Abstract
Inflammation is implicated in placental dysfunction leading to stillbirth, yet evidence of a systemic immune response during parturition remains unclear. Here, we present the first transcriptomic analysis of maternal systemic responses associated with stillbirth, using a minimally invasive finger-prick method for whole blood collection in labouring women in northern Nigeria. This approach facilitated participant recruitment with minimal disruption to care and provided high-quality RNA for transcriptomic profiling. Contrary to expectations, no major differences were observed in systemic immune states between propensity-matched live births and stillbirths. These findings suggest several possibilities, including the dominance of a parturition response masking an underlying immune state, physical protective barriers, or placental tolerance mechanisms. This study offers novel insights into maternal systemic immune health during stillbirth and highlights the need for further research. It also contributes to the broader “Stillbirths in Kano” initiative, aimed at reducing stillbirth rates through enhanced prenatal care.
Similar content being viewed by others
Introduction
The definition of stillbirth varies across the world. The World Health Organization defines stillbirth as ‘A baby who dies after 28 weeks of pregnancy, but before or during birth, is classified as a stillbirth’1. Based on this definition, in 2019 an estimated 2 million babies were stillborn2 with the majority occurring in low- and middle-income countries (LMICs). Within LMICs, the highest proportion of stillbirths was reported to occur in sub-Saharan Africa and South Asia3. Nigeria is reported to have one of the highest stillbirth rates on the African continent and one of six countries that bear the burden of 50% of global stillbirths4.
Over 40% of fetal deaths leading to stillbirths occur during parturition (labour and birth) and are likely preventable with high-quality care during pregnancy and birth5. Antenatal intrauterine fetal deaths are often associated with preventable conditions such as maternal infections and non-communicable diseases6. Other reported risk factors for stillbirths include maternal age (young or advancing), maternal education level, socioeconomic deprivation, hypertension, fetal infection, perinatal asphyxia, previous stillbirth, intrauterine growth restriction, abruptio placenta and placenta praevia, and substandard antenatal care7,8,9,10. Stillbirth rates are commonly used to assess levels and quality of care within a healthcare system, yet in many LMICs stillbirths are often not recorded due to a multitude of barriers in data collection, resulting in inaccurate and unreliable data2,11.
Pregnancy involves complex immunological interactions between the maternal immune system and the developing fetus. These interactions are critical for healthy fetal growth and development. Previous research efforts have tried to better understand the role of immune cell subsets and placental pathology in pregnancy complications, emphasizing the need for careful monitoring of fetal loss and stillbirth in cases of fetal growth restriction, along with a focus on placental pathology to understand its role in adverse pregnancy outcome12,13,14. To date, research into immune cell imbalances in pregnancy complications aims to improve the understanding and potentially develop interventions to mitigate these risks15. Infections are a highly suspected cause for fetal or embryo death especially in early stages of pregnancy16,17. However, investigating the infectious causes and mechanisms contributing to stillbirths in humans is challenging. In this regard, studies in nonhuman primates (NHPs) have provided valuable insights. In the case of teratogenic infections, such as those caused by the Zika virus, NHPs have been shown to closely resemble human responses to infection and pregnancy outcomes. Most notably, the high rates of fetal loss observed in asymptomatic ZIKV-infected NHPs raise concerns about the frequency of infection-associated pregnancy loss in humans18. The central question of whether a circulatory host response change marking either a maternal immune state of infection or fetal death remains unknown.
Here we seek to explore this question in humans, and forms part of a wider feasibility study focussing on stillbirths in Kano. A key output of the feasibility study was to inform further research using a mixed-methods approach and pave the way for future intervention implementation to prevent stillbirth. The primary aims of the immunological component, covered in this report, were to identify whether it was feasible to collect and successfully transfer blood samples taken from the mothers to our laboratory in the UK and successfully perform a transcriptomic analysis, as well as assess if a systemic maternal host response was identifiable in mothers who delivered a stillborn baby. We sought to compare the gene expression profiles of mothers who delivered a liveborn baby and identify any distinct patterns or differences that may be associated with stillbirth. To accomplish this, blood samples were collected from both groups of mothers using a finger-prick method, due to the advantages when compared to a standard venepuncture procedure. Finger-prick collection is a quick and minimally invasive procedure that can be performed at the bedside or in the community, making it suitable for time-sensitive situations such as labour. Finger-prick approaches for gene expression profiling or whole genome transcriptomics have several key advantages that help lower barriers for use, although to date this has not been reportedly used in LMICs19. Accordingly, we have sought to test the feasibility of deploying a finger-prick methodology in a low-resource setting, with the aim of simplifying the handling and processing of samples at the point of collection. Notably, it can be performed by non-phlebotomists, requires less time and resources, avoiding the need for centrifugation, separation, and the immediacy of storage compared to larger blood samples.
By comparing the gene expression profiles between the two groups, we sought to test the possibility of detecting genes and pathways that may be significantly altered in the stillbirth group compared to the live birth group. Such observational transcriptional status could provide insights into the systemic immune and molecular pathways and processes that may be implicated in stillbirth. These studies should help provide insight that may also facilitate the discovery of novel biomarkers or therapeutic targets for the diagnosis, prevention, and management of stillbirth in the future.
Results
Overview of study design
Four hundred and one mothers were approached to participate in the whole blood transcriptomic investigation. All consented to participate, including 59 mothers who were affected by stillbirth. To minimize heterogeneity and reduce selection bias, we propensity score matched mothers with liveborn and stillborn babies. We selected mothers with stillborn babies where the stillbirth was suspected to have been caused by infection or sepsis, or where fetal death occurred prior to labour onset (e.g. foul smelling/offensive odour, macerated stillbirth, signs of maternal infection). The propensity scoring accounted for mothers’ age, monthly household income, area of residence, type of residence, education status, employment status, and parity. The following exclusions for mothers of liveborn infants were made to ensure extreme phenotyping: no previous stillbirths; any chronic or pregnancy-compromising health condition (e.g. not backache or sore throat). Furthermore, exclusions for both live birth and stillbirth include mothers with diabetes, human immunodeficiency virus (HIV), hyperemesis, hypertension, hypotension, malaria, antepartum haemorrhage, pre-eclampsia, sickle cell, typhoid, vaginal bleeding, and health-related birthing complications (e.g. postpartum haemorrhage). Overall, 19 mothers of stillborn babies were matched to 19 mothers of liveborn babies (Fig. 1 and Table 1).
Figure adapted from “Collection and dispensing of small blood samples for Point-of-Care-Testing” SARSTEDT29.
Demographic description of propensity-matched sample population
We identified mothers of 19 stillborn babies and 19 matched controls. Of the 38 mothers, 7.9% (n = 3) were <20 years old, 5.3% (n = 2) were ≥40 years, and 10% (n = 4) were primigravida. The mothers living in extremely low-income households represented 5.3% (n = 2), 42% (n = 16) reported living in low/very low, with 52.6% (n = 20) living at average or above household income. Those without access to water (defined as anything other than a municipal network) represented 84.2% (n = 32), and 65.8% (n = 25) reported living in shacks. Observing deliveries, 76.3% (n = 29) were full-term (37–42 weeks), 20.9% (n = 8) were pre-term (<37 weeks)20, and 2.6% (n = 1) were post-term (>42 weeks)21. There was no significant variation in gender distribution among babies; among male babies, 57.9% (n = 11) were stillborn and 68.4% (n = 13) were liveborn. Among female babies, 36.8% (n = 7) were stillborn and 31.6% (n = 6) were liveborn (Table 1). While our inclusion criteria allowed for stillbirths associated with infection, signs of maternal infection were present in only 31% of cases (Table 1), representing the heterogeneous aetiology of stillbirth in this population.
Quality assessment of extracted finger-prick RNA and genome-wide expression
RNA yield and quality metrics from the 50 µl Minivette protocol are summarized in Table 1. The RNA concentrations ranged from 2 to 26 ng/µl, with integrity values between 5.5 and 8.0. All samples met the established quality assurance thresholds, confirming their suitability for subsequent microarray transcriptomic analysis.
Raw, unprocessed microarray expression data were first subjected to quality assessment of expression values. For these studies, we applied three different quality control (QC) assurance measures: between-array comparison (distances between arrays), array intensity distributions, and individual array quality (MA plots) using a cut-off of two or more fails as the reason for sample removal. Supplementary Figs. 1–4 show the results of quality assurance measurements and where only one sample failed as part of the checks. Bland–Altman ratio (also referred to as MA) plots are one of the quality measures we apply to quantitatively assess the relationship between log fold change and mean expression values across all samples. Data points that deviate significantly along the vertical axis indicate gene expression values of a given sample with markedly distinct expression levels from the overall mean, although not necessarily differential expression. Typically, genes with lower mean expression values exhibit greater variability in log fold change compared to those with higher expression levels. As a result, we observe a spreading-out effect of data points as we move from right to left on the graph. In Supplementary Fig. 1, we can observe that most of the samples display a very good, well-distributed pattern of data points. Outlier detection was performed by computing Hoeffding’s statistic Da on the joint distribution of A and M for each array. Shown are the arrays for all samples (Live and Still). Only 1 array had Da > 0.15 and was marked as an outlier. Sample S013 (highlighted in red) stands as an outlier and was the only sample to fail three out of the five QC assurance checks.
An investigation of differentially expressed genes (DEGs)
First, we assessed the magnitude and extent of variability of gene expression between the live birth and stillbirth groups. In these analyses, 23,707 probes gave a detectable signal for one or more samples. Of these, 11,354 showed a coefficient of variation of >0.1 across all 37 samples. Next, we filtered low-expression genes to improve power for detecting DEGs, further reducing the number of probes from 11,354 to 6845 (Supplementary Fig. 5).
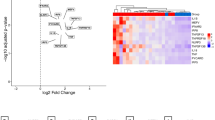
Applying a principal components analysis (PCA) of patient samples based on these 6845 probes and agglomerative unsupervised clustering of normalized unfiltered data did not reveal a separation into live birth and stillbirth groups (Fig. 2a and Supplementary Fig. 4). Unadjusted statistical testing developed 363 differentially expressed probes (P value ≤ 0.05; Fig. 2b). Unsupervised clustering of the top altered unadjusted DEGs showed a separation pattern of live birth and stillbirths (Fig. 2b, c). However, the magnitude of this response was marginal, with only one downregulated gene (SNOR3D) showing an approximate twofold change in expression levels. This gene is a small non-coding nucleolar RNA that has been identified as part of a signature for predicting immunotherapeutic response in human cancer22. Since there are far more variables than observations, it is critical to adjust for multiple hypothesis testing. By adjusting for statistical false discovery and quantitative fold change cut-offs (adjusted P value ≤ 0·05, absolute fold change ≥2), this was reduced to 0 (Fig. 2d). We further applied an independent non-parametric Mann–Whitney test that identified 469 prior to adjustment and that were found to be non-significant after false discovery correction. Figure 3 shows the box–whisker plots of the top ten genes, where nine of these overlap with the top ten DEGs from the parametric testing. These genes show a low-to-moderate correlation (Supplementary Fig. 2).
A Scores plot for PCA analysis of transcriptomic data from Live and Still birth samples (Still – teal and Live – red). B Volcano plot of microarray data (unadjusted p-value). C Hierarchical Clustering Heatmap. D Volcano plot of microarray data (adjusted p-value). Volcano plots Show the differentially expressed genes plotted along dimensions of biological and statistical significance. Genes with the lowest corrected P values are plotted at the top of the graph (e.g., above the line corresponding to P 0.05). Genes overexpressed in Live and Still birth samples are red, respectively (fold-change 2).
Box and Whisker plots for top 10 genes which are statistically significant for livebirth or stillbirth status according to Mann Whitney test.
Additionally, we conducted a focused analysis of 1319 infection and sepsis-related genes (Supplementary Table 3) to determine whether immune-specific signals were obscured in our transcriptome-wide approach. This targeted analysis revealed 76 immune genes with nominal significance (P < 0.05) between stillbirth and live birth groups, with 18 genes demonstrating moderate effect sizes (|fold change| > 0.5). However, consistent with our transcriptome-wide findings, no immune genes achieved statistical significance after multiple testing correction (adj. P > 0.05), and none met stringent effect size thresholds (|fold change| > 1.0) (Supplementary Fig. 6). The identical statistical pattern across both analytical approaches suggests that the absence of significant transcriptomic differences reflects genuine biological and statistical constraints rather than methodological limitations.
Discussion
The transcriptional analysis of maternal blood conducted in this study offers valuable insights into the mother’s immune health status around the time of delivery. We demonstrate the feasibility of using finger-prick transcriptomics in a resource-poor setting to evaluate a mother’s systemic response during labour, comparing mothers who delivered live births and stillbirths. This minimally invasive approach offers significant advantages over traditional venepuncture, including ease of collection, reduced resource requirements, and suitability for community settings. By eliminating the need for centrifugation and immediate storage, finger-prick methodologies enhance scalability and accessibility, making them particularly impactful for low-resource settings.
Blood samples collected from both groups of mothers were successfully transferred to our UK laboratory, maintaining good RNA quality and yield for transcriptomic analysis. While laboratory procedures were centralized for this study, future investigations could adapt these methodologies for local implementation with appropriate resourcing and training. Key challenges include ensuring consistent sample quality in varying environmental conditions, establishing robust cold-chain logistics where necessary, and training personnel in standardized sample collection and handling procedures. Addressing these factors will be crucial for the successful integration of this approach into local healthcare settings. Finger-prick collection is a quick and minimally invasive procedure that can be performed at the bedside or in the community, making it suitable for time-sensitive situations such as labour. However, challenges such as variability in sample volume, potential contamination, and the requirement for careful handling to maintain RNA integrity should be considered to ensure optimal results in diverse settings. Finger-prick approaches for gene expression profiling or whole-genome transcriptomics have several key advantages that help lower barriers for use, although to date, this has not been widely reported in LMICs. Accordingly, we have sought to test the feasibility of deploying a finger-prick methodology in a low-resource setting, with the aim of simplifying the handling and processing of samples at the point of collection. Notably, it can be performed by non-phlebotomists, requires less time and resources, and avoids the need for centrifugation, separation, and immediate storage compared to larger blood samples.
Propensity score matching was employed to control for confounding variables, ensuring comparability between stillbirth cases and live birth controls. Although subgroup sizes for nulliparous women and those with health conditions were small, the diverse cohort reflects real-world populations.
Transcriptional profiling of the systemic immune response to stillbirth has not been reported previously. Against expectations, we found that the maternal systemic transcriptomic immune state did not significantly differ between live birth and stillbirth scenarios, challenging the assumption that intrauterine fetal death would trigger a strong maternal immune response. This finding contrasts with prior studies suggesting systemic inflammatory activation in pregnancy complications and highlights the placenta’s potential role as an immune-regulating organ. It is possible that placental immune tolerance mechanisms suppress maternal responses or that labour-associated immune shifts obscure any stillbirth-specific signatures.
While unsupervised clustering of top-ranked unadjusted DEGs suggested a separation pattern between live birth and stillbirth groups, no individual gene was significant after correction for multiple comparisons. This suggests that the observed separation may be driven by subtle, distributed effects across many genes rather than large effects in individual genes. Given the small sample size and the high dimensionality of transcriptomic data, this pattern could also reflect chance variation or technical artefacts. However, our QC analysis did not reveal systematic technical bias, supporting the interpretation that small, coordinated biological effects may underlie the observed clustering. Further research is needed to dissect these possibilities and determine whether alternative immune markers or functional assays might reveal subtler differences.
Our targeted analysis of infection and sepsis-related genes reinforces the primary finding that maternal systemic immune responses to stillbirth are more subtle than anticipated. The consistent absence of significant transcriptomic differences across both comprehensive and immune-focused approaches supports our interpretation that placental immune tolerance mechanisms may effectively suppress maternal inflammatory responses even in adverse pregnancy outcomes. These findings challenge conventional assumptions about maternal immune activation following fetal death and suggest that the placenta’s role as an immune-regulating organ remains functionally active during labour. The moderate effect sizes observed (0.3–0.7 fold change) are consistent with the tightly regulated nature of pregnancy-related immune adaptations, where dramatic transcriptomic changes might be detrimental to maternal–fetal tolerance. These results highlight the need for larger, adequately powered studies to detect subtle but potentially clinically meaningful immune differences and suggest that alternative approaches such as functional immune assays or single-cell analyses may be required to fully characterize maternal immune responses in stillbirth.
These findings are consistent with the proposed role of the placenta as an immune-regulating organ, acting as a protective barrier between mother and fetus. One explanation could be that the maternal immune system is equipped with protective mechanisms during pregnancy. Alternatively, the immune response to labour may mask subtle transcriptomic changes associated with intrauterine infection or fetal death. Additionally, samples may have been collected before a maternal immune response to fetal death became apparent. While we cannot entirely exclude the inclusion of stillbirths due to non-infectious causes (e.g. hypoxia), efforts were made to enrich the stillbirth cohort with cases more likely associated with infection or sepsis. Signs of maternal and neonatal infection were determined by clinical assessment performed by trained healthcare providers using standardized criteria, including maternal fever, tachycardia, foul-smelling vaginal discharge, and uterine tenderness for maternal infection, and skin discoloration, foul odour, and maceration for neonatal infection.
This study’s limitations include its single-site design and restricted sample size, which may influence the generalizability of the findings and their applicability to broader populations. The single-site focus might limit the variability in environmental or genetic factors captured, while the small sample size reduces statistical power to detect subtle effects. Our comparison included both preterm and term stillbirths compared against term live births, as we did not have sufficient preterm live birth samples for a matched preterm-only analysis. This may potentially mask differences specifically related to gestational age. Reliable data on the precise timing between fetal demise and birth were not available for all cases due to challenges in determining exact time of demise in this setting, which represents another limitation of our study. These limitations emphasize the need for caution when extrapolating results and highlight the importance of multi-site and larger-scale studies in future research.
While our QC measures indicate good RNA preservation across all samples, with consistent RNA Integrity Number (RIN) values between study groups, we acknowledge that subtle RNA degradation could potentially influence our findings. However, the consistent quality metrics across both study groups suggest that the observed lack of major transcriptional differences is unlikely to be solely attributed to technical limitations. After appropriate statistical adjustment for multiple comparisons, no genes were found to be significantly differentially expressed between groups, which precluded performing pathway enrichment analyses such as Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analysis.
Ethical considerations limited our analysis to microarray expression data, which, while beneficial for minimizing technical variation and requiring less input RNA, excludes deeper genomic insights afforded by RNA sequencing or multimodal approaches. Future studies should incorporate larger, multi-cohort designs, RNA sequencing, and single-cell analyses to refine understanding. Our power calculations indicate robust detection capabilities for this setting, but longitudinal studies across pregnancy could capture immune changes beyond labour, providing additional clarity.
In conclusion, this proof-of-concept study provides novel insights into maternal systemic immune responses during labour in live birth and stillbirth contexts. While no significant transcriptional differences were observed, this does not rule out functional differences in specific clinical situations. Our findings highlight the need for further research into systemic, molecular, and cellular mechanisms underlying maternal–fetal immune interactions. We hope this work stimulates further investigation and challenges existing paradigms, ultimately advancing personalized approaches to maternal–fetal health, improved diagnostics, and targeted interventions.
Methods
Setting
This study was conducted in the Murtala Muhammed Specialist Hospital (MMSH), Kano, a tertiary hospital located in northern Nigeria, serving a population of approximately 11 million, and where stillbirths were not being documented at the time of the study. Within the MMSH, there are 17 neonatal intensive care unit beds, 133 maternity beds, and 22 delivery cubicles. Each month, there are around 550 deliveries with 4 midwives on shift at any one time, 2 for complicated deliveries and 2 for uncomplicated deliveries. Preliminary observational work carried out at the MMSH found the incidence of stillbirth to be 180/1000 births.
Study design, ethical approval, informed consent, and procedures
This study is nested as part of a wider feasibility study where the incidence of stillbirth identified was 105/1000 births16. Briefly, we recruited 1998 women, of whom 1789 experienced a live birth and 209 a stillbirth between October 2018 and January 201916. Sample collection was achieved throughout October 2018 from a subset of mothers presenting to MMSH in labour. The mothers were enrolled during labour after informed consent was obtained. An initial pre-tested paper-based questionnaire was completed during labour, and another after delivery, depending on the delivery outcome. This research was performed in accordance with the Declaration of Helsinki, and ethical approval was sought and given by the Health Research Ethics Committee, Kano State of Nigeria Ministry of Health REF: MOH/ Of/797/T.1/950 on 04/09/2018. Mothers were provided with study information in their local dialect, and written informed consent was obtained by trained research nurses.
Exclusion criteria
Participants were excluded from the study if they declined participation, had known HIV infection, or if their samples failed our QC steps. For stillbirth cases, no specific clinical presentation was required for inclusion; cases with and without signs of maternal infection were accepted to represent the diversity of stillbirth aetiologies. Signs of maternal and neonatal infection were determined by clinical assessment performed by trained healthcare providers at MMSH. Standardized criteria for maternal infection included fever (>38 °C), tachycardia, foul-smelling vaginal discharge, and uterine tenderness. Neonatal infection assessment in stillbirths included examination for skin discoloration, foul odour, and the degree of maceration.
Finger-prick collection protocol
In the context of our study, training videos on sample collection were provided to clinical staff at MMSH, and procedures were followed for the collection and processing of blood samples intended for RNA analysis. Briefly, prior to usage, Blood RNA Microfuge Tubes were allowed to equilibrate to room temperature (18–25 °C) and systematically labelled with essential participant information, including unique identifiers and birthdates. The skin surface was disinfected with ethanol prior to procedures involving a finger prick. Thereafter, approximately 50 µl of blood was drawn by capillary action into a Minivette from the site of the finger prick for dispensing to the Blood RNA Tube, vertically positioned within an appropriate rack or tube holder. The Minivette tip was introduced through a hermetic membrane of the tube, and the depressing plunger allowed the collected blood to be judiciously transferred into the Blood RNA tube, after which the Minivette was disengaged from the membrane. Immediately following blood transfer, a gentle manual inversion of the Blood RNA Tube ten times ensued, guaranteeing a comprehensive mixture of the sample. All Blood RNA Tubes were promptly allocated to storage at a temperature of –20 °C within 1–2 h of collection, with an emphasis on maintaining an upright orientation and the use of plastic storage containers to avert potential cracking linked to freezing. Upon the filling of the specified number of tubes, logistical arrangements were initiated for the shipping of these samples on dry ice. Through the execution of these comprehensive protocols, our study ensured the consistent acquisition, treatment, and preservation of blood samples intended for subsequent RNA analysis, thereby safeguarding against contamination and preserving the integrity of these samples throughout the processing and transit phases (Fig. 4).
Finger-prick whole blood transcriptomic study sample flow diagram.
Sample preparation
The samples collected in Africa are frozen to −80 °C and then shipped to the UK on dry ice. On arrival in the UK, the RNA is extracted from the blood and undergoes a sample extraction QC step where the RIN is assessed for quality. Another QC step is performed on the data generated from the microarrays and fails if 2/3 QA steps fail.
Shipped samples were processed for RNA extraction at Cardiff University. Total RNA quality and quantity were assessed using Agilent 4200 TapeStation and a High Sensitivity RNA kit (Agilent Technologies). In all, 2–10 ng of Total RNA with a RIN value >5 was arrayed into a 96-well plate format and sent to Eurofins for processing with the ThermoFisher Clariom D array using the Pico GeneChip labelling kit.
Sample processing: labelling with GeneChip WT Pico Reagent kit
The Affymetrix GeneChip WT Pico kit was used to prep the sample onto the clariom D arrays. First-strand cDNA is synthesized with a combination of a Poly-dT and random primers containing a 5′-adaptor sequence. A 3’-adaptor is added to the single-stranded cDNA followed by low-cycle PCR amplification. The cDNA is used as a template for in vitro transcription that produces amplified amounts of antisense mRNA, (cRNA). The cRNA is then used as input for a second round of first-strand cDNA synthesis, producing single-stranded sense cDNA. After fragmentation and end-labelling, the targets are hybridized to plate arrays, which are stained and imaged on the GeneTitan Multi-Channel Instrument.
Statistics and data analysis methodology
Propensity score matching was used to avoid selection bias and identify likelihood for the best comparable groups without involving the birth outcome. A logistic regression model was fitted to the following demographic characteristics: mothers age, monthly household income, area of residence, education and employment status, and parity.
Quality assurance, normalization, and differential expression analysis
We performed a differential expression analysis using 37 quality-assured finger-prick whole-blood samples from women who had either live birth (n = 19) or stillbirth (n = 18). Expression values were normalized using the robust multi-array average (RMA) method23 implemented in R, and annotated with a NCBI gene name using annotateEset() function from the affycoretools package24 with the Affymetrix clariomdhuman annotation data package clariomdhumantranscriptcluster.db version 8.8.025. Only probesets mapped to a NCBI gene name are retained, and the maximum average intensities are taken over replicate gene names. Mean expression levels were obtained by calculating the geometric means of the RMA-normalized data for live birth and stillbirth groups, respectively. Data was visualized for quality assurance using the R package arrayQualityMetrics. In addition to RMA normalisation, we evaluated potential batch and positional effects, including array row and chip effects, through PCA and visual inspection of QC metrics (Supplementary Figs. 1–3). We observed no systematic batch or positional effects requiring correction. The LIMMA26 package was used, to determine which genes were significantly differentially expressed between the live birth and stillbirth groups. The method implemented in the LIMMA package extends the Empirical Bayes modelling to combine an estimate of variability based on the entire matrix with individual estimates based on each individual value and provides improved error estimates27. The analysis provides the fold change, t-moderated or adjusted P values using the Benjamini–Hochberg procedure28 that are used to order the genes from more to less differentially expressed. As no significantly DEGs were identified after adjustment for multiple comparisons, we did not perform pathway enrichment analyses, including KEGG and GO analysis, as these would not yield meaningful results with the current dataset.
Data availability
Data are provided within the manuscript or Supplementary Information files.
References
Tavares Da Silva, F. et al. Stillbirth: case definition and guidelines for data collection, analysis, and presentation of maternal immunization safety data. Vaccine 34, 6057–6068 (2016).
Hug, L. et al. Global, regional, and national estimates and trends in stillbirths from 2000 to 2019: a systematic assessment. Lancet 398, 772–785 (2021).
McClure, E. M. et al. Stillbirth 2010–2018: a prospective, population-based, multi-country study from the Global Network. Reprod. Health 17, 146 (2020).
Okonofua, F. E. et al. Prevalence and determinants of stillbirth in Nigerian referral hospitals: a multicentre study. BMC Pregnancy Childbirth 19, 533 (2019).
Blencowe, H., Sadoo, S. & Lawn, J. In Oxford Textbook of Global Health of Women, Newborns, Children, and Adolescents (eds Devakumar, D., Hall, J., Qureshi, Z. & Lawn, J.) 140–144 (Oxford University Press, 2018).
Frøen, J. F. et al. Making stillbirths count, making numbers talk - Issues in data collection for stillbirths. BMC Pregnancy Childbirth 9, 58 (2009).
Bhusal, M., Gautam, N., Lim, A. & Tongkumchum, P. Factors associated with stillbirth among pregnant women in Nepal. J. Prev. Med. Public Health 52, 154–160 (2019).
Aminu, M., Bar-Zeev, S., White, S., Mathai, M. & van den Broek, N. Understanding cause of stillbirth: a prospective observational multi-country study from sub-Saharan Africa. BMC Pregnancy Childbirth 19, 470 (2019).
Neogi, S. B. et al. Risk factors for stillbirths: how much can a responsive health system prevent?. BMC Pregnancy Childbirth 18, 33 (2018).
Goldenberg, R. L. et al. Clinical interventions to reduce stillbirths in sub-Saharan Africa: a mathematical model to estimate the potential reduction of stillbirths associated with specific obstetric conditions. BJOG 125, 119–129 (2018).
Lakew, D., Tesfaye, D. & Mekonnen, H. Determinants of stillbirth among women deliveries at Amhara region, Ethiopia. BMC Pregnancy Childbirth 17, 375 (2017).
Ptacek, I., Sebire, N. J., Man, J. A., Brownbill, P. & Heazell, A. E. P. Systematic review of placental pathology reported in association with stillbirth. Placenta 35, 552–562 (2014).
Wicherek, L., Klimek, M., Dutsch-Wicherek, M., Kolodziejski, L. & Skotniczny, K. The molecular changes during placental detachment. Eur. J. Obstet. Gynecol. Reprod. Biol. 125, 171–175 (2006).
Bezemer, R. E. et al. Altered levels of decidual immune cell subsets in fetal growth restriction, stillbirth, and placental pathology. Front. Immunol. 11, 1898 (2020).
Wicherek, L. et al. The placental RCAS1 expression during stillbirth. Reprod. Biol. Endocrinol. 3, 24 (2005).
Milton, R. et al. Incidence and sociodemographic, living environment and maternal health associations with stillbirth in a tertiary healthcare setting in Kano, Northern Nigeria. BMC Pregnancy Childbirth 22, 692 (2022).
Milton, R. et al. Determinants of stillbirth from two observational studies investigating deliveries in Kano, Nigeria. Front. Glob. Womens Health 2, 788157 (2022).
Dudley, D. M. et al. Miscarriage and stillbirth following maternal Zika virus infection in nonhuman primates. Nat. Med. 24, 1104–1107 (2018).
Rinchai, D., Anguiano, E., Nguyen, P. & Chaussabel, D. Finger stick blood collection for gene expression profiling and storage of tempus blood RNA tubes. F1000Res 5, 1385 (2017).
Quinn, J.-A. et al. Preterm birth: case definition & guidelines for data collection, analysis, and presentation of immunisation safety data. Vaccine 34, 6047–6056 (2016).
Caughey, A. B. In Dewhurst’s Textbook of Obstetrics & Gynaecology 269–286 (Wiley-Blackwell, 2012).
Jin, W. et al. Ensemble deep learning enhanced with self-attention for predicting immunotherapeutic responses to cancers. Front. Immunol. 13, 1025330 (2022).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19, 185–93 (2003).
MacDonald, J. W. affycoretools: functions useful for those doing repetitive analyses with Affymetrix GeneChips. R package version 1.81.1. https://bioconductor.org/packages/affycoretools (2023).
MacDonald, J. W. clariomdhumantranscriptcluster.db: Affymetrix Clariom D Human annotation data (chip clariomdhumantranscriptcluster). R package version 8.8.0. Bioconductor. https://org.doi/10.18129/B9.bioc.clariomdhumantranscriptcluster.db (2021).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47 (2015).
Smyth, G. K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 1–25 (2004).
Ferreira, J. A. & Zwinderman, A. H. Approximate power and sample size calculations with the Benjamini-Hochberg method. Int. J. Biostat. 2, 1–36 (2006).
SARSTEDT. Minivette® POCT. https://www.sarstedt.com/en/products/new-products/minivette-poct/ (2025).
Acknowledgements
Firstly, thank you to the mothers and the babies for participating in this research; our heartfelt condolences to those who experienced a stillbirth during this research period. We are so grateful for your participation in this research in such challenging times. Thank you to all at Murtala Mohammed Specialist Hospital, Kano, Nigeria, who made this research possible. We would like to show our appreciation to the Chief Medical Director at the time of the research, Dr Nura Idris, and his successor, Dr Hussaini Mohammed, for supporting and enabling this research. We would like to thank all those involved at the National Hospital Abuja, Nigeria, and the Department of Microbiology, Aminu Kano Teaching Hospital, who made this research possible. We would also like to acknowledge the School of Medicine, Cardiff University, Cardiff, UK, and the Centre for Trials Research, which is funded by Health and Care Research Wales and Cancer Research UK. P.G. was supported by a Ser Cymru Fellowship from the EU-ERDF and Welsh Government throughout this portfolio work. This project was supported using seed funding from Cardiff University’s GCRF QR Funding from the Higher Education Funding Council for Wales. The funder played no role in study design, data collection, analysis and interpretation of data, or the writing of this manuscript.
Author information
Authors and Affiliations
Contributions
Sadly, Professor Peter Ghazal passed away in early April 2024. P.G.’s input and contribution continued up until this point. P.G. contributed to the study design, writing, and development of the manuscript. P.G. reviewed and revised early drafts. P.R.S.R. led the writing of the manuscript, analysed the microarray data, and created the figures. R.M. has contributed to the writing of the manuscript, study design, study management, revision of early drafts, and development of the manuscript. F.M. contributed to study design and delivery. D.G. contributed to the statistical analysis of clinical data, revised early drafts, and developed the manuscript. F.I.A., A.S.M., A.K., F.H.S., F.M.T., R.Y.K. and M.Y.M. conducted the research and data collection. M.B., C.P.E., E.O. and F.J.B. oversaw the delivery of the research. U.U. contributed to the statistical analysis of gene expression and clinical data, developed figures, and contributed to the development of the manuscript. W.J.W. has contributed to the development of the manuscript. S.E. has contributed to the analysis of samples. M.C. developed the extraction of quality RNA from small blood volumes. E.P. assisted with the pre-processing and quality control of the gene expression data. K.C.I. oversaw the delivery of research in Nigeria and has contributed to study design. K.H. contributed to the study design. J.S. and C.S. have contributed to the clinical interpretation of the study. T.R.W. was CI for this study and contributed to the study design. All authors have critically reviewed and approved this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rodrigues, P.R.S., Milton, R., Modibbo, F. et al. Finger-prick transcriptomic profiling in northern Nigeria reveals a muted maternal systemic response in stillbirth. npj Womens Health 4, 8 (2026). https://doi.org/10.1038/s44294-025-00103-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44294-025-00103-w