Abstract
Genomics research is essential for understanding the transmission and evolution of bacterial antimicrobial resistance. The development of nanopore long-read sequencing has revolutionized our understanding of the prevalence, spread, and dynamic evolution of bacterial ARGs. This review discusses the applications and advantages of nanopore sequencing technology in the complete bacterial genome construction, rapid ARG detection, and dynamic evolution analysis of multidrug resistance genetic structure. We emphasize the unique advantages of long-read nanopore sequencing in analyzing the genetic contexts of ARGs in both cultured bacteria and microbiota, as well as the rapid turnaround time for clinical bacterial resistance detection. Technical advancements and the future potential for expanding the applications of nanopore sequencing in the field of antimicrobial resistance are discussed.
Similar content being viewed by others
Introduction
The discovery of antimicrobials has saved countless lives over the past few decades. However, the widespread misuse and overuse of antimicrobials in human medicine, agriculture, and animal husbandry have resulted in the emergence and dissemination of bacterial antimicrobial resistance, threatening the effectiveness of treatments for bacterial infections1,2,3. Many newly developed antimicrobials face the challenge of bacterial resistance within just a few years of their introduction into clinical use1. The rapid development of antimicrobial resistance has significantly diminished the effectiveness of antimicrobials, making human beings face the troubling clinical situation of no effective antimicrobials available4,5. It is estimated that antimicrobial resistance will lead to approximately 1.91 million direct deaths and 8.22 million associated deaths globally by 20506. Multidrug-resistant (MDR) pathogens, such as carbapenem-resistant Acinetobacter baumannii, third-generation cephalosporin-resistant and carbapenem-resistant Enterobacteriaceae, rifampicin-resistant Mycobacterium tuberculosis, are particularly concerning, as they significantly limit therapeutic options for treating infections7,8,9,10. Decoding the genetic features of these MDR bacteria is essential for probing novel resistance mechanisms and the development of effective control strategies.
The rapid development of bacterial resistance is largely driven by the spread of antimicrobial resistance genes (ARGs)11. Mobile genetic elements, such as plasmids, transposons, and integrons, facilitate the horizontal transfer of ARGs across diverse bacterial species11,12. Understanding the detailed genetic structure of ARGs is critical to unraveling their transmission and regulatory mechanisms. Although high-throughput short-read sequencing has been widely used to decode the genome of MDR bacteria, these technologies are unable to accurately resolve complex genomic regions, repetitive sequences, and large mobile genetic elements, leading to incomplete or fragmented assemblies of resistance genes13,14,15,16. Recent advances in long-read sequencing technologies, particularly nanopore sequencing, have revolutionized our ability to characterize antimicrobial resistance mechanisms at a more detailed genomic level17,18,19,20. Nanopore sequencing, with its ability to generate ultra-long reads (N50 > 100 kb) and perform real-time sequencing, enables rapid analysis of bacterial genomes and metagenomes, offering unprecedented insights into the evolution, spread, and ecological impact of ARGs. In this review, we explore how nanopore sequencing has become an indispensable tool in the fight against antimicrobial resistance, offering innovative solutions for surveillance, diagnostics, and evolutionary investigation.
Development of nanopore sequencing and its technical advantage in bacterial resistance research
Historical milestones of nanopore sequencing
Nanopore sequencing is based on the principle of passing DNA strands through a protein nanopore, measuring the changes in electrical current as the DNA passes through the pore, and then using these changes to decipher the DNA sequence. As early as 1989, the concept of nanopore sequencing was documented in the notebook of David Deamer21. Over the course of more than 20 years of dedicated exploration, nanopore sequencing became a reality. In 2014, Oxford Nanopore Technologies (ONT) introduced their groundbreaking nanopore sequencer MinION, and welcomed the first early-stage users before commercialization. The development of nanopore sequencing has been accompanied by the successful resolution of numerous previously unimaginable challenges. These include the ability to thread a DNA molecule through a protein nanopore, the capacity to differentiate between purine and pyrimidine bases using current blockade signals, the recognition of individual bases within a DNA strand, and the precise control of DNA movement through the nanopore facilitated by phi29 DNA Polymerase (Fig. 1A)21,22,23,24. These pivotal technologies facilitate the controlled passage of individual DNA molecules through protein nanopores, resulting in the generation of electrical signal perturbations that can be captured and analyzed.
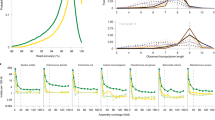
A The major events in the development of nanopore sequencing. It has taken 25 years from the concept of nanopore sequencing to the advent of the first nanopore sequencer ONT MinION. During the development of nanopore sequencing, four major technical points have been overcome, respectively making DNA molecular through the nanopores, distinguishing of purines and pyrimidines, recognizing a single base in an individual DNA strand, and controlling the rate of DNA passage through the nanopore. Since 2014, nanopore sequencing technology has been in a rapid development stage, and the throughput and accuracy of nanopore sequencing have greatly improved. In 2020, QitanTech, a Chinese company, announced China’s first nanopore sequencing platform, QNome-9606. After four years, Chinese MGI released the nanopore sequencing platform CycloneSEQ. B The development of the accuracy of nanopore sequencing. In the early stages, to improve the sequencing accuracy, the library preparation of nanopore sequencing usually uses a 2 d strategy, which means both strands of a dsDNA molecule to be sequenced. With the improvement of sequencing protein, the accuracy of single strand reads gradually catches up to the consensus sequences of the 2 d strategy. Of note, the accuracy of QitanTech nanopore sequencing has a significant improvement within the past two years. C Comparison of the advantages of nanopore sequencing relative to other sequencing platforms. Nanopore sequencing have many technological advantages including flexible, producing ultra-long sequencing reads and real-time sequencing.
In addition to ONT, several Chinese companies are actively developing nanopore sequencing technology. In 2020, QitanTech launched its first nanopore sequencing device, the QNome-9606 (Fig. 1A). QitanTech’s nanopore sequencing technology is based on principles and library preparation methods similar to ONT nanopore sequencing. However, the accuracy of the sequencing reads generated by QitanTech nanopore sequencing remains notably lower than that of ONT25. Despite this limitation, QitanTech nanopore sequencing has already proven sufficient for resolving the genomes of MDR bacteria and for conducting epidemiological studies of resistant bacteria within metagenomic samples25,26,27. In 2024, MGI Tech introduced another Chinese nanopore sequencing platform, CycloneSEQ (Fig. 1A). CycloneSEQ produces long reads with an average length of 11.6 kb and an average quality score of 14.4, and has been successfully applied to the reconstruction of bacterial genomes28. At this stage, nanopore sequencing technology has entered a phase of rapid growth, which is expected to accelerate further advancements, particularly in sequencing throughput and accuracy.
Improvements in throughput and accuracy of nanopore sequencing
Initially, the throughput of a single MinION nanopore sequencing flow cell could only produce a few hundred megabases (Mb) of sequencing data. Due to the limited sequencing throughput, nanopore sequencing was mostly used for microbial research. For example, in 2015, an Escherichia coli genome was successfully assembled into a single contig with an impressive nucleotide accuracy of 99.5% using only nanopore sequencing data (Fig. 1A)19. Following that, nanopore sequencing was applied to many research projects and as the demand for nanopore sequencing increased, the throughput of MinION became insufficient for many studies. To address this limitation, ONT introduced a higher throughput sequencing platform called GridION in the third year after the release of MinION. GridION allowed for simultaneous sequencing of five MinION flow cells, thereby increasing the overall throughput. Subsequently, in 2018, ONT released an ultra-high throughput sequencing platform called PromethION (Fig. 1A). PromethION utilized different flow cells compared to MinION, with a significantly larger number of channels (3000 channels per flow cell). Additionally, PromethION had the capability to support up to 48 flow cells in a single experiment, resulting in the production of several terabases (Tb) of sequencing data. These advancements in sequencing throughput, including the introduction of higher throughput platforms like GridION and PromethION, have significantly expanded the capabilities of long-read sequencing, made nanopore sequencing popular in multiple research fields, including antimicrobial resistance (Fig. 1A).
In addition to sequencing length and throughput, accuracy is another important consideration for the applications of nanopore sequencing. The initial versions of ONT’s flow cells, such as the R6, exhibited an error rate of over 30% (Fig. 1B)29. To generate more accurate sequences, early ONT used a 2 d library preparation method, which allowed both strands of a dsDNA molecule to be sequenced. By this way, a template read, a complement read and a consensus sequence generated by two reads were produced. Compared with template or complement reads, the accuracy of consensus sequences could be improved by about 10% in the R7 version flow cell (accuracy from 62% to 72%)18. Meanwhile, ONT has been continuously improving the nanopore protein, motor protein, and library preparation kit chemistry. As a result, the accuracy of consensus sequences produced by the R7.3 and R9.0 versions of the flow cell has exceeded 90% by 2017 (Fig. 1B)30.
Although the 2 d library preparation method initially provided a significant improvement in the accuracy of nanopore sequencing, it also resulted in the abandonment of many low-quality data, leading to a decrease in throughput. As nanopore and motor proteins continued to be improved, the advantage of the 2 d strategy in terms of accuracy gradually diminished. In the R9 version of the flow cell, the accuracy of consensus sequences could only be improved from 85% to 90% compared to single-strand reads (Fig. 1B)31. To strike a balance between accuracy and throughput, ONT gradually phased out the 2 d library preparation method and replaced it with the 1 d library preparation method, where each strand of a dsDNA is sequenced independently. With the release of the R9.4 version flow cell, the accuracy of nanopore sequencing using the 1 d library preparation method could reach over 90% (Fig. 1B)32,33.
In 2019, the development of nanopore sequencing ushered in a major innovation with the release of a novel version of flow cell R10. Compared to previous nanopores, R10 has a longer barrel and a dual reader head, which could capture two current perturbations of a strand of DNA through the nanopore. Accordingly, the accuracy of nanopore sequencing, particularly in regions with homopolymers, has been greatly enhanced. For example, in targeted sequencing of a two Mb region using human samples, the R10 nanopore demonstrated a high level of accuracy, with consensus sequences showing an identity of approximately 99.96% (Q34), which was comparable to both Illumina and PacBio. Two years after the release of R10, ONT introduced an updated version called R10.4 and announced the “Q20+ plan”. The Q20+ chemistry allows users to generate raw read sequencing data with an accuracy exceeding 99% (Q20) (Fig. 1A). Using ONT R10.4 data alone, with a sequencing coverage of approximately 40-fold, it is possible to obtain microbial genomes with accuracy comparable to those generated by polishing R9.4.1 data with Illumina data34. While there are indications that nanopore sequencing may eventually achieve accuracy comparable to next-generation sequencing technologies, further research is needed to confirm this potential.
Comparative advantages of nanopore sequencing with other sequencing platforms in bacterial antimicrobial resistance research
With the rapid advancement of nanopore sequencing technology, it has been widely used in bacterial antimicrobial resistance research due to its unique ability to generate long reads in real time (Fig. 1C)18,35,36,37. Unlike high-throughput short-read sequencing platforms like Illumina or MGI, which typically produce reads of a few hundreds of base pairs, nanopore sequencing provides long reads that can span entire MDR regions and complex genetic structures38. This capability enables a more comprehensive analysis of the bacterial genome, allowing for precise identification of the genetic structures of ARGs and mobile genetic elements such as plasmids, transposons, and integrons39,40. In addition to generating long reads, nanopore sequencing offers several practical advantages. One of the most notable is its portability and relatively low cost. Devices like the MinION are compact and affordable, making them ideal for on-site sequencing in field studies or clinical settings (Fig. 1C)41,42. In addition, even with ONT’s MinION sequencing flow cells, the throughput can reach 15–30 Gb. By applying ONT’s barcoding library strategy, multiple bacterial genomes can be sequenced in parallel on a single flow cell. At an average coverage of ~100× per genome, this enables the simultaneous reconstruction of 30-60 bacterial genomes, greatly enhancing the efficiency of large-scale strain surveillance and antimicrobial resistance monitoring while substantially reducing per-genome sequencing costs. This feature makes nanopore sequencing especially attractive for rapid, real-time monitoring of antimicrobial resistance, particularly in resource-limited environments or during infectious disease outbreaks43,44. In contrast, high-throughput platforms like Illumina or BGI require more expensive equipment and infrastructure, limiting their accessibility in certain contexts.
When compared to PacBio sequencing, another long-read platform, nanopore sequencing offers significant advantages in cost, portability, and sequencing read length33,45. While PacBio sequencing also produces long reads, it requires expensive equipment and is generally less accessible for smaller labs or field applications. In contrast, nanopore sequencing devices are compact, cost-effective, and capable of real-time operation, delivering immediate results. This feature is particularly valuable for monitoring resistance dynamics during outbreaks or clinical treatment. Furthermore, nanopore sequencing generates significantly longer reads than PacBio sequencing (ranging from tens to hundreds of kb, compared to PacBio’s typical tens of kb), which is critical for resolving complex MDR regions, such as tandem repeats, that are often difficult to assemble with shorter reads46. In terms of sequencing accuracy, PacBio HiFi reads can reach ~99.99%, whereas ONT’s latest Q20 chemistry produces duplex reads with accuracies exceeding 99%47,48. The exceptionally high accuracy of PacBio HiFi reads is particularly advantageous for detecting single-nucleotide variants and small indels with high confidence, which is crucial for characterizing resistance-conferring point mutations. However, the shorter average read length of HiFi reads may limit its capacity to fully resolve repetitive regions and large structural variations underlying ARG mobility. By contrast, ONT duplex reads, although slightly less accurate, can be further corrected using high-accuracy short reads. Consequently, the ability of nanopore sequencing to handle large and complex genomic regions offers unique insights into the evolution and spread of antimicrobial resistance, making it a more powerful tool for combating this growing global health threat than PacBio sequencing.
Rapid identification of bacterial antimicrobial resistance using nanopore sequencing
Rapid genome resolution of MDR bacteria
MDR bacteria pose a serious threat to human health, primarily due to the accumulation of ARGs mediated by mobile genetic elements49. Traditional short-read sequencing often struggles with accurately resolving complex MDR-associated genomic regions, such as antimicrobial resistance islands, genomic rearrangements, and large repetitive elements that are common in the genomes of MDR bacteria18. These unresolved regions can include critical genetic determinants of bacterial resistance, which are essential for understanding the mechanisms of resistance and developing targeted therapies.
Since its introduction in 2014, nanopore long-read sequencing technology has revolutionized bacterial genomics by enabling the direct resolution of complex MDR-associated genomic regions29. By generating long reads capable of spanning repetitive sequences and large genomic segments, nanopore sequencing overcomes many of the inherent limitations of short-read sequencing (Fig. 2A). Hybrid assembly strategies that combine nanopore long reads with short reads have since been widely applied, successfully resolving challenging regions such as chromosomal resistance islands in Salmonella Typhi18. Although nanopore sequencing rapidly produces structurally complete assemblies, earlier generations of the technology suffered from reduced nucleotide accuracy compared with reference genomes (Fig. 2A)19. To address this issue, polishing with short reads or hybrid assemblies was commonly used to improve assembled genome accuracy50,51,52,53,54.
A Application of nanopore sequencing in rapid bacterial genome resolution. Currently, the primary approach for constructing bacterial complete genomes is the hybrid assembly strategy, which combines nanopore long-read sequencing with high-accuracy short-read high-throughput sequencing. With the release of ONT R10.4 nanopore sequencing platform, complete MDR bacterial genomes can now be constructed using only nanopore sequencing data. B Identification and quantification of ARGs from metagenomes using nanopore sequecing. Compared with traditional metagenomic sequencing, nanopore long-read metagenomic sequencing offers more accurate identification of the genetic context of ARGs, enables tracking of ARG host information, and facilitates the assembly of more complete genomes of MDR bacteria. C Application of nanopore target sequencing in low-abundance ARGs detection. There are two primary enrichment strategies for detecting low-abundance ARGs: Cas9-based nanopore targeted sequencing and nanopore adaptive sampling sequencing.
More recently, ONT introduced the Q20+ chemistry with the R10.4 flow cell, achieving raw read accuracies exceeding 99% and enabling assemblies with >99.9% accuracy and fewer than 20 SNPs relative to reference genomes34,55. Importantly, these advances mean that complete and highly accurate bacterial genomes can now be generated solely from nanopore data, reducing turnaround time from ~39 h with hybrid strategies to ~14 h55.
Identification and quantification of ARGs through nanopore long-read metagenomices
In recent years, the spread of environmental ARGs has become increasingly recognized as a significant environmental pollutant56. The presence of ARGs in various ecosystems, including animal and human gut microbiota, soil, water, and wastewater, poses a serious risk to public health, as it can contribute to the spread of antimicrobial resistance in both human, animal and environmental populations56. Therefore, accurate identification and quantification of ARGs in gut and environmental samples are crucial for assessing their potential risk to human health and the environment57. Traditional methods for detecting and quantifying ARGs from uncultured samples predominantly rely on qPCR58,59,60. While qPCR is highly specific and widely used, it requires considerable time and labor for sample preparation, and it also involves high costs for reagents and consumables58. Furthermore, qPCR methods are typically designed to target known ARGs, limiting their ability to identify unknown ARGs.
With the rapid development of technologies, metagenomic sequencing has emerged as a more comprehensive and efficient method for detecting ARGs from complex samples61,62. Metagenomics, particularly through shotgun sequencing, allows for the simultaneous analysis of all microbial DNA present in a sample, providing a broader landscape of the microbial community and its associated resistance profiles63,64. However, it is difficult for short-read metagenomic sequencing to resolve the genetic structure and host information of ARGs effectively65. In addition, short reads may fail to provide sufficient context for understanding how ARGs are integrated into larger genomic contexts or how they are associated with mobile genetic elements, which are critical for the horizontal transfer of ARGs.
In contrast, nanopore long-read metagenomics offers significant advantages over short-read sequencing. Nanopore sequencing generates long reads, with an N50 typically ranging from 3 to 8 kb, depending on the sample quality and sequencing conditions26,65,66,67. These long sequencing reads enable more comprehensive genome assembly and provide detailed resolution of complex genetic structures (Fig. 2B). With nanopore sequencing, researchers can directly observe the relationship between ARGs and other functional genes, such as those involved in gene horizontal transfer or virulence. This capability is crucial for understanding how ARGs are linked to other microbial function genes and for tracking the genetic context in which they exist65. Furthermore, long reads enable more accurate prediction of the genetic location of ARGs within the genome, as well as the identification of their host organisms. This ability to predict ARGs to specific host genomes or plasmids is a significant advantage over short-read sequencing, which often struggles to resolve such complex genomic structures. By identifying the host organisms and genetic contexts of ARGs, nanopore sequencing can provide insights into how ARGs is being transferred between species, such as horizontal gene transfer mediated mobile genetic elements like plasmids, transposons, or integrons66,68. Understanding the host information of ARGs is vital for evaluating their potential to spread across microbial communities or to spread to pathogenic species, which can significantly impact public health.
For example, urban wastewater treatment plants (WWTPs) represent complex ecosystems and serve as critical reservoirs for the exchange of ARGs between humans and the environment. Accurate identification and quantification of ARGs in WWTPs are therefore essential for monitoring their epidemiological prevalence and potential risks to public health. Recent studies have demonstrated that nanopore long-read metagenomic sequencing provides unique advantages for this purpose. By leveraging long-reads, researchers have been able to directly link ARGs to mobile genetic elements, thereby revealing key drivers of ARG dissemination without the need for cultivation65,66,69. Moreover, nanopore metagenomic sequencing enables detailed tracking of ARG dynamics across different treatment stages, for example, from influent sewage to activated sludge, which highlighting the role of mobile elements in mediating these shifts66. Together, these capabilities underscore the power of nanopore long-read metagenomics in elucidating the genetic context and epidemiological dynamics of ARGs in wastewater environments.
Rapid identification of ARGs through nanopore long-read targeted sequencing
Detecting low-abundance ARGs is crucial for the effective treatment of bacterial infections, particularly in complex cases such as lower respiratory tract infections44. In these scenarios, even subtle resistance mechanisms can significantly impact pathogen virulence and lead to treatment failure. Traditional detection methods, such as shotgun metagenomic sequencing and cultivation, often struggle to identify these low-abundance ARGs, as they rely heavily on a high concentration of target DNA. This limitation can result in missed detection of critical resistance determinants, which in turn contributes to delays in treatment and ineffective antimicrobial choices. Nanopore sequencing offers a promising solution to this challenge, particularly through innovative targeted enrichment strategies such as CRISPR/Cas9-based enrichment and adaptive sampling (Fig. 2C)70,71. These techniques have markedly enhanced both the sensitivity and efficiency of ARG detection, even for low-abundance genes.
CRISPR/Cas9-based enrichment works by utilizing programmable guide RNAs to selectively amplify ARGs of interest from complex genomic samples72. This approach enables the precise capture and sequencing of ARGs, even from samples with a low abundance of ARGs, thereby improving detection sensitivity and accuracy72. For example, a method based on nanopore sequencing combined with CRISPR/Cas9 enrichment was demonstrated to identify ARGs in respiratory samples within 10 minutes of real-time sequencing71. This rapid turnaround time is especially important in clinical settings, where time-sensitive decisions need to be made regarding the treatment of infections. When compared to the traditional Illumina-based Cas9 targeting, nanopore sequencing not only enhances the detection of low-abundance ARGs but also reduces the overall turnaround time for ARGs identification to less than 6 h. This reduction in time is critical for informing prompt clinical interventions, particularly for infections caused by highly resistant pathogens.
Moreover, by designing specific guide RNAs, Cas9-based targeted enrichment can also facilitate the identification of the genetic contexts of ARGs, a key factor in understanding how ARGs spread. In particular, nanopore long sequencing reads enables the capture of genetic contexts that are often missed by short-read sequencing methods. A notable application of this approach is Context-Seq, a method that integrates Cas9-targeted enrichment with nanopore sequencing to identify the genetic contexts of key ARGs such as blaTEM and blaCTX-M. These genes, associated with β-lactam resistance, were captured along with their surrounding genomic regions, ranging from 1489 to 23,339 base pairs73. This genomic context information provides valuable insights into the genetic mechanisms of resistance transfer, as many of the captured reads contained genes encoding mobile genetic elements, which are critical in the horizontal transfer of ARGs between bacteria.
Beyond the Cas9-based enrichment methods, nanopore adaptive sampling technique has also shown great feasibility in rapid antimicrobial resistance diagnostics, especially in samples with low prevalence of ARGs74,75. This technique works by selectively removing host DNA during sequencing, effectively enriching microbial DNA and thus enabling more sensitive detection of ARGs. By significantly increasing the yield of microbial DNA, adaptive sampling enhances the detection of ARGs, even in challenging samples where ARGs are present at low levels. Notably, nanopore adaptive sampling can reduce the turnaround time for bacterial pathogen and ARG detection to under 4 h, making it an invaluable tool for clinical diagnostics74. This approach has been successfully used in phenotypic tests for antimicrobial resistance in positive blood cultures. Meanwhile, the accuracy of antimicrobial resistance predictions using this approach showed a categorical agreement rate exceeding 90% for monomicrobial infections, demonstrating its high reliability and potential for use in routine clinical practice70.
Although nanopore targeted sequencing offers significant advantages in terms of turnaround time and detection sensitivity, it also has several limitations that hinder its broader application. Both Cas9-based enrichment and adaptive sampling significantly result sequencing throughput reducing compared to standard nanopore sequencing, leading to higher costs for these methods relative to others. Additionally, the efficiency of Cas9-based enrichment is highly dependent on the design of the guide RNAs. Developing ideal guide RNAs that can both detect a wide range of ARGs and accurately capture their genetic context remains a critical challenge that needs to be addressed.
Structural diversity and dynamic analysis of ARGs and MDR regions
Resolution of genetic structural characteristics of critical ARGs
The emergence of mobile carbapenem resistance genes blaNDM, colistin resistance genes mcr and tigecycline resistance genes tet(X) pose a significant threat to human health76,77,78,79. These genes are often located on plasmids or other mobile genetic elements that facilitate their horizontal transfer between bacteria, thus greatly impairing the usefulness of critical antimicrobials (carbapenemase, colistin and tigecycline) as a clinical last line of defense. Understanding the detailed structural characteristics of these mobile ARGs is essential for elucidating how they spread within microbial populations and across ecosystems. Traditional short-read sequencing technologies often fall short in fully resolving these complex mobile genetic regions due to their inability to assemble large repeat sequences or capture the full context of ARGs. As a result, nanopore long-read sequencing usually was used for the resolving the genetic structural features of the three types of ARGs (Fig. 3A).
A The application of nanopore sequencing in deciphering genetic structural characteristics of critical ARGs. The detailed genetic basis of blaNDM, mcr, and tet(X) has been well decoded with nanopore sequencing. B Single nanopore sequencing reads enable the resolution of genetic diversity and heterogeneity within tandem repeat regions. The ultra-long reads generated by nanopore sequencing can span large MDR repeat regions, which is crucial for studying the genetic heterogeneity of tandem repeats and bacterial antimicrobial heteroresistance. C Nanopore sequencing facilitates the resolution of structural features and the dynamic evolution of MDR plasmids. Nanopore sequencing data can resolve the structural changes of MDR plasmids that occur during conjugation or bacterial division.
In general, blaNDM, mcr, and tet(X) were commonly carried by plasmids. In recent years, many types of blaNDM-, mcr-, and tet(X)-bearing plasmids have been resolved by nanopore long-read sequencing. The blaNDM genes usually located on IncX3, IncC, and IncFII plasmids80,81. The mcr genes usually located on IncX4, IncI2, and IncHI2 plasmids82,83. The tet(X) genes usually located on IncF-hybrid plasmids84. The ability of nanopore sequencing to capture large, contiguous sequences has allowed for a more comprehensive understanding of the mobility and integration mechanisms of these resistance genes. In addition to plasmids, blaDNM have also founded to be involved in some complex genetic structures, such as chromosomally integrated plasmids, large transposons and integrative and conjugative elements (ICEs)85,86,87,88. Similar, many mcr-bearing plasmids have also been found to integrate into bacterial chromosomes using nanopore long-read sequencing, especially in Salmonella species89,90,91. Furthermore, transposon-mediated integration of mcr genes into chromosomes, such as the chromosomal insertion of Tn6330, is frequently observed and can be accurately characterized with nanopore sequencing92,93. Apart from plasmids, the genetic location of tet(X) is primarily found within ICEs, with the tet(X6) variant often carried by these mobile elements84,94.
With the widespread use of antimicrobials, an increasing number of super-bugs have emerged. The generation of these super-bugs is primarily driven by the accumulation of critical ARGs within the same strain95,96,97. Although conventional PCR methods or whole-genome sequencing can identify multiple ARGs in the same strain, the precise genetic location of these ARGs cannot be determined. This limits our ability to understand the reasons behind the clustering of different critical ARGs and to investigate the genomic mechanisms underlying the formation of MDR bacteria. Moreover, clearly decoding the co-existence characteristics of critical ARGs at the genomic level is crucial for studying their horizontal transmission patterns. Currently, many extensively drug-resistant strains have been analyzed using nanopore sequencing. For example, co-existing blaNDM, mcr, and tet(X) genes have been resolved in detail, including reports of mcr-1, tet(X4), and blaNDM-5 occurring together in the same Escherichia coli strain95,96. In this case, nanopore sequencing revealed that mcr-1 was carried by the chromosomal transposon Tn6330, while tet(X4) and blaNDM-5 were located on IncFII and IncX3 plasmids, respectively96. These findings highlight the power of nanopore sequencing in resolving the genetic context of multiple ARGs within single strains, thereby providing critical insights into their horizontal transfer and the emergence of pan-drug resistance.
Tandem amplification of ARGs-bearing genetic structures
In response to environmental pressures, bacteria often develop various adaptive strategies to survive these stresses. A well-known example of this is antimicrobial heteroresistance, where a bacterial population, initially sensitive to antimicrobials, evolves into a mix of subclones with varying degrees of resistance. This phenomenon typically results from the amplification of ARGs within certain subpopulations. Recently, with the help of nanopore long-read sequencing technology, it has become evident that the majority of antimicrobial heteroresistance is linked to the tandem amplification of ARGs within bacterial genomes98. Notably, this type of heteroresistance is commonly observed during clinical antimicrobial treatments99,100,101. For example, tandem amplification of aphA1 in Acinetobacter baumannii can lead to tobramycin resistance, ultimately resulting in treatment failure102. Clinical Escherichia coli could develop piperacillin/tazobactam resistance by IS26-mediated amplification of blaTEM-1B103.
To date, numerous tandem amplification structures have been resolved by nanopore long-read sequencing (Fig. 3B). These amplifications frequently occur in the MDR regions of various plasmids in Enterobacteriaceae. Typically, such tandem repeat structures consist of multiple translocatable units (TUs), each containing a mobile genetic element together with one or more ARGs that drive resistance amplification104,105. A recent identified tigecycline resistance gene, tet(X), has also been reported in tandem repeat form, most commonly as a simple tet(X)-ISCR2 unit77,84,94. Nanopore sequencing can generate ultra-long reads, which are particularly powerful for resolving complex tandem repeats. However, the production of ultra-long reads is often limited by the size of input DNA fragments. In practice, DNA extracted with standard bacterial genomic kits usually yields fragments of ~40–50 kb, which constrains the sequencing of ultra-long reads. In contrast, plasmid extraction methods (such as QIAGEN Plasmid Midi Kit) can recover much larger molecules, often exceeding 100 kb, thereby facilitating ultra-long read generation and enabling resolution of complex multidrug-resistant plasmid structures. Using these approaches, long tandem repeat regions spanning tens of kb in plasmids can be fully deciphered. For example, the tmexCD1-toprJ1 gene cluster is frequently embedded in long tandem repeats of several tens of kb, containing multiple mobile elements as well as additional ARGs106. Moreover, ultra-long reads have provided direct evidence for the involvement of insertion sequences in the formation of tandem repeat regions84,106. Beyond Enterobacteriaceae, tandem repeats have also been identified in Gram-positive bacteria, including poxtA-bearing tandem repeats in Enterococcus faecalis and vanM tandem amplifications in Vancomycin-variable enterococci107,108.
Structural features and dynamic evolution of MDR mobile genetic elements
The formation of MDR regions and fusion plasmids is a complex and dynamic process driven by antimicrobial selection pressures, horizontal gene transfer, and the integration of mobile genetic elements109. These structures present a significant challenge in combating antimicrobial resistance. MDR regions often consist of clusters of ARGs, insertion sequences, integrases, and are located on bacterial chromosomes or mobile genetic elements12. These mobile elements facilitate the rapid horizontal transfer of ARGs between bacterial species12,110,111. Understanding the structures of these complex mobile elements and fusion and recombination plasmids that harbor MDR regions is crucial for investigating the evolution and transmission dynamics of ARGs.
Large mobile elements, such as Salmonella genomic island (SGI), Proteus genomic island (PGI), and ICEs, are typically tens of kb in size and often contain multiple ARGs, as well as other virulence or mobility factors12. Because of frequent recombination mediated by insertion sequences, integrases, and recombinases, these elements often display highly complex architectures that are difficult to resolve with short-read sequencing alone. Nanopore sequencing has greatly improved the resolution of such structures and enabled comprehensive characterization of their genetic composition. In recent years, this approach has facilitated the identification of numerous SGIs, PGIs, and ICEs across diverse bacterial species112,113,114,115,116. These studies have provided a clearer landscape of the genetic features of these regions and their role in the spread of ARGs. As these complex structures are increasingly understood, their contribution to the global dissemination of antimicrobial resistance becomes more apparent. Clinically critical ARGs, such as those encoding carbapenemases and tigecycline resistance, have also been shown to be frequently transmitted through these complex MDR regions86,114,115,117.
Fusion and recombination plasmids are larger and more complex than other large mobile elements, and they often undergo structural changes during bacterial division or conjugative transfer between different bacteria118,119. Before the advent of long-read sequencing technology, plasmid fusion and recombination processes could only be observed through techniques like S1-PFGE or Southern blotting120. However, these methods are limited in their ability to analyze the detailed structure and mechanisms of plasmid fusion and recombination. With the widespread application of nanopore sequencing technology in bacterial genome analysis, it is now easier to resolve fusion and recombination plasmids. This technology also allows us to observe the dynamic structural changes of plasmids during conjugative transfer between bacteria (Fig. 3C). For example, the fusion or recombination of a plasmid carrying carbapenemase genes with a plasmid carrying virulence genes during conjugation can result in the emergence of carbapenem-resistant and hypervirulent Klebsiella pneumoniae121. Furthermore, the dynamic changes in fusion and recombination plasmids accelerate the aggregation and co-transfer of ARGs, contributing to the spread of MDR bacteria40,120,122,123.
Emerging novel application of nanopore sequencing in AMR
Currently, the primary application of nanopore sequencing in bacterial antimicrobial resistance research is DNA sequencing. However, other capabilities of nanopore sequencing, such as direct RNA sequencing and methylation modification detection, have been scarcely applied to the study of bacterial resistance124,125,126. Direct RNA sequencing offers the advantage of analyzing RNA molecules without the need for cDNA synthesis, providing a more accurate and comprehensive view of bacterial gene expression in response to antimicrobial pressure. This could lead to valuable insights into how ARGs are regulated and expressed in real time. Additionally, the ability of nanopore sequencing in detecting DNA methylation modifications directly holds significant potential for understanding epigenetic changes that contribute to antimicrobial resistance. Methylation could play a role in the regulation of resistance genes or the stabilization of mobile genetic elements that carry them. As these capabilities are further developed and integrated into antimicrobial resistance studies, they are likely to offer new avenues for understanding the molecular mechanisms underlying bacterial resistance, enabling more precise diagnostics and novel therapeutic strategies.
Conclusions and future perspectives
Deciphering the genetic structure of MDR bacteria is crucial for understanding the evolution and spread of antimicrobial resistance. Nanopore long-read sequencing technology is revolutionizing bacterial resistance research with two unique advantages: real-time sequencing and ultra-long sequencing reads. Its ability of rapid turnaround time enables quick diagnostics and real-time monitoring of bacterial antimicrobial resistance, particularly in clinical settings. Additionally, its ability to generate ultra-long reads allows for the detailed characterization of complex genetic regions and tracking the dynamic evolution of MDR genetic structures, which were previously difficult to resolve with short-read sequencing technologies. However, the high error rate of sequencing reads and sequencing costs per sample currently limit its broader application. As the further development of nanopore sequencing technology and commercialization in multiple countries, especially in the improvements of both cost reduction and sequencing accuracy, it is poised to play an increasingly pivotal role in addressing the growing global threat of antimicrobial resistance spread.
Data availability
No datasets were generated or analysed during the current study.
References
Wright, G. D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 5, 175–186 (2007).
Dantas, G., Sommer, M. O., Oluwasegun, R. D. & Church, G. M. Bacteria subsisting on antibiotics. Science 320, 100–103 (2008).
Van Boeckel, T. P. et al. Global antibiotic consumption 2000 to 2010: an analysis of national pharmaceutical sales data. Lancet Infect. Dis. 14, 742–750 (2014).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: the WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2018).
Hao, R., Zhao, R., Qiu, S., Wang, L. & Song, H. Antibiotics crisis in China. Science 348, 1100–1101 (2015).
Kariuki, S. Global burden of antimicrobial resistance and forecasts to 2050. Lancet 404, 1172–1173 (2024).
Piperaki, E. T., Tzouvelekis, L. S., Miriagou, V. & Daikos, G. L. Carbapenem-resistant Acinetobacter baumannii: in pursuit of an effective treatment. Clin. Microbiol. Infect. 25, 951–957 (2019).
Ruef, M. et al. Carriage of third-generation cephalosporin-resistant and carbapenem-resistant Enterobacterales among children in sub-Saharan Africa: a systematic review and meta-analysis. EClinicalMedicine 70, 102508 (2024).
Dheda, K. et al. Multidrug-resistant tuberculosis. Nat. Rev. Dis. Prim. 10, 22 (2024).
Jesudason, T. WHO publishes updated list of bacterial priority pathogens. Lancet Microbe 5, 100940 (2024).
Perry, J. A., Westman, E. L. & Wright, G. D. The antibiotic resistome: what’s new? Curr. Opin. Microbiol 21, 45–50 (2014).
Partridge, S. R., Kwong, S. M., Firth, N. & Jensen, S. O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. https://doi.org/10.1128/CMR.00088-17 (2018).
Johnson, A. P. & Woodford, N. Global spread of antibiotic resistance: the example of New Delhi metallo-beta-lactamase (NDM)-mediated carbapenem resistance. J. Med. Microbiol. 62, 499–513 (2013).
Robinson, E. R., Walker, T. M. & Pallen, M. J. Genomics and outbreak investigation: from sequence to consequence. Genome Med. 5, 36. https://doi.org/10.1186/gm440 (2013).
Barrick, J. E. et al. Identifying structural variation in haploid microbial genomes from short-read resequencing data using breseq. BMC Genomics 15 (2014).
Liu, W. et al. Transmission of antimicrobial resistance genes from the environment to human gut is more pronounced in colorectal cancer patients than in healthy subjects. iMeta https://doi.org/10.1002/imt2.70008 (2025).
Laszlo, A. H. et al. Decoding long nanopore sequencing reads of natural DNA. Nat. Biotechnol. 32, 829–833 (2014).
Ashton, P. M. et al. MinION nanopore sequencing identifies the position and structure of a bacterial antibiotic resistance island. Nat. Biotechnol. 33, 296–300 (2015).
Loman, N. J., Quick, J. & Simpson, J. T. A complete bacterial genome assembled de novo using only nanopore sequencing data. Nat. Methods 12, 733–735 (2015).
Gao, Y. et al. The Microbiome Protocols eBook initiative: building a bridge to microbiome research. Imeta 3, e182 (2024).
Deamer, D., Akeson, M. & Branton, D. Three decades of nanopore sequencing. Nat. Biotechnol. 34, 518–524 (2016).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl. Acad. Sci. USA 93, 13770–13773 (1996).
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E. & Deamer, D. W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid, and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 (1999).
Lieberman, K. R. et al. Processive replication of single DNA molecules in a nanopore catalyzed by phi29 DNA polymerase. J. Am. Chem. Soc. 132, 17961–17972 (2010).
Peng, K. et al. QitanTech nanopore long-read sequencing enables rapid resolution of complete genomes of multi-drug resistant pathogens. Front. Microbiol. 13, 778659 (2022).
Peng, K. et al. Long-read metagenomic sequencing reveals that high-copy small plasmids shape the highly prevalent antibiotic resistance genes in animal fecal microbiome. Sci. Total Environ. 893, 164585 (2023).
Liu, C., Guo, J., Lu, M., Shen, N. & Du, P. Dissemination of the mobilised RND efflux pump gene cluster tmexCD-toprJ among Klebsiella pneumoniae. Lancet Microbe 4, e135 (2023).
Liang, H. et al. Efficiently constructing complete genomes with CycloneSEQ to fill gaps in bacterial draft assemblies. GigaByte 2025, gigabyte154 (2025).
Jain, M. et al. Improved data analysis for the MinION nanopore sequencer. Nat. Methods 12, 351–356 (2015).
Jain, M. et al. MinION Analysis and Reference Consortium: Phase 2 data release and analysis of R9.0 chemistry. F1000Res 6, 760 (2017).
Cretu Stancu, M. et al. Mapping and phasing of structural variation in patient genomes using nanopore sequencing. Nat. Commun. 8, 1326 (2017).
Chen, Z. et al. Genomic analyses of multidrug-resistant Salmonella Indiana, Typhimurium, and Enteritidis isolates using MinION and MiSeq sequencing technologies. PLoS One 15, e0235641 (2020).
Karst, S. M. et al. High-accuracy long-read amplicon sequences using unique molecular identifiers with Nanopore or PacBio sequencing. Nat. Methods 18, 165–169 (2021).
Sereika, M. et al. Oxford Nanopore R10.4 long-read sequencing enables the generation of near-finished bacterial genomes from pure cultures and metagenomes without short-read or reference polishing. Nat. Methods 19, 823–826 (2022).
Sheka, D., Alabi, N. & Gordon, P. M. K. Oxford nanopore sequencing in clinical microbiology and infection diagnostics. Brief. Bioinform 22, bbaa403 (2021).
Taxt, A. M., Avershina, E., Frye, S. A., Naseer, U. & Ahmad, R. Rapid identification of pathogens, antibiotic resistance genes and plasmids in blood cultures by nanopore sequencing. Sci. Rep. 10, 7622 (2020).
Sauerborn, E. et al. Detection of hidden antibiotic resistance through real-time genomics. Nat. Commun. 15, 5494 (2024).
Ciuffreda, L., Rodriguez-Perez, H. & Flores, C. Nanopore sequencing and its application to the study of microbial communities. Comput Struct. Biotechnol. J. 19, 1497–1511 (2021).
Yang, R., Tang, J., Niu, J., Hou, B. & Zhang, L. Dissemination mechanisms of unique antibiotic resistance genes from flowback water to soil revealed by combined Illumina and Nanopore sequencing. Water Res. 273, 123030 (2024).
Peter, S. et al. Tracking of antibiotic resistance transfer and rapid plasmid evolution in a hospital setting by nanopore sequencing. mSphere 5, e00525–20 (2020).
Bates, M., Polepole, P., Kapata, N., Loose, M. & O’Grady, J. Application of highly portable MinION nanopore sequencing technology for the monitoring of nosocomial tuberculosis infection. Int. J. Mycobacteriol. 5, S24 (2016).
Nakamura, A. & Komatsu, M. Performance evaluation of whole genome metagenomics sequencing with the MinION nanopore sequencer: Microbial community analysis and antimicrobial resistance gene detection. J. Microbiol. Methods 206, 106688 (2023).
Ko, K. K. K., Chng, K. R. & Nagarajan, N. Metagenomics-enabled microbial surveillance. Nat. Microbiol. 7, 486–496 (2022).
Charalampous, T. et al. Nanopore metagenomics enables rapid clinical diagnosis of bacterial lower respiratory infection. Nat. Biotechnol. 37, 783–792 (2019).
Udaondo, Z. et al. Comparative analysis of PacBio and Oxford nanopore sequencing technologies for transcriptomic landscape identification of Penaeus monodon. Life 11, 862 (2021).
Payne, A., Holmes, N., Rakyan, V. & Loose, M. BulkVis: a graphical viewer for Oxford nanopore bulk FAST5 files. Bioinformatics 35, 2193–2198 (2019).
Eisenhofer, R. et al. A comparison of short-read, HiFi long-read, and hybrid strategies for genome-resolved metagenomics. Microbiol. Spectr. https://doi.org/10.1128/spectrum.03590-23 (2024).
Ni, Y., Liu, X., Simeneh, Z. M., Yang, M. & Li, R. Benchmarking of Nanopore R10.4 and R9.4.1 flow cells in single-cell whole-genome amplification and whole-genome shotgun sequencing. Comput. Struct. Biotechnol. J. 21, 2352–2364 (2023).
Boolchandani, M., D’Souza, A. W. & Dantas, G. Sequencing-based methods and resources to study antimicrobial resistance. Nat. Rev. Genet. 20, 356–370 (2019).
Wick, R. R., Judd, L. M., Gorrie, C. L. & Holt, K. E. Unicycler: resolving bacterial genome assemblies from short and long sequencing reads. PLoS Comput. Biol. 13, e1005595 (2017).
De Maio, N. et al. Comparison of long-read sequencing technologies in the hybrid assembly of complex bacterial genomes. Micro Genom. 5, e000294 (2019).
Chen, Z., Erickson, D. L. & Meng, J. Benchmarking hybrid assembly approaches for genomic analyses of bacterial pathogens using Illumina and Oxford Nanopore sequencing. BMC Genomics 21, 631 (2020).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Zhao, W. et al. Oxford nanopore long-read sequencing enables the generation of complete bacterial and plasmid genomes without short-read sequencing. Front. Microbiol. 14, 1179966 (2023).
Larsson, D. G. J. & Flach, C. F. Antibiotic resistance in the environment. Nat. Rev. Microbiol. 20, 257–269 (2022).
Peng, K. et al. Benchmarking of analysis tools and pipeline development for nanopore long-read metagenomics. Sci. Bull. 70, 1591–1595 (2025).
Abramova, A., Berendonk, T. U. & Bengtsson-Palme, J. A global baseline for qPCR-determined antimicrobial resistance gene prevalence across environments. Environ. Int. 178, 108084 (2023).
Tolosi, R., Carraro, L., Laconi, A. & Piccirillo, A. Optimization of five qPCR protocols toward the detection and the quantification of antimicrobial resistance genes in environmental samples. MethodsX 8, 101488 (2021).
Waseem, H. et al. Contributions and challenges of high throughput qPCR for determining antimicrobial resistance in the environment: a critical review. Molecules 24, 163 (2019).
Hendriksen, R. S. et al. Global monitoring of antimicrobial resistance based on metagenomics analyses of urban sewage. Nat. Commun. 10, 1124 (2019).
Yu, Q. et al. Metagenomics reveals the response of antibiotic resistance genes to elevated temperature in the Yellow River. Sci. Total Environ. 859, 160324 (2023).
He, J., Zhang, N., Shen, X., Muhammad, A. & Shao, Y. Deciphering environmental resistome and mobilome risks on the stone monument: a reservoir of antimicrobial resistance genes. Sci. Total Environ. 838, 156443 (2022).
Baquero, F. Metagenomic epidemiology: a public health need for the control of antimicrobial resistance. Clin. Microbiol. Infect. 18, 67–73 (2012).
Che, Y. et al. Mobile antibiotic resistome in wastewater treatment plants revealed by Nanopore metagenomic sequencing. Microbiome 7, 44 (2019).
Dai, D. et al. Long-read metagenomic sequencing reveals shifts in associations of antibiotic resistance genes with mobile genetic elements from sewage to activated sludge. Microbiome 10, 20 (2022).
Li, L. et al. Short- and long-read metagenomics insight into the genetic contexts and hosts of mobile antibiotic resistome in Chinese swine farms. Sci. Total Environ. 827, 154352 (2022).
Yang, T. et al. Mobility, bacterial hosts, and risks of antibiotic resistome in submicron bioaerosols from a full-scale wastewater treatment plant. J. Environ. Manag. 351, 119771 (2024).
Wu, Z. et al. Nanopore-based long-read metagenomics uncover the resistome intrusion by antibiotic resistant bacteria from treated wastewater in receiving water body. Water Res. 226, 119282 (2022).
Liu, P. Y. et al. Comprehensive pathogen identification and antimicrobial resistance prediction from positive blood cultures using nanopore sequencing technology. Genome Med. 16, 141 (2024).
Serpa, P. H. et al. Metagenomic prediction of antimicrobial resistance in critically ill patients with lower respiratory tract infections. Genome Med. 14, 74 (2022).
Gilpatrick, T. et al. Targeted nanopore sequencing with Cas9-guided adapter ligation. Nat. Biotechnol. 38, 433–438 (2020).
Fuhrmeister, E. R. et al. Context-Seq: CRISPR-Cas9 targeted nanopore sequencing for transmission dynamics of antimicrobial resistance. Nat Commun 16, 5898 (2025).
Cheng, H. et al. A rapid bacterial pathogen and antimicrobial resistance diagnosis workflow using Oxford nanopore adaptive sequencing method. Brief. Bioinform 23, bbac453 (2022).
Wrenn, D. C. & Drown, D. M. Nanopore adaptive sampling enriches for antimicrobial resistance genes in microbial communities. GigaByte 2023, gigabyte103 (2023).
Liu, Y. Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
He, T. et al. Emergence of plasmid-mediated high-level tigecycline resistance genes in animals and humans. Nat. Microbiol. 4, 1450–1456 (2019).
Nordmann, P., Dortet, L. & Poirel, L. Carbapenem resistance in Enterobacteriaceae: here is the storm!. Trends Mol. Med. 18, 263–272 (2012).
Haijie, Z. et al. A novel ABC family protein participates in the regulation of fitness cost caused by tet(X4)-bearing plasmids in Escherichia coli. Fundam. Res. https://doi.org/10.1016/j.fmre.2024.03.020 (2024).
Li, X. et al. Dissemination of blaNDM-5 gene via an IncX3-type plasmid among non-clonal Escherichia coli in China. Antimicrob. Resist. Infect. Control 7, 59 (2018).
Wu, W. et al. NDM Metallo-beta-lactamases and their bacterial producers in health care settings. Clin. Microbiol. Rev. 32, e00115–e00118 (2019).
Naha, S. et al. Carriage and within-host diversity of mcr-1.1-harbouring Escherichia coli from pregnant mothers: inter- and intra-mother transmission dynamics of mcr-1.1. Emerg. Microbes Infect. 12, 2278899 (2023).
Liu, J. H. et al. Plasmid-mediated colistin-resistance genes: mcr. Trends Microbiol. 32, 365–378 (2024).
Li, R. et al. Deciphering the structural diversity and classification of the mobile tigecycline resistance gene tet(X)-bearing plasmidome among bacteria. mSystems 5, e00134–20 (2020).
Reynolds, M. E. et al. Occurrence and characterization of Escherichia coli ST410 co-harbouring blaNDM-5, blaCMY-42 and blaTEM-190 in a dog from the UK. J. Antimicrob. Chemother. 74, 1207–1211 (2019).
He, J. et al. A Novel SXT/R391 integrative and conjugative element carries two copies of the bla(NDM-1) gene in Proteus mirabilis. mSphere 6, e0058821 (2021).
Xiang, R. & Li, M. Identification of Tn6835 and a Novel Genomic Island, MMGI-1, in a pan-resistant Morganella morganii strain. Antimicrob. Agents Chemother. 65, e02524–20 (2021).
Tanabe, M. et al. A novel blaNDM-1-carrying multidrug-resistant genomic island GIMmSU8481 in a faecal Morganella morganii subsp. sibonii isolate from a patient. J. Glob. Antimicrob. Resist. 35, 322–324 (2023).
Yang, C. et al. Genetic and drug susceptibility profiles of mcr-1-bearing foodborne Salmonella strains collected in Shenzhen, China during the period 2014-2017. Microbiol. Res. 265, 127211 (2022).
Chang, M. X. et al. Formation, transmission, and dynamic evolution of a multidrug-resistant chromosomally integrated plasmid in Salmonella Spp. Front. Microbiol. 13, 846954 (2022).
Mei, C. Y. et al. Chromosomally and plasmid-located mcr in Salmonella from animals and food products in China. Microbiol. Spectr. 10, e0277322 (2022).
Goodman, R. N., Tansirichaiya, S., Brouwer, M. S. M. & Roberts, A. P. Intracellular transposition of mobile genetic elements associated with the colistin resistance gene mcr-1. Microbiol. Spectr. 11, e0327822 (2023).
Treilles, M., Chatre, P., Drapeau, A., Madec, J. Y. & Haenni, M. Spread of the mcr-1 colistin-resistance gene in Escherichia coli through plasmid transmission and chromosomal transposition in French goats. Front. Microbiol. 13, 1023403 (2022).
Li, R., Peng, K., Li, Y., Liu, Y. & Wang, Z. Exploring tet(X)-bearing tigecycline-resistant bacteria of swine farming environments. Sci. Total Environ. 733, 139306 (2020).
Shafiq, M. et al. Coexistence of bla (NDM-5) and tet(X4) in international high-risk Escherichia coli clone ST648 of human origin in China. Front Microbiol 13, 1031688 (2022).
Lu, X. Y. et al. Coexistence of tet(X4), mcr-1, and blaNDM-5 in ST6775 Escherichia coli isolates of animal origin in China. Microbiol. Spectr. 10, e0019622 (2022).
Xu, Y., Liu, L., Zhang, H. & Feng, Y. Co-production of Tet(X) and MCR-1, two resistance enzymes by a single plasmid. Environ. Microbiol. 23, 7445–7464 (2021).
Nicoloff, H., Hjort, K., Levin, B. R. & Andersson, D. I. The high prevalence of antibiotic heteroresistance in pathogenic bacteria is mainly caused by gene amplification. Nat. Microbiol. 4, 504–514 (2019).
Nimmo, C. et al. Dynamics of within-host Mycobacterium tuberculosis diversity and heteroresistance during treatment. EBioMedicine 55, 102747 (2020).
Han, X. et al. Genome sequencing unveils bla(KPC-2)-harboring plasmids as drivers of enhanced resistance and virulence in nosocomial Klebsiella pneumoniae. mSystems 9, e0092423 (2024).
Liu, C. et al. Dynamic within-host cefiderocol heteroresistance caused by bla(SHV-12) amplification in pandrug-resistant and hypervirulent Klebsiella pneumoniae sequence type 11. Drug Resist. Updat. 73, 101038 (2024).
Harmer, C. J., Lebreton, F., Stam, J., McGann, P. T. & Hall, R. M. Mechanisms of IS26-mediated amplification of the aphA1 gene leading to tobramycin resistance in an Acinetobacter baumannii isolate. Microbiol. Spectr. 10, e0228722 (2022).
Hubbard, A. T. M. et al. Piperacillin/tazobactam resistance in a clinical isolate of Escherichia coli due to IS26-mediated amplification of bla(TEM-1B). Nat. Commun. 11, 4915 (2020).
Girgis, H. S. et al. Single-molecule nanopore sequencing reveals extreme target copy number heterogeneity in arylomycin-resistant mutants. Proc. Natl. Acad. Sci. USA 118, e2021958118 (2021).
Sandegren, L. & Andersson, D. I. Bacterial gene amplification: implications for the evolution of antibiotic resistance. Nat. Rev. Microbiol. 7, 578–588 (2009).
Peng, K. et al. Plasmids shape the current prevalence of tmexCD1-toprJ1 among Klebsiella pneumoniae in food production chains. mSystems 6, e0070221 (2021).
Sun, L. et al. Tandem amplification of the vanM gene cluster drives vancomycin resistance in vancomycin-variable enterococci. J. Antimicrob. Chemother. 75, 283–291 (2020).
Shan, X. et al. poxtA amplification and mutations in 23S rRNA confer enhanced linezolid resistance in Enterococcus faecalis. J. Antimicrob. Chemother. 79, 3199–3203 (2024).
Liu, Z., Tang, Y., He, M. & Xu, C. Molecular drivers of fusion plasmid: mechanistic insights and evolutionary implications. J. Antimicrob. Chemother. https://doi.org/10.1093/jac/dkaf309 (2025).
Durrant, M. G., Li, M. M., Siranosian, B. A., Montgomery, S. B. & Bhatt, A. S. A bioinformatic analysis of integrative mobile genetic elements highlights their role in bacterial adaptation. Cell Host Microbe 27, 140 (2020).
Lang, A. S., Buchan, A. & Burrus, V. Interactions and evolutionary relationships among bacterial mobile genetic elements. Nat. Rev. Microbiol. 23, 423–438 (2025).
Lei, C. W. et al. Identification of Proteus genomic island 2 variants in two clonal Proteus mirabilis isolates with coexistence of a novel genomic resistance island PmGRI1. J. Antimicrob. Chemother. 75, 2503–2507 (2020).
He, J. et al. Identification of a novel genomic resistance island PmGRI1-STP3 and an SXT/R391 integrative conjugative element in Proteus mirabilis of swine origin in China. J. Glob. Antimicrob. Resist. 25, 77–81 (2021).
Yang, L. et al. Nosocomial outbreak of carbapenemase-producing Proteus mirabilis with two novel Salmonella genomic island 1 variants carrying different bla (NDM-1) gene copies in China. Front. Microbiol. 12, 800938 (2021).
Gao, X. et al. A Salmonella genomic island 1 (SGI1) carries multiple copies of bla(NDM-1) in Vibrio fluvialis of retail razor clam origin. J. Glob. Antimicrob. Resist. 35, 190–192 (2023).
Intuy, R. et al. A novel variant in Salmonella genomic island 1 of multidrug-resistant Salmonella enterica serovar Kentucky ST198. Microbiol. Spectr. 12, e0399423 (2024).
Wang, Q. et al. Characterization of TMexCD3-TOprJ3, an RND-type efflux system conferring resistance to tigecycline in Proteus mirabilis, and its associated Integrative Conjugative Element. Antimicrob. Agents Chemother https://doi.org/10.1128/AAC.02712-20 (2021).
Gao, Y. et al. Dynamic evolution and transmission of a bla(NDM-1)-bearing fusion plasmid in a clinical Escherichia coli. Microbiol. Res. 275, 127450 (2023).
Jones, N. I., Harmer, C. J., Hamidian, M. & Hall, R. M. Evolution of Acinetobacter baumannii plasmids carrying the oxa58 carbapenemase resistance gene via plasmid fusion, IS26-mediated events and dif module shuffling. Plasmid 121, 102628 (2022).
Lu, X., Xiao, X., Liu, Y., Li, R. & Wang, Z. Emerging opportunity and destiny of mcr-1- and tet(X4)-coharboring plasmids in Escherichia coli. Microbiol. Spectr. 9, e0152021 (2021).
Yang, X., Dong, N., Chan, E. W., Zhang, R. & Chen, S. Carbapenem resistance-encoding and virulence-encoding conjugative plasmids in Klebsiella pneumoniae. Trends Microbiol. 29, 65–83 (2021).
Shi, Q. et al. Chromosomal integration and plasmid fusion occurring in ST20 carbapenem-resistant Klebsiella pneumoniae isolates coharboring bla(NDM-1) and bla(IMP-4) induce resistance transmission and fitness variation. Emerg. Microbes Infect. 13, 2339942 (2024).
Wang, X. et al. Co-transfer of mcr-8 with bla(NDM-1) or tmexCD1-toprJ1 by plasmid hybridisation. Int. J. Antimicrob. Agents 60, 106619 (2022).
Pust, M. M., Davenport, C. F., Wiehlmann, L. & Tummler, B. Direct RNA nanopore sequencing of Pseudomonas aeruginosa clone C transcriptomes. J. Bacteriol. 204, e0041821 (2022).
Jain, M., Abu-Shumays, R., Olsen, H. E. & Akeson, M. Advances in nanopore direct RNA sequencing. Nat. Methods 19, 1160–1164 (2022).
Hendra, C. et al. Detection of m6A from direct RNA sequencing using a multiple instance learning framework. Nat. Methods 19, 1590–1598 (2022).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2024YFC3406300), National Key Laboratory of Veterinary Public Health and Safety Open Project Fund (2024SKLVPHS04), the Natural Science Foundation of Jiangsu Province (no. BK20231524), the China Postdoctoral Science Foundation (2024M762745), Jiangsu Funding Program for Excellent Postdoctoral Talent (2025ZB846), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
R.L. and K.P. conceptualized this study, obtained resources, and wrote the main manuscript text. Z.W. edited the draft. C.L. and Q.W. contributed to the writing of the original draft. K.P. and X.X. prepared figures. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Peng, K., Li, C., Wang, Q. et al. The applications and advantages of nanopore sequencing in bacterial antimicrobial resistance surveillance and research. npj Antimicrob Resist 3, 87 (2025). https://doi.org/10.1038/s44259-025-00157-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-025-00157-5