Abstract
Early research has found that biofilms adhered to treatment filtration beds and the inside of drinking water pipes help promote antimicrobial resistance (AMR) in drinking water systems. This study investigates the impact of a sub-minimum inhibitory concentration of ciprofloxacin (10 µg L–1) on AMR and antibiotic resistance genes (ARGs) in multi-species biofilms formed on polyvinyl chloride pipe using a novel bench-scale water distribution reactor. The biofilm was grown from drinking water and contained a highly diverse composition consisting of, but not limited to, Dechloromonas, Pseudomonas, Pseudoxanthomonas, Acidovorax, and Allorhizobium-Neorhizobium-Pararhizobium-Rhizobium. It was found that when the biofilm was exposed to ciprofloxacin, the total cell counts significantly increased, suggesting a potential AMR response. IntI1 and sul1 showed a significant inverse correlation with several genera as well as with increased diversity. Asinibacterium, found at a lower abundance in the biofilms, had a significant positive correlation with intI1, indicating its potential role in ARG promotion and the enhanced risk of a less diverse biofilm. This study highlights the need for targeted control strategies against AMR development in the context of drinking water distribution systems, emphasizing the role of biofilm community composition and residual antibiotic exposure.
Similar content being viewed by others
Introduction
Currently, the global consumption rate of antibiotics is increasing while the rate of their research and development is decreasing1,2,3. With no new antibiotic classes developed since the 1980s, there are limited options when it comes to treating new and evolving bacterial infections1. As a result, antimicrobial resistance (AMR) is accelerated by the overuse of antibiotics4. AMR occurs when microorganisms, such as bacteria, are no longer responsive to antibiotics due to the promotion and/or development of an ability to defeat them. This exacerbates the severity of a bacterial infection, increasing the likelihood of death. Therefore, when antibiotics are prescribed, it is vital that they are taken as directed to ensure that their dosage is above the minimum inhibitory concentration (MIC). The MIC denotes the lowest concentration required to stop the growth of bacteria and thus prevent resistance from occurring5.
Unfortunately, the negative implications of increasing antibiotic consumption further extend into the environment. With the rise in global antibiotic usage, several classes of antibiotics have been detected in water systems, including oceans, lakes, rivers, reservoirs, groundwater, wastewater (WW), and drinking water (DW)6. This is in part owing to the presence of global surveillance systems such as the Global Antimicrobial Resistance and Use Surveillance System (GLASS) initiated by the World Health Organization in 20157. Antibiotics can infiltrate water bodies through multiple routes of entry, while costly and specialized removal treatment options are unavailable to most municipalities8,9. Of interest are the antibiotics that are being detected in DW, such as ciprofloxacin, which has been found to be persistent with concentrations reaching up to 0.492 µg L–110. When present at low, residual concentrations, researchers have argued that the antibiotics are at an acceptable average daily dose (ADD) and health risk quotient (HRQ) for human exposure/ingestion11,12. However, this claim falls short of considering the effect of antibiotics on biofilms coupled with other environmental factors.
When antibiotics are present in DW, they may exist at a sub-MIC or minimal selective concentration (MSC). A sub-MIC is a concentration range that is below the MIC that is nonlethal to bacteria, while the minimum selective concentration (MSC) is the lowest concentration range required for a selection pressure—in both cases leading to AMR13,14. At the same time, biofilms have been found to intrinsically exhibit AMR that is 1000-fold greater than in a single bacterium15. Together, this raises concern for their interactions in a drinking water distribution system (DWDS), which could result in antibiotic-resistant bacteria (ARB) propagation as well as the promotion and transfer of antibiotic resistance genes (ARGs)16. Thus far, DW field research has detected several ARGs in biofilms on DWDS pipes, including ampC, tetA, tetM, intI1, sul1, sul2, mexA, qnrS, qnrA, and qnrB17,18,19. However, this research was limited to investigating ARG occurrence in biofilms formed on cast iron pipes and did not establish whether sub-MICs of antibiotics in DW were responsible for their promotion.
It is important to examine AMR and bacteria in drinking water pipes because they present an emerging public health threat. Thus far, a limited number of early studies have found that surface water sediments and biofilms attached to treatment, pipe, and storage elements of water systems are environmental compartments that enhance horizontal gene transfer (HGT) and recombination and promote antibiotic resistance20,21,22,23. The presence of ARGs in streambed sediment of freshwater supplies and biofilms in treatment and distribution systems presents two emerging threats to public health: (1) a direct route of human ingestion of antibiotic-resistant pathogenic bacteria (e.g., resistant E. coli, Legionella spp., Mycobacterium spp., P. aeruginosa), and (2) the propagation of antibiotic resistance in the broader environment through water use for irrigation, commercial use, handwashing, cleaning, and food preparation.
With the current and projected rise of AMR-associated deaths24, it is important to identify and establish correlations with possible contributors of AMR. This study investigates whether the sub-MIC of ciprofloxacin promotes AMR in biofilms formed on polyvinyl chloride (PVC) DW pipe using a novel bench-scale water distribution reactor (BWDR). PVC pipe is one of the primary materials being used to replace aging pipes and is widely used in North American water distribution and premise plumbing25,26. As pipe material has been found to affect biofilm development and composition25, it is important to assess their respective biofilms in the context of AMR. With this, the objectives of this study are to: (1) determine whether biofilms formed on PVC pipe harbor ARGs, and (2) investigate whether a sub-MIC of ciprofloxacin correlates with the promotion of AMR and ARGs in biofilm through total cell count (TCC) and genetic sequencing, respectively.
Results
Ciprofloxacin concentration and biofilm TCC
The concentration of ciprofloxacin in the bulk water and the TCCs in the biofilm of each BWDR are shown in Fig. 1. As the ciprofloxacin concentration decreases in the bulk water, it sorbs and remains sorbed to/in the biofilm (i.e., no desorption). Therefore, the average concentration in the biofilm increased by 2.46 ± 0.45 μg L–1 after 1 day, 0.71 ± 0.71 μg L–1 by 12 days, and remained constant on/within the biofilm after 12 days. This is because no ciprofloxacin was detected in the bulk water after it was completely removed for the bulk water replacement. Overall, the average TCCs in the BWDR biofilms exposed to ciprofloxacin (i.e., A, B, and C) showed a significant increase, whereas the control (D) that was not exposed to ciprofloxacin did not see a significant increase (p-value < 0.005 and p value > 0.05, respectively). A complete table of the water quality parameters can be found in Table S1 of the Supplementary Material.
The ciprofloxacin (CIP) concentration in the bulk water of BWDRs A, B, and C is connected by solid lines, and its axis is found on the left. The biofilm TCCs in each BWDR, including the control (D) that was not exposed to ciprofloxacin, are connected by dashed lines and its axis is found on the right on the base-10 log scale. After 12 days, the ciprofloxacin was completely removed from the bulk water of BWDRs A, B, and C. Each point represents the mean (n = 3 independent samples), and the error bars indicate the standard deviation. Lines connecting mean values represent the rates of change.
Biofilm genus composition
The relative abundance at the genus level for each BWDR is shown in Fig. 2. Both the maximum genera and the average dominant genera were determined in the BWDRs. Here, the maximum genera was determined by comparing the relative amount of one genus across each BWDR on each day of exposure, and noting the genus with maximum abundance at one time in any given BWDR. The average dominant genera was determined by taking the average relative abundance of the dominant genera across all BWDRs and all times. Presenting both the maximum and average dominant genera communicates the level of variation across BWDRs. The maximum genera versus the average dominant genera (i.e., relative abundance of ≥ 0.05 in at least one BWDR time point) were Dechloromonas (71.5% and 16.7%), Pseudomonas (45.8% and 10.1%), Pseudoxanthomonas (43.8% and 10.7%), Acidovorax (29.0% and 12.5%), Allorhizobium-Neorhizobium-Pararhizobium-Rhizobium (24.2% and 11.1%), Nordella (16.5% and 5.6%), Azospira (7.7% and 1.5%), and Delftia (6.6% and 2.4%). A total of 98 unique genera were identified, and 199 species (including unidentified) were detected. A complete table of the genus compositions can be found in Table S2 of the Supplementary Material. A table indicating the number of species present at a relative abundance of ≥ 0.01 can be found in Table S3 of the Supplementary Material.
Relative abundance of the genus composition in each BWDR over time, where BWDRs A, B, and C were exposed to the sub-MIC of ciprofloxacin, and D is the control with no exposure to ciprofloxacin. 0d is before exposure to ciprofloxacin, *1d and *12d are during exposure to ciprofloxacin, and +14d is an additional 14 days after the ciprofloxacin was removed. Genera with a relative abundance of 0.01 or less were placed in “Others + unassigned”.
ARG quantification by qPCR and RT-qPCR
In total, 6 ARGs (i.e., qnrB, qnrS, mexA, intI1, sul1, and sul2) were selected for qPCR, and 4 ARGs (i.e., qnrB, qnrS, mexA, and intI1) were selected for RT-qPCR to determine their gene copies and transcript copies, respectively, normalized to the corresponding qPCR 16S rRNA. More specifically, qPCR identifies gene abundance, whereas RT-qPCR identifies gene expression, which measures the transcription of genes into mRNA. This can shed light on what genes are actively being expressed and potentially translated into functional proteins to activate AMR mechanisms. The qPCR and RT-qPCR assays were applied to each of the BWDR biofilm samples at each time point (Fig. 3). Figure 3 shows a heatmap of the ARG gene copies per 16S rRNA and transcript copies per 16S rRNA in BWDRs A to D (i.e., a) to d)), where the darker color denotes a greater number of gene/transcript copies per 16S rRNA, and light gray denotes that no copies were detected. Out of the 6 target qPCR genes, only 3 were detected. IntI1 was consistently detected at all time points in each BWDR at concentrations between 1.89 × 10–9 to 2.61 × 10–11 gene copies per 16S rRNA. The ARGs sul1 and sul2 were occasionally detected in all the BWDRs, but not at all time points. When detected, sul1 had a concentration range of 2.69 × 10–10 to 7.11 × 10–11 gene copies per 16S rRNA, and sul2 had a concentration range of 3.24 × 10–6 to 6.79 × 10–10 gene copies per 16S rRNA. The ARGs qnrB, qnrS, and mexA were not detected in any of the BWDRs. Similarly, out of the 4 target RT-qPCR genes, only intI1 was detected. When detected, intI1 had a concentration range of 2.54 × 10–7 to 1.52 × 10–8 transcript copies per 16S rRNA. Overall, the BWDR biofilms exposed to a sub-MIC of ciprofloxacin had a positive correlation with the promotion of each of the detected ARGs; however, none of these correlations were significant (p value > 0.05). Similarly, there were no significant correlations indicating co-occurrence between ARGs. A complete table of the correlations can be found in Table S4 of the Supplementary Material.
The gene copies (qPCR utilizing DNA) and transcript copies (RT-qPCR utilizing RNA) per 16S rRNA biomass obtained from the qPCR at each time point in (a) BWDR A, (b) BWDR B, (c) BWDR C, and (d) BWDR D (control, no ciprofloxacin). 0d is before exposure to ciprofloxacin, *1d and *12d are during exposure to ciprofloxacin, and +14d is after exposure to ciprofloxacin, where the ciprofloxacin was removed after the *12d time point and replaced with fresh water. The color legend indicates the quantity of gene copies per 16S rRNA and transcript copies per 16S rRNA.
Correlation between the sub-MIC of ciprofloxacin-exposed biofilms and ARGs
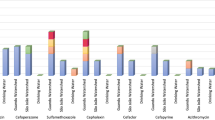
The promotion of the detected ARGs was correlated with the change in genus composition in the BWDRs (A, B, and C) exposed to the sub-MIC of ciprofloxacin (Fig. 4) with Spearman’s rank correlation (ρ). Figure 4 shows a bubble correlation matrix between the detected ARGs with the predominant genera, the Shannon-Wiener diversity, and the Pielou evenness, which showed at least one significant correlation with one ARG, indicated inside the bubble by an asterisk. The size and darkness of the bubble denote the strength of the correlation, and the color of the bubble denotes whether it is a positive or negative correlation, where red is negative, and blue is positive. Dechloromonas and Fonticella had very strong negative correlations with intI1 (ρ = -0.73 and -0.77, respectively, p values = 0.01 and 0.00, respectively), whereas Caulobacter and Sedimentibacter had strong negative correlations with intI1 (ρ = -0.66 and -0.68, respectively, p values = 0.02 and 0.01, respectively). Asinibacterium was the only genus to show a strong positive correlation with intI1 (ρ = 0.63, p value = 0.03). Nordella, Fibrisoma, and Brevundimonas had very strong negative correlations with sul1 (ρ = –0.73, –0.73, and –0.78, respectively, p values = 0.01, 0.01, and 0.00, respectively). Devosia, Asinibacterium, Galbitalea, Reyranella, and Sphingorhabdus had strong negative correlations with sul1 (ρ = –0.60, –0.61, –0.65, –0.68, –0.64, respectively, p values = 0.04, 0.03, 0.02, 0.02, 0.03, respectively). Overall, the majority of the correlations with genera for sul2 were negative, whereas with the RT-qPCR intI1 were positive; however, none of these correlations were significant. The Shannon-Wiener diversity indices for each of the BWDRs exposed to the sub-MIC of ciprofloxacin ranged from 3.44 to 1.70, indicating high to moderate diversity. The Pielou evenness indices were between 0.72 and 0.41, indicating high to moderate evenness. A complete table of the correlations can be found in Table S5 of the Supplementary Material.
Spearman correlation (ρ) between the qPCR intI1, sul1, sul2, and RT-qPCR intI1* ARGs and the genera in the biofilm for BWDRs A, B, and C that had a relative abundance of > 0.01, showing at least one significant correlation with an ARG. The sizes of bubbles indicate the strength of the correlation coefficient (larger bubbles denote a stronger correlation than smaller bubbles). The color of the bubble indicates a positive or negative correlation (blue denotes positive and red denotes negative). * is the significance indicator found inside bubble if *p value < 0.05; **p value < 0.01.
Discussion
It has been well established that biofilms will typically increase their production of extracellular polymeric substances (EPS) as a response to sub-MICs of antibiotics as a first-line intrinsic resistance mechanism27. However, it has been found that sub-MICs of ciprofloxacin are able to rapidly penetrate biofilm and do not remain localized on the surface28. A few studies to date have reported that a sub-MIC level of ciprofloxacin enhances biofilm production and formation29,30,31. Therefore, the observed significant increases in TCCs may have contributed to the AMR response of the biofilm to ciprofloxacin.
Differences in community composition between BWDRs were likely determined by the founder effect. This means that the founding genera of the biofilm at the initial time of attachment had an advantage over the subsequent colonizers and remained dominant. This has been reported in other DWDS biofilm studies32. Therefore, the changes in the community composition were not considered to be caused by the sub-MIC of ciprofloxacin. However, further research is necessary to confirm this. Regarding the predominant genera detected, this study used PVC as the pipe material for the BWDRs. PVC has been found to form substantially different community compositions than other pipe materials33. At the same time, the source water, and thus the native DW bacteria, used to generate DW biofilms in this study originated from Lake Ontario, Canada, which also presents a unique set of species that have not yet been documented with 16S rRNA to the authors' knowledge. Therefore, when making comparisons with other DW biofilm studies, which are primarily conducted in other countries (e.g., China), it is important to note the effects that environmental variables and geographic distance can have on community composition and speciation34. For instance, Vaz-Moreira et al.35 found that while the same species of Pseudomonas could be isolated from different sampling sites (e.g., biofilm versus tap water), their genotypes were never identical.
The qPCR and RT-qPCR results demonstrate both consistencies and divergences compared to the inferred metagenomic predictions and previous ARG studies in DWDS biofilms. For instance, Wang et al.36 demonstrated that a sub-MIC of ciprofloxacin induced the promotion of the ARGs intI1, sul1, sul2, qnrB, qnrS, and mexA. However, their biofilms were formed on cast iron coupons and had dominant genera that were different than those found in this study36. When comparing the results of this study, previous exposure to antibiotic residues, associated fitness costs, and community composition may provide insight into the lack of significant correlation between the presence of the sub-MIC of ciprofloxacin and ARG presence/promotion. In a preliminary study, the Lake Ontario source water and GAC effluent were tested for several antibiotics, including ciprofloxacin residuals, and were not detected. Therefore, the native DW bacteria forming the biofilms in this study may have had a low prior exposure to antibiotic residues. Consequently, maintaining and developing ARGs in the absence of this selective pressure likely imposed a significant fitness cost, thereby limiting their initial presence and abundance37. Fitness cost refers to the reduction in the overall survival or reproductive success of an organism as a result of carrying a genetic trait, such as an ARG. For instance, Flores-Vargas et al.38 found that when riverine biofilms were developed under a sub-MIC exposure to antibiotics, they demonstrated significantly higher absolute numbers of ARGs (e.g., sul1, sul2) with gene richness compared to control samples. Despite this, intI1 was consistently detected and occasionally transcribed in this study, which raises concerns about the potential for a shift in the currently selected resistance mechanisms towards ARG promotion and mobilization should selection pressures arise. This is particularly noteworthy as intI1 has been reported to be strongly associated with the dissemination of ARGs39. For instance, Huang et al.40 reported positive correlations between intI1 and the proliferation and propagation of sul1 and sul2, which is consistent with the ARGs detected in this study. In addition, it has also been found that sul2 has the highest gene copies compared to other ARGs when exposed to a sub-MIC of antibiotics, which is also consistent with the findings of this study38. The presence of sulfonamide resistance genes sul1 and sul2 when the biofilm is exposed to ciprofloxacin might suggest that a co-selection is operating to promote resistance to multiple antibiotic classes.
Overall, the changes in the community composition reflected the ARG patterns that were observed in the biofilm. Specifically, these correlations can be directly compared with the changes in 16S rRNA community composition seen in Fig. 2 and the changes in ARGs seen in Fig. 3. This agrees with previous studies that have also found that the ARG sul1 correlates with the alpha diversity of microbial phylotypes in the sediments of environmental samples41. In addition, it has been found that diverse multi-species biofilms exhibit significantly higher AMR than single-species biofilms42,43,44. However, the present study highlights that this is dependent on the species present, their selected AMR mechanism (e.g., increase in TCCs), as well as the extent of current and emerging selection pressures. More specifically, this study found that there is an increased risk of ARG selection when there is either a less diverse biofilm and/or it contains bacteria such as Asinibacterium. While this study presented a unique consortium of genera native to Lake Ontario sourced DW, studies composed of different genera reported similar findings. For instance, Klumper et al.45 found that higher biodiversity can act as a barrier to the persistence and spread of ARBs and ARGs. In addition, Simoes et al.46 found that the presence or absence of certain species, such as Acinetobacter calcoaceticus, made DW biofilms more resistant. This highlights the need to monitor diversity as an effective control strategy against AMR development. Haenelt et al.47 also came to a similar conclusion and stated that limiting the release of ARG-harboring bacteria may be more crucial than reducing residual antibiotic concentrations in water in controlling AMR. However, where residual concentrations of antibiotics were not attenuated, sub-MICs of antibiotics were found to reduce community composition diversity and, in turn, select for species with ARGs38. It is noted that biofilm diversity can also have an opposite effect on AMR. Biofilms with a high degree of diversity of species and genes can experience higher levels of HGT and thus promote higher levels of AMR. Nevertheless, while community composition may play a greater role in AMR, the presence of a sub-MIC of an antibiotic, such as ciprofloxacin, should not be overlooked.
These results underscore the importance of both monitoring antibiotic residues and biofilm community composition in DWDSs to manage and mitigate the risks associated with AMR. Future studies should explore the long-term implications of sub-MIC antibiotic exposure on biofilm AMR/ARGs, especially in geographic areas that have not yet detected antibiotics in their water, given the potential public health impacts. Overall, this study is the first to characterize biofilms from DW sourced from Lake Ontario and on PVC pipe, provide valuable insights into the mechanisms of multi-species biofilm AMR and ARGs, and emphasize the need for comprehensive strategies to control AMR in DWDSs.
Methods
Reagents and materials
Ciprofloxacin (purity > 98%) was purchased from ThermoFisher Scientific (MA, USA). The stock solution for ciprofloxacin was prepared by weighing and introducing a requisite amount of ~10 g. Ciprofloxacin was then dissolved at room temperature (23 °C) in deionized (DI) water by magnetic stirring at 1600 rpm for 72 h in the dark to yield a concentration below 10 mg L–1. The stock solution was then filtered using a 0.2 μm pore size Nylon syringe filter to remove any solids formed from the liquid phase. The stock solution was then stored in an opaque flask at 4 °C for a maximum of 14 days. A set of 8 serial dilutions were made from samples taken from the stock solution to generate a calibration curve between 0.1 μg L–1 to 500 μg L–1. The ciprofloxacin concentrations were then measured with an LC-MS/MS prior to the desired ciprofloxacin concentration (10 μg L-1) being introduced into the BWDRs.
Experimental set-up and growth period
The water used in this experiment was representative of DW in Kingston, Ontario, Canada, with the exception of having been dechlorinated. Two granular activated carbon (GAC) columns, each composed of 0.86 kg of coconut shell high-activated carbon, were configured in a series arrangement as an open system with continuous flow. The first GAC column was connected directly to an on-site tap and served to dechlorinate the water, while the second GAC column served to reintroduce native DW bacteria, where the effluent from this second GAC was then subsequently collected at the output to supply the BWDRs.
Four BWDRs were constructed to operate as continuous, semi-closed systems and received the effluent of the GAC columns. The bacteria-rich effluent from the GACs was introduced into the BWDRs every two weeks during the biofilm growth period and prior to the injection of the ciprofloxacin. There is a possibility that adding new GAC effluent on a bi-weekly basis introduced new bacteria and genes and increased bacterial diversity in the BWDRs. Despite this, the tap water conditions and the conditions inside the GACs did not vary during the experiment, so the bacterial population between 2-week refresh periods was likely comparable.
During the entirety of the experimental period, the following water quality parameters were continuously measured: temperature and pH (Accumet XL60), turbidity (Hach 2100Q turbidimeter, USA), UV absorbance at 600 nm (OrionTM AquaMate UV-VIS Spectrophotometer, ThermoScientific, USA), total organic carbon (TOC) and total nitrogen (TN) (Shimazdu TOC analyzer, Shimadzu, USA), free chlorine (DR300 Pocket Colorimeter, Hach, USA), adenosine triphosphate (ATP) (Quench-Gone Aqueous (QGA) test kit (LuminUltra Technologies, Canada), with the Lumitester C-110 luminometer (Kikkoman, Japan), and TCCs (Bacterial Counting Kit for flow cytometry, Invitrogen, USA) according to their respective manufacturers’ protocols.
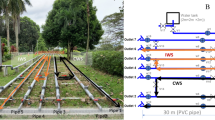
To generate mature, multi-species biofilms, the BWDRs were independently operated at room temperature (20–22 °C) at a turbulent flow rate of 3 litres per minute (Reynolds number = 3000) for 12 weeks. The biofilms were fed with 0.24 g L–1 of nutrient broth (No. 3, NutriSelect® Plus, Millipore Sigma, MO, USA) at the start of each week to expedite bacterial growth and replication. The BWDRs were each composed of 9 meters of looped ¾-inch (1.91 cm) ID PVC pipe with a 4 ¼-inch (10.80 cm) ID PVC reservoir. Each BWDR was comprised of 5 removable sections (surface area 151.29 cm2 each) to obtain sufficient biomass for genetic sequencing and 40 coupons (surface area 0.78 cm2 each) located along the pipe invert for rapid, non-intrusive biofilm monitoring to obtain TCC data (Fig. 5). The coupons were comprised of round disks that were drilled and removed from the same PVC pipes used for constructing the BWDRs. The disks were supported by an outer structure that was 3D-printed (Fig. 5). The disk/support structure was fastened to the pipe by means of vinyl ties to eliminate any leaks from the BWDRs. Ball valves were used to isolate the removable pipe segments from the rest of the network before sampling.
One BWDR consists of a 4 ¼-inch (10.80 cm) ID PVC reservoir and 9 meters of looped ¾-inch (1.91 cm) ID PVC pipe, circulating a total of 7 L of bulk water. Along the pipes are removable sections as well as coupons with a PVC sample disk that fits into holes drilled along the invert of the pipe. Each BWDR has (1) a 5 L PVC reservoir, (2) an adjustable flow meter, (3) a 3D-printed valve to control the flow rate, (4) a centrifugal pump (24 V DC), and (5) ¼-inch (0.64 cm) ID black vinyl tubing connecting the reservoir to the piping. BWDRs were operated for 12 weeks to generate mature biofilm prior to their experimental period54.
Ciprofloxacin experimental period
Following the 12-week biofilm growth period, the 4-week experimental period was initiated to maintain the aforementioned growth conditions. The experimental period consisted of 2 phases: the ciprofloxacin exposure phase (12 days) and the subsequent recovery phase (14 days). Samples were taken at 0d (before exposure), 1d (1 day into exposure), 12d (12 days into exposure), and +14d (14 days post-exposure, where biofilms had 14 days to recover with no ciprofloxacin) (Fig. 6).
During the experimental period, 0d represents the initial conditions prior to the sub-MIC ciprofloxacin exposure, *1d represents the first day of sub-MIC ciprofloxacin exposure after it is initially administered, *12d is the final day of the sub-MIC ciprofloxacin exposure where the bulk water is completely replaced after the final samples are taken, and +14d is 14 days after the bulk water replacement where there is no ciprofloxacin remaining, allowing the biofilm to recover. Red dots represent the ciprofloxacin.
Prior to the experimental period, biofilms were given 2 days to stabilize after their bulk water replacement and nutrient broth administration. Testing showed that after introducing the broth in the BWDRs, the rate of bacterial growth in the bulk water would increase dramatically, but then resume its steady increase after approximately 2 days. The 2-day period was then selected as the stabilization period for these experiments. For the exposure phase, a 10 μg L–1 sub-MIC of ciprofloxacin was administered to three of the BWDRs (i.e., A, B, and C), all considered to be experimental replicates. The sub-inhibitory selective concentration of 10 μg L–1 in ciprofloxacin was chosen on the basis of previous studies that found comparable concentrations of ciprofloxacin in environmental samples13,14,24.
BWDR D was the biofilm control that was not exposed to ciprofloxacin. The ciprofloxacin was administered to the experimental replicates by pipetting the appropriate volume from the stock solution directly into each of the appropriate BWDR reservoirs. For the recovery phase, the bulk water in each BWDR was replaced as per the previously described protocol using the GAC effluent, ensuring that all of the ciprofloxacin was removed from the bulk water. Bulk water samples confirming the ciprofloxacin concentration were taken in triplicate at each time point and immediately tested using a ThermoFisher Exploris 120 Orbitrap mass spectrometer connected to a Vanquish ultra-high-performance liquid chromatography (uHPLC) system, with the autosampler held at 10 °C to preserve sample stability. Analytes were separated using a 50 mm × 2.1 mm × 3.0 μm Zorbax C18 analytical column. Mobile phases consisted of 0.1% formic acid in LC-MS grade water (A) and LC-MS grade acetonitrile (B). A gradient elution profile was used, starting at 90:10 A:B for the first 30 s, before shifting to 10:90 A:B over the next 2.5 min (three minutes total), holding at 10:90 A:B for one minute, before re-equilibrating at starting conditions for three minutes. After separation, analytes were ionized using a heated electrospray ionization source with a static spray voltage, running a positive ion voltage of 2500 V with an Orbitrap resolution of 30,000. Gasses were run in static mode, with a sheath gas of 50 (AU), auxiliary gas of 10 (AU), and sweep gas of 1 (AU). The ion transfer tube temperature and vaporizer temperature were set to 325 °C and 350 °C, respectively. The MS Global settings were set to expect an LC peak width of 10 s, and mass calibration was performed using the RunStart Easy-IC ™ internal calibration system. Analyte peak areas were acquired in full scan mode, with extraction ion chromatograms used to calculate the peak area for quantification purposes. Analyte peak area was determined using XCalibur QuanBrowser, monitoring for m/z 332.1404. Analytes were quantified using an external calibration curve of 0.26 μg L–1 to 52 μg L–1 (R2 = 0.999). The calibration curve was based on a set of serial dilutions created by the stock solution. Quantification limits were set to the lowest concentration of the calibration curve.
Biofilm sample collection
Coupon samples were extracted from each of the BWDRs in triplicate to monitor changes in the TCCs and activity of the biofilms during the growth and experimental periods. The TCC samples were rinsed by submerging a coupon in DI water and swabbed using a sterile swab that was then placed in a prepared fixative solution containing 100 μL of glutaraldehyde (50 wt.%, 340855, Millipore Sigma) and 9.9 mL of DI water. TCC samples were stored in the dark at 4 °C for up to 21 days. For processing, TCC swabs were vortexed at 3000 rpm (Fisherbrand™ Digital Vortex Mixer) for 30 s, then sonicated at a low frequency of 40 kHz (Branson Ultrasonics™ CPX1800, Danbury, CT) for 1 min, then vortexed again for 30 s. To quantify TCCs, the Bacterial Counting Kit for flow cytometry (Invitrogen, Carlsbad, CA USA) was used according to the manufacturer’s protocol and analyzed on a SH800 cell sorter (Sony, Japan). The resulting data analysis was performed using FlowJo™ (BD Biosciences, NJ, USA).
Removable section samples were only extracted during the experimental period. Each removable section contained sufficient biomass for DNA and RNA nucleic acid extractions for 16S rRNA, qPCR, and RT-qPCR. Each removable section was rinsed in DI water by vertical submersion. Sterile swabs were used to remove all visible biofilms from the inner pipe. Swab contents for each removable section were placed in sterile, 15 mL nuclease-free tubes (FroggaBio, ON, Canada) containing 5 mL of autoclaved DI water. The tubes were then vortexed at 3000 rpm (Fisherbrand™ Digital Vortex Mixer) for 30 s, followed by sonication at a low frequency of 40 kHz (Branson Ultrasonics™ CPX1800, Danbury, CT) for 1 min, and then vortexed again for 30 s. Once the biomass was detached from the swabs, they were then removed from the tubes. The tubes were then centrifuged at 3800 rpm for 10 min, creating a pellet. The resulting 4 mL supernatant was discarded. All nuclease-free tubes were stored at –80°C until DNA and RNA extraction. Environmental DNA (eDNA) was extracted from biofilms using the Sox DNA Extraction kit (Metagenom Bio Life Science Inc.), with the exception of ceramic beads that were used for homogenization. Total RNA extraction and genomic DNA removal were performed using RNeasy PowerBiofilm Kit (Qiagen) according to the supplier’s user guide.
16S rRNA
16S rRNA V4 regions were PCR amplified using the forward primer and reverse primer 515FB:5′-GTGYCAGCMGCCGCGGTAA-3′ and 806RB:5′-GGACTACNVGGGTWTCTAAT-3′, respectively48. Illumina sequencing adapters and barcodes were added to the 5′ ends of the primers and each indexed primer pair contained a 5′ random addition of up to six random nucleotides to improve heterogeneity of the sequencing run, improving read quality and yield. PCR was set up in duplicate (25 μL each): 2.5 μL of 10 × standard Taq reaction buffer, 0.5 μL 10 mM dNTP, 0.2 μL Taq DNA polymerase (New England Labs), 0.2 μL BSA (20 mg mL–1, NEB), 5.0 μL of 1 μM forward and reverse primers, respectively, 9.1 μL water, and 2.5 μL eDNA (1–10 ng μL–1). PCR was run as follows: 94 °C for 5 min, 35 cycles of 94 °C for 30 s, 50 °C for 30 s, and 68 °C for 1 min, and finally 68 °C for 5 min. PCR amplicons were pooled and resolved in 2% agarose gel and the gel was purified and quantified using the Qubit dsDNA quantification Assay Kit (ThermoFisher). The 16S rRNA amplicons were diluted to 4 nM, denatured, and sequenced using MiSeq 500-cycle sequencing v2 kit (Illumina, Cat. No. MS-102-2003). Demultiplexed sequences were processed using cutadapt (primer removal49) and DADA2 v.1.22 (sample inference, Callahan et al.50). Reads were truncated at decreasing quality scores and assembled (typically 225 on read R1 and 200 on R2 relative to the original read start position). After quality filtering (chimera, organelles), an amplified sequence variant (ASV) abundance table was constructed. This ASV table records the number of sequence reads for each amplified sequence variant (“species”) for each sample. Taxonomy was assigned to representative sequences using the naive Bayesian classifier implemented in the dada2::assignTaxonomy() function from DADA250 trained against the SILVA reference database release 13851.
Inferred metagenomics
Inferred metagenomics was utilized to estimate ARG presence and abundance based on the 16S rRNA results. An inferred metagenome for each sample was predicted from the 16S rRNA ASV table using PICRUSt2 v.2.5.252. This predicted metagenome was screened for KEGG Orthology Identifiers (KO values) associated with antibiotic resistance (i.e., aminoglycoside, beta-lactamase, macrolide, phenicol, tetracycline, quinolone, sulfonamide, integron, and multidrug efflux pumps). KO values represent standardized functional annotations that group genes from different organisms based on shared biological roles. Total normalized hits for these KO values were processed and plotted using R v.4.1.3 (R Core Team53). The results of the inferred metagenome can be found in Fig. S1 of the Supplementary Material. The results were used to determine which ARGs should be further investigated. The ARGs qnrB, qnrS, mexA, intI1, sul1, and sul2 were selected for investigation due to their consistent abundance prediction and resistance representation.
DNA qPCR and RNA RT-qPCR quantification
qPCR analysis was chosen to identify gene abundance in samples. RT-qPCR analysis was chosen to identify gene expression and measure the transcription of genes into mRNA in the samples. This information indicates which genes are actively being expressed and potentially translated into functional proteins to activate AMR mechanisms. A total of 6 ARGs (i.e., qnrB, qnrS, mexA, intI1, sul1, and sul2) were selected for qPCR, and 4 ARGs (i.e., qnrB, qnrS, mexA, and intI1) were selected for RT-qPCR to determine their gene copies and transcript copies, respectively, normalized to the corresponding qPCR 16S rRNA.
Candidate primers were identified through a literature review and modified for taxonomic coverage where necessary, screening against NCBI genomes and the nt database for common variants. PCR primers and probes (Table 1) were then further refined using PrimerQuest Tool (Integrated DNA Technologies) and purchased from Integrated DNA Technologies, Inc. (IDT, Inc.). eDNA and RNA were quantified using PowerGreen™ SYBR Green 2x qPCR Master Mix (GeneBio Systems, Cat. No. P2101-02) and TaqMan Fast Advanced Master Mix for qPCR (ThermoFisher, Cat. No. 4444556), respectively. Design and Analysis Software 2.6.0 (ThermoFisher) was employed for qPCR running with the QuantStudio 3 instrument and data management.
Statistical analysis
Changes in ciprofloxacin concentration and biofilm TCC
The TCC data was transformed using a base-10 log. Average TCC values were calculated for the experimental replicates (i.e., BWDRs A, B, and C) for an n = 3 values for each time of 0d, *1d, *12d, and +14d. All datasets were subject to the Shapiro-Wilk test to determine whether the data were normally distributed and if the null hypothesis (p-value > 0.05) could be accepted. Since the null hypothesis was accepted, ANOVA was used.
Correlation between the sub-MIC of ciprofloxacin and ARGs
As the concentration of ciprofloxacin decreased in the bulk water, it increased on/in the biofilm. Therefore, the change in the concentration in/on the biofilm and the change in concentration of each ARG were utilized to generate n = 9 matched pairs. The Shapiro-Wilk test was applied and rejected the null hypothesis (p value < 0.05); therefore, the non-parametric Spearman Rank correlation was used.
Correlation between community composition and ARGs
The community composition was grouped by genus and organized by percent of relative abundance. Genera with an average relative abundance over 0.01 were included. Percent relative abundance values were then normalized to the corresponding qPCR 16S rRNA count. For the Shannon-Wiener diversity and Pielou evenness indices, the absolute abundance values of the species were used. All correlations were performed on the experimental replicates (BWDR A, B, and C) to create n = 12 matched pairs. The Shapiro-Wilk test was applied to each dataset to determine normality. All datasets rejected the null hypothesis (p value < 0.05); therefore, the non-parametric Spearman Rank correlation was used. All data analyses were conducted using Jamovi version 2.4.12. The graphs and figures were created with Excel – Microsoft 365 MSO (version 2506 build) and RStudio (version 2023.06.1 + 524).
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information file.
References
Ventola, C. L. The antibiotic resistance crisis: part 1: Causes and threats. Pharm. Ther. 40, 1–7 (2015).
Klein, E. Y. et al. Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proc. Natl. Acad. Sci. 115, E3463–E3470 (2018).
Browne, A. J. et al. Global antibiotic consumption and usage in humans, 2000–18: a spatial modelling study. Lancet Planet. Health 5, e893–e904 (2021).
World Health Organization. Antibiotic resistance. https://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance (2020).
Mouton, J. W. et al. MIC-based dose adjustment: facts and fables. J. Antimicrobial Chemother. 73, 564–568 (2018).
López-Pacheco, I. Y. et al. Anthropogenic contaminants of high concern: Existence in water resources and their adverse effects. Sci. Total Environ. 690, 1068–1088 (2019).
World Health Organization. Global Antimicrobial Resistance and Use Surveillance System (GLASS). https://www.who.int/initiatives/glass (2025).
de Ilurdoz, M. S., Sadhwani, J. J. & Reboso, J. V. Antibiotic removal processes from water & wastewater for the protection of the aquatic environment-a review. J. water process Eng. 45, 102474, 1–14 (2022).
Rogowska, J. & Zimmermann, A. Household pharmaceutical waste disposal as a global problem—a review. Int. J. Environ. Res. Public Health 19, 15798 (2022).
Mahmood, A. R., Al-Haideri, H. H. & Hassan, F. M. Detection of antibiotics in drinking water treatment plants in Baghdad City, Iraq. Adv. Public Health 2019, 1–10 (2019).
Wang, Y. et al. Antibiotic residues of drinking-water and its human exposure risk assessment in rural Eastern China. Water Res. 236, 119940 (2023).
Ben, Y. et al. Efficient detection and assessment of human exposure to trace antibiotic residues in drinking water. Water Res. 175, 1–11 (2020).
Stanton, I. C., Murray, A. K., Zhang, L., Snape, J. & Gaze, W. H. Evolution of antibiotic resistance at low antibiotic concentrations including selection below the minimal selective concentration. Commun. Biol. 3, 1–11 (2020).
Kraupner, N. et al. Selective concentration for ciprofloxacin resistance in Escherichia coli grown in complex aquatic bacterial biofilms. Environ. Int. 116, 255–268 (2018).
Hoyle, B. D. & Costerton, J. W. Bacterial resistance to antibiotics: the role of biofilms. Prog. Drug Res. 37, 91–105 (1991).
Rilstone, V. et al. The role of antibiotics and heavy metals on the development, promotion, and dissemination of antimicrobial resistance in drinking water biofilms. Chemosphere 282, 1–18 (2021).
Chen, Y. et al. Controlling biofilm growth and its antibiotic resistance in drinking water by combined UV and chlorination processes. Water 14, 1–9 (2022).
Kimbell, L. K. et al. Cast iron drinking water pipe biofilms support diverse microbial communities containing antibiotic resistance genes, metal resistance genes, and class 1 integrons. Environ. Sci. Water Res. Technol. 7, 584–598 (2021).
Wang, H., Hu, C., Liu, L. & Xing, X. Interaction of ciprofloxacin chlorination products with bacteria in drinking water distribution systems. J. Hazard. Mater. 339, 174–181 (2017).
Xi, C. et al. Prevalence of antibiotic resistance in drinking water treatment and distribution systems. Appl. Environ. Microbiol. 75, 5714–5718 (2009).
Su, H. C. et al. Persistence of antibiotic resistance genes and bacterial community changes in drinking water treatment system: From drinking water source to tap water. Sci. Total Environ. 616, 453–461 (2018).
Lu, J., Tian, Z., Yu, J., Yang, M. & Zhang, Y. Distribution and abundance of antibiotic resistance genes in sand settling reservoirs and drinking water treatment plants across the Yellow River, China. Water 10, 246 (2018).
Xu, L. et al. High-throughput profiling of antibiotic resistance genes in drinking water treatment plants and distribution systems. Environ. Pollut. 213, 119–126 (2016).
Murray, A. K. et al. A critical meta-analysis of predicted no effect concentrations for antimicrobial resistance selection in the environment. Water Res. 266, 122310 (2024).
Lee, D. et al. The impact of pipe material on the diversity of microbial communities in drinking water distribution systems. Front. Microbiol. 12, 779016 (2021).
Abdulah Shrrat Omar, O. Evaluation of pipe materials in water system networks using the theory of advanced multi-criteria analysis. Sustainability 15, 4491 (2023).
Singh, S., Datta, S., Narayanan, K. B. & Rajnish, K. N. Bacterial exo-polysaccharides in biofilms: role in antimicrobial resistance and treatments. J. Genet. Eng. Biotechnol. 19, 1–19 (2021).
Tseng, B. S. et al. The extracellular matrix protects Pseudomonas aeruginosa biofilms by limiting the penetration of tobramycin. Environ. Microbiol. 15, 2865–2878 (2013).
Rafaque, Z. et al. In vitro investigation of antibiotics efficacy against uropathogenic Escherichia coli biofilms and antibiotic induced biofilm formation at sub-minimum inhibitory concentration of ciprofloxacin. Infect. Drug Resistance, 2801–2810 (2020).
Whelan, S., O’Grady, M. C., Corcoran, G. D., Finn, K. & Lucey, B. Effect of sub-inhibitory concentrations of nitrofurantoin, ciprofloxacin, and trimethoprim on in vitro biofilm formation in uropathogenic Escherichia coli (UPEC). Med. Sci. 11, 1–9 (2022).
Luo, Z. X., Li, Y., Liu, M. F. & Zhao, R. Ciprofloxacin enhances the biofilm formation of Staphylococcus aureus via an agrC-dependent mechanism. Front. Microbiol. 14, 1–11 (2023).
Kelly, J. J., Minalt, N., Culotti, A., Pryor, M. & Packman, A. Temporal variations in the abundance and composition of biofilm communities colonizing drinking water distribution pipes. PloS One 9, 1–11 (2014).
Learbuch, K. L. G., Smidt, H. & Van Der Wielen, P. W. J. J. Influence of pipe materials on the microbial community in unchlorinated drinking water and biofilm. Water Res. 194, 1–13 (2021).
Ma, B., LaPara, T. M., N Evans, A. & Hozalski, R. M. Effects of geographic location and water quality on bacterial communities in full-scale biofilters across North America. FEMS Microbiol. Ecol. 96, 1–9 (2020).
Vaz-Moreira, I., Nunes, O. C. & Manaia, C. M. Diversity and antibiotic resistance in Pseudomonas spp. from drinking water. Sci. Total Environ. 426, 366–374 (2012).
Wang, H. et al. Response of microorganisms in biofilm to sulfadiazine and ciprofloxacin in drinking water distribution systems. Chemosphere 218, 197–204 (2019).
Rajer, F. & Sandegren, L. The role of antibiotic resistance genes in the fitness cost of multiresistance plasmids. MBio 13, e03552–21 (2022).
Flores-Vargas, G., Korber, D. R. & Bergsveinson, J. Sub-MIC antibiotics influence the microbiome, resistome and structure of riverine biofilm communities. Front. Microbiol. 14, 1–19 (2023).
Huyan, J. et al. Dynamics of class 1 integrons in aerobic biofilm reactors spiked with antibiotics. Environ. Int. 140, 1–9 (2020).
Huang, Z., Zhao, W., Xu, T., Zheng, B. & Yin, D. Occurrence and distribution of antibiotic resistance genes in the water and sediments of Qingcaosha Reservoir, Shanghai, China. Environ. Sci. Eur. 31, 1–9 (2019).
Cheng, X. et al. Analysis of Antibiotic Resistance Genes Environmental Factors and Microbial Community From Aquaculture Farms in Five Provinces China. Front. Microbiol. 12, 1–10 (2021).
Whiteley, M., Ott, J. R., Weaver, E. A. & McLean, R. J. Effects of community composition and growth rate on aquifer biofilm bacteria and their susceptibility to betadine disinfection. Environ. Microbiol. 3, 43–52 (2001).
Leriche, V., Briandet, R. & Carpentier, B. Ecology of mixed biofilms subjected daily to a chlorinated alkaline solution: spatial distribution of bacterial species suggests a protective effect of one species to another. Environ. Microbiol. 5, 64–71 (2003).
Burmølle, M. et al. Enhanced biofilm formation and increased resistance to antimicrobial agents and bacterial invasion are caused by synergistic interactions in multispecies biofilms. Appl. Environ. Microbiol. 72, 3916–3923 (2006).
Klümper, U. et al. Environmental microbiome diversity and stability is a barrier to antimicrobial resistance gene accumulation. Commun. Biol. 7, 1–13 (2024).
Simoes, L. C., Simoes, M. & Vieira, M. J. Influence of the diversity of bacterial isolates from drinking water on resistance of biofilms to disinfection. Appl. Environ. Microbiol. 76, 6673–6679 (2010).
Haenelt, S. et al. The fate of sulfonamide resistance genes and anthropogenic pollution marker intI1 after discharge of wastewater into a pristine river stream. Front. Microbiol. 14, 1–12 (2023).
Walters, W. et al. Improved bacterial 16S rRNA gene (V4 and V4-5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. Msystems 1, e00009–15 (2016).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet. J. 17, 10–12 (2011).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–58 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
R. Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/ (2022).
Rilstone, V., Filion, Y. & Champagne, P. Study on the persistence of ciprofloxacin and sulfamethoxazole in simulated drinking water systems. Environ. Syst. Res. 14, 1–12 (2025).
Acknowledgements
This study was supported by the Ontario Research Fund [RE010-095] and the Natural Science and Engineering Research Council [RGPIN-2019-04084]. The funders played no role in the study design, data collection, analysis and interpretation of data, or the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
V.R.—conceptualization, methodology, validation, formal analysis, investigation, writing—original draft, writing—review and editing, and visualization. Y.F.—resources, supervision, writing—review and editing, and funding acquisition. P.C.—resources, supervision, writing—review and editing, and funding acquisition. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rilstone, V., Filion, Y. & Champagne, P. Fate of the resistance profile of drinking water biofilm exposed to a sub-minimum inhibitory concentration of ciprofloxacin. npj Antimicrob Resist 4, 19 (2026). https://doi.org/10.1038/s44259-026-00190-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44259-026-00190-y