Abstract
Early detection of Alzheimer’s disease (AD) is critical yet challenging, particularly in younger individuals. This study leverages artificial intelligence to analyze digital voice recordings from the Craft Story Recall task within the Longitudinal Early-onset AD Study (LEADS) to (1) detect cognitive impairment and (2) differentiate early-onset AD (EOAD) from early onset non-AD cognitive impairment (EOnonAD). Using speech samples from 120 patients and 68 cognitively unimpaired controls, we employed two classification approaches: feature-engineered machine learning and end-to-end deep learning incorporating a Large Language Model. To detect mild cognitive impairment, the feature-engineered model, using acoustic and linguistic features, achieved an AUC of 0.945 on the holdout test set, while the end-to-end model yielded an AUC of 0.988. For differentiating EOAD from EOnonAD, the feature-engineered model achieved an AUC of 0.804, and the end-to-end model yielded an AUC of 0.904 on the holdout set. Explainability analyses revealed reduced linguistic informativeness as a key AD indicator.
Similar content being viewed by others
Introduction
The moments when the insidious cognitive symptoms of Alzheimer’s disease (AD) begin to raise concerns mark the start of a long, tenuous journey, as patients and families navigate the challenge of identifying the cause among a myriad of other factors, such as fatigue, sleep deprivation, or psychiatric disorders1,2,3. These challenges are particularly pronounced in younger individuals, where early signs are often overlooked or misattributed4. This complexity helps explain why the chances of detecting AD at the earliest clinical stage may be as low as 9%5. The fact that patients in the early stages of AD have the lowest chance of receiving a correct diagnosis signals a paradoxical misalignment in care, as this is precisely the time when patients stand to benefit most from new disease-modifying therapies6,7,8,9.
Major barriers to early diagnosis include a paucity of front-line clinician knowledge about dementia10, the insensitivity of standardized screening instruments commonly used in primary care11, the frequent absence of an informant in primary care visits12, the lack of structured cognitive assessments at annual wellness visits13, and the discomfort and even embarrassment among some clinicians in administering cognitive tests14,15. These challenges are further exacerbated for patients in rural or underserved areas, who face significant disparities in accessing timely diagnostic services16. As a result, by the time AD is eventually diagnosed, many patients have already progressed beyond the stage where they could benefit from disease-modifying treatments.
In evaluating the barriers to early AD diagnosis, it immediately becomes clear that many of the longstanding challenges can be mitigated through the advances of artificial intelligence (AI), ideally with expert guidance17. AI enables the accurate and objective measurement of various behavioral domains as well as the detection of subtle patterns of change that may elude human observers18. Many of these advances can be applied remotely, broadening access to diverse populations across geographic locations. These capabilities have already transformed diagnostics in fields such as oncology19,20,21 and cardiovascular diseases22,23, yet they remain underutilized in the realm of neurodegenerative disorders.
One particularly promising domain of behavior for early detection of AD is language, as its production engages multiple cognitive capacities, including memory, executive functioning, and sustained attention24. Speech and language are rich in signals that can indicate mental states in both health and disease, with the capacity to detect neuropsychiatric disorders well in advance of their onset18,25. In this study, we provide proof-of-concept evidence that AI-based analysis of short recordings of speech is a powerful method for the early detection of cognitive impairment and its underlying pathology.
As part of the Longitudinal Early-Onset AD Study (LEADS), we digitally record the National Alzheimer’s Coordinating Center (NACC) Uniform Data Set (UDS) neuropsychological battery in three groups: cognitively unimpaired (CU) controls, patients with early-onset AD (EOAD), and patients with early-onset cognitive impairment unlikely due to AD (EOnonAD). For this study, we selected patients with a cognitive functional status of Mild Cognitive Impairment (MCI)26 and excluded LEADS patients with mild dementia because detection and differential etiological diagnosis at the earliest clinical stage is the most challenging problem. We focused on a test from the battery with rich verbal content—the Craft Story Recall task—to test the hypotheses that AI-based analysis of acoustic and linguistic features from digital voice recordings would be able to 1) detect cognitive impairment and 2) differentiate whether the impairment stems from underlying AD pathology as confirmed by amyloid PET scan. We further hypothesize that this AI-based approach would outperform manually generated cognitive test scores in detecting cognitive decline and the likely etiology of EOAD.
To test these hypotheses, we adopted two broad analytical approaches. The first approach involved feature engineering with machine learning, in which we extracted digital markers of speech and language, such as lexical diversity and syntactic complexity, known for their diagnostic relevance as well as novel features to train a machine learning classifier to detect patients with MCI relative to CU and then also to differentiate EOAD from EOnonAD. In the second approach, the End-to-End deep learning classification, we used a Large Language Model (LLM) directly on raw text without the bias of any predetermined features, allowing the model to autonomously identify linguistic patterns by leveraging its ability to process complex textual data. For each analysis, we applied explainability AI (XAI) methods to identify the most influential features driving classification decisions. This approach enhances model transparency and provides actionable insights into the underlying cognitive mechanisms of AD-related changes in speech and language.
Results
Classifying patients with mild cognitive impairment from cognitively unimpaired individuals
We first aimed at detecting mild cognitive impairment using short speech samples. We tested the hypothesis that computational analysis of acoustic and linguistic features from the Digital Voice samples would perform better than typical manually-derived cognitive test scores at detecting cognitive impairment at a mild clinical stage (in all CDR 0.5 patients compared to CU controls).
In the feature-engineered classification, the predictive performance of XGBoost models was evaluated across distinct feature sets using a nested cross-validation framework on the training data (75% of the cohort), with final validation performed on a held-out test set (25%). The model trained on randomized values yielded a mean AUC of 0.443 (95% CI: 0.308–0.577). The demographics-based model showed a mean AUC of 0.604 (95% CI: 0.488–0.720). The model trained on MoCA scores demonstrated a mean AUC of 0.843 (95% CI: 0.788–0.898). The model based on the Craft Story Recall verbatim score achieved a mean AUC of 0.904 (95% CI: 0.846–0.962). The model employing speech and language features resulted in a mean AUC of 0.881 (95% CI: 0.805–0.956).
The generalizability of each model was assessed on the independent 25% holdout set (Fig. 1). The model trained on randomized values yielded a holdout AUC of 0.597. The demographics-based model showed a holdout AUC of 0.564. The model trained on MoCA scores resulted in a holdout AUC of 0.920. The model based on the Craft Story Recall verbatim score achieved a holdout AUC of 0.863. The model employing speech and language features resulted in a holdout AUC of 0.945 (precision = 0.958, recall = 0.767, and F1-score = 0.852).
The highest numerical value of performance was based on a model using speech and language features, while the model based on demographic data was comparable to one based on a randomized variable.
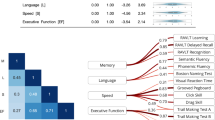
SHAP analysis highlighted the key features contributing to the predictions of various XGBoost models (Fig. 2). Positive SHAP values indicate a higher likelihood of classification as a patient with cognitive impairment; negative SHAP values indicate a higher likelihood of classification as a CU individual. The feature that contributed the most to the differentiation of cognitively impaired patients from CU was the Language Informativeness Index (LII), with CU individuals demonstrating higher LII values. This finding indicates that patients with cognitive impairment produced language during the delayed recall of the Craft story that was less semantically similar to the original story than that produced by CU individuals. The second most distinguishing feature was inter-word pause duration, with cognitively impaired patients exhibiting longer pauses compared to CU individuals. The third influential feature was the number of sentences, with cognitively impaired patients producing more sentences than CU individuals. This increased sentence count may reflect a compensatory strategy to mitigate their difficulty in constructing longer, more complex sentences. Supporting this interpretation, we observed a significant negative correlation between the number of sentences and sentence length across all participants (r = −0.66, p < 0.001). Nonetheless, this compensation appears to be partial, as the total word count among patients (mean = 36.3) remained significantly lower than that of CU individuals (mean = 54.2) (t = −5.838, p < 0.001). The last feature contributing to the classification of cognitive impairment was shorter word length.
Positive SHAP values indicate a higher likelihood of classification as a patient. Red represents higher values for each feature, while blue represents lower feature values. LII was the strongest predictor, with low language informativeness values increasing the probability that the individual was a patient with cognitive impairment. LII was followed by pause duration (high values make cognitive impairment more likely), number of sentences (high values make cognitive impairment more likely), and word length (low values make cognitive impairment more likely).
In the end-to-end deep learning classification for detecting mild cognitive impairment, we fine-tuned RoBERTa-base directly on speech transcripts, eliminating the need for manual feature engineering. The model demonstrated strong performance during hyperparameter optimization via 5-fold cross-validation, achieving a mean AUC of 0.927 (95% CI: 0.892–0.962) with an optimal learning rate of 3e−5 and batch size of 16.
When this best-performing model was evaluated on the completely held-out test set, without any additional training to prevent overfitting, it maintained generalizability. The model achieved an AUC of 0.988 on unseen data (precision = 1.00, recall = 0.93, and F1-score = 0.97).
To better understand the linguistic patterns driving model predictions for RoBERTa, we employed the LIME approach, which identifies word-level contributions to classification decisions (Fig. 3). By aggregating the importance scores of individual words, this method provides a broader understanding of the linguistic patterns that consistently contribute to the model’s predictions. Using LIME, the importance of each word was calculated across all participant texts and aggregated into a collective importance distribution. Positive LIME values indicate a higher likelihood of classification as a patient with cognitive impairment. Words that appeared frequently across participants with consistently high importance scores were identified and visualized. The top 25 influential words, ranked by their average absolute importance, are presented in Fig. 3. The LIME analysis identifies the most influential words that distinguish patients with cognitive impairment from CU individuals include the absence of words indicating specific people (such as “Ricky,” “neighbor,” and “Maria,”), words denoting temporal specificity (such as “Monday” and “afternoon”, ‘3:30”), and specific verbs (“retrieve”, “barking”).
Words with more positive (rightward) values were more frequent in the transcripts of patients with cognitive impairment, whereas words with more negative (leftward) values were more frequent in the transcripts of cognitively unimpaired individuals.
Classifying patients with EOAD from those with EOnonAD
We next classified cognitively impaired patients as EOAD or EOnonAD. We tested the hypothesis that computational analysis of acoustic and linguistic features from the digital voice samples would succeed at the challenge of differential diagnosis, classifying cognitively impaired patients into EOAD or EOnonAD as the likely etiology, which we did not expect traditional cognitive test scores to be able to do very well.
We evaluated the performance of various models using XGBoost on multiple feature sets to distinguish EOAD from EOnonAD (Fig. 4). The model trained on randomized values yielded a mean AUC of 0.584 (95% CI: 0.452−0.715). The demographics-based model showed a mean AUC of 0.616 (95% CI: 0.490−0.742). The model trained on MoCA scores demonstrated a mean AUC of 0.736 (95% CI: 0.564−0.909). The model based on the Craft Story Recall verbatim score achieved a mean AUC of 0.786 (95% CI: 0.676−0.895). The model employing speech and language features resulted in a mean AUC of 0.789 (95% CI: 0.638−0.940).
These data support the hypothesis that speech and language features from digital voice samples of Craft Story delayed recall outperformed standard neuropsychological scoring of this test as well as the MoCA, neither of which performed very well.
The generalizability of each model was assessed on the independent 25% holdout set. The model trained on randomized values yielded a holdout AUC of 0.495. The demographics-based model showed a holdout AUC of 0.538. The model trained on MoCA scores resulted in a holdout AUC of 0.593. The model based on the Craft Story Recall verbatim score achieved a holdout AUC of 0.606. The model employing speech and language features resulted in a holdout AUC of 0.804 (precision = 0.773, recall = 0.895, and F1-score = 0.829).
We used SHAP to determine the importance of each feature in classifying EOAD from EOnonAD (Fig. 5). Positive SHAP values indicate a higher likelihood of classification as EOAD. As shown in Fig. 5, the most influential feature in indicating EOAD was higher word frequency (the use of more common words), lower rate of proper nouns (e.g., “Ricky”), higher adjective modifiers, and lower temporal modifiers.
Positive SHAP values indicate a higher likelihood of classification as EOAD. Red represents high feature values, while blue represents low feature values. EOAD patients tended to use words of higher frequency, fewer proper nouns, more adjectival modifiers, and fewer temporal modifiers.
In the end-to-end deep learning classification to distinguish EOAD from EOnonAD, we employed RoBERTa-base, feeding it the raw transcribed text from the delayed recall task. We performed a hyperparameter search over learning rates (1e−5, 2e−5, 3e−5) and batch sizes (8, 16) using a stratified 5-fold cross-validation on the 75% training set. The best configuration (learning rate: 1e−05, batch size: 16) achieved a mean cross-validation AUC of 0.778 (95% CI: 0.701–0.856). The generalizability of the best model from cross-validation was assessed a single time on the independent 25% holdout set. The model achieved a holdout AUC of 0.904 (precision = 1.000, recall = 0.684, and F1-score = 0.812).
The LIME analysis highlighted the key words driving the end-to-end deep learning classification of EOAD versus EOnonAD patients (Fig. 6). Positive LIME values indicate a higher likelihood of classification as EOAD. EOnonAD patients used more specific words such as “dogs,” and “barking” compared to EOAD patients. EOnonAD were also more likely to use proper nouns such as “Ricky”. EOAD patients used words that did not appear in the story, such as “remember,” often used in sentences such as “I don’t remember.”
Words with more positive (rightward) values were more frequent in the transcripts of patients with EOAD, whereas words with more negative (leftward) values were more frequent in the transcripts of patients with EOnonAD. EOnonAD patients tended to use more specific words (e.g., “dogs” and “Ricky”), whereas EOAD patients tended to use words that were not in the story (e.g., “idea,” “remember”).
Discussion
This proof-of-concept study demonstrates the transformative power of AI analysis on Digital Voice in addressing a pressing challenge in dementia care: the early and accurate detection of cognitive impairment and the differentiation of Alzheimer’s pathology from other etiologies. By analyzing digital voice samples from a brief neuropsychological task rich in information relevant for cognitive function—the Craft Story delayed recall—we achieved high classification performance that exceeded that of traditional MoCA and manually scored Craft Story test scores. Furthermore, our analyses focused on individuals with a Clinical Dementia Rating (CDR) of 0.5 or lower, emphasizing detection at the earliest stages of cognitive decline when disease detection is most challenging and interventions are likely to be most impactful. This performance highlights the fact that there is much more information in the verbal behavior being produced during test performance than that measured by standardized neuropsychological scores, supporting the value of voice recordings as a digital behavioral marker. Findings such as this from our multi-center consortium underscore the remarkable value of digital recordings of cognitive assessments, which is being developed broadly within the Alzheimer’s Disease Centers Network and National Alzheimer’s Coordinating Center.
We applied two approaches: (1) a feature-engineered machine learning model that used extracted digital markers of speech and language, and (2) an end-to-end deep learning model, RoBERTa, which is an LLM that classified raw transcripts without relying on predefined features. While most recent studies investigating this topic have combined hand-selected features with embeddings derived from LLMs to maximize performance27,28,29,30,31, we intentionally kept these approaches separate to allow for a direct comparison between patterns defined by human expertise and those discovered by the model. The two approaches present a tradeoff between performance and transparency. While feature-engineered approaches provide full transparency into the classification task, end-to-end deep learning models generally offer higher performance.
Consistently, across both classification tasks in this study, the end-to-end deep learning approach outperformed the feature-engineered model. These findings align with prior work showing that BERT family models outperformed hand-selected linguistic and acoustic features in detecting clinically-diagnosed AD dementia relative to cognitively unimpaired individuals32. A prior study of a combined group of patients with MCI/mild dementia demonstrated that AI-based analysis of a short story detected cognitive impairment with an AUC of 0.85 and predicted amyloid-PET positivity within the MCI/dementia patients with an AUC of 0.7833. The same group found that selected lexical and temporal speech features could detect MCI with AUCs in the mid-0.70 s34. Thus, we extend prior work by applying these techniques to milder cases and differentiating the etiology of cognitive impairment based on an amyloid PET biomarker35. Furthermore, in the prior study, the average accuracy of feature-engineered detection of clinical AD dementia was 76% and that of the BERT-based procedure was 82%. To our knowledge, none of the prior studies have addressed the challenge of differential etiologic diagnosis based on biomarkers, where our accuracy reached 90%. The superior performance of end-to-end approaches suggests that some indicators of AD either remain undiscovered despite their discrete and human-interpretable nature or are fundamentally latent statistical relationships in language usage that only deep learning models can comprehensively capture.
A highlight of this study is the application of explainability algorithms to probe model decision-making. In the feature-engineered approach, we used SHAP to identify the most influential features driving classification outcomes. For detecting cognitive impairment, the most predictive feature was the Language Informativeness Index (LII), a measure we developed36 of overall semantic similarity between a participant’s retelling and the original story, indicating that reduced semantic fidelity is a hallmark of early impairment. Another robust indicator was shorter word length. In distinguishing AD pathology, the use of less specific nouns, including proper nouns, stood out, consistent with the prior literature36,37,38,39,40. EOAD patients also used fewer temporal modifiers, such as the timing of events. These features, and thereby insights into the underlying cognitive mechanisms, are not captured through traditional approaches.
Although acoustic features such as shimmer, jitter, and harmonic-to-noise ratio were included in the analysis, they did not emerge as significant contributors in the feature selection process and ultimately did not enhance classification performance. Instead, text-based features emerged as robust indicators of early cognitive decline. This finding is consistent with prior work41, where only text-based features remained significant after correction for multiple comparisons. Pause duration is a noteworthy feature. While technically derived from audio and often classified as an acoustic feature, pause duration differs from classic voice-quality metrics such as shimmer and jitter. Rather than reflecting vocal biomechanics, pause duration additionally reflects underlying cognitive or language impairments, such as a dysexecutive loss of train of thought, memory deficits, or lexical retrieval difficulty. For this reason, we separated pause duration as a temporal rate-based feature rather than a purely acoustic one.
To interpret the end-to-end model, we applied LIME, which revealed word-level indicators of group membership. Among cognitively unimpaired individuals, the presence of specific information—such as proper names (“Maria”, “Ricky”) and temporal markers (“Monday”, “afternoon”)—was predictive of preserved cognition. In contrast, patients with EOAD used vague terms such as demonstratives and pronouns instead of specific nouns. EOAD patients also used words that were not in the original story, such as “remember”—typically when remarking that they do not remember something. These words are not captured by conventional scoring systems but clearly contain information useful for classification.
Crucially, both analytical approaches converged on a core factor likely driving classification outcomes: reduced informativeness. Although patients attempted to compensate for their shorter sentences by producing more of them, their language remained vague. They relied on more general terms, increased pronoun usage, and fewer temporal markers, resulting in less specific and less informative communication. Some of these patterns were explicitly captured in the feature-engineered approach using SHAP, while others emerged through the words identified by LIME in the end-to-end model. This interpretation is consistent with our recent cross-linguistic work on Alzheimer’s disease using a picture description task. In both English and Persian, AD patients used more pronouns, higher-frequency words, and shorter sentences, all of which were correlated with reduced LII36. Historically, reduced informativeness has been measured by tallying predefined elements—such as subjects, places, objects, and actions—mentioned in picture descriptions42,43. However, that approach is constrained by its categorical rather than graded scoring system and its reliance on a fixed set of predefined elements. The automated techniques used here overcome these limitations by providing a more nuanced, continuous, and context-sensitive measure of informativeness.
It is important to highlight a key distinction between the explainability methods used in the two classification approaches. In the feature-engineered model, SHAP provides a comprehensive explanation by quantifying the contribution of all features to the XGBoost model’s predictions. In contrast, while explainability methods applied to LLMs, such as LIME, can identify prominent word-level features that influence individual predictions, they overlook the more distributed representations and nonlinear dependencies that transformer models like RoBERTa rely on. As a result, LIME offers only a partial glimpse into the model’s decision-making, leaving many latent cues unexplored44.
Looking forward, expanding this work to cohorts of larger sizes, wider age ranges and socioeconomic backgrounds, languages other than English, and free speech tasks will help test the generalizability of these linguistic markers. In addition, future research should investigate whether these speech-derived indicators can predict disease trajectory. The success of such efforts could enable automated, scalable screening within primary care settings to flag individuals for further evaluation, much like routine vital sign checks or other physical examination measures. An obvious vision for the future would be to incorporate brief real-world valid verbal cognitive tests (such as story retelling) with digital voice recording and AI analytic technology into the primary care cognitive screening component of Medicare annual wellness visits to facilitate the early identification of patients—particularly those with longitudinal change relative to their own baseline—for referral for further evaluation. The use of these methods by specialists to risk-stratify patients for invasive or expensive biomarker tests is also supported by their performance in this study in predicting amyloid PET status in the patient group. Integrating digital voice and AI into routinely obtainable behavioral samples, such as speech during a clinical appointment or even a pre-visit smartphone screen, has the potential to streamline the path from suspicion to timely diagnosis and life-changing interventions.
Methods
Participant information
This study used digital voice samples from participants in LEADS, a prospective, multisite observational study registered on ClinicalTrials.gov (NCT03507257)45. This study is designed to provide a comprehensive characterization of the baseline and longitudinal cognitive and functional characteristics, along with MRI, amyloid PET, tau PET, plasma, and cerebrospinal fluid (CSF) measures in the largest sample of EOAD patients to date. LEADS builds on the infrastructures established by the NACC and the Alzheimer’s Disease Neuroimaging Initiative (ADNI). Biomarker collection in LEADS closely mirrors ADNI’s protocols, leveraging informatics systems developed by the Laboratory of Neuro Imaging (LONI). The study’s framework includes eight specialized cores -- Administrative, Clinical, MRI, PET, Genetics and Biorepository, Biostatistics, Informatics, and Neuropathology -- and operates across 19 clinical sites. The coordinating center, based at the University of Southern California’s Alzheimer’s Therapeutic Research Institute (ATRI), provides support for electronic data capture, regulatory compliance, data management, clinical monitoring, safety oversight, and reporting to the Data and Safety Monitoring Board45.
This research was conducted in accordance with the Declaration of Helsinki. LEADS has designated Indiana University (IU) as the central Institutional Review Board (IRB). The study employs the SMART IRB agreement to streamline the reliance process, with local Human Research Protection Programs (HRPPs) at each site handling state and local policy adherence. These HRPPs relay relevant guidelines to the ATRI Regulatory Team and the central IU IRB, ensuring site-specific documents like informed consent forms and HIPAA authorizations are customized from IU-approved templates. The IU IRB then reviews and approves these documents before site activation. Participants in LEADS provide informed consent in accordance with the Declaration of Helsinki, U.S. federal regulations, state laws, and IU IRB policies. The IU IRB is responsible for overseeing reportable events, study-wide and site-specific amendments, and annual renewals, while local HRPPs manage additional reviews as per the SMART IRB framework. The IRB Reliance Exchange (IREx) online portal is used for the storage and distribution of approved documents across sites.
LEADS enrolls cognitively impaired patients under the age of 65 who must meet National Institute on Aging–Alzheimer’s Association (NIA-AA) criteria for MCI or mild dementia and have a global Clinical Dementia Rating (CDR) score ≤1. LEADS does not exclude individuals with predominantly non-amnestic presentations. Therefore, patients meeting criteria for the dysexecutive, logopenic primary progressive aphasia, or posterior cortical atrophy variants are eligible to enroll. In addition, LEADS enrolls CU individuals under the age of 65 who must have a Mini-Mental State Examination (MMSE) score of ≥24, a global CDR = 0, and a score within the cognitively normal range on neuropsychological testing45.
All LEADS participants undergo comprehensive clinical evaluations using the NACC UDS protocol, neuropsychological assessment, magnetic resonance imaging46, amyloid PET with Florbetaben (FBB) PET, and tau PET with Flortaucipir (FTP) PET47,48. These FDA-approved radiotracers are validated for detecting moderate-to-frequent neuritic plaques and Braak V-VI neurofibrillary tangles, ensuring reliable assessment of Alzheimer’s pathology49. All LEADS participants are invited to consent to a blood draw and lumbar puncture for fluid biomarker evaluation50. A detailed visual read and quantitative assessment of each patient’s amyloid PET scan is performed to classify each patient as EOAD or EOnonAD (see below), and genetic analysis is used to confirm the absence of autosomal dominant pathogenic genetic abnormalities51.
To classify each cognitively impaired participant as EOAD or EOnonAD47, Florbetaben (FBB)-PET scans are visually read as amyloid-positive or amyloid-negative using validated criteria by one of the certified PET core physicians at UCSF. Each FBB scan is first read by a single physician, without access to participants’ clinical information or scan quantification. After reading the scan and saving their interpretation, each physician has access to the SUVRPET-only value. If both classifications are positive, the participant is assigned to the EOAD cohort. If both are negative, the participant is assigned to the EOnonAD cohort. If visual read and quantification-based classification are incongruent, a second visual read is performed by an additional reader who is blind to the previous read and quantification. This second read is used as a tiebreaker.
In this study, we included 120 patients with MCI and 68 CU controls. To focus on the patient group with the mildest impairment, we analyzed individuals with a Clinical Dementia Rating of ≤ 0.5 (excluding those with mild dementia, CDR = 1). Of these patients, 76 participants had a diagnosis of MCI due to EOAD, and 44 had a diagnosis of MCI due to EOnonAD. For clinical and demographic information on the participants, see Table 1. The sample size for each group reflects the digital voice samples available for analysis.
The NACC UDS 3.0 Neuropsychological Test Battery is a standardized set of cognitive assessments designed to evaluate various cognitive domains. The UDS 3.0 battery includes a global cognitive screening test (the Montreal Cognitive Assessment, MoCA), and tests that measure memory (Craft Story 21, Benson Figure Delayed recall), attention/working memory/processing speed (Number Span forward and Backward, Trail-making Test part A), executive function (Trail-making Test part B, verbal fluency), and language (verbal fluency, Multi-lingual Naming Test), providing a reasonably comprehensive assessment of cognitive function at each time point. In LEADS, this battery is supplemented with other tests, including the Mini-Mental State Examination. The NACC UDS 3.0 neuropsychological measures are widely used across Alzheimer’s Disease Research Centers (ADRCs) in the United States, allowing for data harmonization and comparative research across diverse populations.
Digital voice samples
The digital voice samples used in the analyses for the present study were obtained from the delayed recall sections of the Craft Story 21 Recall test52. The Craft Story 21 Recall test evaluates a participant’s ability to remember a short story. In the Immediate Recall phase, the participant retells the story immediately after hearing it. During the Delayed Recall phase, the participant recounts the story after a 20-minute interval. Scoring is conducted separately for verbatim recall (maximum score: 44) and paraphrase recall (maximum score: 25). These verbal responses from LEADS participants are digitally recorded. For this study, we analyzed digital voice samples from the verbatim scoring of the delayed recall phase of the test.
To record verbal neuropsychological assessments, each LEADS site is supplied with an Olympus WS-852 digital recorder. When starting each session, the coordinator administering the testing makes a recording stating the participant identifier (ID) and visit ID to distinguish the new set of recordings from previous recordings (participant name, date of birth, or any other protected health information (PHI) is not recorded). After reading instructions to the participant for each test, the coordinator starts the recording and records each test as a separate file. After each session, recordings are downloaded to a computer connected to a secure shared file directory on the site’s local network. Files are named using a standard naming convention that includes indicators for the study name, site identifier (ID), participant ID, and visit ID (e.g., LDS0370001_Craft_R_V1). The files are then uploaded for permanent storage to IU’s Microsoft OneDrive audio recordings account. Each site is responsible for ensuring that no PHI is included in the recordings prior to uploading the files. The additional activities required for capturing and managing these digital audio files at each site require approximately 10–15 minutes per participant per session.
For this project, we downloaded the digital audio files. To ensure the recordings captured only the participants’ responses, interjections from the coordinator at each site were removed manually, a process that takes about one to two minutes per audio file. The samples were automatically transcribed using Google Cloud Speech-to-Text. Each text file was then manually corrected by listening to the audio file, a process that takes about three to five minutes per file. As part of the preprocessing, the transcripts were converted to lowercase, a standard practice aimed at reducing variability in the input data, such as differences arising from the position of words within a sentence. The final text file included audio transcripts with each line containing one sentence. A sentence was defined as an independent clause and all clauses dependent on it53.
The primary objectives of this study were to test the hypothesis that (1) computational analysis of acoustic and linguistic features from the digital voice samples would perform better than typical manually-generated test scores at detecting cognitive impairment (in very mildly impaired patients compared to CU controls; and (2) acoustic and linguistic features from the digital voice samples would be able to differentiate EOAD from EOnonAD patients.
Classification approaches
To test each hypothesis, we employed two distinct approaches. In the first approach, audio files and transcribed texts were used to extract acoustic features from digital voice audio files and linguistic features from transcribed language samples (Fig. 7A). A machine learning classifier, XGBoost, was trained to perform the classifications based on the extracted features. We refer to this approach as feature-engineered classification. In the second approach, no features were explicitly extracted (Fig. 7B). Instead, the raw transcribed text was directly input into an LLM, RoBERTa54, allowing it to perform the classification without any prior assumptions about the data. We refer to this as the End-to-End deep learning classification. Of the available LLMs, we chose RoBERTa54 as in our recent work that benchmarked various LLMs-- including LLAMA-3-8B-Instruct, Mistral, GPT-3.5-turbo, and GPT-4o-mini-- we found the model has superior performance in classifying texts of patients with neurodegenerative disorders55. All analyses were performed using Python 3.12.3.
A Feature-engineered and B end-to-end deep learning.
We measured various acoustic features, speech rate-based features, lexicosyntactic features, and semantic features from each Digital Voice sample, as detailed below.
Acoustic features were extracted from digital voice audio files. The audio preprocessing pipeline involved volume normalization, noise reduction, and application of a bandpass filter. We included a wide range of voice features, comprising fundamental frequency statistics (meanF0Hz, stdevF0Hz), a measure of vocal breathiness (harmonic-to-noise ratio), various types of frequency perturbation to capture vocal roughness (local jitter, local absolute jitter, relative average perturbation, 5-point period perturbation quotient, difference of differences of periods), distinct types of amplitude perturbation (local shimmer, 3-point amplitude perturbation quotient, 5-point amplitude perturbation quotient, 11-point amplitude perturbation quotient, difference of differences of amplitudes), Mel-frequency cepstral coefficients, and the average of the first four formant frequencies (f1_mean, f2_mean, f3_mean, f4_mean) to characterize vocal tract resonance, using Parselmouth Praat Scripts in Python56.
The rate-based features captured temporal characteristics of speech in audio files: articulation rate (number of syllables per second), speech rate (number of words per second), and pause duration (average word-to-word intervals in seconds). For these measures, we developed a Python-based pipeline that integrates Google Cloud Speech-to-Text API for transcription, and the Syllable counts for each word were obtained using the Carnegie Mellon University Pronouncing Dictionary (cmudict Python package) for syllable counting. Google Cloud Speech-to-Text API was used to generate word-level timestamps. The transcription process included automatic punctuation and word time offsets, enabling precise calculation of word durations and inter-word pauses.
To extract lexicosyntactic features from transcripts of digital voice files, we used our MGH FTD Unit Quantitext toolbox as previously described57,58. These features included word frequency, syntax frequency, sentence length, and word length. Furthermore, part-of-speech tagging and syntactic dependency parsing were performed using the Stanza natural language processing toolkit59, a pipeline for multi-lingual text analysis. Stanza utilizes deep learning-based models trained on Universal Dependencies treebanks to provide robust and accurate linguistic annotations, including tokenization, lemmatization, part-of-speech tagging, and dependency parsing.
As the above features capture specific and localized aspects of the participants’ responses rather than the overall meaning, we also incorporated our Language Informativeness Index (LII), which measures the semantic similarity between the participants’ responses and the original story36. To compute LII, we used a transformer-based model, “bert-base-nli-mean-tokens” from the Sentence Transformers library. This specialized version of the BERT (Bidirectional Encoder Representations from Transformers) model is optimized for generating sentence-level embeddings60. The “bert-base” variant is a compact, efficient architecture with 12 transformer blocks, 768 hidden units, and 12 attention heads, making it ideal for natural language processing tasks. The transcripts of the original story and participant responses were transformed into a high-dimensional vector representation (embedding) using the BERT model, and LII was calculated by measuring the cosine similarity between these embeddings, with a value of 1 indicating maximum similarity.
Machine learning classifier. For feature-based classifications, we employed XGBoost classifiers combined with a nested cross-validation pipeline applied to the 75% training set to ensure unbiased performance estimation. XGBoost (eXtreme Gradient Boosting) is a powerful machine-learning algorithm known for its efficiency, scalability, and ability to capture complex patterns in data61. It builds an ensemble of decision trees trained sequentially, where each new tree corrects the errors made by the previous ones.
To prevent overfitting and over-optimistic results, both feature selection and hyperparameter tuning were performed internally within each fold of the cross-validation on the training data. Specifically, we used a stratified 10-fold cross-validation framework. Within each of the 10 training folds, we performed a sequential forward selection process to identify the most predictive feature set62. Subsequently, a nested hyperparameter grid search was conducted on that same training fold to optimize the model parameters. This ensured that the test fold was completely held out from all aspects of model development to provide a reliable estimate of generalizability.
We evaluated the classification performance of a series of models based on specific variables. The first model was based on a random variable to establish a chance-level baseline. The next model used demographic data, including age, sex, and years of education. We also ran a model using the MoCA score and a model using the Craft Story verbatim score on the delayed story recall. Finally, we ran the feature-engineered model using the speech and language features. To address the class imbalance, each XGBoost model was trained using a scale_pos_weight parameter, which weights the minority class during model training based on the inverse class frequency to avoid biased predictions and improve model calibration63.
The hyperparameter grid search was performed inside the cross-validation loop as described above. We used the scikit-learn package to explore a focused grid of key hyperparameters: maximum tree depth, learning rate, and subsample ratio. This nested approach allowed us to systematically identify optimal hyperparameter configurations while maintaining computational feasibility and ensuring that the reported performance metrics are unbiased.
To understand the contribution of individual features to the predictions, we employed SHAP (SHapley Additive exPlanations) for model interpretation64. SHAP values provide a unified measure of feature importance by quantifying the contribution of each feature to a specific prediction. This approach allowed us to identify which features had the most significant impact (and in what direction) on the model’s decisions.
In our end-to-end deep learning approach, we performed classification directly on speech transcripts without manual feature engineering. We employed the base RoBERTa-base model, a robust transformer architecture pre-trained on a large corpus of text, for binary sequence classification. The model was initialized using the RobertaForSequenceClassification class from the Hugging Face transformers library and fine-tuned for two output classes.
To enhance performance on our imbalanced dataset and improve generalization, we implemented several techniques. We mitigated class imbalance by down-weighting the loss assigned to well-classified examples and focusing training on harder, misclassified instances using Focal Loss. Furthermore, we used a WeightedRandomSampler during training to ensure each batch contained a balanced representation of both classes, preventing the model from being biased toward the majority class.
We also employed data augmentation through synonym replacement to artificially expand the training set. We conducted a systematic grid search to optimize key hyperparameters, evaluating learning rates (1e-5, 2e-5, 3e-5) and batch sizes (8, 16) using a 5-fold stratified cross-validation on the training cohort. This rigorous process identified an optimal learning rate of 3e-5 and a batch size of 16. The model was fine-tuned using the AdamW optimizer with weight decay (0.01) and gradient clipping (max norm=1.0). We incorporated a linear learning rate scheduler with warmup and implemented early stopping with a patience of 3 epochs to prevent overfitting.
The best-performing model from cross-validation was selected and evaluated directly on the held-out test set (25% of the data) without any additional training, ensuring an unbiased assessment of its generalization performance. Predictions were made by applying a softmax function to the model’s output logits to compute class probabilities.
To interpret the predictions of the RoBERTa model, we employed the Local Interpretable Model-agnostic Explanations (LIME) technique65. LIME generates interpretable explanations for individual predictions by perturbing input text and measuring the impact of each token on the model’s output probabilities. The model’s softmax probabilities were passed to LIME, which identified the top influential words for each instance. Feature importance scores were aggregated across all participants to highlight consistent patterns in token contributions to classification decisions. For visualization, the top 50 influential words were selected and plotted, with importance scores distributed along a horizontal axis and words displayed vertically.
For each classification, we employed a stratified holdout validation method, partitioning the data into a 75% training set (used for cross-validation and hyperparameter tuning) and a held-out 25% test set, preserving the original class distribution in both splits. Classification performance was evaluated using the Area Under the Receiver Operating Characteristic curve (AUC-ROC), in addition to precision, recall, and F1-score for language-based models.
Data availability
The data and code associated with this work can be accessed through a request to the senior author, Bradford Dickerson.
Code availability
The data and code associated with this work can be accessed through a request to the senior author, Bradford Dickerson.
Change history
31 December 2025
A Correction to this paper has been published: https://doi.org/10.1038/s44400-025-00055-7
References
Gaugler, J. E. et al. Characteristics of patients misdiagnosed with Alzheimer’s disease and their medication use: an analysis of the NACC-UDS database. BMC Geriatr. 13, 137 (2013).
Becker, J. T., Boller, F., Lopez, O. L., Saxton, J. & McGonigle, K. L. The natural history of Alzheimer’s disease. Description of study cohort and accuracy of diagnosis. Arch. Neurol. 51, 585–594 (1994).
Klatka, L. A., Schiffer, R. B., Powers, J. M. & Kazee, A. M. Incorrect diagnosis of Alzheimer’s Disease: A Clinicopathologic Study. Arch. Neurol. 53, 35–42 (1996).
Tsoukra, P. et al. The Diagnostic Challenge Of Young-onset Dementia Syndromes And Primary Psychiatric Diseases: Results From A Retrospective 20-year Cross-sectional Study. J. Neuropsychiatry Clin. Neurosci. 34, 44–52 (2022).
Valcour, V. G., Masaki, K. H., Curb, J. D. & Blanchette, P. L. The detection of dementia in the primary care setting. Arch. Intern Med 160, 2964–2968 (2000).
McDade, E. et al. Lecanemab in patients with early Alzheimer’s disease: detailed results on biomarker, cognitive, and clinical effects from the randomized and open-label extension of the phase 2 proof-of-concept study. Alzheimers Res Ther. 14, 191 (2022).
van Dyck, C. H. et al. Lecanemab in early Alzheimer’s Disease. N. Engl. J. Med. 388, 9–21 (2023).
Sims, J. R. et al. Donanemab in Early Symptomatic Alzheimer disease: The TRAILBLAZER-ALZ 2 Randomized Clinical Trial. JAMA 330, 512–527 (2023).
Liss, J. L. et al. Practical recommendations for timely, accurate diagnosis of symptomatic Alzheimer’s disease (MCI and dementia) in primary care: a review and synthesis. J. Intern. Med. 290, 310–334 (2021).
Bernstein, A. et al. Dementia assessment and management in primary care settings: a survey of current provider practices in the United States. BMC Health Serv. Res. 19, 919 (2019).
Mitchell, A. J. A meta-analysis of the accuracy of the mini-mental state examination in the detection of dementia and mild cognitive impairment. J. Psychiatr. Res. 43, 411–431 (2009).
Sabbagh, M. N., Malek-Ahmadi, M. & Belden, C. M. The use of informant-based questionnaires in differentiating mild cognitive impairment from normal aging. Expert Rev. Neurother. 12, 637–639 (2012).
Jacobson, M., Thunell, J. & Zissimopoulos, J. Cognitive assessment at medicare’s annual wellness visit in fee-for-service and medicare advantage plans. Health Aff. Proj. Hope 39, 1935–1942 (2020).
Downs, M., Cook, A., Rae, C. & Collins, K. E. Caring for patients with dementia: The GP perspective. Aging Ment. Health 4, 301–304 (2000).
van Hout, H., Vernooij-Dassen, M., Bakker, K., Blom, M. & Grol, R. General practitioners on dementia: tasks, practices and obstacles. Patient Educ. Couns. 39, 219–225 (2000).
Bradford, A., Kunik, M. E., Schulz, P., Williams, S. P. & Singh, H. Missed and delayed diagnosis of dementia in primary care: prevalence and contributing factors. Alzheimer Dis. Assoc. Disord. 23, 306–314 (2009).
Everett S. S., et al. From tool to teammate: a randomized controlled trial of clinician-AI collaborative workflows for diagnosis. medRxiv. Published online June 8, 2025:2025.06.07.25329176. https://doi.org/10.1101/2025.06.07.25329176.
Rezaii, N., Wolff, P. & Price, B. H. Natural language processing in psychiatry: the promises and perils of a transformative approach. Br. J. Psychiatry 220, 251–253 (2022).
Ehteshami Bejnordi, B. et al. Diagnostic assessment of deep learning algorithms for detection of lymph node metastases in women with breast cancer. JAMA 318, 2199–2210 (2017).
Ainscough, B. J. et al. A deep learning approach to automate refinement of somatic variant calling from cancer sequencing data. Nat. Genet 50, 1735–1743 (2018).
Chang, P. et al. Deep-learning convolutional neural networks accurately classify genetic mutations in gliomas. AJNR Am. J. Neuroradiol. 39, 1201–1207 (2018).
Bello, G. A. et al. Deep-learning cardiac motion analysis for human survival prediction. Nat. Mach. Intell. 1, 95–104 (2019).
Poplin, R. et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat. Biomed. Eng. 2, 158–164 (2018).
Levelt W. J. M. Speaking: From Intention to Articulation. A Bradford Book; (1989).
Eyigoz E., Mathur S., Santamaria M., Cecchi G., Naylor M. Linguistic markers predict onset of Alzheimer’s disease. eClinicalMedicine. 28. https://doi.org/10.1016/j.eclinm.2020.100583 (2020).
Atri A., et al. Alzheimer’s Association clinical practice guideline for the diagnostic evaluation, testing, counseling, and disclosure of suspected Alzheimer’s Disease and Related Disorders (DETeCD-ADRD): Executive summary of recommendations for primary care. Alzheimers Dement. https://doi.org/10.1002/alz.14333.
Searle T., Ibrahim Z., Dobson R. Comparing natural language processing techniques for Alzheimer’s Dementia prediction in spontaneous speech. In: Interspeech 2020. ISCA; 2192-2196 (2020).
Martinc M., Pollak S. Tackling the ADReSS challenge: a multimodal approach to the automated recognition of Alzheimer’s Dementia. In: Interspeech 2020. ISCA; 2157-2161 (2020).
Taghibeyglou B., Rudzicz F. Who needs context? Classical techniques for Alzheimer’s disease detection. In: Naumann T., Ben Abacha A., Bethard S., Roberts K., Rumshisky A., eds. Proceedings of the 5th Clinical Natural Language Processing Workshop. Association for Computational Linguistics; 102-107 (2023).
Zolnoori, M., Zolnour, A. & Topaz, M. ADscreen: A speech processing-based screening system for automatic identification of patients with Alzheimer’s disease and related dementia. Artif. Intell. Med. 143, 102624 (2023).
Agbavor, F. & Liang, H. Predicting dementia from spontaneous speech using large language models. PLOS Digit Health 1, e0000168 (2022).
Shakeri, A. & Farmanbar, M. Natural language processing in Alzheimer’s disease research: Systematic review of methods, data, and efficacy. Alzheimer's Dement Diagn. Assess. Dis. Monit. 17, e70082 (2025).
Fristed, E. et al. A remote speech-based AI system to screen for early Alzheimer’s disease via smartphones. Alzheimer's Dement Amst. Neth. 14, e12366 (2022).
Skirrow, C. et al. Storyteller in ADNI4: Application of an early Alzheimer’s disease screening tool using brief, remote, and speech-based testing. Alzheimer's Dement J. Alzheimers Assoc. 20, 7248–7262 (2024).
Balagopalan A., Eyre B., Robin J., Rudzicz F., Novikova J. Comparing Pre-trained and Feature-Based Models for Prediction of Alzheimer’s Disease Based on Speech. Front. Aging Neurosci. 13. https://doi.org/10.3389/fnagi.2021.635945 (2021).
Bayat S., et al. Language abnormalities in Alzheimer’s disease indicate reduced informativeness. Ann. Clin. Transl. Neurol. https://doi.org/10.1002/acn3.52205 (2024).
Williams, E., McAuliffe, M. & Theys, C. Language changes in Alzheimer’s disease: A systematic review of verb processing. Brain Lang. 223, 105041 (2021).
Ahmed, S., de Jager, C. A., Haigh, A. M. & Garrard, P. Semantic processing in connected speech at a uniformly early stage of autopsy-confirmed Alzheimer’s disease. Neuropsychology 27, 79–85 (2013).
Kavé, G. & Goral, M. Word retrieval in picture descriptions produced by individuals with Alzheimer’s disease. J. Clin. Exp. Neuropsychol. 38, 958–966 (2016).
Mohammad Panahi M., et al. Impaired language in Alzheimer’s disease: comparing English and Persian implicates word frequency rather than noun-verb distinction. Aphasiology. 0(0):1–19.
Balagopalan A., Eyre B., Rudzicz F. & Novikova J. To BERT or Not To BERT: Comparing Speech and Language-based Approaches for Alzheimer’s Disease Detection. Interspeech https://doi.org/10.21437/interspeech.2020-2557 (2020).
Croisile, B. et al. Comparative study of oral and written picture description in patients with Alzheimer’s disease. Brain Lang. 53, 1–19 (1996).
Yamada, Y., Shinkawa, K., Nemoto, M., Nemoto, K. & Arai, T. A mobile application using automatic speech analysis for classifying Alzheimer’s disease and mild cognitive impairment. Comput. Speech Lang. 81, 101514 (2023).
Bastings J. & Filippova K. The elephant in the interpretability room: Why use attention as explanation when we have saliency methods? ACL Anthology, https://doi.org/10.18653/v1/2020.blackboxnlp-1.14 (2020).
Apostolova, L. G. et al. The longitudinal early-onset Alzheimer’s disease study (LEADS): Framework and methodology. Alzheimer's Dement J. Alzheimers Assoc. 17, 2043–2055 (2021).
Touroutoglou, A. et al. The sporadic early-onset Alzheimer’s disease signature of atrophy: preliminary findings from the longitudinal early-onset Alzheimer’s Disease Study (LEADS) Cohort. Alzheimer's Dement 19, S74–S88 (2023).
Cho, H. et al. Amyloid and tau-PET in early-onset AD: Baseline data from the Longitudinal Early-onset Alzheimer’s Disease Study (LEADS). Alzheimers Dement 19, S98–S114 (2023).
Fleisher, A. S. et al. Positron Emission Tomography Imaging With [18F]flortaucipir and postmortem assessment of Alzheimer disease neuropathologic changes. JAMA Neurol. 77, 829–839 (2020).
Sabri, O. et al. Florbetaben PET imaging to detect amyloid beta plaques in Alzheimer’s disease: phase 3 study. Alzheimer's Dement J. Alzheimers Assoc. 11, 964–974 (2015).
Dage, J. L. et al. Cerebrospinal fluid biomarkers in the Longitudinal Early-onset Alzheimer’s Disease Study. Alzheimer's Dement 19, S115–S125 (2023).
Nudelman, K. N. H. et al. Pathogenic variants in the Longitudinal Early-onset Alzheimer’s Disease Study cohort. Alzheimer's Dement 19, S64–S73 (2023).
Craft, S. et al. Memory improvement following induced hyperinsulinemia in Alzheimer’s disease. Neurobiol. Aging 17, 123–130 (1996).
Hunt K. W. Grammatical structures written at three grade levels. In: 1965. Accessed January 10, https://www.semanticscholar.org/paper/Grammatical-structures-written-at-three-grade-Hunt/0a86dda1fc0ca4a69332c0e8ef9834d03e7d4587 (2023).
Liu Y., et al. RoBERTa: A Robustly Optimized BERT Pretraining Approach. arXiv. (arXiv:1907.11692). https://doi.org/10.48550/arXiv.1907.11692 (2019).
Merhbene G., et al. Detecting Primary Progressive Aphasia (PPA) from Text: A Benchmarking Study. bioRxiv. Published online: 2025-02. (2025).
Feinberg D. R. Parselmouth Praat Scripts in Python. Published online November 18, https://doi.org/10.17605/OSF.IO/6DWR3 (2018).
Josephy-Hernandez, S. et al. Automated analysis of written language in the three variants of primary progressive aphasia. Brain Commun. 5, fcad202 (2023).
Rezaii, N., Ren, B., Quimby, M., Hochberg, D. & Dickerson, B. C. Less is more in language production: an information-theoretic analysis of agrammatism in primary progressive aphasia. Brain Commun. 5, fcad136 (2023).
Qi P., Zhang Y., Zhang Y., Bolton J., Manning C. D. Stanza: A Python Natural Language Processing Toolkit for Many Human Languages. In: Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics: System Demonstrations. Association for Computational Linguistics; 101–108 (2020).
Devlin J., Chang M. W., Lee K., Toutanova K. BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding. In: Proceedings of the 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies, Volume 1 (Long and Short Papers). Association for Computational Linguistics; 4171-4186 (2019).
Chen T., Guestrin C. XGBoost: A Scalable Tree Boosting System. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining. KDD ’16. Association for Computing Machinery; 785-794. (2016)
Cheng X. A comprehensive study of feature selection techniques in machine learning models. ResearchGate. https://doi.org/10.70088/xpf2b276.
Krawczyk, B. Learning from imbalanced data: open challenges and future directions. Prog. Artif. Intell. 5, 221–232 (2016).
Lundberg S. M., Lee S. I. A Unified Approach to Interpreting Model Predictions. In: Advances in Neural Information Processing Systems. Vol 30. Curran Associates, Inc.; 2017. Accessed November 1, https://papers.nips.cc/paper_files/paper/2017/hash/8a20a8621978632d76c43dfd28b67767-Abstract.html (2023).
Ribeiro M. T., Singh S., Guestrin C. “Why Should I Trust You?”: Explaining the Predictions of Any Classifier. arXiv. 2016;(arXiv:1602.04938). Accessed February 24, http://arxiv.org/abs/1602.04938 (2023).
Acknowledgements
This study was generously supported by Alzheimer’s Association AARG-22-926940, R56 AG057195, U01 AG6057195, U24 AG021886, Alzheimer’s Association LEADS GENETICS-19-639372, U01 AG016976, P30 AG010133, P50 AG008702, P50 AG025688, P50 AG005146, P30 AG062421, P30 AG062422, P50 AG023501, P30 AG010124, P30 AG013854, P50 AG005681, P30AG066506, and P50AG047366 as well as R01 NS131395, R21 DC019567, R21 AG073744, P50 AG005134, and P30 AG062421 and by the Tommy Rickles Chair in Primary Progressive Aphasia Research.
Author information
Authors and Affiliations
Consortia
Contributions
N.R., B.W., and B.C.D. conceptualized the work. NR developed the codes, conducted computational analysis, and wrote the first draft of the paper. N.R., B.W., P.A., L.B., J.L.D., A.E., T.F., K.W., M.C.C., J.H.K., C.R.J., R.K., W.A.K., K.N., A.J.P., R.R., M.R., A.T., M.T., A.T., P.V., A.A., D.C., G.S.D., R.D., N.R.G., I.G., L.S.H., E.C.B.J., D.T.J., J.C.M., M.F.M., E.M., C.U.O., M.R., E.R., S.S., S.S., R.S.T., T.S.W., D.A.W., D.H., K.K., G.D.R., L.G.A., and B.C.D. contributed to writing and editing the manuscript. All authors read and approved the manuscript’s final version.
Corresponding authors
Ethics declarations
Competing interests
Bradford Dickerson is an Associate Editor for npj Dementia. Bradford Dickerson was not involved in the journal’s review of, or decisions related to, this manuscript. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rezaii, N., Wong, B., Aisen, P. et al. Voiceprints of cognitive impairment: analyzing digital voice for early detection of Alzheimer’s and related dementias. npj Dement. 1, 35 (2025). https://doi.org/10.1038/s44400-025-00040-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44400-025-00040-0