Abstract
Individuals with hypoplastic left heart syndrome (HLHS) have an underdeveloped left ventricle and require surgery to reconfigure blood flow for survival. Here we profiled the HLHS right-ventricular microenvironment by single-nucleus RNA sequencing and spatial transcriptomics at birth (before heart failure), after surgery with heart failure and after ventricular assist device (VAD) unloading (reduced hypoxia and volume overload). We show that HLHS cardiomyocytes, both within the heart and when derived from induced pluripotent stem cells, are intrinsically senescent. The HLHS myocardium contained a senescent microvascular niche with endothelial cells, pericytes and YAP-high fibroblasts, consistent with hypoxic and mechanical stress. This senescent niche is similar to adult myocardial infarction but not pediatric dilated cardiomyopathy with heart failure, pointing to a prominent role of hypoxia in senescence. The microvascular senescent niche was improved by VAD, providing insight into the potential to reverse cardiac cell states that lead to heart failure.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
All digitized expression matrices of sequencing data generated in this study were deposited to Zenodo (https://doi.org/10.5281/zenodo.18407403)58. The Texas Genomic Act of 2025 limits human data sharing. Data are available from the corresponding author upon request, subject to completion of appropriate data use agreements and institutional approvals. All other data supporting the findings of this study are available within the article and its Supplementary Information.
Code availability
All code used to generate the figures in this study is available from GitHub (https://github.com/XL-Genomics/2024_Senescent_Niche_in_HLHS).
References
Benson, D. W., Martin, L. J. & Lo, C. W. Genetics of hypoplastic left heart syndrome. J. Pediatr. 173, 25–31 (2016).
Krane, M. et al. Sequential defects in cardiac lineage commitment and maturation cause hypoplastic left hearts syndrome. Circulation 144, 1409–1428 (2021).
Theis, J. L. et al. Genetic association between hypoplastic left heart syndrome and cardiomyopathies. Circ. Genom. Precis. Med. 14, e003126 (2021).
Xu, X. et al. Uncompensated mitochondrial oxidative stress underlies heart failure in an iPSC-derived model of congenital heart disease. Cell Stem Cell 29, 840–855 (2022).
Paige, S. L. et al. Patient-specific induced pluripotent stem cells implicate intrinsic impaired contractility in hypoplastic left heart syndrome. Circulation 142, 1605–1608 (2020).
Wang, Y. J. et al. Systems analysis of de novo mutations in congenital heart diseases identified a protein network in the hypoplastic left heart syndrome. Cell Syst. 13, 895–910 (2022).
Yu, Z. et al. KMT2D–NOTCH mediates coronary abnormalities in hypoplastic left heart syndrome. Circ. Res. 131, 280–282 (2022).
Miao, Y. et al. Intrinsic endocardial defects contribute to hypoplastic left heart syndrome. Cell Stem Cell 27, 574–589 (2020).
Salih, C., Sheppard, M. N. & Ho, S. Y. Morphometry of coronary capillaries in hypoplastic left heart syndrome. Ann. Thorac. Surg. 77, 903–907 (2004).
Chen, J. et al. Molecular and spatial signatures of mouse embryonic endothelial cells at single-cell resolution. Circ. Res. 134, 529–546 (2024).
Farah, E. N. et al. Spatially organized cellular communities form the developing human heart. Nature 627, 854–864 (2024).
Kanemaru, K. et al. Spatially resolved multiomics of human cardiac niches. Nature 619, 801–810 (2023).
Kuppe, C. et al. Spatial multi-omic map of human myocardial infarction. Nature 608, 766–777 (2022).
Li, R. G. et al. YAP induces a neonatal-like pro-renewal niche in the adult heart. Nat. Cardiovasc. Res. 3, 283–300 (2024).
Hill, M. C. et al. Integrated multiomic characterization of congenital heart disease. Nature 606, 181–191 (2022).
Sim, C. B. et al. Sex-specific control of human heart maturation by the progesterone receptor. Circulation 143, 1614–1628 (2021).
Cho, J. et al. Long-term implantable ventricular assist device support in children. J. Thorac. Cardiovasc. Surg. 167, 1417–1426 (2024).
Griveau, A., Wiel, C., Ziegler, D. V., Bergo, M. O. & Bernard, D. The JAK1/2 inhibitor ruxolitinib delays premature aging phenotypes. Aging Cell 19, e13122 (2020).
Martins, R., Lithgow, G. J. & Link, W. Long live FOXO: unraveling the role of FOXO proteins in aging and longevity. Aging Cell 15, 196–207 (2016).
Tripathi, D. & Reddy, S. iPSC model of congenital heart disease predicts disease outcome. Cell Stem Cell 29, 659–660 (2022).
Wong, B. W., Marsch, E., Treps, L., Baes, M. & Carmeliet, P. Endothelial cell metabolism in health and disease: impact of hypoxia. EMBO J. 36, 2187–2203 (2017).
Frangogiannis, N. G. The fate and role of the pericytes in myocardial diseases. Eur. J. Clin. Invest. 54, e14204 (2024).
Buechler, M. B. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021).
Reichart, D. et al. Pathogenic variants damage cell composition and single cell transcription in cardiomyopathies. Science 377, eabo1984 (2022).
Dupont, S. & Wickström, S. A. Mechanical regulation of chromatin and transcription. Nat. Rev. Genet. 23, 624–643 (2022).
Xiao, Y. et al. Hippo pathway deletion in adult resting cardiac fibroblasts initiates a cell state transition with spontaneous and self-sustaining fibrosis. Genes Dev. 33, 1491–1505 (2019).
Nardone, G. et al. YAP regulates cell mechanics by controlling focal adhesion assembly. Nat. Commun. 8, 15321 (2017).
Chen, C., Li, R., Ross, R. S. & Manso, A. M. Integrins and integrin-related proteins in cardiac fibrosis. J. Mol. Cell Cardiol. 93, 162–174 (2016).
Eraslan, G. et al. Single-nucleus cross-tissue molecular reference maps toward understanding disease gene function. Science 376, eabl4290 (2022).
Wang, T. W. & Nakanishi, M. Immune surveillance of senescence: potential application to age-related diseases. Trends Cell Biol. 35, 248–257 (2025).
Romacho, T. et al. Visfatin/eNampt induces endothelial dysfunction in vivo: a role for Toll-like receptor 4 and NLRP3 inflammasome. Sci. Rep. 10, 5386 (2020).
Gasparrini, M. et al. Molecular insights into the interaction between human nicotinamide phosphoribosyltransferase and Toll-like receptor 4. J. Biol. Chem. 298, 101669 (2022).
Gaber, N. et al. Fetal reprogramming and senescence in hypoplastic left heart syndrome and in human pluripotent stem cells during cardiac differentiation. Am. J. Pathol. 183, 720–734 (2013).
Liu, X. et al. The complex genetics of hypoplastic left heart syndrome. Nat. Genet. 49, 1152–1159 (2017).
Kung-Chun Chiu, D. et al. Hypoxia regulates the mitochondrial activity of hepatocellular carcinoma cells through HIF/HEY1/PINK1 pathway. Cell Death Dis. 10, 934 (2019).
Taylor, C. T. & Scholz, C. C. The effect of HIF on metabolism and immunity. Nat. Rev. Nephrol. 18, 573–587 (2022).
Gladyshev, V. N. et al. Molecular damage in aging. Nat. Aging 1, 1096–1106 (2021).
Acosta, J. C. et al. Chemokine signaling via the CXCR2 receptor reinforces senescence. Cell 133, 1006–1018 (2008).
Tsuda, T. & Patel, G. Coronary microvascular dysfunction in childhood: an emerging pathological entity and its clinical implications. Am. Heart J. Plus 42, 100392 (2024).
Rickers, C. et al. Myocardial perfusion in hypoplastic left heart syndrome. Circ. Cardiovasc. Imaging 14, e012468 (2021).
Coppe, J. P., Desprez, P. Y., Krtolica, A. & Campisi, J. The senescence-associated secretory phenotype: the dark side of tumor suppression. Annu. Rev. Pathol. 5, 99–118 (2010).
Moiseeva, V. et al. Senescence atlas reveals an aged-like inflamed niche that blunts muscle regeneration. Nature 613, 169–178 (2023).
Chaib, S., Tchkonia, T. & Kirkland, J. L. Cellular senescence and senolytics: the path to the clinic. Nat. Med. 28, 1556–1568 (2022).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–183 (2011).
Yun, M. H., Davaapil, H. & Brockes, J. P. Recurrent turnover of senescent cells during regeneration of a complex structure. eLife 4, e05505 (2015).
Thorp, E. B. Cardiac macrophages and emerging roles for their metabolism after myocardial infarction. J. Clin. Invest. 133, e171953 (2023).
Zaman, R. & Epelman, S. Resident cardiac macrophages: heterogeneity and function in health and disease. Immunity 55, 1549–1563 (2022).
Vagnozzi, R. J. & Molkentin, J. D. Resident macrophages keep mitochondria running in the heart. Cell Res. 30, 1057–1058 (2020).
Balaban, R. S., Nemoto, S. & Finkel, T. Mitochondria, oxidants, and aging. Cell 120, 483–495 (2005).
Nicolás-Ávila, J. A. et al. A network of macrophages supports mitochondrial homeostasis in the heart. Cell 183, 94–109 (2020).
Hamidzada, H. et al. Primitive macrophages induce sarcomeric maturation and functional enhancement of developing human cardiac microtissues via efferocytic pathways. Nat. Cardiovasc. Res. 3, 567–593 (2024).
Litvinukova, M. et al. Cells of the adult human heart. Nature 588, 466–472 (2020).
Saul, D. et al. A new gene set identifies senescent cells and predicts senescence-associated pathways across tissues. Nat. Commun. 13, 4827 (2022).
Rodriguez Morales, D. et al. Vascular niches are the primary hotspots in cardiac aging. Circ. Res. 137, 1353–1367 (2025).
Liu, X. et al. Distinct human Langerhans cell subsets orchestrate reciprocal functions and require different developmental regulation. Immunity 54, 2305–2320 (2021).
Cousins, R. D. Annotated bibliography of some papers on combining significances or P-values. Preprint at https://doi.org/10.48550/arXiv.0705.2209 (2008).
Amrute, J. M. et al. Defining cardiac functional recovery in end-stage heart failure at single-cell resolution. Nat. Cardiovasc. Res. 2, 399–416 (2023).
Li, X. et al. Ventricular assist device unloading reverses microvascular senescence in single ventricle disease. Zenodo https://doi.org/10.5281/zenodo.18407404 (2026).
Acknowledgements
We thank Scientific Publications at The Texas Heart Institute for manuscript editing, L. Wadhwa and the Texas Children’s Hospital Heart Center Biorepository for technical assistance and E. Klysik and P. Swinton at Baylor College of Medicine for technical assistance. This work was supported by the Don McGill Gene Editing Laboratory of The Texas Heart Institute (X.L. and Y.Z.), the National Institutes of Health (HL118761, HL169511, HL171574 and HL177644 to J.F.M., HL142704 and HL179012 to X.L., T32HL139430 to C.-R.T. and K99HL174827 to R.G.L.), the Additional Ventures Foundation (SVRF 1330043515 to J.F.M. and D.T. and SVRF-EA 988596 to J.F.M. and D.T.), the BCM Cardiovascular Research Institute Pilot Award (J.F.M. and D.T.), the Vivian L. Smith Foundation (J.F.M.), the Levy Logenbaugh Foundation (X.L.), the Graeme McDaniel Foundation (D.T.) and the Mike Hogg Foundation (D.T.).
Author information
Authors and Affiliations
Contributions
Conceptualization, J.F.M., X.L., D.T. and I.A. Methodology, X.L., D.T., Y.Z., C.-R.T., Y.M., R.G.L. and H.J.T. Investigation, D.T., X.L., Y.Z., C.-R.T., Y.M., R.G.L. and H.J.T. Visualization, X.L., Y.Z. and M.A.H.S. Funding acquisition, J.F.M., X.L. and D.T. Project administration, J.F.M., X.L. and D.T. Supervision, J.F.M. Writing—original draft, J.F.M., X.L. and D.T. Writing—review and editing, J.F.M., X.L., D.T., I.A. and M.A.H.S. Writing—revision: J.F.M., X.L. and D.T.
Corresponding author
Ethics declarations
Competing interests
J.F.M. is a cofounder and owns shares in Medley Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Cardiovascular Research thanks Benoit Bruneau, Eldad Tzahor and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cell type annotation, and concordance of snRNA-seq and spatial transcriptomics data.
A, Circulating brain natriuretic peptide (BNP) levels in three HLHS patients, comparing pre- and post-VAD time points. Each line represents an individual patient. B, Heatmap showing the fraction of cell clusters (rows) in the integrated snRNA-seq data predicted as specific cell types (columns) from a reference dataset. HCA: Human Cell Atlas. C, Heatmap of cell type signature gene expression (rows) in all cells, grouped by cell type (columns). A representative signature gene for each cell type is labeled on the left. Cardiomyocytes (CM), endothelial cells (EC), fibroblasts (FB), smooth muscle cells (SMC), pericytes (PC), lymphoid cells (Lym), myeloid cells (Mye), adipocytes (AC), and neuronal cells (NC). D, Principal component analysis of pseudobulk snRNA-seq data showing variance in principal components 1 and 2. Each data point represents an individual donor or HLHS patient, with shapes indicating sex and colors depicting age. E, Scatter plot comparing cell type proportions quantified from matched snRNA-seq (x-axis) and Xenium (y-axis) datasets from the same hearts (C1, H12, H15, H16, V4, and V5).
Extended Data Fig. 2 Cardiomyocyte aging signatures, cell states, and immunofluorescence-based characterization.
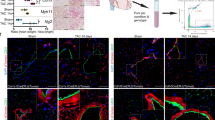
A, Differential expressed genes (DEG) in CMs in HLHS and controls. Each column represents a donor or HLHS patient. Representative genes are listed on the left. B, Cardiomyocyte aging metric signature genes derived from donor CMs from Sim et al, comparing young and adult samples. N denotes the number of donors. wo, weeks old; yo, years old. C, Z-scores of young heart signatures (left) and aged heart signatures (right) in CMs from controls and HLHS hearts. Signature genes are from the GTEx database. Statistical significance was assessed using two-tailed Wilcoxon rank-sum test. *: p < 0.05, **: p < 0.01. D, Expression of CM state signature genes in the integrated data. Representative genes are shown on the left. E, Dot plot showing per-patient quantification of relative CM cell state abundance based on snRNA-seq data across 6 donor controls, 17 stage I-III and 4 post-VAD HLHS patients. The midline indicates the mean abundance within each group, and dashed lines connect paired pre- and post-VAD samples from the same patients. F, Inferred transition from CM1 and CM2 to CM3 in HLHS based on RNA velocity. Arrows overlaid on UMAP embedding represent predicted transitional directions. G, Representative immunofluorescent image of 53BP1 staining (green) in right ventricular sections from a donor control and two HLHS patients, co-stained with DAPI and WGA. Arrowheads indicate representative 53BP1+ CMs. The boxed area in a HLHS heart is enlarged on the bottom panel. H, Representative immunofluorescent image of p16 staining (green) in a Stage III HLHS right ventricular section, co-stained with DAPI and WGA. Arrowheads indicate representative p16⁺ CM3. I, Quantification of cell size of 33 p16⁻ and 27 p16⁺ CMs in the HLHS heart shown in H. J, Representative immunofluorescent images of p16 staining in the paired HLHS heart at post-VAD timepoints, similar to H. K, Quantification of CM cell size in the paired pre- (48 cells) and post-VAD (101 cells) timepoints. L, CM size quantification stratified by p16 signal (26 positive and 22 negative cells in Pre-VAD sample and 34 positive and 67 negative cells in Post-VAD sample). Statistical significance was determined using a Tukey’s multiple comparison test for I, K and L. n.s.: not significant, *p < 0.05, **p < 0.005, ****p < 1×10−4 for all plots.
Extended Data Fig. 3 Identification, spatial validation, and patient-level heterogeneity of VAD-responsive senescent cell states.
A, Differential expression of cell state signature genes (rows) in single cells grouped by cell states: fibroblasts (FBs), endothelial cells (ECs), mural cells, myeloid cells, and lymphoid cells. Representative marker genes are shown on the left. B, Distribution of Milo neighbors for each cell state, corresponding to Figs. 3c and 3d. Nodes represent neighborhoods, colored by log fold-change for stage I-III HLHS versus controls (left) or pre-VAD HLHS versus post-VAD HLHS (right). Neighborhoods with insignificant log fold-changes (FDR > 0.1) are shown in grey. CM3, FB5, EC4, and MP3 cell states are highlighted in boxes. C, Additional representative HLHS Xenium ST sections, showcasing examples of CM3, FB5, EC4, PC2, and MP3 cells, along with their respective marker gene transcripts, similar to Fig. 3g. D, Relative composition of VAD-responsive cells states (CM3, FB5, EC4, PC2, MP3) detected by snRNA-seq (left) and Xenium ST (right) across controls, stage I-III HLHS and post-VAD HLHS samples. E-F, Radar plots showing per-patient relative abundance of VAD-responsive cell states detected by snRNA-seq (E) and Xenium ST (F). Colored boxes indicate paired pre- and post-VAD HLHS samples from the same patient.
Extended Data Fig. 4 Spatial organization and colocalization of senescent cell states defining the HLHS microvascular niche.
A, Correlation of nine key senescence hallmark genes with overall senescence signature expression in cell states. Scatter plot showing mean senescence signature gene expression (x-axis) and each hallmark gene (y-axis). Senescent niche cell states are highlighted in red. B, Heatmap showing expression of marker genes of key senescent cell states (FB5, EC4, PC2, and MP3) across Visium data. C, Pairwise colocalization testing of CM2 and CM3 against all cell states in the integrated data. Senescent niche cell states are highlighted in red. D, Heatmap showing the distribution of individual cell states (rows) across spatial clusters (columns) defined by spatial proximity and neighborhood composition in control, stage III and post-VAD HLHS Xenium samples. Note the absence of a distinct niche enriched for senescent cell states in the control and post-VAD samples (red).
Extended Data Fig. 5 Hypoxic and senescent endothelial and pericyte cell states in HLHS myocardium.
A, Heatmap showing the fraction of EC clusters (rows) in the integrated snRNA-seq data predicted as specific EC subtypes (columns) from reference atlas dataset. B, EC4 (p16+ UEA-1+) percentage relative to all ECs based on immunofluorescent staining in HLHS RV sections. C, UMAP zoomed in on EC4 cells, corresponding to Fig. 5f. Cells are colored by RNA velocity latent time. D, Expression trend of co-expressed genes correlated with the EC4 transition trajectory. Each line represents a gene. E, Correlation of hypoxia response gene expression in ECs and mural cells in HLHS patients. Each dot represents an individual patient, positioned by mean expression z-score in ECs (x-axis) and mural cells (y-axis). The black line represents the generalized additive model (GAM) fit, and the shaded band denotes the 95% confidence interval around the fitted curve. F, Re-embedded UMAP of mural cells, showing two pericyte (PC) and smooth muscle cell (SMC) states. G, Box plot showing PC2 abundance relative to all mural cells across 6 donor controls, 17 stage I-III HLHS, and 4 post-VAD HLHS tissue. Each dot represents an individual donor or patient. Statistical significance was determined using a two-tailed Wilcoxon rank-sum test. *: p < 0.05. In all box plots, the centre line indicates the mean, the box bounds represent the interquartile range, and the whiskers extend to the minimum and maximum values within 1.5× the interquartile range. H, Differential abundance of PC2 in three HLHS patients, comparing post-VAD and pre-VAD time points. I, Quantification of Visium ST section area showing significant IL-6 ligand and receptor colocalization.
Extended Data Fig. 6 Fibroblast senescence and predicted cell state transition.
A, Differential expression of fibroblast genes in HLHS and controls. Each column represents a donor or HLHS patient. Representative genes are listed on the left. B, UMAP showing expression of top principal component 2 genes identified in Fig. 6c. Genes were ranked by loading magnitude, and those contributing most strongly to principal component 2 variance were designated as top-loading genes. The mean expression score of the top 100 genes visualizes their FB5-enriched pattern, with the top 12 representative genes listed on the right. The FB5 state is outlined by a dashed line. C, Box plot showing distribution of senescence signature expression across fibroblasts from pediatric donor controls (19466 cells), stage I-III HLHS (25952 cells), adult donor control (17803 cells), adult DCM heart failure (10347 cells), and adult post-MI heart tissue from myogenic (25465 cells) and ischemic (6626 cells) zones. ****: p < 1×10−4. Statistical significance was assessed using two-tailed t-tests, with the maximal Benjamini-Hochberg-adjusted p-value across leave-one-out iterations reported. In all box plots, the centre line indicates the mean, the box bounds represent the interquartile range, and the whiskers extend to the minimum and maximum values within 1.5× the interquartile range. D, Partition-based graph abstraction (PAGA) showing predicted fibroblast (FB) state transition based on RNA velocity. Nodes represent cell states, and arrows indicate transition direction.
Extended Data Fig. 7 Myeloid cell annotation and senescence signature expression.
A, Expression density overlay on the UMAP depicting patterns of key marker genes such as CCR2, TIMD4, LYVE1, and FOLR2, as well as signatures previously defined for cardiac myeloid cells by Eraslan et al. Density was calculated using the Nebulosa R package. B, Box plot showing distribution of senescence signature expression across myeloid cells from pediatric donor controls (5946 cells), stage I-III HLHS (7431 cells), adult donor controls (17440 cells), adult DCM heart failure (9829 cells), and adult post-MI heart tissue from myogenic (7047 cells) and ischemic (3448 cells) zones. ****: p < 1×10−4. Statistical significance was assessed using two-tailed t-tests, with the maximal Benjamini-Hochberg-adjusted p-value across leave-one-out iterations reported. In all box plots, the centre line indicates the mean, the box bounds represent the interquartile range, and the whiskers extend to the minimum and maximum values within 1.5× the interquartile range.
Extended Data Fig. 8 HLHS senescent niche signaling networks.
A, The complete heatmap showing expression of all ligand genes expressed by senescent cell states (FB5, EC4, PC2 and MP3) in stage I-III HLHS samples. B, Heatmap showing the expression of the POSTN ligand and its receptors, ITGAV and ITGB5 (left), and the NAMPT ligand and its receptor, TLR4 (right), in major cell types across controls, stage I-III HLHS, and post-VAD HLHS samples.
Extended Data Fig. 9 snRNA-seq pipeline, quality control, and robustness analyses.
A, Schematic illustration of the key steps of the computational pipeline used for snRNA-seq data processing. B, Violin plots of key quality control metrics for each snRNA-seq library. Black bars indicate library-specific thresholds. C, UMAP highlighting potential doublets based on Scrublet prediction. D, Joint annotation of doublets or nuclei with high ambient RNA contamination based on Scrublet prediction and manual inspection. These final ambiguous cells are labeled and excluded from all downstream analyses. E, Bar plots showing the maximal contribution by any individual donor or patient in the annotated cell types (left) and cell states (right). Cell populations overrepresented by a single individual above the 50% threshold (dashed line) are marked and excluded from analyses. F, UMAP embeddings from leave-one-out analyses of snRNA-seq data showing that the five HLHS-enriched senescent cell states (CM3, FB5, EC4, PC2, MP3) remain consistently identifiable and form unbiased Louvain clusters across all iterations. G, Leave-one-out Milo-based differential abundance analysis showing that the post-VAD reduction of senescent cell states remains statistically significant across all iterations. Dots represent neighborhoods, colored by log fold-change for pre-VAD HLHS versus post-VAD HLHS. Neighborhoods with insignificant log fold-changes (FDR > 0.1) are shown in grey.
Supplementary information
Supplementary Tables (download XLSX )
Excel sheets of Supplementary Tables 1–8.
Source data
Source Data Fig. 2 (download XLSX )
Quantification for p16, p21 and 53BP1 immunofluorescence.
Source Data Extended Data Fig. 2 (download XLSX )
Quantification for p16 immunofluorescence and CM size.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Turaga, D., Zhao, Y. et al. Ventricular assist device unloading reverses microvascular senescence in single ventricle disease. Nat Cardiovasc Res 5, 262–280 (2026). https://doi.org/10.1038/s44161-026-00790-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s44161-026-00790-x