.32, Fig. 1d), likely due to the small number of data points available for Minas Gerais (n = 4). Sequencing quality varied across samples, with Ct values ranging from 11 to 35. Genome coverage exceeded 70% in 20 of the 25 sequences, and as expected, depth of coverage showed an inverse correlation with Ct values (Table S1).
Phylodynamic insights into viral emergence and spread
All genomes generated in this study were classified within the South American I lineage (Fig. 1e). The sequences from Pará formed a well-supported monophyletic clade, with Bayesian phylogenetic and phylogeographic analyses estimating its emergence around mid-March 2024 (95% highest posterior density [HPD]: early March 2023 to October 2024) and strongly supporting a Northern origin (posterior probability [pp] = 1.0). Notably, this clade is rooted by a basal strain from Pará collected in 2017, suggesting the possibility of undetected viral persistence followed by re-emergence in the region. In contrast, sequences from Minas Gerais clustered with recent genomes from São Paulo1, forming a distinct lineage closely related to a genome from Goiás (Midwest Brazil). Bayesian phylogeographic analysis supported the Midwest as the most likely origin of this clade (posterior probability [pp] = 1.0). The estimated time to the most recent common ancestor (tMRCA) was late February 2019 (95% highest posterior density [HPD]: mid-January 2018 to mid-July 2019). This lineage was genetically distinct from strains associated with the 2016–2018 southeastern outbreak, supporting a scenario of a more recent reintroduction likely occurring around early 2022, followed by sustained local transmission and diversification.
Discussion
This study provides genomic and epidemiological evidence of the co-circulation of distinct yellow fever virus (YFV) lineages across Brazil during a period of renewed viral activity, revealing both undetected persistence in the North—particularly in Pará—and sustained endemic transmission in the Southeast. The observed differences in host type and detection setting—human infections in rural Pará versus NHP detections across rural, peri-urban, and urban areas in Minas Gerais—likely reflect disparities in surveillance coverage and accessibility. In the Amazon region, detection remains hindered by dense forest cover and logistical challenges in reaching remote areas, leading to underreporting of both human and NHP infections¹². By contrast, the more urbanized landscape and established health infrastructure in Minas Gerais facilitate broader surveillance, including systematic monitoring of NHPs in peri-urban zones.
Phylogenetic and spatial analyses indicate an eastward spread of YFV from the Amazon between 2023 and 2025, with additional clusters detected in central and southeastern regions. The mechanisms driving this pattern remain to be fully elucidated but likely involve a combination of factors, including the movement of infected individuals, limited yet ecologically relevant dispersal of NHPs, and potential passive or human-assisted transportation of infected vectors. Although there is currently no direct evidence in Brazil supporting sylvatic vectors transmitting the virus from infected humans to other humans or to NHPs, such events cannot be excluded given the ecological complexity of transmission in forest-edge environments. The transport of infected NHPs over long distances, however, is improbable, suggesting that local viral persistence and short-range dispersal through enzootic corridors represent more plausible explanations. It is notable that the Pará 2025 sequences clustered with a single Pará genome from 2017 but were separated by a long branch. This phylogenetic pattern may indicate undetected viral persistence in the region for multiple years, although incomplete sampling during the intervening period or independent re-emergence events cannot be ruled out. Nevertheless, the formation of a well-supported monophyletic clade suggests localized viral evolution, emphasizing the need for sustained genomic surveillance in historically undersampled areas such as northern Brazil. Our data support the hypothesis that YFV likely circulated cryptically in Pará since at least 2017, while the sequences from Minas Gerais represent a separate introduction from a midwestern lineage dated to 2019, distinct from those associated with the 2016–2018 southeastern outbreaks. These results reinforce the importance of regionally adapted genomic monitoring to clarify lineage dynamics and detect early signs of resurgence. Although Brazil maintains a long-standing national yellow fever immunization program, vaccination coverage remains heterogeneous, with persistently lower rates in remote and rural areas—including parts of northern Brazil—where access to healthcare is limited and outbreak response is delayed20. These disparities leave certain populations at greater risk of spillover and sustained transmission. The continued presence of competent vectors such as Aedes aegypti and suboptimal 17DD vaccination coverage across several regions heightens concerns about the possible re-establishment of an endemic transmission cycle. Importantly, no evidence of YFV infection was detected in Aedes aegypti during the 2023–2025 period, and transmission remained confined to the sylvatic cycle, even near densely populated areas with low vaccination coverage. The absence of Aedes aegypti-borne amplification during this period likely reflects a combination of ecological barriers, limited opportunities for urban transmission, and the effectiveness of early detection and control efforts. Nevertheless, these findings highlight the need to strengthen entomological surveillance and maintain high vaccine coverage in areas at risk of urban spillover. Recent studies have also proposed that cross-reactive immunity from prior dengue or Zika virus infections could influence yellow fever transmission dynamics. While in vitro evidence indicates that sera from individuals previously infected with dengue or Zika can neutralize YFV21, this has not been confirmed in vivo, and there is currently no evidence that such cross-reactivity provides protection in field conditions. Moreover, a recent cohort study in Brazil found no impact of prior DENV or ZIKV infection on YFV vaccine antibody responses22. Thus, although biologically plausible, the role of cross-protective immunity in limiting urban yellow fever transmission remains uncertain and warrants further investigation.
This study is not without limitations. The genomic data analyzed were restricted to samples collected in only two states—Pará and Minas Gerais—and over a limited temporal window (2023–2025). As phylogeographic inferences are highly sensitive to sampling scale, these constraints may limit the detection of intermediary transmission events and broader dissemination patterns across Brazil. These limitations are especially relevant given the proximity of recent cases to densely populated areas with low immunity, where viral spread may go undetected. This underscores the urgent need to strengthen entomological surveillance, improve early warning systems, and expand vaccination coverage, particularly in under-resourced regions. Despite these constraints, the study provides several important and impactful contributions. Most notably, it presents the first genomic data from human YFV cases in northern Brazil—an area historically underrepresented in national surveillance efforts. Prior to this work, only a handful of YFV genomes from the North were publicly available, all from nonhuman primates. By contributing new sequences from both human and NHP cases in Pará, this study fills a critical gap in our understanding of YFV circulation in the Amazon. These data offer unique insights into viral persistence and re-emergence dynamics that would otherwise remain obscured. Through the integration of genomic, spatial, and epidemiological data, this work enhances our resolution of YFV transmission and highlights the continued need for comprehensive and equitable surveillance across diverse ecological settings.
Data availability
All genomes generated in this study have been deposited in the public GenBank database under accession numbers PV710581–PV710608. All source data files have been made available in a public GitHub repository (link: https://github.com/genomicsurveillance/YFV_Reemergence_Brazil_2025).
Code availability
All codes associated with the analysis of the data used for the paper have been made available in a public GitHub repository (link: https://github.com/genomicsurveillance/YFV_Reemergence_Brazil_2025).
References
Lindenbach B. T. et al. Flaviviridae, in Fields Virology, D. M. Knipe, P. M. Howley, Eds. (Lippincott Williams & Wilkins, 2013), 712–746.
Possas, C. et al. Yellow fever outbreak in Brazil: the puzzle of rapid viral spread and challenges for immunisation. Mem. Inst. Oswaldo Cruz. 113, e180278 (2018).
Faria, N. R. et al. Genomic and epidemiological monitoring of yellow fever virus transmission potential. Science 361, 894–899 (2018).
Giovanetti, M. et al. Yellow fever virus reemergence and spread in Southeast Brazil, 2016-2019. J. Virol. 94, e01623–19 (2019).
Giovanetti, M. et al. Genomic epidemiology unveils the dynamics and spatial corridor behind the Yellow Fever virus outbreak in Southern Brazil. Sci. Adv. 9, eadg9204 (2023).
Fernandes, N. C. C. A. et al. Phylogenetic analysis reveals a new introduction of Yellow Fever virus in São Paulo State, Brazil, 2023. Acta Trop. 251, 107110 (2024).
Pan American Health Organization (PAHO). Alerta epidemiológica - Febre amarela na Região das Américas − 26 de março de 2025. Available at https://www.paho.org/pt/documentos/alerta-epidemiologica-febre-amarela-no-regiao-das-americas-26-marco-2025 (accessed on 26 March 2025).
Brazilian Ministry of Health, 2025. https://www.gov.br/saude/pt-br/assuntos/saude-de-a-a-z/f/febre-amarela/publicacoes/guia_vigilancia_epizootias_primatas_entomologia.pdf/view.
Domingo, C. et al. Advanced yellow fever virus genome detection in point-of-care facilities and reference laboratories. J. Clin. Microbiol. 50, 4054–4060 (2012).
Quick, J. et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat. Protoc. 12, 1261–1276 (2017).
Vilsker, M. et al. Genome detective: an automated system for virus identification from high-throughput sequencing data. Bioinformatics 35, 871–873 (2019).
Katoh, K. & Standley, D. M. MAFFT Multiple Sequence Alignment Software Version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Larsson, A. AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30, 3276–3278 (2014).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Rambaut, A., Lam, T. T., Max Carvalho, L. & Pybus, O. G. Exploring the temporal structure of heterochronous sequences using TempEst (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol. 4, vey016 (2018).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901–904 (2018).
Minh B. Q. et al. IQ-TREE 2: New Models and Efficient Methods for Phylogenetic Inference in the Genomic Era. Mol Biol Evol. 2020;37:1530-1534. Erratum in: Mol Biol Evol. 2020 Aug 1;37(8):2461.
Brazilian Ministry of Health, Yellow Fever vaccination coverage, 2025. https://infoms.saude.gov.br/extensions/SEIDIGI_DEMAS_VACINACAO_CALENDARIO_NACIONAL_COBERTURA_RESIDENCIA/SEIDIGI_DEMAS_VACINACAO_CALENDARIO_NACIONAL_COBERTURA_RESIDENCIA.html.
Oliveira, R. A. et al. Previous dengue or Zika virus exposure can drive to infection enhancement or neutralisation of other flaviviruses. Mem. Inst. Oswaldo Cruz. 114, e190098 (2019).
Saucha, C. V. V. et al. Collaborative Group for Yellow Fever Vaccine Studies. Immunogenicity of a single dose of the 17DD yellow fever vaccine in a cohort of adults and children in a non-endemic area, and its association with dengue and Zika seropositivity. PLoS Negl. Trop. Dis. 19, e0012993 (2025).
Acknowledgements
This study was supported by the National Institutes of Health (USA) grant U01 AI151698 for the United World Arbovirus Research Network (UWARN), the Novo Nordisk Foundation (grant NNF24OC0094346), and the Rede Unificada de Análises Integradas de Arbovírus de Minas Gerais (REDE UAI-ARBO-MG), financed by the Fundação de Amparo à Pesquisa do Estado de Minas Gerais (FAPEMIG), grant number RED-00234-23. T.E.R.A. is supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), process number 157696/2025-1. N.R.G. and V.F. receive BDCTI-I scholarships from FAPEMIG.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.G. and L.C.J.A.; Methodology: V.G.D.D.A.; T.E.R.A.; V.F.; K.M.F.M.; L.M.R.T.; L.A.P; D.G.L.D.L.R.; A.M.B.D.F.; D.G.R.; D.B.R.; K.C.L.F.; G.A.L.B.; L.C.M.; L.C.V.F.; L.O.L.; N.R.G.; P.M.S.S.B.; M.S.D.O.; P.S.D.A.; P.E.S.D.S.; R.G.P.; R.G.S.; S.M.D.S.C.; S.H.S.P.P.; S.K.; S.G.P.; V.L.D.S.; J.L.; F.C.D.M.I.; A.S.J.J.; M.G.; L.C.J.A.; Investigation: V.G.D.D.A.; T.E.R.A.; V.F.; K.M.F.M.; L.M.R.T.; L.A.P.; D.G.L.D.L.R.; A.M.B.D.F.; D.G.R.; D.B.R.; K.C.L.F.; G.A.L.B.; L.C.M.; M.S.D.O.; L.C.V.F.; L.O.L.; N.R.G.; P.M.S.S.B.; P.S.D.A.; P.E.S.D.S.; R.G.P.; R.G.S.; S.M.D.S.C.; S.H.S.P.P.; S.K.; S.G.P.; V.L.D.S.; J.L.; F.C.D.M.I.; A.S.J.J.; M.G.; L.C.J.A.; Data curation: V.F., L.M.R.T., and M.G.; Original draft preparation: T.E.R.A., V.F., J.L. and M.G.; Visualization: V.F.; J.L. and M.G. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
das Graças Dantas Andrade, V., Ribeiro Adelino, T.É., Fonseca, V. et al. Reemergence of yellow fever virus in forest and periurban settings in Brazil. Commun Med 6, 35 (2026). https://doi.org/10.1038/s43856-025-01287-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01287-7