Abstract
Alcoholic beverages have been concerned not only for gastronomic delight but also for certain impacts on health, such as obesity, diabetes, and cardiovascular diseases. In this study, we assessed the bioactive functions of 1,1-Diethoxyethane (1,1-DEE), a flavoring compound formed during the aging process of wine by flor yeast, using both cultured cell lines and a high-fat diet (HFD) mouse model. 1,1-DEE was identified in the batches of ethanol that induced oxidation of phosphatase and tensin homolog deleted on chromosome 10 (PTEN) using gel mobility shift assay and gas chromatography-mass spectrometry. PTEN was reversibly oxidized when exposed to 1,1-DEE, but 1,2-DEE did not induce PTEN oxidation. Mechanistically, 1,1-DEE treatment enhanced the production of mitochondrial reactive oxygen species, accompanying by oxidation of PTEN and subsequent activation of Akt signaling. 1,1-DEE treatment elevated Akt activation when combined with insulin, compared with insulin alone, and alleviated palmitate-induced insulin resistance in C2C12 myoblasts. Moreover, the oral administration of 1,1-DEE alleviated glucose intolerance and insulin resistance in HFD-fed mice. 1,1-DEE also mitigated HFD-induced body weight gain and hepatic dyslipidemia without reduction of food intake. Transcriptome analysis revealed significant genes involved in the improvement of insulin sensitivity and dyslipidemia. Thus, 1,1-DEE may serve as a promising therapeutic agent for the intervention of obesity, diabetes, and dyslipidemia.

Similar content being viewed by others
Introduction
Obesity is defined as abnormal weight gain resulting from excessive fat accumulation in adipose and non-adipose tissue. It is frequently associated with insulin resistance, which is a major risk factor for metabolic disorders such as type-2 diabetes, non-alcoholic fatty liver disease, and cardiovascular disease1,2. Western diet is high in saturated fats and fructose, both of which are closely linked to obesity and insulin resistance3. Various animal models have been used to mimic aspects of obesity and obesity-linked metabolic disorders for assessing the applications of therapeutic agents4. However, underlying mechanisms of these diseases are not completely understood, and the existing drugs have certain limitations5,6.
The phosphoinositide-3-kinase (PI3K)/protein kinase B (Akt) signaling pathway is involved in the regulation of insulin signaling7,8,9. Phosphatase and tensin homolog deleted on chromosome 10 (PTEN) acts as a lipid phosphatase that dephosphorylates PIP3 to PIP2, negatively regulating the PI3K/Akt signaling pathway10. When insulin binds to its receptor, the receptor undergoes autophosphorylation on tyrosine residues and phosphorylates insulin receptor substrates, which further favors the activation of the PI3K/Akt signaling pathway9. Insulin-mediated Akt activation promotes the transport of glucose into the intracellular compartment and the synthesis of glycogen while inhibiting gluconeogenesis9. Insulin resistance is associated with impaired PI3K/Akt signaling. Research has revealed that the oxidative inactivation of PTEN by reactive oxygen species (ROS) is associated with increased insulin sensitivity via Akt signaling11,12. Hence, targeting the redox regulation of PTEN may be beneficial for controlling insulin resistance.
Alcoholic beverage consumption is a globally prevalent and often debated phenomenon. While excessive intake in alcoholic beverages can lead to various diseases, moderate consumption (particularly wine) may certainly mitigate body weight gain, insulin insensitivity, and cardiovascular disease13. Research on the benefits of wine consumption has identified various bioactive compounds in wine, with a particular focus on polyphenols such as resveratrol, quercetin, and catechin14. However, little is known about the biological effects of other components in wine. Aroma compounds, commonly present in aged wine, exhibit volatile properties with characteristic odors. Studies have demonstrated that a significant number of aroma compounds are formed during the aging of various types of wine, including sherry, Chinese rice, and potato wines15,16,17,18. These aroma compounds influence the economic value, quality, and consumer preference of wine by imparting alcoholic, fruity, herb, caramel-like, and/or honey-like flavors. These flavors also stimulate olfactory sensors, leaving distinct aromatic impressions in the brain19. Among the aroma compounds formed during the aging process of wine, 1,1-Diethoxyethane (1,1-DEE) is one of the most odor-active compounds associated with fruity and wine flavors. Various studies have revealed a high concentration of 1,1-DEE in wine with a high odor-active index (Table 1). In particular, 1,1-DEE has been reported to be the most prevalent aroma compound in Fino Sherry (448 mg/L)15. The concentration of 1,1-DEE in aged Chinese rice wine is four times higher than that in young rice wine17. 1,1-DEE is also found in potato wine, red wine, wine vinegar, and aged rum18,20,21,22,23. However, its biological functions have not been assessed in vitro or in vivo so far.
In this study, we identified the presence of 1,1-DEE in batches of ethanol obtained from different commercial sources. We also assessed its novel ability to control the redox state of PTEN through ROS generation. In addition, we evaluated the therapeutic effects of 1,1-DEE in controlling obesity, insulin resistance, and dyslipidemia using a high-fat diet (HFD) mouse model.
Results
1,1-DEE was identified in the ethanol batches that induced oxidation of PTEN
For the study of redox status of PTEN, we developed a gel mobility shift assay as described previously24,25,26. We performed cell lysis with NEM, a well-known alkylating reagent, that can bind to the thiol groups of reduced PTEN. This process prevents various redox reactions and results in a higher molecular weight band on a western blot compared with the oxidized PTEN band. In the present study, the regulation of PTEN’s redox state by different batches of ethanol from commercial sources (Supplementary Table 1) was investigated in HepG2 cells. Intriguingly, we found that the oxidation rate of PTEN significantly varied across different ethanol batches, even when using the same concentration of ethanol. The E1 batch induced greater PTEN oxidation than the E2 and E3. Combining E1 with E2 or E3 reduced the PTEN oxidation level (Fig. 1A), suggesting that E1’s oxidative effect was stronger than the other two batches. In addition, we observed that E1’s odor differed from those of the other ethanol batches. Specifically, E1 displayed a much stronger alcoholic and fruity odor than the other batches, suggesting that there may be an E1-specific compound involved in the redox regulation of PTEN.
A Immunoblotting of protein samples from the treatment of HepG2 cells with 100 mM E1, E2, or E3 for 10 min. B Workflow for the freeze-drying of 100 mM E1, E2, and E4. This model was created in BioRender. Lee, S. (2025) http://BioRender.com/2yu3squi. C The cells were treated with 100 mM E1, E2, or E4 or their corresponding dried samples described in (B) for 10 min. The cell extracts were alkylated with 10 mM NEM and subjected to non-reducing or reducing electrophoresis, followed by immunoblotting with PTEN and β-actin antibodies. D GS-MS analysis of 1,1-DEE fragmentation in E1. E The chemical structure of 1,1-DEE. This figure was created in BioRender. Lee, S. (2025) http://BioRender.com/lc7iuk5. Data were presented as mean ± SEM. The differences between two groups were assessed by unpaired two-tailed Student’s t-tests (A, C), with the following significance levels: nsp > 0.05, ***p ≤ 0.001, and ****p ≤ 0.0001.
To investigate the properties of this unknown compound, we analyzed the E1, E2, and E4 batches. E4 displayed a similar odor profile to E1 (Fig. 1B). In each batch, we freeze-dried 100 mM of ethanol to evaporate the volatile compounds. Dried samples were reconstituted with either PBS or 100 mM of E2. We then treated HepG2 cells with either 100 mM of ethanol (E1, E2, or E4) or the corresponding dried samples for 10 min. E1 and E4 induced greater PTEN oxidation than E2 and the control. However, the dried samples from E1, E2, and E4 did not exhibit similar effects, regardless of the reconstitution status (PBS or E2; Fig. 1C). This result suggests that the unknown compound is volatile and evaporates during the freeze-drying process.
Next, we used GC-MS to detect the compounds in the ethanol batches. Interestingly, a high concentration of 1,1-DEE was found in E1 and E4 but not in E2 or E3 (Supplementary Tables 2, 3, 4, 5). GC-MS-based fragmentation analysis of 1,1-DEE from E1 and E4 and the identified chemical structure are shown in Fig. 1D, E. Altogether, the unknown compound was identified as 1,1-DEE, and the results highlight its novel role in cellular functions via the redox regulation of PTEN.
1,1-DEE induced reversible oxidative inactivation of PTEN and activation of Akt
To confirm whether 1,1-DEE can exhibit its molecular effects via the regulation of PTEN’s redox state, cells were treated with various concentrations of 1,1-DEE, after which the PTEN oxidation level was assessed using a previously described method25. Within 10 min of treatment, we observed 1,1-DEE-mediated PTEN oxidation in various cell lines (Fig. 2A and Supplementary Fig. 1A, B). This 1,1-DEE-mediated oxidation occurred in a concentration-dependent manner, with higher concentrations inducing higher oxidation levels. We also assessed the effect of 1,1-DEE-induced PTEN oxidation on Akt activation. The phosphorylation of Akt at Ser473 and Thr308 was enhanced after 10 min of treatment with 1,1-DEE, starting at 1 mM, indicating increased Akt activity. Akt phosphorylation showed a concentration-dependent increase with 1,1-DEE, indicating the negative regulation of PTEN in the Akt signaling pathway (Fig. 2A).
A C2C12 cells were treated with various concentrations of 1,1-DEE (0 to 50 mM) for 10 min. B C2C12 cells were treated with 20 mM of 1,1-DEE for various durations over a 120-min period. C Chemical structure of 1,2-DEE. This figure was created in BioRender. Lee, S. (2025) http://BioRender.com/v87h58o. D, E Cells were treated with similar conditions as in (A) and (B) using 1,2-DEE. The cell lysates were mixed with sample buffer and then subjected to non-reducing or reducing electrophoresis, followed by immunoblotting with antibodies against PTEN, p-AktSer473, p-AktThr308, Akt, and GAPDH. Data were presented as mean ± SEM. Statistical analysis was assessed by unpaired two-tailed Student’s t-tests (A, B), with the following significance levels: nsp > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001. F HepG2 cells were transfected with an HA-tag pCGN containing either PTEN WT, C71S, C124S, or C71S/C124S vector. After the transfection, the cells were treated with 20 mM 1,1-DEE for 10 min. The cell lysates were subjected to non-reducing or reducing electrophoresis, followed by immunoblotting with antibodies against HA-tag.
We next sought to elucidate the pattern of 1,1-DEE-mediated regulation of PTEN’s redox state. PTEN oxidation levels started to increase as early as 5 min after 1,1-DEE treatment, peaked at 10 min, gradually returned to the basal level after 120 min (Fig. 2B and Supplementary Fig. 1C, D). Furthermore, similar to the reversible oxidation of PTEN mediated by 1,1-DEE, Akt phosphorylation at Ser473 and Thr308 was also found to be reversible (Fig. 2B), thus suggesting that the oxidative inhibition of PTEN by 1,1-DEE causes the reversible activation of Akt. On the other hand, 1,2-DEE, an isomer of 1,1-DEE, did not induce PTEN oxidation in C2C12 cells at different concentrations and time points, suggesting a specific conformation-dependent function of 1,1-DEE in the redox regulation of PTEN (Fig. 2C, D, E).
The 1,1-DEE-mediated oxidation of PTEN showed a similar pattern to that of H2O2-induced PTEN oxidation24. It has been demonstrated that H2O2-mediated peroxymonocarbonate (HCO4-) generation leads to the oxidation of PTEN at the Cys124 residue and the formation of a disulfide bond with the Cys71 residue, which then inactivates PTEN’s phosphatase function24,27. To investigate the mechanism underlying 1,1-DEE-mediated PTEN oxidation, HepG2 cells transfected with an HA tag-pCGN vector containing either PTEN WT or PTEN mutants (PTEN C71S, PTEN C124S, or PTEN C71/124S) were treated with 1,1-DEE for 10 min. Accordingly, PTEN oxidation was detected in cells transfected with WT PTEN but not in those transfected with PTEN C71S, PTEN C124S, or PTEN C71/124S (Fig. 2F). This indicates that 1,1-DEE mediates the oxidative inactivation of PTEN by inducing the formation of a disulfide bond between its Cys124 and Cys71 residues.
1,1-DEE enhanced production of mitochondrial reactive oxygen species
ROS are by-products of cellular redox reactions that can contribute to both physiological and pathophysiological conditions28,29. At low and controlled levels, ROS can modulate redox-sensitive signaling pathways to maintain cellular homeostasis. DCFH-DA staining of cells exposed to 1,1-DEE revealed an increase in ROS levels compared with the untreated group (Fig. 3A). Moreover, flow cytometry analysis of the DCFH-DA-stained cells revealed that the ROS levels increased as early as 5 min into the treatment, peaked at 10 min, and decreased after 30 min (Fig. 3B). This suggests that 1,1-DEE induces a reversible increase in ROS generation.
A, B HepG2 cells were treated with either 20 mM 1,1-DEE for various durations over a 30-min period or 0.5 mM H2O2 for 10 min. The cells were stained with 10 µM DCFH.DA for 30 min. Representative images of the stained cells were obtained using fluorescence microscopy at 10x magnification (A). Flow cytometry was performed using FITC channel with 10,000 events per sample (B). C Flow cytometry analysis of MitoSOX Red staining after treatment of AC16 cells with 15 mM of 1,1-DEE for various time points up to 30 min. Data were observed using the Pe-A channel with 10,000 events per sample. D HepG2 cells were pretreated with 10 mM of NAC for 120 min. Then, cells were treated with 20 mM 1,1-DEE for various durations over a 120 min period. Data were presented as mean ± SEM. Statistical analysis was assessed by unpaired two-tailed Student’s t-tests (B) and two-way ANOVA (D), with the following significance levels: nsp > 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
Mitochondria are major sources of ROS generation, especially superoxide anion (O2 ∙ -), which is predominant through electron leakage from the mitochondrial respiratory complexes I and III during oxidative phosphorylation30. In a parallel study, mitochondrial respiration was transiently inhibited in the cells upon exposure to 1,1-DEE, suggesting that 1,1-DEE may favor mitochondrial ROS generation through this inhibition. Molecular simulation revealed that 1,1-DEE can dock into the ubiquinone-binding site of human respiratory complex I, thereby interrupting the electron transfer from NADH to ubiquinone (Supplementary Fig. 2). This interruption leads to leakage of electrons from the respiratory chain. These leaked electrons can reduce molecular oxygen (O2) to form O2 ∙ -, which is then dismutated into H2O2. In fact, MitoSox staining revealed that 1,1-DEE induced mitochondrial ROS generation during 30 min of treatment (Fig. 3C), indicating that mitochondria are major sources for the ROS generation by 1,1-DEE.
To understand the relationship between ROS generation and 1,1-DEE-induced PTEN oxidation, cells were pretreated with NAC, a ROS scavenger, for 120 min and treated with 1,1-DEE for various durations over a 120-min period. NAC pretreatment reduced 1,1-DEE-induced PTEN oxidation, indicating that the ROS generation induced by 1,1-DEE is responsible for the oxidative inactivation of PTEN. In addition, the phosphorylation of Akt at Ser473 and Thr308 was also reduced by NAC pretreatment (Fig. 3D), suggesting the regulatory mechanism of 1,1-DEE-induced ROS generation in the PTEN/Akt signaling pathway.
1,1-DEE treatment improved insulin sensitivity through regulation of PTEN/Akt signaling
The PI3K/PTEN/Akt signaling pathway is crucial in regulating glucose metabolism and insulin resistance in mice fed with the HFD. Given that 1,1-DEE treatment results in PTEN oxidation and Akt activation, we hypothesized that 1,1-DEE increases insulin sensitivity through regulation of the PTEN/Akt signaling pathway. Combining 10 nM of insulin and various concentrations of 1,1-DEE (1–10 mM) significantly enhanced Akt activation compared with treatment with insulin alone (Fig. 4A). We next established an in vitro cellular model for insulin resistance using treatment with palmitate as described previously31. It was found that Akt phosphorylation at Ser473 and Thr308 was significantly reduced in insulin-resistant cells compared with the control cells (Fig. 4B), thus confirming the successful generation of an insulin-resistant model through palmitate treatment. Importantly, co-treatment with 1,1-DEE and insulin re-upregulated Akt phosphorylation in insulin-resistant cells (Fig. 4B). This suggests that 1,1-DEE is a candidate material for mitigating palmitate-induced insulin resistance by oxidative inactivation of PTEN and activation of Akt.
A C2C12 cells were treated with various concentrations of 1,1-DEE (0 to 10 mM) in the absence or presence of 10 nM insulin for 10 min. B C2C12 cells were pretreated with 500 µM palmitate for 24 h. The cells were then treated with either 10 mM of insulin or a combination of 5 mM of 1,1-DEE and 10 nM of insulin for 10 min. The cell lysates were mixed with sample buffer and then subjected to non-reducing or reducing electrophoresis, followed by immunoblotting with antibodies against PTEN, p-AktSer473, p-AktThr308, Akt, and β-actin. Data were presented as mean ± SEM. Statistical analysis was assessed by unpaired two-tailed Student’s t-tests (A) and one-way ANOVA (B), with the following significance levels: nsp > 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
1,1-DEE treatment attenuated glucose intolerance and insulin resistance in mice fed the HFD
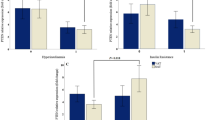
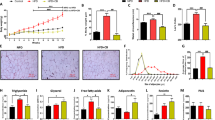
HFD is associated with various metabolic dysregulations, including obesity, hyperglycemia, and insulin resistance. We next investigated the ability of 1,1-DEE to control glucose uptake and insulin sensitivity using the HFD-fed mice model. The HFD group displayed a higher level of fasting blood glucose than the ND group, indicating a hyperglycemic condition. Interestingly, the HFD supplemented with 1,1-DEE group had a lower fasting blood glucose level than the HFD (Fig. 5A). As shown in Fig. 5B, C, the HFD group exhibited impaired glucose tolerance after 120 min, as indicated by a high intraperitoneal GTT area under the curve (AUC) level. Meanwhile, the HFD + D mice displayed greater glucose tolerance, as indicated by a lower GTT AUC value compared with that for the HFD group (p = 0.1846). These data reveal the ability of 1,1-DEE to control the glucose level. HFD-induced obesity is associated with insulin resistance. After 8 weeks of HFD feeding, the mice had a higher serum insulin level than the ND control group, whereas the HFD + D group had a lower serum insulin level than the HFD group (p < 0.05) (Fig. 5D). Next, we tested insulin tolerance after the mice had fasted for 4 h followed by being intraperitoneally injected with insulin. The HFD group showed a high insulin ITT AUC value, thus indicating a higher insulin intolerance than the ND group. However, the HFD + D group showed a slight decrease in ITT AUC value compared with the HFD group, thus suggesting an amelioration of insulin tolerance induced by 1,1-DEE treatment (p = 0.3514) (Fig. 5E, F).
A–F 16-week-old mice were fasted for 12 h overnight. Fasting blood glucose levels were measured (A). Intraperitoneal glucose tolerance test was performed with a duration of 120 min (B). C The area under curve (AUC) of (B) was measured. D Fasting serum insulin levels were determined. E Intraperitoneal insulin tolerance test was performed with a duration of 120 min after fasting mice for 4 h. F AUC levels of (E) were calculated. Data were presented as mean ± SEM (n = 4). Statistical analysis was assessed by one-way ANOVA (A, C, D, F), and multiple t-tests (B, E), with the following significance levels: nsp > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001. G, H GSEA preranked enrichment plots for the “GOBP response to insulin” (G) and “GOBP insulin-like growth factor receptor signaling pathway” (H), which were enriched in the ND group but not in the HFD group. I–K GSEA preranked enrichment plots for the “GOBP response to insulin” (I) and “GOBP insulin-like growth factor receptor signaling pathway” (J), which were enriched in the HFD + D group but not in the HFD group. K A heatmap for the leading gene expressions in the “GOBP response to insulin” in comparison between the HFD and the HFD + D groups.
Given that redox regulation of PTEN was implicated in amelioration of insulin resistance in the HFD + D group, we validated differences in oxidation levels of PTEN between the HFD and HFD + D groups in white adipose tissue (WAT), liver, and heart. Interestingly, we observed an increase in PTEN oxidation in WAT of the HFD + D group compared with the HFD control group (Supplementary Fig. 3A). This accompanied by an enhancement in Akt phosphorylation in the HFD + D group compared with the HFD group. No differences in PTEN redox state were found between tested groups in liver and heart tissues (Supplementary Fig. 3B, C). These data suggest that 1,1-DEE specifically induces oxidative inactivation of PTEN in WAT, leading to enhanced insulin sensitivity by activation of AKT signaling pathway.
We performed the RNA-seq analysis on mouse liver tissue to assess the differences in transcriptomic profiles among tested groups. The GSEA preranked enrichment analysis revealed downregulations in insulin metabolic processes of “GOBP response to insulin” (NES = −1.37, FDR = 0.3, nom p-value = 0.002), “GOBP insulin-like growth factor receptor signaling pathway” (NES = −1.57, FDR = 0.17, nom p-value = 0.006), and “GOBP cellular response to insulin stimulus” (Fig. 5G, H, Supplementary Fig. 4A). However, treatment of 1,1-DEE significantly enriched these processes, with the NES = 2.1, FDR = 0.012, and nom p-value = 0.0 in “GOBP response to insulin”, and the NES = 2.6, FDR = 0.0, and nom p-value = 0.00 in “GOBP insulin-like growth factor receptor signaling pathway” (Fig. 5I, J). Additionally, treatment of 1,1-DEE enhanced other insulin sensitivity regulation processes such as “GOBP insulin receptor signaling pathway”, “GOBP cellular response to insulin stimulus”, “GOBP insulin secretion”, and “GOBP regulation of insulin secretion” (Supplementary Fig. 4B, C, D, E). A heatmap showed differences in leading transcriptional expressions of “GOBP response to insulin” between the HFD and the HFD + D groups, including genes involved in insulin signaling pathway (insulin receptor (Insr), insulin receptor substrate 2 (irs2)), growth factor signaling pathway (insulin-like growth factor binding protein 1 (igfbp1), insulin-like growth factor 1 receptor (Igf1r), egfr, fibroblast growth factor receptor 1 (fgfr1), fgfr2, erbb4), transportation (slc39a14) (Fig. 5K). Thus, 1,1-DEE could improve insulin resistance through transcriptional regulation of these genes.
1,1-DEE treatment alleviated body weight gain and dyslipidemia in mice fed the HFD
As can be seen in Fig. 6A presenting representative mice in each group, the HFD group exhibited greater body weight gain than the ND group. However, supplementation with 1,1-DEE relatively suppressed HFD-induced body weight gain, and there was no significant difference in body weight between the ND and ND + D groups (Fig. 6B). The ND + D group showed higher food intake than the ND group, while food consumption by the HFD and HFD + D groups was similar (Fig. 6C). In addition, the HFD group exhibited an increase in eWAT and iWAT weight compared with the ND group. H&E staining revealed larger adipose sizes in the HFD group than the ND group. Supplementation with 1,1-DEE attenuated these changes (Fig. 6D, E, F).
A–H 8-week-old mice in each group were fed with indicated diets for 8 weeks. Representative images of 16-week-old mice in each group (A). Body weight (B). Food intake (C). eWAT weight (D). iWAT weight (E). Representative H&E staining of eWAT and iWAT, scale bar: 100 µM (F). Liver weight (G). Representative images and H&E staining of liver, scale bar: 100 µM (H). I–L Serum biomarkers were measured, including T-chol (I), HDL-C (J), LDL-C (K), TG (L). Data were presented as mean ± SEM (n = 4). Statistical analysis was performed using one-way ANOVA (C, D, E, G, I–L), with the following significance levels: nsp > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, and ****p ≤ 0.0001.
We assessed the effects of 1,1-DEE on the regulation of dyslipidemia induced by the HFD. While no significant difference was observed in liver weight between the ND and HFD groups with or without 1,1-DEE (Fig. 6G), H&E staining of liver sections revealed increased cytoplasmic vacuoles in the hepatocytes of the HFD group compared with the ND group, thereby suggesting the sign of steatosis in the HFD group. However, treatment with 1,1-DEE prevented this change (Fig. 6H). These results suggest that 1,1-DEE treatment can mitigate HFD-induced liver steatosis, as evidenced by decreased liver vacuolation.
Next, we sought to examine the effects of 1,1-DEE on serum biomarkers indicating liver damage, such as ALT, AST, and LHD. All of these were relatively increased in the HFD group compared with the ND group. Interestingly, the addition of 1,1-DEE reduced the levels of these biomarkers compared with its counterpart (Supplementary Fig. 5A, B, C). No significant differences were found in serum ALP, T-Bil, and Alb levels across the four groups (Supplementary Fig. 5D, E, F). Serum biomarkers indicating lipogenesis and adipogenesis, such as TG, T-chol, HDL-C, and LDL-C (a “bad” cholesterol involved in plaque buildup in the arteries), were also investigated. The HFD group exhibited enhancements in T-chol, HDL-C, and LDL-C levels but not TG levels compared with the ND group (Fig. 6I, J, K, L). Interestingly, the HFD + D group showed reduced LDL-C levels compared with the HFD group (Fig. 6K). Additionally, 1,1-DEE supplementation decreased the TG level compared with the corresponding counterpart HFD and ND groups (Fig. 6L). Collectively, based on the changes in multiple parameters, 1,1-DEE shows effects consistent with prevention of liver damage and lipogenesis in HFD-induced obese mice.
1,1-DEE treatment restored transcriptional expression changes in mice fed the HFD
We performed RNA-seq analysis using the liver tissues to delineate the transcriptome expression between groups of mice. The heatmaps demonstrated DEGs in the comparisons between the HFD and ND groups, as well as between the HFD + D and HFD groups (Supplementary Fig. 6A, B). Volcano plots displayed DEGs (pink and blue dots) and non-statistically significant genes (gray dots) (Fig. 7A, B). We next listed the most upregulated genes after the HFD (yellow dots), including Fgf21, Gm15441, Cfd, derlin-3 (Derl3), cyclin dependent kinase inhibitor 1 A (Cdkn1a), Cidec, Rec114, Cytochrome P450 genes (Cyp4a14, Cyp4a10, Cyp2b9). The most downregulated genes consisted of Chrna4, stearoyl-CoA desaturase (Scd1), Gm34654, Xlr4a, Cib3, Cyp2b10, Cyp2c53-ps, Cyp2c55, Cyp2g1, Cyp3a11 (Fig. 7A). After the HFD + D, the most upregulated genes included chymotrypsin like elastase 2 A (Cela2a), Cela1, Nbl1, Upk3b, scavenger receptor class A member 5 (Scara5), adhesion G protein-coupled receptor F1 (Adgrf1), Large2, Slc9a7, Fkbp5, Chmp4c. The most decreased genes included MYC proto-oncogene (Myc), Mir670hg, Gm15441, Cd3d, Themis, Gm38467, 9430085M18Rik, Gm52341, Zfp979, Rad51b, and Obp2a (Fig. 7B). After the HFD, there were 1117 DEGs with 396 upregulated genes and 720 downregulated genes compared with the ND. 1,1-DEE treatment revealed 432 DEGs in the comparison between the HFD + D and HFD groups, including 175 increased genes and 257 decreased genes (Fig. 7C).
A–E Transcriptome analysis in the liver tissue of ND, HFD, and HFD + D fed mice. Volcano plot showed DEGs for the comparison between the ND and HFD groups (A). Volcano plot illustrated DEGs for the comparison between the HFD and HFD + D groups. Yellow dots: most upregulated or downregulated genes, pink dots: upregulated genes, blue dots: downregulated genes, gray dots: non-significant genes; the x axis is log2(FC), and the y axis is -log10(p-value) (B). The bar graph showed the number of upregulated and downregulated genes in the HFD/ND and HFD + D/HFD (C). The Venn diagram showed overlapping genes in the comparison between the HFD/ND and HFD + D/HFD (D). A heatmap showed fully rescued genes after 1,1-DEE supplement (E).
To investigate overlapping genes between DEGs of HFD/ND and HFD + D/HFD, we performed the Venn diagram analysis. In these two DEGs, 90 genes were overlapping, suggesting that there were rescued genes after the treatment of 1,1-DEE (Fig. 7D). Rescued scatter plot of these overlapping genes revealed 81 fully rescued genes and 9 partially rescued genes. Fully rescued genes are divided into 43 rescued downregulated (genes are decreased in HFD and reverted after the supplement of 1,1-DEE), and 38 rescued upregulated genes (genes are increased in HFD and reverted in the HFD + D) (Supplementary Fig. 6C). The PCA plot displayed a slight shift of the HFD + D group toward the ND group, suggesting a partial effect of 1,1-DEE at transcriptome level (Supplementary Fig. 6D). A heatmap demonstrated fully rescued gene expressions and gene names (Fig. 7E). In the rescued downregulated list, there were notable genes that were downregulated in HFD/ND and were reverted in HFD + D/HFD, including scd1, pcsk9, LDL receptor related protein 1 (lrp1), leptin receptor (lepr), ppargc1b, and esr1. Additionally, prominent rescued upregulated genes are acyl-CoA thioesterase 1 (acot1), acot2, fatty acid-binding protein 1 (fabp1), and fabp2 (Fig. 7E).
Discussion
Epidemiological research has suggested the health-promoting effects of moderate wine consumption, in particular, it can reduce the risk of type-2 diabetes32,33,34, cardiovascular diseases35,36,37, and neurodegenerative diseases38,39,40, and promote longevity35,41,42,43. In this study, we unraveled the novel bioactive function of 1,1-DEE in regulation of insulin resistance and metabolic dysregulations using HFD-fed mice.
In wine, various aroma compounds are generated during aging, when developed yeast strains form biofilms (flor yeast) on the wine surface following alcoholic fermentation44,45. As aging progresses, the abundance of aroma compounds increases. This results in the development of fruity, caramel-like, or alcoholic flavors. 1,1-DEE is a key aroma compound in wine, known for its high concentration and odor level (Table 1). We identified 1,1-DEE in several batches of ethanol obtained from commercial sources, as revealed by high area values in GC–MS analysis. Additionally, because of the presence of 1,1-DEE, E1 and E4 exhibited higher PTEN oxidation than the other ethanol batches, suggesting that 1,1-DEE plays a bioactive role in addition to conferring odor and flavor.
PTEN is a negative regulator in the PI3K/Akt signaling pathway10. We observed the oxidative inactivation of PTEN after 1,1-DEE treatment, resulting in the activation of Akt through increased Akt phosphorylation in concentration- and time-dependent manners. Studies have demonstrated that the oxidative inactivation of PTEN is regulated by H2O2, a major ROS product derived from different intracellular organelles and systems. In this process, PTEN is oxidized at Cys124 to form a disulfide bond with Cys7125,27,46. Mitochondrial-derived H2O2 has been shown to regulate the redox state of PTEN through oxidation, thereby regulating angiogenesis, vascular endothelial growth factor signaling, and muscle differentiation47,48. H2O2 derived from NADPH oxidases also plays a key role in PTEN oxidation49,50. In this study, we found that 1,1-DEE increased mitochondrial ROS levels, thereby accompanying by the oxidative inactivation of PTEN. Mechanistically, by binding at the quinone (Q) site of mitochondrial complex I, 1,1-DEE partially inhibits the electron transfer from NADH to ubiquinone, causing electron leakage at the flavin mononucleotide (FMN) and iron-sulfur clusters in the NDUFS2 subunit. These leaked electrons reduce O2, generating O2 ∙ -, which is then dismutated into H2O2. This inhibition of the electron transfer is similar to the action of other well-characterized complex I inhibitors like Rotenone and Piericidin A, which also induce electron back-pressure and ROS production at the FMN/Fe-S cluster level51. Collectively, our findings indicate that 1,1-DEE mediates the production of ROS, including H2O2. This further induces the production of HCO4−, promoting PTEN oxidation.
The relationship between insulin sensitivity, glucose transportation, and metabolism has been well-established in various studies7,8,9. The PI3K/Akt signaling pathway significantly contributes to both insulin sensitivity and resistance. Dysfunction in this pathway in the skeletal muscle, adipose tissue, liver, and brain can impair Akt-mediated processes, such as glucose transportation, glycogen synthesis, and lipid synthesis, all of which contribute to insulin resistance7. We observed that supplementation with 1,1-DEE reduced HFD-induced glucose intolerance and insulin resistance by ameliorating fasting blood glucose levels, glucose tolerance, insulin tolerance, and serum insulin levels. This suggests that 1.1-DEE has a role in regulating glucose metabolism and insulin sensitivity. As PTEN/Akt signaling is activated by 1,1-DEE, there is a high possibility that 1,1-DEE regulates insulin sensitivity through the activation of the PTEN/Akt axis. We observed that Akt signaling was enhanced through 1,1-DEE-mediated oxidative inactivation of PTEN using several experimental methods, including co-treatment with 1,1-DEE and insulin, in vitro palmitate-induced insulin resistance. Thus, 1,1-DEE can enhance insulin sensitivity by activating the PTEN/Akt axis, thereby increasing glucose transportation from the extracellular space into the cytosol. A previous study described that adipocyte-specific deletion of PTEN enhanced insulin sensitivity, decreased adipose tissue inflammation, and reduced hepatic steatosis, suggesting that selective loss of PTEN in mature adipocytes is sufficient to confer systemic metabolic improvements52. Additionally, although hepatic PTEN deficiency also improved insulin sensitivity, it contributed to steatosis through increased fatty acid uptake and de novo lipogenesis53. Since 1,1-DEE selectively oxidized PTEN and activated Akt signaling in WAT but not in the liver, this effect may be sufficient for attenuation of insulin resistance without inducing liver lipotoxicity. In addition, transcriptome analysis of the liver of mice displayed enhanced enrichments in processes related to insulin signaling and growth factor signaling in the HFD + D group compared with the HFD group. Studies demonstrated that insr and irs2 genes were downregulated during the HFD54,55, leading to severe insulin resistance. Also, other genes, such as igf1r56 and fgfr157, involved in growth factor signaling can influence insulin sensitivity and glucose tolerance. cela2a encodes an enzyme responsible for reducing platelet formation and increasing insulin secretion, degradation, and sensitivity58. In previous studies, igfbp1 and igfbp5, contributing to insulin resistance and glucose intolerance, were found to be downregulated in obese conditions59,60. By improving these genes, 1,1-DEE can potentially restore glucose uptake and insulin sensitivity in addition to reactivating the PI3K/Akt axis by oxidative inactivation of PTEN.
Studies have demonstrated that HFD induces considerable weight gain in mouse models, as represented by increased adipose tissue weight and cell size61,62,63. In this study, 1,1-DEE treatment resulted in considerably lower body weight gain in the HFD + D group than in the HFD group, as demonstrated by the decrease in the weight and size of eWAT and iWAT tissues. This suggests that 1,1-DEE can control body weight gain induced by the HFD. Current weight loss drugs such as metformin and glucagon-like peptide-1 (GLP-1) receptor agonists (GLP-1 RAs) exert the anti-obesity effect by reducing food intake and appetite through several mechanisms. Metformin can trigger the production and secretion of the growth/differentiation factor 15 in peripheral tissues such as liver, distal intestine, and kidney64,65,66. This is a stress-responsive cytokine that can act via a GDNF family receptor α-like on the brainstem to suppress appetite and induce weight loss65. GLP-1 RAs, which mimic the action of endogenous GLP-1, have been widely studied in recent years for their anti-obesity effect67,68. By binding and activating GLP-1 receptors on the hypothalamus and brainstem, they promoted satiety, reduced hunger, and delayed gastric emptying, leading to the reduction in food intake69. Our results showed that the food intake levels were similar between the HFD and HFD + D groups, suggesting that 1,1-DEE prevented weight gain in HFD-fed mice independently of food intake. Additionally, 1,1-DEE treatment enhanced food intake in the ND + D group compared with the ND group without differing in body weight gain, indicating that 1,1-DEE may enhance energy expenditure to regulate weight gain. Our parallel study indicated that 1,1-DEE can activate the AMP-activated protein kinase (AMPK), resulting in enhancement in fatty acid oxidation, glycolysis, and mitochondrial biogenesis. This enhanced mitochondrial aerobic oxidative phosphorylation and ATP generation, supporting enhanced energy expenditure and ameliorated obesity induced by the HFD. AMPK activation plays a critical role in the regulation of insulin resistance70,71,72, obesity73,74, and lipid metabolism75,76. Further studies are necessary to validate the direct role of 1,1-DEE–induced AMPK activation in controlling HFD-induced obesity and diabetes.
Dyslipidemia, referring to an abnormality in the synthesis and metabolism of lipids, is also strongly associated with HFD consumption63. Histological analysis of the liver of mice in the HFD group revealed an abnormal increase in cytoplasmic vacuoles, suggesting an increase in lipid accumulation and hepatic steatosis. This was supported by increased levels of serum biomarkers associated with liver injury, triglyceride synthesis, and cholesterol synthesis, including ALT, AST, LDH, TG, T-chol, HDL-C, and LDL-C. HDL-C is considered a “good” source of cholesterol and is normally expected to decrease in individuals with obesity and metabolic syndrome. The increased HDL-C level in the HFD group compared with the ND group could be attributed to several reasons. First, mice lack cholesteryl ester transfer protein, which facilitates the transfer of cholesterol esters from HDL to LDL/VLDL77. Second, we observed a significant increase in the T-chol level, indicating that increased cholesterol flux upregulates both HDL-C and LDL-C levels. However, 1,1-DEE treatment significantly suppressed vacuolation in the liver of mice in the HFD + D group compared with the HFD group. 1,1-DEE also ameliorated the levels of serum biomarkers, including ALT, AST, LDH, TG, and LDL-C, indicating reduced hepatocyte damage and triglyceride synthesis. The T-chol level did not differ significantly between the HFD and HFD + D groups, being higher than that in the ND group. However, the LDL-C level was significantly lower in the HFD + D group than in the HFD group, suggesting that 1,1-DEE is involved in the reduction of LDL-C biosynthesis and/or increase in LDL-C clearance. In summary, 1,1-DEE can ameliorate HFD-induced body weight gain, liver damage, and dyslipidemia.
The liver plays a central role in glucose and lipid metabolism. Transcriptome analysis of the liver of mice revealed upregulated and downregulated genes in comparison between the HFD and ND groups. Derl3 encodes a component of the endoplasmic reticulum (ER)-associated degradation machinery for unfolded or misfolded glycoproteins78. Increased Derl3 in the HFD group suggests an ER stress condition, which can contribute to insulin resistance and fatty liver disease79,80,81. Cdkn1a is a cellular senescence marker that is correlated with hyperinsulinemia and fatty liver disease82,83. Additionally, cytochrome P450s (CYPs) families include enzymes that function as monooxygenases. Among these, the CYP4 family, including cyp4a10 and cyp4a14, is responsible for the ω-hydroxylation of endogenous fatty acids, which is associated with lipid accumulation (such as arachidonic acid and 20-hydroxy-eicosatetraenoic) and liver inflammation84. On the other hand, CYP2 and CYP3 families mainly function in the xenobiotic process, which is used for the elimination and modification of external particles. Therefore, the downregulation of CYP2 or CYP3 genes in the HFD mice, including Cyp2b10, Cyp2c53-ps, Cyp2c55, Cyp2g1, and Cyp3a11, suggests impaired liver detoxification85,86. The concurrent upregulation and downregulation of these genes, as shown in Fig. 7A, may reflect hepatic stress or an adaptive metabolic shift resulting in dyslipidemia and fatty liver disease. However, transcriptome analysis of liver mice in comparison between the HFD + D and HFD exhibited genes involved in the amelioration of HFD-induced dyslipidemia. Treatment with 1,1-DEE restored the key genes involved in the clearance of LDL (lrp187, and abca1 (log2FC = 0.57, p-value < 0.05, data not shown)88,89), increase in fatty acid oxidation and decrease in TG synthesis (pparδ (log2FC = 0.69, p-value < 0.05, data not shown)90,91, and lepr92), and enhancement in HDL formation (pparδ91, and abca189). It also restored several downregulated genes that participated in dyslipidemia, including acot193, and fabp194. These findings suggest that treatment with 1,1-DEE can help rescue lipid metabolic dysfunction and mitigate fatty liver disease induced by the HFD.
In this study, we elucidated the bioactive functions of 1,1-DEE in the amelioration of HFD-induced obesity, insulin resistance, and dyslipidemia. 1,1-DEE can serve as a potential therapeutic agent to prevent obesity, type-2 diabetes, and dyslipidemia.
Methods
Materials and reagents
Ethanol batches (E1, E2, E3, and E4) used in this study are listed in Supplementary Table 1. 1,1-Diethoxyethane (A902), N-ethyl maleimide (NEM, E3876), N-Acetyl-L-Cysteine (NAC, A9165), 2′,7′-Dichlorofluorescein diacetate (DCFH.DA) (35845), and hydrogen peroxide (H2O2) solution (88579) were purchased from Sigma Aldrich. 1,2-Diethoxyethane (D0456) was obtained from the Division of Tokyo Chemical Industry. MitoSOX®Red CMXRos (M36008) was purchased from Invitrogen.
Antibodies used in this study were as follows: Akt (#9272S, 1:1000), phospho-AktSer473 (#9271S, 1:1000), phospho-AktThr308 (#9275S, 1:1000), HA-tag (#3724, 1:1000), β-Actin (#4970, 1:1000) and anti-rabbit IgG horseradish peroxidase-conjugated (#7074,1:5000) antibodies were obtained from Cell Signaling Technology (Danvers, MA, USA). GAPDH antibody (LF-PA0212, 1:1000) was obtained from Ab Frontier (Daejeon, Republic of Korea).
Animal models and 1,1-DEE oral administration
Eight-week-old male C57BL-6J mice were obtained from Damool Science (Daejeon, Korea) and housed under specific pathogen-free conditions in the animal facility at the Hwasun Biomedical Convergence Center, Chonnam National University Medical School, South Korea. All animal experiments were reviewed and approved by the Animal Use and Care Committees at Chonnam National University Medical School and in accordance with institution’s guidelines for animal care and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. Mice were randomly divided into 4 groups (n = 4 per group) and fed different diets: a normal diet (ND) group, an ND with 1,1-DEE administration (ND + D) group, an HFD group, and an HFD with 1,1-DEE administration (HFD + D) group. ND contains 10 kcal% fat, 70 kcal% carbohydrate, and 20 kcal% protein, while HFD includes 60 kcal% fat, 20 kcal% carbohydrate, and 20 kcal% protein. Mice were orally treated with 100 mg/kg of 1,1-DEE every other day, whereas the control groups received equivalent amounts of water. After 8 weeks of treatment, mice were used for glucose tolerance and insulin tolerance tests. Then, mice were euthanized under deep anesthesia using isoflurane with no distress. Blood samples were obtained and kept at room temperature (RT) for 30 min. Liver and WAT tissues were collected and quickly immersed in liquid nitrogen before being stored at −80 °C for further investigations.
Cell culture, 1,1-DEE and 1,2-DEE treatments, and transient transfection of HA-tagged pCGN vector
C2C12 (CRL-1772, ATCC), HepG2 (HB-8065, ATCC), Ea.hy926 (CRL-2922, ATCC), and MEF (Dr. Eui-Ju Yeo, Gachon University) cells were cultured in Dulbeco-Modified Eagle’s Medium (DMEM, LM001-05), supplemented with 10% (v/v) fetal bovine serum (FBS, S001-01), and 1% (v/v) penicillin/streptomycin (P/S, LS 202-02) obtained from Welgene (Gyeongsan, Republic of Korea). AC16 cells (#SCC109, Millipore, CA, USA) were maintained in Dulbecco’s modified Eagle medium/nutrient mixture F-12 (DMEM/F12, LM002-05), supplemented with 12.5% (v/v) FBS, and 1% P/S (Welgene). Upon reaching 70% confluency, the cells were seeded in 6-well plates and allowed to adhere overnight. 1,1-DEE or 1,2-DEE was diluted in serum-free DMEM to predetermined concentrations before cell treatment.
For the transient transfection, the cells were seeded in 6-well plates and allowed to adhere overnight. The cells were then transfected with a HA-tagged pCGN PTEN vector containing wild-type (WT), C71S, C124S, or C71S/C124S using Lipofectamine 2000 (#11608-027, Invitrogen, Waltham, MA, USA) following the manufacturer’s protocol. In brief, DNA and Lipofectamine 2000 were mixed separately with Opti-MEM (Ref: 31985-070, Gibco Life Technologies, Grand Island, NY, USA) and incubated at room temperature (RT) for 5 min. The two mixtures were then combined and incubated at RT for another 20 min. The cells were then treated with this mixture, which was incubated for 6 h at 37 °C in 5% CO2. Next, the medium was replaced with serum-containing DMEM, and the cells were incubated further. One day after the transfection, the cells were treated with the predetermined concentrations of 1,1-DEE and subsequently used for various analyses.
Gas chromatography-mass spectrometry (GC-MS)
For gas sampling, we used a continuous irrigation system for transurethral resection of the prostate (TURP) and vaporization. The gas that developed during the cutting and cauterization was gathered in the upper portion of the bladder. The mixture of the solution used for the irrigation and gas was aspirated through the tubing into the large bottle of the vacuum pump. One part of the gas above the fluid in the bottle was directed from the out-gas vent to the tubing connected to a Tenax absorber (Tenax GR; Japan Analytical Industry, Tokyo, Japan). The flow rate of the gas directed to the Tenax absorber was maintained at 0.05 L/min using a gas flow pump (MP-S30; SIBATA, Tokyo, Japan). Approximately 45 L of gas and room air mixture was produced per hour, as measured using the Tedlar Bag (Sigma-Aldrich). For quantification, 1 L of surgical gas was directed to the Tenax absorber at a 0.05 L/min flow rate.
For gas analysis, we used the Automated Purge & Trap Sampler JTD-505III (Japan Analytical Industry) for purging and trap sampling of gas. For quantification and qualification analyses, GC/MS QP 2010 Plus (Shimadzu, Kyoto, Japan) was used.
The purge and trap conditions were as follows: 280 °C, desorption temperature; 30 min, desorption time; 50 mL/min, desorption gas flow rate; −40 °C, cold-trap temperature for sample trapping; 280 °C, pyrolysis temperature; 280 °C, transfer-line temperature; 280 °C, needle heater temper; 200 °C, cold-trap heater; 86 MPa, head press; 1.0 mL/min, column flow; and l/100, split ratio.
The GC/MS analysis conditions were as follows: DB-624 column (30 m x 0.251 mm x 1.40 mm; Agilent Technologies, Wilmington, DE, USA); 30–600 mass scan; 0–30 min, oven temperature program (40 °C for 3 min hold, 10 ml/min up to 260 °C, 5 min hold); 200 °C, ion source temperature; 250 °C, transfer line temperature; and 70 eV, EM voltage.
Freeze-drying
Ethanol samples were stored in 15 ml Falcon tubes and frozen at −80 °C for 5 h. Then, a small hole was created on every falcon tube cap. The frozen samples were freeze-dried overnight to evaporate all volatile and water-containing components. The next day, each falcon tube was then reconstituted with PBS or E2 before cell treatment.
DCFH-DA staining for ROS detection
Total intracellular ROS levels were assessed according to a previous report95. In brief, cells were seeded in a 6-well plate and incubated overnight at 37 °C in 5% CO2. After treatment, cells were washed once with serum-free DMEM and incubated with 10 µM DCFH-DA for 30 min at 37 °C. Then, cells were washed once with serum-free DMEM and twice with PBS. The cells were quickly imaged with a fluorescence microscope.
Flow cytometry analysis
Intracellular ROS and mitochondrial superoxide levels were determined using flow cytometry (FACS CANTO II, BD Biosciences, NJ, USA). After treatment with 1,1-DEE, cells were incubated with predetermined concentrations of DCFH-DA for 30 min at 37 °C. Cells were then collected using Trypsin-EDTA (#25300-062, Gibco) and washed with cold PBS. The intracellular ROS level was immediately quantified with excitation at 485 nm and emission at 530 nm. In terms of mitochondrial ROS detection, the cells were incubated with MitoSOX®Red CMXRos, collected by trypsinization, and fixed with 70% cold ethanol at 4 °C for 30 min. MitoSOX Red signal was detected at excitation and emission wavelengths of 488 and 695 nm, respectively.
Mice serum chemical analysis
After coagulating the blood at RT, blood samples were centrifuged at 3000 rpm, 4 °C for 20 min, and serum samples were transferred to Eppendorf tubes. Serum chemical analysis was performed using an automated blood chemistry analyzer (Duyeol Biotech Company, Seoul, Republic of Korea), including markers: aspartate transferase (AST), alanine transaminase (ALT), alkaline phosphatase (ALP), Lactate dehydrogenase (LDH), Total Bilirubin (T-Bili), Total cholesterol (T-chol), Triglycerides (TG), Albumin (Alb), LDL-cholesterol (LDL-C), and HDL-cholesterol (HDL-C). Serum insulin levels were measured using a mouse insulin ELISA kit (#292-89401, FUJIFILM Wako Pure Chemical Corporation, Japan).
Intraperitoneal glucose tolerance (GTT) and insulin tolerance test (ITT)
For the glucose tolerance test, mice were fasted for 12 h overnight by transferring them to clean cages without access to food. Fasting blood glucose was measured using a glucose meter. Then, each mouse was intraperitoneally injected with a dose of 2 g/kg of D-glucose. Blood glucose levels were monitored at 15, 30, 60, 90, and 120 min post-injection.
For the insulin tolerance test, mice were starved for 4 h. Mice were intraperitoneally injected with insulin (Humulin R) with a dose of 0.6 U/kg. The blood glucose levels were recorded at 15, 30, 60, 90, and 120 min after insulin injection using the glucose meter.
Histological analysis
Liver and WAT tissues from each mouse were obtained and fixed in 4% of paraformaldehyde. Tissues were cut into 5-mm-thick sections before being embedded in paraffin. Paraffin sections were stained with hematoxylin and eosin (H&E). Representative images were obtained at 20X magnification using Leica ProgRes microscope, Aperio Digital Pathology Slide Scanner (Leica Microsystems, CMS GmbH, Wetzlar, Germany) (scale bars: 100 µM).
Western blot analysis and antibodies
The redox state of PTEN was analyzed as previously described in ref. 24. In brief, cells were lysed with a lysis buffer containing 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 5% glycerol, 0.1% NP-40, phosphatase inhibitors, protease inhibitors, and 10 mM NEM. Cell lysates were sonicated and centrifuged at 13,000 rpm for 10 min. Next, supernatants were collected, and protein concentrations were measured using the PierceTM BCA Protein Assay Kit (Thermo Fisher Scientific, Waltham, MA, USA). The lysates were mixed with either a reducing sample buffer containing 60 mM Tris (pH 6.8), 25% glycerol, 2% SDS, 5% 2-mercaptoethanol, and 0.5% bromophenol or a non-reducing sample buffer with the same composition, excluding 2-mercaptoethanol. Samples were then subjected to SDS-PAGE and immunoblotted with a PTEN-specific antibody. To investigate the expressions of other proteins, cell lysates were mixed with the described reducing sample buffer before being subjected to SDS-PAGE and immunoblotted with different specific antibodies.
Tissues were lysed using NP-40 lysis buffer (RIPA, 1% NP-40, 10 mM NEM). Tissue extracts were kept on ice for 1 h, followed by centrifugation at 13,000 rpm for 30 min. Supernatants were collected, and protein concentrations were measured using BCA kit. Reducing and non-reducing sample buffers were added to make corresponding protein samples. Samples were then subjected to SDS-PAGE and immunoblotted with specific antibodies.
RNA-sequencing (RNA-seq) analysis
The quality of total isolated RNA was assessed using the Agilent 2100 Bioanalyzer (Agilent Technologies), and only samples with an RNA Integrity Number (RIN) ≥ 8.0 were used for library preparation. We used the TruSeq mRNA Sample Preparation Kit (Illumina) for mRNA-seq library, following the manufacturer’s instructions (TruSeq Stranded mRNA Reference Guide #1000000040498 v00). The constructed library was sequenced using a HiSeq 4000 sequencing system, resulting in 2 × 100 bp sequencing reads.
Bioinformatic analysis of RNA-seq data
Sequencing reads were assessed for quality and checked for primer/adaptor sequence contamination using FastQC (v0.11.7). The reads were trimmed using TrimGalore using a threshold of an average sequence quality threshold of > 30. The resulting trimmed reads were aligned with the mouse reference genome GRCm38 using STAR (STAR-2.7.0). The HTseq-count algorithm and ensemble Gencode m25 gene annotations were used to profile the gene expression from the raw read counts. Finally, EdgeR was used to convert the count data to counts per million (cpm). If multiple entries mapped to the same gene symbol, redundant genes were combined by averaging their CPM values. RNA expression data were normalized as previously described in ref. 96. In this study, differentially expressed genes (DEGs) were filtered as having a p-value ≤ 0.05, and log2 fold change (log2FC) ≤ −0.5 or ≥ 0.5. Heatmaps, volcano plots, Venn diagram, and rescued scatter plots were generated using R packages. Several packages were used such as pheatmap, ggplot2, VennDiagram, principal component analysis (PCA), and others. Gene set enrichment analysis (GSEA) was performed with GSEA desktop v4.4.0 application using preranked lists generated from log2FC values. The analysis concentrated on Gene Ontology biological process (GOBP), using the m5.go.bp.v2025.1.Mm.symbols.gmt gene set.
Molecular docking simulation
A cryo-EM structure of human respiratory complex I (hRCI) (PDB ID: 5XTD) was selected for molecular docking analysis. Three domains of hRCI configuring the binding site of piericidin A, were used for the docking analysis as macromolecular docking targets on AutoDock Vina software, including NADH dehydrogenase [ubiquinone] iron-sulfur protein 2 (NDUFS2), NADH dehydrogenase [ubiquinone] iron-sulfur protein 7 (NDUFS7), and NADH-ubiquinone oxidoreductase chain 1 (ND1). The ligand (1,1-DEE) was retrieved from ChEBI database. The docking grid was centered on the corresponding region of the piericidin A binding site and set with the following grid parameters: 30 Å × 30 Å × 30 Å with the sampling exhaustivity of 4.
Quantification and statistical analysis
The western blot protein bands were quantified via densitometry using ImageJ 1.50i (National Institutes of Health, Bethesda, MD, USA). All values are expressed as mean ± standard error of the mean (SEM). Statistical significance was analyzed via unpaired two-tailed Student’s t-tests for two-group comparisons, and one-way or two-way ANOVA for multiple-group comparisons using GraphPad Prism (GraphPad, San Diego, CA, USA). A p-value of < 0.05 was considered statistically significant.
Data availability
The data generated in this study are available within the paper and Supplementary information. The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Code availability
The code using in the current study is available from the corresponding author upon reasonable request.
References
Piché, M. E., Tchernof, A. & Després, J. P. Obesity Phenotypes, Diabetes, and Cardiovascular Diseases. Circ. Res 126, 1477–1500 (2020).
Kahn, B. B. & Flier, J. S. Obesity and insulin resistance. J. Clin. Invest 106, 473–481 (2000).
Clemente-Suárez, V. J., Beltrán-Velasco, A. I., Redondo-Flórez, L., Martín-Rodríguez, A. & Tornero-Aguilera, J. F. Global Impacts of Western Diet and Its Effects on Metabolism and Health: A Narrative Review. Nutrients 15, 2749 (2023).
Kleinert, M. et al. Animal models of obesity and diabetes mellitus. Nat. Rev. Endocrinol. 14, 140–162 (2018).
Kokkorakis, M. et al. Emerging pharmacotherapies for obesity: A systematic review. Pharm. Rev. 77, 100002 (2025).
Henderson Lewis, K., Sloan, C. E., Bessesen, D. H. & Arterburn, D. Effectiveness and safety of drugs for obesity. Bmj 384, e072686 (2024).
Huang, X., Liu, G., Guo, J. & Su, Z. The PI3K/AKT pathway in obesity and type 2 diabetes. Int J. Biol. Sci. 14, 1483–1496 (2018).
Savova, M. S., Mihaylova, L. V., Tews, D., Wabitsch, M. & Georgiev, M. I. Targeting PI3K/AKT signaling pathway in obesity. Biomed. Pharmacother. 159, 114244 (2023).
Hopkins, B. D., Goncalves, M. D. & Cantley, L. C. Insulin-PI3K signalling: an evolutionarily insulated metabolic driver of cancer. Nat. Rev. Endocrinol. 16, 276–283 (2020).
Lee, Y. R., Chen, M. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor: new modes and prospects. Nat. Rev. Mol. Cell Biol. 19, 547–562 (2018).
Loh, K. et al. Reactive oxygen species enhance insulin sensitivity. Cell Metab. 10, 260–272 (2009).
Tiganis, T. Reactive oxygen species and insulin resistance: the good, the bad and the ugly. Trends Pharm. Sci. 32, 82–89 (2011).
Giacosa, A. et al. Mediterranean Way of Drinking and Longevity. Crit. Rev. Food Sci. Nutr. 56, 635–640 (2016).
Hrelia, S. et al. Moderate Wine Consumption and Health: A Narrative Review. Nutrients 15, 175 (2022).
Marcq, P. & Schieberle, P. Characterization of the Key Aroma Compounds in a Commercial Fino and a Commercial Pedro Ximénez Sherry Wine by Application of the Sensomics Approach. J. Agric Food Chem. 69, 5125–5133 (2021).
Marcq, P. & Schieberle, P. Characterization of the key aroma compounds in a commercial Amontillado sherry wine by means of the sensomics approach. J. Agric Food Chem. 63, 4761–4770 (2015).
Chen, S., Wang, C., Qian, M., Li, Z. & Xu, Y. Characterization of the Key Aroma Compounds in Aged Chinese Rice Wine by Comparative Aroma Extract Dilution Analysis, Quantitative Measurements, Aroma Recombination, and Omission Studies. J. Agric Food Chem. 67, 4876–4884 (2019).
Tan, C. et al. The Effects of Ultrasonic and Gamma Irradiation on the Flavor of Potato Wines Investigated by Sensory Omics. Foods 12, https://doi.org/10.3390/foods12152821 (2023).
Dunkel, A. et al. Nature’s chemical signatures in human olfaction: a foodborne perspective for future biotechnology. Angew. Chem. Int Ed. Engl. 53, 7124–7143 (2014).
Moreno, J., Moreno-García, J., López-Muñoz, B., Mauricio, J. C. & García-Martínez, T. Use of a flor velum yeast for modulating colour, ethanol and major aroma compound contents in red wine. Food Chem. 213, 90–97 (2016).
Callejón, R. M., Morales, M. L., Troncoso, A. M. & Silva Ferreira, A. C. Targeting key aromatic substances on the typical aroma of sherry vinegar. J. Agric Food Chem. 56, 6631–6639 (2008).
Pino, J. A., Tolle, S., Gök, R. & Winterhalter, P. Characterisation of odour-active compounds in aged rum. Food Chem. 132, 1436–1441 (2012).
Franitza, L., Granvogl, M. & Schieberle, P. Characterization of the Key Aroma Compounds in Two Commercial Rums by Means of the Sensomics Approach. J. Agric Food Chem. 64, 637–645 (2016).
Lee, S. R. et al. Reversible inactivation of the tumor suppressor PTEN by H2O2. J. Biol. Chem. 277, 20336–20342 (2002).
Han, S. J. et al. Assay of the redox state of the tumor suppressor PTEN by mobility shift. Methods 77-78, 58–62 (2015).
Zhang, Y. et al. Redox regulation of tumor suppressor PTEN in cell signaling. Redox Biol. 34, 101553 (2020).
Trinh, V. H. et al. Redox Regulation of Phosphatase and Tensin Homolog by Bicarbonate and Hydrogen Peroxide: Implication of Peroxymonocarbonate in Cell Signaling. Antioxid. ((Basel)) 13, 473 (2024).
Sies, H. Hydrogen peroxide as a central redox signaling molecule in physiologic al oxidative stress: Oxidative eustress. Redox Biol. 11, 613–619 (2017).
Schieber, M. & Chandel, N. S. ROS function in redox signaling and oxidative stress. Curr. Biol. 24, R453-R462 (2014).
Suomalainen, A. & Nunnari, J. Mitochondria at the crossroads of health and disease. Cell 187, 2601–2627 (2024).
Krako Jakovljevic, N. et al. In vitro models of insulin resistance: Mitochondrial coupling is differently affected in liver and muscle cells. Mitochondrion 61, 165–173 (2021).
Baliunas, D. O. et al. Alcohol as a risk factor for type 2 diabetes: A systematic review and meta-analysis. Diab. Care 32, 2123–2132 (2009).
Chiva-Blanch, G. et al. Effects of red wine polyphenols and alcohol on glucose metabolism and the lipid profile: a randomized clinical trial. Clin. Nutr. 32, 200–206 (2013).
da Luz, P. L. et al. Coronary artery plaque burden and calcium scores in healthy men adhering to long-term wine drinking or alcohol abstinence. Braz. J. Med Biol. Res 47, 697–705 (2014).
Gronbaek, M. et al. Mortality associated with moderate intakes of wine, beer, or spirits. Bmj 310, 1165–1169 (1995).
Grønbæk, M. et al. Type of alcohol consumed and mortality from all causes, coronary heart disease, and cancer. Ann. Intern Med 133, 411–419 (2000).
Levantesi, G. et al. Wine consumption and risk of cardiovascular events after myocardial infarction: results from the GISSI-Prevenzione trial. Int J. Cardiol. 163, 282–287 (2013).
Letenneur, L. Risk of dementia and alcohol and wine consumption: a review of recent results. Biol. Res 37, 189–193 (2004).
Lucerón-Lucas-Torres, M. et al. Association Between Wine Consumption and Cognitive Decline in Older People: A Systematic Review and Meta-Analysis of Longitudinal Studies. Front Nutr. 9, 863059 (2022).
Truelsen, T., Grønbæk, M., Schnohr, P. & Boysen, G. Intake of beer, wine, and spirits and risk of stroke: the Copenhagen City Heart Study. Stroke 29, 2467–2472 (1998).
Gea, A. et al. Mediterranean alcohol-drinking pattern and mortality in the SUN (Seguimiento Universidad de Navarra) Project: a prospective cohort study. Br. J. Nutr. 111, 1871–1880 (2014).
Jayasekara, H., English, D. R., Room, R. & MacInnis, R. J. Alcohol consumption over time and risk of death: a systematic review and meta-analysis. Am. J. Epidemiol. 179, 1049–1059 (2014).
Grønbæk, M. et al. Changes in alcohol intake and mortality: a longitudinal population-based study. Epidemiology 15, 222–228 (2004).
Lyu, J., Chen, S., Nie, Y., Xu, Y. & Tang, K. Aroma release during wine consumption: Factors and analytical approaches. Food Chem. 346, 128957 (2021).
David-Vaizant, V. & Alexandre, H. Flor Yeast Diversity and Dynamics in Biologically Aged Wines. Front Microbiol 9, 2235 (2018).
Davies, M. J. et al. Effects of moderate alcohol intake on fasting insulin and glucose concentrations and insulin sensitivity in postmenopausal women: a randomized controlled trial. JAMA 287, 2559–2562 (2002).
Connor, K. M. et al. Mitochondrial H2O2 regulates the angiogenic phenotype via PTEN oxidation. J. Biol. Chem. 280, 16916–16924 (2005).
Kim, J. H. et al. Mitochondrial ROS-derived PTEN oxidation activates PI3K pathway for mTOR-induced myogenic autophagy. Cell Death Differ. 25, 1921–1937 (2018).
Seo, J. H., Ahn, Y., Lee, S. R., Yeo, C. Y. & Hur, K. C. The major target of the endogenously generated reactive oxygen species in response to insulin stimulation is phosphatase and tensin homolog and not phosphoinositide-3 kinase (PI-3 kinase) in the PI-3 kinase/Akt pathway. Mol. Biol. Cell 16, 348–357 (2005).
Cai, B., Liu, M., Li, J., Xu, D. & Li, J. Cigarette smoke extract amplifies NADPH oxidase-dependent ROS production to inactivate PTEN by oxidation in BEAS-2B cells. Food Chem. Toxicol. 150, 112050 (2021).
Fato, R. et al. Differential effects of mitochondrial Complex I inhibitors on production of reactive oxygen species. Biochim Biophys. Acta 1787, 384–392 (2009).
Morley, T. S., Xia, J. Y. & Scherer, P. E. Selective enhancement of insulin sensitivity in the mature adipocyte is sufficient for systemic metabolic improvements. Nat. Commun. 6, 7906 (2015).
Peyrou, M. et al. Hepatic PTEN deficiency improves muscle insulin sensitivity and decreases adiposity in mice. J. Hepatol. 62, 421–429 (2015).
Gan, K.-X. Mitofusin-2 ameliorates high-fat diet-induced insulin resistance in li ver of rats. World J. Gastroenterol. 19, 1572–1581 (2013).
Thakur, S., Rawat, P., Dehury, B. & Mondal, P. TRIM32 regulates insulin sensitivity by controlling insulin receptor degradation in the liver. EMBO Rep. 26, 791–809 (2025).
Haywood, N. J. et al. Endothelial IGF-1 receptor mediates crosstalk with the gut wall to regulate microbiota in obesity. EMBO Rep. 22, e50767 (2021).
Geng, L. et al. Exercise Alleviates Obesity-Induced Metabolic Dysfunction via Enhancing FGF21 Sensitivity in Adipose Tissues. Cell Rep. 26, 2738–2752.e4 (2019).
Esteghamat, F. et al. CELA2A mutations predispose to early-onset atherosclerosis and metabolic syndrome and affect plasma insulin and platelet activation. Nat. Genet. 51, 1233–1243 (2019).
Minchenko, D. O., Tsymbal, D. O., Davydov, V. V. & Minchenko, O. H. Expression of genes encoding IGF1, IGF2, and IGFBPs in blood of obese adolescents with insulin resistance. Endocr. Regul. 53, 34–45 (2019).
Rhee, J. S. et al. Diet-induced obesity impairs endometrial stromal cell decidualization: a potential role for impaired autophagy. Hum. Reprod. 31, 1315–1326 (2016).
Zhai, Z., Yang, Y., Chen, S. & Wu, Z. Long-Term Exposure to Polystyrene Microspheres and High-Fat Diet-Induced Obesity in Mice: Evaluating a Role for Microbiota Dysbiosis. Environ. Health Perspect. 132, 97002 (2024).
Gao, Y., Guo, M., Chen, J., Sun, Y. & Wang, M. A ginseng polysaccharide protects intestinal barrier integrity in high-fat diet-fed obese mice. Int J. Biol. Macromol. 277, 133976 (2024).
Wei, B. et al. Probiotic-fermented tomato alleviates high-fat diet-induced obesity in mice: Insights from microbiome and metabolomics. Food Chem. 436, 137719 (2024).
Day, E. A. et al. Metformin-induced increases in GDF15 are important for suppressing appetite and promoting weight loss. Nat. Metab. 1, 1202–1208 (2019).
Coll, A. P. et al. GDF15 mediates the effects of metformin on body weight and energy balance. Nature 578, 444–448 (2020).
Zhang, S.-Y. et al. Metformin triggers a kidney GDF15-dependent area postrema axis to regulate food intake and body weight. Cell Metab. 35, 875–886.e5 (2023).
Wong, H. J. et al. Efficacy of GLP-1 Receptor Agonists on Weight Loss, BMI, and Waist Circumference for Patients With Obesity or Overweight: A Systematic Review, Meta-analysis, and Meta-regression of 47 Randomized Controlled Trials. Diab. Care 48, 292−300 (2025).
Alfaris, N. et al. GLP-1 single, dual, and triple receptor agonists for treating type 2 diabetes and obesity: a narrative review. EClin. Med. 75, 102782 (2024).
Moiz, A. et al. Mechanisms of GLP-1 Receptor Agonist-Induced Weight Loss: A Review of Central and Peripheral Pathways in Appetite and Energy Regulation. Am. J. Med 138, 934–940 (2025).
Yan, J. et al. Catalpol ameliorates hepatic insulin resistance in type 2 diabetes through acting on AMPK/NOX4/PI3K/AKT pathway. Pharm. Res 130, 466–480 (2018).
Zhang, Y. et al. Sulforaphane alleviates high fat diet-induced insulin resistance via AMPK/Nrf2/GPx4 axis. Biomed. Pharmacother. 152, 113273 (2022).
Entezari, M. et al. AMPK signaling in diabetes mellitus, insulin resistance and diabetic complications: A pre-clinical and clinical investigation. Biomed. Pharmacother. 146, 112563 (2022).
Peng, C. et al. Macrophage SUCLA2 coupled glutaminolysis manipulates obesity through AMPK. Nat. Commun. 16, 1738 (2025).
Fang, C. et al. The AMPK pathway in fatty liver disease. Front Physiol. 13, 970292 (2022).
Dong, J. et al. ACACA reduces lipid accumulation through dual regulation of lipid metabolism and mitochondrial function via AMPK- PPARα- CPT1A axis. J. Transl. Med 22, 196 (2024).
Gao, L. et al. CPI-613 rewires lipid metabolism to enhance pancreatic cancer apoptosis via the AMPK-ACC signaling. J. Exp. Clin. Cancer Res 39, 73 (2020).
Barter, P. J. et al. Cholesteryl ester transfer protein: a novel target for raising HDL and inhibiting atherosclerosis. Arterioscler Thromb. Vasc. Biol. 23, 160–167 (2003).
Eura, Y., Miyata, T. & Kokame, K. Derlin-3 Is Required for Changes in ERAD Complex Formation under ER Stress. Int. J. Mol. Sci. 21, https://doi.org/10.3390/ijms21176146 (2020).
Pagliassotti, M. J. Endoplasmic reticulum stress in nonalcoholic fatty liver disease. Annu Rev. Nutr. 32, 17–33 (2012).
Gentile, C. L., Frye, M. & Pagliassotti, M. J. Endoplasmic reticulum stress and the unfolded protein response in nonalcoholic fatty liver disease. Antioxid. Redox Signal 15, 505–521 (2011).
Belmont, P. J. et al. Roles for endoplasmic reticulum-associated degradation and the novel endoplasmic reticulum stress response gene Derlin-3 in the ischemic heart. Circ. Res 106, 307–316 (2010).
Aravinthan, A. et al. Gene polymorphisms of cellular senescence marker p21 and disease progression in non-alcohol-related fatty liver disease. Cell Cycle 13, 1489–1494 (2014).
Meijnikman, A. S. et al. Hyperinsulinemia Is Highly Associated With Markers of Hepatocytic Senescence in Two Independent Cohorts. Diabetes 71, 1929–1936 (2022).
Zhou, M., Li, J., Xu, J., Zheng, L. & Xu, S. Exploring human CYP4 enzymes: Physiological roles, function in diseases and focus on inhibitors. Drug Discov. Today 28, 103560 (2023).
Sun, S. et al. E4bp4-Cyp3a11 axis in high-fat diet-induced obese mice with weight fluctuation. Nutr. Metab. ((Lond.)) 21, 30 (2024).
He, Y. et al. High fat diet significantly changed the global gene expression profile involved in hepatic drug metabolism and pharmacokinetic system in mice. Nutr. Metab. ((Lond.)) 17, 37 (2020).
Rohlmann, A., Gotthardt, M., Hammer, R. E. & Herz, J. Inducible inactivation of hepatic LRP gene by cre-mediated recombination confirms role of LRP in clearance of chylomicron remnants. J. Clin. Investig. 101, 689–695 (1998).
Bodzioch, M. et al. The gene encoding ATP-binding cassette transporter 1 is mutated in Tangier disease. Nat. Genet. 22, 347–351 (1999).
Oram, J. F. & Heinecke, J. W. ATP-binding cassette transporter A1: a cell cholesterol exporter that protects against cardiovascular disease. Physiol. Rev. 85, 1343–1372 (2005).
Oliver, W. R. Jr. et al. A selective peroxisome proliferator-activated receptor delta agonist promotes reverse cholesterol transport. Proc. Natl. Acad. Sci. USA. 98, 5306–5311 (2001).
Barish, G. D. PPAR delta: a dagger in the heart of the metabolic syndrome. J. Clin. Investig. 116, 590–597 (2006).
Shimabukuro, M. et al. Direct antidiabetic effect of leptin through triglyceride depletion of tissues. Proc. Natl. Acad. Sci. USA. 94, 4637–4641 (1997).
Heden, T. D. et al. ACOT1 deficiency attenuates high-fat diet-induced fat mass gain by increasing energy expenditure. JCI Insight 8, e160987 (2023).
Yu, M. et al. Berberine alleviates lipid metabolism disorders via inhibition of mitochondrial complex I in gut and liver. Int. J. Biol. Sci. 17, 1693–1707 (2021).
Kim, H. & Xue, X. Detection of Total Reactive Oxygen Species in Adherent Cells by 2’,7’-Dichlorodihydrofluorescein Diacetate Staining. J. Vis. Exp. https://doi.org/10.3791/60682 (2020).
Park, S. J. et al. Proteogenomic Characterization Reveals Estrogen Signaling as a Target for Never-Smoker Lung Adenocarcinoma Patients without EGFR or ALK Alterations. Cancer Res 84, 1491–1503 (2024).
Huang, Q. et al. Effects of storage time on flavor characteristics of bran-free fermented Baijiu by using electronic sensory, descriptive sensory analysis, GC × GC-MS, and ICP-MS. Food Chem. X 23, 101667 (2024).
Cicha-Wojciechowicz, D., Frank, S., Steinhaus, M. & Majcher, M. A. Key Odorants Forming Aroma of Polish Mead: Influence of the Raw Material and Manufacturing Processes. J. Agric Food Chem. 72, 10548–10557 (2024).
Willner, B., Granvogl, M. & Schieberle, P. Characterization of the key aroma compounds in Bartlett pear brandies by means of the sensomics concept. J. Agric Food Chem. 61, 9583–9593 (2013).
Wang, J. et al. Succession patterns of aroma components during brewing process of broomcorn millet (Panicum miliaceum L.) Huangjiu. Food Res Int 154, 110982 (2022).
Acknowledgements
This study was supported by the Basic Research Program of National Research Foundation of Korea (RS-2018-NR033232 and RS-2022-NR067329). Thang Nguyen Huu is also supported in part by the Center for Global Future Biomedical Scientists at Chonnam National University. We are grateful to Professor Jae-Sang Kim for mentorship.
Author information
Authors and Affiliations
Contributions
S.-R.L. and T.N.H. designed the study. T.N.H. and H.D.T. performed the experiments. M-K.K. conducted the molecular simulation. T.N.H., H.D.T., M-K.K. analyzed the data. T.N.H. wrote the manuscript. All authors read and revised the manuscript, figures, and tables prior to submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nguyen Huu, T., Duong Thanh, H., Kim, MK. et al. 1,1-Diethoxyethane increases insulin sensitivity and ameliorates obesity and dyslipidemia in mice fed high-fat diet. npj Metab Health Dis 4, 2 (2026). https://doi.org/10.1038/s44324-025-00094-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44324-025-00094-x
This article is cited by
-
1,1-Diethoxyethane enhances aerobic respiration in human mitochondria via activation of AMP-activated protein kinase
Communications Biology (2026)