Abstract
Single-cell sequencing (scRNA-seq) is transforming our ability to dissect cell type-specific gene regulatory networks underlying developmental processes and cellular differentiation with unprecedented resolution. By leveraging single-cell transcriptomes of Arabidopsis thaliana shoot and root apical meristems, we aimed to dissect the complex spatiotemporal interplay between stem cell identity and hypoxia. Single-cell transcriptomic profiling demonstrated a substantial overlap between hypoxia-responsive gene expression and stem cell markers across shoot and root meristems. Trajectory inference coupled with RNA velocity revealed that hypoxia-related transcriptional signatures are enriched in undifferentiated stem cells and gradually decline as differentiation progresses, supporting a critical role for hypoxia in cell fate determination. Our findings suggest that the activation of hypoxia signaling may be a conserved feature in the shoot and root apical meristems of Arabidopsis thaliana and could play a crucial role in regulating stem cell fate decisions during development.
Similar content being viewed by others
Introduction
Complex multicellular organisms depend on oxygen for growth and development1. Oxygen is required for ATP production through cellular respiration, and its scarcity poses a significant challenge to plant survival. However, oxygen deprivation (hypoxia) also functions as a developmental signal that drives cell fate and tissue patterning. In animal systems, hypoxia regulates stem cell maintenance and differentiation2. Stem cells are undifferentiated cells that can self-renew and differentiate into specialized cell types. In plants, stem cells reside within specialized tissues known as meristems, which enable the continuous generation of new organs throughout the plant’s life. In the shoot apical meristem (SAM), the population of stem cells is located in the upper three cell layers at the central zone3. In the root apical meristem (RAM), the stem cell niche surrounds the quiescent center (QC) and sustains the formation of all root tissues. Stem cell division in plant meristems produces daughter cells that proliferate and eventually differentiate, generating new tissues and organs4.
Molecular oxygen is emerging as a signaling molecule essential for proper plant meristematic activity and development5. Recent research has demonstrated the role of hypoxia in the activity of meristems and stem cell identity in plant systems6,7. In vivo imaging using fluorescence-based hypoxia-responsive reporters has revealed that stem cells located in both the SAM and lateral root primordia (LRP) display a transcriptional activation indicative of hypoxic responses, even under normoxic environmental conditions6,7. Direct oxygen tension measurements revealed that the central zone of the SAM resides within a hypoxic niche6. Within this spatial domain, characterized by an oxygen tension of ~5%, an oxygen-labile regulatory protein known as LITTLE ZIPPER 2 (ZPR2) evades proteasomal degradation through the PROTEOLYSIS6 (PRT6) N-degron pathway6. Stabilization of ZPR2 enables the repression of stem cell differentiation and preserves stemness. To date, ZPR2 is the only known oxygen-sensitive protein that regulates the function of the Arabidopsis shoot apical meristem. Furthermore, another oxygen-labile protein, VERNALIZATION 2 (VRN2), is stabilized in the hypoxic shoot apex and emerging leaves of Arabidopsis, where it negatively regulates cell expansion8,9. These recent findings suggest that oxygen gradients within plant meristems are key factors for proper plant development and could direct stem cell fate. However, it remains unclear how, and to what extent, hypoxia shapes the spatiotemporal regulation of gene expression involved in stem cell identity and differentiation in plant meristems. This gap is mainly due to the challenge of isolating individual transcriptomes of stem cells, which hinders our understanding of the role of hypoxia signaling in plant meristems.
Single-cell RNA sequencing (scRNA-seq) has revolutionized the study of gene regulatory networks at a cellular level10. scRNA-seq allows for the capture of individual cell transcriptomes, revealing cellular heterogeneity within tissues and organs. As a result, it enables the identification of novel cell types and facilitates the dissection of complex cell type-specific gene regulatory networks that regulate development. Therefore, scRNA-seq presents an exciting opportunity to explore the emerging role of hypoxia in plant meristems at an unprecedented resolution. Single-cell approaches require plant cells to dissociate from their rigid cell walls, which is mainly achieved through protoplast preparation or nuclei isolation protocols11. Cell isolation procedures expose cells that are otherwise part of a hypoxic niche to atmospheric oxygen, which may result in the loss of hypoxia-associated transcriptional signatures found in stem cells or other cells that typically reside in hypoxic tissue niches.
Here, we investigated whether hypoxia-associated transcriptional signatures in plants could be captured and analyzed through the lens of single-cell genomics. To this end, we leveraged available scRNA-seq datasets from Arabidopsis shoot and root apical meristems12,13 to explore the relationship between stemness and hypoxia at single-cell resolution. By integrating trajectory analysis, pseudotime reconstruction, and RNA velocity, we examined the transcriptional dynamics of hypoxia-responsive genes across different cell lineages of shoot and root apices. Our study demonstrates that scRNA-seq is a promising technology for exploring hypoxia signaling at single-cell resolution and identifying key hypoxia-responsive regulators of plant development.
Results
Hypoxia-responsive genes are enriched in the stem cells of shoot and root apices
To explore hypoxia-related transcriptional signatures in Arabidopsis stem cells, we leveraged single-cell RNA sequencing datasets from two independent studies that characterized the transcriptional landscapes of the vegetative shoot and root apices12,13. Both experiments relied on protoplasting for cellular isolation. To ensure analytical consistency, the datasets were reanalyzed de novo, starting from the original FASTQ files12,13. Following quality control and filtering, dimensionality reduction using Uniform Manifold Approximation and Projection (UMAP), coupled with unsupervised clustering, resolved discrete cell populations within both shoot and root datasets. After clustering, we reconstructed the cellular complexity of the shoot and root apices of Arabidopsis, identifying 19 and 20 distinct clusters for the SAM and RAM, respectively (Supplementary Fig. 1a and b).
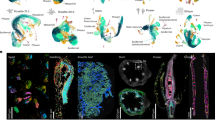
Cell clusters were annotated based on genes previously characterized to be specifically expressed in a specific cell-type or tissue12,13 (Fig. 1b and e, Supplementary Table 1). For instance, PROTODERMAL FACTOR 1 (PDF1), a gene expressed in the shoot epidermis13, shows high expression levels in clusters 1, 9, 12, and 15. This indicates that these clusters are part of the shoot epidermis (Fig. 1a and b). In the root apex, expression of DOF AFFECTING GERMINATION 1 (DAG1), a phloem marker gene14, was localized in clusters 0, 3, 6, and 9, indicating that these clusters are part of the root phloem (Fig. 1d and e). Following this strategy and based on the expression of several marker genes (Fig. 1c and f; Supplementary Tables 2 and 3), cluster annotation led to the identification of the expected different cell types previously found in the shoot and root apices12,13 (Fig. 1a and b). The 19 clusters in the shoot apex were attributed to 9 distinct cell types (Fig. 1a and c), whereas the 20 clusters identified for the root apex belonged to 10 different cell populations.
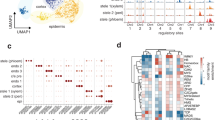
a, d UMAP plots showing the distribution of the 41270 cells for the (a) shoot apex and 14501 for the (b) root apex, which were grouped in 19 and 20 clusters for the shoot and root apices, respectively. Clusters were grouped and colored according to their annotation to a specific tissue or cell type. b, e Feature plots showing the spatial expression patterns of representative cell-type marker genes plotted on UMAPs of the (b) shoot and (e) root apices. c, f Expression of known cell-type marker genes in the Arabidopsis (c) shoot and (f) root apices. The dot diameter represents the percentage of cluster cells expressing a cell-type marker gene. The complete list of marker genes used for shoot and root apices cluster annotations is in Supplementary Tables 1, 2, and 3. Dot plots illustrating marker gene expression across clusters. Abbreviations for shoot apex tissues or cell-types: Epidermis (Ep); Guard cells (Gc); Proliferating cells (Pc); Meristem (M); Mesophyll (Mp); Phloem (Ph); Xylem (Xm); Companion cells (Cc); Stem cells (St). Abbreviations for root apex tissues or cell-types: Root cap (Rc); Root hairs (Rh); Epidermis (Ep); Meristem (M); Root cortex (Rx); Stem cells (St); Endodermis (En); Xylem (Xm); Phloem (Ph); Pericycle (Pc); Sieve elements (Se).
Stem cells reside within plant meristems and, if hypoxic, are expected to activate hypoxia-responsive gene expression6,7. Thus, after identifying stem cell populations within the shoot and root apices, we analyzed whether stem cell marker genes could be co-expressed with the core of “hypoxia responsive genes” (HRGs)15 (Supplementary Table 4). Co-expression analyses between HRGs and stem cell markers (SCMs; Supplementary Table 5) revealed an overlap between HRGs and SCMs, as indicated by the percentage of cells simultaneously expressing both SCM-HRG pair (Fig. 2a and b). Notably, the proportion of cells co-expressing both SCMs and HRGs was higher in the root compared to the shoot apex (Fig. 2a and b). On the contrary, HRGs showed lower co-expression with markers of differentiated tissues in both shoot and root apices compared with SCMs (Supplementary Fig. 2). Moreover, expression of HRGs in stem cells, defined as cells expressing ≥10 out of 30 stem cell markers (Supplementary Table 6), revealed that 27 and 35 out of 49 HRGs are expressed in stem cells of shoot and root apices, respectively (Supplementary Fig. 3). Spatial distribution of representative SCMs and HRGs on the UMAP (Fig. 2c-f) and their expression levels across the clusters (Supplementary Fig. 4) showed a broad overlap between SCMs and HRGs, confirming that HRGs are abundantly expressed in the stem cells of shoot and root apices. Moreover, violin plots of representative HRGs showed how hypoxia responses are not uniquely activated in stem cells of shoot and root apices (Supplementary Fig. 4). Indeed, HRG expression was also high in the vascular tissues of the root apex, in line with previous reports that measured low oxygen levels in the vasculature of different plant species16,17,18.
a, b Heatmaps showing the overlap of SCMs and HRGs in (a) shoot and (b) root apices. 45 and 49 HRGs were co-expressed with at least one SCM in the shoot and root apices, respectively. Scale bar indicates the proportion of cells expressing each SCM that also express the corresponding HRG relative to the whole cell population expressing a given SCM. A darker red color indicates high co-expression of the HRG-SCM pair. c–f UMAP feature plots displaying the spatial expression of representative SCMs and HRGs in the (c, e) shoot and (d, f) root apices.
Pseudotime analysis reveals early activation of hypoxia-related genes during differentiation
To investigate the relationship between stemness and hypoxia during cell differentiation in shoot and root apices, we employed pseudotime analysis, which enables the ordering of cells based on the gradual transition of their transcriptomes, placing them along a trajectory of cell differentiation (Fig. 3a and b, Supplementary Fig. 5). For pseudotime analysis, the complete single-cell transcriptomic datasets from the shoot and root apices (Fig. 1) were used to capture the transcriptional transitions across all cell types present in the apices. We first defined the origin of the developmental trajectory based on the expression of SCMs. As expected, SCMs are highly expressed at the beginning of the trajectory, with their expression progressively declining as the cells transition toward differentiation. Indeed, SCMs such as WRKY DNA-BINDING PROTEIN 33 (WRKY33) and EARLY RESPONSE TO DEHYDRATION 6 (ERD6) exhibited a gradual decrease in expression along the pseudotime axis. In contrast, markers of mature tissues, such as PDF1 and DAG1, either showed the opposite trend, with a progressively increasing expression along the trajectory, or displayed lineage-specific patterns indicative of cell-type specialization (Fig. 3c and d; Supplementary Figs. 6, 7). Similarly to SCMs, the expression of HRGs, such as CALMODULIN-LIKE 38 (CML38) and SIMILAR TO RCD ONE 5 (SRO5), declined along the pseudotime trajectory (Fig. 3c and d; Supplementary Fig. 6). This suggests a coordinated downregulation of both stemness and hypoxia-related genes as cells transition toward differentiated states.
a, b Pseudotime trajectories of shoot (a) and root (b) apices visualized in UMAP space. Cells with low pseudotime values (purple) correspond to stem cell populations, while higher values (yellow) indicate progressive differentiation. Black lines represent the principal graph inferred by Monocle3, outlining the main developmental trajectories. c, d Expression patterns of SCMs, HRGs, and markers of differentiated tissues along the pseudotime axis. Gene modules for (e) shoot and (g) root apices plotted along pseudotime bins. f Comparative dot plot showing the GO enrichment of hypoxia-related processes found in early modules of the pseudotime trajectories of shoot and root apices. Dot size indicates the number of genes associated with each GO term.
To further explore the spatiotemporal dynamics of hypoxia responses during cell differentiation in plant meristems, we identified several gene modules, each defined by groups of genes with similar expression patterns along the trajectory. These modules were defined by clustering genes with similar expression patterns over pseudotime. We identified 37 gene modules in the shoot apex (Fig. 3e) and 27 gene modules in the root apex (Fig. 3g), each representing distinct gene activation patterns through specific cell differentiation phases. These modules were grouped into two differentiation phases: early-phase modules, enriched in genes highly active at the beginning of the trajectory, and late-phase modules, with genes activated later along the trajectory (Fig. 3e and g). Gene Ontology (GO) enrichment analysis of early-phase modules showed a highly significant enrichment for hypoxia-related GO terms for both the shoot and root apices (Fig. 3f; Supplementary Table 7). Contrarily, GO enrichment analysis of late-phase modules revealed the abundance of cellular and molecular processes associated with cell growth and differentiation, such as plastid organization, photosynthesis, and ribosome biogenesis (Supplementary Table 8). The enrichment of hypoxia-related GO terms in early-phase modules indicates the activation of hypoxia-associated gene expression in pluripotent undifferentiated cells, suggesting a pivotal role for hypoxia signaling in meristem function and cell fate determination.
Expression dynamics along the root differentiation trajectory reveal a link between stemness and response to hypoxia
To further explore the expression of SCMs and HRGs along the cell differentiation trajectory, we reconstructed a pseudotime bifurcated trajectory by integrating the root stem cell cluster with that of differentiated tissues, specifically the root cap and root cortex (Fig. 4a). The root differentiation trajectory inference showed that stem cells, which reside at the onset of the bifurcated trajectory (the stem cell cluster was manually assigned as the root of the trajectory), co-express SCMs and HRGs (Fig. 4b). To validate the computational attribution of root stem cells in vivo, we first identified the 100 most upregulated genes in the stem cells compared with the other clusters of the root differentiation trajectory (Supplementary Table 9). Among these genes, we identified GRETCHEN HAGEN3-type 3.3 (GH3.3; AT2G23170), a gene previously characterized as expressed in the lateral root meristem of Arabidopsis19. According to our computational gene expression trajectory, GH3.3 is predominantly expressed at the origin of the trajectory, specifically in the stem cells (Fig. 4b). Thus, GH3.3 was chosen as a candidate gene to verify whether the attribution of root stem cells in the root apical meristem could be experimentally confirmed by analyzing its spatial expression in planta. We generated an Arabidopsis stable transgenic reporter line using the GH3.3 promoter fused to three times repeated GFP coding sequence (pGH3.3::3xGFP). Spatial imaging of pGH3.3::3xGFP activity in the root apical meristem showed that, as predicted by the single-cell data, GH3.3 is expressed in the QC and stem cell initials of the columella, while its expression declines as cells progress into columella differentiation (Fig. 4c). The expression of CML38, an HRG, showed a spatial distribution in the bifurcated trajectory that largely overlapped with GH3.3 (Fig. 4b). In contrast, EXTENSIN 3 (EXT3), a marker of mature cortical tissues, and TYROSINE TRANSAMINASE FAMILY PROTEIN 4 (TAT4), a marker of the root cap, were expressed after the bifurcation point specifically in the branches belonging to the root cortex and cap, respectively (Fig. 4b).
a Diffusion map pseudotime showing bifurcation of stem cells into cortex and cap differentiation trajectories. b Expression of GH3.3, CML38, EXT3, and TAT4 across the diffusion map. c pGH3.3::3xGFP activity in the root apical meristem showing GH3.3 expression in QC and columella initials. Green and magenta represent fluorescence signal emitted by GFP and propidium iodide, respectively. Scale bar = 50 μm. d RNA velocity streamlines projected onto UMAP, illustrating gene expression dynamics based on splicing kinetics. The arrow direction indicates the future state of cells, while the arrow length represents the velocity of gene expression dynamics. e Latent time across root stem cells and cortex. Large circular arrows depict the self-renewal loop in stem cells and the differentiation path towards the root cortex. Arrows show the direction from stem cells to differentiated cell states. e Latent time across root stem cells and cortex. f Phase portraits and UMAP feature plots for GH3.3 and CML38 showing their splicing kinetics in root stem cells and cortex.
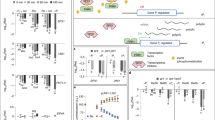
RNA velocity methods20 allow inference of the future state of individual cells by using the abundance of unspliced (pre-mRNA) to spliced RNA transcripts (mature RNA). RNA velocity values assume that the relationship between spliced mRNA expression and unspliced pre-mRNA expression can be used to predict whether a gene is in the process of being upregulated, downregulated, or in a steady expression state. The ratio of unspliced to spliced RNA for a given gene reflects its transcriptional dynamics within a cell population and can thus provide insights into cell fate decisions and differentiation20. Therefore, to investigate hypoxia-related transcriptional dynamics during stem cell differentiation, we applied RNA velocity analysis to root stem cells and the root cortex, which was selected as a representative mature tissue, using the same clustered dataset described in Fig. 1. RNA velocity analysis revealed directional streamlines indicating a clear trajectory from stem cells toward the root cortex, consistent with the differentiation axis (Fig. 4d). This inference was independently corroborated by latent time estimation (Fig. 4e), a model-derived continuum that orders cells based on their internal transcriptional dynamics, which provides a refined view of gene regulation during cell fate transitions. Indeed, stem cells showed the lowest latent time values, typically attributed to undifferentiated cell states21. Dynamic modeling of GH3.3 revealed a complete transcriptional cycle characterized by elevated levels of unspliced transcripts and high velocity in stem cells (Fig. 4f), indicating active transcriptional activity and a rapid change in gene expression state, respectively. As stem cells progress into the root cortex, the velocity and abundance of GH3.3 decrease (Fig. 4f), in line with the spatial activity of pGH3.3::3xGFP, which is primarily localized in the stem cell initials (Fig. 4c). Notably, CML38 displayed a canonical “almond‑shaped” dynamic profile in the phase plot, showing high velocity and abundance of transcripts at the interface of the transition from stem cells to root cortex. The expression profile of CML38 indicates that this hypoxia-responsive gene is dynamically regulated during stem cell differentiation. Since one hundred eighty genes belonging to the GO term “response to hypoxia” were identified after GO enrichment analysis within the early-phase modules (Fig. 3f; Supplementary Table 7), we decided to further investigate hypoxia-related transcriptional dynamics during stem cell differentiation by analyzing which of these 180 genes showed profiles similar to CML38. Among these genes, in addition to CML38, we identified 12 genes exhibiting a distinct almond-shaped trajectory in unspliced vs. spliced space (Fig. 5a; Supplementary Fig. 8), a hallmark of transcriptional cycling along latent time. The observed loop-shaped patterns suggest that these genes undergo highly dynamic transcriptional regulation, characterized by transient upregulation in undifferentiated cellular states, followed by transcriptional downregulation as cells progress toward differentiation (Supplementary Fig. 9). Such dynamics are characteristic of regulatory genes that may act as modulators of cell fate decisions. These findings suggested that a subset of hypoxia-responsive genes is spatiotemporally regulated during stem cell transitions. Within this group, WRKY22 regulates root development by controlling ethylene biosynthesis22. GLYCEROPHOSPHODIESTER PHOSPHODIESTERASE 2 (GDPD2) is strongly downregulated in the Arabidopsis double knockout mutant for NAP1-RELATED PROTEINS 1 (NRP1) and 2 (NRP2)23. This mutant exhibited cell cycle arrest in the RAM, suggesting that GDPD2, among other genes, may be involved in this process. Notably, genes such as DEHYDRATION-RESPONSIVE ELEMENT BINDING PROTEIN 2 (DREB2A), HEAT SHOCK TRANSCRIPTION FACTOR A2 (HSFA2), and GDPD2 displayed particularly well-defined transcriptional dynamics, with strong induction followed by repression over latent time (Fig. 5a), indicating their potential role as key gene drivers at the interface between stemness and differentiation.
a Latent time plots of 12 selected genes belonging to the Gene Ontology category “response to hypoxia”, found after GO enrichment analysis in early-phase modules. b Heatmap showing the expression levels (log₂ fold change) from ref. 24 of the 12 genes in (a) and CML38. c Average of expression patterns of the 12 genes in (a) along pseudotime (Fig. 2b). d Graph showing how expression of SCMs and HRGs declines as cells progress along the differentiation trajectory, where blue and yellow indicate stem cells and differentiated cell types, respectively. e Schematic representation of a bifurcated trajectory from (blue) root stem cells to (magenta and yellow) differentiated cells. Each branch of the trajectory represents a cell lineage. Oxygen gradients may play a crucial role in controlling stem cell state transitions.
Since these genes belonged to the GO term “response to hypoxia”, we evaluated whether they were actually responsive to hypoxia by mining their expression in an available bulk RNA-seq dataset of Arabidopsis seedlings exposed to hypoxia for 2 and 9 h24. All 13 genes were upregulated in at least one condition, both in nuclear and polyA-enriched RNA (Fig. 5b), confirming their transcriptional responsiveness to low oxygen. Interestingly, seven genes out of 13 either harbor in their locus the hypoxia responsive promoter element (HRPE) recognized by ERFVIIs25 or were found to be associated with HYPOXIA RESPONSIVE ERF 2 (HRE2), one of the five Arabidopsis ERFVIIs, in a ChIP-seq experiment24 (Supplementary Table 10). These genes included GDPD2, TANDEM ZINC FINGER 11 (TZF11), WRKY22, ACYL-COA-BINDING DOMAIN 3 (ACBP3), CYSTEINE-RICH TRANSMEMBRANE MODULE 6 (ATHCYSTM6) and AT3G04640. Importantly, all 13 genes showed a consistent decrease in expression as the cells progressed along the root differentiation trajectory (Fig. 5c). This decline in hypoxia responses along the differentiation trajectory supports a model in which hypoxia-responsive genes may be crucial for maintaining pluripotency or controlling cell state transitions (Fig. 5d and e).
Discussion
Multicellular organisms depend on oxygen to sustain growth and survival. In both plants and animals, oxygen is not only involved in cellular respiration but also serves as a signaling molecule that modulates development5,26. Hypoxic niches are a proven component of animal stem cell systems, with lower oxygen tension maintaining stemness and regulating the self-renewal/differentiation balance of the stem cell pool3. More recent research has now established that similar principles apply to plant meristems6,7. The SAM contains a hypoxic niche where the stabilization of oxygen-labile proteins is crucial for stem cell maintenance6. However, the extent to which hypoxia signaling contributes to stem cell identity in plant meristems, and how it is spatiotemporally modulated during cell differentiation, remains to be determined.
Our study addresses these questions by applying single-cell genomics approaches. By reanalyzing scRNA-seq data from shoot and root apices12,13, we found that hypoxia-associated transcriptional signatures overlapped with stemness. Stem cell markers and hypoxia-responsive genes were co-expressed in shoot and root apices, with a higher overlap in the root apex (Fig. 2). Although a hypoxic environment is established during LRP development7, it is still unclear whether a hypoxic niche exists within the RAM. Expression of HRGs within stem cells of the root apex suggests that a hypoxic niche may exist within the RAM, opening the possibility that root stem cells, similar to their shoot counterparts6, may reside in a low-oxygen microenvironment. Integrative single-cell transcriptomics across stem cell populations from various plant meristems (e.g., SAM, RAM, LRP, procambium, and axillary meristems) will shed light on the possible conservation and role of hypoxia signaling for plant meristem functioning. While our study is limited to Arabidopsis thaliana, cross-species scRNAseq integration analyses27,28 could reveal the extent to which the activation of hypoxia-related pathways is conserved in the plant meristems of different species. Indeed, a recent cross-species comparison using single-cell genomics between maize and Arabidopsis identified conserved stem cell regulators in the SAM29. Interestingly, the authors found, among others, stem cell regulators belonging to a family of sugar kinase genes related to hypoxia signaling in the SAM of maize29. Moreover, in the same study, a maize homolog of Arabidopsis ZPR2 was identified29, which might be involved in regulating the function of maize SAM as it is in Arabidopsis6. These results suggest that the role of oxygen-sensing pathways might be conserved in the shoot apical meristems of different plant species.
Reconstruction of developmental trajectories from stem cells to differentiated tissues in both shoot and root apical meristems revealed that SCMs and HRGs were highly expressed at early pseudotime, reducing gradually as differentiation continued (Fig. 3). This simultaneous decline of SCMs and HRGs along the cell differentiation trajectory suggests that hypoxia signaling may be actively involved in maintaining pluripotency and/or controlling cell state transitions. Gene module separation followed by GO enrichment analysis also supported this hypothesis, which showed that early modules were strongly enriched for biological processes linked to hypoxia. In contrast, late modules were characterized by growth- and differentiation-related processes, such as plastid organization, photosynthesis, and ribosome biogenesis. These findings suggest that hypoxia-associated transcriptional programs are a distinctive feature of undifferentiated cell states and are progressively repressed as cell differentiation progresses. Consistent with our results, a recent study on leaf morphogenesis demonstrated that developing leaves initiate in a hypoxic state and subsequently establish a distal-to-proximal oxygenation gradient, which influences cell fate specification and tissue differentiation through the action of ERFVIIs30. Thus, oxygen gradients might guide the developmental trajectories of individual cell lineages not only in plant meristems but also in undifferentiated proliferative tissues.
Co-expression analyses between HRGs and SCMs (Fig. 2a and b), together with HRG expression in stem cells (Supplementary Fig. 3), indicate that hypoxia responses are part of the transcriptional program of stem cells. Moreover, co-expression between HRGs and markers of differentiated tissues shows that different cell types and tissues might have different responsiveness to hypoxia (Supplementary Fig. 2).
An unresolved question remains whether hypoxia is necessary to determine stemness or is a consequence of the metabolic activity of stem cells and/or other factors that cause the activation of HRGs. While this work is unable to answer this question, future characterization of mutants with altered stem cell homeostasis, such as wuschel and clavata331, or mutants with impaired metabolic activity of stem cells, through the lens of single-cell transcriptomics, will advance our understanding of how hypoxia is generated and which role it plays in plant meristems.
RNA velocity analysis uncovered that a portion of genes belonging to the GO term “response to hypoxia” found in the early-phase modules of the root apex possess dynamic loops of expression in stem cells and early progenitor groups (Fig. 5). The transcriptional dynamics observed in these genes may be functionally relevant, contributing to the regulation of cell fate decisions in the RAM. Indeed, among these genes, WRKY22 and GDPD2 were previously associated with the function of the root apical meristem and root development22,23. Therefore, these hypoxia-responsive genes might be important regulators of cell fate and root development. Moreover, these results support the reliability of RNA velocity methods as tools for identifying key regulators of cell fate. Since these genes were upregulated by hypoxia (Fig. 5b), their expression could also be regulated by the endogenous hypoxia established within the RAM. However, the presence of a hypoxic RAM niche remains to be experimentally confirmed. Alternatively, these hypoxia-responsive genes may be crucial for the developmental reprogramming of plant meristems that occurs in response to environmental hypoxia following waterlogging or submergence.
Interestingly, among the 13 hypoxia-responsive genes showing high transcriptional dynamics (Fig. 5), seven genes harbor the HRPE within their promoters or gene sequences, which are recognized by ERFVII transcription factors (Supplementary Table 10). Therefore, the transcriptional activity of these genes might be regulated through ERFVII stabilization and action in stem cells during plant development or in response to hypoxia, while the others are regulated ERFVII-independent oxygen-sensing pathways.
Collectively, scRNA-seq represents a promising approach for exploring the function of hypoxia signaling at single-cell resolution and identifying key hypoxia-responsive regulators of plant development. The expression of several HRGs, such as ALCOHOL DEHYDROGENASE and PYRUVATE DECARBOXYLASE, was barely detected, possibly highlighting a decline in the expression of some HRGs following reoxygenation during protoplast preparation. Alternatively, the hypoxic cell niches in shoot and root apices may selectively express a subset of HRGs, as observed in hypoxic niches originating after pathogen infection32. Our work identifies hypoxia-responsive genes that might be important for the differentiation of stem cells in the RAM, and future functional studies of these genes will determine whether this is the case. Importantly, expanding single-cell transcriptomics to oxygen-sensing mutants, such as erfVII33, vrn28, zpr26 and prt634, will determine the extent to which oxygen-sensing mechanisms are actively involved in controlling cell fate decisions in plant meristems.
Methods
Cloning and plant transformation
GH3.3 promoter (2531 bp upstream of the start codon) was amplified from genomic DNA using Phusion® High-Fidelity DNA Polymerase (New England Biolabs). Restriction sites required for GreenGate cloning were added to the primers used to amplify GH3.3 promoter (pGH3.3_forward: aacaggtctcaacctggaggaaataactattt; pGH3.3_reverse: aacaggtctcatgttatcaacggtcatgatta), which was subsequently cloned into pCR2.1 vector (TA Cloning™ Kit, Thermo Fisher Scientific). GreenGate Cloning System method35 was used to assemble the construct pGH3.3::3xGFP::tUBQ10 together with the BastaR cassette (pGGF002) into the destination vector (pGGZ001) for plant transformation. The destination vector carrying both pGH3.3::3xGFP::tUBQ10 and p35S::BastaR::t35S was transformed into Agrobacterium tumefaciens (GV3101). Arabidopsis thaliana Columbia-0 (Col-0) was transformed with Agrobacterium following the floral dipping method36. Selection of transgenic plants was carried out on medium containing 25 μg/ml of ammonium glufosinate. Homozygous plants were obtained before performing spatial imaging of pGH3.3::3xGFP.
Plant growth conditions
To assess the spatial expression of GH3.3, pGH3.3::3xGFP seeds were surface-sterilized using sequential incubations in 70% (v/v) ethanol, 5% (v/v) hypochlorite solution, and then washed repeatedly with sterile water. Seeds were stratified at 4 °C in the dark for 48 h and then germinated on vertical plates (Greiner Bio-One) containing Murashige and Skoog half-strength medium, 0.5% w/v sucrose, 0.9% w/v phytoagar, pH 5.7. pGH3.3::3xGFP seedlings were grown under constant a temperature of 22 °C, with a photoperiod of 16 h/8 h light/dark cycles and light intensity of 100 μmol m−2 s−1 PPFD (Photosynthetic Photon Flux Density). 5-day-old pGH3.3::3xGFP seedlings were used to analyze the spatial expression of GH3.3 by monitoring GFP signal in the RAM.
Confocal imaging
pGH3.3::3xGFP roots were stained in 10 μg/mL propidium iodide (Sigma) before observation to visualize the cell walls of living cells. Fluorescence imaging was performed using the confocal microscope Stellaris 8 (Leica Microsystems, Germany) equipped with a pulsed white light laser tunable from 440 to 800 nm and operating at 80 MHz repetition rate. Fluorescence signals from GFP-tagged proteins and propidium iodide were acquired in sequential mode, with excitation of the apical root at 488 nm and 543 nm, respectively. Emission signals were collected using 40× HC PL APO CS2 oil immersion objective (N.A. = 1.30) (Leica microsystems) in the 495–545 nm range for GFP and in the 589-781 nm range for propidium iodide. A 2048 × 2048- pixel image was generated with no digital zoom, and the signal-to-noise ratio (SNR) was improved by averaging two frames.
Pre-processing of raw scRNA-seq data
Single-cell RNA sequencing data analyzed in this study were derived from ref. 13 for the vegetative shoot apex and ref. 12 for the root apex. Raw sequencing data were downloaded using the “prefetch” command within sratools toolkit (https://trace.ncbi.nlm.nih.gov/Traces/sra/sra.cgi?view=software). Data for the SAM were downloaded from the Beijing Institute of Genomics Data Center (BioProject PRJCA003094) and for the RAM from the NCBI SRA database (BioProject PRJNA517021). FASTQ files were generated using the “fastq-dump” function. The FASTQ files were processed with Cell Ranger version 3.0.1 (10x Genomics). The genome sequence and annotation (GTF) files for Arabidopsis thaliana were downloaded from the TAIR10 database (https://www.arabidopsis.org/). A custom reference genome was built using the “cellranger mkref” function with the “--genome”, “--fasta”, and “--genes” arguments. Gene expression matrices were generated using the “cellranger count” command with the arguments “--id”, “--transcriptome”, “--fastqs”, and “--sample”. These matrices served as raw input for downstream analyses. Each step was independently performed for the shoot apex and root tip datasets.
Data integration, clustering, and annotation
Further analyses were conducted using the Seurat package v.5.1.037. Data processing was performed sequentially, starting with quality control to remove low-quality cells and genes. Specifically, cells expressing fewer than 200 genes were removed, and genes expressed in fewer than three cells were excluded. Data normalization was carried out using the NormalizeData function with the LogNormalize method and a scaling factor of 10,000. Highly variable genes were identified using the FindVariableGenes function with the vst method, selecting the top 2000 features and then scaled with the ScaleData function.
Principal component analysis (PCA) was performed using the RunPCA function on the identified variable genes, retaining the first 50 principal components. A shared nearest neighbor (SNN) graph was constructed, and cells were clustered using Louvain community detection through the FindNeighbors and FindClusters functions, by testing different resolution values (ranging from 0.4 to 1) and selecting 0.4 as the setting that provided the most biologically consistent clustering for both SAM and RAM. Visualization of clustering results was performed using the non-linear dimensional reduction method RunUMAP. Unlike the original analyses by refs. 12,13, we reprocessed the raw FASTQ files from both datasets using a unified pipeline, ensuring identical quality control thresholds, normalization, and clustering parameters. This analysis resulted in 19 clusters for the shoot apex and 20 for the root apex, rather than the 23 and 24 clusters originally reported by refs. 12,13, and enabled direct cross-tissue comparisons. To ensure consistency in cluster annotation, we relied on a dual approach by visualizing both the marker genes reported by Zhang et al.12,13 and the cluster-enriched genes identified through the Seurat function FindAllMarkers. The resulting marker sets for each cluster were subsequently analyzed using the Cell Type Predictor tool available in the scPlantDB platform, enabling an automated comparison with known cell type expression profiles. The complete list of genes is provided in Supplementary Tables 1–3. Clusters were then merged based on tissue-specific annotations.
To identify marker genes for each cell type, we leveraged the scPlantDB database38. For each annotated cell type, we ranked marker genes based on the percentage difference (PctDiff) metric, which quantifies the difference in the proportion of cells expressing a given gene in the target cell type relative to all other cell types. Only markers with a PctDiff ≥ 0.65 were considered, ensuring high specificity and minimal cross-lineage expression. From this filtered set, the top ten marker genes for each cell type were selected and used for co-expression analyses with HRGs (Fig. 2a, b; Supplementary Fig. 2; Supplementary Table 4). HRGs (Supplementary Table 6) were obtained from Mustroph et al. (2010)15. To quantify the overlap between SCMs, markers of differentiated tissues and HRGs, we calculated the proportion of cells expressing each HRG within the population of cells expressing a given SCM. Specifically, for each SCM–HRG pair, we determined the number of cells co-expressing both genes (expression ≥10 counts per gene) and divided this value by the total number of cells expressing the corresponding SCM (Fig. 2a, b). To calculate the percentage of stem cells expressing HRGs in the shoot and root apices, we selected the top 30 SCMs (Supplementary Table 5) to predict stem cells as cells expressing at least 10 of these 30 markers. Subsequently, % of stem cells expressing a given HRG in the whole stem cell population was calculated and represented as heatmaps (Supplementary Fig. 3).
Differential trajectory and cell lineage analysis
Trajectory and lineage differentiation analyses were performed using Monocle39,40. For pseudotime inference, we used the entire single-cell transcriptomic datasets of the shoot and root apices described in Fig. 1, without applying additional subsetting. The Seurat-integrated objects were converted into Monocle3 cell_data_set objects using the as.cell_data_set function. Within Monocle, we performed dimensionality reduction de novo, first by principal component analysis (PCA), followed by UMAP using the preprocess_cds and reduce_dimension functions. Cells were then reclustered in Monocle using the cluster_cells function to ensure compatibility with the learn_graph trajectory construction algorithm. Importantly, the biological composition of the datasets remained unchanged, as all cell populations identified in the Seurat analysis were retained. Individual pseudotime expression profiles for SCMs and HRGs were visualized using the plot_genes_in_pseudotime function. To analyze transcriptional dynamics along pseudotime, cells were ordered according to the trajectory inferred by Monocle3 and then divided into ten equal intervals, referred as “bins”, using the cut() function in R. Each bin contains the cells whose pseudotime values fall within the corresponding interval, thereby preserving their relative position along the developmental path. Average expression levels were calculated for gene modules previously identified with the find_gene_modules function. Gene expression modules along pseudotime were defined using the aggregate_gene_expression function. Early and late pseudotime gene modules were enriched in Gene Ontology “Biological Function” via enrichGO. Monocle2 was then utilized to delineate cellular differentiation trajectories within subclusters. For both shoot apex and root tip datasets, stem cell clusters and two clusters representing mature tissues were selected for analysis. Using DDRTree dimensionality reduction and the orderCells function, pseudotime-based differentiation trajectories were generated, assigning the stem cell clusters as the initial “State”. Highly expressed genes within the stem cell clusters for both tissues were compared with bulk RNA-seq datasets previously published24. Figures were generated in R.
RNA velocity analysis
RNA velocity analysis was performed to model the transcriptional dynamics and infer the future states of individual cells. Spliced and unspliced transcript counts were generated using the velocyto pipeline41. BAM alignment files produced by Cell Ranger were processed with velocyto.py using the TAIR10 Arabidopsis genome annotation to generate loom files containing spliced, unspliced, and ambiguous counts for each cell.
Downstream analyses were conducted using the scVelo Python package21. Loom files were imported into scVelo and integrated with metadata and dimensionality reduction coordinates (UMAP) obtained from the single-cell RNA-seq analysis. Cells were filtered and normalized following scVelo’s recommended preprocessing pipeline. The first- and second-order moments were computed based on the neighborhood graph constructed from the UMAP embedding. RNA velocities were estimated using both the stochastic and dynamical models implemented in scVelo.
The velocity graph and velocity streamlines were projected onto the UMAP embedding to visualize the directionality of gene expression changes. Latent time, representing the internal transcriptional progression of each cell, was inferred using the dynamical model. Gene-specific transcriptional kinetics, including splicing rates and transcriptional induction/repression dynamics, were modeled and visualized using scVelo plotting functions.
Data availability
Single-cell RNA sequencing data analyzed in this study were derived from ref. 13 for the vegetative shoot apex and ref. 12 for the root apex. Raw sequencing data for the SAM were downloaded from the Beijing Institute of Genomics Data Center (BioProject PRJCA003094; https://ngdc.cncb.ac.cn/search/specific?db=bioproject&q=PRJCA003094) and for the RAM from the NCBI SRA database (BioProject PRJNA517021; https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA517021).
Code availability
All scripts and workflows used in this study are publicly available on GitHub (https://github.com/SimoneFromThePlantLab/scRNAseq-hypoxia-SAM-RAM) and have been archived on Zenodo (https://doi.org/10.5281/zenodo.16743581) for long-term accessibility.
References
Loreti, E. & Perata, P. The many facets of hypoxia in plants. Plants 9, 745 (2020).
Yoshida, Y., Takahashi, K., Okita, K., Ichisaka, T. & Yamanaka, S. Hypoxia enhances the generation of induced pluripotent stem cells. Cell Stem Cell 5, 237–241 (2009).
Greb, T. & Lohmann, J. U. Plant Stem Cells Curr. Biol. 26, R816–R821 (2016).
Soyars, C. L., James, S. R. & Nimchuk, Z. L. Ready, aim, shoot: stem cell regulation of the shoot apical meristem. Curr. Opin. Plant Biol. 29, 163–168 (2016).
Weits, D. A., Dongen, J. T. & Licausi, F. Molecular oxygen as a signaling component in plant development. N. Phytol. 229, 24–35 (2021).
Weits, D. A. et al. An apical hypoxic niche sets the pace of shoot meristem activity. Nature 569, 714–717 (2019).
Shukla, V. et al. Endogenous hypoxia in lateral root primordia controls root architecture by antagonizing auxin signaling in Arabidopsis. Mol. Plant 12, 538–551 (2019).
Gibbs, D. J. et al. Oxygen-dependent proteolysis regulates the stability of angiosperm polycomb repressive complex 2 subunit VERNALIZATION 2. Nat. Commun. 9, 5438 (2018).
Osborne, R. et al. VRN2-PRC2 facilitates light-triggered repression of PIF signaling to coordinate growth in Arabidopsis. Dev. Cell S1534580725001224 https://doi.org/10.1016/j.devcel.2025.03.001(2025).
Zhu, M., Taylor, I. W. & Benfey, P. N. Single-cell genomics revolutionizes plant development studies across scales. Development 149, dev200179 (2022).
Cervantes-Pérez, S. A. et al. Cell-specific pathways recruited for symbiotic nodulation in the Medicago truncatula legume. Mol. Plant 15, 1868–1888 (2022).
Zhang, T.-Q., Xu, Z.-G., Shang, G.-D. & Wang, J.-W. A single-cell RNA sequencing profiles the developmental landscape of Arabidopsis root. Mol. Plant 12, 648–660 (2019).
Zhang, T.-Q., Chen, Y. & Wang, J.-W. A single-cell analysis of the Arabidopsis vegetative shoot apex. Dev. Cell 56, 1056–1074.e8 (2021).
Lee, J.-Y. et al. Transcriptional and posttranscriptional regulation of transcription factor expression in Arabidopsis roots. Proc. Natl. Acad. Sci. USA 103, 6055–6060 (2006).
Mustroph, A. et al. Cross-kingdom comparison of transcriptomic adjustments to low-oxygen stress highlights conserved and plant-specific responses. Plant Physiol. 152, 1484–1500 (2010).
Armstrong, W. Microelectrode and modelling study of oxygen distribution in roots. Ann. Bot. 74, 287–299 (1994).
Thomson, C. J. & Greenway, H. Metabolic evidence for stelar anoxia in maize roots exposed to low O2 concentrations. Plant Physiol. 96, 1294–1301 (1991).
Van Dongen, J. T., Schurr, U., Pfister, M. & Geigenberger, P. Phloem metabolism and function have to cope with low internal oxygen. Plant Physiol. 131, 1529–1543 (2003).
Wang, Q. et al. GH3-mediated auxin inactivation attenuates multiple stages of lateral root development. N. Phytol. 240, 1900–1912 (2023).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
Wang, Z. et al. The transcription factor WRKY22 modulates ethylene biosynthesis and root development through transactivating the transcription of ACS5 and ACO5 in Arabidopsis. Physiol. Plant. 176, e14371 (2024).
Zhu, Y. et al. Arabidopsis NRP1 and NRP2 Encode Histone Chaperones and Are Required for Maintaining Postembryonic Root Growth. Plant Cell 18, 2879–2892 (2006).
Lee, T. A. & Bailey-Serres, J. Integrative analysis from the epigenome to translatome uncovers patterns of dominant nuclear regulation during transient stress. Plant Cell tpc.00463.2019 https://doi.org/10.1105/tpc.19.00463 (2019).
Gasch, P. et al. Redundant ERF-VII transcription factors bind to an evolutionarily conserved cis-Motif to regulate hypoxia-responsive gene expression in Arabidopsis. Plant Cell 28, 160–180 (2016).
Ratcliffe, P. J. Oxygen sensing and hypoxia signalling pathways in animals: the implications of physiology for cancer. J. Physiol. 591, 2027–2042 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Conde, D. et al. Single-nuclei transcriptome analysis of the shoot apex vascular system differentiation in Populus. Development 149, dev200632 (2022).
Xu, X. et al. Large-scale single-cell profiling of stem cells identifies redundant regulators of shoot development and yield trait variation. Dev. Cell S1534580725005003 https://doi.org/10.1016/j.devcel.2025.07.024 (2025).
Panicucci, G. et al. Progressive oxygenation of developing leaves directs morphogenesis. Preprint at https://doi.org/10.1101/2025.09.21.677629 (2025).
Yadav, R. K. et al. WUSCHEL protein movement mediates stem cell homeostasis in the Arabidopsis shoot apex. Genes Dev. 25, 2025–2030 (2011).
Brunello, L. et al. The transcription factor ORA59 represses hypoxia responses during Botrytis cinerea infection and reoxygenation. Plant Physiol. 197, kiae677 (2024).
Abbas, M. et al. Oxygen sensing coordinates photomorphogenesis to facilitate seedling survival. Curr. Biol. 25, 1483–1488 (2015).
Zubrycka, A. et al. ERFVII action and modulation through oxygen-sensing in Arabidopsis thaliana. Nat. Commun. 14, 4665 (2023).
Lampropoulos, A. et al. GreenGate—a novel, versatile, and efficient cloning system for plant transgenesis. PLoS ONE 8, e83043 (2013).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana: Floral dip transformation of Arabidopsis. Plant J. 16, 735–743 (1998).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2024).
He, Z. et al. scPlantDB: a comprehensive database for exploring cell types and markers of plant cell atlases. Nucleic Acids Res. 52, D1629–D1638 (2024).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Cao, J. et al. Comprehensive single-cell transcriptional profiling of a multicellular organism. Science 357, 661–667 (2017).
Klingler, D., Schleicher, J. T. & Claassen, M. Quantification of spliced and unspliced transcripts by velocyto is inaccurate for 5’-sequencing data. Preprint at https://doi.org/10.1101/2025.01.17.633503 (2025).
Acknowledgements
This work was supported by Scuola Superiore Sant'Anna and by MUR-PRIN2022 (PRIN 2022-2022YHWH9R; Next Generation EU) to P.P. and E.L. The authors acknowledge the support from Project ECS00000017 ‘Ecosistema dell’Innovazione’ Tuscany Health Ecosystem (THE, PNRR, Spoke 4) from the European Union-Next Generation EU.
Author information
Authors and Affiliations
Contributions
S.C., P.P. and P.M.T. conceived and designed the research. S.C., F.F., M.M., and P.M.T. conducted the analyses and experiments. S.C., E.L., P.P. and P.M.T. interpreted the results. S.C., E.L., P.P. and P.M.T. wrote the manuscript. All the authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Castellana, S., Fioriti, F., Marazzini, M. et al. Inferring hypoxia-responsive regulators of cell fate in plant meristems through single-cell transcriptomics. npj Sci. Plants 2, 2 (2026). https://doi.org/10.1038/s44383-025-00015-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44383-025-00015-3