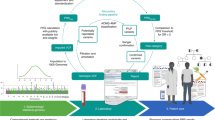
Abstract
Background
At present, there is no consensus on which genotyping platform should serve as the standard for clinical polygenic risk score (PRS) implementation. Previous studies have compared the overall performance and concordance of different genotyping and sequencing technologies; however, these analyses have generally averaged the results over the whole genome. We evaluated differences in a 313-variant breast cancer PRS (PRS313) across genomic platforms and their impact on risk stratification.
Methods
We compare PRS313 derived from genotyping arrays (Global Screening Array [GSA], OncoArray-500K [OncoArray], Global Diversity Array [GDA], custom Axiom_PrecipV1 array [ThermoFisher]) and low-coverage genome sequencing (lc-WGS) in 2 cell lines and 92 individuals. Probes are designed for all variants on ThermoFisher (success rate: 259/313). Sanger sequencing profiles indels. Concordance of high-risk classification (PRS313scoresum > 0.6) across platforms is assessed using Kappa statistics.
Results
In saliva samples, indel concordance with Sanger sequencing varies widely (Kappa: 0.007-1.000). PRS313-ThermoFisher is predictable from other platforms using linear models, despite systematic differences. Greater agreement is observed between arrays with high imputation overlap (e.g., GDA ~ GSA slope=0.986). Agreement in high-risk classification before mean correction is moderate (Fleiss’s Kappa=0.552) and improves after mean correction (Kappa=0.650). Arrays with similar designs show higher agreement before mean correction (Kappa=0.745). Mean correction narrows high-risk proportions from 4-45% to 15-21%. Overall, 26 of 92 samples are classified as high risk on at least one platform, but only 7 are high risk across all. When restricting to identical variants across all platforms for PRS313 calculation, the corresponding number of high-risk individuals are 24 and 11.
Conclusion
Our findings demonstrate that platform-specific variability can influence PRS313 estimates to potentially reclassify individuals around clinically relevant thresholds.
Plain Language Summary
This study looks at how different DNA sequencing methods can affect estimates of genetic risk for breast cancer. DNA sequencing is a process that determines the exact order of the building blocks (nucleotides) in a person’s genes. Different sequencing technologies and platforms may read this information slightly differently, leading to variations in the data even when analyzing the same individual. These differences can be important because they might change a person’s estimated risk for breast cancer, especially if the risk score is close to a level where doctors would take action. To study this, we analyzed a previously validated breast cancer risk model based on 313 genetic variants, called a polygenic risk score (PRS). A PRS combines the effects of many small genetic differences across the genome to estimate a person’s overall risk for developing a disease, rather than focusing on a single high-risk gene. We found that the estimated risk could shift depending on the sequencing platform used, revealing systematic biases in how people are classified into risk groups. We also identified concerns with certain types of genetic changes, called insertions and deletions (indels), which are sometimes inconsistently detected across platforms and could affect the reliability of the risk score.
Similar content being viewed by others
Introduction
Interest in using polygenic risk scores (PRS) in clinical practice for better disease screening and diagnosis is steadily increasing1,2,3. Although their validity is well-supported, the practical benefits of PRS in real-world settings remain insufficiently explored4,5,6. In breast cancer risk prediction, PRS alone performs poorly for individual risk prediction and population stratification, indicating the need for integration with other risk factors7,8,9,10,11. Another challenge is the variability in PRS distribution across populations, which may benefit from population-specific calibration12,13,14,15.
This study examines another source of variability: the impact of sequencing technology on PRS-derived breast cancer risk stratification. Sanger sequencing is recognized as the gold standard in many research and clinical applications due to its low error rate and high accuracy in detecting single-nucleotide variants (SNVs) and indels16. Arrays are key tools in PRS studies, offering cost-effective, high-accuracy genotyping for common variants17. Recent advancements in low-coverage genome sequencing (lc-WGS) and improved imputation methods, such as GLIMPSE2, have improved rare variant imputation (minor allele frequency [MAF] < 1%), making lc-WGS a strong alternative to arrays. In addition, many countries are investing in efforts to sequence their populations1,2,3. Previous studies have compared the overall performance and concordance of different genotyping and sequencing technologies; however, these analyses have generally averaged the results over the whole genome18,19. It remains unclear whether these findings can be generalized to specific sets of variants.
As PRS313 is increasingly used in clinical settings, it is important to understand whether sequencing platform differences could meaningfully alter risk estimates for these variants. Our study aims to fill this gap by focusing on a targeted 313-variant PRS (PRS313), the most widely studied breast cancer PRS20,21. We assessed how different genomic platforms impact the accuracy and reliability of risk stratification for this specific set of variants. Genome-wide heritability estimates suggest that PRS loci account for approximately 40–45% of the heritability attributed to all common variants captured by genome-wide SNV arrays20. Discrimination statistics (AUC) for PRS313 within White European populations are approximately 0.63–0.65321,22. PRS313 has been implemented in pilot risk-based breast cancer screening programs such as the Personalized Risk Assessment for Prevention and Early Detection of Breast Cancer: Integration and Implementation (PERSPECTIVE I&I) study23 and the BREAst cancer screening Tailored for HEr (BREATHE) study24. To ensure reliable risk prediction and clinical application, we evaluate the degree of agreement between array and lc-WGS results, as the breast cancer PRS313 is primarily developed and validated on arrays25.
At present, there is no consensus on which genotyping platform should serve as the standard for clinical PRS313 implementation. Our findings demonstrate that platform-specific variability can influence PRS313 estimates, potentially reclassifying individuals around clinically relevant thresholds. Women near the high-risk cut-off are particularly affected, as small differences in measurement can shift risk classification. This phenomenon is analogous to clinical metrics such as blood pressure or hemoglobin A1c, where minor differences between devices or assays can alter whether a patient crosses a diagnostic threshold and thus influence treatment decisions. Our study demonstrates that sequencing and genotyping platforms introduce systematic differences in PRS313-based breast cancer risk estimates. While PRS313 is generally predictable across platforms, variability in indel detection and differences in variant coverage can lead to inconsistent high-risk classifications. Correcting for platform-specific mean differences improves agreement but does not eliminate discrepancies entirely. These findings highlight that platform choice can influence clinical risk stratification and emphasize the importance of standardizing PRS calculation or calibrating scores across platforms to ensure reliable risk prediction.
Methods
Study population
Volunteers aged 21–80 years, who identified as Chinese, Malay, or Indian descent, were eligible for our study. Between 12 December 2022 and 16 June 2023, 100 individuals signed up, of whom 94 completed informed consent. Two individuals withdrew before providing saliva samples. Our analytical cohort includes 92 individuals who provided ~4 ml of saliva each (PAXgene Saliva Collectors from Qiagen, Part number: 769040, two tubes of 2 ml per individual), of whom 90 agreed to share their genetic data for future research beyond this study.
This study was approved by the A*STAR ethics board (reference number: 2022-062, approval date: 14 September 2022).
Cell lines
We included cell lines (to minimize the potential influence of DNA quality) on each array (GM18592, female; GM18609, male). Both cell lines were of cell type B-lymphocyte, ethnicity Han Chinese, from the repository NHGRI Sample Repository for Human Genetic Research. Each cell line underwent a single DNA extraction and library preparation; replicates represent repeated measurements of the same prepared sample in different wells, primarily to fill array slots. Genotyping chips have a fixed capacity. After assigning slots to the 92 unique samples, the remaining slots allowed for two replicates (Repeats 1–2) per cell line on GSA, OncoArray, and GDA. For ThermoFisher, the fixed capacity allowed for the inclusion of only one replicate per cell line. We note that this design does not capture variability from independent sample processing or library preparation, and thus, the replicates are not intended to assess full platform reproducibility. Instead, they provide a controlled means to evaluate measurement consistency under identical library preparation conditions.
DNA extraction
Saliva samples were pre-heated at 50 °C in an air incubator overnight before the purification of genomic DNA using prepIT•L2P reagent (Part number: PT-L2P-45) from DNA Genotek, according to the manufacturer’s recommendations.
“Ground truth”: direct genotyping and Sanger sequencing
We designed primers for all 313 variants in the breast cancer PRS313 panel using ThermoFisher Scientific’s Applied Biosystems™ Axiom™ arrays (Axiom_PrecipV1). However, 54 primers (17%) failed quality checks. In total, 259 variants (235/265 SNVs and 24/48 indels) were successfully genotyped on ThermoFisher. While primers for all 48 indels in the PRS313 panel were designed, four indels could not be successfully designed for Sanger sequencing validation (Supplementary Data 1). The remaining 44 indels were validated through Sanger sequencing.
Due to the large number of experiments required per individual and the limited amount of saliva samples available, we prioritized genotyping and lc-WGS experiments over Sanger sequencing validation. As a result, Sanger sequencing was performed for only 87–89 individuals (3 had no DNA for Sanger sequencing; 2 had insufficient DNA for all indels).
Genotyping and imputation on commercially available arrays
In addition to the custom ThermoFisher array, genotyping was done on three Illumina arrays: (1) Infinium Global Screening Array v3.0 (GSA), (2) Infinium OncoArray-500K v1.0 (Rev. C) (OncoArray), and (3) Infinium Global Diversity Array v1.0 (GDA). Genotype calling was conducted using the genotyping module v2.0.5 in the GenomeStudio 2.0.5 (Illumina, San Diego, CA, USA). Manifest files were downloaded from Illumina’s product website and requested from the ThermoFisher’s team. The number of unique variants (by chromosome number and position only) from chromosomes 1–22, X and Y: 646,728 on GSA, 497,147 on OncoArray, 1,943,749 on GDA, and 574,005 on ThermoFisher (Supplementary Data 2). The number of overlapping variants genotyped on the arrays is presented in Supplementary Fig. 1.
For Infinium arrays, liftover was done from build GRCh37 to GRCh38 using the script developed by Neil Robertson (https://www.chg.ox.ac.uk/~wrayner/strand/, accessed in February 2024). Genotyping strand and position files were downloaded from https://www.strand.org.uk/ (accessed February 2024): source (forward) strand was available for OncoArray-500K-C, and TOP strand files were used for GSA-24v3-0_A2 and GDA-8v1-0_A1.
Basic quality control and extraction of single nucleotide polymorphisms (SNPs) were done for the arrays (plink1.9 --geno 0.05 --maf 0.01 --hwe 0.0000001 --snps-only just-acgt) were done before alignment with reference panel 1000_Genomes_phase3_v5. Conversion to vcf files was done with PLINK 1.9. Sorting and strand alignment were done with bcftools (version 1.6) and java script (conform-gt.24May16.cee.jar, source https://faculty.washington.edu/browning/conform-gt/conform-gt.24May16.cee.jar, accessed in February 2024), respectively. Imputation using Minimac4 v1.7.4 was done on the Michigan Imputation Server (https://imputationserver.sph.umich.edu/index.html#!, accessed in February 2024)26,27. The imputation settings are detailed in Supplementary Data 3.
Low-coverage genome sequencing (lc-WGS)
Lc-WGS was done on NovaSeq using Novaseq 6000 S4 reagent with read length of 2 × 150 bp, with 94 samples per Flowcell/4 lanes. HapMap cell lines (GM18592 and GM18609) were included in each lane as controls. Briefly, 300–400 ng of genomic DNA was fragmented using Covaris S2 Focused-ultrasonicator with microTUBE AFA Fiber Snap-Cap (Part No. PN 520045, Covaris) based on the 50-µl sonication protocol, to generate peak size of 400-bp fragments. Fragmented samples were library prepared using NEBNext® Ultra™ II DNA Library Prep Kit for Illumina® (Part No. E7645L, NEB) and NEBNext® Multiplex Oligos for Illumina® 96 Unique Dual Index Primer Pairs (Part No. E6440, NEB). Purifications of samples when required were done using AMPure XP Beads for DNA Cleanup (Part No. A63882, Beckman Coulter) with a sample-to-beads ratio of 1:1. PCR enrichment with 7 cycles was performed using BioRad T100. PCR-enriched library products were quantified using the Agilent TapeStation 4200 system and DNA1000 ScreenTape assay (Part No. 5067-5583 and 5067-5582, Agilent Technologies). Libraries were quantified by qPCR using the KAPA library quantification kit (Part No. KK4854–07960298001, Roche) using a Roche Lightcycler 480 II real-time PCR system. Samples were normalized and pooled by equal volume before submission for sequencing.
The raw sequencing data generated from lc-WGS were processed using the DRAGEN germline workflow v3.7.8 (https://developer.illumina.com/dragen/dragen-popgen).
Sequencing reads were aligned to the GRCh38 reference genome. Imputation of lc-WGS data was done using the 1000 Genome Projects Phase 3 reference panel (same as for the arrays, downloaded from http://ftp.1000genomes.ebi.ac.uk/vol1/ftp/data_collections/1000G_2504_high_coverage/working/20201028_3202_phased/).
Using the 1000G phased reference panel, we first define chromosomal chunks using GLIMPSE2_chunk with a minimal window size of 6 Mb using the sequential chunk model. We downloaded the b38 genetic map for GLIMPSE2_chunk from https://github.com/odelaneau/shapeit4/blob/master/maps/genetic_maps.b38.tar.gz. Next, we convert the reference panel into the binary format using GLIMPSE2_split_reference using the default parameters. We used the single-sample variant calls from Dragen v3.7.8 for the lc-WGS imputation. GLIMPSE2_phase was used to impute genotypes for each imputation region as defined by the binary imputation panel using the default parameters. The imputed VCFs were index using bcftools prior to GLIMPSE2_ligate. Lastly, we use GLIMPSE2_ligate with the default parameters to ligate all chromosomal chunks together.
Supplementary Data 4 presents the proportion of variants that passed the quality checks. Supplementary Data 5 shows the coverage information for each sample. The average alignment coverage over the whole genome ranged from 2.00 to 13.05 for saliva samples, with a mean of 6.91 (standard deviation [SD] of 2.76); and 9.44 to 18.12 for cell lines, with a mean of 13.10 (SD of 3.19).
Polygenic risk score (PRS313) calculation
PRS313 was developed using genotype data from 94,075 invasive breast cancer cases and 75,017 controls of European ancestry from 69 BCAC studies, genotyped with either the iCOGS or OncoArray arrays20. Imputation was carried out separately for the iCOGS and OncoArray datasets using the Phase 3 (October 2014) release of the 1000 Genomes Project as the reference. Among the 313 variants, 55 were genotyped on iCOGS and 113 on OncoArray.
The dataset was randomly divided into training and validation sets; the validation set comprised ~10% of OncoArray samples after excluding studies of bilateral breast cancer, studies or sub-studies oversampling for family history, and individuals with in situ disease or unknown ER status. The best-performing PRSs were further evaluated in an independent test dataset of 11,428 cases and 18,323 controls from ten cohort-based studies (all OncoArray-genotyped). External validation was conducted in the UK Biobank, including 190,040 women of European ancestry without prior cancer or mastectomy at recruitment. Incident invasive breast cancers (n = 3215) were ascertained through cancer registries over 1,381,019 person-years of follow-up, with follow-up censored at risk-reducing mastectomy, cancer diagnosis, death, or January 15, 2017.
PRS313 was calculated as the weighted sum of effect alleles from 313 breast cancer variants, as previously described (Supplementary Data 6)20. This was performed with PLINK 1.9 using the “scoresum” option28. Calculations were performed using various subsets of variants at different imputation quality thresholds (rsq ≥0, > 0.3, > 0.8, and > 0.9). Lc-WGS used a different method for imputation (GLIMPSE2), where changing the imputation quality will not change the subset of variants consistently across all individuals.
Statistics and reproducibility
Assessing agreement, correlation, linear predictions, and high-risk classification in saliva samples
Fleiss’s Kappa statistic was used to assess the agreement in variant calling between the different platforms. For indels, Kappa was calculated between each platform and the results from Sanger sequencing. We assessed the correlations and linear predictions of PRS313 calculated from different platforms to evaluate the need for mean correction.
Risk threshold selection
In our hypothetical scenario, not representative of the general population, we compared high-risk classification differences between PRS313 derived from the different platforms. The proportion of high-risk individuals identified depends on the mean and SD of the PRS31322. The threshold for high risk was determined based on the PRS313 distribution in non-breast cancer controls, which followed a Gaussian distribution with a mean 0.130 and SD of 0.565)8. The 80th percentile of the PRS313 distribution in these controls was ~0.6. Hence, in our analyses, individuals with PRS313 (scoresum) greater than 0.6 were classified as high-risk.
Mean correction
To align our PRS313 distribution with a validated reference and to improve interpretability, we applied mean correction. Specifically, we adjusted the mean of the PRS313 values in our study population to match the reported mean of 0.130 from a previously published study of a Singaporean population with similar East Asian ancestry8. This approach assumes that the Singaporean dataset, drawn from a population with similar East Asian ancestry, provides an appropriate reference. Mean correction is not a universal requirement for PRS313 interpretation, but rather a dataset-specific correction. In cohorts where the PRS313 distribution closely matches that of the reference population, mean correction may not be needed.
Variations in scores near the thresholds potentially leading to reclassification
For each individual, 17 PRS313 values were generated; 16 from four genotyping arrays (OncoArray, GSA, GDA, ThermoFisher) using four imputation filters (rsq ≥0, ≥0.3, ≥0.8, ≥0.9), and one from lc-WGS. We quantified the within-individual variation as the difference between the maximum and minimum across the 17 scores for each individual. The OncoArray score without filtering (rsq≥0) was the reference “true” PRS313. Other scores were compared against this to determine concordance in high-/low-risk classification, stratified by percentile of the “true” score (<75th, 75th–84th, ≥85th). While the point 80th percentile (score = 0.606) was taken as the high-risk threshold, we expect natural variation to result in misclassification within the interval around this point. Here, we selected a ±5 percentile interval.
Subset analysis excluding indels
This subset excludes indels to focus on SNVs, which are more consistently genotyped and imputed across platforms.
Subset analysis restricting PRS313 to identical variants found across all platforms
As different variants contribute different effect sizes to the PRS313. For a comparable assessment of post-imputation PRS313 linear model predictions and concordance in high-risk individuals identified, we restricted the variants used in the PRS313 calculation to be the same across all platforms. Analysis was repeated across four levels of imputation quality (rsq ≥0, >0.3, >0.8, and >0.9).
Subset analysis restricting PRS313 to identical directly genotyped variants found across all arrays
We further restricted the dataset to 45 variants directly typed in all four arrays. In this case, the threshold for high risk was selected to be the average of the PRS45 at the 80th centile of the four arrays.
Subset analysis restricting to Chinese females
Applying PRS313 parameters derived from European populations to non-European samples can lead to a systematic shift in PRS313 distribution due to differences in allele frequencies and linkage disequilibrium. To account for this expected systematic shift, we included an additional analysis using mean-corrected PRS313. However, our study population comprises multiple ethnic groups, which may result in variability in PRS313 distribution. To minimize variability attributable to population heterogeneity, we restricted the analytical sample to a single ethnicity. Given the limited sample size (n = 92), we selected the largest ethnic-gender subgroup, Chinese females (n = 47). We evaluated correlations and linear predictions of PRS313 across different genotyping platforms to assess the need for mean correction. Between-platform agreement in risk classification was quantified using Fleiss’s Kappa statistic. Mean correction was performed by aligning the mean PRS313 score with that of the reference population (0.158), corresponding to the mean of Chinese controls reported previously8.
The GRCh38 reference genome was used for all analyses. Statistical analysis was done using R version 4.2.2. All reported p-values are two-sided.
Ethics approval and consent to participate
This study was approved by the A*STAR ethics board (reference number: 2022-062, approval date: 14 September 2022).
Results
Characteristics of 92 individuals
A total of 92 participants donated saliva samples. The median age was 32 years (interquartile range [IQR] of 26–38 years), 71% (n = 65) were females, 68% (n = 63) identified themselves as Chinese, 22% (n = 20) identified as Indian and 10% (n = 9) identified as Malay (Supplementary Data 7).
Directly typed variants
Among 2,734,610 unique variants typed across the different arrays, 28,646 (1%) were common across all four (Supplementary Fig. 1). Less than 50% of the PRS313 variants were present on any of the Infinium arrays: 64 (20%) on GSA, 99 (32%) on OncoArray, and 82 (26%) on GDA (Supplementary Fig. 2). All variants found were SNVs. Among the 313 variants, 259 (83%) were successfully designed on ThermoFisher, including 235 SNVs and 24 indels.
PRS313 using cell lines
Supplementary Fig. 3 shows variations in calculated PRS313 (y-axis) for 1–4 repeats per cell line. Four replicates per cell line were performed for lc-WGS (Repeats 1–4). Supplementary Fig. 2 shows the number of genotyped/imputed SNVs/indels at selected imputation quality levels (rsq ≥0, >0.3, >0.8, or >0.9). Post-imputation, between 86% and 91% of the 313 variants were available for analysis (rsq≥0). Specifically, 269 (86%) variants were available for GDA, 270 (86%) for GSA and OncoArray, 285 (91%) for ThermoFisher, and 300 (96%, by GLIMPSE2) for lc-WGS (Supplementary Fig. 2). lc-WGS yielded identical PRS313 values (to four decimal places) across four repeats. ThermoFisher’s PRS313, primarily based on directly genotyped variants (91%), was systematically lower than that from Infinium arrays or lc-WGS when using the same imputation quality threshold. PRS313 results from Infinium arrays were similar when all variants were included (rsq≥0) but diverged when restricted to high imputation quality variants, likely due to differences in variant subsets (Supplementary Fig. 3).
Sanger sequencing for indels
Sanger sequencing was performed for 87-89 individuals (Supplementary Data 8, Fig. 1). Primers could not be designed successfully for (chr:position:A1:A2) chr1:168201814:C:CA, chr4:125831837:AAT:A, chr16:3958541:C:CAAAAA, and chr19:19406245:CGGGCG:C (Supplementary Data 1). However, chr1:168201814:C:CA was typed on ThermoFisher. chr1:168201814:C:CA and chr19:19406245:CGGGCG:C were available in the post-imputation lc-WGS data. Four indels were not present/ imputed on all platforms (chr1:145830809:CT:C, chr2:217091173:G:GA, chr4:83448971:TA:TAA, chr22:38187308:AAAAGAAAG:AAAAG). We did not observe variations in 17/44 indels that was Sanger sequenced, hence pairwise agreement with other platforms cannot be calculated (NA).
Of the 48 indels, primers were designed successfully for 44. Primer design failed for (chr:position:A1:A2) chr1:168201814:C:CA, chr4:125831837:AAT:A, chr16:3958541:C:CAAAAA, and chr19:19406245:CGGGCG:C. Four additional indels were not present/imputed on all platforms (chr1:145830809:CT:C, chr2:217091173:G:GA, chr4:83448971:TA:TAA, chr22:38187308:AAAAGAAAG:AAAAG). Pairwise agreement (Kappa) of indels between Sanger and platforms (ThermoFisher, lc-WGS, GSA, OncoArray and GDA) is presented. We did not observe variations in 17/44 indels that were Sanger sequenced, hence pairwise agreement with other platforms cannot be calculated (NA). No indels were genotyped on the Infinium arrays, but 18 indels (41% of the 44 indels) were imputed. Agreement between Sanger sequencing and imputed Infinium variants ranged between 0.515 (chr8:127201316:C:CA on GSA) to 1. Of the 30 indels imputed in lc-WGS (MAF > 1%), 7 indels had Kappa < 0.5 and Kappa was not calculated for 8 indels (i.e., no variation in Sanger).
No indels were genotyped on the Infinium arrays, but 18 indels (41% of the 44 indels) were imputed (rsq ≥ 0). Agreement between Sanger sequencing and imputed Infinium variants ranged between 0.515 (chr8:127201316:C:CA on GSA) to 1 (Supplementary Data 8). Of the 30 indels imputed in lc-WGS (MAF > 1%), 7 indels had Kappa < 0.5 and Kappa was not calculated for 8 indels (i.e., no variation in Sanger) (Supplementary Data 8).
Concordance of variants across platforms
Among the PRS313 variants (rsq ≥ 0) in saliva samples, 266 (85%) were genotyped or imputed across all platforms (arrays and lc-WGS). Almost perfect agreement (Kappa > 0.8)29 between all platforms was observed for 78% (n = 206) of these 266 variants (Supplementary Data 9). However, 15 SNV showed slight to no agreement (Kappa ≤ 0.2). The median concordance (Fleiss’s Kappa) between all platforms was 0.962 [IQR: 0.860–0.993]. Between arrays, it was 0.972 [IQR: 0.846–1]. Excluding ThermoFisher, median concordance between arrays increased to 0.984 [IQR: 0.885 to 1] (Supplementary Data 9).
Effect of imputation quality and variant frequency on concordance of variants
Fleiss’s Kappa (using all 5 platforms where available) was calculated for 273 variants (87% of 313). Seven variants were not typed or imputed on any platform. For nine variants where Fleiss’s Kappa could not be calculated but more than one platform had typed or imputed the variant, the MAF was 0 across all available platforms. Variants were observed only on ThermoFisher (n = 3) or lc-WGS (n = 21). For ThermoFisher, the MAF was 0 for all three variants, whereas for lc-WGS the MAF ranged from 0 to 0.480, with a median of 0.150. Supplementary Fig. 4 shows the platform-specific MAF by Fleiss’s Kappa, for the remaining 273 variants (i.e., those imputed on one or more arrays). More variants with low MAF tended to be associated with Kappa ≤ 0.5.
Forty-five variants were typed on all arrays, while 28 variants were not typed or imputed on any array. Among the 16 variants that were not typed by all arrays or imputed by any array, 14 were typed only by ThermoFisher, one by both ThermoFisher and OncoArray, and one by all arrays except OncoArray. Supplementary Fig. 5 shows the imputation quality by Fleiss’s Kappa (using all 5 platforms where available), for the remaining 224 variants (i.e., those imputed on one or more arrays). Variants with Kappa ≤ 0.5 were called from a range of imputation quality scores.
High PRS313 correlation across platforms, but different risk scores are obtained for the same individuals
Pairwise correlation between PRS313 derived from different platforms (rsq ≥ 0) ranged from 0.754 (lc-WGS ~ GSA) to 0.940 (GDA~OncoArray) (Fig. 2, Supplementary Data 10). PRS313 calculated from ThermoFisher (91% directly typed variants), could be predicted using PRS313 from the Infinium arrays and lc-WGS via linear models. However, PRS313 with a high proportion of imputed variants (Infinium arrays and lc-WGS) tended to overestimate PRS313. The GDA ~ GSA pair exhibited a slight shift (intercept = −0.013) in mean but maintained a slope close to 1. (Fig. 2). When restricting the PRS313 calculation to variants with higher imputation quality (rsq >0.3, >0.8, or >0.9), similar linear relationships were observed (Supplementary Fig. 6).
A Using lc-WGS to predict ThermoFisher (ground truth). B GDA (185/313 variants imputed) prediction of the ground truth (ThermoFisher). C GDA PRS313 (185/313 variants imputed) prediction of OncoArray PRS313 (171/313 variants imputed). The intercept and slope with respective 95% confidence intervals (denoted by blue line), and correlation (r-square) are reported.
The chosen platform affects who is identified as high risk, especially those not at the extremes of the risk spectrum
PRS313 variability across all platforms and imputation quality thresholds ranged from 0.464 to 0.565 (SD) (Fig. 3, Supplementary Data 11). Without mean correction, at rsq ≥ 0, 42 (46% of 92) were high-risk by any platform, with 4 (4%) identified as high-risk by all five platforms. Without mean correction, substantial agreement (Fleiss’s Kappa: 0.61–0.80) among arrays and lc-WGS was observed when the classification of high-risk was performed without the restriction of imputation quality (rsq ≥ 0). Moderate agreement (Fleiss’s Kappa: 0.41–0.60) was observed when imputation quality was implemented (lowest Fleiss’s Kappa rsq>0.9, infinium = 0.381 to highest Fleiss’s Kapparsq>0.8, infinium = 0.559) (Supplementary Data 12).
The reference mean value of 0.130 (based on the PRS313 distribution in non-breast cancer controls of a previously published dataset) and the 80th percentile with the corresponding score of ~0.6 are indicated.
After mean correction (centering the mean to that of the reference population of 0.130), a total of 26 (28% of 92) unique individuals were high-risk by any platform, with 7 (identified as high-risk by all five platforms (Fig. 4). The high-risk proportion ranged between 12% and 21% across platforms and imputation quality, closer to the expected 20% based on the threshold selected from the reference population (Supplementary Data 11). Agreement improved (lowest Fleiss’s Kapparsq>0.9, infinium = 0.599 to highest Fleiss’s Kapparsq>0.8, infinium = 0.808) after mean correction (centering the mean to that of the reference population of 0.130) (Supplementary Data 12).
A PRS313 post-mean correction, allowing the number of variants used to differ with array/ lc-WGS, using the maximum number of variants with rsq ≥ 0, and B including only variants found across all platforms. Left: A matrix layout for intersections that has at least one individual. Set size denotes the number of high-risk individuals identified on each array. Dark circles indicate sets that are part of the intersection. Right: The same information on the overlaps of high-risk individuals identified depicted in a Venn diagram.
Within-individual variation resulted in greater misclassification near the chosen high-risk threshold
Substantial variation was observed in PRS313 derived from all available variants without mean correction, with within-individual values ranging from 0.764 to 1.482. After applying mean correction to each score, within-individual variation was reduced (minimum 0.181, maximum 0.970). Before mean correction, 63 individuals were classified into the <75th percentile group (score < 0.511), 14 into the 75th–84th percentile group (score 0.511–0.714), and 15 into the ≥85th percentile group (score ≥ 0.715) (Supplementary Data 13). As expected, the proportion correctly classified was higher for individuals further from the threshold (Supplementary Data 14). The mean correct classification rates were 92, 56, and 82% for the <75th, 75th–84th, and ≥85th percentile groups, respectively. With mean correction applied to both the “true” and comparison scores, these rates improved to 99%, 78%, and 84%, respectively. Similar trends were observed when scores were based only on identical variants across all platforms.
Subset analysis excluding indels
Agreement of indels across platforms is poor (Fig. 1). We thus excluded indels and used 251 (GDA), 252 (GSA and OncoArray), 260 (ThermoFisher), and 256 (lc-WGS) SNVs (rsq ≥ 0) for PRS313 calculation (Supplementary Data 10). Removing indels resulted in fewer individuals identified as high-risk (13–18%, compared to 15–21% previously) (Supplementary Data 11). Correspondingly, agreement in the high-risk individuals identified by the different arrays was improved (Kappaall_arrays = 0.708, previously 0.668) (Supplementary Data 12). In contrast, among the Infinium arrays, the agreement in the high-risk individuals decreased (KappaInfinium = 0.697, previously 0.739; at rsq ≥ 0) (Supplementary Data 12).
Subset analysis restricting PRS313 to identical variants found across all platforms
There is a systematic difference between the breast cancer PRS313 derived from the commercially available OncoArray and ThermoFisher arrays (Supplementary Data 15). With identical variants, Infinium arrays (GSA, OncoArray, GDA) showed less over- or underestimation, regardless of the imputation quality threshold applied. Comparisons by pairs of arrays are presented in Supplementary Fig. 7.
PRS313 distributions among Infinium arrays were more similar than with ThermoFisher (Supplementary Fig. 8). Interestingly, the distributions were right-shifted when imputation quality threshold was set at rsq>0.3. Notably, 3 of the 4 variants with low imputation quality rsq≤0.3 had negative beta estimates with effect allele corresponding to the major allele (chr11_433617, beta = −0.0437; chr2_240449440, beta = −0.1232; chr20_11399194, beta = 0.0844; chr22_29155884, beta = −0.1716).
After mean correction, a total of 24 (26%) unique individuals were high-risk by any platform, with 46% (11 of 24) identified as high-risk by all five platforms (Fig. 4). This is larger than the 27% (7 of 26) when the number of variants was varied between the platforms. The high-risk proportion ranged between 11 and 20% across platforms and imputation quality (Supplementary Data 16). Agreement ranged between Fleiss’s Kapparsq>0.3, arrays = 0.675 to the highest Fleiss’s Kapparsq≥0, Infinium = 0.784) after mean correction (Supplementary Data 16).
Subset analysis restricting PRS313 to identical directly genotyped variants found across all arrays
PRS45 (rsq ≥ 0) was identical across platforms for each cell line. In the 92 saliva samples, there is no observable difference between the breast cancer PRS45 derived from the commercially available arrays (Supplementary Fig. 9). As expected with the fewer variants included the SD for PRS45 was lower than that of PRS313. Using OncoArray to predict ThermoFisher (simple linear regression), the fitted line (slope = 1, intercept = 0) falls on the line that indicates perfect prediction (i.e., x = y) (Supplementary Fig. 10). This was similarly observed for the GSA ~ GDA pair. For all other pairs, the intercept was <0.1 with a correlation >0.9.
Using the mean 80th percentile as the high-risk threshold (PRS45 > −0.124), the 80th percentile of PRS45 was −0.157 for GSA, −0.095 for OncoArray, −0.157 for GDA, and −0.088 for ThermoFisher.
Without mean correction, a total of 26 (28% of 92) unique individuals were high-risk by any platform, with 58% (15 of 26) identified as high-risk by all five platforms (Supplementary Fig. 11). Agreement among platforms was almost perfect; Fleiss’s Kappaarrays+lc-WGS = 0.821, Fleiss’s Kappaarrays = 0.854, Fleiss’s KappaInfinium = 0.847).
Subset analysis restricting to Chinese females
PRS313 distributions based on all available variants and identical variants across platforms are shown in Supplementary Fig. 12 and Supplementary Fig. 13, respectively. Patterns of platform-specific shifts were similar to those observed in the full sample of 92 individuals. Pairwise linear associations between breast cancer PRS313 scores across arrays are presented in Supplementary Fig. 14. Due to the small overall sample size, restricting the analysis to Chinese females further limited our ability to quantify any change in correlations between PRS313 scores from different platforms. Agreement in risk classification was not higher in this homogeneous subset compared with the full study population (Supplementary Data 17).
Discussion
In an ideal setting, PRS313 is calculated using directly genotyped variants for both the reference population and the population PRS313 is applied to, as this would provide the most accurate and reliable measurement for making personalized healthcare decisions. However, due to practical constraints such as cost and logistics, selective genotyping and imputation of missing genotypes are often performed30,31. While these methods help increase the number of available variants, they also introduce variability in the proportion of directly genotyped variants and imputation success rates. Our study highlights key issues and opportunities in the comparability and utility of PRS313 across different genotyping arrays and sequencing platforms. We observed variability in concordance between genotyped indels and Sanger sequencing. Although PRS313 values from different platforms are highly correlated, varying risk scores were obtained for the same individuals. Mean correction improved risk classification agreement, reducing variability in the proportion of high-risk individuals identified across platforms (from 4 to 45% pre-mean correction to 15–21% post-mean correction), aligning more closely with the expected 20% from a reference population. However, high-risk individuals identified across different platforms remained largely inconsistent.
Specifically, 269 (86%) variants were available for GDA, 270 (86%) for GSA and OncoArray, 285 (91%) for ThermoFisher, and 300 (96%, by GLIMPSE2). We first examined PRS313 generated from cell lines which experience fewer issues with variability in DNA quality. The availability of PRS313 variants post-imputation varied across platforms, with lc-WGS showing the highest proportion (96%) of available variants, followed by ThermoFisher (91%), GDA (86%), GSA (86%), and OncoArray (86%). Besides high variant availability, lc-WGS demonstrated excellent reproducibility, producing identical PRS313 values across four replicates. It is important to note that the discrepancy in variant availability will affect PRS313 calculation, particularly if variants with higher weightage for risk prediction are unavailable or of lower imputation quality on certain platforms.
When examining PRS313 in 92 real-world biological samples, a key issue we revealed is the inadequate capture of PRS313 indels using commercial off-the-shelf arrays. Our study found that none of the commercially available arrays included any of the 48 indels from PRS313. Only half of the indels (24/48) passed probe design on the custom ThermoFisher array, and 44 primers were successfully designed for Sanger sequencing. Even between the directly genotyped indels, there was considerable variability in agreement. On the other hand, the high agreement observed with Infinium arrays (where all indels were imputed) may reflect the influence of the common imputation reference panel, rather than the inherent accuracy of array-based methods32,33. These difficulties highlight the importance of carefully considering the inclusion of indels, especially when PRS313 includes a substantial proportion of indels, as this may significantly affect the accuracy of breast cancer risk prediction.
The variability in PRS313 across different platforms has clinical implications for risk stratification. Although there is a strong correlation between platforms, the risk scores can differ, especially for individuals not at the extremes of the risk spectrum25. Agreement is better without restricting imputation quality, presumably because more data points are included. However, inaccuracies may result due to less reliable variants. In contrast, restricting to high-quality variants may improve prediction accuracy but result in fewer variants, thus reducing overall agreement in risk classification. While mean correction improves consistency and reduces discrepancies by aligning risk classifications closer to the reference population, the results still indicate that 28% of individuals were identified as high-risk by at least one platform, and only 8% were identified by all five platforms. This suggests that while mean correction helps, platform-specific PRS313 variability remains a challenge. To the layperson, this result matters because it shows that the test used to estimate a woman’s breast cancer risk can give different results depending on which method is used.
This study is the first to examine the consistency of PRS313 called across platforms, especially indels that were previously mostly imputed and not genotyped. The combination of Sanger sequencing, three commercial Illumina Infinium arrays, one custom ThermoFisher array, and lc-WGS provides a diverse and comprehensive selection of technologies for genotyping variants for PRS313 calculation. Efforts were made to design all PRS313 variants on the ThermoFisher array; Sanger sequencing was performed to validate all indels.
A limitation of our study is the reliance on the 1000 Genomes Project reference panel for imputation. While widely used, it may be suboptimal for the Asian population included here, and imputation uncertainty could propagate into PRS estimates, particularly for indels and population-specific variants. Use of more ancestry-matched reference panels may improve accuracy and should be considered in future studies12,13. While the study evaluates agreement across platforms, it does not address how closely any of these platforms reflect the underlying biological truth. Without a definitive reference (e.g., high coverage WGS from blood-matched DNA), it remains unclear whether high concordance simply reflects technical consistency rather than biological accuracy. Another consideration in this study is the use of saliva-derived DNA, which can exhibit greater variability in yield, purity, and integrity compared to blood-derived DNA. Such variability has the potential to influence probe performance, genotyping fidelity, and downstream analyses such as imputation. However, in real-world implementation, saliva offers practical advantages, particularly in studies involving healthy individuals, where non-invasive collection can improve participation rates and overall sample accessibility. For these reasons, we selected saliva as the DNA source for this study. Future studies focused specifically on platform reproducibility, including library preparation variability, could incorporate multiple independent extractions and library preparations per sample.
Conclusion
In summary, our evaluation underscores the importance of assessing PRS within the context of technical, biological, and analytical variability. While PRS313 remains a valuable tool for breast cancer risk stratification, its translation into clinical practice requires rigorous validation, harmonization of analytical pipelines, and ongoing calibration against diverse reference populations.
Availability of data and material
The PRS313 values to support the analyses are available as Supplementary Data 18. Consent for the use of participants’ genetic data in future research was obtained. Due to the sensitive nature of the genotyping and low-coverage whole genome sequencing (lc-WGS) data, the individual-level datasets are not publicly available. Access will be granted to qualified researchers upon reasonable request and approval. Researchers interested in accessing the data should contact the Principal Investigator at lijm1@a-star.edu.sg. Requests will typically be reviewed and responded to within 4–6 weeks. Approved users must comply with a data use agreement, which includes the following restrictions: the data may only be used for the specific research purposes approved in the request, cannot be shared with third parties, must be stored securely, and must not be used to attempt to identify individual participants. Any publications or presentations resulting from the use of the data must acknowledge the source and comply with applicable ethical and legal guidelines.
References
Ayoub, A., Mchugh, J., Hayward, J., Rafi, I. & Qureshi, N. Polygenic risk scores: improving the prediction of future disease or added complexity? Br. J. Gen. Pr. 72, 396–398 (2022).
Wong, E. et al. The Singapore national precision medicine strategy. Nat. Genet 55, 178–186 (2023).
All of Us Research Program Genomics, I Genomic data in the All of US Research Program. Nature 627, 340–346 (2024).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Sud, A. et al. Realistic expectations are key to realising the benefits of polygenic scores. BMJ 380, e073149 (2023).
Kumuthini, J. et al. The clinical utility of polygenic risk scores in genomic medicine practices: a systematic review. Hum. Genet. 141, 1697–1704 (2022).
Hingorani, A. D. et al. Performance of polygenic risk scores in screening, prediction, and risk stratification: secondary analysis of data in the polygenic score Catalog. BMJ Med. 2, e000554 (2023).
Ho, P. J. et al. Overlap of high-risk individuals predicted by family history, and genetic and non-genetic breast cancer risk prediction models: implications for risk stratification. BMC Med. 20, 150 (2022).
Ho, P. J., Lim, E. H., Hartman, M., Wong, F. Y. & Li, J. Breast cancer risk stratification using genetic and non-genetic risk assessment tools for 246,142 women in the UK Biobank. Genet. Med. 25, 100917 (2023).
Carver, T. et al. CanRisk Tool-A Web interface for the prediction of breast and ovarian cancer risk and the likelihood of carrying genetic pathogenic variants. Cancer Epidemiol. Biomark. Prev. 30, 469–473 (2021).
Xiang, R. et al. Recent advances in polygenic scores: translation, equitability, methods and FAIR tools. Genome Med. 16, 33 (2024).
Liu, C. et al. Generalizability of polygenic risk scores for breast cancer among women with European, African, and Latinx Ancestry. JAMA Netw. Open 4, e2119084 (2021).
Yiangou, K. et al. Polygenic score distribution differences across European ancestry populations: implications for breast cancer risk prediction. Breast Cancer Res. 26, 189 (2024).
Hou, K. et al. Calibrated prediction intervals for polygenic scores across diverse contexts. Nat. Genet. 56, 1386–1396 (2024).
Roberts, E. et al. Breast cancer polygenic risk scores derived in White European populations are not calibrated for women of Ashkenazi Jewish descent. Genet. Med. 25, 100846 (2023).
Sanger, F., Nicklen, S. & Coulson, A. R. DNA sequencing with chain-terminating inhibitors. Proc. Natl. Acad. Sci. USA 74, 5463–5467 (1977).
Lamy, P., Grove, J. & Wiuf, C. A review of software for microarray genotyping. Hum. Genom. 5, 304–309 (2011).
Verlouw, J. A. M. et al. A comparison of genotyping arrays. Eur. J. Hum. Genet. 29, 1611–1624 (2021).
Nguyen, D. T. et al. A comprehensive evaluation of polygenic score and genotype imputation performances of human SNP arrays in diverse populations. Sci. Rep. 12, 17556 (2022).
Mavaddat, N. et al. Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am. J. Hum. Genet. 104, 21–34 (2019).
Roberts, E., Howell, S. & Evans, D. G. Polygenic risk scores and breast cancer risk prediction. Breast 67, 71–77 (2023).
Ho, W. K. et al. Polygenic risk scores for prediction of breast cancer risk in Asian populations. Genet. Med. 24, 586–600 (2022).
Brooks, J. D. et al. Personalized risk assessment for prevention and early detection of breast cancer: integration and implementation (PERSPECTIVE I&I). J. Pers. Med. 11, 511 (2021).
Liu, J. et al. BREAst screening Tailored for HEr (BREATHE)-A study protocol on personalised risk-based breast cancer screening programme. PLoS ONE 17, e0265965 (2022).
Chen, S. F. et al. Genotype imputation and variability in polygenic risk score estimation. Genome Med. 12, 100 (2020).
Michigan Imputation Server 2. Vol. 2024.
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Landis, J. R. & Koch, G. G. The measurement of observer agreement for categorical data. Biometrics 33, 159–174 (1977).
Montpetit, A. et al. An evaluation of the performance of tag SNPs derived from HapMap in a Caucasian population. PLoS Genet. 2, e27 (2006).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Duan, Q., Liu, E. Y., Croteau-Chonka, D. C., Mohlke, K. L. & Li, Y. A comprehensive SNP and indel imputability database. Bioinformatics 29, 528–531 (2013).
Nguyen, T. V. et al. Empirical versus estimated accuracy of imputation: optimising filtering thresholds for sequence imputation. Genet. Sel. Evol. 56, 72 (2024).
Acknowledgements
This project is supported by A*STAR, Precision Health Research Singapore—Clinical Implementation Pilot (PRECISE BREATHE CIP) and JurongHealth Fund (JHF-20-RE-003), and Open Fund-Large Collaborative Grant (OFLCG23may-0031). This work was supported by the A*STAR Computational Resource Centre with its high-performance computing facilities.
Author information
Authors and Affiliations
Contributions
Conceptualization: J.L. Project administration: A.J.K., P.J.H., and K.A.K. Methodology: P.J.H., J.H.J.T., P.A.V.G., Z.L., R.D., W.K.H., and N.B. Software: P.A.V.G. and N.B. Formal analysis: P.J.H. Investigation: K.A.K., E.L., L.L.G., and D.Q.C. Resources: E.L., L.L.G., H.L.W., M.H., D.Q.C., and I.B.H.T. Data Curation: P.J.H., J.H.J.T., and Z.L. Writing—original draft: P.J.H., A.J.K., and J.L. Writing—review and editing: All authors. Supervision: J.L., R.D., and N.B. Funding acquisition: J.L., M.H., and N.B.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Declaration of AI and AI-assisted technologies in the writing process During the preparation of this work the authors used ChatGPT and Grammarly to improve readability and language. After using this tool/service, the author reviewed and edited the content as needed and take full responsibility for the content of the publication.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ho, P.J., Khng, A.J., Tan, J.H.J. et al. Genomic platform specific polygenic risk scores impact breast cancer risk stratification. Commun Med 6, 41 (2026). https://doi.org/10.1038/s43856-025-01298-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01298-4