Abstract
Background:
Long COVID is a complex condition where symptoms persist for more than 3 months after SARS-CoV-2 infection and affects an estimated 5-30% of individuals. While persistent inflammation has emerged as an important feature of this condition, it is unclear if immune responses from COVID-19 vaccination or SARS-CoV-2 re-infection exacerbate or mirror the initial inflammatory responses.
Methods:
We quantified 182 inflammatory and neurology-related proteins in plasma using multiplexed affinity proteomics. Plasma samples from the COVID PROFILE cohort conducted in Victoria, Australia, were collected 6-9 months after first infection, but before COVID-19 vaccination from individuals who had recovered from COVID-19 (n = 21) or from individuals with long COVID (n = 12). To establish baseline plasma profiles, protein levels were benchmarked against unvaccinated, SARS-CoV-2 naive individuals (n = 24). In addition, we performed longitudinal analysis in a subset of individuals (n = 34), where paired samples collected 2-4 weeks after a third COVID-19 vaccine dose and after SARS-CoV-2 breakthrough infection were available to assess inflammatory and neurology protein plasma levels after antigen exposure in these contexts.
Results:
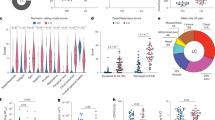
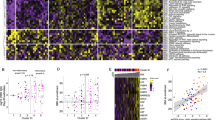
In this cohort Boruta feature selection and lasso regression models identified IL-20, HAGH, NAAA, CLEC10A, LXN, and MCP-1, TRAIL, G-CSF, NBL1, and CCL23 as best discriminating proteins when comparing the long COVID group to groups of either healthy or COVID-19 recovered. Notably, longitudinal analysis indicated differences in the levels of a subset of plasma proteins following primary infection compared to after COVID-19 booster vaccination and breakthrough infection within the groups.
Conclusions:
These findings suggest that there is an altered immune response outcome primarily observed in individuals with long COVID upon re-exposure.
Plain language summary
Long COVID is a condition in which people continue to have symptoms for more than three months after SARS-CoV-2 infection. Ongoing inflammation is thought to contribute to long COVID, but it is unclear whether COVID-19 vaccination or re-infection with SARS-CoV-2 lead to similar, worse or different inflammatory responses compared with the initial infection in people with this condition. We examined blood proteins linked to inflammation and the nervous system to better understand these responses in people with long COVID, individuals that had completely recovered from the first infection and healthy controls. We found that in both long COVID individuals and completely recovered people there were different changes in the level of some immune-related proteins after vaccination or re-infection compared with the response after the original infection suggesting a different immune response from the initial infection upon re-exposure.
Similar content being viewed by others
Data availability
To protect patient privacy, this study uses anonymised data from the COVID PROFILE study. Data have been deposited into Zenodo https://doi.org/10.5281/zenodo.15237087 and are available as Supplementary Data 2 and 3. The source data for Figs. 2, 3 and 4 is in Supplementary Data 2. The source data for Figs. 4 and 5 is in Supplementary Data 3.
Code availability
R code has been deposited into Zenodo https://doi.org/10.5281/zenodo.15237087. This paper does not report original code. This project leverages open-source R code. We have documented the specific R packages and functions used in relevant sections.
References
WHO. Post COVID-19 condition (long COVID). https://www.who.int/europe/news-room/fact-sheets/item/post-covid-19-condition (2022)
Davis, H. E., McCorkell, L., Vogel, J. M. & Topol, E. J. Long COVID: major findings, mechanisms and recommendations. Nat. Rev. Microbiol. 21, 133–146 (2023).
Ceban, F. et al. Fatigue and cognitive impairment in Post-COVID-19 Syndrome: a systematic review and meta-analysis. Brain Behav. Immun. 101, 93–135 (2022).
Al-Aly, Z., Bowe, B. & Xie, Y. Long COVID after breakthrough SARS-CoV-2 infection. Nat. Med. 28, 1461–1467 (2022).
Ayoubkhani, D. et al. Risk of long COVID in people infected with severe acute respiratory syndrome coronavirus 2 after 2 doses of a coronavirus disease 2019 vaccine: community-based, matched cohort study. Open Forum Infect. Dis. 9, ofac464 (2022).
Cervia-Hasler, C. et al. Persistent complement dysregulation with signs of thromboinflammation in active long Covid. Science 383, eadg7942 (2024).
Greene, C. et al. Blood-brain barrier disruption and sustained systemic inflammation in individuals with long COVID-associated cognitive impairment. Nat. Neurosci. 27, 421–432 (2024).
Hanson, A. L. et al. Iron dysregulation and inflammatory stress erythropoiesis associates with long-term outcome of COVID-19. Nat. Immunol. 25, 471–482 (2024).
Klein, J. et al. Distinguishing features of long COVID identified through immune profiling. Nature 623, 139–148 (2023).
Phetsouphanh, C. et al. Immunological dysfunction persists for 8 months following initial mild-to-moderate SARS-CoV-2 infection. Nat. Immunol. 23, 210–216 (2022).
Eaton-Fitch, N. et al. Immune exhaustion in ME/CFS and long COVID. JCI Insight 9, e183810 (2024).
Yin, K. et al. Long COVID manifests with T cell dysregulation, inflammation and an uncoordinated adaptive immune response to SARS-CoV-2. Nat. Immunol. 25, 218–225 (2024).
Group, P.-C. C. Clinical characteristics with inflammation profiling of long COVID and association with 1-year recovery following hospitalisation in the UK: a prospective observational study. Lancet Respir. Med. 10, 761–775 (2022).
Peluso, M. J. et al. Plasma markers of neurologic injury and inflammation in people with self-reported neurologic postacute sequelae of SARS-CoV-2 infection. Neurol. Neuroimmunol. Neuroinflamm. 9, e200003 (2022).
Gu, X. et al. Probing long COVID through a proteomic lens: a comprehensive two-year longitudinal cohort study of hospitalised survivors. EBioMedicine 98, 104851 (2023).
Etter, M. M. et al. Severe neuro-COVID is associated with peripheral immune signatures, autoimmunity and neurodegeneration: a prospective cross-sectional study. Nat. Commun. 13, 6777 (2022).
Tandon, P. et al. Unraveling links between chronic inflammation and long COVID: workshop report. J. Immunol. 212, 505–512 (2024).
Phetsouphanh, C. et al. Improvement of immune dysregulation in individuals with long COVID at 24-months following SARS-CoV-2 infection. Nat. Commun. 15, 3315 (2024).
Eriksson, E. M. et al. Cohort profile: a longitudinal Victorian COVID-19 cohort (COVID PROFILE). Preprint at medRxiv https://doi.org/10.1101/2023.04.27.23289157 (2023).
Ernster, V. L. Nested case-control studies. Prev. Med. 23, 587–590 (1994).
Mazhari, R. et al. SARS-CoV-2 multi-antigen serology assay. Methods Protoc. 4, 72 (2021).
Assarsson, E. et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 9, e95192 (2014).
Nevola, K. et al. OlinkAnalyze: facilitate analysis of proteomic data from Olink. R package version 3.6.2 (2024).
Olink. Introduction to bridging Olink® NPX datasets. https://cran.r-project.org/web/packages/OlinkAnalyze/vignettes/bridging_introduction.html (2025).
Olink. Calculating LOD from Olink® Explore data, <https://cran.r-project.org/web//packages/OlinkAnalyze/vignettes/LOD.html#introduction> (2025).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Tay, J. K., Narasimhan, B. & Hastie, T. Elastic net regularization paths for all generalized linear models. J. Stat. Softw. 106, 1 (2023).
Kursa, M. B. & Rudnicki, W. R. Feature selection with the Boruta package. J. Stat. Softw. 36, 1–13 (2010).
Meyer, D., Dimitriadou, E., Hornik, K., Weingessel, A. & F, L. _e1071: misc functions of the Department of Statistics, Probability Theory Group (Formerly: E1071), TU Wien_. R. Package Vers. 1, 7–14 (2023).
Gianfagna, F. et al. Anti-SARS-CoV-2 antibody levels and kinetics of vaccine response: potential role for unresolved inflammation following recovery from SARS-CoV-2 infection. Sci. Rep. 12, 385 (2022).
Koutsakos, M. et al. The magnitude and timing of recalled immunity after breakthrough infection is shaped by SARS-CoV-2 variants. Immunity 55, 1316–1326 e1314 (2022).
Zhang, Z. et al. Humoral and cellular immune memory to four COVID-19 vaccines. Cell 185, 2434–2451 e2417 (2022).
Grady, C. B. et al. Impact of COVID-19 vaccination on symptoms and immune phenotypes in vaccine-naive individuals with Long COVID. Commun. Med. 5, 163 (2025).
Hamzaraj, K. et al. Impact of circulating anti-spike protein antibody levels on multi-organ long COVID symptoms. Vaccines 12, 610 (2024).
Joung, S. et al. Serological response to vaccination in post-acute sequelae of COVID. BMC Infect. Dis. 23, 97 (2023).
Madenbayeva, A. M. et al. Impact of QazVac vaccination on clinical manifestations and immune responses in post-COVID syndrome: a cross-sectional study. Front. Med. 12, 1556623 (2025).
Tsuchida, T. et al. Relationship between changes in symptoms and antibody titers after a single vaccination in patients with Long COVID. J. Med. Virol. 94, 3416–3420 (2022).
Iosef, C. et al. Plasma proteome of Long-COVID patients indicates HIF-mediated vasculo-proliferative disease with impact on brain and heart function. J. Transl. Med. 21, 377 (2023).
Liew, F. et al. Large-scale phenotyping of patients with long COVID post-hospitalization reveals mechanistic subtypes of disease. Nat. Immunol. 25, 607–621 (2024).
Patel, M. A. et al. Organ and cell-specific biomarkers of Long-COVID identified with targeted proteomics and machine learning. Mol. Med. 29, 26 (2023).
Zhao, J. et al. Plasma biomarkers for systemic inflammation in COVID-19 survivors. Proteom. Clin. Appl. 16, e2200031 (2022).
Rajamanickam, A. et al. Characterization of IL-10 family of cytokines in acute and convalescent COVID-19 individuals. J. Interferon Cytokine Res. 43, 469–477 (2023).
Chen, J., Caspi, R. R. & Chong, W. P. IL-20 receptor cytokines in autoimmune diseases. J. Leukoc. Biol. 104, 953–959 (2018).
Kragstrup, T. W. et al. The interleukin-20 receptor axis in early rheumatoid arthritis: novel links between disease-associated autoantibodies and radiographic progression. Arthritis Res. Ther. 18, 61 (2016).
Chen, Y. et al. New-onset autoimmune phenomena post-COVID-19 vaccination. Immunology 165, 386–401 (2022).
Wang, E. Y. et al. Diverse functional autoantibodies in patients with COVID-19. Nature 595, 283–288 (2021).
Hsu, Y. H. et al. Anti-IL-20 monoclonal antibody inhibited inflammation and protected against cartilage destruction in murine models of osteoarthritis. PLoS ONE 12, e0175802 (2017).
Kragstrup, T. W. et al. Increased interleukin (IL)-20 and IL-24 target osteoblasts and synovial monocytes in spondyloarthritis. Clin. Exp. Immunol. 189, 342–351 (2017).
Llaurador-Coll, M. et al. Plasma levels of neurology-related proteins are associated with cognitive performance in an older population with overweight/obesity and metabolic syndrome. Geroscience 45, 2457–2470 (2023).
Kindrachuk, J. et al. Antiviral potential of ERK/MAPK and PI3K/AKT/mTOR signaling modulation for Middle East respiratory syndrome coronavirus infection as identified by temporal kinome analysis. Antimicrob. Agents Chemother. 59, 1088–1099 (2015).
Kong, F. et al. Sirtuins as potential therapeutic targets for hepatitis B virus infection. Front. Med. 8, 751516 (2021).
Roche, K. L. et al. An allosteric inhibitor of sirtuin 2 deacetylase activity exhibits broad-spectrum antiviral activity. J. Clin. Invest. 133, e158978 (2023).
Acosta-Ampudia, Y. et al. Persistent autoimmune activation and proinflammatory state in post-coronavirus disease 2019 syndrome. J. Infect. Dis. 225, 2155–2162 (2022).
Blomberg, B. et al. Long COVID in a prospective cohort of home-isolated patients. Nat. Med. 27, 1607–1613 (2021).
Ghafari, M. et al. Prevalence of persistent SARS-CoV-2 in a large community surveillance study. Nature 626, 1094–1101 (2024).
Zollner, A. et al. Postacute COVID-19 is characterized by gut viral antigen persistence in inflammatory bowel diseases. Gastroenterology 163, 495–506 e498 (2022).
Augustin, M. et al. Immunological fingerprint in coronavirus disease-19 convalescents with and without post-COVID syndrome. Front. Med. 10, 1129288 (2023).
Kervevan, J. et al. Divergent adaptive immune responses define two types of long COVID. Front. Immunol. 14, 1221961 (2023).
COVID-19 Australia: Epidemiology Report 57 https://www1.health.gov.au/internet/main/publishing.nsf/Content/C50CAE02452A48A7CA2587320081F7BF/<span>$File/covid_19_australia_epidemiology_report_57_reporting_period_ending_16_january_2022.pdf (2022).
COVID-19 Australia: Epidemiology Report 60. https://www1.health.gov.au/internet/main/publishing.nsf/Content/C50CAE02452A48A7CA2587320081F7BF/<span>$File/covid_19_australia_epidemiology_report_60_reporting_period_ending_10_april_2022.pdf (2022).
Wik, L. et al. Proximity extension assay in combination with next-generation sequencing for high-throughput proteome-wide analysis. Mol. Cell Proteom. 20, 100168 (2021).
Therneau T, A. B. rpart: recursive partitioning and regression trees. R package version 4.1.23. https://CRAN.R-project.org/package=rpart (2023).
Williams, G. J. Data Mining with Rattle and R: The Art of Excavating Data for Knowledge Discovery. Series Use R! (Springer, 2011).
Lenth, R. emmeans: estimated marginal means, aka least-squares means. R package version 1.10.0. (2024).
Acknowledgements
We wish to thank Anne Hart, Maureen Ford and all members of the COVID PROFILE consortium and the study participants in the COVID PROFILE study. We also wish to thank Dr. Lauren Smith for her valuable discussion and input around data analysis. Dr. Bansal received funding from the University of Bergen, The National Graduate School in Infection Biology and Antimicrobials (or IBA) and Pasteur legatet & Thjøtta’s legat, University of Oslo, Norway [101563]. The COVID PROFILE study was supported by WHO Unity funds (2020/1085469-0), and WEHI philanthropic funds. I.M. is supported by an NHMRC Senior Research Fellowship (#1043345). This work was made possible through Victorian State Government Operational Infrastructure Support and Australian Government NHMRC IRIISS.
Author information
Authors and Affiliations
Consortia
Contributions
S.W.Z.O and E.M.E. conceptualised the study. A.B., S.W.Z.O., J.C., I.A. and E.M.E. performed the data analysis. S.W.Z.O., N.W.K. and R.M. generated the data. A.B., S.W.Z.O., N.W.K., R.B. and E.M.E. designed experimental work; A.B., S.W.Z.O. and E.M.E. wrote the first draft of the manuscript. R.B., I.M., I.A. and R.J.C. provided guidance and discussion and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Medicine thanks Ruth Drury, Gerald Mboowa, Lalit Batra, Guo-Lin Wang, Andreza Lemos Salvio and the other, anonymous, reviewer for their contribution to the peer review of this work. Peer review reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bansal, A., Olechnowicz, S.W.Z., Kiernan-Walker, N. et al. Divergent inflammatory and neurology-related protein levels in long COVID following primary and breakthrough SARS-CoV-2 infections. Commun Med (2026). https://doi.org/10.1038/s43856-026-01541-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s43856-026-01541-6