Abstract
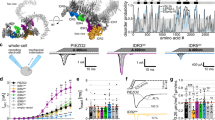
Mechanosensitive ion channels translate physical forces into biochemical signals, enabling cells to sense pressure, tension and tissue deformation. Piezo1, a trimeric mechanogated ion channel, is essential in processes such as vascular regulation and immune function, yet the force required for its activation has remained unclear because existing approaches couple applied force with membrane deformation, preventing direct calibration. Here we present a DNA-tethered extracellular force sensing platform that delivers calibrated piconewton forces directly to Piezo1 while simultaneously monitoring channel activity via a genetically encoded calcium reporter. Combining micropipette manipulation and DNA hairpin-based force calibration, we show that Piezo1 opens approximately at 15.0 pN, providing a direct quantification of its activation threshold and demonstrating that Piezo1 can be gated by tether-mediated forces independent of membrane tension, supporting a force from filament mechanism. This approach offers a generalizable strategy for precisely probing mechanotransduction pathways at the localized molecular sites and can be extended to diverse force responsive systems across biology and materials science.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are present in this Article and its Supplementary Information. The full image dataset with a large size is available from the corresponding author upon request. Source data are provided with this paper.
References
Wu, J., Lewis, A. H. & Grandl, J. Touch, tension, and transduction—the function and regulation of Piezoion channels. Trends Biochem. Sci. 42, 57–71 (2017).
Wang, S. et al. Endothelial cation channel PIEZO1 controls blood pressure by mediating flow-induced ATP release. J. Clin. Invest. 126, 4527–4536 (2016).
Lim, X. R. et al. Endothelial Piezo1 channel mediates mechano-feedback control of brain blood flow. Nat. Commun. 15, 8686 (2024).
Liu, C. S. C. et al. Piezo1 mechanosensing regulates integrin-dependent chemotactic migration in human T cells. eLife 12, RP91903 (2024).
Yang, Z. et al. Nano-mechanical immunoengineering: nanoparticle elasticity reprograms tumor-associated macrophages via Piezo1. ACS Nano 18, 21221–21235 (2024).
Zarychanski, R. et al. Mutations in the mechanotransduction protein PIEZO1 are associated with hereditary xerocytosis. Blood 120, 1908–1915 (2012).
Alper, S. L. Genetic diseases of PIEZO1 and PIEZO2 dysfunction. Curr. Top. Membr. 79, 97–134 (2017).
Zhao, Q. et al. Structure and mechanogating mechanism of the Piezo1 channel. Nature 554, 487–492 (2018).
Saotome, K. et al. Structure of the mechanically activated ion channel Piezo1. Nature 554, 481–486 (2018).
Ge, J. et al. Architecture of the mammalian mechanosensitive Piezo1 channel. Nature 527, 64–69 (2015).
Kefauver, J. M., Ward, A. B. & Patapoutian, A. Discoveries in structure and physiology of mechanically activated ion channels. Nature 587, 567–576 (2020).
Lin, Y. C. et al. Force-induced conformational changes in PIEZO1. Nature 573, 230–234 (2019).
Lewis, A. H. & Grandl, J. Mechanical sensitivity of Piezo1 ion channels can be tuned by cellular membrane tension. eLife 4, e12088 (2015).
Yang, S. et al. Membrane curvature governs the distribution of Piezo1 in live cells. Nat. Commun. 13, 7467 (2022).
Yang, X. et al. Structure deformation and curvature sensing of PIEZO1 in lipid membranes. Nature 604, 377–383 (2022).
Luchtefeld, I. et al. Dissecting cell membrane tension dynamics and its effect on Piezo1-mediated cellular mechanosensitivity using force-controlled nanopipettes. Nat. Methods 21, 1063–1073 (2024).
Mulhall, E. M. et al. Direct observation of the conformational states of PIEZO1. Nature 620, 1117–1125 (2023).
Wang, J. et al. Tethering Piezo channels to the actin cytoskeleton for mechanogating via the cadherin–beta-catenin mechanotransduction complex. Cell Rep. 38, 110342 (2022).
Bustamante, C., Chemla, Y. R., Forde, N. R. & Izhaky, D. Mechanical processes in biochemistry. Annu. Rev. Biochem. 73, 705–748 (2004).
Evans, E. A. & Calderwood, D. A. Forces and bond dynamics in cell adhesion. Science 316, 1148–1153 (2007).
Murthy, S. E., Dubin, A. E. & Patapoutian, A. Piezos thrive under pressure: mechanically activated ion channels in health and disease. Nat. Rev. Mol. Cell Biol. 18, 771–783 (2017).
Leckband, D. E. & de Rooij, J. Cadherin adhesion and mechanotransduction. Annu. Rev. Cell Dev. Biol. 30, 291–315 (2014).
Wang, J. et al. PIEZO1-mediated calcium signaling reinforces mechanical properties of hair follicle stem cells to promote quiescence. Sci. Adv. 11, eadt2771 (2025).
Coste, B. et al. Piezo proteins are pore-forming subunits of mechanically activated channels. Nature 483, 176–181 (2012).
Lewis, A. H. & Grandl, J. Stretch and poke stimulation for characterizing mechanically activated ion channels. Methods Enzymol. 654, 225–253 (2021).
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010).
Wang, X. et al. Mechanical stability of the cell nucleus—roles played by the cytoskeleton in nuclear deformation and strain recovery. J. Cell Sci. 131, jcs209627 (2018).
Ye, J. et al. Ultrasonic control of neural activity through activation of the mechanosensitive channel MscL. Nano Lett. 18, 4148–4155 (2018).
Peralta, F. A. et al. Optical control of PIEZO1 channels. Nat. Commun. 14, 1269 (2023).
Dumitru, A. C. et al. Probing PIEZO1 localization upon activation using high-resolution atomic force and confocal microscopy. Nano Lett. 21, 4950–4958 (2021).
Shin, W. et al. Magnetogenetics with Piezo1 mechanosensitive ion channel for CRISPR gene editing. Nano Lett. 22, 7415–7422 (2022).
Lee, J. U. et al. Non-contact long-range magnetic stimulation of mechanosensitive ion channels in freely moving animals. Nat. Mater. 20, 1029–1036 (2021).
Wu, J., Goyal, R. & Grandl, J. Localized force application reveals mechanically sensitive domains of Piezo1. Nat. Commun. 7, 12939 (2016).
Basu, A., Lagier, S., Vologodskaia, M., Fabella, B. A. & Hudspeth, A. J. Direct mechanical stimulation oftip links in hair cells through DNA tethers. eLife 5, e16041 (2016).
MacHattie, L. A. & Thomas, C. A. DNA from bacteriophage lambda: molecular length and conformation. Science 144, 1142–1144 (1964).
Yaganoglu, S. et al. Highly specific and non-invasive imaging of Piezo1-dependent activity across scales using GenEPi. Nat. Commun. 14, 4352 (2023).
Gudimchuk, N. B. et al. Mechanisms of microtubule dynamics and force generation examined with computational modeling and electron cryotomography. Nat. Commun. 11, 3765 (2020).
Wiegand, T. et al. Forces during cellular uptake of viruses and nanoparticles at the ventral side. Nat. Commun. 11, 32 (2020).
Huang, W., Fu, C. & Yan, J. Single-cell quantification of the mechanical stability of cell-cell adherens junction using glass micropipettes. Methods Mol. Biol. 2600, 267–280 (2023).
Huang, W., Fu, C. & Yan, J. in Mechanobiology: Methods and Protocols (ed. Zaidel-Bar, R.) 267–280 (Springer, 2023).
Xiao, B. Mechanisms of mechanotransduction and physiological roles of PIEZO channels. Nat. Rev. Mol. Cell Biol. 25, 886–903 (2024).
del Rio, A. et al. Stretching single talin rod molecules activates vinculin binding. Science 323, 638–641 (2009).
Hoffman, B. D., Grashoff, C. & Schwartz, M. A. Dynamic molecular processes mediate cellular mechanotransduction. Nature 475, 316–323 (2011).
Elosegui-Artola, A. et al. Force triggers YAP nuclear entry by regulating transport across nuclear pores. Cell 171, 1397–1410 (2017).
Yao, M. et al. The mechanical response of talin. Nat. Commun. 7, 11966 (2016).
Zhang, Y. et al. Multi-domain interaction mediated strength-building in human α-actinin dimers unveiled by direct single-molecule quantification. Nat. Commun. 15, 6151 (2024).
Zhou, T. et al. Piezo1/2 mediate mechanotransduction essential for bone formation through concerted activation of NFAT-YAP1-β-catenin. eLife 9, e52779 (2020).
Poole, K., Herget, R., Lapatsina, L., Ngo, H. D. & Lewin, G. R. Tuning Piezo ion channels to detect molecular-scale movements relevant for fine touch. Nat. Commun. 5, 3520 (2014).
Servin-Vences, M. R., Moroni, M., Lewin, G. R. & Poole, K. Direct measurement of TRPV4 and PIEZO1 activity reveals multiple mechanotransduction pathways in chondrocytes. eLife 6, e21074 (2017).
Roeterink, R. M. A., Casadevall, I. S. X., Collins, D. J. & Scott, D. Force versus response: methods for activating and characterizing mechanosensitive ion channels and GPCRs. Adv. Healthc. Mater. 13, 2402167 (2024).
Andreu, I. et al. The force loading rate drives cell mechanosensing through both reinforcement and cytoskeletal softening. Nat. Commun. 12, 4229 (2021).
Hu, Y. et al. DNA-based ForceChrono probes for deciphering single-molecule force dynamics in living cells. Cell 187, 3445–3459.e15 (2024).
Acknowledgements
This work was supported by the National Research Foundation, Prime Minister’s Office, Singapore, under the NRF Investigators Program (award no. NRF-NRFI052019-0003 to X.L.) and NUS NANONASH Programme (NUHSRO/2020/002/NanoNash/LOA; award no. R143000B43114 to X.L.). J.Y. received support from the Singapore Ministry of Education Academic Research Fund Tier 2 (MOE; grant no. MOE-T2EP50123-0008), National Research Foundation, Prime Minister’s Office and the Mid-Sized Grant (grant no. NRF-MSG-2023-0001). We thank J. Grandl (Duke University) for generously providing plasmids encoding wild-type mouse Piezo1 and the bungarotoxin-binding site-inserted constructs.
Author information
Authors and Affiliations
Contributions
X.L., J.Y., M.S. and Y.L. conceived and designed the project. X.L., J.Y. and Y.L. supervised the project, providing conceptual and experimental guidance. M.S. carried our molecular cloning, electrophysiology, imaging and micropipette experiments. J.L. performed λ-DNA modifications and contributed to molecular cloning. C.F. assisted with micropipette experiments. M.S. and Y.L. drafted the original paper. X.L. and J.Y. edited the paper. All authors participated in the discussion and interpretation of the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Sensors thanks Isaac Li, Jisook Moon and Kate Poole for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Effects of ionomycin treatment.
a, GFP fluorescence response to ionomycin treatment. b, Fluorescence intensity changes in HEK293T cells expressing BBS-Piezo1-jGCaMP8m upon addition of ionomycin. c, Quantification of relative fluorescence before and after ionomycin treatment (n = 24 cells from three independent tests). Statistical analysis was performed using a two-tailed paired t-test (P < 0.0001), data are presented as mean ± s.e.m.
Extended Data Fig. 2 Micropipette-enabled mechanical perturbation of GsMTx4-treated cells.
a, Representative images of HEK293T cells expressing BBS-Piezo1-jGCaMP8m during direct micropipette perturbation after GsMTx4 treatment. The dashed line indicates the position and direction of micropipette movement. b, Representative fluorescence intensity trace of GsMTx4-treated cells under micropipette perturbation. Experimental tests were repeated for three times.
Extended Data Fig. 3 Micropipette-based perturbation with DSPE-beads.
a, Bright-field images illustrating removal of untethered beads by micropipette flow. b, Representative GFP channel images showing fluorescence changes during tethered bead pulling. Experimental tests were repeated for four times.
Extended Data Fig. 4 Schematic and synthesis of the 1000 bp hairpin DNA.
The construct comprises five functional segments: a hairpin stem, a biotinylated region, a loop, a linker, and a thiol-modified end. The synthesis protocol involves four steps: (1) PCR amplification of the core hairpin sequence; (2) restriction enzyme digestion to generate cohesive ends; (3) annealing to form the hairpin structure; and (4) ligation to assemble the complete construct.
Extended Data Fig. 5 Verification of micropipette suction force using magnetic tweezers and DNA hairpin probes.
a, Force-distance calibration curve of the micropipette, with blue and green circles indicating verified force values Fclose and Fopen at corresponding pipette distances. b, Schematic of magnetic tweezer setup used to quantify hairpin closing force and opening forces. c, Bead height changes during force-jump mode (switching between two constant-force levels). d, Bead height changes during force-ramp mode (linearly increasing force). e, Schematic of micropipette suction applying lateral tension to monitor DNA hairpin opening. f, Representative trace of bead displacement versus suction pressure, showing hairpin opening under increasing force.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–12 and Tables 1 and 2.
Supplementary Movie 1 (download MP4 )
Tracking of tethered GCaMP in a transfected cell.
Supplementary Movie 2 (download MP4 )
Force application via grabbing DNA-bead to activate Piezo1.
Supplementary Movie 3 (download MP4 )
Force application via remote suction on DNA-bead to activate Piezo1.
Source data
Source Data Fig. 2 (download XLSX )
Source data and statistics.
Source Data Fig. 3 (download XLSX )
Source data and statistics.
Source Data Fig. 4 (download XLSX )
Source data and statistics.
Source Data Extended Data Fig. 1 (download XLSX )
Source data and statistics.
Source Data Extended Data Fig. 2 (download XLSX )
Source data and statistics.
Source Data Extended Data Fig. 5 (download XLSX )
Source data and statistics.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sui, M., Liu, J., Fu, C. et al. Direct quantification of Piezo1 activation threshold through DNA-tethered extracellular force sensing. Nat. Sens. (2026). https://doi.org/10.1038/s44460-026-00060-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s44460-026-00060-0