Abstract
Epigenetic patterns including DNA methylation are known to vary between distantly related species, but it is not clear how these patterns differ at an intraspecific level. The sweetpotato whitefly, Bemisia tabaci (Gennadius) (Aleyrodidae; Hemiptera), encompasses several cryptic species. These cryptic species possess highly similar genomes but exhibit substantial biological and physiological differences. B. tabaci cryptic species are invasive, highly polyphagous, and transmit an array of plant infecting single stranded DNA viruses (ssDNA) –begomoviruses. In this study, DNA methylation patterns around genes and genomic features of two prominent B. tabaci cryptic species were investigated following acquisition of a monopartite ssDNA virus –tomato yellow curl virus. The cryptic species investigated included: B (also known as Middle East Asia Minor 1) and Q (also known as Mediterranean). Genomic features, such as promoters, gene bodies, and transposable elements were assessed for methylation levels in both B and Q cryptic species. Despite overall similar trends, both cryptic species showed differences in methylation levels between these genomic features. Virus induced differentially methylated regions were associated with predominantly distinct genes in B and Q cryptic species. All differentially methylated regions were assessed for differential gene expression and alternative splicing events with and without virus acquisition. DNA methylation levels were found to have a negative correlation with differential gene expression in both B and Q cryptic species. The differentially expressed genes were further grouped into hyper- and hypomethylated clusters. These clusters included genes with implications for virus-vector interactions including immune functions and xenobiotics’ detoxification. The observed DNA methylation pattern differences within each cryptic species could, in part, explain some of the biological and physiological differences between them.
Similar content being viewed by others
Introduction
Epigenetic mechanisms play an important role in the phenotypic diversity and adaptation of eukaryotes to their biotic environments, including interactions with microbes. Among those mechanisms, DNA methylation has emerged as a widely studied epigenetic modification1,2,3,4. These epigenetic modifications are regulated by enzymes known as DNA methyltransferases (DNMTs)5. DNA methylation in eukaryotes alters transcription factor binding and thus directly or indirectly contributes to variation in gene expression, alternative splicing (AS) events, and transposable element (TE) silencing6,7,8,9,10. In the vertebrate genomes, DNA methylation is globally targeted, present in many gene promoters, and enriched in transposable elements11. However, there is currently no robust evidence among insects for DNA methylation enrichment in TEs or in promoters12. Gene regulation in insects is believed to be mediated primarily by intragenic DNA methylation, which in turn is known to influence AS and reduce transcriptional noise10,13. Comparison of major insect groups revealed that DNA methylation occurs in a larger proportion of the genomes of hemimetabolous insect orders such as Hemiptera than in holometabolous insect orders1,14. Global methylation patterns following interactions with pathogens also may vary across insect orders and may relate to the evolution of insect immune responses15,16.
The sweetpotato whitefly, Bemisia tabaci (Gennadius) (family Aleyrodidae; order Hemiptera), is an agricultural pest with global importance and serves as a vector for over 300 diverse plant pathogenic viruses including single stranded DNA (ssDNA) viruses belonging to the genus Begomovirus17. Begomoviruses cause significant losses to global food production18,19. These viruses have high mutation rates and exist as quasispecies with variant genomes within a population20. For this investigation, a common monopartite Begomovirus viz., tomato yellow leaf curl virus (TYLCV) was chosen. TYLCV is transmitted by B. tabaci in a persistent-circulative, non-propogative manner21,22,23. Viruses transmitted in this manner are typically acquired by phloem-feeding insects within several hours of feeding, and the acquired viruses persists within insects for life24,25,26,27. The virus upon acquisition traverses between membranes at the midgut into the hemolymph and then from the hemolymph into the salivary glands via highly specific receptor-mediated endocytosis28,29. The viruses that traverse into the salivary glands can then be reinoculated into susceptible plants via feeding28,29. There also is a latent period between virus acquisition and inoculation24,25,26,27. The process of TYLCV transmission involves several complex interactions between the virus and whitefly proteins30, and it is not clear if these complex interactions would have any effect on DNA methylation.
B. tabaci species complex comprises 5 to 34 different cryptic species with each exhibiting unique biological and physiological characteristics31,32,33,34,35. Notable members of this species complex include the B cryptic species, which also is referred to as Middle East Asia Minor 1, and the Q cryptic species, which also is referred to as Mediterranean (MED)36,37. Both B and Q, as well as the Sub-Saharan Africa-East and Central Africa (SSA-ECA) cryptic species have genomic resources available38,39,40,41,42,43. B and Q cryptic species were selected for comparative analysis in this study, as they are both present in the United States24,44,45,46. The B cryptic species is widespread, whereas Q cryptic species is a relatively recent introduction into the United States46,47. Both B and Q cryptic species efficiently transmit TYLCV; however, the B cryptic species has been observed to transmit a wider array of begomoviruses than the Q cryptic species48.
Differences in virus transmission capabilities between the cryptic species, as consequences of differential interactions of cryptic species with begomoviruses49, could variably impact their DNA methylation profiles and defenses against the virus. The global and gene level DNA methylation patterns were profiled in response to virus infection in B and Q cryptic species to shed light on potential evolutionary variations in gene regulation between the cryptic species of B. tabaci. DNA methylation patterns within whitefly TEs were investigated to contribute to further describing the genetic landscape of B. tabaci50,51,52,53. The relationship between differential methylation, differential expression, and AS was assessed following virus acquisition27. Protein clusters (gene families) were characterized to assess the functions associated with virus response related to differential methylation and differential expression. Several candidate protein clusters related to phytovirus acquisition and retention were identified.
Results
DNA methylation in proximity to genomic features
Global DNA methylation patterns were assessed within the B. tabaci B and Q cryptic species genomes (~650 Mb), which had assemblies with Benchmarking Universal Single Copy Orthologs (BUSCO) scores of 94.6% and 93.4% completeness, respectively38,41,54,55. To explore the DNA methylation landscape in these whitefly cryptic species, this study analyzed the global average methylation levels and the percentage of 5-methylcytosine (5mC) in the CpG (Cytosine-phosphate-Guanine) context within the coding DNA sequences (CDS) of their genomes. Global DNA methylation levels were comparable between both B and Q cryptic species. The non-viruliferous (n = 5 for both cryptic species) and viruliferous (n = 5 for both cryptic species) groups showed relative consistency of global DNA methylation patterns in both whitefly cryptic species (Additional file 1: Fig. S1). Spearman’s rank correlation coefficient (rho) computations were found to be ~0.40 and ~0.60 in B and Q cryptic species, respectively, from all pairwise comparisons of CpG sites of non-viruliferous and viruliferous samples, indicating overall global similarities in DNA methylation (Additional file 1: Fig. S2 & S3). In both cryptic species, expected methylation patterns were observed around the CDS and within genomic features such as promoters, 5’- untranslated regions (UTRs), exons, 3’-UTRs, and introns (Fig. 1a–d; Additional file 2: Tables S1–S6; Additional file 3: Tables S7–S12). Kolmogorov-Smirnov tests were statistically significant with respect to methylation levels of the genomic features, indicating that the data were not normally distributed (B cryptic species D = 0.985, P < 2.2e−16; Q cryptic species D = 0.970, P < 2.2e−16).The promoter regions as well as the 5’-UTR had low methylation levels compared with exon and intron regions. Pairwise statistically significant differences, via Wilcoxon signed-rank tests, were observed between all genomic features in both cryptic species (Additional file 1: Table S13) and between non-viruliferous and viruliferous groups within genomic features in the Q cryptic species (Fig. 1d). Using Wilcoxon signed-rank tests, the Q cryptic species was found to exhibit slightly, but significantly lower levels of CDS, promoter, 5’-UTR, and 3’-UTR methylation compared with the B cryptic species (P < 2.2e−16).
B cryptic species methylation patterns within (a) the coding sequences (CDS) and 5 kb upstream and downstream of the translation start site (TSS) and translation termination site (TTS), respectively, and (b) the promoters, 5’-UTRs, exons, introns, and 3’-UTRs. Q cryptic species methylation pattern within (c) the CDS and 5 kb upstream and downstream of the TSS and TTS, respectively, and (d) the promoters, 5’-UTRs, exons, introns, and 3’-UTRs. B cryptic species methylation patterns within (e) the transposable elements (TEs) and 5 kb upstream and downstream of the TSS and TTS, respectively, and (f) various transposable element superfamilies. Q cryptic species methylation pattern within (g) the TEs and 5 kb upstream and downstream of the TSS and TTS, respectively, and (h) various transposable element superfamilies. Comparisons between non-viruliferous and viruliferous groups were conducted via Wilcoxon signed-rank tests. Statistical significance levels are indicated by asterisks in the figure: ns (not significant) for p > 0.05, * for p ≤ 0.05, ** for p ≤ 0.01, *** for p ≤ 0.001, and **** for p ≤ 0.0001.
DNA methylation was next considered in proximity to TEs in B. tabaci. TE counts associated with both cryptic species were enumerated (Additional file 1: Table S14). The global TE methylation patterns were below the levels observed in flanking regions, indicating that TEs were depleted for DNA methylation overall (Fig. 1e–h; Additional file 4: Tables S15 and S16). Kolmogorov-Smirnov tests were statistically significant with respect to methylation levels of the TEs, indicating the data were not normally distributed (B cryptic species D = 0.977, P < 2.2e−16; Q cryptic species D = 0.949, P < 2.2e−16). The TE methylation levels were found to be similar between the two cryptic species. Nevertheless, variations were observed between TE types, for example, the copia retrotransposons were the most highly methylated superfamily of TEs (Fig. 1f, h). Other superfamilies, such as gypsy retrotransposons, also were found to be methylated above background (Fig. 1f, h). Using pairwise Wilcoxon signed-rank tests, the methylation levels of copia and gypsy retrotransposons were found to have a statistically significant difference in comparison with all other TE superfamilies in both cryptic species (Additional file 1: Tables S17). However, there were no statistically significant differences between methylation levels of non-viruliferous and viruliferous groups in copia or gypsy retrotransposons (Fig. 1f, h).
The methylation patterns were compared with the CpG observed-to-expected (CpGo/e) ratio, which assesses the relative abundance of CpG sites56. Regions with low CpGo/e may indicate that a given site could potentially be DNA methylated. Bimodality in DNA methylation and CpGo/e ratios in the promoter regions was noticed in both cryptic species (Fig. 2a, c) and gene bodies (Fig. 2b, d; Additional file 5: Tables S18 and S19). The comparison of promoter and gene body methylation levels revealed clusters at varying ranges (Fig. 2e, f). Promoter and gene body methylation levels were found to be positively correlated with one another in both cryptic species (B cryptic species Spearman’s rho = 0.545, P < 2.2e−16; Q cryptic species Spearman’s rho = 0.676, P < 2.2e−16). Additionally, CpGo/e and methylation levels in both the promoter and gene body were found to be positively correlated in both cryptic species (Additional file 1: Fig. S4a–d).
a The CpGo/e comparison reveals for B and Q, 4190 and 4636 promoters, respectively, were below 0.8 (indicated by the dotted line). b The CpGo/e comparison reveals that for B and Q cryptic species, 7654 and 7816 gene bodies, respectively, were below 0.8 (indicated by the dotted line). c The methylation level analysis shows that for B and Q cryptic species, 5059 and 5353 promoters, respectively, exceeded 20% (marked by the dotted line). d The methylation level analysis shows that for B and Q cryptic species, 7580 and 8240 gene bodies, respectively, exceeded 20% (marked by the dotted line). Contour density plots showing the relationship between gene body methylation levels (x-axis) and promoter methylation levels (y-axis) of (e) B and (f) Q cryptic species. Contour density lines show gene body methylation levels represented by different line types, categorized into distinct, non-overlapping intervals: 0.0–0.2, 0.2–0.4, 0.4–0.6, 0.6–0.8, and 0.8–1.0.
Identification of differentially methylated regions in response to virus acquisition
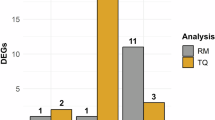
Differentially methylated regions (DMRs) were investigated in response to virus acquisition in B. tabaci B and Q cryptic species using a 1 kb sliding window approach. Several potential candidate sites with significant differences in methylation were detected (Additional file 6: Tables S20 and S21). Genes within 1.5 kb of a DMR were classified as differentially methylated. In B cryptic species, 57 genes were identified with DMRs (Additional file 6: Table S22; Top 20 genes can be found in Table 1). Similarly, in Q cryptic species, 58 genes were identified with DMRs (Additional file 6: Table S23; Top 20 genes can be found in Table 2).
Comparison of DNA methylation, gene expression, and alternative splicing in relation to virus acquisition
This study focused on determining the methylation levels concerning virus acquisition in B. tabaci B and Q cryptic species. In general, higher DNA methylation levels were found to be associated with lower differential expression45. The expression analysis in B cryptic species identified a total of 328 differentially expressed genes (DEGs) (Additional file 7: Table S24), and in Q cryptic species, a total of 1616 DEGs were identified (Additional file 7: Table S25). Both B and Q cryptic species showed a non-monotonic association between DNA methylation and gene expression quantiles (Fig. 3a, c). Kruskal-Wallis tests were conducted to examine the methylation level differences among quantiles in both cryptic species (B cryptic species Chi-square = 287, P < 2.2e−16, df = 9; Q cryptic species Chi-square = 826, P < 2.2e−16, df = 9). Pairwise Wilcoxon signed-rank tests also showed statistically significant differences between quantiles for both B and Q cryptic species (Additional file 1: Tables S26). Regardless of the direction of expression, differentially expressed genes were more often hypomethylated than hypermethylated in both B and Q cryptic species, and significant negative linear correlations (B cryptic species Spearman’s rho = −0.62, P < 2.2e−16; Q cryptic species Spearman’s rho = −0.48, P < 2.2e−16) were observed between methylation levels and the degree of differential gene expression (Fig. 3b, d).
a Expression quantiles in relation to methylation levels in B cryptic species. b The absolute Log2 Fold Change (LFC) of differential expression in relation to methylation levels in B cryptic species. c Expression quantiles in relation to methylation levels in Q cryptic species. d The absolute LFC of differential expression in relation to methylation levels in Q cryptic species. Comparisons between non-viruliferous and viruliferous groups were conducted via Wilcoxon signed-rank tests. Statistical significance levels are indicated by asterisks in the figure: ns (not significant) for p > 0.05, * for p ≤ 0.05, ** for p ≤ 0.01, *** for p ≤ 0.001, and **** for p ≤ 0.0001.
The association between DNA methylation, gene expression, and AS events (Additional file 8: Tables S27 and S28) was investigated in both whitefly cryptic species. The promoters and gene bodies showed similar methylation patterns in relation to differential expression and in relation to AS events for both cryptic species (Fig. 4; Additional file 1: Table S29). Significant differences between promoter and gene body methylation were found when methylation was below 0.2 in B cryptic species (Fig. 4a) and below 0.8 in Q cryptic species (Fig. 4c). There were no significant differences between promoter and gene body methylation in relation to AS in both cryptic species; however, there was a significant difference between methylation levels 0.2–0.4 and 0.4–0.6 as well as 0.2–0.4 and 0.6–0.8 in B cryptic species (Fig. 4b). Also, there were significant differences between methylation levels 0.2–0.4 and high methylation levels 0.8–1.0 in relation to AS in Q cryptic species (Fig. 4d).
The B cryptic species methylation levels of promoters and gene bodies corresponding to (a) gene expression and (b) alternative splicing. The Q cryptic species methylation levels of promoters and gene bodies corresponding to (c) gene expression and (d) alternative splicing. Counts of genes can be found under their respective boxes. Comparisons between non-viruliferous and viruliferous groups were conducted via Wilcoxon signed-rank tests. Statistical significance levels are indicated by asterisks in the figure: ns (not significant) for p > 0.05, * for p ≤ 0.05, ** for p ≤ 0.01, *** for p ≤ 0.001, and **** for p ≤ 0.0001.
Differential methylation was compared with differential expression and AS events (Additional file 9: Tables S30 and S31). The genes with differential methylation, differential expression, and AS were compared in cryptic species B (Fig. 5a), Q (Fig. 5b), and orthologs between them (Fig. 5c). The analysis revealed that there were low numbers of overlapping genes showing differential methylation and differential expression in both cryptic species, which was found to be insignificant. The overlap of differential methylation and AS in Q also was not found to be significant. When comparing single copy orthologous genes between the two genome assemblies (Additional file 9: Table S32), a high number of AS events was found (Fig. 5c). However, no single copy orthologs were found to be both differentially methylated and alternatively spliced.
Similar distributions of overlapping and independent genomic events can be seen in the (a) B cryptic species (# of genes = 2503), (b) Q cryptic species (# of genes = 5069), and (c) one-to-one orthologs (# of genes = 357).
Protein clustering of DMRs and DEGs
A within- and between-cryptic species gene family approach was taken to characterize the function of genes that may be functionally similar but display divergence in DEG and/or DMR patterns. Five gene families containing 10 genes were found to be exclusive to the B cryptic species, 183 gene families containing 266 genes were found to be exclusive to the Q cryptic species, and 102 gene families containing 483 genes were found in both cryptic species (Fig. 6; Additional file 1: Figs. S5a, S5b, S6). This approach provided a comprehensive view of how genes with similar functions are collectively regulated in response to virus acquisition. By examining the Gene Ontology (GO) terms linked with gene families displaying varying methylation and expression patterns (Additional file 9: Table S33), the functional roles of these proteins within the hierarchical GO categorization were determined. Protein clusters (gene families) meeting the minimum inclusion criteria, being that the gene family contained at least one differentially methylated gene from B and/or Q cryptic species, were explored further for possible associations with virus acquisition.
Upset plot (subset includes only groups with at least one methylated set) of gene families within Bemisia tabaci B and Q cryptic species relating to the numbers of differentially expressed genes (high and low expression; FDR < 0.05) and differentially methylated genes (hyper- and hypomethylated; q < 0.05).
The most frequent GO terms, associated with clusters meeting the minimum inclusion criteria were investigated, as they may be related to virus acquisiton. These GO terms were: “DNA integration” (GO:0015074; n = 3 clusters) and “transposition, DNA-mediated” (GO:0006313; n = 2 clusters). Clusters 7, 163, and 172 were characterized as being associated with the GO term “DNA integration” (GO:0015074). However, begomoviruses such as TYLCV are not expected to integrate into vector or host genomes57. Clusters 18 and 72 were characterized as being associated with the GO term “DNA-mediated transposition” (GO:0006313), which relates to the movement of transposable elements from one part of the genome to another58. The observed changes in methylation and gene expression in clusters 18 and 72 suggest their potential roles in modulating transposon activity during virus acquisition.
Additional protein clusters, meeting the minimum inclusion criteria, with unique GO terms were investigated. Cluster 31 was characterized as a Biological Process (BP) associated with the GO term “translation” (GO:0006412), suggesting its involvement in protein synthesis, which may relate to virus acquisition59. Cluster 71 was characterized as a BP associated with the GO term “regulation of protein tyrosine kinase activity” (GO:0061097), which is involved in modulating the activity of tyrosine kinase and indirectly relates to antiviral activity60,61,62. The observed changes in gene expression and DNA methylation in clusters 31 and 71 point to their potential roles in influencing cellular signaling pathways and regulating protein production during virus acquisition, respectively.
Cluster 168 was characterized as a BP associated with the GO term “protein localization to nuclear pore” (GO:0090204), indicating its involvement in mediating protein transport to the nuclear pore complex63. The observed changes in DNA methylation and gene expression in cluster 168 suggest its potential role in regulating nuclear import and export during virus acquisition. Cluster 170 was characterized as a BP associated with the GO term “ubiquitin-dependent ERAD pathway” (GO:0030433), suggesting its potential involvement in ER-associated protein degradation via the ubiquitin-proteasome system64. The observed changes in DNA methylation and gene expression in cluster 170 indicate its potential role in regulating protein degradation following virus acquisition. Cluster 171 was characterized as a BP associated with the GO term “spliceosomal snRNP assembly” (GO:0000387), indicating its involvement in the assembly of spliceosomal small nuclear ribonucleoproteins65.
Discussion
Epigenetic regulation is known to shape the genetic landscape of organisms at varying intensities. This study examined the influence of DNA methylation on two cryptic species (B and Q) of B. tabaci36,37. These two cryptic species are of particular interest due to their differences in biological traits, behavior, xenobiotics’ detoxification (including insecticides used in agricultural landscape), and virus transmission abilities31,32. Both B and Q cryptic species of B. tabaci exist in the Southeastern United States and have previously been shown to be efficient transmitters of the monopartite ssDNA virus –TYLCV24,44. Closely related species can exhibit distinct DNA methylation patterns, such as the observed differences between species within genus Triportheus (Characiformes fishes)66. However, intraspecific differences, such as those in the B. tabaci cryptic species complex, have not been widely investigated.
Previous studies have shown that insects exhibit significantly lower levels of DNA methylation compared with mammals, and the distribution of DNA methylation across the genome also is variable67. Low genomic DNA methylation in B. tabaci was expected based on other previous studies14,68. Genes frequently subjected to methylation tend be more highly conserved across invertebrate species, indicating slower evolutionary rates and stronger selective constraint69. Hypermethylated regions tend to be found within proximity to more highly expressed genes such as housekeeping genes70,71,72,73,74. In this study, promoter and gene body methylation levels were compared between B and Q cryptic species. The methylation levels were observed to have a similar pattern of bimodality within both cryptic species. However, this bimodality of the promoter region is not common among invertebrates75, suggesting that B. tabaci might exhibit unique patterns of promoter methylation. Bimodal promoter methylation has been studied extensively in vertebrate systems, where genes with high DNA methylation levels have been found to be more conserved than those with low DNA methylation levels76,77. The methylation of promoters in mammalian systems may serve to reduce the activity of TEs and repress transcription activity78. High DNA methylation in promoter regions of B. tabaci may be related to stress responses or other functionally significant events, similar to the promoter DNA methylation patterns observed in Ciona savignyi (Herdman)79. In B. tabaci, promoters that were lowly methylated were generally associated with metabolism, signaling, structural, and transport proteins. It also is worth acknowledging that promoter regions’ annotations are dependent on completeness and accuracy of their respective gene body annotations.
Low methylation levels also were observed across most TE superfamilies. However, copia retrotransposons, one of the largest TE superfamilies in eukaryotes80,81, were found to be targeted more by DNA methylation than other TE superfamilies. Sequence homology between copia retrotransposons and the promoter regions of heat shock proteins (HSPs) may indicate that this class of TEs could contribute towards whitefly’s response to biotic stressors such as pathogens82,83. HSPs, broadly, are involved in homeostasis and immune response to phytoviruses in several insects including B. tabaci84. While there was a low number of copia retrotransposons found within the genome assemblies of both cryptic species, these TEs also have been observed to be preferentially methylated and accumulate in regions of high CpG density in other arthropod species50,51. Members of another major TE superfamily found in eukaryotes, known as gypsy retrotransposons80,81, were also found to be likely targets of DNA methylation compared with other TE superfamilies in B. tabaci. Gypsy retrotransposons activation has been found to be involved in Drosophila development and immune responses85. DNA methylation patterns also were altered in other TE types following virus acquisition within the following TE superfamilies: CACTA TIR transposons, hAT TIR transposons, Mutator TIR transposons, and LTR retrotransposons.
The relationship between DNA methylation and gene expression can vary among different insect species and can be context dependent1. Previously reported transcript-derived expression differences in response to virus acquisition in B and Q cryptic species of B. tabaci45 were compared to the methylation events. The aim was to uncover key methylation patterns affecting virus acquisition and subsequent inoculation in both B. tabaci cryptic species. Gene expression levels could potentially be influenced by DNA methylation; however, differential promoter methylation was not found to be associated with gene expression differences in either cryptic species in the current study.
Between the two cryptic species, two pairs of orthologs were identified where members from B and Q each exhibited DMRs, whereas 55 and 56 genes exhibited DMRs in only B or Q cryptic species, respectively. In the B cryptic species, genes with DMRs such as Bta00119 (Tyrosine-protein kinase) suggests potential immune signaling pathways in response to virus acquisition86. In the Q cryptic species, genes with DMRs such as BTA018500.1 (40 S ribosomal protein S28) and may relate to regulation of apoptosis following virus acquisition87. The first pair of orthologs with DMRs were: Bta03757 (Brinker) in B cryptic species and BTA018959.1 (unnamed protein product) in Q cryptic species. The second pair that had genes with DMRs and were also found to be DEGs viz., Bta06672 (Threonine--tRNA ligase) in B cryptic species and BTA026677.3 (39 S ribosomal protein L39, mitochondrial) in Q cryptic species. Threonine--tRNA ligase, which plays an important role in protein biosynthesis, has been previously identified as a candidate for pest control via RNA interference (RNAi) in three rice planthopper species88. These distinct genes could indicate differences in biological and physiological responses of the two cryptic species. Additionally, differential methylation may be weakly related to the presence of AS events in B. tabaci.
The integration of gene expression analysis with the identification of specific protein clusters associated with GO terms provided valuable insights into B. tabaci-virus interactions. Protein clusters (gene families) composed of similar genes and enriched in GO terms such as “DNA integration”, “transposition, DNA-mediated”, “regulation of protein tyrosine kinase activity”, “protein localization to nuclear pore”, “ubiquitin-dependent ERAD pathway”, and “spliceosomal snRNP assembly” are of particular interest mainly because of their relevance to phytovirus acquisition and retention.
The low number of genes with DMRs were expected by random chance, suggesting a lack of conservation of site-specific DNA methylation. As this work delves into the epigenetic regulation of gene expression in B. tabaci and its modulation by virus acquisition, it contributes to a broader understanding of insect-vector interactions and the molecular basis of vector-borne pathogen dynamics. This study provides an overview of baseline DNA methylation levels in B. tabaci B and Q cryptic species, as well as shows that non-propagative viruses may have minimal, but differential, impact on the global or targeted methylation patterns of their insect vector. It also may be of interest to look at temporal differences in DNA methylation following acquisition of other persistently transmitted viruses in the two cryptic species. Beyond DNA methylation, the study of other forms of epigenetic alterations in B. tabaci, such as histone modifications and non-coding RNAs (ncRNAs), may further elucidate the intricate epigenetic regulation underlying their response to virus acquisition.
Methods
Bemisia tabaci rearing and TYLCV acquisition
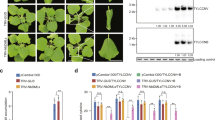
Both B and Q cryptic species were reared in separate greenhouses on non-infected tomato, Solanum lycopersicum cv. Florida 47 (Seminis Vegetable Seeds, St. Louis, MO, USA) plants, for two generations according to protocols outlined in ref. 48. Newly emerged adults were collected and released on TYLCV-infected and non-infected tomato plants for a 72-h acquisition access period (AAP). TYLCV was maintained on tomato plants via B. tabaci-mediated inoculation. After 72 h, specimens were collected by removal of leaf tissue. The excised tissues were dipped into a falcon tube to release the whiteflies. The collected specimens were frozen and stored at −20 °C. This set up was replicated five times per each treatment (non-viruliferous and virulifeorus) for both B and Q cryptic species. Single viruliferous adults tested via endpoint PCR has revealed that 90–100% were positive for TYLCV following acquisition45,89.
DNA extraction and confirmation of TYLCV acquisition
Total DNA was extracted from 30 females from each replicate and homogenized using Cetyltrimethylammonium bromide90. The B and Q cryptic species and virus acquisition status was confirmed by use of PCR. The specific cryptic species was confirmed with the use of BEM23 primers at an annealing temperature of 55 °C—B cryptic species = 200 bp and Q cryptic species = 400 bp91. The presence of TYLCV was confirmed with V2 primers, which encode for the pre-coat protein92. Quality control was performed with the use of NanoDrop® Microvolume Spectrophotometer and Qubit® Fluorometer (ThermoFisher Scientific, Waltham, MA, USA).
Library preparation and sequencing
The extracted DNA was sent to Novogene for bisulfite conversion sequencing. Upon amplification, those sites were read as thiamine and were sequenced with an Illumina HiSeq X to produce short read data. The quality of the raw sequence data was checked with FastQC v0.11.9 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) on a per sample basis. Sequencing information derived from FastQC for all samples was organized and visualized using the multiQC v1.11 tool93.
Sequence data processing
Adapter contamination of raw reads was removed via Trimmomatic v0.3994 with the following parameters: PE, -phred33, ILLUMINACLIP: NexteraPE-PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36. Following the trimming step, the sequence quality was checked again with FastQC. Sequencing information from all samples was organized and visualized using multiQC93. This was done to ensure that the adapters have been removed and that only high-quality reads remained. The overall bioinformatics pipeline is shown in Additional file 1: Fig. S7, where the first steps are quality control and trimming of the raw reads. Genome quality was checked with the use of BUSCO v4.0.6 using the Insecta odb10 lineage54. The lineage to measure the quality of B. tabaci genomes was Insecta odb10, which is a set of 1367 core genes from over 70 insect species and eight insect orders.
High quality trimmed reads were mapped to the respective genomes from B. tabaci B and Q cryptic species with the use of Bowtie2 v2.4.195, which was built into Bismarck v0.23.096. The read alignments can be found in Additional file 1: Tables S34 and S35. With bisulfite-converted reads, unmethylated cytosines were converted to uracil. When mapping the reads, methylated cytosines could only map to the reference cytosines, but improper mapping due to single nucleotide polymorphisms were considered by Bowtie2. Cytosines were considered methylated if they were above the methylation difference threshold of five. Thiamine sites in bisulfite-converted reads can align to the reference cytosines; however, the reverse is not true, this mapping asymmetry provided the basis for determining methylated cytosine pileup97. The set of raw reads was deduplicated to eliminate the potential PCR contamination.
Methylation status
As a part of the bioinformatics pipeline, the amount of cytosine in CpG, CHG, and CHH context was determined by Bismarck. Investigation of differential methylation in various genomics regions of B. tabaci genome was conducted. The read alignments can be found in Additional file 1: Tables S36 and S37. The R v4.1.0 (https://www.r-project.org/contributors.html) tool to determine differential methylation was methylKit98, that implements the Dispersion Shrinkage for Sequencing (DSS) methylation calling software99 for determining regions with methylated sites. A sliding window approach was used, as detection of a signal from any individual site was too weak. The Bismark reports were filtered with the methylKit function filterByCoverage() using the parameters: lo.count = 10, lo.perc = NULL, hi.count = NULL, hi.perc = 99.9, pipeline = “bismarkCytosineReport”. The filtered Bismark reports were tiled with the methylKit function tileMethylCounts() using the following parameters: win.size = 1000, step.size = 1000, cov.bases = 5, mc.cores = 24, pipeline = “bismarkCytosineReport”. A comparison of the mapped reads from non-viruliferous and viruliferous B. tabaci adults of both cryptic species was generated. The threshold for methylation differential and the qvalue was 10 and 0.05, respectively. Using custom R script (found in file “Figure_3.R” in the Dryad repository https://doi.org/10.5061/dryad.jdfn2z3jn), genes were called if they were within 1.5 kb of a methylated region. SeqMonk (https://www.bioinformatics.babraham.ac.uk/projects/seqmonk/) was used to view the methylation patterns 5 kb upstream and downstream of within both the B and Q cryptic species genomes as well as within various genomic features. The CpG observed/expected (CpGo/e), which serves as an approximation for the number of cytosines in CpG context derived from the evolutionary buildup of methylated cytosines was determined for coding sequences using Notos100. Additionally, transposable element determination was conducted with Extensive De Novo Annotator (EDTA) v2.1.0101,102.
Transcript expression and alternative splicing events
Previously published RNA sequence data relating to TYLCV-acquisition45 were used for downstream analysis. Associated RNA extraction protocols and sequencing methods can be found in ref. 45. Adapter contamination of raw reads was removed via Trimmomatic v0.3994 with the following parameters: PE, -phred33, ILLUMINACLIP: NexteraPE-PE.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36. Trimmed reads were mapped to the respective B and Q cryptic species genomes by using the program Spliced Transcripts Alignment to Reference (STAR) v2.7.10a103 with the following parameters: --outFilterType BySJout --outSAMattributes NH HI AS NM MD --outFilterMultimapNmax 20 --outFilterMismatchNmax 999 --outFilterMismatchNoverLmax 0.04 --alignIntronMin 20 --alignIntronMax 1000000 --alignMatesGapMax 1000000 --alignSJoverhangMin 8 --alignSJDBoverhangMin 1 --sjdbScore 1. Mapped read counts were quantitated with the use of RNA-Seq by Expectation Maximization (RSEM) v1.3.3104. DESeq2 was used to determine the expression values of the mapped reads, with a false discovery rate (FDR) < 0.05105. The differential exon usage of STAR mapped reads were determined by using DEXSeq106. The genomic format files were processed using Another Gtf/Gff Analysis Toolkit (AGAT) v0.8.1 (https://zenodo.org/record/7950165).
GO-Term analysis of DMR and DEG protein clusters
Gene family orthologs between methylated genes and differentially expressed genes were determined using OrthoVenn2107 and OrthoFinder2 v2.5.2108,109,110 with setting -M msa to specify the usage of the multiple alignment using fast Fourier transformation (MAFFT) tool111. Alignment statistics were determined with Clustal Omega112 and the European Molecular Biology Open Software Suite (EMBOSS) infoalign113. The gene families were visualized using the R package ComplexHeatmap v3.17114.
Additional statistical analyses
Statistical analyses were conducted using R v4.1.0, employing the following tests as appropriate: the Kolmogorov-Smirnov test to assess normality or skewness of the data, the Kruskal-Wallis test for non-parametric comparisons among multiple independent groups, the Wilcoxon signed-rank test for non-parametric comparisons between two means within a single group, and Spearman’s rank correlation for assessing monotonic relationships.
LLM technology implementation and assistance
During the preparation of this work, the authors minimally used ChatGPT 3.5 for assistance with coding and editing. After using this tool/service, the authors reviewed and edited the content as needed and take full responsibility for the content of the publication.
Data availability
The bisulfite sequence data generated in this study have been submitted to the National Center for Biotechnology Information (NCBI; https://www.ncbi.nlm.nih.gov/). The data can be accessed under the BioProject accession number PRJNA813453, with BioSamples accession numbers SAMN26503470, SAMN26503469, SAMN26503468, and SAMN26503371. The corresponding Sequence Read Archive (SRA) accession numbers are SRR18266360 through SRR18266379. Supplementary information, additional files, and software can be found in the Dryad repository https://doi.org/10.5061/dryad.jdfn2z3jn. Additional file 1: Tables S1–S37, Figs. S1–S7. Respective tables can be found in Additional file 2: Tables S1–S6; Additional file 3: Tables S7–S12; Additional file 4: Tables S15 and S16; Additional file 5: Tables S18 and S19; Additional file 6: Tables S20–S23; Additional file 7: Tables S24 and S25; Additional file 8: Tables S27 and S28; Additional file 9: Tables S30–S33.
References
Bewick, A. J., Vogel, K. J., Moore, A. J. & Schmitz, R. J. Evolution of DNA methylation across insects. Mol. Biol. Evol. 34, 654–665 (2017).
Bewick, A. J. et al. Diversity of cytosine methylation across the fungal tree of life. Nat. Ecol. Evol. 3, 479–490 (2019).
Zhang, H., Lang, Z. & Zhu, J.-K. Dynamics and function of DNA methylation in plants. Nat. Rev. Mol. Cell Biol. 19, 489–506 (2018).
Arneson, A. et al. A mammalian methylation array for profiling methylation levels at conserved sequences. Nat. Commun. 13, 783 (2022).
Klose, R. J. & Bird, A. P. Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci. 31, 89–97 (2006).
Zhu, H., Wang, G. & Qian, J. Transcription factors as readers and effectors of DNA methylation. Nat. Rev. Genet 17, 551–565 (2016).
Flores, K. et al. Genome-wide association between DNA methylation and alternative splicing in an invertebrate. BMC Genom. 13, 480 (2012).
Glastad, K. M., Gokhale, K., Liebig, J. & Goodisman, M. A. The caste- and sex-specific DNA methylome of the termite Zootermopsis nevadensis. Sci. Rep. 6, 37110 (2016).
Yin, Y. et al. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
Shukla, S. et al. CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing. Nature 479, 74–79 (2011).
Zeng, J. et al. Divergent whole-genome methylation maps of human and chimpanzee brains reveal epigenetic basis of human regulatory evolution. Am. J. Hum. Genet. 91, 455–465 (2012).
Glastad, K. M. et al. Epigenetic regulator CoREST controls social behavior in ants. Mol. Cell 77, 338–351.e336 (2020).
Huh, I., Zeng, J., Park, T. & Yi, S. V. DNA methylation and transcriptional noise. Epigenetics Chromatin 6, 9 (2013).
Glastad, K. M., Hunt, B. G. & Goodisman, M. A. D. Epigenetics in insects: genome regulation and the generation of phenotypic diversity. Annu. Rev. Entomol. 64, 185–203 (2019).
Mukherjee, K. & Dobrindt, U. The emerging role of epigenetic mechanisms in insect defense against pathogens. Curr. Opin. Insect Sci. 49, 8–14 (2022).
Mukherjee, K., Dubovskiy, I., Grizanova, E., Lehmann, R. & Vilcinskas, A. Epigenetic mechanisms mediate the experimental evolution of resistance against parasitic fungi in the greater wax moth Galleria mellonella. Sci. Rep. 9, 1626 (2019).
Gilbertson, R. L., Batuman, O., Webster, C. G. & Adkins, S. Role of the insect supervectors Bemisia tabaci and Frankliniella occidentalis in the emergence and global spread of plant viruses. Annu Rev. Virol. 2, 67–93 (2015).
Leke, W. N., Mignouna, D. B., Brown, J. K. & Kvarnheden, A. Begomovirus disease complex: emerging threat to vegetable production systems of West and Central Africa. Agric. Food Secur. 4, https://doi.org/10.1186/s40066-014-0020-2 (2015).
Saurabh, S. et al. Tiny Flies: A mighty pest that threatens agricultural productivity-a case for next-generation control strategies of whiteflies. Insects 12, https://doi.org/10.3390/insects12070585 (2021).
Domingo, E. & Perales, C. Viral quasispecies. PLoS Genet. 15, e1008271 (2019).
Pakkianathan, B. C. et al. Replication of Tomato yellow leaf curl virus in its whitefly vector, Bemisia tabaci. J. Virol. 89, 9791–9803 (2015).
Becker, N. et al. Rapid accumulation and low degradation: key parameters of Tomato yellow leaf curl virus persistence in its insect vector Bemisia tabaci. Sci. Rep. 5, 17696 (2015).
Sanchez-Campos, S. et al. Tomato yellow leaf curl virus: no evidence for replication in the insect vector Bemisia tabaci. Sci. Rep. 6, 30942 (2016).
Rosen, R. et al. Persistent, circulative transmission of begomoviruses by whitefly vectors. Curr. Opin. Virol. 15, 1–8 (2015).
Hogenhout, S. A., Ammar el, D., Whitfield, A. E. & Redinbaugh, M. G. Insect vector interactions with persistently transmitted viruses. Annu. Rev. Phytopathol. 46, 327–359 (2008).
Eigenbrode, S. D., Bosque-Perez, N. A. & Davis, T. S. Insect-borne plant pathogens and their vectors: ecology, evolution, and complex interactions. Annu. Rev. Entomol. 63, 169–191 (2018).
Catto, M. A. et al. A review on transcriptional responses of interactions between insect vectors and plant viruses. Cells 11, 693 (2022).
Stafford, C. A., Walker, G. P. & Ullman, D. E. Hitching a ride: vector feeding and virus transmission. Commun. Integr. Biol. 5, 43–49 (2012).
Zhao, J. et al. A vector whitefly endocytic receptor facilitates the entry of begomoviruses into its midgut cells via binding to virion capsid proteins. PLoS Pathog. 16, e1009053 (2020).
Czosnek, H., Hariton-Shalev, A., Sobol, I., Gorovits, R. & Ghanim, M. The incredible journey of begomoviruses in their whitefly vector. Viruses 9, https://doi.org/10.3390/v9100273 (2017).
Hsieh, C.-H., Wang, C.-H. & Ko, C.-C. Evidence from molecular markers and population genetic analyses suggests recent invasions of the Western North Pacific region by biotypes B and Q of Bemisia tabaci (Gennadius). Environ. Entomol. 36, 952–961 (2007).
Elfekih, S. et al. Genome-wide analyses of the Bemisia tabaci species complex reveal contrasting patterns of admixture and complex demographic histories. PLoS One 13, e0190555 (2018).
MacLeod, N., Canty, R. J. & Polaszek, A. Morphology-based identification of Bemisia tabaci cryptic species puparia via embedded group-contrast convolution neural network analysis. Syst. Biol. 71, 1095–1109 (2022).
Brown, J. K., Paredes-Montero, J. R. & Stocks, I. C. The Bemisia tabaci cryptic (sibling) species group—imperative for a taxonomic reassessment. Curr. Opin. Insect Sci. 57, 101032 (2023).
de Moraes, L. A. et al. Distribution and phylogenetics of whiteflies and their endosymbiont relationships after the Mediterranean species invasion in Brazil. Sci. Rep. 8, 14589 (2018).
Hamon, A. & Salguero, V. Bemisia tabaci, Sweetpotato Whitefly in Florida (Homoptera: Aleyrodidae: Aleyrodinae). Fla. Dept. of Agric. & Consumer Ser. Entomology Circular No. 292. 2 p. (1987).
Dennehy, T. J. et al. New challenges to management of whitefly resistance to insecticides in Arizona. University of Arizona Cooperative Extension, Vegetable Report. 31 pp. Series P-144. (eds D. N. Byrne and P. Baciewicz). https://cals.arizona.edu/pubs/crops/az1382/index.html (2005).
Xie, W. et al. Genome sequencing of the sweetpotato whitefly Bemisia tabaci MED/Q. Gigascience 6, 1–7 (2017).
Li, H. et al. Invasion genomics uncover complex introduction patterns of the globally invasive whitefly, Bemisia tabaciMED. Divers. Distrib. https://doi.org/10.1111/ddi.13751 (2023).
Chen, W. et al. Genome of the African cassava whitefly Bemisia tabaci and distribution and genetic diversity of cassava-colonizing whiteflies in Africa. Insect Biochem Mol. Biol. 110, 112–120 (2019).
Chen, W. et al. The draft genome of whitefly Bemisia tabaci MEAM1, a global crop pest, provides novel insights into virus transmission, host adaptation, and insecticide resistance. BMC Biol. 14, 110 (2016).
Campbell, L. I. et al. Comparative evolutionary analyses of eight whitefly Bemisia tabaci sensu lato genomes: cryptic species, agricultural pests and plant-virus vectors. BMC Genom. 24, 408 (2023).
Xie, W. et al. The invasive MED/Q Bemisia tabaci genome: a tale of gene loss and gene gain. BMC Genom. 19, 68 (2018).
Gautam, S. et al. Effects of host plants and their infection status on acquisition and inoculation of A plant virus by its hemipteran vector. Pathogens 12, https://doi.org/10.3390/pathogens12091119 (2023).
Mugerwa, H. et al. Differential transcriptional responses in two old world Bemisia tabaci cryptic species post acquisition of old and new world begomoviruses. Cells 11, https://doi.org/10.3390/cells11132060 (2022).
Li, Y., Mbata, G. N., Punnuri, S., Simmons, A. M. & Shapiro-Ilan, D. I. Bemisia tabaci on vegetables in the Southern United States: incidence, impact, and management. Insects 12, https://doi.org/10.3390/insects12030198 (2021).
McKenzie, C. L., Sparks, A. N., Roberts, P., Oetting, R. D. & Osborne, L. S. Survey of Bemisia tabaci (Hemiptera: Aleyrodidae) in Agricultural Ecosystems in Georgia. J. Entomol. Sci. 55, https://doi.org/10.18474/0749-8004-55.2.163 (2020).
Gautam, S. et al. Differential Transmission of Old and New World Begomoviruses by Middle East-Asia Minor 1 (MEAM1) and Mediterranean (MED) Cryptic Species of Bemisia tabaci. Viruses 14, https://doi.org/10.3390/v14051104 (2022).
Polston, J. E., De Barro, P. & Boykin, L. M. Transmission specificities of plant viruses with the newly identified species of the Bemisia tabaci species complex. Pest. Manag. Sci. 70, 1547–1552 (2014).
de Mendoza, A., Pflueger, J. & Lister, R. Capture of a functionally active methyl-CpG binding domain by an arthropod retrotransposon family. Genome Res. 29, 1277–1286 (2019).
Yu, X. et al. Sex-specific transcription and DNA methylation landscapes of the Asian citrus psyllid, a vector of huanglongbing pathogens. Evolution 77, 1203–1215 (2023).
Sicat, J. P. A., Visendi, P., Sewe, S. O., Bouvaine, S. & Seal, S. E. Characterization of transposable elements within the Bemisia tabaci species complex. Mob. DNA 13, 12 (2022).
Zidi, M. et al. Genome-wide screening of transposable elements in the whitefly, Bemisia tabaci (Hemiptera: Aleyrodidae), revealed insertions with potential insecticide resistance implications. Insects 13, https://doi.org/10.3390/insects13050396 (2022).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Gordon, S. P. et al. Widespread polycistronic transcripts in fungi revealed by single-molecule mRNA sequencing. PLoS One 10, e0132628 (2015).
Yi, S. V. & Goodisman, M. A. Computational approaches for understanding the evolution of DNA methylation in animals. Epigenetics 4, 551–556 (2009).
Bhattacharjee, B. & Hallan, V. Geminivirus-derived vectors as tools for functional genomics. Front. Microbiol. 13, 799345 (2022).
Coates, B. S. Horizontal transfer of a non-autonomous Helitron among insect and viral genomes. BMC Genom. 16, 137 (2015).
Kiser, L. M., Sokoloski, K. J. & Hardy, R. W. Interactions between capsid and viral RNA regulate Chikungunya virus translation in a host-specific manner. Virology 560, 34–42 (2021).
Ahlers, L. R. H. et al. Insulin potentiates JAK/STAT signaling to broadly inhibit flavivirus replication in insect vectors. Cell Rep. 29, 1946–1960.e1945 (2019).
Kemp, C. et al. Broad RNA interference-mediated antiviral immunity and virus-specific inducible responses in Drosophila. J. Immunol. 190, 650–658 (2013).
Dostert, C. et al. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat. Immunol. 6, 946–953 (2005).
Chen, L. L. et al. Identification of a nucleocapsid protein (VP35) gene of shrimp white spot syndrome virus and characterization of the motif important for targeting VP35 to the nuclei of transfected insect cells. Virology 293, 44–53 (2002).
Li, P. et al. Plant begomoviruses subvert ubiquitination to suppress plant defenses against insect vectors. PLoS Pathog. 15, e1007607 (2019).
Yang, C., Kang, L. & Zhao, Q. Comparative transcriptomic analysis of the l-4i silkworm (Lepidoptera: Bombyx mori) mutants and its wild-type strain P33 by RNA-Seq. Comp. Biochem Physiol. Part D. Genom. Proteom. 38, 100800 (2021).
Schmid, M., Steinlein, C., Yano, C. F. & Cioffi, M. B. Hypermethylated chromosome regions in nine fish species with heteromorphic sex chromosomes. Cytogenet. Genome Res. 147, 169–178 (2015).
Field, L. M., Lyko, F., Mandrioli, M. & Prantera, G. DNA methylation in insects. Insect Mol. Biol. 13, 109–115 (2004).
Cunningham, C. B. et al. An association between Dnmt1 and Wnt in the production of oocytes in the whitefly Bemisia tabaci. Insect. Mol. Biol. https://doi.org/10.1111/imb.12893 (2024).
Ylla, G. et al. Insights into the genomic evolution of insects from cricket genomes. Commun. Biol. 4, 733 (2021).
Sun, D., Li, Q. & Yu, H. DNA methylation differences between male and female gonads of the oyster reveal the role of epigenetics in sex determination. Gene 820, 146260 (2022).
Marshall, H. et al. DNA methylation is associated with codon degeneracy in a species of bumblebee. Heredity 130, 188–195 (2023).
Elango, N., Hunt, B. G., Goodisman, M. A. & Yi, S. V. DNA methylation is widespread and associated with differential gene expression in castes of the honeybee, Apis mellifera. Proc. Natl Acad. Sci. USA 106, 11206–11211 (2009).
Provataris, P., Meusemann, K., Niehuis, O., Grath, S. & Misof, B. Signatures of DNA methylation across insects suggest reduced DNA methylation levels in holometabola. Genome Biol. Evol. 10, 1185–1197 (2018).
Foret, S., Kucharski, R., Pittelkow, Y., Lockett, G. A. & Maleszka, R. Epigenetic regulation of the honey bee transcriptome: unravelling the nature of methylated genes. BMC Genom. 10, 472 (2009).
Kvist, J. et al. Pattern of DNA methylation in daphnia: evolutionary perspective. Genome Biol. Evol. 10, 1988–2007 (2018).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat. Genet 39, 457–466 (2007).
Jiang, N. et al. Conserved and divergent patterns of DNA methylation in higher vertebrates. Genome Biol. Evol. 6, 2998–3014 (2014).
Greenberg, M. V. C. & Bourc’his, D. The diverse roles of DNA methylation in mammalian development and disease. Nat. Rev. Mol. Cell Biol. 20, 590–607 (2019).
Fu, R., Huang, X., Chen, Y., Chen, Z. & Zhan, A. Interactive regulations of dynamic methylation and transcriptional responses to recurring environmental stresses during biological invasions. Front. Mar. Sci. 8, https://doi.org/10.3389/fmars.2021.800745 (2021).
Bourque, G. et al. Ten things you should know about transposable elements. Genome Biol. 19, 199 (2018).
Malik, H. S. & Eickbush, T. H. Phylogenetic analysis of ribonuclease H domains suggests a late, chimeric origin of LTR retrotransposable elements and retroviruses. Genome Res. 11, 1187–1197 (2001).
Merel, V., Boulesteix, M., Fablet, M. & Vieira, C. Transposable elements in Drosophila. Mob. DNA 11, 23 (2020).
Strand, D. J. & McDonald, J. F. Copia is transcriptionally responsive to environmental stress. Nucleic Acids Res. 13, 4401–4410 (1985).
Merkling, S. H. et al. The heat shock response restricts virus infection in Drosophila. Sci. Rep. 5, 12758 (2015).
Wang, L. et al. Retrotransposon activation during Drosophila metamorphosis conditions adult antiviral responses. Nat. Genet. 54, 1933–1945 (2022).
Jiang, L. et al. Distinct functions of Bombyx mori peptidoglycan recognition protein 2 in immune responses to bacteria and viruses. Front. Immunol. 10, 776 (2019).
Badillo-Vargas, I. E. et al. Proteomic analysis of Frankliniella occidentalis and differentially expressed proteins in response to tomato spotted wilt virus infection. J. Virol. 86, 8793–8809 (2012).
Li, H. J., Zhang, H. H., Lu, J. B. & Zhang, C. X. Threonyl-tRNA synthetase gene, a potential target for RNAi-based control of three rice planthoppers. Pest Manag. Sci. 78, 4589–4598 (2022).
Gautam, S. et al. Virus-virus interactions in a plant host and in a hemipteran vector: Implications for vector fitness and virus epidemics. Virus Res. 286, 198069 (2020).
Ghosh, S., Bouvaine, S., Richardson, S. C. W., Ghanim, M. & Maruthi, M. N. Fitness costs associated with infections of secondary endosymbionts in the cassava whitefly species Bemisia tabaci. J. Pest Sci. (2004) 91, 17–28 (2018).
De Barro, P. J. et al. Isolation and characterization of microsatellite loci in Bemisia tabaci. Mol. Ecol. Notes 3, 40–43 (2003).
Marchant, W. G., Gautam, S., Hutton, S. F. & Srinivasan, R. Tomato yellow leaf curl virus-resistant and -susceptible tomato genotypes similarly impact the virus population genetics. Front. Plant Sci. 11, 599697 (2020).
Ewels, P., Magnusson, M., Lundin, S. & Kaller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Xi, Y. & Li, W. BSMAP: whole genome bisulfite sequence MAPping program. BMC Bioinform. 10, 232 (2009).
Akalin, A. et al. methylKit: a comprehensive R package for the analysis of genome-wide DNA methylation profiles. Genome Biol. 13, R87 (2012).
Park, Y. & Wu, H. Differential methylation analysis for BS-seq data under general experimental design. Bioinformatics 32, 1446–1453 (2016).
Bulla, I. et al. Notos - a galaxy tool to analyze CpN observed expected ratios for inferring DNA methylation types. BMC Bioinform. 19, 105 (2018).
Su, W., Ou, S., Hufford, M. B. & Peterson, T. A tutorial of EDTA: extensive de novo TE annotator. Methods Mol. Biol. 2250, 55–67 (2021).
Bell, E. A. et al. Transposable element annotation in non-model species: the benefits of species-specific repeat libraries using semi-automated EDTA and DeepTE de novo pipelines. Mol. Ecol. Resour. 22, 823–833 (2022).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Anders, S., Reyes, A. & Huber, W. Detecting differential usage of exons from RNA-seq data. Genome Res. 22, 2008–2017 (2012).
Xu, L. et al. OrthoVenn2: a web server for whole-genome comparison and annotation of orthologous clusters across multiple species. Nucleic Acids Res. 47, W52–W58 (2019).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Emms, D. M. & Kelly, S. STRIDE: species tree root inference from gene duplication events. Mol. Biol. Evol. 34, 3267–3278 (2017).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Sievers, F. & Higgins, D. G. Clustal omega. Curr. Protoc. Bioinforma. 48, 3 13 11–13 13 16 (2014).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Acknowledgements
Thank you to Yi-Ju Chen, Alex Waugh, Sarah Orr, Mike Goodisman, and Ken Ross for additional feedback and discussions on the manuscript.
Author information
Authors and Affiliations
Contributions
M.A.C: Data curation; formal analysis; visualization; writing—original draft; writing—review and editing. S.G: Methodology; writing—review and editing. B.M.: Methodology; writing—review and editing. S.P.: Writing—review and editing. A.S.: Funding acquisition. B.H.: Resources; supervision; writing—review and editing. R.S.: Conceptualization; funding acquisition; resources; supervision; writing—review and editing. All authors have reviewed and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The study was conducted with all biosafety guidelines or regulations approved by the University of Georgia. All experiments were carried out in accordance with those regulations. This study only used adult whiteflies. No vertebrates were used in this study, and no ethical concerns were encountered.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Catto, M.A., Ghosh, S., Pandey, S. et al. A plant virus differentially alters DNA methylation in two cryptic species of a hemipteran vector. npj Viruses 2, 35 (2024). https://doi.org/10.1038/s44298-024-00044-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44298-024-00044-2