Abstract
Aspartic proteinases (APs) are essential proteolytic enzymes involved in plant growth, development, and defence. This study investigates PSI-containing Aspartic Proteinases (APs) in Solanum lycopersicum (tomato plant), focusing on their molecular characteristics, tissue-specific expression, and response to abiotic stress. While 58 APs were identified in S. lycopersicum, only five (AP V, AP W, AP X, AP Y, and AP Z) contained a C-terminal vacuolar sorting domain and the Plant Specific Insert (PSI) domain, a unique domain implicated in vacuolar trafficking and stress responses. These PSI-containing APs were further analyzed through phylogenetics, revealing their evolutionary relationships with homologous APs from other species. Expression profiling showed that most PSI-containing APs are highly expressed in cotyledons and roots, with AP Y being distinctively predominant in reproductive tissues. Under osmotic and saline stress, expression patterns varied, with AP Z exhibiting a broader stress sensitivity. Biochemical analyses confirmed that severe osmotic stress led to increased oxidative stress markers and antioxidant responses, suggestive of activation of the plant-adaptive mechanisms to abiotic stress. As a whole, it is shown that PSI-containing APs from Solanum lycopersicum are strictly regulated within the plant and exhibit varied responses to environmental stressors.
Similar content being viewed by others
Introduction
Aspartic proteases (APs) (EC 3.4.23) are a diverse and evolutionarily conserved group of proteolytic enzymes in plants, playing critical roles in various physiological and developmental processes such as seed germination, defense mechanisms, programmed cell death (PCD), and hormone regulation. Structurally, APs belong predominantly to the pepsin-like A1 family, characterized by the conserved Asp-Gly-Thr motif at their catalytic sites. The catalytic mechanism involves two Asp residues that promote the hydrolyzation of peptide bonds in an acidic pH environment1,2,3. Many plant APs, such as cardosins from Cynara cardunculus, also contain unique structural features, including plant-specific inserts (PSIs) and C-terminal vacuolar sorting determinants (ctVSDs), which enable the precise targeting of APs to various cellular locations, such as vacuoles and lytic compartments1,2,3. Moreover, APs exhibit significant evolutionary conservation across species, evidenced by their presence in model plants like Arabidopsis thaliana, cereal crops such as Oryza sativa (rice) and Triticum aestivum (wheat), and specially plants like C. cardunculus and Solanum lycopersicum (tomato). In Arabidopsis, over 50 AP genes have been identified, many of which are categorized into typical (PSI-containing) and atypical groups based on structural features4,5. Rice harbors 96 AP genes, while Vitis vinifera (grapevine) and Populus trichocarpa (poplar) possess 50 and 67 AP genes, respectively, reflecting significant genomic expansion6,7. Despite this diversity, the catalytic core of APs is highly conserved, suggesting a strong evolutionary pressure to maintain their enzymatic function across plant lineages7.
APs are expressed in a variety of plant tissues, with different isoforms exhibiting tissue-specific functions. For example, during seed germination, APs help break down storage proteins in the endosperm to provide nutrients for the growing embryo5,8. They are also involved in leaf senescence and PCD, where they degrade cellular components as part of plant aging and development9. In reproductive tissues, APs are thought to play a role in pollen-stigma interactions, embryo development, and post-fertilization events, being involved in cellular remodeling and PCD8. Moreover, the bifunctional role of PSI-containing APs, such as membrane destabilization and protein degradation, suggests they play a crucial role in seed germination and development10.
In addition to their involvement in growth and development, APs play an essential role in plant defense. Many APs are upregulated during pathogen infection, contributing to the hypersensitive response and systemic acquired resistance (SAR)11. For example, the constitutive disease resistance 1 from Arabidopsis is involved in generating peptide elicitors that activate SAR and enhance plant immunity12. Similarly, AP in guard cells 1 (ASPG1) regulates drought tolerance by influencing seed dormancy and antioxidant defenses13. Atypical APs, such as nucellin-like APs, exhibit unique structural features that diverge from conventional PSI-containing enzymes and have emerged as key players in developmental processes and stress responses7. Interestingly, pathogens have evolved strategies to overcome plant defenses by secreting their own APs, which can degrade host proteins to facilitate infection11.
PSIs are unique protein domains found exclusively in plant APs. These approximately 100-amino-acid-long domains function as VSDs and contribute to plant defense14. Positioned between the heavy and light chains of AP proenzymes, PSIs play crucial roles in protein maturation and intracellular trafficking. For example, during the maturation process of cardosins, precursor proteins are directed to the vacuole via two distinct VSDs: the ctVSD, a conventional VSD, and the PSI, classified as an unconventional VSD, as it does not fit into established VSD classifications, such as ssVSDs, ctVSDs, or psVSDs2,3. Despite this classification, PSIs mediate both conventional and unconventional trafficking depending on their glycosylation status and structural context2,14. Despite being excised during maturation, PSIs retain functional properties, including antimicrobial activity, suggesting that their role extends beyond vacuolar trafficking. This multifunctionality highlights their significant contributions to plant defense mechanisms11,14. Evolutionary conservation of PSIs spans monocots, dicots, and green algae, underscoring their fundamental importance in plant biology14. Chlamydomonas reinhardtii, a green alga, contains a PSI-like domain in chlapsin, which may represent an ancestral form of PSIs15. While PSIs in monocots like Hordeum vulgare (barley) primarily mediate vacuolar trafficking, those in dicots like Solanum tuberosum (potato) and Cirsium vulgare prioritize antimicrobial defense, reflecting species-specific adaptations15,16. In dicots, the roles of PSIs extend to abiotic stress responses, including potential contributions to drought and salinity tolerance17. Non-glycosylated PSIs, such as those in cardosin A, are hypothesized to mediate rapid Golgi-bypass trafficking, delivering stress-responsive proteins to vacuoles under challenging environmental conditions3,18. While these hypotheses remain to be fully validated, they highlight the potential involvement of PSIs in hormone signaling and stress adaptation. The ability of PSIs to balance trafficking and defense roles underscores their multifunctionality and evolutionary adaptability.
The dwarf tomato cultivar Micro-Tom (S. lycopersicum cv. Micro-Tom) serves as an excellent model system for studying plant stress responses due to its compact size, short life cycle, and well-characterized genetics19. These features, combined with its genetic tractability and the availability of mutant collections, allow for efficient analysis of gene function, endomembrane dynamics, and stress adaptation mechanisms. This work identified AP sequences in S. lycopersicum, highlighting those that are homologous to cardosin A and B, i.e., PSI-containing APs. It then examined the organ-specific expression of these PSI-containing APs in tomato along with stress responses, particularly under drought and salinity conditions.
Methods
Protein and genomic sequences
Protein sequences of Cynara cardunculus cardosin A (AJ132884) and cardosin B (AJ237674), Glycine max soyAP1 (NM_001250416) and soyAP2 (NM_001250135), Solanum tuberosum StAP (AY672651), Cirsium vulgare cirsin (JN703462), Hordeum vulgare phytepsin (P42210), and Chlamydomonas reinhardtii chlapsin (AJ579366) were retrieved from the National Center for Biotechnology Information database (https://www.ncbi.nlm.nih.gov/). Genomic sequence from Arabidopsis thaliana AtAP A1 (AT1G11910.1) was retrieved from The Arabidopsis Information Resource (TAIR; www.arabidopsis.org). Using the Sol Genomics Network (https://solgenomics.net), 58 Solanum lycopersicum protein and genomic sequences corresponding to APs were retrieved through a targeted search for the “aspartic protease” keyword.
Phylogenetic and sequence analysis
A comparative analysis was then conducted with InterPro software (https://www.ebi.ac.uk/interpro/) to identify the presence of saposin-like domains within the retrieved 58 S. lycopersicum protein sequences. Five sequences containing saposin-like domains were identified and designated as AP V (Solyc03g005280.2.1), AP W (Solyc07g051850.2.1), AP X (Solyc09g082760.2.1), AP Y (Solyc02g032940.2.1), and AP Z (Solyc01g101240.2.1). Protein sequences of cardosin A, cardosin B, soyAP1, soyAP2, StAP, cirsin, phytepsin, and 58 S. lycopersicum APs were aligned using the T-Coffee multiple sequence alignment (MSA) tool20 (https://tcoffee.crg.eu). PSI-containing APs alignment was exported to Jalview software21 for analysis. Phylogenetic trees were constructed using the Molecular Evolutionary Genetics Analysis X software22, employing the Maximum Likelihood method with the Jones-Taylor-Thornton (JTT) substitution model23. Evolutionary relationships among the analyzed taxa were visualized through bootstrap consensus trees, derived from 1000 replicates for statistical robustness. APs’ domains were depicted using InterPro24—classification of protein families tool (https://www.ebi.ac.uk/interpro/).
Plant growth
Solanum lycopersicum cv. Micro-Tom seeds were surface-sterilized in 70% (v/v) ethanol for 1 min to remove superficial contaminants, followed by a 15-min immersion in 20% (v/v) sodium hypochlorite solution supplemented with 0.001% (v/v) Tween 20 (Sigma-Aldrich, USA) as a surfactant. After the bleach treatment, the seeds were rinsed with sterile deionized water, completing a total of five washes. The sterilized seeds were then air-dried on sterile filter paper and germinated in vitro on Murashige and Skoog (MS) medium (Duchefa Biochemie, Netherlands), supplemented with 1.5% (w/v) sucrose and 0.7% (w/v) Bactoagar (Duchefa Biochemie, Netherlands). Depending on the purpose, two different experiments were performed with tomato plants: (i) in order to obtain an insight into the expression profile of the tomato APs across different organs and tissues, after 14 days of germination, the plants were transferred to pots with growth substrate [SiroPlant and allowed to grow till fruitification]; (ii) to study the possible relationship between APs and abiotic stress conditions, an in vitro experiment was conducted. Here, specific concentrations of stress-inducing agents were added to the medium: 50 mM sodium chloride (NaCl - S1) to induce saline stress, and 50 mM and 100 mM mannitol (H1 and H2) to simulate osmotic (drought-like) stress. Seeds were germinated (as before), and seedlings were maintained in vitro for 14 and 25 days, when samples were collected for RT-qPCR and biochemical assays.
Micro-Tom tomato (Solanum lycopersicum var. Micro-Tom) and wild-type Nicotiana tabacum plants were grown under controlled conditions using substrates from the Siro® brand (Siro Group, Albergaria-a-Velha, Portugal). Micro-Tom plants were cultivated in Siro® Plant substrate composed of humus of pine bark, sphagnum peat, and wood fiber (Siro® Fibra), supplemented with a slow-release mineral fertilizer (NPK 16-8-14, 2.0 g L−1) lasting up to 9 months. The substrate has a pH of 5.5 ± 0.5, electrical conductivity of 75 µS cm−1 (±30%), bulk density of 400 g L−1 (±20%), and organic matter content > 70%. Plants were watered every 2 days with 50 mL of distilled water and maintained at 23 °C under a 16-h light photoperiod with average humidity levels between 50% and 60% and a light intensity of 180 µmol m−2 s−1. For the in vitro stress experiment, seedlings were collected at 14 and 25 days after germination, corresponding to plants with approximately 3–4 true leaves (~2 cm height) and 7–8 true leaves (~6 cm height), respectively. Three biological replicates were prepared for each treatment condition and experiment.
Nicotiana tabacum plants were grown in Plant Universal substrate (SIRO®, Portugal) under continuous light at 24 °C with 50–60% relative humidity and the same light intensity of 180 µmol m−2 s−1.
Gene expression analysis
Total RNA was extracted from 200 mg of various tissues of S. lycopersicum cv. Micro-Tom was grown under different growth conditions composing two sets of samples. The first set included: cotyledons from 14-day-old seedlings, roots, stem, leaf, flower, green tomato, and ripe tomato from 8-week-old plants grown under normal conditions. The second set included 14-day-old and 25-day-old seedlings and shoots from plants grown under stress conditions. RNA was extracted using the NZY Total RNA Isolation Kit (NZYTech, Portugal). Three biological replicates were prepared for each condition, and RNA concentration and purity were measured using a DS-11 Spectrophotometer/Fluorometer (DeNovix, USA). For cDNA synthesis, 100 ng of RNA samples of the first set and 1 µg of RNA from samples of the second set were reverse transcribed using the NZY First-Strand cDNA Synthesis Kit (NZYTech, Portugal).
For qPCR analysis, primers for S. lycopersicum AP V, AP W, AP X, AP Y, and AP Z (Supplementary Table 1) were designed using Primer3Plus (https://www.primer3plus.com/index.html) with default qPCR settings. The primer pairs were tested and validated by generating a standard curve from a tenfold dilution series (1:10, 1:100, and 1:1000) of cDNA. Three biological replicates and three technical replicates were performed for each experimental condition. Reactions were performed using NZYSupreme qPCR Green Master Mix (NZYTech, Portugal), following manufacturer’s instructions, using 1:10 diluted cDNA template, and caried on a CFX96 Real-Time System (Bio-Rad, USA). A melt curve analysis was performed at the end of the PCR from 65 to 95 °C, with fluorescence measurements taken every 0.5 °C to confirm the specificity of the amplicons. The cycle threshold (Ct) values, efficiency, and expression levels were analyzed using Bio-Rad CFX Maestro software (version 1.0; Bio-Rad, USA). The expression levels of the Sl APs were normalized using two reference genes: Ubiquitin 3 (Genbank: X58253) and Elongation factor 1α (Genbank: BT013246), validated in the study of Lacerda and co-workers25. The normalized relative gene expression was calculated using the 2−ΔΔCt method26.
Quantification of H2O2
Hydrogen peroxide (H2O2) levels were quantified according to Alexieva and co-workers27 with modifications for adaptation to a microplate assay. Frozen tissue aliquots were homogenized in 1 mL of 0.1% (w/v) trichloroacetic acid (TCA; Sigma-Aldrich, USA) using a bead mill homogenizer. To prevent sample overheating, homogenization was performed in three cycles of 5 m/s for 20 s, with a 1-min interval on ice between each cycle. The homogenates were centrifuged at 12,000 × g for 15 min at 4 °C, and the resulting supernatant was collected for subsequent analysis. For the assay, 50 μL of the supernatant was transferred to each well of a microplate, followed by the addition of 50 μL of 100 mM potassium phosphate buffer, pH 7.0, and 200 μL of 1 M potassium iodide. The reaction mixture was incubated at room temperature (RT) in the darkness for 1 h, and the absorbance was measured at 390 nm using a microplate reader. H2O2 levels were calculated from a standard curve, prepared with stock solutions of H2O2, and expressed as nanomoles of H2O2 per gram of fresh weight (nmol g−1 F.W.).
Determination of lipid peroxidation levels
Lipid peroxidation was evaluated by quantifying malondialdehyde (MDA) content through the thiobarbituric acid reactive substances assay, as described by Heath and Packer28. Aliquots of 100 mg tissue were homogenized in 1 mL of 0.1% (w/v) TCA on ice. The homogenate was centrifuged at 12,000 × g for 15 min at 4 °C to remove debris. The resulting supernatant (500 µL) was mixed with 1.5 mL of 0.5% (w/v) thiobarbituric acid dissolved in 20% (w/v) TCA. The reaction mixture was incubated at 95 °C for 30 min to allow color development and subsequently cooled rapidly on ice to terminate the reaction. After cooling, the samples were centrifuged again at 10,000 × g for 10 min to clarify the solution. The absorbance of the supernatant was recorded at 532 nm using a spectrophotometer, with nonspecific absorbance corrections measured at 600 nm. The MDA concentration was calculated using an extinction coefficient of 155 mM−1 cm−1. Results were expressed as nanomoles of MDA per gram of fresh weight.
Determination of proline levels
Proline (Pro) levels were quantified according to Bates and co-workers29. Frozen aerial tissue samples were homogenized in 3% (w/v) sulfosalicylic acid using a bead mill homogenizer. The homogenates were centrifuged at 16,000 × g for 10 min at 4 °C, and the resulting supernatant was collected. A 200 μL aliquot of each sample was mixed with 200 μL of glacial acetic acid and 200 μL of acid ninhydrin reagent. The reaction mixture was incubated at 95 °C for 1 h and then cooled on ice. To extract the reaction product, 1 mL of toluene was added to the cooled mixture, which was vortexed for 15 s to separate the organic (red upper phase) and aqueous (whitish lower phase) layers. The red organic phase was collected, and absorbance was measured at 520 nm. Proline concentration was determined using a standard curve generated with known concentrations of L-proline. Results were expressed as milligrams of proline per gram of fresh weight (mg g−1 F.W.).
Quantification of reduced glutathione
Glutathione (GSH) levels were quantified using a protocol adapted from Soares and co-workers30. Frozen tissue samples (100 mg) were homogenized in 1 mL of 3% (w/v) sulfosalicylic acid using a bead mill homogenizer. The homogenates were centrifuged at 16,000 × g for 10 min at 4 °C, and the resulting supernatant was collected. A 50 μL aliquot of the supernatant was mixed with 200 μL of deionized water and 750 μL of a reaction mixture containing 100 mM potassium phosphate buffer (pH 7.0), 10 mM ethylenediaminetetraacetic acid (EDTA), and 1.5 mg mL−1 5,5′-dithiobis-(2-nitrobenzoic acid) (DTNB). The mixture was vortexed briefly and incubated in the dark for 10 min to allow for the reaction. The absorbance was measured at 412 nm, and GSH levels were quantified based on a standard curve prepared with known concentrations of reduced glutathione. Results were expressed as micromoles of GSH per gram of fresh weight (μmol g−1 F.W.).
Statistical analysis
All data were statistically analyzed using GraphPad Prism 10 software (www.graphpad.com). For biochemical analysis, stress-induced differences were compared to the control using a one-way analysis of variance (ANOVA) followed by Dunnett’s multiple comparison test. For qPCR data, gene expression levels were analyzed using a one-way ANOVA, followed by Tukey’s multiple comparison test, or a two-way ANOVA, followed by Dunnett’s multiple comparison test. Statistically significant differences were identified with an α threshold of 0.05 and a 95% confidence interval, with specific p values (*p < 0.0332; **p < 0.0021; ***p < 0.0002; ****p < 0.0001).
Constructs design and assembly for the transient transformation of Nicotiana tabacum
Solanum lycopersicum PSI W, PSI X, and PSI Z sequences were amplified from cDNA of 14-day-old seedlings grown under normal conditions. Sequences were amplified using Phusion High-Fidelity PCR Master Mix (Thermo Fisher Scientific, USA), according to the manufacturer’s instructions, and with primers listed in Supplementary Table 2. All PCR products were purified using GeneJET PCR Purification Kit (Thermo Fisher Scientific, USA) in line with the manufacturer’s procedure. Purified PCR products were cloned into pMiniT 2.0 using NEB® PCR Cloning Kit (NEB, USA), according to the manufacturer’s instructions. Then, inserts were transferred into a modified version of pMDC83 destination vector31, containing a SP from N. tabacum chitinase A and a mCherry sequence, using Gibson assembly32, generating pMDC83::SP-PSI W-mCherry, pMDC83::SP-PSI X-mCherry, and pMDC83::SP-PSI Z-mCherry. All constructs were verified by sequencing.
Agrobacterium-mediated transient transformation of Nicotiana tabacum
Agrobacterium tumefaciens GV3101 was used for the transient transformation of N. tabacum leaves, following the method described by Sparkes and co-workers33. A 5 mL liquid culture of LB medium, supplemented with 50 µg/mL kanamycin, 30 μg/mL gentamicin, and 10 μg/mL rifampicin, was prepared for A. tumefaciens containing the specified constructs and incubated overnight at 28 °C with shaking at 200 rpm. One mL of culture was centrifuged at 3500 × g for 3 min at RT, and the pellet was resuspended in 2 mL of infiltration medium (10 mM MgCl2, 10 mM MES-KOH pH 5.6, and 150 μM acetosyringone). The suspension was then centrifuged and resuspended again in infiltration buffer. The suspension was diluted to an OD600 of 0.3 and infiltrated into the abaxial side of 3- to 4-week-old leaves. Co-expression with the dominant-negative mutant SarIH74L-GFP was performed using an OD600 of 0.1 in combination with SP-PSI W-mCherry, SP-PSI X-mCherry, or SP-mCherry-Z, all at an OD600 of 0.3.
Nicotiana tabacum leaves were visualized 72 h post-infiltration using a Leica TCS SP5 II confocal microscope system (Leica Microsystems, Germany) with a HC Plan Apochromat (PL APO) confocal scanning (CS) 40x/1.10 NA correction ring (CORR) water-immersion objective (Leica Microsystems, Germany). To detect GFP and RFP fluorescence, laser lines of 488 nm and 561 nm were used, respectively. Emission was collected at 500–530 nm for GFP and 590–620 nm for mCherry, using a sequential acquisition mode, optimizing signal specificity and minimizing spectral overlap. Images were analyzed using ImageJ software (https://imagej.net/ij/).
Results
Identification of APs in Solanum lycopersicum
Since no PSI-containing APs have been identified in S. lycopersicum so far, a comprehensive analysis was performed using the Solgenomics database to identify APs in this species. A search for APs in S. lycopersicum revealed 58 candidate proteins (Table 1), which were subsequently examined for characteristic features of AP homologs, including saposin-like domains, ctVSDs, and glycosylation motifs. InterPro was utilized to identify saposin-like domains, which were regarded as PSIs in this study. Among the 58 identified APs in S. lycopersicum, only five contained PSIs, which were designated as AP V (Solyc03g005280.2.1), AP W (Solyc07g051850.2.1), AP X (Solyc09g082760.2.1), AP Y (Solyc02g032940.2.1), and AP Z (Solyc01g101240.2.1).
Notably, all PSI-containing APs also possessed a ctVSD, similar to those described for cardosins. The ctVSD sequence VGFAEAA, found in cardosin A2, was identified in AP W, AP Y, and AP Z, as well as in AtAP A1 and cirsin. The ctVSD sequences from AP V (VGFAESA) and AP X (IGFAESA) exhibited some degree of similarity to the ctVSD of cardosin A, with some differences. Specifically, AP V and AP X contain a Ser (S) residue in place of the Ala (A) residue, while AP X exhibits an Ile (I) residue instead of the Val (V) residue. Cardosin B presents a VGFAEAV ctVSD sequence2, which is also present in soyAP2. Additionally, some ctVSD sequences from phytepsin (IGFAKAA), StAP (VGFAEVHNSL), soyAP1 (VGFADAA), and chlapsin (LGFANAA) showed unique variations; however, the GFA motif is consistently shared among these sequences, a feature also common to the other PSI-containing APs (Table 1).
Earlier studies have demonstrated that PSI B is glycosylated and contains the NETE glycosylation motif3. Based on this knowledge, the analysis of S. lycopersicum APs focused on identifying glycosylation motifs such as NET, NGT, and NQT within the identified PSIs. This approach enabled the identification of glycosylated PSIs. In detail, AP V contained the NGT motif, AP W contained the NETA motif, AP X contained the NET motif, and AP Y contained the NQT motif. Similarly, soyAP1 and cirsin contained the NETE motif, AtAP A1 and phytepsin contained the NMTQ motif, and chlapsin contained the NSTI motif. In contrast, PSI Z lacks a glycosylation motif, as do soyAP2 and StAP. Based on their glycosylation motifs, PSI V, PSI W, PSI X, and PSI Y were classified as PSI B-like due to their similarity to the glycosylation profile of PSI B from C. cardunculus, while PSI Z is classified as PSI A-like, as it does not have the glycosylation motif.
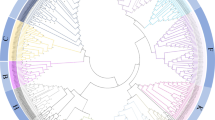
APs from Table 1 were further aligned in an MSA, conducted with T-Coffee, a precise alignment tool20, and a Maximum Likelihood phylogenetic tree was created with MEGA-X22 to infer evolutionary relationships (Fig. 1). PSI-containing APs from S. lycopersicum are closely related to each other and show strong similarity to PSI-containing APs from other species. Specifically, AP Y is most closely related to soyAP1, while also showing similarity to AtAP A1 and H. vulgare phytepsin. AP W is the closest S. lycopersicum AP to cardosin A and B, as well as C. vulgare cirsin. AP Z is closely associated with StAP. In contrast, AP V and AP X form a paralogous pair that clusters near soyAP2. Notably, C. reinhardtii chlapsin is phylogenetically distant from other PSI-containing APs, reflecting its evolutionary divergence as a green alga. The phylogenetic tree reveals a distinct clustering of these PSI-containing APs, suggesting a conserved evolutionary lineage among them.
Phylogenetic tree of 58 S. lycopersicum APs and known APs such as C. cardunculus cardosin A and cardosin B (purple), A. thaliana AtAP A1 (orange), H. vulgare phytepsin (pink), S. tuberosum StAP (brown), G. max soyAP1 and soyAP2 (blue), C. vulgare cirsin (red), and C. reinhardtii chlapsin (green). PSI-containing APs in S. lycopersicum (AP V, AP W, AP X, AP Y, and AP Z) are highlighted in bold. The phylogenetic tree was obtained with MEGA-X22 using the Maximum Likelihood method and the JTT matrix-based model23. The percentage of replicate trees in which the associated taxa clustered together in the bootstrap test (1000 replicates) is shown next to the branches. This analysis involved 67 amino acid sequences, with a total of 1485 positions in the final dataset.
To have a deeper insight into this cluster formed by APs containing the PSI domain, a new alignment using only these sequences was obtained (Supplementary Fig. 1A). The results show patches with a high degree of conservation (Fig. 1A, yellow rectangles) contrasting with regions with low amino acidic conservation (Fig. 1A, brown rectangles). Regarding the PSI domain, it is clear that amino acidic residues at the N- and C-terminal ends are highly conserved between the different APs. Domain analysis focusing on S. lycopersicum APs harboring the PSI domain (Supplementary Fig. 1B) clearly identified these sequences as belonging to the beta-site APP-cleaving enzyme, with the peptidase A1 domain and a saposin-like domain in the PSI region. Furthermore, the two catalytic sites were identified in all APs (ACT-site), and for some APs, like AP W and AP Z, a conserved disulfide site was identified in the terminal region of the protein (Supplementary Fig. 1B).
PSI-containing APs in S. lycopersicum are tissue-specific
To investigate the tissue-specific expression patterns of the identified PSI-containing APs in S. lycopersicum, plants were grown for 8 weeks under standard growth conditions for the Micro-Tom variety until the fruiting stage. RNA was extracted from seven different tissues, such as cotyledons from 14-day-old seedlings, root, stem, leaf, flower, green tomato, and ripe tomato from 8-week-old plants grown under normal conditions. Expression levels were analyzed using qPCR, and data were normalized using the ΔΔCq method. The results revealed that AP V exhibited its highest expression in cotyledons, with levels significantly higher compared to all other tissues, except for leaves (Fig. 2A). AP W also showed the highest expression in cotyledons, followed by roots. Its expression was notably increased in cotyledons compared to all tissues except roots, while levels in roots were significantly higher than in leaves, flowers, and ripe tomatoes (Fig. 2B).
Expression levels of AP V (A), AP W (B), AP X (C), AP Y (D), and AP Z (E) were normalized using the ΔΔCq method. Tissues analyzed included cotyledons, root, stem, green leaf, flower, green tomato, and ripe tomato. Error bars represent the standard error of the mean (SEM) from three biological replicates. Statistical analyses were conducted using a one-way ANOVA followed by Tukey’s multiple comparison test using the GraphPad Prism 10 software (http://graphpad.com). Asterisks represent statistically significant differences compared to the tissue with higher expression, unless otherwise specified, with an α threshold of 0.05 and a 95% confidence interval (*p < 0.03; **p < 0.002; ***p < 0.0002; ****p < 0.0001).
Similarly, AP X was most highly expressed in cotyledons, with elevated expression levels compared to all other tissues analyzed. Additionally, AP X showed increased expression in roots compared to stems and green tomatoes (Fig. 2C). AP Y exhibited the highest expression levels in green tomato and flower tissues, though no statistical differences were detected between tissues (Fig. 2D). AP Z showed high expression in cotyledons, followed by roots, with expression levels in both cotyledon and roots significantly higher than in ripe tomato (Fig. 2E). Overall, the qPCR results reveal distinct tissue-specific expression profiles for the five PSI-containing APs. Cotyledons serve as a primary site of expression for four (AP V, AP W, AP X, and AP Z) of the five genes (Fig. 2). AP Y was the only identified PSI-containing AP to present higher expression levels in reproductive tissues instead of cotyledons. Additionally, unique patterns, such as the high expression of AP Y in green tomato and flower tissues and the root-specific expression of AP W and AP Z (Fig. 2), suggest diverse functional roles for these APs.
PSI-containing APs respond differently to stress
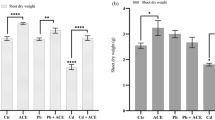
To determine the role of PSI-containing APs in stress responses in S. lycopersicum, their expression levels were evaluated under various abiotic stress conditions. Plants were subjected to osmotic stress with 50 mM mannitol (H1) and 100 mM mannitol (H2), and saline stress with 50 mM NaCl (S1) to simulate drought and salinity conditions, respectively. Visible differences in 25-day-old plant growth were observed across the different stress treatments, in relation to unstressed plants (Fig. 3A). Plants grown under control conditions (C) exhibited significantly higher growth compared to plants exposed to osmotic (H1 and H2) and saline stress (S1; Fig. 3B). Plants under both osmotic (H1 and H2) and saline (S1) stress showed stunted growth (Fig. 3B), indicating the detrimental effects of these stress conditions on overall plant development. With the intention of confirming the oxidative stress status on this 25-day-old S. lycopersicum plants, H2O2, lipid peroxidation (MDA), Proline (Pro), and GSH levels were measured. Under severe osmotic stress (H2), H2O2 (~0.8 nmol g−1 F.W.), MDA (~20 nmol g−1 F.W.), Pro (~500 nmol g−1 F.W.), and GSH (~200 nmol g−1 F.W.) significantly increased compared to control, while in mild osmotic stress (H1) and saline stress (S1), these markers remained similar to control (Supplementary Fig. 2).
A Seedlings were grown under control conditions (C), osmotic stress with 50 mM mannitol (H1) and 100 mM mannitol (H2), and saline stress with 50 mM sodium chloride (S1), for 25 days post-germination. B Representative images highlight the reduced growth of 25-day-old plants under osmotic stress (H1 and H2) and saline (S1) conditions compared to the control (C).
Next, to investigate the molecular responses of PSI-containing APs (AP V, AP W, AP X, AP Y, and AP Z) from S. lycopersicum, qPCR analysis was performed at two different time points. First, the expression levels of each target AP were evaluated in seedlings grown under control and stress conditions for 14 days (Fig. 4A). qPCR results revealed distinct expression patterns for the five genes. AP V expression was downregulated under saline stress (S1). AP W expression was significantly reduced under osmotic stress (H2). AP X showed a notable decrease in expression under saline stress (S1), while AP Z exhibited a significant decrease in expression under both osmotic (H2) and saline (S1) stress conditions. AP Y did not display any significant reduction in expression across the stress treatments (Fig. 4A). Second, the same expression analysis was performed with 25-day-old plants grown under control and stress conditions (Fig. 4B). AP V was upregulated under both osmotic stress conditions (H1 and H2). AP W, AP X, AP Y, and AP Z showed no notable decrease in expression under the stress treatments (Fig. 4B). These findings indicate that PSI-containing APs exhibit distinct and time-dependent transcriptional responses to salt and osmotic stress, which may reflect differential regulation of these genes at specific developmental stages in S. lycopersicum. Specifically, AP V appears to play a dynamic role, being downregulated in seedlings under saline stress but upregulated in older plants under osmotic stress. In contrast, AP W, AP X, AP Y, and AP Z demonstrate more stable expression patterns, with AP Z showing a broader sensitivity to stress across different conditions and time points.
Normalized expression levels (ΔΔCq) of AP V, AP W, AP X, AP Y, and AP Z) of 14-day-old seedlings (A—orange box) and 25-day-old plants (B—blue box) grown under control conditions, osmotic stress with 50 mM mannitol (H1) and 100 mM mannitol (H2), and saline stress with 50 mM sodium chloride (S1). Error bars represent the standard error of the mean (SEM) from three biological replicates. Statistical analyses were conducted using a two-way ANOVA followed by Dunnett’s multiple comparison test using the GraphPad Prism 10 software (http://graphpad.com). Asterisks represent statistically significant differences compared to the control conditions (C) with an α threshold of 0.05 and a 95% confidence interval (*p < 0.03; **p < 0.002; ***p < 0.0002; ****p < 0.0001).
Solanum lycopersicum PSIs can mediate trafficking to the vacuole
To investigate the subcellular localization of S. lycopersicum PSIs and to compare them with the subcellular localizations of PSI A and PSI B from C. cardunculus, the protein sequences of PSI W, PSI X, and PSI Z were cloned into a destination vector containing a signal peptide (SP) for ER targeting and a mCherry fluorescent protein. These PSIs were chosen due to their glycosylation pattern, mimicking either PSI B (PSI W and X) or PSI A (PSI Z). All three PSIs localized to the vacuole (Fig. 5A), with some fluorescence also being detected at the cell periphery and surrounding the nucleus. To have a more detailed view of the PSI’s trafficking route, a co-expression assay with the dominant negative form of SarIH74L-GFP, blocking the ER-to-Golgi pathway, was performed. All PSIs became blocked in the ER after co-expression with the SarI mutant, despite some fluorescence still being detected in the vacuole for PSI W and PSI X (Fig. 5B). PSI Z fluorescence is entirely detected in the ER.
Transient expression of SP-PSI W-mCherry, SP-PSI X-mCherry, and SP-PSI Z-mCherry (A) in N. tabacum leaves was recorded 72 h post-infiltration in the control situation. Co-expression of the different PSIs with the dominant negative mutant SarIH74L-GFP (B) in N. tabacum leaves was recorded 72 h post-infiltration.
Discussion
APs are vital enzymes in plants, contributing to protein degradation, vacuolar trafficking, stress responses, and reproduction. These enzymes are classified into various subfamilies based on their structural features and functional roles, with some possessing unique domains like PSIs and ctVSDs. The identification, in S. lycopersicum, of a subset of APs containing PSIs and ctVSDs provided a first insight into the functional and structural complexity of this enzyme family. This study analyzed the structure, expression, and potential roles of these APs, comparing them to the well-characterized cardosins A and B from C. cardunculus.
From the 58 identified APs in the S. lycopersicum genome, five PSI-containing APs were found: AP V (PSI V), AP W (PSI W), AP X (PSI X), AP Y (PSI Y), and AP Z (PSI Z). Notably, all five PSI-containing APs also possessed VGFAEA-like ctVSDs, similar to cardosin A and cardosin B, as well as A. thaliana AtAP A1, G. max soyAP2, and C. vulgare cirsin, while none of the APs lacking PSIs exhibits such ctVSDs. This strong association between PSIs and ctVSDs suggests a coordinated role in vacuolar targeting. The PSI likely facilitates membrane interactions, vesicle formation, and vacuolar sorting under specific conditions, while the ctVSD serves as the primary signal for vacuolar sorting under normal circumstances. According to Pereira et al.2, both the PSI and the ctVSD are independently sufficient for vacuolar targeting. However, when both signals are present, and in vegetative tissues, the ctVSD acts as the dominant sorting signal, dictating the protein trafficking pathway to the vacuole. Despite this dominance, the PSI offers an alternative Golgi-independent pathway, bypassing conventional ER-to-Golgi trafficking and potentially providing redundancy under specific conditions2. This interplay between dominant and alternative vacuolar sorting signals underscores the complexity of trafficking mechanisms in PSI-containing APs, which may exploit distinct pathways depending on developmental or stress-related cues.
The phylogenetic analysis of S. lycopersicum APs and known APs from other species highlighted the evolutionary relationships among PSI-containing APs. The close clustering of S. lycopersicum PSI-containing APs with homologous enzymes from C. cardunculus (cardosin A and cardosin B), A. thaliana (AtAP A1), H. vulgare (phytepsin), S. tuberosum (StAP), G. max (soyAP1 and soyAP2), and C. vulgare (cirsin) underscored their evolutionary conservation across diverse plant species and fundamental roles in critical cellular processes such as protein degradation and vacuolar trafficking. Structural similarities among these APs suggested that the PSI domain is crucial for their proper function. Distinct clustering patterns further supported functional similarities, such as AP W grouping with cardosins and cirsin, reinforcing their role in vacuolar trafficking. Cirsin shows a high sequence similarity to cardosin B, with 95% sequence identity, also possessing a glycosylated PSI. Studies have confirmed that both glycosylated and non-glycosylated PSI variants of cirsin exhibit antifungal activity against phytopathogenic fungi. Additionally, recombinant procirsin also exhibits milk-clotting activity, suggesting potential use as a vegetable rennet. The PSI domain appears to play a critical role in the proper folding and function of cirsin, as its deletion led to an inactive protein34,35. Likewise, the proximity of AP to soyAP1, AtAP A1, and phytepsin suggested that these APs may serve similar roles in their respective species, potentially reflecting an adaptation to specific physiological needs. SoyAP1 is highly expressed in seeds, particularly in dry seeds and early germination stages, where it plays a key role in storage protein mobilization36. A. thaliana AtAP A1 has been successfully expressed and characterized as a heterodimeric glycosylated enzyme. AtAP A1 is localized in the MVBs, where it undergoes activation as the luminal pH gradually decreases, supporting its involvement in vacuolar trafficking. However, the precise role of its PSI in vacuolar targeting and enzymatic regulation remains to be fully elucidated37. Phytepsin is involved in protein processing, metabolic turnover, and PCD. It ultimately localizes to vacuoles in roots and grains, where it functions in protein degradation. The PSI domain of phytepsin has been suggested to play a role in vacuolar targeting, although its exact function remains unclear. Glycosylation at the PSI domain was found to slow down proteolytic maturation, suggesting a protective role against premature degradation. Additionally, phytepsin can undergo autoactivation at acidic pH, further confirming its function in the vacuolar environment38. The pairing of AP V and AP X as paralogs, clustering near soyAP2, indicates a potential functional divergence within this group of APs, due to gene duplication, enabling specialization under different cellular or stress conditions. SoyAP2 is expressed more broadly across various tissues, including roots, stems, leaves, and flowers, but is not found in dry seeds, suggesting it plays a more general role in protein degradation and cellular homeostasis36. AP Z paired with StAP, which also lacks a glycosylation motif on the PSI domain14. A key feature of StAP is its membrane-interacting ability, which is largely attributed to its PSI. The PSI domain facilitates membrane interactions, bilayer fusion, and antimicrobial activity39. StAP has been shown to directly interact with fungal pathogens such as Fusarium solani and Phytophthora infestans, leading to membrane permeabilization and cytotoxic effects40. Furthermore, the transgenic expression of StAP-PSI in A. thaliana significantly enhanced resistance against Botrytis cinerea, suggesting a potential application in plant biotechnology and pathogen resistance strategies41. StAP is upregulated in potato tubers and leaves following wounding or pathogen infection, reinforcing their involvement in defense mechanisms42. The phylogenetic distance of C. reinhardtii chlapsin from other PSI-containing APs highlighted the evolutionary divergence of green algae from land plants. Unlike most plant APs, which are typically vacuolar or secreted, chlapsin is localized in the chloroplast. It shares structural characteristics with typical plant APs, such as catalytic DTG motifs and optimal enzymatic activity at acidic pH, but also possesses distinct features, including an unusually long PSI with an additional 80 amino acids rich in alanine residues. Chlapsin PSI domain contains six conserved cysteine residues and a putative glycosylation site, but it remains unclear if it has any role in vacuolar trafficking15. Despite this, the presence of a PSI domain in chlapsin suggested that the essential role of PSIs in protein sorting and vacuolar targeting may be an ancient feature conserved across plant lineages.
In line with the descriptions for APs from other species, the distinct expression patterns of PSI-containing APs in S. lycopersicum suggested diverse functional roles. AP V, AP W, AP X, and AP Z showed high expression in cotyledons, with AP W and AP Z also being elevated in roots, indicating roles in nutrient storage and seedling development, similar to APs in other species8. The broad expression of AP Z across tissues indicated a constitutive role in maintaining cellular homeostasis. In contrast, AP Y stood out as the only PSI-containing AP predominantly expressed in reproductive tissues, with high levels in green tomatoes and flowers. While no statistical differences were detected among tissues, this pattern implied a role in reproductive processes, such as flower or fruit development, but further insights are needed to clarify this hypothesis. Among the PSI-containing APs analyzed, AP Y showed markedly high expression in reproductive organs, particularly flowers and green fruits. The presence of a putative N-glycosylation motif (NQT) within its PSI suggests that AP Y may undergo post-translational modification, influencing its stability and trafficking. Similar PSI-containing APs from Cynara cardunculus8 and Arabidopsis thaliana43 have been linked to protein degradation and cellular remodeling events during flower and fruit development. Therefore, the elevated expression of AP Y in reproductive tissues may reflect a conserved role in proteolytic remodeling associated with organ differentiation and maturation, potentially mediated by PSI-dependent targeting to the endomembrane system. Similarly, cardosin B localizes to the extracellular matrix (ECM) of the style transmitting tissue, where its glycosylated PSI enhances trafficking efficiency and stability, enabling ECM remodeling and pollen tube guidance during fertilization8. In contrast, APs with non-glycosylated PSIs may function similarly to cardosin A, which primarily localizes to PSVs in stigmatic papillae, facilitating nutrient storage and protein turnover during reproductive development2,8. Although the subcellular localization of AP Y was not experimentally assessed, its potential intracellular function can be indirectly supported by conservation analysis of the PSI domain and by comparison with homologous proteins from other plant species. Phylogenetically, AP Y clusters with well-characterized PSI-containing APs such as Arabidopsis thaliana AtAP A1, Hordeum vulgare phytepsin, and Glycine max soyAP1, which have been reported to localize to endomembrane compartments and/or the vacuole3,37,38. Moreover, AP Y retains a conserved PSI domain together with a canonical ctVSD, strongly suggesting that its functional context is linked to vacuolar trafficking and regulated proteolysis, in line with mechanisms described for other typical plant APs.
Complementary stress studies in S. lycopersicum revealed distinct responses of PSI-containing APs under osmotic and saline stress conditions, with notable differences in expression patterns between 14-day-old seedlings and 25-day-old plants. In 14-day-old seedlings, AP V, AP X, and AP Z showed significant downregulation under saline stress, suggesting a specific involvement in salt-induced responses, potentially allowing other proteins to take their place in compensatory roles. Similarly, AP W and AP Z exhibited marked downregulation under severe osmotic stress, indicating their involvement in processes sensitive to osmotic balance and water availability. Notably, the transcriptional modulation of PSI-containing APs under severe osmotic stress coincided with a pronounced increase in oxidative stress markers, including H₂O₂ accumulation, lipid peroxidation, and enhanced antioxidant responses. This association suggests that PSI-containing AP expression is coordinated with the cellular redox status, supporting a functional link between stress-induced transcriptional regulation of APs and oxidative stress signaling. Such coordination is consistent with a role for PSI-containing APs in proteolytic remodeling during oxidative stress conditions.
Interestingly, AP V displayed contrasting transcriptional responses under salt and osmotic stress at different developmental stages, being downregulated in 14-day-old seedlings under saline stress but upregulated in 25-day-old plants under osmotic stress. This divergence likely reflects developmental stage–dependent activation of distinct stress signaling pathways: saline stress in young seedlings predominantly triggers ionic toxicityand MAPK-mediated signaling44, whereas osmotic stress in mature plants is more strongly associated with ABA-dependent pathways45. The upregulation of AP V in older plants may therefore reflect activation of ABA-regulated proteolytic processes contributing to vacuolar remodeling and cellular homeostasis during prolonged water deficit. Such divergence likely reflects age-dependent modulation of stress signaling networks. Young seedlings primarily rely on basal antioxidant defenses and osmolyte accumulation for rapid protection, whereas older plants exhibit more specialized, organ-level adjustments involving vacuolar proteolysis and remodeling of endomembrane trafficking46,47,48. The enhanced AP V expression in 25-day-old plants may therefore indicate activation of proteolytic pathways supporting cellular homeostasis during prolonged stress. In contrast, AP Y maintained stable expression across both stress conditions in both 14-day-old seedlings and 25-day-old plants, suggesting its critical role in reproductive and/or metabolic processes that are not significantly impacted by osmotic or saline stress. This stability is consistent with its high expression in reproductive tissues, potentially explaining why AP Y expression remained unaffected. AP Z showed significant downregulation under both osmotic and saline stress conditions, suggesting its broader sensitivity to environmental stressors compared to other APs. Notably, the downregulation of most PSI-containing APs under stress conditions points to their responsiveness to abiotic stress, with other genes likely being activated to enhance plant survival. Insights from A. thaliana further highlight the functional roles of APs under stress conditions. For example, ASPG1, an Arabidopsis AP, has been shown to regulate stomatal aperture and enhance drought tolerance, illustrating the adaptive versatility of APs in abiotic stress responses13. Other Arabidopsis APs, such as At1g11910 and At1g62290, exhibit differential expression under hydric and saline stress, with the former consistently upregulated and the latter displaying responsiveness only under extreme stress conditions17. These findings suggest that APs have specialized roles in proteostasis and stress adaptation. In wheat, APs exhibit transcriptional upregulation during biotic stress, such as powdery mildew infection, indicating their involvement in immune responses and pathogen defense49. Similarly, rice APs contribute to PCD, pollen viability, and reproductive tissue development under abiotic stress11. Studies on atypical APs, such as nucellin-like proteases in Arabidopsis and Oryza sativa, reveal additional regulatory roles in development and stress responses. These enzymes lack conventional catalytic sites and the PSI domain but retain regulatory functions in PCD and vacuolar degradation7. The functional parallels with PSI-containing APs in tomato suggest that these enzymes could play similar roles in balancing protein turnover and developmental processes under stress and normal conditions.
This study not only contributed valuable insights into gene expression and stress responses but also highlighted the conserved role of PSIs in mediating vacuolar trafficking. The subcellular localization of SP-PSI W-mCherry, SP-PSI X-mCherry, and SP-PSI Z-mCherry in N. tabacum provided valuable preliminary insights into the trafficking behavior of S. lycopersicum PSIs. All three PSIs showed strong localization to the vacuole and at the cell periphery, probably corresponding to intermediate trafficking steps, like the ER and Golgi. These observations resemble C. cardunculus PSI A and PSI B and G. max PSI1 and PSI2 localizations in previous studies2,3, highlighting the conserved role of PSIs in mediating vacuolar trafficking. Vieira and co-workers3 showed that different PSIs can mediate vacuolar trafficking through different routes, depending on the glycosylation state of the PSIs: through the Golgi (glycosylated - conventional route) and bypassing the Golgi (non-glycosylated – unconventional route). Here, the same ER-to-Golgi blockage was tested using the dominant negative form of SarI (SarIH74L). Surprisingly, both PSI W, X, and Z were blocked in the ER after co-expression with SarIH74L. This indicates that glycosylation per se is not the only feature to determine the PSI-mediated route to the vacuole, as PSI Z, not having the glycosylation motif, was expected to be able to bypass the blockage and still accumulate in the vacuole, as observed for PSI A3. This observation reopens the discussion on the mechanisms behind PSI-mediated trafficking, as no receptors or interacting partners have been disclosed so far. A recent study from Sampaio and colleagues50 discuss the possibility of the PSI dimerization and its ability to interact with membranes, being involved in the process. They show that PSI A and B are able to dimerize and identify an internal loop region with potential for direct membrane interaction, which becomes exposed upon dimerization. Further research in this area would be fundamental to understanding the mechanisms behind PSIs’ function as vacuolar determinants.
At present, the molecular determinants that govern vacuolar trafficking of PSI-containing APs remain unresolved. Available evidence indicates that PSIs are dynamic modulators of vacuolar sorting rather than simple binary “signals.” PSIs can form homo- and heterodimers, and dimer formation increases membrane-binding affinity, a phenomenon demonstrated in vitro (via liposome binding/fusion assays) and in vivo (via mbSUS/FRET-FLIM)50,51. When glycosylation motifs were swapped between PSI A (glycosylated) and PSI B (non-glycosylated), Vieira and co-workers3 observed differences in trafficking behavior—glycosylated forms showed greater sensitivity to pathway perturbation, yet most PSIs still followed the conventional Golgi-dependent route. Taken together, these data indicate that glycosylation is likely one contributing recognition cue, but it is not the sole regulator of PSI trafficking—and neither is dimerization. Instead, PSI trafficking appears combinatorial and context-dependent, influenced by oligomeric state, transient membrane interactions, and the physiological or pharmacological state of the secretory pathway and/or other unknown factors. Clarifying this multilayered regulation will require an integrated approach combining mutational analysis (glycosylation site removal/addition; helix/disordered region swaps), in vivo and in vitro dimerization assays, membrane-binding (liposome) studies, and live-cell FRET/FLIM to monitor PSI oligomerization under trafficking perturbations.
This research highlights the diversity and functional significance of PSI-containing APs in S. lycopersicum, particularly their roles in protein degradation, vacuolar targeting, and stress adaptation. The findings suggest that these APs are tightly regulated at the tissue level and respond differentially to environmental stressors. The observed correlation between PSI glycosylation and potential trafficking pathways adds another layer of complexity to AP function. Future studies should explore the precise molecular mechanisms underlying PSI-mediated trafficking and stress response, which could have implications for improving stress resilience in crop species. Future verification of AP Y subcellular localization should be conducted using experimental systems that preserve the physiological context of reproductive organs, such as protoplasts derived from floral tissues or tissue-specific expression approaches.
Data availability
All data generated or analyzed during this study are included in this manuscript and its supplementary material.
References
Egas, C. et al. The saposin-like domain of the plant aspartic proteinase precursor is a potent inducer of vesicle leakage*. J. Biol. Chem. 275, 38190–38196 (2000).
Pereira, C., Pereira, S., Satiat-Jeunemaitre, B. & Pissarra, J. Cardosin A contains two vacuolar sorting signals using different vacuolar routes in tobacco epidermal cells. Plant J. 76, 87–100 (2013).
Vieira, V. et al. N-linked glycosylation modulates golgi-independent vacuolar sorting mediated by the plant specific insert. Plants 8, 312 (2019).
Faro, C. & Gal, S. Aspartic proteinase content of the Arabidopsis genome. Curr. Protein Pept. Sci. 6, 493–500 (2005).
Darabi, M. & Seddigh, S. Bioinformatic characterization of aspartic protease (AP) enzyme in seed plants. Plant Syst. Evol. 301, 2399–2417 (2015).
Guo, R. et al. Genome-wide identification, evolutionary and expression analysis of the aspartic protease gene superfamily in grape. BMC Genom. 14, 554 (2013).
Soares, A., Ribeiro Carlton, S. M. & Simões, I. Atypical and nucellin-like aspartic proteases: emerging players in plant developmental processes and stress responses. J. Exp. Bot. 70, 2059–2076 (2019).
Pissarra, J. et al. From flower to seed germination in Cynara cardunculus: a role for aspartic proteinases. https://repositorio-aberto.up.pt/handle/10216/82523 (2007).
Beers, E. P., Woffenden, B. J. & Zhao, C. Plant proteolytic enzymes: possible roles during programmed cell death. Plant Mol. Biol. 44, 399–415 (2000).
Pereira, C. S. et al. Cardosins in postembryonic development of cardoon: towards an elucidation of the biological function of plant aspartic proteinases. Protoplasma 232, 203–213 (2008).
Figueiredo, L., Santos, R. B. & Figueiredo, A. Defense and offense strategies: the role of aspartic proteases in plant–pathogen interactions. Biology 10, 75 (2021).
Simões, I., Faro, R., Bur, D. & Faro, C. Characterization of recombinant CDR1, an Arabidopsis aspartic proteinase involved in disease resistance. J. Biol. Chem. 282, 31358–31365 (2007).
Yao, X., Xiong, W., Ye, T. & Wu, Y. Overexpression of the aspartic protease ASPG1 gene confers drought avoidance in Arabidopsis. J. Exp. Bot. 63, 2579–2593 (2012).
Cheung, L. K. Y., Dupuis, J. H., Dee, D. R., Bryksa, B. C. & Yada, R. Y. Roles of plant-specific inserts in plant defense. Trends Plant Sci. 25, 682–694 (2020).
Almeida, C. M. et al. Chlapsin, a chloroplastidial aspartic proteinase from the green algae Chlamydomonas reinhardtii. Planta 236, 283–296 (2012).
Muñoz, F. F. et al. The swaposin-like domain of potato aspartic protease (StAsp-PSI) exerts antimicrobial activity on plant and human pathogens. Peptides 31, 777–785 (2010).
Neves, J., Séneca, A., Pereira, S., Pissarra, J. & Pereira, C. Abiotic stress upregulates the expression of genes involved in psv and autophagy routes. Biol. Life Sci. Forum 4, 40 (2020).
Pereira, C., Vieira, V., Pissarra, J. & Pereira, S. The journey of cardosin A in young Arabidopsis seedlings leads to evidence of a Golgi-independent pathway to the protein storage vacuole. Front. Plant Sci. 14, 1085898 (2023).
Shikata, M. & Ezura, H. Micro-Tom Tomato as an alternative plant model system: mutant collection and efficient transformation. Methods Mol. Biol. 1363, 47–55 (2016).
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39, W13–17 (2011).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2—a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Bioinformatics 8, 275–282 (1992).
Blum, M. et al. InterPro: the protein sequence classification resource in 2025. Nucleic Acids Res. 53, D444–D456 (2025).
Lacerda, A. L. M. et al. Reference gene selection for qPCR analysis in tomato-bipartite begomovirus interaction and validation in additional tomato-virus pathosystems. PLoS ONE 10, e0136820 (2015).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Alexieva, V., Sergiev, I., Mapelli, S. & Karanov, E. The effect of drought and ultraviolet radiation on growth and stress markers in pea and wheat. Plant Cell Environ. 24, 1337–1344 (2001).
Heath, R. L. & Packer, L. Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. Arch. Biochem. Biophys. 125, 189–198 (1968).
Bates, L. S., Waldren, R. P. & Teare, I. D. Rapid determination of free proline for water-stress studies. Plant Soil 39, 205–207 (1973).
Soares, C., Carvalho, M. E. A., Azevedo, R. A. & Fidalgo, F. Plants facing oxidative challenges—a little help from the antioxidant networks. Environ. Exp. Bot. 161, 4–25 (2019).
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in Planta. Plant Physiol. 133, 462–469 (2003).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Sparkes, I. A., Teanby, N. A. & Hawes, C. Truncated myosin XI tail fusions inhibit peroxisome, Golgi, and mitochondrial movement in tobacco leaf epidermal cells: a genetic tool for the next generation. J. Exp. Bot. 59, 2499–2512 (2008).
Lufrano, D. et al. Molecular cloning and characterization of procirsin, an active aspartic protease precursor from Cirsium vulgare (Asteraceae). Phytochemistry 81, 7–18 (2012).
Curto, P. et al. Establishing the yeast Kluyveromyces lactis as an expression host for production of the saposin-like domain of the aspartic protease cirsin. Appl. Environ. Microbiol. 80, 86–96 (2013).
Terauchi, K., Asakura, T., Nishizawa, N. K., Matsumoto, I. & Abe, K. Characterization of the genes for two soybean aspartic proteinases and analysis of their different tissue-dependent expression. Planta 218, 947–957 (2004).
Mazorra-Manzano, M. A. & Yada, R. Y. Expression and characterization of the recombinant aspartic proteinase A1 from Arabidopsis thaliana. Phytochemistry 69, 2439–2448 (2008).
Glathe, S. et al. Transport and activation of the vacuolar aspartic proteinase phytepsin in barley (Hordeum vulgare L. J. Biol. Chem. 273, 31230–31236 (1998). .
Bryksa, B. C. & Yada, R. Protein structure insights into the bilayer interactions of the saposin-like domain of Solanum tuberosum aspartic protease. Sci. Rep. 7, 16911 (2017).
Mendieta, J. R., Pagano, M. R., Muñoz, F. F., Daleo, G. R. & Guevara, M. G. Antimicrobial activity of potato aspartic proteases (StAPs) involves membrane permeabilization. Microbiology 152, 2039–2047 (2006).
Frey, M. E., D’Ippolito, S., Pepe, A., Daleo, G. R. & Guevara, M. G. Transgenic expression of plant-specific insert of potato aspartic proteases (StAP-PSI) confers enhanced resistance to Botrytis cinerea in Arabidopsis thaliana. Phytochemistry 149, 1–11 (2018).
Guevara, M. G., Veríssimo, P., Pires, E., Faro, C. & Daleo, G. R. Potato aspartic proteases: induction, antimicrobial activity and substrate specificity. J. Plant Pathol. 86, 233–238 (2004).
Gao, C. et al. A unique plant ESCRT component, FREE1, regulates multivesicular body protein sorting and plant growth. Curr. Biol. 24, 2556–2563 (2014).
Zhu, J.-K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Danquah, A., de Zelicourt, A., Colcombet, J. & Hirt, H. The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol. Adv. 32, 40–52 (2014).
Kanojia, A. et al. Developmentally controlled changes during Arabidopsis leaf development indicate causes for loss of stress tolerance with age. J. Exp. Bot. 71, 6340–6354 (2020).
AbdElgawad, H. et al. High salinity induces different oxidative stress and antioxidant responses in maize seedlings organs. Front. Plant Sci. 7, 276 (2016).
Miller, G., Suzuki, N., Ciftci-Yilmaz, S. & Mittler, R. Reactive oxygen species homeostasis and signalling during drought and salinity stresses. Plant Cell Environ. 33, 453–467 (2010).
Yang, Y. & Feng, D. Genome-wide identification of the aspartic protease gene family and their response under powdery mildew stress in wheat. Mol. Biol. Rep. 47, 8949–8961 (2020).
Sampaio, M. et al. Interaction dynamics of plant-specific insert domains from Cynara cardunculus: a study of homo- and heterodimer formation. Molecules 29, 5139 (2024).
Zhao, X. et al. Insights into the mechanism of membrane fusion induced by the plant defense element, plant-specific insert. J. Biol. Chem. 295, 14548–14562 (2020).
Acknowledgements
This work was supported by UID/05748/2025 (https://doi.org/10.54499/UID/05748/2025) and UID/PRR/05748/2025 (https://doi.org/10.54499/UID/PRR/05748/2025). Miguel Sampaio and João Neves are the recipients of a FCT fellowship: UI/BD/151042/2021 (https://doi.org/10.54499/UI/BD/151042/2021) and PRT/BD/154965/2023 (https://doi.org/10.54499/PRT/BD/154965/2023), respectively.
Author information
Authors and Affiliations
Contributions
C.P., G.P.S., J.A., C.S., and M.S. conceptualized and designed the study; C.P., J.P., and F.F. acquired funding and curated data; M.S., J.N., and J.M. performed the experiments and data analysis; M.S. and C.P. wrote the original draft, with all authors contributing to the review and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Sampaio, M., Neves, J., Monteiro, J. et al. Tomato aspartic proteinases harbouring PSI domains reveal stress responsiveness, organ specificity, and conserved features. npj Sci. Plants 2, 8 (2026). https://doi.org/10.1038/s44383-026-00023-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44383-026-00023-x