Abstract
Microsporidia, a large group of fungal-related intracellular parasites, infect several economically significant animals, leading to substantial economic losses. As currently available anti-microsporidia therapies are either ineffective or come with numerous adverse effects, there is a need for alternative microsporidia inhibitors. Here we screen a subset of the ChemBridge DIVERset library, comprising 2500 diverse compounds, using Caenorhabditis elegans infected with its natural microsporidian parasite, Nematocida parisii. By testing these compounds at 60 μM in 96-well assay plates, we identified 26 hits that restored the ability of C. elegans to produce progeny in the presence of N. parisii. We confirmed that out of 20 tested compounds, 18 ChemBridge compounds effectively inhibit N. parisii infection in C. elegans. Of these 18, 10 were benzenesulfonamide derivatives which inhibit microsporidia infection by inactivating spores. We screened an additional 475-compound benzenesulfonamide library, successfully identifying three compounds that are effective at a lower concentration than the initial hits. We further show that one benzenesulfonamide compound displays inhibitory activity against several species of microsporidia, inhibiting infection of species belonging to the Nematocida, Enterocytozoon, and Encephalitozoon genera. Together our results suggest that benzenesulfonamides are a potential scaffold for the development of microsporidia antiseptics.
Similar content being viewed by others
Introduction
Microsporidia are obligate intracellular pathogens that infect most kinds of animal groups1,2,3. Microsporidia were previously classified as protozoa, but they are now regarded as fungi4. Over 1700 species of microsporidia have been identified and they are widespread in the natural environment5,6. Microsporidian infections in host organisms are usually deleterious and can cause mortality7,8. A variety of economically significant animals, such as farmed penaeid shrimp, honey bees, and silkworms are at risk from these parasites9,10. Microsporidiosis outbreaks, especially in hatcheries, may collapse industries11. Additionally, 17 species of microsporidia have been reported to infect humans12. Patients with compromised immune systems, such as HIV-positive individuals, organ transplant recipients, or cancer patients, are more likely to be infected by this emerging pathogen12.
Anti-microsporidia therapies currently available are either ineffective or associated with several adverse effects13. Albendazole, a benzimidazole derivative, is one of the most common treatments for microsporidiosis. Other benzimidazole derivatives, such as MMV1782387, carbendazim, fenbendazole, and oxfendazole also show anti-microsporidia activity14,15. Benzimidazole analogs are effective in controlling microsporidiosis during the proliferative phase, however, they were not observed to enhance parasite clearance15. Benzimidazole analogs are effective against the intracellular stages of microsporidia, but ineffective on spores16. Albendazole is also less effective at inhibiting the human-infecting microsporidia Encephalitozoon bieneusi and Vittaforma corneae17,18. Albendazole is thought to bind to microsporidian beta tubulin, inhibit tubulin polymerization, and these microsporidian species contain variants that have been shown to cause albendazole resistance in other organisms14. The other most commonly used inhibitor of microsporidia is fumagillin, which binds specifically and covalently to methionine aminopeptidase type 2 (MetAP2)19,20,21. Fumagillin is not approved for human and agricultural use due to concerns regarding host toxicity14,22. Other inhibitors such as orlistat and dexrazoxane have been explored to a limited extent23,24. There remains a need to identify other inhibitors to treat microsporidia infections in both animals and agriculturally relevant species15.
Microsporidia can only grow inside of host cells, and it has been challenging to screen for inhibitors of this parasite. The model organism Caenorhabditis elegans has proven to be an effective model organism for drug discovery25. Several species of microsporidia infect this nematode, and studies using C. elegans have contributed to our understanding of microsporidian infection mechanisms and host immunity26. The most common species found infecting C. elegans is Nematocida parisii, which infects intestinal cells and leads to a decrease in size, reproductive capacity, and life expectancy27,28,29,30. N. parisii infection begins when spores are ingested and then use their unique infectious apparatus called the polar tube to deposit sporoplasms inside of intestinal cells. The sporoplasms then grow into meronts which differentiate into spores which are then defecated from the worms26,31. We recently developed a high-efficiency C. elegans-N. parisii drug screening system for the discovery and characterization of novel inhibitors of microsporidia15,24. Infection with N. parisii reduces the ability of C. elegans to produce progeny, allowing for a quantifiable phenotype that can be reversed with microsporidia inhibitors.
To identify novel microsporidia inhibitors, we screened 2500 compounds from the ChemBridge DIVERSet library. We identified and validated 18 compounds which could reduce the levels of N. parisii infection in C. elegans. Most of the inhibitors we identified were halide-containing benzenesulfonamides which inhibited microsporidia infection by inactivating spores. We show that one of these benzenesulfonamides exhibits broad-spectrum inhibitory effects on microsporidia, affecting species within the Nematocida, Enterocytozoon, and Encephalitozoon clades. We performed a targeted screen of other benzenesulfonamides, identifying several with increased affinity. Overall, this study identifies benzenesulfonamides as broad-spectrum inhibitors of microsporidian spores.
Results
Screen of ChemBridge DIVERset library identifies microsporidia inhibitors
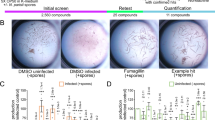
To identify novel microsporidia inhibitors we screened 2500 small-molecule compounds from the ChemBridge DIVERset library collection32. Using our previously described 96-well infection assay, compounds were incubated with N. parisii spores (Fig. 1A). C. elegans at the earliest larval (L1) stage were added one hour later and then cultured in liquid at 21 °C for five days15,24. As a result of microsporidia infection, C. elegans produce fewer embryos. Compounds which inhibit microsporidia can restore the ability of the host animals to produce progeny. In order to quantify offspring production after incubation with spores and each compound, each well was stained with rose bengal and scanned using a flatbed scanner, followed by automated quantification of progeny number33 (Fig. 1B). Each compound was screened in triplicate at a concentration of 60 µm (Data S1). Compared to uninfected controls, 26 compounds increased progeny production by 40% in infected worms (Fig. 1C). These compounds increased the ability to make progeny 2.4-17-fold compared to infected controls (Supplementary Fig. 1 and Supplementary Data 1).
A Schematic of N. parisii infection of C. elegans. The process of infection begins when the worms ingest microsporidia spores. These spores then germinate inside of the intestinal lumen of the worm, depositing the sporoplasm inside an intestinal cell. The sporoplasm then differentiates into a meront and differentiates into spores which then exit the worm. B Schematic of assay to identify microsporidia inhibitors. Compounds, microsporidia spores, and E. coli (food source of C. elegans) are incubated together and the C. elegans animals at the earliest larval stage are added. After 5 days of incubation, the worms are stained with rose bengal, imaged using a flatbed scanner, and quantified using image analysis. C Compounds at a final concentration of 60 µM were incubated with N. parisii. L1 stage C. elegans were added one hour later, cultured for five days, and progeny numbers quantified. Points represent mean progeny production of a compound as a percentage of DMSO uninfected controls. Compounds with an activity less than 40% are colored blue, compounds with an activity of at least 40% are colored red, and compound-IDs are shown. Three independent biological replicates were performed for each compound. D Clustered heat map of structural similarity of the 26 compounds with at least 40% activity. The scale indicates compound similarity with 0 (blue) being most similar and 1 (red) being least similar. Compounds tested in subsequent experiments are indicated by the dark blue dots. E Structures of benzenesulfonamides with inhibitory activity against N. parisii.
Hierarchical clustering analysis was used to determine the structural similarity of the identified microsporidia inhibitors (Fig. 1D). This analysis revealed a large cluster of ten compounds which are all benzenesulfonamide derivatives. All of these compounds contained both nitrogen- and sulfur-linked benzene rings. These 10 compounds also all contained at least 1 halide connected to a benzene ring (Fig. 1E). We also observed a second large cluster of compounds that contained either an aromatic imine or a pyrazole ring (Supplementary Fig. 2). The other 8 compounds were mostly dissimilar from each other.
Validation of compounds which inhibit N. parisii infection of C. elegans
To validate inhibitors identified in our screen, we selected 20 out of the 26 hits and quantified if the compounds could inhibit the infection of N. parisii. In a similar manner to our initial screen, N. parisii spores were cultured in the presence of compounds for one hour in 24-well plates, then L1 stage worms were added and cultured in liquid at 21 °C for four days. Afterwards, worms were fixed and stained with direct yellow 96 (DY96) which binds to both microsporidia spores and to C. elegans embryos34 (Fig. 2A). In the presence of N. parisii spores, 18 of the 20 compounds, as well as the known microsporidia inhibitor dexrazoxane, significantly increased the proportion of adult worms containing embryos (which we refer to as gravid) (Fig. 2B)24. To determine the relationship between embryo numbers per worm from our validation experiment and progeny production from our screen, we performed a linear correlation which shows a moderate correlation with an R2 of 0.3044 (Supplementary Fig. 3). The lack of strong correlation between the assays may be because the initial screen is quantifying total progeny production compared to the static quantification of embryo number. Among the newly identified compounds, 16 of them significantly reduce the number of animals displaying newly formed N. parisii spores (Fig. 2C). Together our data shows that our high-throughput screen can identify inhibitors with high confidence. Additionally, all ten of the benzenesulfonamide inhibitors we identified significantly restore progeny production in infected C. elegans and 9 of them significantly inhibit N. parisii infection.
A–C N. parisii spores were incubated with compounds at a final concentration of 100 μM. One hour later, L1 stage worms were added and cultured for 4 days. Worms were then fixed and stained with DY96 to detect microsporidian spores and nematode embryos. A Representative images of continuous infection assay with the conditions DMSO uninfected (negative control), DMSO infection (positive control for infection), dexrazoxane (positive control for inhibition), and 5357859 (benzenesulfonamide compound). DAPI (top row) stains worm nuclei and DY96 (middle row) stains microsporidia spores and C. elegans embryos. Merged images (bottom row) show worms that contain microsporidia spores and/or embryos. Scale bars are 500 μm. B The percentage of animals which contain embryos. C The percentage of animals with newly formed spores. Benzenesulfonamide compounds are colored red. n = 3 biological replicates, N = ≥ 100 worms counted per biological replicate. The P values were determined by one-way ANOVA with post hoc test, with all comparisons to DMSO infected. Means ± SD (horizontal bars) are shown. (*p < 0.05, **p < 0.01, ****p < 0.0001, ns means not significant).
Sulfonamides are a class of compounds that have been used as drugs in humans and animals to treat infectious disease35. We tested five medicinal sulfonamides for their activity against N. parisii. We did not observe any significant effect of these compounds in restoring embryo production of infected C. elegans or reducing infection levels (Supplementary Fig. 4).
Characterization of the mechanisms by which compounds inhibit N. parisii
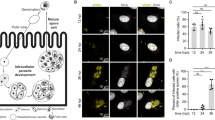
Inhibitors of microsporidia could function by blocking infection or the ability of the intracellular stages of the parasite to proliferate inside of host cells. To determine if these compounds reduced microsporidia proliferation, we performed pulse-chase experiments. We infected the worms with spores, then washed them three hours later to remove non-ingested spores. Next, the infected worms were incubated with each of the 20 ChemBridge compounds, or dexrazoxane which has been previously shown to limit N. parisii growth24 (Fig. 3A). One of the ChemBridge compounds, 5357859, increased the proportion of gravid worms, though to a much lower extent than dexrazoxane (Fig. 3B). None of the other 19 ChemBridge compounds reduced the percentage of worms displaying newly generated spores (Fig. 3C). To examine if ChemBridge compounds displayed inhibition of the meront stage of microsporidia growth, we stained meronts using Fluorescence in situ hybridization (FISH) at two days (before spore formation) or four days (after spore formation). Of the ChemBridge compounds, only 5357859 significantly reduced the amount of meronts in the worms, and to a much lesser extent than dexrazoxane (Fig. 3D, E). Similar to dexrazoxane, 5357859 did not decrease the rate of infection among animals, demonstrating that this compound does not promote pathogen clearance (Fig. 3F).
A–F L1 stage worms were incubated for 3 h in the presence of N. parisii spores and then washed to remove undigested spores. Infected worms were incubated with compounds for 2 (D) or 4 (A–C, E) days, fixed, and stained with DY96 and a FISH probe specific to the N. parisii 18S rRNA. A Representative images of pulse infection assay with the conditions DMSO infection (positive control for infection), 5357859 (benzenesulfonamide compound), and dexrazoxane (positive control for inhibition). DAPI (top row) stains worm nuclei and FISH (middle row) stains microsporidia meronts. Merged images (bottom row) show worms that contain microsporidia meronts. B The percentage of animals which contain embryos. C The percentage of animals which contain newly formed spores. D, E Quantification of the pathogen load at 2 (D) or 4 (E) days following infection. F The percentage of animals with FISH signal. Benzenesulfonamide compounds are colored red. n = 3 biological replicates, N = ≥ 100 worms (B, C, F) or N = 10 (D, E) counted per biological replicate. The P values were determined by one-way ANOVA with post hoc test, with all comparisons to DMSO infected, except for those indicated by brackets. Means ± SD (horizontal bars) are shown. (**p < 0.01, ****p < 0.0001, ns means not significant).
Dormant microsporidian spores can germinate (also called spore firing) within seconds of exposure to environmental stimuli36. In the intestinal lumen, many microsporidia, including N. parisii, germinate, depositing their sporoplasms inside of host cells, resulting in empty spores that no longer contain sporoplasms37,38. To determine whether any of the 18 ChemBridge compounds that had a significant effect on infected C. elegans progeny production also affect spore germination, we conducted spore firing assays24. Compounds were incubated with N. parisii spores for 24 h, and then the spores were washed to remove the compounds. These treated spores were then used to infect L1 stage C. elegans for three hours. To visualize the sporoplasms and spores in the intestinal lumen, they were stained with both FISH and DY96. To determine the spore empty rate, we counted the number of empty spores divided by the total number of spores. ZPCK, a compound previously shown to inhibit spore germination24, and two ChemBridge compounds, 5313458 and 5343775, significantly reduced spore empty rates (Fig. 4A). Surprisingly, even though most of the compounds did not affect spore germination, the 18 ChemBridge compounds, as well as ZPCK, significantly decreased the amount of sporoplasms that invaded each worm (Fig. 4B). One possibility that would explain this incongruity is that the compounds could potentially decrease animal infection by inducing spore firing outside of the host. In order to determine whether any of the 18 ChemBridge compounds could induce spore firing in vitro, we incubated compounds with N. parisii spores for 24 h, fixed and stained with DY96 and a N. parisii 18S rRNA FISH probe. None of the compounds significantly triggered spore germination in vitro (Supplementary Fig. 5).
A, B After incubating N. parisii spores with compounds for 24 h, spores were washed and added to L1 stage worms. After incubation for 3 h, worms were fixed and the spores (DY96) and sporoplasms (FISH) were stained. A Percentage of empty spores in the intestinal lumen. B The mean number of sporoplasms per worm. Benzenesulfonamide compounds are colored red. (n = 3, N = ≥ 50 spores counted per biological replicate). The P values were determined by one-way ANOVA with post hoc test. Means ± SD (horizontal bars) are shown. (**p < 0.01, ***p < 0.001, ****p < 0.0001, ns means not significant).
N. parisii spores are inactivated by benzenesulfonamide derivatives
One possibility that could explain how the identified compounds could block microsporidia invasion without altering the rate of empty spores, is if the compounds were lethal to the spores and facilitated the lysis of their sporoplasms by the intestinal contents of C. elegans. To test this hypothesis, we first performed mortality assays using the 18 Chembridge compounds and heat treatment as a control. After incubating the compounds with N. parisii spores for 24 h, spores were stained with Calcofluor White M2R, which binds to the spores, and Sytox Green, which only stains the DNA of inviable spores (Fig. 5A). Heat treatment and 12 of the ChemBridge compounds, including all ten of the benzenesulfonamide derivatives, significantly increased spore mortality (Fig. 5B). We then tested if spores treated with either acetone or with several ChemBridge compounds could cause the spores to no longer have their sporoplasm. Spores were treated with acetone alone, compounds alone, or compounds followed by treatment with acetone. These spores were then either incubated with L1 stage C. elegans for three hours or analyzed in vitro for spore mortality. The treatment of spores with acetone or benzenesulfonamide derivatives results in a significant increase in spore mortality (Fig. 5C). In intestinal lumen, both 5343775 and 5357859 as well as their acetone co-treatment groups, show higher empty rates than the acetone treatment group (Fig. 5D). These results suggest that spores treated with the compound can lead to sporoplasm destruction in the intestinal lumen.
A, B N. parisii spores were treated with the indicated compounds for 24 h, followed by Sytox Green and Calcofluor White M2R staining. A Representative images of spore mortality assay with the conditions spore prep. (negative control), heat-treated (positive control) and 5357859 (benzenesulfonamide compound). Calcofluor White M2R (top row) stains spores and Sytox Green (middle row) stains the DNA of spores that are no longer viable. Spores that are dual stained (marked by arrowheads) in the merged image (bottom row) are counted as inviable, and those that are viable (marked by arrows) are only stained blue. Scale bars are 50 μm. B Percentage of non-viable spores. C, D N. parisii spores were incubated with the indicated compounds, acetone, or both for 24 h. Spores were then washed and stained with Sytox Green and Calcofluor White M2R (C) or mixed with L1 stage worms incubated for 3 h, fixed, and stained with DY96 and a FISH probe (D). C Percentage of non-viable spores. D Percentage of empty spores in the intestinal lumen. N = ≥ 100 spores (B, C) or N = ≥ 50 worms (D) counted per biological replicate. The P values were determined by one-way ANOVA with post hoc test. Means ± SD (horizontal bars) are shown. (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, ns means not significant).
Multiple species of microsporidia are inhibited by benzenesulfonamides
In order to determine whether the compounds we identified are effective against other microsporidia species, we first tested several strong N. parisii inhibitors of different chemical structures against P. epiphaga, which infects the epidermis and muscle of C. elegans. This species belongs to the Enterocytozoonida clade, which also contains the human-infecting species V. cornea and E. bieneusi29,39,40. FISH staining was used to determine whether the five selected ChemBridge inhibitors, including two benzenesulfonamides, could inhibit P. epiphaga infection in C. elegans. All of the tested compounds significantly inhibit P. epiphaga infection (Fig. 6A).
A P. epiphaga spores were incubated with compounds at a concentration of 200 μM for 1 h and then L1 stage C. elegans were added and cultured for 4 days. Animals were then fixed and stained with DY96 and a P. epiphaga 18S rRNA FISH probe. Quantification of pathogen load. B E. cuniculi spores were incubated with 5337895, acetone, or 1% DMSO for 24 h, followed by staining with Fluorescent Brightener 28, Sytox Green, and propidium iodine. Quantification of spore mortality. C, D E. cuniculi spores were incubated with 5337895, acetone, or 1% DMSO for 24 h. Spores were then added to RK-13 cells and incubated for 3 days. C Cells were fixed, stained with Fluorescent Brightener 28 and parasitophorous vacuole number counted. D Quantitation of E. cuniculi SSU concentration. E, F RK-13 cells were incubated with E. cuniculi spores and either 5 μM dexrazoxane, 25 μM 5357859, or 200 nM albendazole for either 3 or 5 days. E Cells were fixed, stained with Fluorescent Brightener 28, and parasitophorous vacuole number counted. F Quantitation of E. cuniculi SSU concentration. Benzenesulfonamide compounds are colored red. n = 3 biological replicates, N = 10 animals (A), N = ≥ 100 spores (B), N > = 10 figure (C, E), or N = three well samples (D, F) qquantified per biological replicate. The P values were determined by one-way ANOVA (A, B) or two-way ANOVA(C–F) with post hoc test. Means ± SD (horizontal bars) are shown (*p < 0.05, **p < 0.01, ****p < 0.0001, ns means not significant).
We then tested the ability of the benzenesulfonamide 5357859 to inhibit the microsporidian Encephalitozoon cuniculi, which can infect various hosts including farm and companion animals, as well as humans41. We first used mortality assays to determine that 5357859 effectively inactivates mature E. cuniculi spores in vitro (Fig. 6B). We treated spores with 5357859 and used them to infect RK-13 cells. We then measured infection levels using parasitophorous vacuole counting and real-time PCR analyses and found that these compounds significantly reduce the pathogen load after incubation for 3- or 5-days (Fig. 6C, D). To determine if treating E. cuniculi and cells at the same time could reduce infection, we incubated RK-13 cells with 5357859, albendazole, or dexrazoxane at the maximum safe dose we determined from cytotoxicity experiments (Supplementary Fig. 6). After 3 days of incubation, we did not observe any significant decreases in parasitophorous vacuoles or Small Subunit (SSU) ribosomal RNA copies when treated with any of the compounds. After 5 days of incubation, we observed a significant decrease in the parasite by both measurements for dexrazoxane, albendazole, and 5357859 (Fig. 6E, F).
Identification of higher affinity benzenesulfonamide analogs that inhibit N. parisii infection
To identify higher affinity benzenesulfonamide inhibitors, we screened a library of 475 ChemBridge benzenesulfonamide analogs. To identify benzenesulfonamide analogs with stronger inhibitory effects, we decreased the compounds concentration to 15 μM from the 60 μM that we used in the original screen. There were 7 benzenesulfonamides from the previous screen that served as negative controls, and none of them increased progeny production in infected worms by more than 18% (Data S1). We identified three compounds that increased progeny production in infected worms by more than 40% compared to the uninfected controls (Fig. 7A). Two of these compounds, 5351234 and 5354953, differ from 5357859 only by the position of the chlorines on the nitrogen-bound benzene. The other compound, 7819020, contains two fused benzene rings, with one of them connected to a bromine (Fig. 7B).
A Compounds at a final concentration of 15 µM were incubated with N. parisii. L1 stage C. elegans were added one hour later and then cultured for five days. The number of worms in each well were quantified and the progeny production of each compound as a percentage of the uninfected controls is shown. Three independent biological replicates were performed for each compound. The green sulfonamide-ID represents sulfonamides that exhibited an activity of at least 40%. The red sulfonamide-ID represents the activity of the previously identified inhibitor 5357859. B The chemical structures of 5357859 and the 3 ChemBridge benzenesulfonamides with higher affinity.
To validate and characterize the higher affinity benzenesulfonamide inhibitors, we treated both infected and uninfected worms with 5357859 and the three higher affinity benzenesulfonamides at a concentration of 15, 60 or 100 μM. To determine toxicity to C. elegans, we measured the proportion of uninfected animals that were gravid. None of the compounds significantly reduced the proportion of gravid animals at 15 μM (Fig. 8A). However, 5351234 at 60 μM or higher, and 5354953 at 100 μM significantly reduced population gravidity (Fig. 8E, I). We then determined the ability of these compounds to restore embryo production in animals infected with N. parisii. All of the compounds increased the percentage of gravid worms in the presence of N. parisii at 15, 60 or 100 μM (Fig. 8B, F, J). At the lowest concentration 5357859 did not rescue gravidity to uninfected levels (Fig. 8B). At the highest concentration 5351234 and 5354953 did not rescue gravidity (Fig. 8J). We also evaluated whether these compounds could inhibit N. parisii infection, observing that all compounds mostly eliminated infection at the highest concentration, and that the three higher affinity benzenesulfonamides had higher activity than 5357859 at 15 μM (Fig. 8C, G, K). Finally, we tested the ability of these compounds to cause spore mortality, observing that all compounds could and at 15 μM, 5357859 demonstrated a significantly reduced ability to inactivate spores than the other three (Fig. 8D, H, L).
A–C, E–G, I–K N. parisii spores were incubated with the indicated benzenesulfonamides at different concentrations for 1 h. L1 stage C. elegans either without spore or mixed with the treated spores were cultured for four days. Worms were then fixed and stained with DY96. Effect of the indicated benzenesulfonamides at 15 µM (A–C), 60 µM (E–G) or 100 µM (I–K) on the percentage of worms with embryos in uninfected worms (A, E, I), infected worms (B, F, J) and the percentage of worms with newly formed spores (C, F, K). N. parisii spores were incubated with 15 µM (D) 60 µM (H) and 100 µM (L) compound for 24 h and stained with Sytox Green and Calcofluor White M2R. The percentage of spores that showed Sytox Green staining. n = 3, N = ≥ 100 worms (A–C, E–G, I–K) or N = ≥ 100 spores (D, H, L) counted per biological replicate). The P values were determined by one-way ANOVA with post hoc test. Means ± SD (horizontal bars) are shown. (*p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001, ns means not significant).
Discussion
Through two rounds of high-throughput screening of a total of ~3000 compounds, we identified 13 halide-substituted benzenesulfonamides, some of which can deactivate spores at low micromolar concentrations. We find that these compounds predominantly function by making spores non-viable and preventing invasion of host cells. We also find that these compounds have activity against several species of microsporidia. Although these compounds could potentially be used as antiseptics, at higher concentrations that cause nearly complete protection against infection, slight host toxicity is observed.
There has been effort to identify compounds that can be used as disinfectants or antiseptics to treat microsporidia spores42. Treatments such as hydrogen peroxide, ethanol, bleach, ozone, chlorine have been demonstrated to be effective against Encephalitozoon species43,44,45,46. However, some treatments, such as levulinic acid in combination with SDS, that are very effect against bacteria, do not inactivate microsporidia spores47. Several types of compounds have been shown to have activity on spores, but limited toxicity to hosts, such as porphyrins with Nosema ceranae or massetolides with N. parisii48,49. Most of the benzenesulfonamides we identified cause spore mortality, but do not induce or block germination. Almost all the benzenesulfonamides appear to only act on spores, but 5357859, also has a slight infect on N. parisii proliferation. The data we have generated on a large number of benzenesulfonamide compounds could be useful for generating models to predict higher affinity compounds50.
Sulfonamides are an important class of medicinally active compounds, displaying a wide range of activities. Studies have indicated that sulfonamides act as antibacterial51,52, antifungal51, antiviral53, anti-tumour54, antimalarial55, anti-ulcer and anti-inflammatory agents56, as well as having anti-convulsant57 and anti-oxidant effects58,59. There are several case studies reporting treating microsporidia infections of humans with the sulfonamides sulfadiazine or sulfamethoxazole60,61,62. As these compounds were included as part of multi-drug regimens, their effectiveness in clinically treating microsporidia is unclear. We observe no activity of clinically used sulfonamides against microsporidia. Additionally, sulfonamides were reported to not have activity against E. cuniculi63. The benzenesulfonamides we identified in this study all contain halogen functional groups, whereas the clinically used sulfonamides we tested do not contain halides, but instead the benzene groups are functionalized with various nitrogen and oxygen modifications.
In addition to benzenesulfonamides, we found several other types of compounds that were capable of inactivating N. parisii. A series of diaryl derivatives, such as (E)-2-{[(3-ethynylphenyl)imino]methyl}-4-nitrophenol), an analog of 5461570, was shown to specifically inhibit the interaction of NusB-NusE, decrease bacterial rRNA levels, and demonstrate antimicrobial activity against a variety of clinical methicillin-resistant Staphylococcus aureus strains without significant cytotoxicity against human cell lines64. A high-throughput screening process targeting Leishmania major revealed that 5469758 inhibits the parasite’s growth and vitality65.
In conclusion, we have identified benzenesulfonamides with the potential to be used as antiseptics. Being able to treat spores without host toxicity would have potential application against microsporidia that infect agriculturally important animals such as with honey bees, shrimp, and silkworms9,10. To further develop these compounds as antiseptics, it will be important to determine the mechanism by which these molecules inactivate spores, which could be done by selecting for drug resistance spores66 and to identify molecules with enhanced selectivity for microsporidia.
Methods
C. elegans maintenance
Escherichia coli OP50-1, the food source for C. elegans, was grown for 18 h at 37 °C in lysogeny broth (LB) and then concentrated by centrifugation. The wild-type C. elegans strain N2 was cultivated as a mixed population, then L4 stage worms were picked onto nematode growth media (NGM) plates, which were seeded with 10x OP50-1 E. coli and kept at 21 °C for 4 days67. To synchronize the worms, M9 solution was used to wash them from the NGM plates, and sodium hypochlorite and sodium hydroxide were applied to bleach the worms. As soon as the embryos had been released from the gravid adults, they were washed and incubated at 21 °C for ~18 h to allow the embryos to hatch.
N. parisii spore preparation
Stocks of N. parisii (ERTm1) spores were prepared as previously described28,34. On NGM plates, C. elegans N2 worms were infected with N. parisii spores. After incubating the infected worms for several days, they were harvested and frozen at −80 °C. Infected worms were mechanically disrupted using zirconia beads (2 mm diameter), followed by the removal of animal debris using a 5 μm filter (Millipore). Preparations of N. parisii spores were confirmed to be free of bacterial contamination and stored at −80 °C. The concentration of spores in the sample was determined by counting DY96-stained spores using a sperm counting slide (Cell-VU).
Source of chemicals
ChemBridge DIVERSet screening compounds were acquired from SPARC Drug Discovery at the Hospital for Sick Children. These compounds were dissolved in 0.3 μl of DMSO at a concentration of 10 mM and arrayed into 96-well plates. For retesting, solid compounds were bought from ChemBridge. All compounds were stored at a temperature of −80 °C.
Screening of microsporidia inhibitors in 96-well plates
Quantifying the ability of compounds to restore the ability of C. elegans to produce progeny in the presence of N. parisii was performed as previously described15,24. 96-well plates containing ChemBridge compounds in columns 2 to 11 were filled with 25 μl of K-medium (51 mM NaCl, 32 mM KCl, 3 mM CaCl2, 3 mM MgSO4, 3.25 μM cholesterol) containing 5x OP-50-1 and N. parisii spores, except for column 1, where spores were not added. 500 nl of DMSO was added to columns 1 and 12 for the DMSO uninfected and infected controls. We then added 25 μl of K-medium containing 24,000 spores/μl of N. parisii and plates were incubated at room temperature (~21 °C) for 1 h. Then, 25 μl of K-medium containing 100 bleach-synchronized L1 worms was added. After all the additions, each well contained 100 bleach-synchronized L1 worms, 1% DMSO, 12,000 spores/μl N. parisii (except for uninfected controls) and 60 μM of compounds (except for infected and uninfected controls). This concentration had been previously used in a prior study on identification of microsporidia inhibitors in C. elegans24. Each 96-well plate was covered with a breathable adhesive porous film and placed inside humidity boxes wrapped in parafilm and incubated for five days at 21 °C on a shaker with 180 revolutions per minute. Three independent biological replicates were performed for each compound, for a total of 96 plates being screened.
Quantification of progeny production
After 5 days of incubation, a VIAFLO 96 Electronic pipette was used to add 10 μl of 0.3125 mg/mL Rose Bengal. Plates were wrapped in parafilm and incubated for 24 h at 37 °C, resulting in magenta staining of the worms. To each well, 240 μl of M9/0.1% Tween-20 was added and the plate was centrifuged for 1 min at 2200 x g. Wells were washed again by removing 200 μl of supernatant from each well and adding 150 μl of M9/0.1% Tween-20. After mixing the worms in the plate by repeated pipetting, 25 μl of the liquid was transferred to another 96-well white clear bottom plates containing 300 μl M9/0.1%Tween-20. Worms in the plate were allowed to settle for 30 min and then the plates were scanned using an Epson Perfection V850 Pro flat-bed scanner with the following settings: positive film holder, 4800 DPI, and 24-bit color. The images were modified using GIMP version 2.10.18, with horizontal and vertical gridlines positioned such that each well is separated by a grid. HTML color codes #000000 and #FFC9AF were removed. Additionally, the images were modified by applying unsharp masking with the following parameters (radius = 10, effect = 10, threshold = 0.5). Yellow, blue, cyan, and green hue saturation were altered by adjusting lightness to 100 and saturation to −100. Red and magenta were adjusted to have a lightness of −100 and a saturation of 100. Using LZW compression, each well was exported as a single. tiff image. The number of worms was counted using WorMachine33 running in MATLAB, with the pixel binarization threshold set to 30, the neighboring threshold set to 1, and the maximum object area was set to 0.003%.
Structural similarity measurements
Atom Pair fingerprints (APfp) were used to calculate the structural similarity between the 26 compounds meeting our threshold for displaying anti-microsporidia activity68,69 The APfp of these compounds was determined using ChemMine tools70. Z-score values were used as the similarity measure and hierarchical clustering was performed71.
Continuous infection assays
Compounds at a concentration of 100 μM, except for dexrazoxane which was at 60 μM, were incubated in 200 μl of K-medium containing 24,000 spores/μl of N. parisii for 1 h. We then added 200 μl of K-medium containing 800 bleach-synchronized L1 worms to each well. Each well of 24-well assay plates contained a total volume of 400 μl including 800 L1 worms and 12,000 N. parisii spores/μl. Plates were then covered with adhesive porous film, placed inside a box, and the boxes were enclosed in parafilm. Plates were incubated for four days at 21 °C using a shaker with 180 rpm. After incubation, the samples were washed twice with M9/0.1% Tween-20, fixed with acetone, stained with DY96, and analyzed by fluorescence microscopy.
Pulse infection assays
To generate populations of infected worms, ~8000 bleach-synchronized L1-stage worms, 30 million N. parisii spores, and 5 μl 10x OP50-1 were mixed and added to a 6-cm NGM plate. These plates were dried in a clean cabinet and then incubated for three hours at 21 °C. Worms were washed off of plates with M9/0.1% Tween-20, and then washed twice to remove undigested spores. Worms were added to 24-well plates. Each well of 24-well assay plates contained a total volume of 400 μl including 800 L1 worms with compounds present at a concentration of 100 μM, except for dexrazoxane which was at 60 μM. Three wells were tested for each biological replicate. Samples were incubated as described above, for either two or four days. Worms were then fixed in acetone and stained with DY96 and a FISH probe as described below.
Spore firing assays
Spores at a concentration of 24,000 spores/μl, were incubated with compounds in 2% DMSO at a concentration of 200 μM except for ZPCK, which was at 120 μM, for 24 h at 21 °C. The 24-well assay plate was prepared as described above after the spores were washed three times with K-medium. The assays concentrations were 12,000 spores/μl, 100 μM compounds except ZPCK (60 μM), and 1% DMSO. After 3 h of incubation, samples were fixed in acetone, stained with FISH and DY96, and observed with fluorescence microscopy.
Mortality assays
N. parisii spores were incubated at a concentration of 24,000 spores/μl with the compounds at 200 μM in 2% DMSO for 24 h at 21 °C. In the heat treatment group, the spores were heated at 100 °C for 10 min. The spores were washed twice with 1 ml H2O, resuspended in 100 μl solution containing 2 μg/ml Calcofluor White M2R and 8 μM Sytox Green nucleic acid stain, and incubated at room temperature (~21 °C for 10 min). A total of 2.5 μl of each mixture was then spotted onto slides, mixed with 2.5 μl of 2% agar, covered with a glass slide, and examined using fluorescence microscopy.
E. cuniculi spores were incubated at a concentration of 10,000 spores/μl with 200 μM compounds in 2% DMSO, 2% DMSO only, or acetone for 24 h at 37 °C. The spores were washed twice with 1 ml PBS/0.1% Tween-20, resuspended in 100 μl solution of 0.5 μg/ml Fluorescent Brightener 28, 2.5 μM Sytox Green nucleic acid and 0.5 μM propidium iodine. The mixture was incubated for 10 min to facilitate staining, followed by examination using the Olympus BX53 microscope (Olympus, Tokyo, Japan).
DY96 staining, fluorescence in situ hybridization (FISH), and fluorescence microscopy
After incubation, samples were washed twice with M9/0.1% Tween-20 to remove any remaining bacteria. Then the animals were fixed in 700 µl of acetone for 15 min and washed twice with PBS/0.1% Tween-20. DY96 solution (10 µg/ml DY96, 0.1% SDS in 1x PBS + 0.1% Tween-20) was added and samples were rotated for 30 minutes. The DY96 solution was then removed and EverBriteTM Mounting Medium with DAPI was added. For FISH staining, acetone-fixed samples were washed once with PBS/0.1% Tween-20 and then once with hybridization buffer (900 mM NaCl, 20 mM Tris HCl, 0.01% SDS) Samples were then incubated in the hybridization buffer containing 5 ng/µl microB N. parisii 18S rRNA FISH probe72 (ctctcggcactcctcctg) conjugated to Cal Fluor Red 610 (LGC Biosearch Technologies) for 24 h at 46 °C. Samples were washed once with 1 ml wash buffer (50 ml hybridization buffer + 5 mM EDTA). Samples were then stained with 20 µg/µl DY96. Zeiss Zen 2.3 was used to obtain images of the samples using a ZEISS Axio Imager 2 at magnifications ranging from 5x to 63x. Animals with any number of embryos were counted as gravid worms. An infected worm was defined as an animal that exhibits any newly formed spores.
P. epiphaga infection assays
Stocks of P. epiphaga (JUm1396) spores were prepared as described above for N. parisii. The infection experiments with P. epiphaga were conducted using 24-well assay plates, with each well containing a final volume of 400 μl in K-medium, 800 L1 worms, and 60,000 P. epiphaga spores/μl. Except for dexrazoxane, which was used at 60 μM, all compounds were used at a concentration of 100 μM. The DMSO was used at a final concentration of 1%. Plates were incubated as described above for 4 days. The samples were then fixed in acetone and washed twice, stained with a CAL Fluor Red 610 FISH probe specific to P. epiphaga 18S rRNA (CTCTATACTGTGCGCACGG). Worms were imaged as described above and FISH fluorescence was quantified using the threshold and measure tools in ImageJ 2.9.073.
Cell culture and preparation of E. cuniculi spores
E. cuniculi spores (GB-M1) were propagated in RK-13 (rabbit kidney) cells as previously described74. The RK-13 cells were cultured in minimal essential medium supplemented with 10% fetal bovine serum and maintained at a constant temperature of 37 °C with 5% CO2. Following a cultivation period of 7 days, the RK-13 cells and spores were harvested. Cells were lysed by osmotic shock using water and pipetting to facilitate the release of spores. Cellular debris was removed using a 5 μm filter. Spore concentration was determined using a serial dilution method, followed by enumeration of the spores75.
Infection of cells with pretreated E. cuniculi spores
100 μl containing 10 million E. cuniculi spores were incubated with 200 μM 5357859, 2% DMSO, or acetone for 24 h at 37 °C. The spores were then washed twice with 1 ml PBS/0.1% Tween-20. We then mixed 1 × 106 spores with 1 × 105 seeded RK-13 cells which were then added into each well of 12-well plate. The medium in the well was refreshed every 3 days. At 0, 3 or 5 days after incubation, the culture media was removed and washed twice with PBS and the cells were harvested for microscopy or qPCR detection.
Continues infection with E. cuniculi spores
After seeding 5 × 104 RK-13 cells in a 24-well plate, medium containing 5 μM dexrazoxane, 25 μM 5357859, or 200 nM albendazole was added into wells. After 8 hours, 2.5 × 105 E. cuniculi spores were added to each well. The medium in the wells was refreshed every 3 days. After either 3 or 5 days after incubation with spores, the culture media was removed and the wells were washed three times with PBS, and the cells were harvested for microscopy or qPCR detection.
Parasitophorous vacuoles counting
After infection of RK-13 cells with E. cuniculi spores either 3 or 5 days, the culture medium was discarded, and the cells were washed three times with PBS/0.1% Tween-20. Cells were then fixed in 4% paraformaldehyde for 15 min and washed 3 times with PBS/0.1% Tween-20 to remove excess fixative. 0.5 μg/ml Fluorescent Brightener 28 working solution was then added to the samples and incubated in the dark for 30 min. The cells were then washed three times with PBS/0.1% Tween-20. The number of parasitophorous vacuoles was then examined and counted under a AX10 fluorescence microscope (Germany).
SYBR Green Real-time PCR
DNA was extracted from cells utilizing the ONEGA E.Z.N.A. Tissue DNA Kit. Absolute fluorescence quantification PCR was conducted in accordance with previously described methods (Forward primer ECUNF1: 5’-TCCTAGTAATAGCGGCTGACGAA-3’, Reverse primer ECUNR2: 5’-ACTCAGGACTCAGACCTTCCGA-3’)76. The qPCR reactions were carried out using the SYBR Green qPCR Kit. All real-time PCR experiments were conducted in triplicate using the FQD-96A Fluorescence Quantitative Polymerase Chain Reaction Detection System (Bioer Technology, Hangzhou, China).
Analyses of cell viability
Cell viability was assessed using the Cell Counting Kit-8 (CCK-8). A total of 10,000 cells per well, along with the compounds, were seeded into 96-well plates. At the tested time points, 10 μl of CCK-8 reagent (C0041, Beyotime, China) was added to every well as per the manufacturer’s instructions. The cells were then incubated for an additional hour at 37 °C with 5% CO2. Finally, the optical density was measured at 450 nm.
Statistical analyses
The data were collected from three independent biological repeats and analyzed by GraphPad Prism 8. P-values were determined by ANOVA. Statistical significance was defined as p < 0.05, *p < 0.01, ***p < 0.001, and ****p < 0.0001.
Data availability
Data is provided within the manuscript and in Data S1.
References
Han, B. & Weiss, L. M. Microsporidia: obligate intracellular pathogens within the fungal kingdom. Microbiol. Spectr. 5, 5.2.03 (2017).
Keeling, P. J. & Fast, N. M. Microsporidia: biology and evolution of highly reduced intracellular parasites. Annu. Rev. Microbiol. 56, 93–116 (2002).
Willis, A. R. & Reinke, A. W. Factors that determine microsporidia infection and host specificity. Exp. Suppl. 2012 114, 91–114 (2022).
Vossbrinck, C. R., Andreadis, T. G. & Weiss, L. M. Phylogenetics: taxonomy and the microsporidia as derived fungi. in Opportunistic Infections: Toxoplasma, Sarcocystis, and Microsporidia (eds. Lindsay, D. S. & Weiss, L. M.) vol. 9 189–213 (Springer, 2004).
Han, B., Takvorian, P. M. & Weiss, L. M. Invasion of host cells by microsporidia. Front. Microbiol. 11, 172 (2020).
Murareanu, B. M., Sukhdeo, R., Qu, R., Jiang, J. & Reinke, A. W. Generation of a microsporidia species attribute database and analysis of the extensive ecological and phenotypic diversity of microsporidia. mBio 12, https://doi.org/10.1128/mbio.01490-21 (2021).
Luallen, R. J. et al. Discovery of a natural microsporidian pathogen with a broad tissue tropism in Caenorhabditis elegans. PLOS Pathog. 12, e1005724 (2016).
Meng, X.-Z. et al. Pathological analysis of silkworm infected by two microsporidia Nosema bombycis CQ1 and Vairimorpha necatrix BM. J. Invertebr. Pathol. 153, 75–84 (2018).
Patil, P. K. et al. Economic loss due to diseases in Indian shrimp farming with special reference to Enterocytozoon hepatopenaei (EHP) and white spot syndrome virus (WSSV). Aquaculture 533, 736231 (2021).
Botías, C. et al. The growing prevalence of Nosema ceranae in honey bees in Spain, an emerging problem for the last decade. Res. Vet. Sci. 93, 150–155 (2012).
Franzen, C. Microsporidia: a review of 150 Years of Research. Open Parasitol. J. 2, 1–34 (2008).
Han, B., Pan, G. & Weiss, L. M. Microsporidiosis in Humans. Clin. Microbiol. Rev. 34, e00010–e00020 (2021).
Han, B. & Weiss, L. Therapeutic targets for the treatment of microsporidiosis in humans. Expert Opin. Ther. Targets 22, 903−915 (2018).
Wei, J., Fei, Z., Pan, G., Weiss, L. M. & Zhou, Z. Current therapy and therapeutic targets for microsporidiosis. Front. Microbiol. 13, 835390 (2022).
Huang, Q., Chen, J., Pan, G. & Reinke, A. W. Screening of the Pandemic Response Box identifies anti-microsporidia compounds. PLoS Negl. Trop. Dis. 17, e0011806 (2023).
Jyothi, N. B. & Patil, C. S. Chemotherapeutic effect of benzimidazole derivatives on Nosema bombycis naegeli in silkworm Bombyx mori l-an ultrastructural study. Biospectra 9, 49–54 (2014).
Akiyoshi, D. E. et al. Analysis of the β-Tubulin Genes from Enterocytozoon bieneusi Isolates from a Human and Rhesus Macaque. J. Eukaryot. Microbiol. 54, 38–41 (2007).
Franzen, C. & Salzberger, B. Analysis of the β-Tubulin Gene from Vittaforma corneae Suggests Benzimidazole Resistance. Antimicrob. Agents Chemother. 52, 790–793 (2008).
Lefkove, B., Govindarajan, B. & Arbiser, J. L. Fumagillin: an anti-infective as a parent molecule for novel angiogenesis inhibitors. Expert Rev. Anti Infect. Ther. 5, 573–579 (2007).
Sin, N. et al. The anti-angiogenic agent fumagillin covalently binds and inhibits the methionine aminopeptidase, MetAP-2. Proc. Natl. Acad. Sci. USA 94, 6099–6103 (1997).
Das, B. C. et al. Methionine aminopeptidases: potential therapeutic target for microsporidia and other microbes. J. Eukaryot. Microbiol. 71, e13036 (2024).
No, C. R. On pharmacologically active substances and their classification regarding maximum residue limits in foodstuffs of animal origin. Off. J. Eur. Communities 50, 1–72 (2010).
Abou-El-Naga, I. F., Said, D. E., Gaafar, M. R., Ahmed, S. M. & El-Deeb, S. A. A new scope for orlistat: effect of approved anti-obesity drug against experimental microsporidiosis. Med. Mycol. 57, 181–195 (2019).
Murareanu, B. M. et al. High-throughput small molecule screen identifies inhibitors of microsporidia invasion and proliferation in C. elegans. Nat. Commun. 13, 5653 (2022).
O’Reilly, L. P., Luke, C. J., Perlmutter, D. H., Silverman, G. A. & Pak, S. C. C. elegans in high-throughput drug discovery. Adv. Drug Deliv. Rev. 69–70, 247–253 (2014).
Gang, S. S. & Lažetić, V. Microsporidia: pervasive natural pathogens of Caenorhabditis elegans and related nematodes. J. Eukaryot. Microbiol. 71, e13027 (2024).
Balla, K. M., Luallen, R. J., Bakowski, M. A. & Troemel, E. R. Cell-to-cell spread of microsporidia causes Caenorhabditis elegans organs to form syncytia. Nat. Microbiol. 1, 1–6 (2016).
Willis, A. R. et al. A parental transcriptional response to microsporidia infection induces inherited immunity in offspring. Sci. Adv. 7, eabf3114 (2021).
Zhang, G. et al. A large collection of novel nematode-infecting microsporidia and their diverse interactions with Caenorhabditis elegans and other related nematodes. PLoS Pathog. 12, e1006093 (2016).
Tecle, E. et al. The purine nucleoside phosphorylase pnp-1 regulates epithelial cell resistance to infection in C. elegans. PLOS Pathog. 17, e1009350 (2021).
Jarkass, H. T. E. & Reinke, A. W. The ins and outs of host-microsporidia interactions during invasion, proliferation and exit. Cell. Microbiol. 22, e13247 (2020).
Gilad, Y., Nadassy, K. & Senderowitz, H. A reliable computational workflow for the selection of optimal screening libraries. J. Cheminform.7, 61 (2015).
Hakim, A. et al. WorMachine: machine learning-based phenotypic analysis tool for worms. BMC Biol. 16, 8 (2018).
Willis, A. R., El Jarkass, H. T. & Reinke, A. W. Studying inherited immunity in a caenorhabditis elegans model of microsporidia infection. J. Vis. Exp. e63636, https://doi.org/10.3791/63636 (2022).
Ovung, A. & Bhattacharyya, J. Sulfonamide drugs: structure, antibacterial property, toxicity, and biophysical interactions. Biophys. Rev. 13, 259–272 (2021).
Huang, Q. et al. Germination of microsporidian spores: the known and unknown. J. Fungi 9, 774 (2023).
Fries, I., Feng, F., da Silva, A., Slemenda, S. B. & Pieniazek, N. J. Nosema ceranae n. sp.(Microspora, Nosematidae), morphological and molecular characterization of a microsporidian parasite of the Asian honey bee Apis cerana (Hymenoptera, Apidae). Eur. J. Protistol. 32, 356–365 (1996).
Tamim El Jarkass, H. et al. An intestinally secreted host factor promotes microsporidia invasion of C. elegans. eLife 11, e72458 (2022).
Bojko, J. et al. Microsporidia: a new taxonomic, evolutionary, and ecological synthesis. Trends Parasitol. 38, 642−659 (2022).
Wadi, L. et al. Genomic and phenotypic evolution of nematode-infecting microsporidia. PLOS Pathog. 19, e1011510 (2023).
Magalhães, T. R., Pinto, F. F. & Queiroga, F. L. A multidisciplinary review about Encephalitozoon cuniculi in a One Health perspective. Parasitol. Res. 121, 2463–2479 (2022).
Leiro, J. M., Piazzon, C., Domínguez, B., Mallo, N. & Lamas, J. Evaluation of some physical and chemical treatments for inactivating microsporidian spores isolated from fish. Int. J. Food Microbiol. 156, 152–160 (2012).
Ortega, Y. R., Torres, M. P., Van Exel, S., Moss, L. & Cama, V. Efficacy of a sanitizer and disinfectants to inactivate Encephalitozoon intestinalis Spores. J. Food Prot. 70, 681–684 (2007).
John, D. E., Haas, C. N., Nwachuku, N. & Gerba, C. P. Chlorine and ozone disinfection of Encephalitozoon intestinalis spores. Water Res. 39, 2369–2375 (2005).
Huffman, D. E., Gennaccaro, A., Rose, J. B. & Dussert, B. W. Low- and medium-pressure UV inactivation of microsporidia Encephalitozoon intestinalis. Water Res. 36, 3161–3164 (2002).
Jordan, C. N., DiCristina, J. A. & Lindsay, D. S. Activity of bleach, ethanol and two commercial disinfectants against spores of Encephalitozoon cuniculi. Vet. Parasitol. 136, 343–346 (2006).
Ortega, Y. R., Torres, M. P. & Tatum, J. M. Efficacy of Levulinic Acid-Sodium Dodecyl Sulfate against Encephalitozoon intestinalis, Escherichia coli O157:H7, and Cryptosporidium parvum. J. Food Prot. 74, 140–144 (2011).
Jarkass, H. T. E. et al. The Caenorhabditis elegans bacterial microbiome influences microsporidia infection through nutrient limitation and inhibiting parasite invasion. 2024.06.05.597580 Preprint at https://doi.org/10.1101/2024.06.05.597580 (2024).
Buczek, K. et al. Bioactivity studies of porphyrinoids against microsporidia isolated from honeybees. Sci. Rep. 10, 1–16 (2020).
Stokes, J. M. et al. A deep learning approach to antibiotic discovery. Cell 180, 688–702.e13 (2020).
Krátký, M. et al. Antimicrobial activity of sulfonamides containing 5-chloro-2-hydroxybenzaldehyde and 5-chloro-2-hydroxybenzoic acid scaffold. Eur. J. Med. Chem. 50, 433–440 (2012).
Supuran, C. T., Casini, A. & Scozzafava, A. Protease inhibitors of the sulfonamide type: anticancer, antiinflammatory, and antiviral agents. Med. Res. Rev. 23, 535–558 (2003).
Supuran, C. T., Innocenti, A., Mastrolorenzo, A. & Scozzafava, A. Antiviral sulfonamide derivatives. Mini Rev. Med. Chem. 4, 189–200 (2004).
Huang, Z., Lin, Z. & Huang, J. A novel kind of antitumour drugs using sulfonamide as parent compound. Eur. J. Med. Chem. 36, 863–872 (2001).
Domínguez, J. N. et al. Synthesis and antimalarial activity of sulfonamide chalcone derivatives. Il Farm. 60, 307–311 (2005).
Chazalette, C. et al. Carbonic anhydrase inhibitors. Design of anticonvulsant sulfonamides incorporating indane moieties. Bioorg. Med. Chem. Lett. 14, 5781–5786 (2004).
Siddiqui, N. et al. Synthesis and anticonvulsant activity of sulfonamide derivatives-hydrophobic domain. Bioorg. Med. Chem. Lett. 17, 255–259 (2007).
Lahtinen, M. et al. Synthesis, characterization, thermal and antimicrobial studies of N-substituted Sulfanilamide derivatives. J. Mol. Struct. 1060, 280–290 (2014).
Alexiou, P. & Demopoulos, V. J. A diverse series of substituted benzenesulfonamides as aldose reductase inhibitors with antioxidant activity: design, synthesis, and in vitro activity. J. Med. Chem. 53, 7756–7766 (2010).
Field, A. S. et al. Myositis associated with a newly described microsporidian, Trachipleistophora hominis, in a patient with AIDS. J. Clin. Microbiol. 34, 2803–2811 (1996).
Coyle, C. M. et al. Fatal myositis due to the microsporidian brachiola algerae, a mosquito pathogen. N. Engl. J. Med. 351, 42–47 (2004).
Martins, S. A. R., Muccioli, C., Belfort, R. & Castelo, A. Resolution of microsporidial keratoconjunctivitis in an AIDS patient treated with highly active antiretroviral therapy. Am. J. Ophthalmol. 131, 378–379 (2001).
Beauvais, B., Sarfati, C., Challier, S. & Derouin, F. In vitro model to assess effect of antimicrobial agents on Encephalitozoon cuniculi. Antimicrob. Agents Chemother. 38, 2440–2448 (1994).
Qiu, Y. et al. Design, synthesis and biological evaluation of antimicrobial diarylimine and –amine compounds targeting the interaction between the bacterial NusB and NusE proteins. Eur. J. Med. Chem. 178, 214–231 (2019).
AID 1063 - Leishmania major promastigote HTS - PubChem. https://pubchem.ncbi.nlm.nih.gov/bioassay/1063.
Cowell, A. N. et al. Mapping the malaria parasite druggable genome by using in vitro evolution and chemogenomics. Science 359, 191–199 (2018).
Lewis, J. A. & Fleming, J. T. Basic culture methods. Methods Cell Biol. 48, 3–29 (1995).
O’Boyle, N. M. & Sayle, R. A. Comparing structural fingerprints using a literature-based similarity benchmark. J. Cheminform.8, 36 (2016).
Bajusz, D., Rácz, A. & Héberger, K. Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations?. J. Cheminform.7, 20 (2015).
Backman, T. W., Cao, Y. & Girke, T. ChemMine tools: an online service for analyzing and clustering small molecules. Nucleic Acids Res. 39, W486–W491 (2011).
Cao, Y., Charisi, A., Cheng, L.-C., Jiang, T. & Girke, T. ChemmineR: a compound mining framework for R. Bioinformatics 24, 1733–1734 (2008).
Troemel, E. R., Félix, M.-A., Whiteman, N. K., Barrière, A. & Ausubel, F. M. Microsporidia are natural intracellular parasites of the nematode caenorhabditis elegans. PLoS Biol. 6, e309 (2008).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Han, B., Moretto, M. & Weiss, L. Encephalitozoon: tissue culture, cryopreservation and murine infection. Curr. Protoc. Microbiol. 52, e72 (2019).
Moss, D. M., Croppo, G. P., Wallace, S. & Visvesvara, G. S. Flow cytometric analysis of microsporidia belonging to the genus encephalitozoon. J. Clin. Microbiol. 37, 371–375 (1999).
Reabel, S. Molecular diagnostic methods for Detection of Encephalitozoon cuniculi in pet rabbits (Diss. University of Guelph, 2012).
Acknowledgements
We thank Meng Xiao and Yin Chen Wan for helpful comments on this manuscript. We thank Dr. Han Bing for kindly providing GB-M1 E. cuniculi spores. This work was supported by Canadian Institutes of Health Research grant (no. 461807 to A. W. R.) and Q. H. was supported by an award from the China Scholarship Council.
Author information
Authors and Affiliations
Contributions
Contributions: Q. H. and A.R. conceived of the project. Q. H. performed the small-molecule screens and all experiments with N. parisii. H. J. and Y. D. performed experiments on E. cuniculi. J. W., G. P., J. C., and A. R. provided supervision. All authors analyzed the data. Q. H. and A. R. co-wrote the paper with edits from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, Q., Jiang, H., Wei, J. et al. Small-molecule screen in C. elegans identifies benzenesulfonamides as inhibitors of microsporidia spores. npj Antimicrob Resist 3, 41 (2025). https://doi.org/10.1038/s44259-025-00116-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00116-0