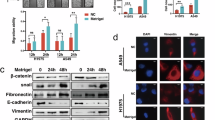
Abstract
Background
Before surgical resection of lung tumor, intraoperative biopsy is needed for cancer diagnosis, while current techniques that guide biopsy have limited performance in tumor identification and boundary determination. Remodeling of extracellular matrix (ECM), mainly collagen and elastin fibers, is an emerging hallmark of tumorigenesis.
Methods
Herein, we establish a quantitative multiphoton microscopy (MPM) imaging method for time-efficient, highly-sensitive lung cancer detection via characterization of ECM remodeling. From label-free images of collagen and elastin fibers acquired simultaneously, we construct a similarity coefficient (SC) metric to describe their interaction, and further develop an artificial intelligence (AI)-ECM framework by producing a fiber voxel dictionary via unsupervised learning of morpho-structural features for explainable and visible assessments of cancer risk.
Results
The application is demonstrated by ex vivo human lung cancer diagnosis with a sensitivity of 99.37%, and recognizing the tumor boundary. The translational potential is further revealed via in vivo imaging of a murine model harboring human lung cancer.
Conclusions
This technology can help surgeons perform more precise biopsies and surgeries by providing explainable visual cues, thus leading to better outcomes for lung cancer patients.
Plain language summary
During lung cancer surgery, doctors need to quickly find the exact tumor location and its edges to take a suitable sample for diagnosis. Current methods can be slow and sometimes miss the tumor. We develop an artificial intelligence (AI)-assisted fast imaging method to see the microscopic fiber structures surrounding lung cells. In cancer, these fibers change their layout. Our system images these changes without needing to add any additional dyes or labels and uses AI to instantly analyze them, creating easy-to-understand color-coded maps that highlight cancerous areas. For human lung tissues, our method identifies cancer with over 99% accuracy and clearly shows the tumor boundary. We also prove it works in living animals. This technology can help surgeons perform more precise biopsies and surgeries, leading to better outcomes for lung cancer patients in the future.
Similar content being viewed by others
Introduction
Lung cancer is the leading cause of cancer-related deaths globally1,2, causing the highest cancer mortality in men, and secondly in women, only behind breast cancer3,4. Radical operation is the most effective method to treat lung cancer at early stages, and precise localization of cancerous lesions intraoperatively is required for tissue sampling and subsequent biopsy, which is crucial for the correct assessment and staging of lung cancer to determine therapy and prognosis of the patient. Thoracotomy is a routine option yet requires a 30–40 cm incision, which causes issues of disfigurement, increased risk of postoperative complications and decrease of pulmonary function. Minimally invasive modalities including video-assisted thoracic surgery (VATS), endobronchial ultrasound-guided fine-needle aspiration (EBUS-FNA) and transesophageal endoscopic ultrasound-guided fine-needle aspiration (EUS-FNA) allow sampling of lesions in a visually guided manner5, but with limitations. Although thoracoscopy-based VATS allows surgeons to visualize the lung structure intraoperatively6,7, visually distinguishing early-stage tumors from normal tissues remains a challenge due to the limited optical performance of thoracoscopy and subtle alterations from tumorigenesis. Meanwhile, training and cost issues, as well as insufficient sensitivity (88% to malignancy8) hinder the widespread application of VATS9. EBUS-FNA and EUS-FNA are used to detect the presence of mediastinal lymph node metastasis10, yet have low sensitivity when applied alone (69% of either one11). Inappropriate tissue sampling can lead to incorrect assessments in biopsy and prolong the surgery time. Cryosectioning is the gold standard for intraoperative diagnosis12, but it might take 30 min or more to include tissue sampling, freezing, sectioning, hematoxylin and eosin (H&E) staining, and pathologists’ review and report, which makes in-time feedback during surgery very challenging. Therefore, an intraoperative tool capable of time-efficient, highly-sensitive and quantitative tumor identification with visual cues is needed.
Extracellular matrix (ECM) is a key element of lung microenvironment and its dynamic remodeling plays an important role in tumor development13,14, making it an emerging cancer hallmark even at early stages. Collagen and elastin fibers are two main components within the lung ECM constituting approximately 60% and 24% of dry lung mass, respectively15,16, and their proper regulation is closely associated with the elastic recoil, mechanical stability and airway patency of lung17. Collagen fibers provide high tensile strength but low elasticity, ensuring the overarching architecture, while elastin fibers with strong elasticity but low tensile strength contribute to the compliance and elastic recoil of lung18. Enzymes including lysyl oxidases (LOX) and matrix metalloproteinases (MMPs) are critical for the deposition and stabilization of mature collagen and elastin fibers19. Meanwhile, transforming growth factor-beta (TGF-β) activates fibroblasts, promotes the production of ECM, and makes the tumor microenvironment more supportive and aggressive20,21. Fiber remodeling caused by dysregulation of these molecular mechanisms has been linked to progression and metastasis of a variety of malignant neoplasms through altering cell migration and invasion, affecting angiogenesis and therapy resistance and increasing intratumoural fluid pressure. Previous studies have demonstrated that progression of lung diseases including asthma, idiopathic fibrosis, pulmonary arterial hypertension, and chronic obstructive pulmonary disease, is accompanied by lung dysfunction along with alterations in mechanical and biochemical status of ECM15,16,22,23,24,25. ECM remodeling has been suggested to play an important role in tumor progression26,27,28,29, including lung cancer30,31,32,33,34, yet few studies have provided a clear picture regarding the interaction between collagen and elastin fibers during the progression of cancers through a way of simultaneous acquisition of image signals from two fiber types.
Nonlinear imaging techniques35, including two-photon excited fluorescence (TPEF), second harmonic generation (SHG), coherent anti-stokes Raman scattering (CARS), stimulated Raman scattering (SRS), third harmonic generation (THG), etc., rely on endogenous multiphoton processes of near-infrared light, offering deep tissue penetration, label-free imaging ability and sub-micron resolution. Among them, multiphoton microscopy (MPM), including SHG and TPEF, has shown great advantages in ECM imaging. SHG is applicable to structures with non-centrosymmetric nature, such as collagen fibers. TPEF is suitable for imaging of elastin fibers and can be detected simultaneously with SHG signals. Further, we have established an analysis platform for fibrous tissues, such as collagen and elastin fibers, by developing voxel-wise morpho-structural parameters of orientation36, alignment37, waviness38, thickness39 and local coverage40, in a truly three-dimensional (3D) context. These efforts have made an impact in uncovering ECM remodeling in a variety of diseases including osteoarthritis41 and neurodegenerative diseases37. At the same time, MPM-based analysis has also shown potential in the diagnosis of cancers including skin cancer42, breast cancer29, ovarian cancer40, gastric cancer28, and so on.
In this study, we establish a quantitative MPM imaging system for assessments of ECM remodeling, aiming at time-efficient intraoperative applications by identifying the correct tumor location and accurate tumor boundary. A specific imaging strategy is developed according to measured fluorescence spectra of collagen and elastin to enable concurrent image acquisition with minimum crosstalk. Based on our recently proposed 3D morpho-structural parameters, we construct an optical metric termed similarity coefficient (SC) to describe the interaction between these two fiber components during tumor progression. Then we develop the artificial intelligence (AI)-ECM framework by producing a fiber voxel dictionary which fully acknowledges the information abundance from voxel-wise level measurements, and achieve highly-sensitive, explainable and visible assessments of cancer risk. The application of this quantitative imaging system is demonstrated by classifying cancerous and normal human lung tissues ex vivo of a total of 222 patients from two hospitals, and accurately identifying tumor boundary with visual cues. Finally, the translational potential of this method is revealed via in vivo imaging of a murine model harboring human lung cancer.
Methods
Multiphoton microscopy (MPM) system
MPM images of lung tissues were obtained using a custom-built microscope with a 25× water dipping objective (NA 0.95, free working distance: 2.5 mm) equipped with a tunable (680–1300 nm) fs laser (InSight DeepSee; Spectra Physics; Mountain View, CA). Emission photons were collected by photomultiplier tube (PMT) detectors. In this study, 800 nm was used as the excitation wavelength for both SHG imaging of collagen fibers and TPEF imaging of elastin fibers. Specifically, TPEF images were collected using the 525 (±25) nm emission filter (Chroma), and SHG images were acquired using the 400 (±10) nm emission filter (Chroma). Both TPEF and SHG signals were collected in epi-direction. According to the fluorescence spectra of collagen and elastin at 800 nm excitation, these settings were proven to efficiently harvest fluorescence signals from elastin fibers while induce minimal crosstalk between these two fiber components. The MPM method was established by obtaining TPEF and SHG images (512 × 512 pixels; 465 × 465 μm; i.e., with a voxel size of 0.91 μm in transverse dimension) simultaneously under the same excitation wavelength of 800 nm for highly time-efficient image collection which was key to intraoperative applications, using a laser power on the sample of ~65 mW for ex vivo imaging and ~72 mW for in vivo imaging based on the experience from our previous work43,44. Exposure of lung tissues to 60–70 mW NIR fs pulses was estimated to be equivalent to a dose of about 0.6 minimal erythema dose of UV exposure45; meanwhile, image acquisition was monitored and no image quality problems were found caused by photodamage. Adenocarcinoma originated primarily from the epithelium surrounding the smaller bronchi and alveoli, and the ECM near the epithelium was remodeled most dramatically during cancer progression. Therefore, 3D imaging was performed starting from the sample surface, with a depth of 70–100 μm and interslice distance of 0.90 μm. The gain of PMT was kept constant throughout this study, and the image intensity was normalized by the laser power recorded for each imaging session.
Preparation and ex vivo imaging of human lung tissues
Lung tissue specimens were harvested from patients with non-small cell lung cancer (NSCLC). The size might vary among tissue samples, typically with 2–10 mm in x, y dimensions and 1–3 mm in z dimension. Excised tissue samples were imaged freshly following extraction, without any treatment. A total of 222 patients from two hospitals (the First Affiliated Hospital, Zhejiang University School of Medicine, n = 75; Fujian Provincial Hospital, n = 147) were involved in this study, and informed consent was obtained from all patients and/or their legal guardians. Confirmed from histology by authorized pathologists, a total of 484 lung samples, including 226 normal and 258 lesion-level tumor tissues, were collected for ex vivo imaging. Specifically, 86 patients provided only one tumor tissues; 136 patients provided two tissue samples, with 100 of them providing one normal and one tumor tissue, and the rest of them providing two tumor tissues. To assess the ability of our quantitative imaging method in tumor boundary identification, additional 23 boundary tissues, 25 cancerous tissues and 20 normal tissues from extra 20 patients were added to the sample pool. Samples were collected during resection surgery. The collected samples were about 2–10 mm in length and width, and about 1–3 mm in thickness. For each tissue sample, three 3D image stacks were obtained from the MPM system and used for the subsequent AI-ECM analysis with imaging fields marked. Post imaging, formalin fixation and H&E histology were performed according to standard protocols and three authorized pathologists assessed specimens to confirm the sample type of the corresponding imaging fields. If the pathologists disagreed on the sample type, then this imaging field would not be used for the subsequent analysis. After determining the normal and cancerous areas, pathologists would delineate a transitional zone, with a width of 2 mm. When the imaging field was located in this transitional zone, it would be considered from a boundary sample. Immunohistochemistry and molecular analysis were performed subsequently to help determine the type and degree of differentiation of the tumor. All procedures were performed in accordance with relevant guidelines/regulations and approved by the institutional review board at Zhejiang University (ZJU21410).
Time efficiency assessment
For each MPM imaging site (512 × 512 pixels, 70–100 μm depth), the acquisition step required 1.5 min on average (range: 1–2 min), while computational processing (AI-ECM) took 2.5 min per site (range: 2–3 min). Crucially, our workflow enabled parallel imaging and computation, i.e., the system processed and diagnosed the obtained image stack while at the same time acquired the next image. This parallelization indicated that after the initial non-parallelizable acquisition of an image stack (1.5 min), each additional site primarily consumed the longer computation time (2.5 min). Therefore, for cancer diagnosis each lung sample containing three adjacent imaging sites would need theoretically 9 min (1.5 + 3 × 2.5 min). For intraoperative applications with sparse imaging of 10 sites on the tumor boundary, it required about 26.5 min (1.5 + 10 × 2.5 min), while further optimization through computing pipelining (parallel computing multiple imaging stacks) might reduce this to approximately 20–25 min.
Preparation and in vivo imaging of a murine model harboring human lung cancer
To test the translational potential of the MPM imaging system and ECM feature extraction, we prepared a murine orthotopic model of human non-small lung cancer through direct thoracic implantation of lung cancer cells46,47 for in vivo imaging. We selected H1299 cell line (Procell Life Science & Technology, Wuhan, China), which was suitable for the generation of lung orthotopic tumors. The H1299 cell line was authenticated using short tandem repeat (STR) profiling and confirmed to be free of mycoplasma contamination. We grew the cell line in growth medium of RPMI 1640 supplemented with 10% fetal bovine serum and 1% penicillin-streptomycin in a 37 °C, 5% CO2 incubator. H1299 cells were mixed with trypan blue and counted with hemocytometer. Suspension with a concentration of 1 × 106 cells/50 μL was prepared for injection at 50 μL per mouse. Specifically, 8-week-old BALB/c nude mice mouse was firstly anesthetized through intraperitoneal injection of ketamine/xylazine mixture. A transverse incision of ~1 cm was made ~0.5 cm below the inferior border of the left scapula along the midline long axis of the left lateral side of the mouse chest and respiring lung could be observed as a pale pink structure after removing soft tissue. Cancer cell suspension was then injected to a depth of approximately 5 mm into the left lobe of the lung between the 6th and 7th rib where mouse lung was wider and thicker at this level. All procedures were conducted with approval from the Animal Use and Care Committee at Zhejiang University (ZJU20190076) and in accordance with relevant guidelines/regulations.
The in vivo imaging of the mouse lung tissue was performed 3 weeks after the cancer cell implantation when the primary tumor had been established but metastatic spread had not yet begun. Then in vivo imaging was conducted under isoflurane anesthesia, and surgical exposure procedure was the same as that during cancer cell injection. The skin, muscle and ribs were removed exposing the lung tissue for imaging. Care was taken to avoid damaging the surrounding tissue, including blood vessels and nerves. Using blunt forceps, the lungs were gently retracted to expose the region of interest containing the cancer tissue. A glass coverslip was then placed over the exposed area, and a minimal amount of cyanoacrylate-latex glue was used to secure the mouse’s skin surrounding the exposed tissues to the coverslip. Special attention was given to ensure that cancer tissues were protected from direct contact with the glue. The mouse was then positioned and secured on a custom-built, inverted MPM stage. The careful retraction of lung tissue, along with the skin-coverslip adhesion and the weight of the mouse, helped minimize motion artifacts caused by breathing. Mouse body temperature was maintained using a heated stage. Imaging commenced within 10 min of anesthesia induction and typically continued for an additional 30 min. Anesthesia was induced with 3% isoflurane for 2–3 min, then maintained at 1.5%. Three fields were imaged per animal. During the imaging process, if the image quality, observable on the screen, degraded because of breathing motion, we stopped the image capture and started a new acquisition to guarantee optimal contrast and resolution. After imaging, the mice were euthanized using isoflurane anesthesia followed by cervical dislocation.
A total of 8 BALB/c nude mice were involved in this study, with half of them randomly assigned to be implanted with H1299 cells as cancer group and the other half kept as control group, using a computer-generated random number sequence. Note that for imaging of normal controls, the imaged regions were located at similar areas as tumor locations in cancer group for reliable comparison. After in vivo imaging, lung tissues were harvested and sectioned for histology to validate cancerous or normal ones by pathologists blinded to the treatment groups, with sample labels coded by another independent researcher. No animals were excluded from this study.
3D orientation algorithm
3D orientation was defined by the azimuthal angle \(\theta \) and the polar angle \(\varphi \), both ranging from 0° to 180°, and \(\varphi \) could be further resolved by the equation:
where \(\beta \) and \(\gamma \) were two extra azimuthal angles (Supplementary Fig. 1)36. In this way, 3D orientation was defined by three azimuthal angles, which could be regarded as the projection of this orientation on three planes in a Cartesian coordinate system. For orientation calculation, the thickness (or diameter) of the fiber was a crucial factor when determining the window size. Taking the average diameter of elastin fibers as an example, which was approximately 7 voxels, we generated a series of simulated 3D fibers and evaluated the calculation accuracy using windows of different sizes (Supplementary Fig. 2). Specifically, we obtained the orientation (including \(\theta \) and \(\varphi \), Supplementary Fig. 2a) and compared it with the ground truth to obtain the error level (Supplementary Fig. 2b). As can be seen, the calculation accuracy remained basically unchanged after the window size exceeded twice the diameter (i.e., ~ 15 voxels in this case). To balance accuracy and computational efficiency, and also because the diameter of elastin fibers was generally larger than that of collagen fibers, we chose 15 voxels as the window size for orientation calculation. Then all vectors passing through the center voxel were generated and weighted by two factors \({W}_{1}\) and \({W}_{2}\):
where \({W}_{1}\) weighted the vector by the inverse of the vector length \(L\). \({W}_{2}\) weighted the vector by the intensity variations, where \({a}_{1}\), \({a}_{2}\), \({a}_{3}\) were intensities of the central voxel and the two symmetrical voxels (to the central one) along the vector, and \(\bar{a}\) was the average intensity of the three voxels36. The azimuthal orientation of the central voxel was defined as the direction of the sum of all weighted vectors, and 3D orientation was obtained after the determination of the three azimuthal angles.
3D directional variance algorithm
3D directional variance measured alignment of fiber structures in a certain region around the center voxel based on 3D orientation information37. Directional variance was a normalized metric ranging from 0 to 1, with 0 revealing perfectly parallel alignment, while 1 corresponding to complete disorder. Directional variance \({\bar{D}}_{3D}\) was defined as:
where:
and:
where \(\varphi \), \(\theta \), \(\beta \) and \(\gamma \) were orientations described above, and \(k\) was the number of fiber voxels in the region. In this study, we chose the same window size as for orientation quantification to calculate directional variance for optimal performance37.
Waviness algorithm
Waviness measured the bending degree of the fibrous structure based on the 3D orientation information as well38. It ranged from 0 to 1 with higher values corresponding to curvier morphology. To determine waviness of a certain voxel, a region of interest (generally a 3D cube window) was generated first around the voxel and the orientation difference \({\delta }_{\theta }\), \({\delta }_{\beta }\) and \({\delta }_{\gamma }\) were calculated between the central voxel and non-central voxels for \(\theta \), \(\beta \) and \(\gamma \) orientation. The optimal window size38 for waviness calculation was between 3 and 4 times the fiber diameter and we chose a size of 25 × 25 × 25 voxels (considering similar voxel size along xy and z directions). Orientation difference \(\delta \) needed to be modified to its absolute value when it was negative and modified to its supplementary angle when it was over 90°. Finally, the waviness \({W}_{3D}\) of the central voxel was calculated as:
where \(k\) was the number of non-central fiber voxels within the region. Calculation of waviness was based on orientation information; therefore, the accuracy of waviness depended basically on the accuracy of the orientation. It was worth mentioning that the calculation window needed to contain enough fiber information to truly reflect the characteristics such as the bending degree of the fiber (waviness). Supplementary Fig. 2c shows different window sizes compared with fiber diameter in a simulated fiber image. As can be seen from this illustration, a window 3–4 times the fiber diameter can contain relatively complete information at the bending site, consistent with our previous choice for waviness determination38. Therefore, we used 25 voxels (about 3.5 times the elastin fiber diameter) as the window size for waviness calculation.
Local coverage algorithm
Local coverage quantified the localized voxel-wise distribution of fibrous structure40. Segmentation of fibrous structure in MPM images needed to be implemented before local coverage quantification. As a pre-processing step, we applied intensity normalization to all the MPM stacks. Owing to the purely endogenous contrast of SHG and TPEF, background noise was very weak. Therefore, we set the threshold to be 0.1 for these normalized SHG and TPEF image stacks and generated the binary mask accordingly for the calculation of local coverage. Local coverage \({L}_{3D}\) was defined as:
where \(M\) was the segmented mask of fibrous structure in the local region around the center voxel, \(A\) was the entire region, and \(N\) was the number of voxels within the region. The size of the local region was chosen as the same as that for calculating waviness for optimal performance.
Thickness algorithm
Thickness measured the diameter of the fiber structure39, and was quantified for elastin only in this study because the overlapping and intertwining of collagen made it difficult to extract individual fibers. Thickness quantification was also based on the segmentation of fibrous structure. To calculate thickness, minimum distance matrix of each voxel to the background was firstly generated. Then adaptive distance transmission was performed so that voxels perpendicular to the fiber at each distinct location had approximately the same value. Next, smooth correlation operator of segmented voxels was used to smooth the distance transmission matrix to finally obtain thickness for all the fiber voxels.
Similarity coefficient (SC)
The development of similarity coefficient (SC) was inspired from a previous study which measured how vimentin intermediate filaments templated microtubule networks to enhance persistence in directed cell migration48, and we extended such quantification from previous 2D to 3D context. SC characterized resemblance between collagen and elastin fibers based on the spatial location and 3D orientation information of voxels from both fiber components. Generally, we assumed that fibers were similar if they had close spatial locations and consistent orientations. To quantify SC, orientation matrices were firstly obtained for collagen and elastin fibers, including azimuthal angle \(\theta \) and polar angle \(\varphi \). Next, the closest elastin voxel in the 3D stack was found for every collagen voxel and the minimum distance matrix \(d\) was generated. Notably, we could also center on elastin voxels and find the nearest elastin voxel. Then included angle \(\alpha \) was calculated between the paired collagen and elastin voxels based on the voxel-wise orientation information, defined as:
where \({\theta }_{e}\) and \({\varphi }_{e}\) were azimuthal and polar angle of the elastin fiber, while \({\theta }_{c}\) and \({\varphi }_{c}\) were that of the collagen fiber. \(\alpha \) needed to be modified to its absolute value when it was negative and modified to its supplementary angle when it was over 90°. After the determination of \(\alpha \) and \(d\) matrix for all the collagen voxels, we then defined SC as:
where \(e\) was the natural constant and \(D\) was the maximum distance for the two fiber components to be assumed similar, set at 20 voxels. When the minimum distance \(d\) was over \(D\), SC would be 0. SC was also a voxel-wise metric ranging from 0 to 1, and a higher value revealed a more similar structure between collagen and elastin fibers.
Training of the fiber voxel dictionary
To take the advantage of complementary insights and information abundance provided by voxel-wise fiber metrics, we drew on and refined a method from texture analysis49,50,51 to construct fiber voxel dictionary via K-means clustering. Each collagen voxel corresponded to a 4 × 1 numerical vector with directional variance, waviness, local coverage and SC while each elastin voxel corresponded to a numerical 4 × 1 vector with directional variance, waviness, local coverage and thickness. The thickness metric was normalized to the maximum value. A total of 1 million voxels were randomly extracted from 100 normal and 100 cancerous samples of 94 patients. To ensure data balance during the training of fiber voxel dictionary, the number of voxels from normal and cancer tissues was the same (i.e., 500,000 voxels). Then all the voxels were implemented with K-means clustering using MATLAB. K-means ++ algorithm was employed for cluster center initialization and Euclidean distance was used to measure distance between voxels since all metrics were normalized.
The selection of K value, which was also the number of fiber vocabularies, was important for its potential impact on the diagnosis performance of the fiber voxel dictionary. We tested different K values from 100 to 1050 and compared the diagnosis performance with a series of indexes including accuracy, AUC, sensitivity, specificity, precision and F1-score (Supplementary Tables 1–3). We found that the impact of K value on classification performance was not significant in the tested range and chose K = 250 for both collagen and elastin fiber voxel vocabularies considering relatively good classification performance and model complexity.
Cancer risk index (CRI) quantification
CRI reflected the lung cancer risk of fiber vocabularies. To get CRI, we firstly obtained the averaged fiber vocabulary distributions \(V\). We assumed that if a vocabulary appeared frequently in cancer samples and had a low probability present in normal samples, it carried a risk of cancer, and conversely it processed a nature of safety. To meet this end, we employed Spearman’s rank correlation coefficient between averaged vocabulary distributions (\(V\)) and tissue state (\(S\), binary value with −1 for normal tissues and 1 for cancer tissues). Spearman’s rank correlation coefficient could measure the correlation between two variables and was suitable for data with non-normal distribution. For each vocabulary, CRI was defined as:
where \(n\) was the number of tissue samples, and \({d}_{i}\) was the absolute difference between the rank of \({V}_{i}\) and that of \({S}_{i}\).
Correlation assessment between fiber vocabularies
We analyzed correlation between different fiber vocabularies and generated a color map to demonstrate the correlation. To get this result, we firstly got an \(n\times 1\) vector for every fiber vocabulary and values in the vector represented the voxel proportions of this vocabulary in all lung tissue samples, where \(n\) was the number of samples and \(n=358\). Then we got an \(n\times k\) matrix where \(k\) was the number of fiber vocabularies and \(k=250\) for both collagen and elastin fibers. Next, we calculated correlation pairwise Pearson’s correlation coefficient matrix (with a size of \(k\times k\)) between columns in the input \(n\times k\) matrix \(I\). The formula of Pearson’s correlation coefficient matrix \(P\) was:
Where \(i\) and \(j\) were two column indices (also the fiber vocabulary indices) in the input \(I\) matrix. \(P(i,j)\) represented the correlation of frequency of appearance in the same sample between two vocabularies and ranged from −1 to 1. The larger the value, the more likely the two vocabularies were to appear in the same sample. Color maps were then generated based on the matrix \(P\).
Spectra measurement of collagen and elastin fibers
Spectra measurement was based on an established protocol52. Specifically, collagen fibrillar gels were fabricated using Type I collagen isolated from rat tail tendons, serving as the collagen sample. Elastin powder (E7152, Sigma-Aldrich) derived from human lung tissues through non-degradative extraction was utilized as the elastin sample. The elastin powder was rehydrated with a single drop of water and subsequently mounted between a microscope slide and a coverslip for further analysis. The experimental setup for spectra measurement was shown in Supplementary Fig. 3. The spectra of these samples were measured using a central wavelength of 800 nm (the same wavelength as that used for imaging). Laser power was controlled by λ/2 plate and Glan prism. TPEF signals were obtained and assessed, and spectrum was normalized to the maximum intensity for comparison.
Statistics and reproducibility
Two-tailed Mann–Whitney tests were used to determine significant differences. Linear SVM model was used to construct diagnosis tool of lung cancer tissues based on vocabulary distributions obtained through fiber voxel dictionary. For the binary classifier of lung cancer and normal samples, training set included 200 tissue samples (100 normal and 100 tumor samples) and test set included 284 samples (126 normal and 158 tumor samples). For the three-way classifiers of normal, cancer and boundary samples, considering the relatively small number of the boundary samples, we applied the leave-one-out method for cross validation and ROC curves were obtained in a one-vs.-all way.
Results
Workflow of AI-ECM assessment
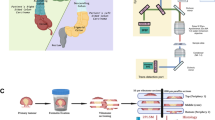
We proposed an AI-ECM workflow to gain insights into ECM (mainly collagen and elastin fibers) remodeling during lung cancer progression and demonstrated its potential in intraoperative assessments (Fig. 1). We developed a parallel-acquisition MPM system which optimized excitation/emission settings to enable simultaneous image acquisition of both fiber components to minimize time burden. Specifically, SHG images of collagen fibers were harvested at 800 nm excitation and 400 ± 10 nm emission, and TPEF images were collected at 800 nm excitation and 525 ± 25 nm emission. Such settings guaranteed efficient collection of auto-fluorescence from elastin fibers while neglecting fluorescence from collagen fibers, according to measured spectra at 800 nm excitation (Fig. 1b).
a AI-ECM is proposed for rapid intraoperative assessments. The feasibility of this method is tested from surgically excised lung tissues from two hospitals. b An MPM system is developed for rapid concurrent SHG imaging of collagen fibers and TPEF imaging of elastin fibers. Then different fiber metrics are extracted from MPM images for cancer risk assessments through fiber voxel dictionary proposed in this study. Cancer risk maps of collagen and elastin are then generated, with red hues revealing risk of lung cancer and blue hues corresponding to normal ECM structure. Merged cancer risk map is then prepared by combining contributions from two fiber components, which provides further optimism of intraoperative application in tumor identification, margin detection and surgery guidance. c Fiber metrics include directional variance, waviness, local coverage, thickness and similarity coefficient (SC) for collagen and elastin fibers. Representative 3D stacks of different features are displayed along with distributions of metric values. Scale bar: 100 μm. d Rapid AI-ECM results can be used to guide the subsequent biopsy and cryosectioning for improved intraoperative diagnosis. Scale bar: 100 μm.
With the approval of the institutional review board, ex vivo MPM imaging was performed on human lung tissue specimens of 222 patients from 2 hospitals (Fig. 1a), with over 60% of cancer samples being localized cases at early stages. To gain a comprehensive understanding in morphology and organization of both fiber components, we performed multi-parametric analysis (Fig. 1c) based on MPM images by extracting voxel-wise features including orientation36, alignment (represented by directional variance metric)37,53, waviness38,54, density (represented by local coverage metric)40 and thickness39 (Supplementary Figs. 4 and 5). All the metrics mentioned above offered voxel-wise features, and applied to images of both collagen and elastin fibers, except that thickness was not quantified for collagen fibers since intertwining of collagen bundles made it difficult to resolve the exact diameter of individual fibers. Next, we developed a voxel-wise quantitative metric, similarity coefficient (SC), to describe the interaction between collagen and elastin during the remodeling process caused by tumor progression (Fig. 1c). Then we integrated SC and other voxel-wise fiber features to produce a voxel dictionary for MPM images through unsupervised learning to achieve explainable assessments of lung cancer risk. Rapid AI-ECM results might guide subsequent biopsies or surgeries for improved intraoperative applications (Fig. 1d).
Similarity coefficient for assessing interaction between collagen and elastin
Spatial overlap of collagen and elastin fibers and the resemblance in orientation were found in certain regions of MPM images, which implied potential interaction between the two fibers. Inspired by a previous parameter which quantified how intra-cellular components templated each other in directed migration within 2D images48, we developed a metric, termed 3D similarity coefficient (SC), to quantitatively evaluate collagen-elastin resemblance in a 3D context (Fig. 2a). To quantify 3D SC, voxel-wise orientations (including azimuthal angle \(\theta \) and polar angle \(\varphi \)) were obtained for collagen and elastin fibers (Fig. 2b and Supplementary Fig. 6). Then the nearest elastin voxel in the 3D stack was found for every collagen voxel and the included angle was calculated between the paired voxels based on the orientation information. Next, 3D SC value was determined by the product of the negative correlation linear function of voxel distance and the logarithmic decay function of the included angle (detailed in “Methods”). 3D SC was also a normalized voxel-wise metric, with the color-coded map enabling direct visualization of the similar region (Fig. 2c and Supplementary Video 1).
a Quantification process of 3D SC. b \(\theta \) distributions of collagen and elastin fibers within dotted area in (a). c Representative 3D stacks of MPM intensity and corresponding SC maps. Scale bar: 100 μm. d Box plots comparing SC in normal and cancer samples. n = 222 patients. ***p = 4.8e−14. e Demonstration of similar region (SR) and non-similar region (NSR) with corresponding SC map. f \(\theta \) distributions of both fiber components within SR. g \(\theta \) distributions within NSR. Box plots comparing h collagen directional variance (***p = 2.6e−16), i collagen local coverage (**p = 0.006), j elastin waviness (***p = 5.2e−15), and k elastin local coverage (***p = 1.8e−9) between NSR and SR. Each point corresponds to a tissue sample. n = 222 patients. l Scatter plot showing correlation between collagen alignment and SC. DV directional variance, Wav waviness, LC local coverage, Thi thickness, Ela elastin, Col collagen. Two-tailed Mann–Whitney tests were used to determine significant differences.
We quantified 3D SC for all the lung samples and compared mean SC between normal and cancerous tissues. A significant higher SC was found in cancerous tissues, implying that collagen and elastin resembled each other with tumor progression (Fig. 2d). We further assessed alignment, distribution and waviness features in similar region (SR) and non-similar region (NSR) for both normal and cancer samples. Specifically, SR was defined as regions with SC over 0, while SC was equal to 0 in NSR caused either by large distance between them (over 18 μm) or included angles being over 60° (see “Methods”, with representative regions shown in Fig. 2e). In most areas of NSR, the non-overlap of collagen and elastin fibers made them hard to interact with each other; while in certain NSR like the one marked in the dotted box (Fig. 2e), both collagen and elastin existed yet their orientations did not show uniformity in contrast to that within SR, as evident from \(\theta \) distributions (Fig. 2f, g), which also indicated weak connections between the two fibers. Collagen and elastin features including directional variance, waviness, local coverage and thickness were compared between SR and NSR for both normal and cancerous samples. We observed significant difference in collagen alignment, elastin waviness, and density of both fibers between SR and NSR in tumor tissues (Fig. 2h–k and Supplementary Fig. 7), revealing that morphological changes of two fibers was influenced by the mutual interaction between them. However, this phenomenon was not found in normal tissues (Supplementary Fig. 8). Meanwhile, trends of change in fiber morphology from normal to tumor tissues were identical in SR and NSR (Supplementary Fig. 9). These results revealed that the remodeling of ECM might be influenced by dual factors including tumor progression and interaction between the two fiber components. Elastin waviness was found to be lower in SR (Fig. 2j), indicating that straighter elastin was more prone to interact with collagen. We also observed a higher elastin density in SR compared with that in NSR (Fig. 2k), yet this density difference was less significant than that of collagen (Fig. 2i), probably due to the discrepancy in turnover rates between them22. Further we assessed the correlation between SC and other metrics (Supplementary Fig. 10), and found significant negative correlation between SC and collagen alignment only (Fig. 2l), implying that the mutual interaction influenced greatly on the rearrangement of collagen fibers.
Construction of fiber voxel dictionary via unsupervised learning
So far, we obtained a series of metrics to quantify morpho-structural characteristics of collagen and elastin fibers, and developed SC to establish a connection between them (more examples shown in Supplementary Fig. 11). To better analyze ECM remodeling caused by tumor progression, fiber metrics needed to be integrated to provide complementary information from different perspectives to ensure the accuracy and comprehensiveness. Meanwhile, the advantage of information abundance from voxel-wise manner was also needed to be fully utilized for an improved sensitivity and specificity in tumor identification. To this end, we constructed a fiber voxel dictionary through unsupervised learning of voxel-wise metrics extracted from lung samples (Fig. 3). A total of 100 normal and 100 cancerous samples from a total of 94 patients were prepared for the training of voxel dictionary, with a total of 1 million voxels from both collagen and elastin fibers forming the training set. Metrics were categorized as collagen or elastin-related ones, and SC was included in the collagen category for convenience, although it contained information of both components (Fig. 3a). Then all the voxels were processed with K-means clustering to obtain 250 cluster centers for collagen and elastin fibers respectively (Fig. 3b), termed as fiber vocabulary which built up the fiber voxel dictionary (Fig. 3c). For a new MPM image stack (Fig. 3d), each voxel was subsumed into a certain vocabulary based on its voxel-wise metric values according to the dictionary, and then this image stack could be translated into a distribution of fiber vocabularies (Fig. 3e).
a Data preparation process for dictionary training including fiber metric calculation and voxel extraction. b Voxel clustering through K-means method (the actual number of fiber metrics is greater than three and here 3D metric space is used as a representative). c Fiber voxel dictionary is constructed from fiber vocabularies. d Preparation of a new sample image stack for the translation through fiber voxel dictionary. Scale bar: 100 μm. e Translated fiber vocabulary histograms of a representative lung tissue image stack. f SVM model designed for lung cancer diagnosis based on fiber vocabulary histogram results. g ROC and AUC (area under the curve) results of lung cancer diagnosis SVM model using collagen/elastin dictionary alone and both. h Classification performance of SVM model including accuracy, sensitivity and specificity with 95% confidence interval (CI), using features of both fiber components. DV directional variance, Wav waviness, LC local coverage, Thi thickness, Ela elastin, Col collagen.
We further tested the potential of fiber voxel dictionary in lung cancer diagnosis. A total of 484 lung samples from 222 patients, including 226 normal and 258 lesion-level tumor tissues, were translated to distributions of fiber vocabularies according to the voxel dictionary. These samples were further divided into training set (100 normal samples and 100 tumor samples from 94 patients) and test set (126 normal samples and 158 tumor samples from 128 patients) to be adopted in a support vector machine (SVM) (Fig. 3f). Patient information in training and test set is detailed in Table 1. There were no significant differences in tumor status (including primary site, tumor size and lymph node invasion) and mean fiber characteristics (directional variance, waviness, local coverage, thickness and SC) between training and test set.
Diagnosis results from fiber voxel dictionary were illustrated by receiver operating characteristic (ROC) curves (Fig. 3g) and outputs in accuracy, sensitivity and specificity for data from two hospitals by combining features from collagen and elastin fibers (Fig. 3h), indicating that the proposed dictionary-based method led to excellent performance in lung cancer diagnosis and complementary insights from both fiber components are helpful. Notably, a sensitivity of 99.37% (95.32–99.98, 95% confidence interval) was achieved. The average time of lung cancer diagnosis for each sample was below 10 min including MPM imaging and AI-ECM (fiber metric extraction and translation via fiber voxel dictionary) analysis of three 3D image stacks. Considering different imaging depths of 3D stacks (70–100 μm), there might be a time variation of about 2 min.
Cancer risk assessment from AI-ECM
To maximize the voxel-wise translation capability of AI-ECM for intuitive and user-friendly cancer risk assessment, we developed cancer risk index (CRI) to evaluate the response of different fiber vocabularies to cancerous and normal states. Fiber vocabulary distributions of all the samples were collected, and then Spearman’s rank correlation analysis between proportion of each fiber vocabulary and the binary tissue state (cancer or normal) was implemented to get CRI ranging from −1 to 1 for all collagen and elastin vocabularies (see “Methods”). Positive CRI values indicated that the corresponding vocabulary was more responsive to cancer state (Fig. 4a), while negative CRI ones implied a lower probability of occurrence in cancerous tissues thereby possessing a safety nature (Fig. 4a). To demonstrate the response of vocabularies, we obtained the average voxel proportion of all vocabularies for normal and cancer samples, with vocabularies sequenced and colored according to its CRI value for collagen (Fig. 4b) and elastin (Fig. 4c). For both vocabularies, a higher CRI (closer to 1) indicated that the corresponding vocabulary appeared more frequently in cancer samples and less frequently in normal sample, while vocabularies with CRI close to −1 showed the opposite. Voxel proportions of vocabularies with CRI near 0 were similar in cancer and normal tissues, revealing no obvious inclination to either state. Based on these results, we sorted vocabularies into cancer risk category with CRI higher than 0.5, safety category with CRI lower than −0.5 and non-significant one of the rest, and obtained correlation of voxel proportion between vocabularies and results (Fig. 4d, e). Strong positive correlation was found among cancer risk vocabularies, which implied that these vocabularies often appeared simultaneously in cancer samples, and similar results were also found in safety vocabularies.
a Demonstration of color map for cancer risk index (CRI). Average voxel proportion of collagen (b) and elastin (c) vocabularies, sequenced and colored according to CRI. Correlation analysis of voxel proportion between different vocabularies in collagen (d) and elastin (e), with correlation coefficients color-coded. f Comparison of average metric value between cancer risk and safety vocabularies in collagen, including directional variance (***p = 2.7e−5), waviness (**p = 0.0017), local coverage (***p = 1.2e−20), and SC (*p = 0.024). n = 222 patients. g Comparison of average metric value between cancer risk and safety vocabularies in elastin, including directional variance (***p = 5.2e−9), waviness, local coverage (***p = 7.8e−18), and thickness (***p = 1.7e−17). n = 222 patients. Mean and standard deviation are calculated. Two-tailed Mann–Whitney tests were used to determine significant differences. h Typical MPM images and corresponding CRI maps of cancer and normal lung tissues. Scale bar: 100 μm. DV directional variance, Wav waviness, LC local coverage, Thi thickness, Ela elastin, Col collagen.
To assess the structural remodeling of collagen and elastin from tumor progression, we extracted the mean metric value by cancer risk and safety vocabularies (Fig. 4f, g). In collagen fibers, directional variance was higher, while waviness and local coverage were lower in cancer risk vocabularies, revealing that collagen fibers became less aligned, straighter and sparser with lung tumorigenesis (Fig. 4f). In elastin fibers, local coverage and thickness were lower while directional variance was higher in cancer vocabularies, indicating sparser, thinner and less aligned elastin fibers were exclusive in cancer samples (Fig. 4g). SC in cancer vocabularies was higher, indicating that an inclination of resembling each other between these two fibrous components was also an important hallmark of cancer risk (Fig. 4f), consistent with above-mentioned mean SC results (Fig. 2d). It is worth mentioning that the change trends of some fiber metrics from safety to cancer risk vocabularies were identical to that from SR to NSR (Fig. 2h–k), including LC for both fibers and directional variance in collagen, which again revealed that the ECM remodeling was influenced by dual factors including tumor progression and interaction between the two fiber components.
We further generated CRI maps for understandable and explainable visualization to clinicians to highlight translational potential of AI-ECM. Every voxel within a lung tissue image could be translated into a certain vocabulary through fiber dictionary and color-coded according to CRI. From representative MPM images and corresponding CRI maps of cancer and normal lung tissues (Fig. 4h and Supplementary Videos 2 and 3), CRI values of almost the whole image in normal tissues were negative (blue hues) while major areas in cancer samples were covered by positive CRI values (red hues). Such contrasts ensured feasibility of CRI assessment and provided intuitive cues of cancer risk. Through CRI maps, risky regions could be highlighted to aid assessment of tissue remodeling from lung cancer progression; meanwhile, CRI maps offered potential explainability for diagnosis decisions of AI-ECM and further assisted clinicians to evaluate potential malignant regions if necessary. By combining fiber voxel dictionary and CRI assessment, with the former offering diagnosis results with high sensitivity and specificity and the latter offering user-friendly intuitive cancer risk visualization, further optimism was expected for application of AI-ECM into lung cancer detection.
Detection of tumor-normal boundary with AI-ECM
To assess the ability of AI-ECM in tumor boundary identification, newly-collected 23 boundary samples, 25 cancerous samples and 20 normal samples from extra 20 patients were adopted in the study and analyzed using pre-trained fiber voxel dictionary. Based on the vocabulary distribution, we constructed one versus all SVM model, and leave-one-out method was used for classification among boundary, normal and cancer samples considering the relatively small data size of boundary tissues (Fig. 5a). High AUC values in identifying each tissue type were obtained, indicating that AI-ECM could sensitively uncover subtle structural remodeling of ECM in tumor-normal border areas. Boundary samples were not involved in training of the dictionary yet were still well-classified based on the vocabulary distribution, verifying the generalizability of fiber voxel dictionary.
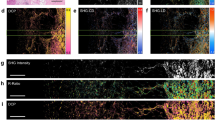
a ROC curve of one vs. all SVM model. b Vocabulary distributions for collagen and elastin fibers in boundary. c Representative H&E stained histological image of lung cancer tissues. Scale bar: 1 mm. Solid black line marks the border between cancerous (upper left) and normal (lower right) regions, and dashed box represents one MPM image field. d The MPM image corresponding to the dashed box in (c). Scale bar: 100 μm. CRI maps of collagen (e) and elastin (f) fibers corresponding to the MPM image. g The merged collagen and elastin CRI map. h Establishment of murine orthotopic model through implantation of human lung cancer cells. i Schematic of in vivo imaging protocol. j Typical 3D MPM stack obtained from in vivo imaging. Scale bar: 100 μm. k Boxplots showing comparison of fiber metrics between normal and cancer samples, including collagen directional variance (***p = 1.0e−4), collagen waviness (***p = 2.0e−4), collagen local coverage (***p = 7.4e−7), SC (***p = 1.0e−6), elastin directional variance (***p = 1.4e−4), elastin waviness (***p = 2.0e−4), elastin local coverage (***p = 5.0e−6), and elastin thickness (***p = 2.2e−5). A total of 12 MPM volumes for normal tissues and 12 ones for cancer tissues are captured with n = 4 mice for each group. DV directional variance, Wav waviness, LC local coverage, Thi thickness, Ela elastin, Col collagen. Two-tailed Mann–Whitney tests were used to determine significant differences.
We then established responses of different vocabularies to boundary, and found that neither risk nor safe vocabularies were dominant in boundary samples as evident from relatively uniform vocabulary distribution (Fig. 5b), in contrast to that in normal or cancer samples (Fig. 4b, c). Also we noticed that safe vocabularies were more responsive compared with risk ones in collagen fibers, indicating that collagen characteristics in boundary areas were closer to normal rather than cancerous tissues. This phenomenon was not found in elastin fibers, revealing that tumor progression might affect elastin remodeling at a higher level in boundary areas.
Next, we obtained CRI maps at boundary region. On a representative H&E stained histological image, the surgeon marked the border between normal and cancerous regions (Fig. 5c). A typical field along the border was then imaged by MPM (Fig. 5d) with the border line highlighted. According to CRI maps of collagen (Fig. 5e) and elastin (Fig. 5f) fibers, as well as the merged one (Fig. 5g), different CRI values were observed on two sides of the border line, revealing that tissues on one side was at risk of cancer (red hues) while the other side tended to be at normal state (blue hues). Meanwhile, tissues with gray hues (no inclination) also occupied a considerable portion in the map. Determination of the tumor boundary was important yet challenging for clinicians to decide the extent of the excision, and advantages of high resolution of MPM, high sensitivity of AI-ECM and intuitiveness of CRI map provided further optimism for accurate boundary identification.
In vivo imaging of a murine model harboring human lung cancer
To test the translational potential of MPM and ECM feature extraction, we implanted H1299 cells, a kind of human non-small cancer cell line, into mouse lung to establish murine orthotopic model as object of in vivo imaging (Fig. 5h). A total of eight mice were involved, with half of them implanted with H1299 cells as cancer group and the other half as normal controls. Following thoracotomy surgery, we performed in vivo imaging using the developed MPM system for rapid data collection (see “Methods”). During the imaging process, we used combined strategies including anesthesia and tape-based fixation to minimize motion artifacts (Fig. 5i). Then we extracted collagen and elastin features from MPM volume images (Fig. 5j). Statistical analysis results revealed that although there were small differences in the absolute values of each metric, identical trends were found from in vivo study of the murine model as that obtained from ex vivo imaging of human lung tissues (Fig. 5k). Specifically, collagen and elastin fibers were less aligned, straighter, sparser while more similar in cancerous tissues than normal ones. Significance in waviness of elastin fibers was obtained from in vivo study, while not found in ex vivo case. These preliminary in vivo imaging and quantification results demonstrated that MPM imaging along with quantitative ECM characterization might be potentially applicable to intraoperative scenarios.
Discussion
Due to the limitations of existing techniques in the context of guiding clinicians accurately to the correct tumor location and identifying the exact tumor boundary for biopsy intraoperatively, in this study we develop MPM system that enables fast, label-free and high-resolution imaging of collagen and elastin fibers. Fine structures of these fiber components can be resolved at sub-micron level, without any exogenous agents55,56. Specifically, MPM is able to capture images from collagen and elastin simultaneously instead of sequentially, while with minimum crosstalk by optimizing the excitation/emission settings according to measured fluorescence spectra. Especially, the entire time needed, including the imaging and AI-ECM analysis of three 3D image stacks, can be controlled below 10 min in this work. For intraoperative applications, sparse imaging of 8–12 sites on the tumor boundary can be completed in 20–25 min. Moreover, the time cost can be further optimized by using detectors with higher sensitivity and/or by reducing analysis time through parallel computing. Therefore, our method might be more time efficient in tumor detection and boundary identification than cryosectioning, which typically requires 30 min or more. Considering the translational potential, we propose a combined use of MPM and existing techniques such as thoracoscopy, with the latter offering insights into areas that are grossly abnormal from large-field view and the former confirming the target area for subsequent biopsy or surgery with a high accuracy.
Biomechanical properties of connective tissues are regulated via cellular signaling to provide normal function57, especially in lung where delicate architecture needs to be maintained to provide optimized interface for efficient gas exchange. Based on the simultaneously present collagen and elastin signals, we develop similarity coefficient (SC) in this study to assess the resemblance level in morpho-structural characteristics based on their spatial distance and orientation information. Notably, SC can be assessed in a voxel-wise way to sensitively identify the subtle alterations within local regions. We find that the similarity is significantly higher in cancerous tissues in contrast to normal ones, revealing that lung cancer progression accompanies remodeling of these two fiber components in a way towards resembling and templating each other. Elastin is an important component of blood vessels and a higher SC in tumor tissues might be possibly related to tumor-associated angiogenesis. Additionally, collagen-elastin similarity is also potentially associated with stromal stiffening, which is a cardinal alteration of biomechanical properties in tumor tissues26. Fragmented elastin architecture revealed by lower thickness (Fig. 4g) in our study diminishes elastic energy storage, and collagen fibers that proliferate with elastic fibers as anchor points and templates (high SC) provide physical buffering, further weakening tissue rebound and leading to increased stiffness, as commonly found in lung cancer58. Stromal stiffening would further activate mechanosensitive pathways (e.g., YAP/TAZ) in cancer-associated fibroblasts, perpetuating a feedforward loop of ECM remodeling and tumor progression59. Tumor progression is accompanied by abnormal formation of blood vessels, which not only promotes tumor growth, but also enhances cancer cell invasion and metastasis. Collagen fragments such as endostatin, tumstatin, canstatin, arresten, and hexastatin, have shown stimulatory or inhibitory effects on blood vessel generation60. Meanwhile, ECM-related enzymes such as MMPs61 and LOXs62 alter the physicochemical state and morphological characteristics of the ECM to initiate vascular branching. The high homogeneity of collagen and elastin fibers might be the result of ECM remodeling to promote angiogenesis. Previous studies implied that the stiffness of lung tissues also increased with tumor progression58,63, and future efforts will be taken to assess in detail such correlation.
Besides information offered from SC, collagen and elastin fibers reorganize themselves in response to tumorigenesis. Specifically, both fiber components become straighter and less aligned within cancerous tissues in contrast to normal ones. It is worth mentioning that previous study showed different changes of collagen organization in other cancer types. Collagen fibers became more aligned in ovarian cancer64; in breast cancer, orientation of collagen fibers gradually became perpendicular to the tumor border to facilitate the invasion of cancer cells65,66. However, collagen fibers became more disorganized in colon cancer67, which was identical with our observations in lung cancer. Collagen deposition, as represented by an increase in density, was found in breast cancer68 and renal cell carcinoma69 while in ovarian cancer70 and lung cancer in this study collagen density decreased. These results mentioned above revealed that changes in collagen organization were likely cancer-dependent. It was worth mentioning that previous study reported that liver kinase B1 (LKB1)-depletion-driven collagen fibers became more aligned at tumor invasion fronts in 3D spheroid models of lung tumor cells71 while our clinical tissue analysis demonstrated global collagen disorder in non-small cell lung cancer (NSCLC). The observed differences might originate from the study model (in vitro model vs. human tissues), observation scale (local cellular microenvironment vs. organizational level) and the complexity of influencing factors (simplified mechanism vs. combined effects from multiple mechanisms). Specifically, although we obtained statistically decreased alignment (higher directional variance) in collagen fibers from tumor tissues, it was possible that there were certain locations from tumor samples where collagen fibers exhibited comparable or even increased alignment relative to that from normal tissues (Supplementary Fig. 12). Moreover, elastin fibers within cancerous tissues are more fragmented with smaller thickness, probably due to the fiber proliferation caused by ECM disorder72,73. Currently, characteristics represented by quantitative metrics are typically assessed in a way by averaging values within a custom-determined field, which might mask the functional/structural heterogeneity, considered as a very important hallmark of malignant neoplasm74. To fully acknowledge the information abundance from voxel-wise measurements including SC and other morpho-structural metrics, and account for tumor-induced heterogeneity, we draw on and refine a method from texture analysis to construct a fiber voxel dictionary via unsupervised learning49,50,51. Each voxel in the image stack can be translated into a certain fiber vocabulary through the dictionary according to its fiber metrics, and then the image stack can be transferred into a distribution of fiber vocabularies for diagnosis of lung cancer, which exhibits excellent performance in classifying cancerous tissues from normal ones. Different from texture analysis, a main advantage of our method is that it is completely explainable, with each individual metric clearly representing a specific morpho-structural feature or collagen-elastin interaction.
For machine learning based tools, black box problem has been a major obstacle for its translational application in clinical practice75 and enhancing confidence of clinicians in using it as a basis of intraoperative decision-making is a challenge. This is especially critical for AI-ECM workflow since ECM remodeling has not been a key point of clinical assessment, although it is already widely-accepted as an emerging hallmark of cancer and attracts increasing attention. In this context, we develop CRI to assess the lung cancer risk of fiber vocabularies and evaluate the ECM remodeling caused by tumor progression in a visible way. Owing to the voxel-wise nature, we are able to observe heterogeneous remodeling patterns and enhance explainability of diagnosis decisions made by AI-ECM. CRI maps not only provide intuitive visual cues for tumorigenesis, highlighting the risky regions and making the diagnostic decision understandable to clinicians, but also offer potential to spatially locate lesions and tumor margins, as verified through successfully defining the tumor-normal border. Accurate definition of tumor boundary intraoperatively from thoracoscopy is challenging, and clinicians might rely on experience to decide the location of biopsy, inevitably leading to inaccuracy and subjectivity. Collectively, AI-ECM workflow can provide timely feedback and reliable guidance to ensure proper biopsy decision. Besides, our method also exhibits tremendous potential in determining the extent and scope of surgical resection following diagnosis from cryosectioning.
AI-ECM, based on MPM’s rapid high-resolution imaging capabilities and visible cancer risk maps (CRI), holds important potential for preoperative and intraoperative applications. For preoperative applications, by miniaturizing MPM probes for compatibility with navigational forceps or fine-needle aspiration (FNA) tools, AI-ECM could enable real-time, label-free imaging of ECM remodeling during biopsy sampling. For instance, a fiber-optic MPM probe76,77 could be integrated into the biopsy needle to visualize ECM status at the target site during FNA procedures. This would allow surgeons to dynamically assess cancer risk in suspicious regions, ensuring accurate sampling of high-risk areas and reducing false negatives. Similarly, in navigational bronchoscopy78, AI-ECM-generated CRI maps could be superimposed onto real-time endoscopic images, providing visual cues for tumor boundary identification and guiding targeted cryo-biopsy sampling. For intraoperative applications such as minimally invasive lung resections, AI-ECM could rapidly scan and quantitatively assess the resection margin to distinguish cancerous from normal tissues, which would enhance precision in tumor boundary determination and reduce reliance on postoperative histopathology for margin assessment. For intraoperative frozen section diagnosis, AI-ECM could quickly evaluate ECM characteristics of tissue samples before biopsy to prioritize high-risk areas for frozen sectioning, shorten intraoperative waiting time and improve the efficiency of pathological diagnosis.
The widespread adoption of AI-ECM by pathologists relies on its interpretability, compatibility with existing workflows, and cost-effectiveness. First, the color-coded CRI maps provide intuitive visualizations of ECM remodeling, aligning with pathologists’ familiarity with H&E-stained slides. This user-centric design minimizes the learning cost and allows AI-ECM to complement conventional histopathology. Second, the fully automated analysis pipeline ensures standardized outputs that can be directly integrated into digital pathology platforms or hospital information systems79,80. Finally, cost barriers may be mitigated through scalable MPM technologies (e.g., shared laser systems) and prioritizing high-value applications, such as guiding cryo-biopsies or reducing reoperation rates through tumor boundary analysis.
While our study demonstrates the potential of AI-ECM for intraoperative lung cancer detection, several limitations should be acknowledged. First, our study was limited to lung adenocarcinoma, which is one of the most common subtypes of NSCLC. However, lung cancer is a heterogeneous disease with various histological subtypes81, each of which might exhibit unique ECM remodeling patterns. Expanding the scope of our research to include other subtypes would enhance the generalizability of our findings and provide a more complete picture of lung cancer-induced ECM changes. Besides, inclusion of pre-invasive lesions would be helpful as well, as there is more disagreement among pathologists in diagnosing these samples. Second, since lung adenocarcinoma originates primarily from the epithelium surrounding alveoli, therefore imaging and analysis in this work were primarily focused on alveolar regions. However, assessments of other tissue components within periphery lung, such as larger blood vessels and bronchioles with distinct collagen and elastin fiber organization78, could provide additional insights into lung cancer development. Future work would extend to these components to achieve a more comprehensive understanding of the ECM remodeling. Third, while our AI-ECM framework provides a powerful tool for assessing tumor progression, it is helpful to integrate other biomarkers that are commonly used in lung cancer diagnosis, such as blood-based markers, genetic mutations, or immunohistochemical data. In addition, MPM can obtain label-free fluorescence signals from metabolic coenzymes in cells, such as NADH (reduced form of nicotinamide adenine dinucleotide) and FAD (flavin adenine dinucleotide)43, and previous studies have shown that the extracted metabolic information from them serve as a sensitive biomarker for skin cancer diagnosis42. Combining AI-ECM with these additional biomarkers could potentially improve the sensitivity and specificity of lung cancer detection. Future studies would explore the integration of AI-ECM with multi-omics approaches to develop a more holistic diagnostic framework that leverages both structural and molecular information. Fourth, potential confounding effects to ECM remodeling of comorbid airway diseases, such as chronic obstructive pulmonary disease (COPD) which is prevalent among lung cancer patients82, were not involved in this study. Further studies are needed to validate cancer diagnostic specificity in patients with overlapping COPD-related changes83, which will enhance the robustness and clinical applicability of AI-ECM in diverse patient populations. Finally, while our current study employed standardized imaging volumes (465 × 465 μm FOV, 70–100 μm depth) to establish baseline diagnostic performance, we recognize the critical need to investigate parameter optimization to meet time requirements of intraoperative applications. Future work will systematically explore the resolution-FOV-accuracy tradeoff through assessing resolution-dependent diagnostic performance and establishing adaptive resolution requirements for specific diagnostic tasks.
In summary, we have proposed a framework which integrates MPM imaging and AI-ECM workflow for quantitative characterization of ECM remodeling during tumor progression and verified its potential in intraoperative application of tumor identification and accurate boundary detection for subsequent biopsy. This study paves the way for future clinical trials and sets the stage for technological refinement, which will lead to a promising modality for time-efficient, highly-sensitive, explainable cancer detection with visual cues, improving identification of malignant lesions in lung at early stages and thus patient survival.
Data availability
The source data that make up all graphs in the paper are shown in Supplementary Data 1. Any further data that support the findings of this study are available from the corresponding authors upon request.
Code availability
The custom MATLAB source code for structural assessment of fiber-like structures is available at: https://github.com/ShuhaoQian/Waviness-DirectionalVariance-Orientation_Data_Availability, as well as the Zenodo platform84.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Schabath, M. B. & Cote, M. L. Cancer progress and priorities: lung cancer. Cancer Epidemiol. Biomark. Prev. 28, 1563–1579 (2019).
Leiter, A., Veluswamy, R. R. & Wisnivesky, J. P. The global burden of lung cancer: current status and future trends. Nat. Rev. Clin. Oncol. 20, 624–639 (2023).
Xia, C. et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin. Med. J. 135, 584–590 (2022).
Yasufuku, K. & Fujisawa, T. Staging and diagnosis of non-small cell lung cancer: invasive modalities. Respirology 12, 173–183 (2007).
Landreneau, R. J. et al. Video-assisted thoracic surgery: basic technical concepts and intercostal approach strategies. Ann. Thorac. Surg. 54, 800–807 (1992).
Sihoe, A. D. L. Video-assisted thoracoscopic surgery as the gold standard for lung cancer surgery. Respirology 25, 49–60 (2020).
Hansen, M., Faurschou, P. & Clementsen, P. Medical thoracoscopy, results and complications in 146 patients: a retrospective study. Resp. Med. 92, 228–232 (1998).
Lin, J. & Iannettoni, M. D. The role of thoracoscopy in the management of lung cancer. Surg. Oncol. 12, 195–200 (2003).
Annema, J. T. et al. Mediastinoscopy vs endosonography for mediastinal nodal staging of lung cancer a randomized trial. J. Am. Med Assoc. 304, 2245–2252 (2010).
Wallace, M. B. et al. Minimally invasive endoscopic staging of suspected lung cancer. J. Am. Med Assoc. 299, 540–546 (2008).
Ozyoruk, K. B. et al. A deep-learning model for transforming the style of tissue images from cryosectioned to formalin-fixed and paraffin-embedded. Nat. Biomed. Eng. 6, 1407–1419 (2022).
Lu, P., Weaver, V. M. & Werb, Z. The extracellular matrix: a dynamic niche in cancer progression. J. Cell Biol. 196, 395–406 (2012).
Cox, T. R. & Erler, J. T. Remodeling and homeostasis of the extracellular matrix: implications for fibrotic diseases and cancer. Dis. Model Mech. 4, 165–178 (2011).
Gu, B.-H., Madison, M. C., Corry, D. & Kheradmand, F. Matrix remodeling in chronic lung diseases. Matrix Biol. 73, 52–63 (2018).
Zhou, Y. et al. Extracellular matrix in lung development, homeostasis and disease. Matrix Biol. 73, 77–104 (2018).
Mižíková, I. et al. Collagen and elastin cross-linking is altered during aberrant late lung development associated with hyperoxia. Am. J. Physiol. Lung Cell. Mol. Physiol. 308, L1145–L1158 (2015).
White, E. S. Lung extracellular matrix and fibroblast function. Ann. Am. Thorac. Soc. 12, S30–S33 (2015).
Cox, T. R. The matrix in cancer. Nat. Rev. Cancer 21, 217–238 (2021).
Noda, K. et al. Latent TGF-β binding protein 4 promotes elastic fiber assembly by interacting with fibulin-5. Proc. Natl. Acad. Sci. USA 110, 2852–2857 (2013).
Robertson, I. B. et al. Latent TGF-β-binding proteins. Matrix Biol. 47, 44–53 (2015).
Burgess, J. K., Mauad, T., Tjin, G., Karlsson, J. C. & Westergren-Thorsson, G. The extracellular matrix—the under-recognized element in lung disease?. J. Pathol. 240, 397–409 (2016).
James, D. et al. Probing ECM remodeling in idiopathic pulmonary fibrosis via second harmonic generation microscopy analysis of macro/supramolecular collagen structure. J. Biomed. Opt. 25, 014505 (2019).
James, D., Brereton, C., Davies, D., Jones, M. & Campagnola, P. Examining lysyl oxidase-like modulation of collagen architecture in 3D spheroid models of idiopathic pulmonary fibrosis via second-harmonic generation microscopy. J. Biomed. Opt. 26, 066501 (2021).
Tisler, M. et al. Analysis of fibroblast migration dynamics in idiopathic pulmonary fibrosis using image-based scaffolds of the lung extracellular matrix. Am. J. Physiol. Lung Cell. Mol. Physiol. 318, L276–L286 (2020).
Levental, K. R. et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell 139, 891–906 (2009).
Chen, D. et al. Predicting postoperative peritoneal metastasis in gastric cancer with serosal invasion using a collagen nomogram. Nat. Commun. 12, 179 (2021).
Chen, D. et al. Association of the collagen signature in the tumor microenvironment with lymph node metastasis in early gastric cancer. JAMA Surg. 154, e185249 (2019).
Xi, G. et al. Large-scale tumor-associated collagen signatures identify high-risk breast cancer patients. Theranostics 11, 3229 (2021).
Jain, M. et al. Multiphoton microscopy: a potential “Optical Biopsy” tool for real-time evaluation of lung tumors without the need for exogenous contrast agents. Arch. Pathol. Lab. Med. 138, 1037–1047 (2013).
van Huizen, L. M. G. et al. Compact portable multiphoton microscopy reveals histopathological hallmarks of unprocessed lung tumor tissue in real time. Transl. Biophotonics 2, e202000009 (2020).
Golaraei, A. et al. Characterization of collagen in non-small cell lung carcinoma with second harmonic polarization microscopy. Biomed. Opt. Express 5, 3562–3567 (2014).
Mirsanaye, K. et al. Unsupervised determination of lung tumor margin with widefield polarimetric second-harmonic generation microscopy. Sci. Rep. 12, 20713 (2022).
van Huizen, L. M. G. et al. Rapid on-site histology of lung and pleural biopsies using higher harmonic generation microscopy and artificial intelligence analysis. Mod. Pathol. 38, 100633 (2025).
Rui, L. et al. Advances in nonlinear optical microscopy for biophotonics. J. Nanophotonics 12, 033007 (2018).
Liu, Z. et al. Rapid three-dimensional quantification of voxel-wise collagen fiber orientation. Biomed. Opt. Express 6, 2294–2310 (2015).
Liu, Z. et al. Automated quantification of three-dimensional organization of fiber-like structures in biological tissues. Biomaterials 116, 34–47 (2017).
Qian, S. et al. Mapping organizational changes of fiber-like structures in disease progression by multiparametric. Quant. Imaging Laser Photonics Rev. 16, 2100576 (2022).
Meng, J. et al. Mapping physiological and pathological functions of cortical vasculature through aggregation-induced emission nanoprobes assisted quantitative, in vivo NIR-II imaging. Biomater. Adv. 136, 212760 (2022).
Qian, S. H. et al. Identification of human ovarian cancer relying on collagen fiber coverage features by quantitative second harmonic generation imaging. Opt. Express 30, 25718–25733 (2022).
Liu, Z. Y. et al. Label-free, multi-parametric assessments of cell metabolism and matrix remodeling within human and early-stage murine osteoarthritic articular cartilage. Commun. Biol. 6, 405 (2023).
Pouli, D. et al. Imaging mitochondrial dynamics in human skin reveals depth-dependent hypoxia and malignant potential for diagnosis. Sci. Transl. Med. 8, 367ra169–367ra169 (2016).
Liu, Z. et al. Mapping metabolic changes by noninvasive, multiparametric, high-resolution imaging using endogenous contrast. Sci. Adv. 4, eaap9302 (2018).
Pouli, D. et al. Label-free, high-resolution optical metabolic imaging of human cervical precancers reveals potential for intraepithelial neoplasia diagnosis. Cell Rep. Med. 1, 100017 (2020).
Fischer, F. et al. Assessing the risk of skin damage due to femtosecond laser irradiation. J. Biophotonics 1, 470–477 (2008).
Wilson, A. N., Chen, B., Liu, X., Kurie, J. M. & Kim, J. A method for orthotopic transplantation of lung cancer in mice. Methods Mol. Biol. 2374, 231–242 (2022).
Justilien, V. & Fields, A. P. Utility and applications of orthotopic models of human non-small cell lung cancer (NSCLC) for the evaluation of novel and emerging cancer therapeutics. Curr. Protoc. Pharmacol. 62, Unit-14.27 (2013).
Gan, Z. et al. Vimentin intermediate filaments template microtubule networks to enhance persistence in cell polarity and directed migration. Cell Syst. 3, 252–263 (2016).
Leung, T. & Malik, J. Representing and recognizing the visual appearance of materials using three-dimensional textons. Int. J. Comput. Vis. 43, 29–44 (2001).
Wen, B. et al. 3D texture analysis for classification of second harmonic generation images of human ovarian cancer. Sci. Rep. 6, 35734 (2016).
Wen, B. L. et al. Texture analysis applied to second harmonic generation image data for ovarian cancer classification. J. Biomed. Opt. 19, 096007 (2014).
Peyrot, D. A. et al. Development of a nonlinear fiber-optic spectrometer for human lung tissue exploration. Biomed. Opt. Express 3, 840–853 (2012).
Liu, Z. et al. 3D organizational mapping of collagen fibers elucidates matrix remodeling in a hormone-sensitive 3D breast tissue model. Biomaterials 179, 96–108 (2018).
Qian, S. et al. Identification of endoplasmic reticulum formation mechanism by multi-parametric, quantitative super-resolution imaging. Opt. Lett. 47, 357–360 (2022).
Campagnola, P. J. & Dong, C. Y. Second harmonic generation microscopy: principles and applications to disease diagnosis. Laser Photonics Rev. 5, 13–26 (2011).
Zipfel, W. R., Williams, R. M. & Webb, W. W. Nonlinear magic: multiphoton microscopy in the biosciences. Nat. Biotechnol. 21, 1369–1377 (2003).
Huang, J. et al. Extracellular matrix and its therapeutic potential for cancer treatment. Signal Transduct. Target. Ther. 6, 153 (2021).
Guo, T., He, C., Venado, A. & Zhou, Y. Extracellular matrix stiffness in lung health and disease. Compr. Physiol. 12, 3523–3558 (2022).
Piccolo, S., Cordenonsi, M. & Dupont, S. Molecular pathways: YAP and TAZ take center stage in organ growth and tumorigenesis. Clin. Cancer Res. 19, 4925–4930 (2013).
van Hinsbergh, V. W. M. & Koolwijk, P. Endothelial sprouting and angiogenesis: matrix metalloproteinases in the lead. Cardiovasc. Res. 78, 203–212 (2008).
Mott, J. D. & Werb, Z. Regulation of matrix biology by matrix metalloproteinases. Curr. Opin. Cell Biol. 16, 558–564 (2004).
Bignon, M. et al. Lysyl oxidase-like protein-2 regulates sprouting angiogenesis and type IV collagen assembly in the endothelial basement membrane. Blood 118, 3979–3989 (2011).
Jiang, Y. F. et al. Targeting extracellular matrix stiffness and mechanotransducers to improve cancer therapy. J. Hematol. Oncol. 15, 34 (2022).
Nadiarnykh, O., LaComb, R. B., Brewer, M. A. & Campagnola, P. J. Alterations of the extracellular matrix in ovarian cancer studied by Second Harmonic Generation imaging microscopy. BMC Cancer 10, 94 (2010).
Provenzano, P. P. et al. Collagen reorganization at the tumor-stromal interface facilitates local invasion. BMC Med. 4, 38 (2006).
Provenzano, P. P. et al. Collagen density promotes mammary tumor initiation and progression. BMC Med. 6, 11 (2008).
Birk, J. W. et al. Second harmonic generation imaging distinguishes both high-grade dysplasia and cancer from normal colonic mucosa. Dig. Dis. Sci. 59, 1529–1534 (2014).
Samata, M. K. et al. Collagen I fiber density increases in lymph node positive breast cancers: pilot study. J. Biomed. Opt. 17, 116017 (2012).
Best, S. L. et al. Collagen organization of renal cell carcinoma differs between low and high grade tumors. BMC Cancer 19, 490 (2019).
Kauppila, S., Bode, M. K., Stenbäck, F., Risteli, L. & Risteli, J. Cross-linked telopeptides of type I and III collagens in malignant ovarian tumours in vivo. Br. J. Cancer 81, 654–661 (1999).
Konen, J. et al. LKB1 kinase-dependent and -independent defects disrupt polarity and adhesion signaling to drive collagen remodeling during invasion. Mol. Biol. Cell 27, 1069–1084 (2016).
Thorlacius-Ussing, J., Kehlet, S. N., Rønnow, S. R., Karsdal, M. A. & Willumsen, N. Non-invasive profiling of protease-specific elastin turnover in lung cancer: biomarker potential. J. Cancer Res. Clin. Oncol. 145, 383–392 (2019).
Toupance, S. et al. Elastin-derived peptides increase invasive capacities of lung cancer cells by post-transcriptional regulation of MMP-2 and uPA. Clin. Exp. Metastasis 29, 511–522 (2012).
Dagogo-Jack, I. & Shaw, A. T. Tumour heterogeneity and resistance to cancer therapies. Nat. Rev. Clin. Oncol. 15, 81–94 (2018).
Castelvecchi, D. Can we open the black box of AI?. Nat. N. 538, 20 (2016).
Lukic, A. et al. Endoscopic fiber probe for nonlinear spectroscopic imaging. Optica 4, 496–501 (2017).
Lombardini, A. et al. High-resolution multimodal flexible coherent Raman endoscope. Light Sci. Appl. 7, 10 (2018).
Harris, K., Puchalski, J. & Sterman, D. Recent advances in bronchoscopic treatment of peripheral lung cancers. Chest 151, 674–685 (2017).
Jahn, S. W., Plass, M. & Moinfar, F. Digital pathology: advantages, limitations and emerging perspectives. J. Clin. Med. 9, 3697 (2020).
Baxi, V., Edwards, R., Montalto, M. & Saha, S. Digital pathology and artificial intelligence in translational medicine and clinical practice. Mod. Pathol. 35, 23–32 (2022).
Travis, W. D. Pathology of lung cancer. Clin. Chest Med. 32, 669–692 (2011).
Hogg, J. C. & Timens, W. The pathology of chronic obstructive pulmonary disease. Annu. Rev. Pathol. 4, 435–459 (2009).
Stolz, D. et al. Systemic biomarkers of collagen and elastin turnover are associated with clinically relevant outcomes in COPD. Chest 151, 47–59 (2017).
Qian, S. et al. Code for waviness, directional variance and orientation. Zenodo https://doi.org/10.5281/zenodo.17973526 (2025).
Acknowledgements
We thank Prof. Chen-Yuan Dong at Taiwan University and Prof. Shean-Jen Chen at Yang Ming Chiao Tung University for insightful discussion. This work was supported by the National Natural Science Foundation of China (62275232 to Z.L., 62375104 to S.Z., and 62575254 to Z.D.), and the Natural Science Foundation of Zhejiang Province (LZ25F050007).
Author information
Authors and Affiliations
Contributions
Z.L., S.Z., S.Q., Z.D., K.S., and J.L. conceived the idea. S.Q., L.Y., J.M., L.Z., T.H., L.C., and G.X. performed the experiments and imaging. S.Q., L.Y., J.M., L.Z., T.H., R.J., C.W., and B.N. analyzed the data with the conceptual advice from all the authors. S.Q., Z.L., and S.Z. wrote the manuscript with input from all the authors. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Yi Jiang, Joan Nichols, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qian, S., Yang, L., Meng, J. et al. Intraoperative biopsy imaging of lung cancer risk. Commun Med 6, 143 (2026). https://doi.org/10.1038/s43856-026-01406-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s43856-026-01406-y