Abstract
Background
The transcription factor Hypoxia-Inducible Factor 2α (HIF2α) plays a crucial role in cancer cell adaptation to hypoxic conditions, particularly in clear cell renal cell carcinoma, promoting tumor growth and angiogenesis. Targeting HIF2α through pharmacologic inhibition offers a promising therapeutic strategy for HIF2α-driven cancers.
Methods
An in silico docking study using 10,000 drug-like compounds was performed using the previously solved X-ray crystal structure of HIF2α. Select compounds predicted to bind to the Per-Arnt-Sim-A (PAS-A) and PAS-B domains of HIF2α were further evaluated for biological activity in clear cell renal cell carcinoma and normal kidney cell lines. Biochemical and cell-based assays were performed to define the mechanism of action for a lead compound.
Results
Here, we identify Compound-c2 as a selective HIF2α inhibitor that binds to the PAS-B domain of HIF2α. Notably, Compound-c2 disrupts the interaction between HIF2α and the molecular chaperone Hsp70, leading to proteasomal degradation of HIF2α and the induction of apoptosis in ccRCC.
Conclusions
The distinctive inhibitory mechanism of the HIF2α inhibitor identified here, Compound-c2, sets it apart from previous HIF2α antagonists. This positions Compound-c2 as a promising alternative with potential applications in addressing drug resistance, providing a unique approach to inhibit HIF2α-related processes.

Plain language summary
Cancer occurs when cells grow out of control. In the most common type of kidney cancer, this is caused by abnormal activity of a specific part of cellular machinery. In this study, we used computer simulations to find drugs that can stop this growth-causing activity. By testing drugs identified from these simulations, we found one drug that kills kidney cancer cells, but not normal cells. We show that this drug works differently than another drug used to kill kidney cancer. These results show that in the future, this drug could be used to treat kidney cancer, particularly in patients where other available drugs have not worked.
Similar content being viewed by others
Introduction
Hypoxia-Inducible Factor 2 alpha (HIF2α) is a transcription factor that plays an essential role in mediating cellular responses to hypoxia1,2,3,4. HIF2α structure and function have attracted significant interest, as misregulation of the physiological and pathological processes influenced by HIF2α are implicated in the development and progression of several forms of cancer, including renal cancer5,6,7,8. HIF2α belongs to the basic helix-loop-helix (bHLH) Per-Arnt-Sim (PAS) family of transcription factors9. The protein consists of several functional domains, including the N-terminal bHLH domain that enables DNA binding, two PAS domains (PAS-A and PAS-B) involved in heterodimerization with its partner aryl hydrocarbon receptor nuclear translocator (ARNT), and a C-terminal transactivation domain that facilitates transcriptional activity. Notably, the PAS-B domain has been identified as a regulatory site susceptible to ligand interactions, offering a potential target for therapeutic interventions4,10.
Under normoxic conditions, HIF2α is hydroxylated by prolyl hydroxylase enzymes, marking it for proteasomal degradation11,12. However, during hypoxia, these hydroxylations are impaired, allowing HIF2α to evade degradation11,12. Stabilized HIF2α forms a heterodimer with ARNT and translocates to the nucleus, where it binds to hypoxia-responsive elements (HREs) in the promoters of genes involved in angiogenesis, erythropoiesis, glucose metabolism, and cell survival13. Tumor cells depend on these genes to ensure their growth and survival in the hostile tumor microenvironment14. HIF2α also contributes to the epithelial-mesenchymal transition (EMT), a process facilitating metastasis by enhancing cell mobility and invasiveness.
Clear cell renal cell carcinoma (ccRCC) serves as a great example of HIF2α playing a role in cancer progression. Mutations in the tumor suppressor Von Hippel-Lindau (VHL) gene are prevalent in ccRCC. Resultant mutant VHL protein is unable to exert the regulatory function of targeting HIF2α for degradation, resulting in the accumulation of HIF2α. This leads to the aberrant expression of HIF2α target genes, fueling tumor development1,15. A number of pharmaceutical agents have been developed for targeting HIF2α to treat cancer. For example, Belzutifan (MK-6482 also known as WELIREG, Merck), is a U.S. Food and Drug Administration (FDA) approved HIF2α inhibitor16. It disrupts HIF2α activity by binding to its PAS-B domain, inhibiting downstream gene expression crucial for tumor growth5,6,7. Belzutifan is intended for adult patients with VHL disease who need treatment for associated conditions like renal cell carcinoma (RCC), central nervous system (CNS) hemangioblastomas, or pancreatic neuroendocrine tumors (pNET) that do not necessitate immediate surgery7, and has recently be shown to be superior to mTOR inhibition in patients with metastatic ccRCC who failed immune checkpoint and anti-angiogenic therapies17. Therefore, HIF2α inhibition through pharmacologic small molecules offers an attractive therapeutic approach for HIF2α-driven cancers.
In this study, we perform an in silico screen18,19 using the PAS-B domain of HIF2α. We identify Compound-c2 as an antagonist of HIF2α and demonstrate that it specifically binds to and disrupts HIF2α interaction with the molecular chaperone Heat shock protein-70 (Hsp70). This consequently leads to the proteasomal degradation of HIF2α and induces apoptosis in ccRCC.
Methods
Mammalian cell culture
Cultured human embryonic kidney (HEK293) (ATCC, Cat# CRL-1573; RRID:CVCL_0045), (Human, unknown) and HK-2 (ATCC, Cat# CRL-2190; RRID:CVCL_0302), (Human, Male) cells were grown in Dulbecco’s Modified Eagle Medium (DMEM, Sigma-Aldrich), 786-O cells (ATCC Cat# CRL-1932; RRID:CVCL_1051), (Human, Male) OS-RC-2 (RIKEN Cell Bank, Tsukuba, Japan, RRID:CVCL_1626) cells (Human, Male), and TUHR14TKB (RIKEN Cell Bank, Tsukuba, Japan, RRID:CVCL_5953) cells (Human, Male) in Roswell Park Memorial Institute (RPMI)1640 Medium (Sigma-Aldrich), A498 cells (ATCC Cat# CRL-44; RRID:CVCL_1056), (Human, Female) in Minimum Essential Medium (MEM, Sigma-Aldrich), Caki-1 (ATCC Cat# HTB-46; RRID:CVCL_0234), (Human, Male) and Caki-2 cells (ATCC Cat# HTB-47; RRID:CVCL_0235), (Human, Male) in McCoy’s 5 A Medium (Sigma-Aldrich) all supplemented with 10% fetal bovine serum (FBS, Sigma-Aldrich). Cells were maintained in a CellQ incubator (Panasonic Healthcare) at 37 °C in an atmosphere containing 5% CO2.
Generation of Belzutifan-resistant 786-O cell line
786-O cells were treated with 10 µM Belzutifan (MedChemExpress Cat# HY-125840, CAS: 1672668-24-4) in Roswell Park Memorial Institute (RPMI)1640 Medium (Sigma-Aldrich). This allowed approximately 10% survival, and the cells required 120 h before approximately 70% confluence was achieved. Once at 70% confluence, the cells were split and treated with 10 µM Belzutifan again. This process was repeated until the cells appeared healthy and grew at a similar rate to 786-O cells. Belzutifan was removed from the cells for at least 24 h before additional drug treatments and/or protein extraction.
Plasmids
For mammalian expression HA-HIF2α -pcDNA3 was a gift from William Kaelin (Addgene plasmid # 18950; RRID:Addgene_18950). Point mutations F244A, H248A, M252A, F254A, Y281A, Y307A, M309A, T321A, and N341A, S246A, A277E, F280A, I337A, and C339A were created by site-directed mutagenesis (see Supplemental Data 1), mCherry-HIF2α was subcloned using HindIII and BamHI to insert mCherry into the HA-HIF2α-pcDNA3 backbone (see Supplemental Data 1) and confirmed by DNA sequencing. The pcDNA3-FLAG-Hsp70 and sub-domains (NTB and SBD) were cloned using primers in Supplemental Data 1. pcDNA5/FRT/TO GFP HSPA1A was a gift from Harm Kampinga (Addgene plasmid # 19483; RRID:Addgene_19483)20.
Cell transfection and treatment
Cultured cells were split and then transfected the following day when about 40% confluent with each construct using Mirus TransIT-2020 (MirusBio) according to manufacturer’s protocol. Cells were transfected using 2 µg (HEK293) or 4 µg (786-O) of plasmids for transient transfection. Cells were incubated at 37 °C and then extracted or collected for analysis after 24 h (HEK293) or 48 h (786-O). Short interfering RNA (siRNA) scramble control and EPAS1 (HIF2α) targeting duplexes were purchased from OriGene (SKU: SR320042A, SR320042B, and SR320042C). Indicated cells were transiently transfected with the siRNA using Mirus TransIT-2020. For HIF2α knock-down, either 30 nM of control siRNA or 10 nM of each HIF2α siRNA duplex (A, B and C) were mixed prior to transfection. Cells were incubated at 37 °C for 72 h in the transfection media, then harvested for protein extraction. Transfection efficiency was monitored via protein levels of the appropriate epitope tag attached to the protein of interest. For every experiment involving transient transfections, Western blots of input levels of each protein epitope tag have been provided.
Protein extraction, immunoprecipitation, and immunoblotting
Protein extraction from mammalian cells was carried out using methods previously described21,22. Cell lysates were quantified using 1X Bradford reagent (BioRad). For Western blotting, 25 µg protein lysate was used. For immunoprecipitation, 500 µg cell lysates were incubated with 30 µl anti-FLAG antibody conjugated beads (Sigma-Aldrich, Cat# A2220) at 4 °C for 2h. Endogenous IPs were achieved by incubating 1500 µg lysate with 1.5 µl anti-HIF2α antibody (Cell Signaling Technology, Cat#7096; RRID: AB_10898028) overnight followed by 25 µl protein G agarose (ThermoFisher Scientific, Cat# 15-920-010) at 4 °C for 2h. Immunopellets were washed 4 times with fresh lysis buffer (20 mM Tris-HCl (pH 7.4), 100 mM NaCl, 1 mM MgCl2, 0.1% NP40, protease inhibitor cocktail (Roche, Cat#11873580001), and PhosSTOP (Roche, 4906845001)) and eluted in 5x Laemmli buffer. Precipitated proteins were separated by SDS-PAGE and transferred to nitrocellulose membranes. Co-immunoprecipitated proteins or proteins from cell lysate were detected with antibodies recognizing: 1:4,000 Rabbit anti-FLAG tag (ThermoFisher Scientific, Cat# PA1-984B; RRID:AB_347227), 1:4,000 Rabbit anti-HA tag (C29F4) (Cell Signaling Technology, Cat# 3724; RRID:AB_1549585), 1:1,000 Rabbit anti-glucocorticoid receptor (Cell Signaling Technology, Cat# 7076; RRID: AB_330924), 1:8,000 Rabbit anti-Beta Actin (GeneTex, Cat# GTX637675, RRID: AB_3073746), 1:4,000 Mouse anti-GAPDH (1D4) (Enzo Life Sciences, Cat# ADI-CSA-335; RRID:AB_10617247), 1:1,000 Rabbit anti-HIF2α (Cell Signaling Technology, Cat#7096; RRID: AB_10898028), 1:1,000 Rabbit anti-HIF1α (Cell Signaling Technology, Cat#14179; RRID: AB_2622225), 1:4,000 Rabbit anti-HSP70 (StressMarq Biosciences, Cat#SPC-103; RRID:AB_2570584), 1:4,000 Rabbit anti-AKT (Cell Signaling Technology, Cat#9272; RRID: AB_329827), 1:1,000 Rabbit anti-phospho-AKT (S473) (Cell Signaling Technology, Cat#4060; RRID: AB_2315049), 1:1,000 Rabbit anti-cleaved caspase-3 (Cell Signaling Technology, Cat# 9664; RRID:AB_2070042), 1:1,000 Rabbit anti-cleaved caspase-7 (Cell Signaling Technology, Cat# 9491; RRID:AB_2068144), 1:1,000 Cleaved PARP (Cell Signaling Technology, Cat# 5625; RRID:AB_10699459). Secondary antibodies raised against mouse (Cell Signaling Technology, Cat# 7076; RRID:AB_330924) and rabbit (Cell Signaling Technology, Cat# 7074; RRID:AB_2099233) were used at 1:4,000 dilution.
FRET
Cultured cells were split and then transfected the following day when about 40% confluent with each construct using Mirus TransIT-2020 (MirusBio) according to manufacturer’s protocol. Cells were co-transfected using 1 µg of GFP-HSP70 and 4 µg of mCherry-HIF2α for transient transfection. Cells transfected with either mCherry-HIF2α or GFP-HSP70 alone were used as controls. Cells were incubated at 37 °C for 24 h prior to treatment with 30 µM Compound-c2 (4 h), 30 µM Compound-y (16 h), 30 µM Compound-z (16 h), 30 µM Belzutifan (2 h), or DMSO control. After treatment, growth media was removed and cells were washed twice with PBS. Cells were briefly trypsinized, then resuspended in PBS. Cells were pelleted by gentle centrifugation (2 min, 1200 rpm) and washed with PBS once more. Cells were then resuspended in 500 µL of PBS. Cells were plated in triplicate in a Nunc™ F96 MicroWell™ Black plate and fluorescence was read immediately on Tecan SPARK plate reader. GFP signal was detected by excitation and emission wavelength of 478 nm and 523 nm, respectively. mCherry signal was detected by excitation and emission wavelength of 585 nm and 630 nm, respectively. FRET signal was detected by excitation and emission wavelength of 478 nm and 610 nm, respectively. Excitation and emission bandwidth were 20 nm for all conditions. The average value of mCherry signal from cells expressing GFP-HSP70 alone was considered to background signal, and this value was subtracted from the signal values of all samples. The FRET signal was then normalized to mCherry-HIF2α expression by dividing FRET signal values for each sample by mCherry signal values for each sample. Corrected-FRET values were then normalized to the DMSO (NT) control. An unpaired two-tailed t-test was used to determine statistical significance between normalized FRET values of DMSO or Compound-c2, -y, -z, or Belzutifan treated cells.
Cell viability assay
Renal cancer cell lines 786-O, Caki-1, A498 cells as well as HEK293 and HK-2 cells were plated at 10,000 cells per well in 96-well plates. Cells were treated with different amounts of inhibitors and DMSO was used as control. After 24 h, cell viability assay was performed using the Quick Cell Proliferation Kit Plus using a reference wavelength of 650 nm (BioVision, Cat# K302-500; CAS# 150849-52-8) according to the manufacturer’s protocol. The absorbance at 450 nm was measured on a Tecan Infinite M200 Pro and proliferation rate was calculated.
In silico docking
Virtual screening was performed using MTiOpenScreen19 with the diverse chemical compound collection (Diverse-lib), which is curated by the Ressource Parisienne en BioInformatique Structurale (RPBS). The Diverse-lib library is derived from the PubChem BioAssay Database23 of ~3,574,000 molecules (names provided in pubchem SID). After removing the redundant molecules, the data set filtered to remove molecules with undesireable ADME properties, predicted toxic and Pan Assay Interference Compounds (PAINS), and clustered to maximize diversity. This led to a focused library of 99,288 diverse drug-like PubChem molecules. A library of 3D conformations of these structures was then generated using Frog224 keeping a maximum of one stereoisomer per compound without generating multiple ring conformations. The molecules were then protonated at pH 7 using the major macrospecies option of the ChemAxon (www.chemaxon.com) plugin to provide the Diverse-lib dataset.
The protein structure of the HIF2-ARNT complex was downloaded from the PDB (PDB:4ZQD9), and water molecules and the ligand were removed. The binding sites were then defined with key amino acid residues (Tyr307, Tyr281 and Phe254 for site 2 and Met108, Met172, Phe134 for site 5). The MTiOpenServer then uses Autodock Vina18 to perform rigid docking and score 10,000 molecules randomly chosen from the diverse-lib library. AutoDock Vina employs a gradient-based conformational search approach and defines the search space by a grid box defined by the box center coordinates and its dimensions of x, y and z. A grid box with dimensions of 75 Å on each side is used for the docking. In AutoDock Vina the grid resolution is internally assigned to 1 Å, the number of binding modes of 10 and exhaustiveness of 8. The grid center is automatically calculated based on the list of protein residues of the binding site provided by the user. The scoring of the generated docking poses and ranking of the ligands is based on the Vina empirical scoring function approximating the binding affinity in kcal/mol. The three best scored poses of the 500 top ranked compounds as ranked by AutoDock Vina is then retrieved. The 23 selected compounds had the best binding scores from the commercially available compounds in each data set, and these were then purchased and used for further evaluation.
Modeling of compound-z in site 2 of the HIF2-ARNT complex was accomplished using the same crystal structure from above, where water molecules were removed. Docking was then performed with Autodock Vina18 on the MTiAutodock server using the key amino acid residues (Tyr307, Tyr281 and Phe254) to define the binding pocket. AutoDock Vina employs a gradient-based conformational search approach and defines the search space by a grid box defined by the box center coordinates and its dimensions of x, y and z. A grid box with dimensions of 75 Å on each side is used for the docking. In AutoDock Vina the grid resolution is internally assigned to 1 Å, the number of binding modes of 10 and exhaustiveness of 8. The grid center is automatically calculated based on the list of protein residues of the binding site provided by the user. The scoring of the generated docking poses and ranking of the ligands is based on the Vina empirical scoring function approximating the binding affinity in kcal/mol. The ten best docking poses are then downloaded by the user, the docking pose with highest predicted binding affinity was used for the model.
RT-qPCR
RNA was extracted using RNA extraction mini-kit and quantified using NanoQuant (TECAN). RNA (1 µg) from each sample was reverse transcribed into cDNA using iScript cDNA synthesis kit (BioRad, Cat# 1708890) as previously described25. RT-qPCR analysis was performed using nucleic acid stain SYBR-GREEN (BioRad, Cat# 1725121) and specific qPCR oligonucleotide primers: VEGF1, NDRG2, CCND1, GLUT1, and ACTB control. VEGF1, NDRG2, CCND1, GLUT1 mRNA levels were normalized to ACTB control.
Primer Sequences
Sequences of oligonucleotides are provided in Supplemental Data 1.
VEGF-luciferase reporter
786-O cells were transiently transfected with VEGF-luciferase reporter for 24 h as previously described26. Cells were plated in a 96-well plate for 24 h followed by treatment with 30 µM of Compound-c2 for an additional 8 h. siRNA targeting HIF2α was used as a control. One-Glo Luciferase reagent (Promega, Cat# E6110) was added for 30 min and luminescence was read using Tecan SPARK. Data was normalized to DMSO control.
Synthesis of Biotin-Compound-c2
2-(3-Ethylureido)benzonitrile (2)
A mixture of 2-aminobenzonitrile 1 (5.0 g, 42.3 mmol) and ethyl isocyanate (3.35 mL, 42.3 mmol) were warmed to form a clear solution. The reaction mixture was stirred for 26 h under argon, resulting in the formation of a precipitate. The precipitate was recrystallized from MeOH to afford the urea product 2 (3.9 g, 48%) as a white fluffy solid. This compound has been previously reported. TLC Rf = 0.38 (30% EA/hexanes); 1H NMR (400 MHz, DMSO-d6) δ 8.43 (s, 1H), 8.01 (d, J = 8.4 Hz, 1H), 7.67 (d, J = 7.6 Hz, 1H), 7.57 (t, J = 8.0 Hz, 1H), 7.08 (t, J = 7.6 Hz, 1H), 6.95 (t, J = 4.7 Hz, 1H), 3.16-3.09 (m, 2H), 1.07 (t, J = 7.2 Hz, 3H);); 13C NMR (100 MHz, DMSO-d6) δ 154.3, 142.8, 133.8, 132.9, 122.1, 120.6, 117.0, 100.9, 34.0, 15.1.
3,4-Dihydro-4-imino-3-ethylquinazolin-2(1H)-one (3), (Compound-w)
The benzonitrile 2 (1.68 g, 8.87 mmol) was dissolved in EtOH (15 mL) with heating. Ammonium hydroxide (8.9 mL) was added and the reaction was refluxed for 1 h. The reaction mixture was cooled to rt and DI water (30 mL) was added. The precipitate was vacuum filtered, washed with DI water, and dried using an azeotrope of EtOH to yield the product 3 (1.48 g, 88%) as a white fluffy solid. This compound has been previously reported. TLC Rf = 0.31 (100% EA); 1H NMR (400 MHz, DMSO-d6) δ 10.68 (s, 1H), 8.85 (s, 1H), 8.05 (d, J = 8.0 Hz, 1H), 7.46 (t, J = 8.4 Hz, 1H), 7.09 (t, J = 7.6 Hz, 1H), 7.03 (d, J = 8.4 Hz, 1H), 4.06 (q, J = 6.18 Hz, 2H), 1.14 (t, J = 6.18 Hz, 3H); 13C NMR (100 MHz, DMSO-d6) δ 155.2, 149.8, 136.9, 132.7, 125.9, 122.0, 115.1, 113.0, 35.7, 12.3.
4-Bromo-N-[3-ethyl-2-oxo-2,3-dihydro-4(1H)quinazolinyliden]benzenecarboxamide (4), (Compound-y)
NaH (2 equiv.) was suspended in dry DMA and stirred for 15 min. A solution of quinazoline 3 (1 equiv.) dissolved in dry DMA (0.1 M) was added dropwise to the reaction, followed by addition of 4-bromobenzoyl chloride (1.5 equiv). The reaction was stirred for 24 h. Ethyl acetate (EA) and DI water were added and the layers were separated. The aqueous layer was extracted with EA (2x). The combined organic layers were washed with DI water (2x) and 2% LiCl solution, dried over Na2SO4, filtered, and concentrated under vacuum.The residue was purified via silica gel chromatography (50% EA/hexanes) to yield the bromide 4 (393 mg, 21%) as a white solid; mp = 142–145 °C; TLC Rf = 0.54 (50% EA/hexanes); IR (ATR) 3304, 3284, 2978, 1713, 1681, 1163, 673 cm-1; 1H NMR (400 MHz, CDCl3) δ 7.93 (d, J = 7.5 Hz, 1H), 7.80 (d, J = 9.1 Hz, 2H), 7.64 (d, J = 8.6, 2H), 7.45 (t, J = 8.3 Hz, 1H), 7.26 (t, J = 8.0 Hz, 1H), 6.97 (d, J = 8.1 Hz, 1H), 4.19 (q, J = 14.0, 6.9 Hz, 2H), 1.30 (t, J = 7.0 Hz, 3H); 13C NMR (100 MHz, CDCl3) δ 170.2, 155.2, 149.0, 135.8, 133.5, 132.8, 131.6, 130.6, 128.2, 126.4, 124.4, 115.2, 114.3, 37.7, 12.3; Anal. Calcd for C17H14BrN3O2: C, 54.86; H, 3.79; N, 11.29. Found: C, 54.56; H, 3.71; N, 11.20.
4-Iodo-N-[3-ethyl-2-oxo-2,3-dihydro-4(1H)-quinazolinyliden]benzenecarboxamide (5)
NaH (2 equiv.) was suspended in dry DMA and stirred for 15 min. A solution of quinazoline 3 (1 equiv.) dissolved in dry DMA (0.1 M) was added dropwise to the reaction, followed by addition of 4-iodobenzoyl chloride (1.5 equiv.). The reaction was stirred for 24 h. Ethyl acetate (EA) and water were added and the layers were separated. The aqueous layer was extracted with EA (2x). The combined organic layers were washed with DI water (2x) and 2% LiCl, dried over Na2SO4, filtered, and concentrated under vacuum. The residue was purified via silica gel chromatography (3% MeOH/DCM) to yield the iodide 5 (161 mg, 36%) as a white solid; mp = 160–163 °C; TLC Rf = 0.40 (1% MeOH/DCM); IR (ATR) 3304, 3285, 2978, 1712, 1670, 1471, 598 cm−1; 1H NMR (400 MHz, CD3CN) δ 8.88 (s, 1H), 7.73 (d, J = 8.4 Hz, 1H), 7.42 (d, J = 8.8 Hz, 2H), 7.18 (d, J = 8.0 Hz, 2H), 6.92 (t, J = 7.6 Hz, 1H), 6.73 (t, J = 6.8 Hz, 1H), 6.32 (d, J = 7.6 Hz, 1H), 3.51-3.50 (m, 2H), 0.60 (t, J = 7.2 Hz, 3H); 13C NMR (100 MHz, DMSO-d6) δ 170.8, 153.6, 138.5, 137.1, 134.9, 133.1, 131.9, 131.7, 129.0, 126.9, 123.8, 114.5, 104.7, 36.5, 12.1; Anal. Calcd for C17H14IN3O2: C, 48.71; H, 3.37; N, 10.02. Found: C, 49.06; H, 3.39; N, 9.81; HRMS (ESI) m/z calcd for C17H15IN3O2 [M + H]+ 420.0203, found: 420.0201.
N-(Prop-2-ynyl)-d-biotinamide (7)
Propargylamine (0.34 mL, 5.32 mmol) and N-methylmorpholine (0.59 mL, 5.32 mmol) were dissolved in dry DMF (15 mL) at 0 °C and stirred for 30 min under argon. EDC (826 mg, 5.32 mmol) and d-biotin 5 (1.0 g, 4.09 mmol) were added and the reaction was stirred for 2 h and then warmed to rt and stirred for 24 h. The DMF was distilled off and the residue was purified via silica gel chromatography (15% MeOH/CHCl3) to afford the product 7 (694 mg, 60%) as a white solid. mp = 162–165 °C; TLC Rf = 0.27 (10% MeOH/CHCl3); IR (ATR) 3422, 3270, 2929, 2860, 1693, 1637 cm−1; 1H NMR (400 MHz, DMSO-d6) δ 8.21 (t, J = 5.2 Hz, 1H), 6.41 (s, 1H), 6.35 (s, 1H), 4.32-4.28 (m, 1H), 4.14-4.11 (m, 1H), 3.83 (dd, J = 5.4, 2.4 Hz, 2H), 3.12-3.07 (m, 2H), 2.82 (dd, J = 12.8, 4.9 Hz, 1H), 2.57 (d, J = 12.0 Hz, 1H), 2.08 (t, J = 7.0 Hz, 2H), 1.65-1.24 (m, 6H); 13C NMR (100 MHz, DMSO-d6) δ 171.8, 162.7, 81.3, 72.8, 61.0, 59.2, 55.4, 39.8, 34.8, 28.1, 28.0, 27.7, 25.1; Anal. Calcd for C13H19N3O2S: C, 55.49; H, 6.81; N, 14.93. Found: C, 55.62; H, 6.87; N, 14.68.
1’N-(4,4’-Dimethoxytrityl)−11’N-(prop-2-ynyl)-d-biotinamide (8)
Biotin amide 7 (519 mg, 1.84 mmol) was dissolved in dry pyridine (10 mL). DMTrCl (750 mg, 2.21 mmol) and DMAP (23 mg, 0.184 mmol) were added and the reaction was stirred under argon at 40 ˚C for 24 h. Methanol was added (5 mL) and stirred for 5 min under argon. The reaction mixture was concentrated under vacuum and the residue was dissolved in DCM (40 mL) and washed with NaHCO3 (50 mL). The aqueous layer was extracted with DCM (3 × 20 mL). The organic layers were collected, washed with brine (40 mL), dried over Na2SO4, and concentrated under vacuum. The crude residue was purified via silica gel chromatography (1% MeOH/CHCl3 to 4% MeOH/CHCl3) to afford the product 8 (730 mg, 68%) as a white foam. mp = 87–90 °C; TLC Rf = 0.27 (4% MeOH/CHCl3); IR (ATR) 3420, 3271, 3066, 2929, 2860, 1692, 1647, 1507, 1249 cm−1; 1H NMR (400 MHz, CD3OD) δ 7.88 (s, 1H), 7.29-7.22 (m, 5H), 7.17-7.11 (m, 4H), 6.82 (d, J = 8.8 Hz, 4H), 4.54-4.50 (m, 1H), 4.37-4.34 (m, 1H), 3.92 (d, J = 2.5 Hz, 2H), 3.76 (s, 6H), 3.16-3.12 (m, 1H), 2.56 (t, J = 2.4 Hz, 1H), 2.31 (d, J = 13.6 Hz, 1H), 2.24 (dd, J = 12.8, 5.5 Hz, 1H), 2.17 (t, J = 7.2 Hz, 2H), 1.77-1.53 (m, 4H), 1.45-1.37 (m, 2H); 13C NMR (100 MHz, DMSO-d6) δ 171.8, 160.8, 157.8, 141.1, 135.9, 131.0, 129.4, 127.3, 112.6, 81.4, 79.2, 72.8, 71.8, 64.5, 59.3, 55.0, 54.3, 38.6, 34.9, 28.4, 28.1, 27.7, 25.1; Anal. Calcd for C34H37N3O4S: C, 69.96; H, 6.39; N, 7.20. Found: C, 69.58; H, 6.39; N, 6.83.
1’N-(4,4’-Dimethoxytrityl)-N-(prop-1-ynyl)-d-biotinamide-N-(3-ethyl-2-oxo-1,2,3,4-tetrahydroquinazolin-4-ylidene)benzamide (9)
Pd(PPh3)2Cl2 (12 mg, 0.0171 mmol), CuI (7 mg, 0.0343 mmol), biotin amide 8 (100 mg, 0.171 mmol) and iodide 5 (86 mg, 0.206 mmol) were added to a round bottom flask and purged with argon. Dry DMA (1.7 mL) and dry TEA (3.4 mL) were added and the reaction mixture was stirred for 24 h under argon. DI water (10 mL) and EA (10 mL) were added and the layers were separated. The organic layer was washed with DI water (2 ×10 mL), dried over Na2SO4, and concentrated under vacuum. The crude residue was purified via silica gel chromatography (5% MeOH/DCM) to give the coupled product 9 (83 mg, 55%) as a pale yellow solid. mp = 148–150 °C; TLC Rf = 0.49 (10% MeOH/CHCl3); IR (ATR) 2980, 2359, 1676, 1603, 1507, 1245 cm−1; 1H NMR (400 MHz, Acetone-d6) δ 8.21 (d, J = 7.3 Hz, 1H), 8.01 (d, J = 8.4 Hz, 2H), 7.57 (d, J = 8.4 Hz, 2H), 7.52-7.44 (m, 2H), 7.35-7.23 (m, 6H), 7.16 (t, J = 9.2 Hz, 4H), 6.96 (d, J = 8.4 Hz, 1H), 6.84 (d, J = 8.0 Hz, 4H), 6.03 (bs, 1H), 4.48-4.38 (m, 2H), 4.22 (d, J = 4.2 Hz, 2H), 4.15 (q, J = 6.5 Hz, 2H), 3.78 (s, 6H), 3.20-3.16 (m, 1H), 2.81 (bs, 1H), 2.36 (d, J = 11.1 Hz, 1H), 2.28 (dd, J = 12.7, 5.3 Hz, 1H), 2.17-2.13 (m, 2H), 1.78-1.36 (m, 6H), 1.22 (t, J = 6.7 Hz, 3H); 13C NMR (100 MHz, DMSO-d6) δ 172.7, 169.9, 161.8, 158.4, 155.1, 148.8, 143.8, 135.8, 135.7, 133.5, 132.5, 131.3, 130.1, 129.7, 127.6, 127.0, 126.3, 124.2, 115.0, 112.8, 90.6, 81.9, 72.8, 65.4, 59.7, 55.2, 54.4, 39.3, 37.6, 35.2, 31.6, 29.7, 28.2, 27.8, 25.0, 22.7, 14.1, 12.1; HRMS (ESI) m/z calcd for C51H50N6O6SNa [M +Na]+ 897.3405, found: 897.3406.
Biotin-Compound-C2 (10)
Protected biotin conjugate 9 (15 mg, 0.131 mmol) was dissolved in 3% TFA in DCM (1 mL). The reaction mixture was stirred for 2 h. MeOH (3 mL) was added and the mixture was concentrated under vacuum. The crude residue was purified via silica gel chromatography (5% MeOH/DCM to 4% MeOH: 95% DCM: 1% TEA) to yield the Biotin-Compound-C2 conjugate 10 (46 mg, 61%) as a pale yellow solid. mp = 159–162 °C; TLC Rf = 0.63 (20% MeOH/CHCl3); IR (ATR) 2926, 2019, 1701, 1602, 1232 cm−1; 1H NMR (400 MHz, DMSO-d6) δ 9.45 (bs, 1H), 8.37 (t, J = 5.3 Hz, 1H), 8.29 (d, J = 7.9 Hz, 1H), 7.98 (d, J = 8.2 Hz, 2H), 7.58 (d, J = 8.2 Hz, 2H), 7.50 (t, J = 7.9 Hz, 1H), 7.29 (t, J = 7.9 Hz, 1H), 6.87 (d, J = 8.2 Hz, 1H), 6.41 (s, 1H), 6.34 (s, 1H), 4.29-4.26 (m, 1H), 4.15 (d, J = 5.2 Hz, 2H), 4.12-4.05 (m, 3H), 3.10-3.05 (m, 1H), 2.79 (dd, J = 13.1, 5.4 Hz, 1H), 2.55 (d, J = 12.4 Hz, 1H), 2.12 (t, J = 7.7 Hz, 2H), 1.65-1.29 (m, 6H), 1.16 (t, J = 6.2 Hz, 3H); 13C NMR (100 MHz, DMSO-d6) δ 172.4, 172.3, 170.4, 163.0, 148.7, 135.2, 133.6, 132.5, 131.9, 130.8, 129.3, 127.2, 124.2, 114.6, 114.1, 92.5, 80.7, 61.3, 59.4, 55.6, 36.8, 35.1, 28.8, 28.7, 28.3, 28.2, 25.3, 12.2; HRMS (ESI) m/z calcd for C31H32N6O4S [M + H]+ 573.2279, found: 573.2272.
tert-Butyl N-[3-(4-{[(4E)−3-ethyl-2-oxo-1,2,3,4-tetrahydroquinazolin-4-ylidene] carbamoyl} phenyl)prop-2-yn-1-yl]carbamate (11), (Compound-z)
Boc protected propargyl amine (15 mg, 0.099 mmol), iodide 5 (50 mg, 0.119 mmol), Pd(PPh3)2Cl2 (7 mg, 0.010 mmol), and CuI (4 mg, 0.019 mmol) were purged with argon. Dry THF (0.9 mL) and dry TEA (2 mL) were added and the reaction was stirred for 24 h under argon. DI water (5 mL) and EA (3 mL) were then added. The aqueous layer was extracted with EA (2 × 3 mL). The combined organic layers were washed with brine (10 mL), dried over Na2SO4, and concentrated under vacuum. The crude residue was purified via silica gel chromatography (40% EA/hexanes) to afford the alkyne 11 (35 mg, 81%) as a yellow solid. mp = 135–137 °C; TLC Rf = 0.11 (40% EA/hexanes); IR (ATR, cm−1) 2980, 2932, 2168, 1672, 1162; 1H NMR (400 MHz, acetone-d6) δ 8.94 (bs, 1H), 8.21 (d, J = 8.4 Hz, 1H), 8.02 (d, J = 7.8 Hz, 2H), 7.58 (d, J = 7.8 Hz, 2H), 7.51 (t, J = 7.8 Hz, 1H), 7.29 (t, J = 7.8 Hz, 1H), 6.97 (d, J = 8.4 Hz, 1H), 6.48 (bs, 1H), 4.19-4.14 (m, 4H), 1.43 (s, 9H), 1.23 (t, J = 6.8 Hz, 3H); 13C NMR (100 MHz, acetone-d6) δ 171.3, 156.3, 154.9, 149.9, 136.5, 133.8, 133.5, 133.0, 131.2, 130.4, 127.4, 124.6, 115.6, 115.5, 92.6, 81.7, 79.4, 37.5, 31.3, 28.6, 12.5; HRMS (ESI) m/z calcd for C25H26N4O4Na [M + Na]+ 469.1846, found: 469.1846.
3-(4-{[(4E)−3-Ethyl-2-oxo-1,2,3,4-tetrahydroquinazolin-4-ylidene]carbomoyl} phenyl)prop-2-yn-1-ammonium trifluoroacetate (12)
The Boc protected amine 11 (43 mg, 0.0963 mmol) was dissolved in TFA (1 mL) and stirred for 2 h. The reaction mixture was then concentrated under vacuum. The residue was dissolved in MeOH (3 × 10 mL) and then concentrated, followed by DCM (3 × 10 mL) to yield the TFA salt 12 (44 mg, 100%) as a light brown solid. mp = 98–101 °C; TLC Rf = 0.38 (10% MeOH/DCM); IR (ATR) 2996, 1973, 1669, 1601, 1129 cm-1; 1H NMR (400 MHz, CD3OD) δ 8.43 (d, J = 8.4 Hz, 1H), 8.13 (d, J = 8.42 Hz, 2H), 7.86 (t, J = 8.0 Hz, 1H), 7.68 (d, J = 8.4 Hz, 2H), 7.56 (t, J = 8.0 Hz, 1H), 7.19 (d, J = 8.4 Hz, 1H), 4.25 (q, J = 7.0 Hz, 2H), 4.10 (s, 2H), 1.38 (t, J = 7.3 Hz, 3H); 13C NMR (100 MHz, CD3OD) δ 169.0, 160.3, 147.5, 139.4, 139.0, 133.8, 132.8, 132.2, 130.4, 127.7, 126.6, 116.7, 110.4, 86.7, 86.3, 41.3, 30.7, 11.5; HRMS (ESI) m/z calcd for C20H19N4O2+ [M]+ 347.1503, found: 347.1501.
Biotin-Compound-c2 pulldown
HEK293 cells were transiently transfected with HA-HIF2α or active site point mutants and protein lysate extracted. Lysate was incubated with indicated amounts of Biotin-Compound-c2 (10) at 4 °C for 1 h then added to 20 µl streptavidin-conjugated agarose and incubated at 4 °C for 1 h. Following three washes with fresh extraction buffer, bound proteins were eluted in 5x Laemmli buffer. Samples were run by SDS PAGE, transferred to nitrocellulose membrane, and analyzed by Western blot.
Statistics and reproducibility
Presented data are representative of three biological replicates, unless otherwise specified. Densitometry was performed using Photoshop v.23.5.1 to quantify Western blot band signal intensity. All statistical analysis were performed using GraphPad Prism version 9.5.0 for Windows (GraphPad Software, https://www.graphpad.com). Statistical significance was ascertained between individual samples using a parametric unpaired two-tailed t-test or one- or two-way analysis of variance (ANOVA) with Tukey’s post hoc test where indicated. Exact p-values are provided on each figure where possible, and significance is denoted by asterisks: *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. Error bars represent the standard deviation for three independent experiments, unless otherwise indicated.
Preparation of Fig
Some Fig. panels were prepared using BioRender software (https://biorender.com/).
Results
Screening and identification of a HIF2α antagonist
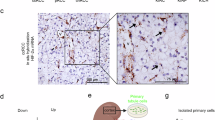
The HIF2α–ARNT complex has a complicated architecture where ARNT spirals around the outside of each HIF subunit9. This leads to a complex heterodimeric structure with five distinct small-molecule binding pockets (binding pockets 1-5) (Fig. 1a). These sites can be further subdivided into PAS domain encapsulated sites (labeled (1, 2, 4, 5) in Fig. 1a) as well as an interfacial cavity formed by the heterodimerization of the two subunits (labeled 3 in Fig. 1a). HIF2α PAS-B domain (labeled (2) in Fig. 1a, Fig. 1b) has proven to be an attractive drug target5,6,7,9,10. We therefore used this domain to conduct an in silico docking study using the previously solved X-ray crystal structure of the HIF2α–ARNT complex (PDB:4ZP4)9, (Fig. 1a). We also performed docking at the HIF2α PAS-A domain (labeled (5) Fig. 1a, 1c). Since this domain has only 60% amino acid sequence identity with HIF2α PAS-B domain9, it may provide selectivity towards structurally distinct inhibitors of HIF2α. A virtual screen using 10,000 drug-like compounds (Fig. 1d) was performed using Autodock Vina18 employing MTiOpenScreen19 at sites 2 and 5 (Fig. 1a). This provided two sets of potential inhibitors, Compounds-c (from site 2, HIF2α PAS-B) and Compounds-i (from site 5, HIF2α PAS-A). From these, the top 23 commercially available compounds with the lowest predicted binding energies from each set were chosen for further evaluation (Supplementary Fig. 1a, 1b). We examined the effect of these 46-compounds at a single dose (10 µM) for 24 h on proliferation of ccRCC (VHL-null) 786-O cells by the 3-(4, 5-dimethylthiazolyl-2)−2, 5-diphenyltetrazolium bromide (MTT) assay. These cells have elevated levels of HIF2α due to the lack of VHL gene14. We set the inhibitory threshold to 60% viability compared to cells with treated with DMSO (vehicle control) and identified Compounds c2 and c15 with the most effective impact on 786-O cells (Supplementary Fig. 1c). Compounds i15 and i19 (that target HIF2α PAS-A) also showed modest inhibitory effect in 786-O cells (Supplementary Fig. 1d). To further assess these four compounds identified from the initial screen (Compounds c2, c15, i15, and i19), we repeated MTT assays in control cell lines HEK293 and VHL-positive ccRCC cell line Caki-1 in addition to VHL-null 786-O cells (Supplementary Fig. 1e). Our data confirmed our earlier results that Compounds-c2 and c15 have a ~ 50% inhibitory effect on 786-O cell proliferation (Supplementary Fig. 1e). Since Compounds-i15 and i19 did not reach this inhibitory threshold, we decided not to further examine them.
a Intramolecular interfaces 1, 2, 4, and 5 on ARNT (blue) and HIF2α (green) (PDB:4ZP4). Interface 3 is the intermolecular junction between ARNT and HIF2α. b Detailed surface of the binding pocket 2 of the HIF2α PAS-B domain used in the docking run. c Detailed surface of the binding pocket 5 of the HIF2α PAS-A domain used in the docking run. d Schematic workflow of in silico screening of a library of drug like compounds (diverse-lib, ~10,000 compounds) using Autodock Vina on the MTiOpenScreen server. Potential HIF2α PAS-B and PAS-A inhibitors (c-compounds and i-compounds, respectively) were identified and further analyzed in cell based assays. This schematic was prepared using BioRender software (https://biorender.com/) and the HIF2α structure (PDB:4ZP4).
Compound-c2 mediated inhibition and proteasomal degradation of HIF2α
To examine the efficacy of our compounds in VHL-null ccRCC cells that are dependent on HIF2α, we treated a second VHL-null ccRCC cell line, A49827,28,29,30, with Compound-c2 and c15. Our data showed that Compound-c2 inhibited the growth of both ccRCC cell lines 786-O and A498 (Fig. 2a). Conversely, this compound had no effect on normal HEK293 cells or ccRCC cells with intact VHL (Caki-1), suggesting its specificity for cells which are VHL-null and have high HIF2α levels (Fig. 2a). In contrast, Compound-c15 had growth inhibitory effect on our tested cell lines irrespective of presence or absence of the VHL gene, suggesting Compound-c15 may have off target effects in cells (Supplementary Fig. 1e, Supplementary Fig. 2). We therefore decided to further analyze only Compound-c2.
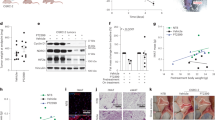
a Two VHL-null ccRCC cell lines (A498, 786-O), VHL-positive ccRCC cell line (Caki-1), and normal kidney HEK293 cells were treated with the indicated amounts of Compound-c2 for 24 h. The effect of Compound-c2 on cell viability was assessed by MTT assay for three independent experiments. A Student’s t test (two-tailed) was performed to assess statistical significance compared to DMSO vehicle control (**P < 0.01). b 786-O cells were treated with the indicated amounts of Compound-c2 for 24 h. Induction of apoptosis was assessed by immunoblotting. β-actin was used as a loading control. c Luminescence from 786-O cells transiently expressing VEGF-luciferase reporter for 24 h, treated with 30 µM of Compound-c2 for an additional 8 h. siRNA targeting HIF2α was used as a control. Error bars represent the standard deviation (S.D.) of three independent experiments. One-way analysis of variance (ANOVA) with Tukey’s post hoc test was used to determine statistical significance (*P < 0.05, ***P < 0.001). d VEGF, NDRG2, CCND1, GLUT1, and ACTB mRNA expression was determined by RT-qPCR in 786-O cells following treatment with the indicated amounts of Compound-c2 for 24 h. Bars represent the ratio of HIF2α target mRNA:ACTB mRNA normalized to the NT control. Error bars represent the standard deviation (S.D.) of three replicates. Two-way analysis of variance (ANOVA) with Tukey’s post hoc test was used to determine statistical significance. e 786-O cells were treated with the indicated amounts of Compound-c2 for 24 h. The stability of HIF2α was assessed by immunoblotting. β-actin was used as a loading control. f 786-O cells were treated with 30 μM Compound-c2 for the indicated durations. The stability of HIF2α was assessed by immunoblotting. β-actin was used as a loading control. g 30 μM Compound-c2 was used to treat OS-RC-2 cells for the indicated length of time. The stability of HIF2α was assessed by immunoblotting. β-actin was used as a loading control. h 30 μM Compound-c2 was used to treat TUHR14TKB cells for the indicated length of time. The stability of HIF2α was assessed by immunoblotting. β-actin was used as a loading control. i 50 nM Bortezomib was added to 786-O cells for 1 h followed by addition of 30 µM Compound-c2 for 4 h. 786-O cells were also treated individually with Bortezomib or Compound-c2. The stability of HIF2α was assessed by immunoblotting. β-actin was used as a loading control. LE, long exposure. SE, short exposure.
We next treated 786-O cells with Compound-c2 and observed a marked increase in cleaved caspase-3 and cleaved caspase-7 (an indication of apoptosis) (Fig. 2b). To determine the effect of Compound-c2 on HIF2α function, we examined HIF2α activity by VEGF-luciferase reporter assay26. We found that treatment of 786-O cells with 30 μM Compound-c2 led to significant downregulation of HIF2α activity (Fig. 2c). We further showed that Compound-c2 also downregulates mRNA expression of the HIF2α targets VEGF, NDRG2, CCND1, and GLUT1 (Fig. 2d)6,31. Interestingly, treatment of 786-O cells with different amounts of Compound-c2 for 24 h reduced levels of HIF2α (Fig. 2e). We further showed that diminished levels of HIF2α occurs within 4 h treatment of 786-O (Fig. 2f). We observed the same effect in two other RCC cell lines with high levels of HIF2α32, OS-RC-2 (Fig. 2g), and TUHR14TKB31 (Fig. 2h). To determine if Compound-c2 caused HIF2α degradation via the proteosome, we co-treated 786-O cells with Compound-c2 and the proteasome inhibitor Bortezomib and demonstrated stabilization of HIF2α protein level in these cells even in the presence of Compound-c2 (Fig. 2i). Taken together, our data demonstrates that Compound-c2 treatment inhibits HIF2α and leads to its proteasomal degradation.
Specificity and selectivity of Compound-c2 for HIF2α
HIF exists in two main isoforms: HIF1α and HIF2α15. In ccRCC, HIF1α has been shown to have tumor suppressor properties, while HIF2α acts to promote tumor growth33. The two isoforms share similar amino acid sequence identity in both the PAS A (68%) and PAS B (73%) domains34. To determine whether Compound-c2 is specific for HIF2α, we first treated Caki-2 cells, which only express HIF1α15, with Compound-c2. We found that Compound-c2 did not induce degradation of HIF1α (Fig. 3a).
a Caki-2 cells were treated with 30 μM Compound-c2 for the indicated durations. The stability of HIF1α was assessed by immunoblotting. β-actin was used as a loading control. LE Long Exposure, SE Short Exposure. b Lysate from 786-O cells was challenged with the indicated amounts of Biotin-Compound-c2 for 1 h. HIF2α binding and input was assessed by immunoblotting. c Empty vector (EV) and HA-HIF2α were transiently expressed in HEK293 cells and lysates were challenged with the indicated amounts of Biotin-Compound-c2 for 1 h. HIF2α binding to Compound-c2 was assessed by immunoblotting. d Amino acid sites of the HIF2α PAS-B domain predicted to contact Compound-c2. Residues shown are those within 3.5 Å of the docked model of Compound-c2. e HEK293 cells were transiently transfected with indicated HA-HIF2α mutants. Lysates were challenged with 1 µM of Biotin-Compound-c2 for 1 h and HIF2α binding to Compound-c2 was assessed by immunoblotting. Densitometry was performed using Photoshop v.23.5.1 to quantify Western blot band signal intensity. Bars represent the mean signal intensity values of HIF2α pulldown:HIF2α input compared to that of WT for three independent measurements. Error bars represent the standard deviation (S.D.) of three independent measurements. A Student’s t test (two-sided) was performed to assess statistical significance compared to WT control (NS- not significant). (See also Fig. S3B-C).
To provide further support that Compound-c2 is binding HIF2α, a biotinylated version of Compound-c2 was developed (Supplementary Fig. 3a). This involved the coupling of an alkyne tagged biotin to an iodinated version of Compound-c2 (Supplementary Fig. 3a). Different amounts of Biotin-Compound-c2 (10) (Supplementary Fig. 3a) were then used to challenge protein lysate from 786-O cells. Our data showed HIF2α binding to 100 nM Biotin-Compound-c2 (Fig. 3b, Supplementary Fig. 3b). Additionally, we confirmed our observation by transiently expressing HA-HIF2α in HEK293 cells and demonstrated binding of HIF2α to 1.0 µM Biotin-Compound-c2 (Fig. 3c).
The docked model of Compound-c2 was then used to predict the amino acids that potentially contact Compound-c2 within the HIF2α PAS-B pocket (residues within 3.5 Å) (Fig. 3d). A hydrogen bond is likely present between the central amide of Compound-c2 and Y307 (the amide carbonyl is 2.8 Å from the oxygen on the tyrosine phenol), while the benzamide aryl ring rests in a hydrophobic pocket near M309 and the heterocycle rests in another hydrophobic pocket near M252 (Fig. 3d). HIF2α residues potentially contacting Compound-c2 were individually mutated to alanine (F244A, S246A H248A, M252A, F254A, Y281A, Y307A, M309A, T321A, I337A, C339A and N341A) (Fig. 3d). As Belzutifan also binds to the HIF2α PAS-B pocket, we also mutated residues within the PAS-B pocket known to contribute to Belzutifan binding (A277E, F280A) to verify that Compound-c2 to has a distinct binding mechanism to HIF2α35. These mutants were transiently expressed in HEK293 cells and their lysate were challenged by Biotin-Compound-c2. Our data showed that all mutants listed above disrupted binding to Compound-c2, with an exception of HIF2α- F244A, A277E, and Y281A (Fig. 3e and Supplementary Fig. 3c, d). Notably, while this method may be prone to introducing artifacts by disrupting protein folding, no substantial effects in protein stability of the point mutants—which would be consistent with disruptions in protein folding—were observed (Supplementary Fig. 3c). This suggests that the constructed point mutations of HIF2α are driving the observed effects on binding to Compound-c2. Taken together, our data confirms the specificity of Compound-c2 for the PAS-B pocket of HIF2α.
Identification of the Compound-c2 pharmacophore
To identify the pharmacophore of Compound-c2, we tested some of the intermediate compounds that were prepared during the incorporation of biotin (Supplementary Fig. 3a) for biological activity. Intermediates 1 (Compound-v), 3 (Compound-w), 2 (Compound-x), 4 (Compound-y), and 11 (Compound-z) (Fig. 4a and Supplementary Fig. 3a) were evaluated by treating 786-O cells at 10 µM for 24 h (Fig. 4b). Cells were then assessed for induction of apoptosis by immunoblotting. Our data showed that both Compound-y and Compound-z caused a reduction of HIF2α levels, as well as an increase in apoptotic markers cleaved caspase-3 and cleaved PARP (particularly upon treatment with Compound-z) (Fig. 4b). We found a similar induction of the apoptotic marker cleaved caspase-3 in a second VHL-null ccRCC cell line, A498, treated with Compound-y and Compound-z, with a reduction in HIF2α levels similar to that caused by Compound-c2 treatment (Fig. 4c). This indicates that both the quinazolinone and the benzamide groups are necessary for inducing apoptosis in 786-O cells.
a Chemical structures of the parent Compound-c2 (center circle) and intermediate compounds (outer circle, Compounds-v, w, x, y, and z). b 786-O cells were treated for 24 h with DMSO (vehicle control) or 10 μM of the Compounds-v, w, x, y, and z. Induction of apoptosis was assessed by immunoblotting. β-actin was used as a loading control. c A498 cells were treated for 24 h with DMSO (vehicle control) or 10 μM of the Compounds-y, z, or c2. Induction of apoptosis was assessed by immunoblotting. β-actin was used as a loading control. d VHL-null ccRCC cell lines (A498, 786-O), VHL-positive ccRCC cell line (Caki-1), and normal renal epithelial HK-2 cells were treated with the indicated amounts of Compound-y for 24 h. The effect of Compound-y on cell viability was assessed by MTT assay for three independent experiments. One-way analysis of variance (ANOVA) with Tukey’s post hoc test was used to determine statistical significance compared to DMSO vehicle control (*P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001). e A498, 786-O VHL-null ccRCC cell lines, Caki-1 VHL-positive ccRCC cell line, and normal renal epithelial HK-2 cells were treated with the indicated amounts of Compound-z for 24 h. The effect of Compound-z on cell viability was assessed by MTT assay for three independent experiments. One-way analysis of variance (ANOVA) with Tukey’s post hoc test was used to determine statistical significance compared to DMSO vehicle control (*P < 0.05; **P < 0.01, ***P < 0.001). f Amino acid sites of the HIF2α PAS-B domain predicted to contact Compound-z. Residues shown are those within 3.5 Å of the docked model of Compound-z.
To examine the specificity of Compound-y and Compound-z for HIF2α, we tested the effect of these compounds for 24 h on the proliferation of VHL-null ccRCC cells (786-O, A498) and VHL-positive ccRCC cells (Caki-1) by MTT assay with HK-2 (normal renal proximal tubule cells) cells as a control (Fig. 4d, e). Our data showed that Compound-y had a growth inhibitory effect on our tested cell lines irrespective of presence or absence of the VHL gene, suggesting Compound-y may have off target effects in cells (Fig. 4d). In contrast, Compound-z inhibited the growth of both ccRCC cell lines 786-O and A498, but had no effect in ccRCC cells with intact VHL (Caki-1) or normal HK-2 cells, suggesting its specificity for cells which are VHL-null and have high HIF2α levels (Fig. 4e). These findings suggest that the substitution pattern on the benzamide is critical to specific inhibition of HIF2α. Based on this information, we have modeled the predicted binding of Compound-z to the HIF2α PAS-B pocket (Fig. 4f). This model predicts that the hydrogen bond between central amide of Compound-z and Y307 is maintained, although the amide carbonyl is further (3.4 Å) from the oxygen on the tyrosine phenol. Additionally, Compound-c2 must twist slightly to fit in the binding site, which places the aromatic rings near M309 and near M252 in slightly different conformations; however, the pocket is large enough to accommodate these changes. Collectively, our data demonstrated that the two intermediates of Biotin-Compound-c2 (Compounds-y and -z) have biological activity in VHL-null ccRCC cells; however, only Compound-z is specific toward HIF2α.
Compound-c2 disrupts Hsp70 chaperoning of HIF2α
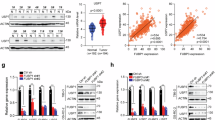
Compound-c2 leads to proteasomal degradation of HIF2α, even in the absence of the VHL-E3 ubiquitin ligase. Therefore, this raised the possibility of a chaperone-dependent pathway for the elimination of HIF2α. The molecular chaperone Hsp70 is involved in the folding and stability of many proteins also known as clients and has been previously shown to interact with HIF2α36,37,38. Here, we also provide evidence that HIF2α binds to Hsp70 by immunoprecipitating endogenous HIF2α and co-immunoprecipitating Hsp70 from 786-O cells (Fig. 5a). We next transiently co-expressed HA-HIF2α with different FLAG tagged domains of Hsp70, nucleotide binding domain (NTB) and substrate binding domain (SBD), in HEK293 cells (Fig. 5b). Immunoprecipitation of FLAG-Hsp70-SBD appears to preferentially co-immunoprecipitate HA-HIF2α, suggesting that HIF2α is a client of Hsp70 (Fig. 5b). To evaluate the effect of Hsp70 activity on HIF2α, we treated 786-O cells with the Hsp70 inhibitor, JG9839. Inhibition of Hsp70 led to the degradation of HIF2α (Fig. 5c), where co-treatment with Bortezomib stabilized HIF2α level (Fig. 5d), suggesting HIF2α relies on Hsp70 for its stability and activity. It is noteworthy that Compound-c2 does not lead to changes in Hsp70 protein levels or Hsp70 client levels, indicating that Compound-c2 does not inhibit Hsp70 activity (Supplementary Fig. 4a, b). Since both the disruption of the Hsp70:HIF2α complex by the Hsp70 inhibitor JG98 and treatment with Compound-c2 results in HIF2α degradation, we asked if Compound-c2 disrupts its interaction with Hsp70. Our data showed that immunoprecipitation of endogenous HIF2α followed by brief (2 h) treatment of 780-O cells with Compound-c2 caused dissociation of the HIF2α:Hsp70 complex (Fig. 5e). Furthermore, using HIF2α-mCherry and HSP70-eGFP expression plasmids to transiently co-express these constructs in HEK293 cells, we demonstrated that 30 µM Compound-c2 for 4 h caused a significant reduction in FRET signal between HIF2α-mCherry and HSP70-eGFP (Fig. 5f, g). Our data suggests that Compound-c2 disrupts the HIF2α:Hsp70 complex (Fig. 5e–g). Additionally, we tested the effect of intermediate Compounds-y and -z to toward HIF2α and Hsp70 binding. Our results via immunoprecipitation and Western blotting (Fig. 5h) as well as FRET (Supplementary Fig. 5a) confirmed that treatment with both Compound-y and Compound-z disrupted the Hsp70:HIF2α complex.
a HIF2α was immunoprecipitated (IP) from 786-O cell lysates using anti-HIF2α antibody (Ab). Interaction with Hsp70 was assessed by immunoblotting. Immunoglobulin G (IgG) was used as a control. β-actin was used as a loading control. b) HA-HIF2α was co-expressed with FLAG tagged domains of Hsp70 depicted in the schematic (left) in HEK293 cells: Hsp70 full length (FL), nucleotide binding domain (NTB), and substrate binding domain (SBD), or empty vector (EV) as a control. Binding of Hsp70 domains with HIF2α was assessed by immunoblotting. GAPDH was used as a loading control. c 786-O cells were treated with 10 μM of the Hsp70 inhibitor JG98 for the indicated times. The stability of HIF2α was assessed by immunoblotting. Phospho-AKT and AKT client levels were used as a positive control for the Hsp70 inhibitor. GAPDH was used as a loading control. d 50 nM Bortezomib (BZ) was added to 786-O cells for 1 h followed by addition of 10 µM JG98 for 4 h. Additionally, 786-O cells were treated individually with Bortezomib or JG98. The stability of HIF2α was assessed by immunoblotting. GAPDH was used as a loading control. All samples were run on a single gel; however, internal samples were removed from the center lanes. e HIF2α was immunoprecipitated (IP) from untreated and treated 786-O cells with 30 µM Compound-c2 for 2 h. Anti-HIF2α and immunoglobulin G (IgG) (control) antibodies (Ab) were used to examine HIF2α interaction with Hsp70 by immunoblotting. β-actin was used as a loading control. f HEK293 cells were co-transfected with mCherry-HIF2α and GFP-HSP70 for 24 h. Cells were then treated with 30 µM Compound-c2 or DMSO control (NT) for 4 hours. FRET was quantified by fluorescence signal of excitation and emission wavelength of 478 nm and 610 nm, respectively. This schematic was prepared using BioRender software (https://biorender.com/). g Data presented as mean value of normalized FRET signal is shown. A Student’s t test (two-tailed) was performed to assess statistical significance. h HIF2α was immunoprecipitated (IP) from 786-O cells treated with DMSO (vehicle control) or 10 μM of Compound-y or z for 24 h. Interaction of HIF2α with Hsp70 as well as induction of apoptosis was assessed by immunoblotting. β-actin was used as a loading control. i 786-O or Belzutifan (Belz) Resistant 786-O cells were treated with DMSO (vehicle control) or 10 μM of Belzutifan for 24 h. HIF2α levels were assessed by immunoblotting. β-actin was used as a loading control. j Lysate from Belzutifan (Belz) Resistant 786-O cells was challenged with the indicated amounts of Biotin-Compound-c2 for 1 h. HIF2α binding and input was assessed by immunoblotting. k Belzutifan (Belz) Resistant 786-O cells were treated with DMSO (vehicle control) or 30 μM of Compound-c2, -y, or z for 24 h. Levels of HIF2α and induction of apoptosis were assessed by immunoblotting. β-actin was used as a loading control. l Belzutifan (Belz) Resistant 786-O cells were treated with 10 μM of the Hsp70 inhibitor JG98 for the indicated times. The stability of HIF2α was assessed by immunoblotting. Phospho-AKT and AKT client levels were used as a positive control for the Hsp70 inhibitor. GAPDH was used as a loading control. m HEK293 cells transiently expressing WT HA-HIF2α and S246A, H248A, and F280A mutants were treated with 10 µM of Hsp70 inhibitor, JG98, for the indicated time points. HA-HIF2α protein levels were examine by immunoblotting. β-actin was used as a loading control.
Belzutifan (Welireg, Merck & Co., Inc.) is an FDA approved HIF2α inhibitor for adult patients with VHL disease, including those with advanced renal cell carcinoma16. As an initial step to compare the effect of our identified compounds with that of Belzutifan, we first investigated the impact of Belzutifan treatment on the Hsp70:HIF2α complex. No significant reduction in FRET signal between HIF2α-mCherry and HSP70-eGFP was observed upon Belzutifan treatment (Supplementary Fig. 5b). Furthermore, no disruption of the Hsp70:HIF2α complex was observed via immunoprecipitation and Western blotting (Supplementary Fig. 5c). Our data demonstrates that Belzutifan does not disrupt the Hsp70:HIF2α complex (Supplementary Fig. 5b, c). This is in contrast to Compounds-c2, -y, and -z (Fig. 5e–h and Supplementary Fig. 5a), which suggests a distinct mechanism of action of our identified compounds from Belzutifan. Furthermore, Compounds-c2 and -z caused a stronger induction of apoptosis compared to Belzutifan in 786-O cells, but not in HEK293 cells (Supplementary Fig. 5d-f). This cell-specific effect was associated with an enhanced reduction of HIF2α protein levels upon treatment with Compounds-c2 and -z as compared to Belzutifan (Supplementary Fig. 5d,e,g).
Notably, patients that receive Belzutifan can potentially develop resistance, therefore requiring a second line of therapy to combat Belzutifan-resistant disease. We have developed a Belzutifan-resistant 786-O cell line that were cultured in 10 µM Belzutifan until survival in the drug treatment was maintained. Treatment of 786-O cells with 10 µM of Belzutifan for 24 h, as expected, led to reduction of HIF2α (Fig. 5i). However, reintroduction of 10 µM Belzutifan in the Belzutifan-resistant 786-O cells did not further affect HIF2α levels (Fig. 5i). Surprisingly, HIF2α levels in these Belzutifan-resistant 786-O cells have been significantly reduced compared to 786-O cells (Fig. 5i). We next examined the effect of our HIF2α inhibitors in this Belzutifan-resistant cell line by first examining the binding of biotin-Compound-c2 to HIF2α (Fig. 5j). Our data showed that despite exhibiting Belzutifan resistance, HIF2α from these cells retained the ability to bind Biotin-Compound-c2 (1 µM) (Fig. 5j). Additionally, treatment of these cells with Compound-c2, -y, or -z caused apoptosis, but did not cause lower levels of HIF2α (Fig. 5k). Lastly, we wanted to examine whether the stability of HIF2α from Belzutifan-resistant 786-O cells also depended on the molecular chaperone Hsp70. We tested this by treating these cells with 10 µM of the Hsp70 inhibitor JG98 (Fig. 5l)39. Our data show that inhibition of Hsp70 led to destabilization of HIF2α from Belzutifan-resistant 786-O cells (Fig. 5l). Our earlier data (Fig. 3e) suggested that HIF2α-S246A, or H248A, or F280A mutants had reduced affinity towards Biotin-Compound-c2. As an alternative approach for targeting these mutants, we tested whether they are sensitive to Hsp70 inhibition. Our data indicates that treatment of HEK293 cells transiently expressing the above mutants with 10 µM of JG98 led to degradation of HIF2α (Fig. 5m).
Taken together, our data suggest that HIF2α is a client of the molecular chaperone Hsp70. Compound-c2 and Compound-z specifically binds to HIF2α and disrupts its interaction with Hsp70 to cause its proteasomal degradation. Lastly, Compound-c2 -z and -y targeted HIF2α in Belzutifan-resistant 786-O cells and caused apoptosis, therefore providing a potential alternative to combat drug resistance in ccRCC (Fig. 6).
Schematic representation of targeting HIF2α by Compounds-c2 and -z. VHL-null kidney tumors exhibit elevated HIF2α levels, which is protected from proteasomal degradation by the molecular chaperone Hsp70 (left). Stable HIF2α translocates to the nucleus to upregulate genes associated with tumor survival. Treatment with Compound-c2 or Compound-z (right) dissociates the HIF2α:Hsp70 complex, causing HIF2α proteasomal degradation and subsequent apoptosis of cancer cells. This schematic was prepared using BioRender software (https://biorender.com/).
Discussion
Belzutifan (Welireg, Merck & Co., Inc.) is an FDA approved HIF2α inhibitor for patients with advanced renal cell carcinoma (RCC) following treatment with a programmed death receptor-1 (PD-1) or programmed death-ligand 1 (PD-L1) inhibitor and a vascular endothelial growth factor tyrosine kinase inhibitor (VEGF-TKI)16,40. This achievement was based on foundational work to understand the structure and function of HIF2α, which led to the identification and further development of small molecule HIF2α inhibitors10. However, targeted therapy by any small molecules in cancer eventually leads to acquired drug resistance. One mechanism of acquired resistance is mutation of the drug target. Previous studies in Patient-Derived Xenograft (PDX) models have demonstrated that prolonged exposure of ccRCC tumors to PT2385 (precursor to Belzutifan) acquired resistance as the result of a mutation (G323E) in HIF2α5. Although G323 is in the HIF2α PAS-B domain5, in silico modeling using the docked model of Compound-c2 and HIF2α did not predict Compound-c2 to make contact with the G323 residue (Fig. 3d). This provides a compelling rationale that Compound-c2, the alternative HIF2α inhibitor identified in this work, may be a promising lead as a candidate therapy in patients who develop resistance to Belzutifan. Given that Belzutifan elicits an objective response in only ~49% of patients and has no reported complete responses40, there remains substantial unmet need for complementary or second-line strategies. Compound-c2 may fill this therapeutic niche, particularly in Belzutifan-refractory disease. Furthermore, the Compound-c2 mode of action that results in inhibition of HIF2α appears to be different from Belzutifan and its analogs. Compound-c2 binding to HIF2α PAS-B caused its dissociation from the molecular chaperone Hsp70 (Fig. 5e–g), which we did not observe upon treatment with Belzutifan (Supplementary Fig. 5b, c). This disruption of the HIF2α:Hsp70 complex led to degradation of HIF2α in the proteasome (Fig. 6). Our data based on the Belzutifan-resistant 786-O cell line suggested that our compounds can cause apoptosis in this cell line; however, they did not negatively impact HIF2α levels. Interestingly, HIF2α from Belzutifan-resistant 786-O cells still depend on the molecular chaperone Hsp70.
Our findings showed that HIF2α is indeed a client of Hsp70, and it preferentially binds to the substrate binding domain of Hsp70. This suggests that perhaps in the presence of oxygen, HIF2α must initially dissociate from the molecular chaperone Hsp70, which allows for prolyl-hydroxylation and VHL mediated ubiquitination and degradation of HIF2α. Our data indicate that inhibition of Hsp70 reduces HIF2α levels, even in Belzutifan-resistant 786-O cells (Fig. 5l) and in HEK293 cells overexpressing mutant HIF2α that cannot bind to Compound-c2 (Fig. 5m). Therefore, targeting the molecular chaperones, and consequently activating the VHL-independent HIF2α -degradative pathway, may be an attractive therapeutic approach for the treatment of ccRCC. This also raises the possibility of the involvement of other molecular chaperones such as Hsp90 towards HIF2α stability. Previous work has shown the involvement of Hsp90 in the chaperoning and turnover of HIF1α41,42,43. Further work is necessary to explore the therapeutic potential of molecular chaperone inhibitors in VHL-null ccRCC.
Although we have provided a body of evidence that suggests Compound-c2 specifically binds and inhibits HIF2α in VHL-null ccRCC, some outstanding questions warrant further studies. These include in vivo analysis of our compound, which requires detailed pharmacokinetic and pharmacodynamic analysis. Further analysis of small molecules based on Compound-c2 suggested that the quinazolinone and the benzamide groups provided the pro-apoptotic activity of these molecules, with the substitution on the benzamide being critical to selective HIF2α inhibition. These components can potentially serve as attractive scaffolds for the design and synthesis of compounds similar to Compound-c2 with improved binding affinity. Compound-c2 and its derivatives (Compound-y and -z) caused an induction of apoptosis in VHL-null ccRCC; however, it remains to be determined whether this is activation of extrinsic or intrinsic pathways of apoptosis. HIF2α upregulation has also been reported previously in other maladies, and the efficacy of Compound-c2 and its derivatives in targeting HIF2α in those diseases remains to be examined.
Here, we utilized the crystal structure of HIF2α and an in silico drug screening approach to identify HIF2α inhibitors. Docking a library of small druglike molecules in both the PAS-A and PAS-B pockets of HIF2α led to the identification of Compound-c2 that specifically binds to the PAS-B pocket. Our data based on mutagenesis of HIF2α suggests that Compound-c2 makes contacts with 11 residues in the PAS-B pocket. Given the pro-survival role of HIF2α in VHL-null ccRCC, Compound-c2 caused apoptosis only in ccRCC cell lines that lacked the VHL gene. Our data, based on binding and pharmacologic inhibition of Hsp70 in VHL-null ccRCC cells, suggest that HIF2α is a client of Hsp70. Therefore, we have provided a model in which Compound-c2 binds to the HIF2α PAS-B pocket, causing the dissociation of HIF2α from Hsp70 and its consequent ubiquitination and degradation in the proteasome. Collectively, this unique mechanism of action positions Compound-c2 as a promising alternative to inhibit HIF2α-related processes, with potential future applications in addressing drug resistance for kidney cancer treatment.
Data availability
This study did not generate any datasets and was not deposited in any database. Supplemental Figs. 1–5 may be accessed in the Supplementary Information file. Uncropped western blots for Figs. 2b, 2e–i, 3a–c, 4b, c, 5a–e, 5h-m, Supplementary Fig. 3b–d, Supplementary Fig. 4, Supplementary Fig. 5c-e may be accessed in the Supplementary Information file as Supplemental Figs. 6−15. Sequences of oligonucleotides are provided in Supplemental Data 1. The 1H NMR, 13 C NMR, and HRMS spectra of the synthesized target compounds may be accessed in the Supplemental Data 2. The numerical results underlying graphs for Figs. 2a, 2c, d, 3e, 4d, e, 5g, Supplementary Fig. 1c–e, Supplementary Fig. 2, Supplementary Fig. 5a, b, Supplementary Fig. 5f, g may be accessed in the Supplemental Data 3. The predicted binding energies towards HIF2α for the top 500 compounds, ranked by binding energies on 3 scored poses generated using the MTiOpenscreen server, may be accessed in Supplemental Data 3. This includes binding energies for Site 2 and Site 5 on HIF2α (see Fig. 1a).
References
Kondo, K., Kim, W. Y., Lechpammer, M. & Kaelin, W. G. Jr. Inhibition of HIF2alpha is sufficient to suppress pVHL-defective tumor growth. PLoS Biol. 1, E83 (2003).
Wang, G. L., Jiang, B. H., Rue, E. A. & Semenza, G. L. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc. Natl. Acad. Sci. USA 92, 5510–5514 (1995).
Loboda, A., Jozkowicz, A. & Dulak, J. HIF-1 and HIF-2 transcription factors-similar but not identical. Mol. Cells 29, 435–442 (2010).
Scheuermann, T. H. et al. Artificial ligand binding within the HIF2alpha PAS-B domain of the HIF2 transcription factor. Proc. Natl. Acad. Sci. USA 106, 450–455 (2009).
Chen, W. et al. Targeting renal cell carcinoma with a HIF-2 antagonist. Nature, 539, 112–117 (2016).
Cho, H. et al. On-target efficacy of a HIF2alpha antagonist in preclinical kidney cancer models. Nature, 539, 107–111 (2016).
Choueiri, T. K. et al. Inhibition of hypoxia-inducible factor-2alpha in renal cell carcinoma with belzutifan: a phase 1 trial and biomarker analysis. Nat. Med 27, 802–805 (2021).
Choueiri, T. K. et al. From basic science to clinical translation in kidney cancer: a report from the second kidney cancer research summit. Clin. Cancer Res. 28, 831–839 (2022).
Wu, D., Potluri, N., Lu, J., Kim, Y. & Rastinejad, F. Structural integration in hypoxia-inducible factors. Nature 524, 303–308 (2015).
Wallace, E. M. et al. A small-molecule antagonist of HIF2alpha is efficacious in preclinical models of renal cell carcinoma. Cancer Res. 76, 5491–5500 (2016).
Maxwell, P. H. et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature 399, 271–275 (1999).
Kaelin, W. G. Jr. The von Hippel-Lindau tumour suppressor protein: O2 sensing and cancer. Nat. Rev. Cancer 8, 865–873 (2008).
Duan, D. R. et al. Characterization of the VHL tumor suppressor gene product: localization, complex formation, and the effect of natural inactivating mutations. Proc. Natl. Acad. Sci. USA 92, 6459–6463 (1995).
Iliopoulos, O., Kibel, A., Gray, S. & Kaelin, W. G. Jr. Tumour suppression by the human von Hippel-Lindau gene product. Nat. Med. 1, 822–826 (1995).
Kondo, K., Klco, J., Nakamura, E., Lechpammer, M. & Kaelin, W. G. Jr. Inhibition of HIF is necessary for tumor suppression by the von Hippel-Lindau protein. Cancer Cell 1, 237–246 (2002).
Fallah, J. et al. FDA approval summary: belzutifan for von hippel-lindau disease-associated tumors. Clin. Cancer Res. 28, 4843–4848 (2022).
Albiges, L. et al. Pembrolizumab plus lenvatinib as first-line therapy for advanced non-clear-cell renal cell carcinoma (KEYNOTE-B61): a single-arm, multicentre, phase 2 trial. Lancet Oncol. 24, 881–891 (2023).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Labbe, C. M. et al. MTiOpenScreen: a web server for structure-based virtual screening. Nucleic Acids Res 43, W448–W454 (2015).
Hageman, J. & Kampinga, H. H. Computational analysis of the human HSPH/HSPA/DNAJ family and cloning of a human HSPH/HSPA/DNAJ expression library. Cell Stress Chaperones 14, 1–21 (2009).
Mollapour, M. et al. Swe1Wee1-dependent tyrosine phosphorylation of Hsp90 regulates distinct facets of chaperone function. Mol. Cell 37, 333–343 (2010).
Woodford, M. R. et al. The FNIP co-chaperones decelerate the Hsp90 chaperone cycle and enhance drug binding. Nat. Commun. 7, 12037 (2016).
Wang, Y. et al. PubChem BioAssay: 2014 update. Nucleic Acids Res. 42, D1075–D1082 (2014).
Miteva, M. A., Guyon, F. & Tuffery, P. Frog2: Efficient 3D conformation ensemble generator for small compounds. Nucleic Acids Res. 38, W622–W627 (2010).
Baker-Williams, A. J. et al. Co-chaperones TIMP2 and AHA1 competitively regulate extracellular HSP90:client MMP2 activity and matrix proteolysis. Cell Rep. 28, 1894–1906 e1896 (2019).
Post, D. E. & Van Meir, E. G. Generation of bidirectional hypoxia/HIF-responsive expression vectors to target gene expression to hypoxic cells. Gene Ther. 8, 1801–1807 (2001).
DepMap B. DepMap 24Q4 Public. In: Figshare (2024).
Ding, X. F., Zhou, J., Chen, G. & Wu, Y. L. VHL loss predicts response to Aurora kinase A inhibitor in renal cell carcinoma cells. Mol. Med Rep. 18, 1206–1210 (2018).
Latif, F. et al. Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 260, 1317–1320 (1993).
Ikediobi, O. N. et al. Mutation analysis of 24 known cancer genes in the NCI-60 cell line set. Mol. Cancer Ther. 5, 2606–2612 (2006).
Jiang, Q. et al. HIF regulates multiple translated endogenous retroviruses: Implications for cancer immunotherapy. Cell 188, 1807–1827 e1834 (2025).
Stransky, L. A. et al. Sensitivity of VHL mutant kidney cancers to HIF2 inhibitors does not require an intact p53 pathway. Proc. Natl. Acad. Sci. USA 119, e2120403119 (2022).
Shen, C. & Kaelin, W. G. Jr. The VHL/HIF axis in clear cell renal carcinoma. Semin. Cancer Biol. 23, 18–25 (2013).
Tian, H., McKnight, S. L. & Russell, D. W. Endothelial PAS domain protein 1 (EPAS1), a transcription factor selectively expressed in endothelial cells. Genes Dev. 11, 72–82 (1997).
Ren, X., Diao, X., Zhuang, J. & Wu, D. Structural basis for the allosteric inhibition of hypoxia-inducible factor (HIF)-2 by belzutifan. Mol. Pharmacol. 102, p240–247 (2022).
Mayer, M. P. & Gierasch, L. M. Recent advances in the structural and mechanistic aspects of Hsp70 molecular chaperones. J. Biol. Chem. 294, 2085–2097 (2019).
Zuiderweg, E. R., Hightower, L. E. & Gestwicki, J. E. The remarkable multivalency of the Hsp70 chaperones. Cell Stress Chaperones 22, 173–189 (2017).
Luo, W. et al. Hsp70 and CHIP selectively mediate ubiquitination and degradation of hypoxia-inducible factor (HIF)-1alpha but Not HIF-2alpha. J. Biol. Chem. 285, 3651–3663 (2010).
Li, X. et al. Analogs of the allosteric heat shock protein 70 (Hsp70) inhibitor, MKT-077, as anti-cancer agents. ACS Med. Chem. Lett. 4, 1042–1047 (2013).
Jonasch, E. et al. Belzutifan for renal cell carcinoma in von Hippel-Lindau disease. N. Engl. J. Med. 385, 2036–2046 (2021).
Isaacs, J. S. et al. Hsp90 regulates a von Hippel Lindau-independent hypoxia-inducible factor-1 alpha-degradative pathway. J. Biol. Chem. 277, 29936–29944 (2002).
Hubbi, M. E. et al. Chaperone-mediated autophagy targets hypoxia-inducible factor-1alpha (HIF-1alpha) for lysosomal degradation. J. Biol. Chem. 288, 10703–10714 (2013).
Tang, X. et al. Heat shock protein-90alpha (Hsp90alpha) stabilizes hypoxia-inducible factor-1alpha (HIF-1alpha) in support of spermatogenesis and tumorigenesis. Cancer Gene Ther. 28, 1058–1070 (2021).
Acknowledgements
We are grateful for constructive scientific discussion with both Drs. W. Marston Linehan and Len Neckers. This work was supported by the National Institute of General Medical Sciences of the National Institutes of Health under award number R35GM139584 (M.M.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This work was also supported with funds from the SUNY Upstate Medical University and Upstate Foundation. We thank Dr. Jason Gestwicki for providing JG-98. Schematics in Figs. 1 and 5 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Conceptualization of the project G.B., J.D.C., and M.M.; experimental design, investigation, data analysis and presentation, J.A.H., S.J.B., A.P., S.C., M.F.B., B.S., R.A.S., M.A.D., D.E.P., M.R.W., D.B., J.D.C., M.M., and G.B.; original draft, J.A.H. and M.M.; contributions to manuscript writing, review, and editing J.A.H., S.J.B., J.D.C., M.M., and G.B.; and supervision, J.D.C., M.M., and G.B. All authors read the manuscript and provided their final approval for the content.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Xiaojin Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Heritz, J.A., Backe, S.J., Pacherille, A. et al. Targeting and dissociating HIF2α from the molecular chaperone Hsp70 triggers apoptosis in kidney cancer. Commun Med 6, 91 (2026). https://doi.org/10.1038/s43856-025-01356-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-025-01356-x