Abstract
Inborn errors of tyrosine metabolism result in patient’s inability to degrade tyrosine. Current treatment consists of a phenylalanine and tyrosine restricted diet and nitisinone, causing a block in the tyrosine degradation pathway. However, tyrosine levels will increase, leading to acquired hypertyrosinemia, implying the need for an add-on treatment. Tyrosine ammonia lyases (TAL) can provide such an add-on treatment as they catalyze the deamination of tyrosine into p-coumaric acid and ammonia. In this study, we developed a robust high-throughput screening (HTS) assay to assess the capacity of bacterial TAL enzymes to decrease excessive tyrosine. The assay is based on the spectrophotometric quantification of p-coumaric acid after conversion of tyrosine by bacterial TAL. As a benchmark, TAL from Flavobacterium johnsoniae (FjTAL) was used to optimize the assay. Optimal growth conditions for high-level protein expression were determined by incubating transformed Escherichia coli BL21 (DE3) cells at different temperatures during various incubation times. Subsequently, assay temperature and pH were optimized followed by testing different ratios of tyrosine assay mixes to bacterial lysate. Finally, assay robustness and functionality were evaluated. Optimal FjTAL expression was obtained after incubation for 24 h at 22 °C. Ideal assay conditions consist of a 80/20 ratio of 1 mM tyrosine assay mix to FjTAL lysate performed at pH 9.2 and 37 °C. The robustness test showed Z′ values > 0.4 and signal window values > 2 without edge or drift effects. As proof-of-principle, we successfully determined the catalytic activity of two other bacterial TAL enzymes RsTAL (5.718.10–3 ± 0.21.10–3) and SeSAM8 (4.658.10–3 ± 0.37.10–3). A robust, simple and reliable HTS assay was thus developed to evaluate the tyrosine degradation capacity of bacterial TAL enzymes.
Similar content being viewed by others
Introduction
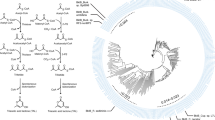
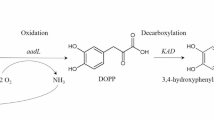
Tyrosine inherited metabolic disorders (TIMD) are a group of inborn errors of amino acid metabolism where a deficiency occurs in one of the enzymes involved in the degradation of tyrosine (Tyr)1,2, a semi-essential amino acid derived from either diet or the hydroxylation of phenylalanine (Phe). Tyr catabolism proceeds predominantly in the liver cytosol. The Phe/Tyr-degradation pathway, shown in Fig. 1, consists of 6 enzymatic steps and results in the formation of fumarate and acetoacetate2. Five different TIMD have been identified at four steps of the degradation pathway and are characterized by the patient’s inability to completely break down Tyr3.
The degradation pathway of tyrosine. At four steps of the pathway, tyrosine inherited metabolic disorders have been identified. Nitisinone, also known as NTBC, a substrate reduction therapy for hereditary tyrosinemia type 1 and alkaptonuria, inhibits the conversion of 4-hydroxyphenylpyruvate into homogentisate and thus decreases the production of the toxic metabolites in hereditary tyrosinemia type 1 and alkaptonuria2,3. However, tyrosine levels increase upon nitisinone administration, leaving patients at risk for ocular, cutaneous and neurological side effects2,3,4.
In this study, we developed a high-throughput screening (HTS) assay that is of particular interest for four of the TIMD, namely hereditary tyrosinemia type 1 (HT1), alkaptonuria (AKU), hereditary tyrosinemia type 2 (HT2) and hereditary tyrosinemia type 3 (HT3). HT1 and AKU are caused by a deficiency of respectively fumarylacetoacetate hydrolase and 4-hydroxyphenylpyruvate dioxygenase (HPD) and are currently treated by dietary restriction of Phe and Tyr intake with the administration of a daily dose of 2-(2-nitro-4-trifluoro-methylbenzoyl)-1,3-cyclohexane-dione (NTBC)2,3,4,5. Despite clinical improvement, NTBC increases blood Tyr levels up to 800–1000 µM resulting in acquired hypertyrosinemia leading to ocular, cutaneous and possible neurological complications2,3,4,6,7. A diet low in Phe and Tyr could ameliorate the complications associated with NTBC-related acquired hypertyrosinemia, however, dietary restrictions are very demanding and developmental delay as well as impaired cognitive functions are still observed in some patients treated with both NTBC and a protein-restricted diet8. Next to HT1 and AKU, HT2 is caused by a defect of the tyrosine aminotransferase (TAT) enzyme, resulting in markedly elevated Tyr concentrations in serum and cerebrospinal fluid with clinical manifestations including eye and skin lesions and highly variable neurological complications2,5,9. HT3 on the other hand is caused by a deficiency of the HPD enzyme. Due to this enzyme block, plasma Tyr concentrations increase2,5,9. Patients with HT3 present with neurological symptoms including ataxia, intellectual impairment, tremors, increased tendon reflexes, microcephaly and seizures2.
In the context of hypertyrosinemia in TIMD, aromatic amino acid ammonia lyase enzymes, and more specifically tyrosine ammonia lyase (TAL), can provide a therapy to decrease excessive Tyr levels. Aromatic amino acid ammonia lyases, including histidine ammonia lyase (HAL), TAL and phenylalanine ammonia lyase (PAL), catalyze the non-oxidative deamination of L-amino acids by reversibly cleaving a carbon–nitrogen bond releasing ammonia which can be rapidly metabolized via the urea cycle. Corresponding aryl acids are formed during this process10,11.
Similar to the treatment of phenylketonuria with Pegvaliase, a bacterial PAL enzyme12, bacterial TAL enzymes can provide an alternative degradation route, independent of the Tyr degradation pathway shown in Fig. 1, for the excessive Tyr levels in TIMD. TAL enzymes could thus possibly reduce Tyr levels to within the normal range by directly converting Tyr into p-coumaric acid (pCA) and ammonia13. In addition, the formed metabolite pCA has strong anti-oxidant and protective properties against reactive oxygen species induced apoptosis13,14. Furthermore, it is able to reduce lipid peroxidation in hepatic steatosis and shows protective effects against acute liver and kidney damage15. Next to a decrease of the Tyr levels, these beneficial properties of pCA can provide a desirable supplementary effect of a TAL-based therapeutic approach in patients suffering from TIMD. TAL enzymes do not exist in mammals16, however, they can be found in bacteria such as Flavobacterium johnsoniae, Rhodobacter capsulatus and Rhodobacter sphaeroides17,18. In analogy to treatment with Pegvaliase, immunogenic reactions and side effects of the same order of magnitude, ranging from local injection site reactions to acute systemic hypersensitivity reactions, could thus be expected for the here proposed TAL-based therapy19. Besides their possible therapeutic application, TAL variants are used as biocatalyst for the production of aromatic compounds including pCA, stilbenes, plastic precursors and flavonoids16,20.
The goal of this study is to develop a HTS assay that allows to evaluate the catalytic activity of bacterial TAL enzymes in the context of hypertyrosinemia in TIMD. To achieve this, we chose the bacterial host organism Escherichia coli (E. coli) BL21 (DE3) as expression system. Next to rapid growth under inexpensive culturing conditions, E. coli cells are easy to manipulate and transform with exogenous DNA plasmids after which they allow fast expression and proper folding of the recombinant enzymes21,22,23. The capacity to decrease Tyr of the recombinant TAL enzymes, expressed in E. coli cells, can then be analyzed in the here developed HTS assay by spectrophotometrically following the overtime production of pCA.
TAL screening assays focusing on TALs originating from fruit juices, animal tissues and plants are already commercially available. Both available screening systems are based on the colorimetric detection of TAL activity by performing an endpoint measurement after 45 min of reaction time24,25. Yet, we intended to develop a HTS screening system especially for TAL enzymes from bacterial origin based on overtime absorption measurement of the reaction product pCA during 30 min. Besides, we aimed to obtain more versatile and easy to adjust assay parameters such as assay pH, necessary for the different pH optima of various TAL enzymes as well as for further downstream therapeutic approaches, while allowing overtime spectrophotometric measurement of pCA production to gain better insights in TAL characteristics compared to endpoint measurement of these commercially available TAL screening systems13,24,25.
In addition to the commercially available assays, various analysis systems have been reported in scientific literature to characterize TAL enzymes and assess their activity. Analysis systems developed by Jendresen et al., Virklund et al. and Brack et al. all required protein purification13,26,27. Moreover, Jendresen et al. analyzed TAL activity both by quantification of produced pCA through HPLC analysis as well as by spectrophotometrically measuring the increase in absorption at 310 nm at 30 °C with a Tyr concentration of 1 mM13. Virklund et al. quantified TAL activity by adding 1 mM Tyr to 1 µg of the purified TAL enzyme and measuring the absorbance spectrum between 220 and 700 nm during 1 h. Production rate of pCA was determined by calculating the slope of absorbance overtime at three different wavelengths, resulting in a more complex assessment of TAL activity26. Furthermore, Brack et al. characterized TAL pH and temperature optima by spectrophotometrically measuring pCA production at 310 nm during 10 min using 2 mM Tyr. They demonstrated increased TAL activity at temperatures above 40 °C with optimal TAL activity at 65 °C. In depth characterization of biotransformation capability of TAL was performed through HPLC analysis using the purified enzyme with 30 mM Tyr at 65 °C and pH 9.527. The here developed TAL activity screening simplifies the analysis of TAL enzymes compared to the already established analysis systems. Although protein purification is required for in depth characterization of TAL enzymes, this step significantly reduces the high-throughput application of these analysis systems, a drawback that we overcome with the here developed HTS assay based on the crude cell lysate. Additionally, spectrophotometric measurement of pCA production at one single wavelength, determining biotransformation capability of TAL enzymes, is a more simple and cost-effective analysis method compared to the HPLC analysis performed in the methods developed by Jendresen et al. and Brack et al. as well as compared to the more complex analysis method of Virklund et al. where pCA production is calculated based on three different wavelengths13,26,27. Besides, we aimed to obtain more versatile and easy to adjust assay parameters such as assay pH, necessary for the different pH optima of various TAL enzymes and their implications in various future downstream therapeutic approaches. Thus, although TAL activity seems to increase at temperatures above 40 °C with an optimum around 65 °C26,27, our assay was specifically developed to assess TAL activity at physiological temperature to allow analysis under conditions necessary for further downstream therapeutic applications of TAL enzymes whereas the focus in previous analysis of TAL enzymes has been on either their characterization or their industrial applications.
Materials and methods
Materials
All chemicals were purchased from Merck KGaA (Darmstadt, Germany) and were of analytical grade or higher. Lysogeny Broth (LB) medium with or without 1% glucose (LBG) supplement was used as standard growth medium. Growth medium was supplemented with 50 µg/ml kanamycin (KANA) for bacterial selection. OD600 was measured on an Eppendorf Biophotometer D30 (Hamburg, Germany). TAL proteins were expressed in 96-well deep well conical bottom MTP plates from Greiner Bio-one GmbH (Kremsmünster, Austria) by incubating in an Eppendorf New Brunswick™ Innova 42 R incubator shaker series (Hamburg, Germany). pCA absorbance measurements were performed on a Fluostar optima spectrophotometer (BMG Labtech, Ortenberg, Germany). Plasmid isolation kits were purchased from Merck KGaA (Darmstadt, Germany).
Bacterial strains, plasmids and target gene
Assay optimization was performed with the TAL enzyme derived from Flavobacterium johnsoniae (FjTAL) whereas assay functionality was assessed with two additional TAL enzymes from Rhodobacter sphaeroides (RsTAL) and Saccharothrix espanaensis (SeSAM8). cDNA of the three bacterial TAL enzymes was obtained from Geneart in standard pMA-T-based cloning plasmids (Thermo Scientific, Chicago, IL, USA). Codon optimized cDNA sequences for all bacterial TAL enzymes can be found in the Supplementary Information in Supplementary Table 1. To obtain high-level protein expression in E. coli, the bacterial TAL cDNA was subcloned into a pET42b( +) plasmid from Novagen (Darmstadt, Germany), containing a gene for KANA resistance, by using NdeI and EcoRI as restriction enzymes for ligase-based cloning in the multiple cloning site of the pET42b( +) vector. The pET42b( +) plasmid without bacterial TAL cDNA, called the empty vector (EV) plasmid, was used as a negative control. E. coli DH5α cells (Stratagene, La Jolla, CA, USA) were used for plasmid maintenance and propagation. For protein expression, E. coli BL21 (DE3) cells (Agilent Technologies, Santa Clara, CA, USA) were used. Both strains were transformed by performing heat shock.
Optimal expression conditions
LBGKANA medium was inoculated with the E. coli BL21 (DE3) cells containing the pET42b( +)-FjTAL plasmid followed by overnight incubation in a shaking incubator at 37 °C and 250 rpm. To allow protein expression, overnight cultures were diluted in LBKANA medium to an OD600 of 0.1 and incubated at 30 °C and 250 rpm. Once an OD600 between 0.6 and 0.8 was reached, overexpression of the FjTAL protein was induced by adding 1 mM isopropyl-β-D-1-thiogalactopyranoside (IPTG) to the cultures after which they were incubated under different expression temperatures (16 °C, 22 °C, 30 °C and 37 °C). At various time points (0 h, 2 h, 4 h, 6 h, 24 h and 48 h), samples for protein extraction were taken from independently cultured flasks in such a way that three biological replicates for each condition were obtained. Bacterial cell concentration was normalized between samples by measuring the OD600 followed by centrifugation for 5 min at 3000 g and 4 °C to acquire a bacterial cell pellet. Cell pellets were subsequently washed in cool and sterile HEPES buffer (20 mM, pH 7.0) and thereafter centrifuged for 2 min at 2350 g and 4 °C. After discarding the supernatant, the pellet fractions were stored at -20 °C until analysis of protein expression.
Protein extraction and SDS-PAGE
Frozen pellets were thawed and prepared for sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) to analyze protein expression. Thawed pellets were resuspended in 485 µl HEPES buffer (20 mM, pH 7.0), cooled on ice and sonicated at an amplitude of 40% during 3 times 10 s with an interval of 30 s to disrupt the E. coli cells. Thereafter, samples were centrifuged for 10 min at maximum speed and 4 °C after which pellet and supernatant fractions were separated. Supernatant fractions, containing the soluble FjTAL protein, were subsequently analyzed by SDS-PAGE to determine optimal conditions for protein expression. In summary, proteins were separated by loading the supernatant samples on a 12% Mini-Protean® TGX Stain-free™ gel (Bio-Rad, Hercules, CA, USA) and subjecting them to electrophoresis for 1 h at 150 V. Gels were then visualized using a ChemiDoc MP imaging system (Bio-Rad, Hercules, CA, USA) and FjTAL expression was quantified using Bio-Rad ImageLab software (v.6.1) after normalization of the sample pixel density data by subtracting the average pixel density of three background signals.
Spectrophotometric HTS assay and ideal Tyr concentration
E. coli BL21 (DE3) pET42b( +)-FjTAL cells were cultured and FjTAL expression was induced as described above. Post-IPTG induction, 600 µl of the LBKANA culture, containing E. coli BL21 (DE3) pET42b( +)-FjTAL cells, was transferred to a 96-well deep well conical bottom MTP plate and incubated for 24 h at 22 °C and 400 rpm. Afterwards, 200 µl of each well was transferred into a 96-well conical bottom MTP plate followed by centrifugation for 10 min at 3200 g and 4 °C. Supernatant was discarded and the plate containing the bacterial cell pellets was frozen at -20 °C for at least 1 h to allow ice crystal formation and ensure proper cell lysis. Plates were then thawed and cell pellets were lysed in a ThermoMixer® C (Eppendorf, Hamburg, Germany), using 10 µl CelLytic B Cell Lysis reagent per mg cell pellet, during 1 h at 4 °C and 400 rpm. Thereafter, plates were centrifuged for 20 min at 3250 g and 4 °C. Supernatant containing FjTAL was transferred to a 96-well flat bottom MTP plate to which assay mix, consisting of Tyr dissolved in a 50 mM Tris HCl buffer, was added after which the absorption was measured at 310 nm during 30 min to follow the overtime production of pCA. To determine ideal assay conditions, the assay was performed at both 25 °C and 37 °C as well as at pH 7.4 and 9.2. Besides, the assay mix contained various concentrations Tyr between 0.125 mM and 1.5 mM and different ratios of cell lysate to assay mix were tested (5/95, 10/90, 15/85, 20/80 and 25/75). Experiments were performed in such a way to obtain three biological repeats where each repeat consisted of four technical replicates. E. coli BL21 (DE3) pET42b( +)-EV cells were used as negative control as well as to correct for possible background signals. The assay set up is visualized in Fig. 2.
Methodology of the optimized HTS assay. E. coli BL21 (DE3) pET42b( +)-FjTAL cells were cultured and FjTAL expression was induced followed by incubation under the conditions that yield the highest FjTAL concentration. Afterwards, bacteria were lysed and the bacterial cell lysate, containing the FjTAL enzyme, was used to perform the HTS assay.
Assay robustness
Assay robustness experiments were performed on three consecutive days to test plate uniformity. Plates were filled in an interleaved signal format consisting of a high (H; 20 µl FjTAL cell lysate), middle (M; 10 µl FjTAL cell lysate) and low (L; 20 µl EV cell lysate) signal. Each day, three plates with a different plate layout (plate 1: HML, plate 2: MLH and plate 3: LHM) were run while plate formats were unaltered between days. The 96-well MTP plates were filled with 200 µl bacterial culture according to the different plate layouts followed by protein expression in all wells. Absorbance measurements for the high and low signal were performed using a 20/80 ratio of cell lysate to assay mix containing 1 mM Tyr. For the middle signal, absorbance measurements were performed by adding 10 µl FjTAL cell lysate to 90 µl assay mix containing 1 mM Tyr. To assess the robustness, mean, standard deviation (SD), coefficient of variation (CV), middle signal percent activity (Mid%), Z’ factor and signal window (SW) of the assay were calculated. Mid%, Z’ factor and SW were calculated using the following Equations28:
where n is the number of replicates used for the HTS assay.
Assay functionality
Functionality of the HTS assay was tested by performing the assay with two additional bacterial TAL enzymes: RsTAL and SeSAM8. Protein expression and absorbance measurements were performed as described above under optimized assay conditions.
Results
Optimal expression conditions
Optimal conditions for FjTAL expression using E. coli BL21 (DE3) cells as host organism were determined by incubating E. coli BL21 (DE3) pET42b( +)-FjTAL cells post-IPTG at different temperatures during various time periods as described in the materials and methods section. Normalization between samples was done by performing OD600 measurements followed by bacterial cell lysis and the evaluation of FjTAL production in the supernatant fraction by SDS-PAGE. At circa 57 kDa, a band for FjTAL can be detected, correlating to its molecular weight29. As observed in Fig. 3a, FjTAL expression could be clearly observed 2 h post-induction at 16 °C, 22 °C and 30 °C whilst almost no expression is detected in the supernatant fraction at 37 °C. Although FjTAL expression at 37 °C is stable over time, increasing expression levels at 16 °C, 22 °C and 30 °C were observed 24 h after IPTG addition where the expression is the highest at 22 °C (Fig. 3b). After 48 h, expression levels decreased and therefore ideal expression conditions to yield the highest FjTAL production were set at incubation for 24 h at 22 °C post-induction.
(a) Analysis of the supernatant fraction of the bacterial cell lysate, containing FjTAL, by SDS-PAGE. E. coli BL21 (DE3) pET42b( +)-FjTAL cultures were incubated at four different temperatures (16 °C, 22 °C, 30 °C and 37 °C) post-induction of FjTAL expression followed by sampling after 0 h, 2 h, 4 h, 6 h, 24 h and 48 h. FjTAL expression could be detected 2 h post IPTG addition at 16 °C, 22 °C and 30 °C. Almost no expression was observed after incubation at 37 °C. While the expression of FjTAL was stable at 37 °C, increased levels of protein expression were observed at the three other temperatures. Uncropped gels are available as Supplementary Information in Supplementary Fig. 1. (b) Analysis of the detected FjTAL bands on the SDS-PAGE gels. Normalized pixel intensity of the FjTAL band was calculated for each condition. After comparison, incubation for 24 h at 22 °C post induction of protein expression were chosen as ideal expression conditions.
Spectrophotometric HTS assay and ideal Tyr concentration
After establishing the optimal FjTAL expression conditions, assay temperature and pH were optimized by culturing bacteria and inducing FjTAL expression followed by bacterial lysis using CelLytic B Reagent after which the assay was performed as described in the materials and methods section at both 25 °C and 37 °C as well as at pH 7.4 and 9.2. As shown in Fig. 4a, pH has a higher impact on FjTAL activity than assay temperature. Overall, FjTAL was significantly more active at pH 9.2 and 37 °C (P < 0.0001) compared to the three other conditions tested. Once assay pH and temperature were determined, the ratio of cell lysate containing FjTAL to assay mix containing Tyr was optimized by testing five different ratio’s with 12 assay mixes, each with a final Tyr concentration between 0.125 mM and 1.5 mM. This allowed the set up of a Michaelis Menten graph in Fig. 4b and the determination of the Vmax for each ratio in Fig. 4c. As observed, the determined Vmax increased with increasing cell lysate volumes and was the highest at a 25/75 cell lysate to assay mix ratio. However, there was no significant difference between the ratio of 20/80 and 25/75. Besides, no plateau was reached with the tested Tyr concentrations in the assay mix under these conditions, implying that the FjTAL enzyme was not yet saturated using a 25/75 ratio. Therefore, optimal assay conditions consisted of a 20/80 ratio of bacterial cell lysate to assay mix containing 1 mM Tyr as here the highest Vmax was obtained while the FjTAL enzyme was saturated as this allowed for easy comparison of bacterial TAL activity between different variants.
(a) Influence of pH and temperature on bacterial TAL activity. There was a significant difference between both tested conditions at pH 9.2 as well as between both conditions tested at pH 7.4 (P < 0.0001). Furthermore, assay performance was significantly better at 37 °C and pH 9.2 compared to testing at 25 °C at pH 7.4 (P < 0.0001) and 37 °C at pH 7.4 (P < 0.0001). (b) Michaelis–Menten graphs for each ratio of cell lysate containing FjTAL to assay mix containing various concentrations Tyr. (c) Analysis of the Vmax values for each ratio of cell lysate containing FjTAL to assay mix containing Tyr. Based on (b) and (c), a 20/80 ratio of cell lysate to assay mix containing 1 mM Tyr were chosen as ideal assay conditions as here the highest Vmax was obtained while the FjTAL enzyme was saturated.
Assay robustness
To analyze the plate uniformity and signal separation of the screening platform, the assay was performed on three different days where on each day, three plates were measured. Sufficient SW, and thus signal separation, was assessed by performing the assay with a high signal, consisting of the optimized assay signal (20 µl FjTAL cell lysate), a low signal which resembles the background signal of the assay (20 µl EV cell lysate) and a middle signal resembling the activity of a 50% active FjTAL enzyme at pH 9.2 (10 µl FjTAL cell lysate). Plates were filled in such a way that over the three plates, the high, middle, and low signal were measured once at each position of the 96-well plate on a given day. Filling the plates according to this interleaved format furthermore allowed to detect edge or drift effects.
Plate uniformity results according to a 96-well microplate format are shown in Table 1. Based on these data, overall assay quality, signal separation as well as inter-plate and inter-day variations were evaluated. To resemble real-life HTS assay conditions, one single replicate was taken into account per signal to determine CVs, Mid%, Z’ factor and SW. It is generally accepted that all CV values should be ≤ 20% in order to obtain significant separation between the high and low signals28. High signal CVs ranged between 5.1–9.5% where the middle signal CV interval was 7.2–10.9%. However, low signal CVs were between 8.1%—24.1% . Here, the CVs from plate 1, day 1 and plate 2, day 2 exceeded the ≤ 20% criterion. Often, the low signal CV fails to meet this criterion, particularly when its mean is very low as was the case here. Therefore, the alternative criterion that within each plate, the SD of the low signal ≤ both the SD of the middle signal and the SD of the high signal, is accepted as well28. As all low SD ≤ both middle SD and high SD, the assay passed this alternative criterion. Besides CVs, it should be considered that the SD of Mid% ≤ 20 on all plates and either all Z’ factor ≥ 0.4 or all SW ≥ 2. With all Mid% SD ≤ 20%, an average Z′ factor of 0.6411 ± 0.9701.10–1 and average SW of 8.1467 ± 2.9855, the assay passed these criteria. Inter-plate and inter-day fold shifts are represented in Table 2 and were < 2. Therefore, the assay could be considered excellent and robust under the optimized assay conditions.
Furthermore, the spatial uniformity of the assay was assessed. This allowed to detect potential row and column edge or drift effects along with other systematic sources of variability. These are often caused by (i) evaporation during longer incubation periods as well as plate stacking and (ii) short incubation times as the outer wells reach the incubation temperature faster under these conditions. Overall, plates should not exhibit material edge and drift effects > 20%. Effects < 20% or effects only occurring in a single or a few plates, are regarded as insignificant28. Row and column effects of the high and middle signal were evaluated from left-to-right and top-to-bottom. Middle signal drift effects, with a maximum of 17.29%, did not exceed 20% where for the high signal, only one drift effect > 20% was detected on day 2, plate 2 (21.6468%). However, as this drift effect only occurred in one single plate, it does not point to a predominant pattern and was therefore considered insignificant28. Furthermore, data was visualized by plotting against well number where the wells were ordered by row first followed by ordering by column. This way, scatter plots, revealing additional patterns of edge and drift effects, were obtained. Scatter plots are shown in Fig. 5 and no additional edge or drift effects were observed over the three plates of each day.
To reveal possible patterns of edge and drift effects, spatial uniformity of the assay was assessed by visualizing the data through scatter plots. Signals were plotted against well number, first by row, then by column. Calculated edge and drift effects for high and middle signal only exceeded the 20% limit once in a single plate which can be regarded as insignificant.
Assay functionality
Assay functionality was tested by performing the assay under optimized conditions (pH 9.2 and 37 °C) with FjTAL, RsTAL and SeSAM8. Bacterial TAL activity was detected for FjTAL (1.943.10–2 ± 0.30.10–2), RsTAL (5.718.10–3 ± 0.21.10–3) and SeSAM8 (4.658.10–3 ± 0.37.10–3), proving the functionality of the optimized HTS system (Fig. 6).
Results of the HTS assay performed with FjTAL, RsTAL and SeSAM8 to analyze the functionality of the assay. In graph (a), the measured absorption over time is displayed whereas the increase in absorption per minute, correlating to the TAL activity, is visualized in graph (b). As TAL activity was detected for the three investigated enzymes, this proves that the developed screening platform can broadly be applied.
Discussion
Increasing Tyr levels are a result of the mutated TAT and HPD enzyme in respectively HT2 and HT35. Additionally, this is also an inevitable consequence of the administration of the lifesaving drug NTBC in HT1 and AKU patients7. However, to day, these excessive Tyr levels can only be treated by installing a Tyr restricted diet, impacting patients quality of life and resulting in poor dietary compliance30. Therefore, there is a serious need for a new treatment providing an alternative degradation route to control the increased Tyr levels in hypertyrosinemia. Here, we hypothesize that bacterial TAL enzymes, in analogy to the use of a bacterial PAL enzyme for the treatment of phenylketonuria, could decrease these excessive Tyr levels12. In this paper, a HTS assay was developed in E. coli bacteria to assess the fitness of bacterial TAL enzymes to metabolize Tyr by evaluating their catalytic activity.
E. coli allows for easy transformation and manipulation while providing a fast and cheap expression system21,22,23. Solubility and stability problems of the expressed protein might however occur at higher expression temperatures such as 37 °C. Due to the increased translation rate under these conditions, saturation of the protein folding mechanisms occurs which results in incorrect protein folding and the formation of inclusion bodies. These processes are favored at higher expression temperatures and can explain the low expression of FjTAL in the supernatant fraction observed at 37 °C31,32,33. On the other hand, temperatures below 15 °C result in decreased fatty acid saturation as well as inhibition of DNA, RNA and protein synthesis34. Therefore, ideal expression temperatures, using E. coli as expression system, lie between 15 °C and 25 °C as here the solubility and stability of the recombinant protein are improved while protein degradation and aggregation into inclusion bodies is diminished23,33,34,35. As E. coli cells tend to grow optimal at higher temperatures, in this HTS assay, precultures were grown at 37 °C followed by a temperature switch to 22 °C to allow ideal expression of a soluble and stable recombinant FjTAL protein36. Furthermore, the more optimal assay temperature of 37 °C compared to 25 °C corresponds to the study results of Virklund et al. where they observed higher FjTAL activity with increasing temperatures between 30 °C and 45 °C. This could be explained by the increased solubility of Tyr at higher temperatures and the accompanying decrease in product inhibition26. Indeed, Brack et al. showed even more increased TAL activity at temperatures above 40 °C and ideally at 50 °C–65 °C, however, the here optimized assay temperature is justified as measurement at 37 °C allows analysis of TAL enzymes with regard to their possible therapeutic implications in the context of TIMD27. Additionally, the here developed screenings system is highly relevant for further downstream therapeutic applications of TAL enzymes as the optimized Tyr concentration of 1 mM perfectly correlates to the Tyr-levels in HT1 and AKU patients treated with NTBC6.
The bacterial TAL enzymes evaluated in our developed bioassay all showed TAL activity at pH 9.2 thus proving the assay’s applicability. However, FjTAL proved to be three to four times more active compared to RsTAL and SeSAM8. This difference in TAL activity for the tested enzymes in the HTS assay can be explained by a difference in their pH optima. Where the pH optimum of FjTAL is around 9.2, RsTAL and SeSAM8 tend to be more active at higher pH levels of respectively 10 and 10.513. This higher activity for FjTAL compared to other bacterial TAL enzymes confirms the results of two previous studies performed by Jendresen et al. and Virklund et al.13,26. Besides, the estimated Km value of 33.98 ± 1.70 µM for FjTAL under optimized assay conditions correlates to the Km value of 33 ± 5 µM found in the study of Virklund et al.26. Next to its higher activity, scientific literature shows that FjTAL has a stronger substrate specificity compared to the two other bacterial TAL enzymes included in this study. With nearly no affinity for Phe, FjTAL shows almost no PAL activity13. This high TAL activity in combination with its specific substrate affinity makes FjTAL an ideal candidate for future exploration to combat the excessive Tyr levels in TIMD patients suffering from hypertyrosinemia.
The robustness and reliability of the developed HTS system was validated according to the NCGC guidelines28. With a Z′ factor, a characteristic parameter for the quality of the assay itself and preferred for the evaluation of HTS systems37,38, ≥ 0.4 and no edge or drift effects under the manually optimized conditions, the assay fulfills the acceptance criteria for a robust and reliable screening assay. In addition, the assay is performed in a 96-well microplate format using only one replicate, allowing for the high-throughput screening of bacterial TAL enzymes. Compared to the methods used by Jendresen et al., Virklund et al. and Brack et al., the here developed assay is based on the crude cell lysate without the need for protein extraction or clean up, thus reducing assay time and costs13,26,27. Additionally, in contrast to Jendresen et al. and Virklund et al., absorbance measurement of the produced pCA is only performed at one wavelength and the catalytic parameters of the TAL enzymes can be determined over a shorter period of time as here pCA production is followed over 30 min13,26. Besides, the here developed HTS system has several advantages over the commercially available TAL assay kits. Firstly, the commercially available TAL assay kits are developed to detect TAL activity in fruit juices, plants and animal tissues whereas our assay was optimized specifically for bacterial TAL enzymes24,25. Secondly, in contrast to its commercially available variants where TAL activity is assessed based on an endpoint measurement after 45 min, our HTS system measures the production of pCA during 30 min, allowing to assess the activity of the tested TAL enzymes overtime24. On top of that, assay parameters such as the assay pH can be adjusted according to the pH optimum of various TAL enzymes whilst in the commercially available TAL assay kits, all parameters are set24,25. Additionally, adjustment of the assay pH allows to directly evaluate the activity of TAL enzymes at the pH level necessary for various therapeutic administration routes, pointing once again to the broad applicability of the here developed screening system. Therefore, the here developed simple, fast, robust, reliable and cost-effective HTS system allows for a better and more complete estimation of the fitness of various bacterial TAL enzymes, including their catalytic activity, for further downstream therapeutic applications whereas previously developed assays had been focusing on, and where thus optimized for, industrial application of TAL enzymes.
The HTS assay was specifically optimized to screen future bacterial TAL variants, more specifically but not exclusively FjTAL variants, for their catalytic activity and fitness for further downstream therapeutic applications. FjTAL variants with improved catalytic activity can be developed using directed protein evolution techniques to create a library of FjTAL variants with a possibly improved catalytic activity39,40. This library of FjTAL variants can then be transformed into E. coli BL21 (DE3) cells followed by induction of the expression of the variants and screening for TAL activity using the here developed HTS system. Variants with improved activity could then be tested for their capacity to decrease Tyr and could be used, by analogy for the use of a bacterial PAL in phenylketonuria, to resolve the acquired hypertyrosinemia in HT1 and AKU patients treated with NTBC well as to decrease Tyr levels in patients suffering from HT2 and HT3. Although the assay may be indicative of improved TAL variants to decrease Tyr levels, it should be noted that it concerns a bacterial assay and is thus not representative of the effect of these TAL variants in human cell systems or of the in vivo situation in patients. To make an estimation of the real-life impact of the improved TAL variants, additional in vitro tests in human based cell models of HT1 and AKU as well as in vivo experiments on HT1 and AKU mouse models would be required. Additionally, the improved TAL variants could also be used for the production of various previously mentioned aromatic compounds, as the TAL reaction is currently the rate limiting step in these processes16,20
In conclusion, in this study, we developed a fast, simple, cost-effective, robust and reliable HTS assay for the evaluation of bacterial TAL enzymes, which can be employed to evaluate the activity of a library of bacterial TAL enzymes in the context of developing new therapies to treat hypertyrosinemia in TIMD patients. Assay parameters such as assay pH are easily adjustable and thus allow for the evaluation of various TAL enzymes or to screen for TAL activity at the required pH necessary for the desired therapeutic application. However, it is important to bear in mind that although the assay allows to identify potent TAL enzymes, it will only be informative on the potential therapeutic effect in patients but does not necessarily reflect on the real-life situation.
Data availability
The datasets generated and/or analyzed during this study are available from the corresponding author on reasonable request.
References
Ghadimi, H. Diagnosis of inborn errors of amino acid metabolism. Arch. Pediatr. Adolesc. Med. 114(4), 433. https://doi.org/10.1001/archpedi.1967.02090250131016 (1967).
Chakrapani, A., Gissen, P. & McKiernan, P. Disorders of tyrosine metabolism. Inborn Metab. Dis. https://doi.org/10.1007/978-3-642-15720-2 (2012).
Holme, E. & Mitchell, G. A. Tyrosine Metabolism. Phys. Guide Diagnos. Treat. Follow-Up Inherit. Metab. Dis. https://doi.org/10.1007/978-3-642-40337-8 (2014).
Grompe, M. The pathophysiology and treatment of hereditary tyrosinemia type 1. Semin. Liver Dis. 21(4), 563–572. https://doi.org/10.1055/s-2001-19035 (2001).
Kvittingen, E. A. & Holme, E. Disorders of tyrosine metabolism. In Inborn metabolic diseases 186–194 (Springer, 2000).
Olsson, B. et al. Relationship between serum concentrations of nitisinone and its effect on homogentisic acid and tyrosine in patients with alkaptonuria. JIMD Rep. 24, 21–27. https://doi.org/10.1007/8904_2015_412 (2015).
Davison, A. S. et al. Assessment of the effect of once daily nitisinone therapy on 24-h urinary metadrenalines and 5-hydroxyindole acetic acid excretion in patients with alkaptonuria after 4 weeks of treatment. J. Inherit. Metab. Dis. https://doi.org/10.1007/8904 (2017).
García, M. I., de la Parra, A., Arias, C., Arredondo, M. & Cabello, J. F. Long-term cognitive functioning in individuals with tyrosinemia type 1 treated with nitisinone and protein-restricted diet. Mol. Genet. Metab. Rep 11, 12–16. https://doi.org/10.1016/j.ymgmr.2017.01.016 (2017).
Scott, C. R. The genetic tyrosinemias. Am. J. Med. Genet. C Semin. Med. Genet. 126, 121–126. https://doi.org/10.1002/ajmg.c (2006).
Viola, R. E. The ammonia-lyases: Enzymes that use a wide range of approaches to catalyze the same type of reaction catalyze the same type of reaction. Crit. Rev. Biochem. Mol. Biol. https://doi.org/10.1080/10409238.2019.1708261 (2020).
Parmeggiani, F., Weise, N. J., Ahmed, S. T. & Turner, N. J. Synthetic and Therapeutic applications of ammonia-lyases and aminomutases. Chem. Rev. https://doi.org/10.1021/acs.chemrev.6b00824 (2018).
Thomas, J. et al. Pegvaliase for the treatment of phenylketonuria: Results of a long-term phase 3 clinical trial program (PRISM). Mol. Genet. Metab. 124(1), 27–38. https://doi.org/10.1016/j.ymgme.2018.03.006 (2018).
Jendresen, C. B. et al. Highly active and specific tyrosine ammonia-lyases from diverse origins enable enhanced production of aromatic compounds in bacteria and Saccharomyces cerevisiae. Appl. Environ. Microbiol. 81(13), 4458–4476. https://doi.org/10.1128/AEM.00405-15 (2015).
Cha, H., Lee, S., Lee, J. H. & Park, J. Protective effects of p-coumaric acid against acetaminophen-induced hepatotoxicity in mice. Food Chem. Toxicol. https://doi.org/10.1016/j.fct.2018.08.060 (2018).
Akdemir, F. N. E., Albayrak, M., Çalik, M., Bayir, Y. & Gülçin, I. The protective effects of p -coumaric acid on acute liver and kidney damages induced by cisplatin. Biomedicines 5(18), 1–11. https://doi.org/10.3390/biomedicines5020018 (2017).
Barros, J. & Dixon, R. A. Plant phenylalanine/tyrosine ammonia-lyases. Trends Plant Sci. 25(1), 66–79. https://doi.org/10.1016/j.tplants.2019.09.011 (2020).
Kyndt, J. A., Meyer, T. E., Cusanovich, M. A. & Van Beumen, J. J. Characterization of a bacterial tyrosine ammonia lyase, a biosynthetic enzyme for the photoactive yellow protein. FEBS Lett. 512, 240–244. https://doi.org/10.1016/s0014-5793(02)02272-x (2002).
Nishiyama, Y. et al. Expression of bacterial tyrosine ammonia-lyase creates a novel p -coumaric acid pathway in the biosynthesis of phenylpropanoids in Arabidopsis. Planta 232(1), 209–218. https://doi.org/10.1007/s00425-010-1166-1 (2010).
Hausmann, O. et al. Pegvaliase: Immunological profile and recommendations for the clinical management of hypersensitivity reactions in patients with phenylketonuria treated with this enzyme substitution therapy. Mol. Genet. Metab. 128(1–2), 84–91. https://doi.org/10.1016/j.ymgme.2019.05.006 (2019).
Eudes, A. et al. Production of hydroxycinnamoyl anthranilates from glucose in Escherichia coli. Microb. Cell. Fact. 12(1), 62. https://doi.org/10.1186/1475-2859-12-62 (2013).
Rosano, G. L. & Ceccarelli, E. A. Recombinant protein expression in Escherichia coli: Advances and challenges. Front. Microbiol. https://doi.org/10.3389/fmicb.2014.00172 (2014).
Gopal, G. J. & Kumar, A. Strategies for the production of recombinant protein in Escherichia coli. Protein J. 32(6), 419–425. https://doi.org/10.1007/s10930-013-9502-5 (2013).
Jia, B. & Jeon, C. O. High-throughput recombinant protein expression in Escherichia coli : Current status and future perspectives. Open Biol. 6(8), 160196. https://doi.org/10.1098/rsob.160196 (2016).
Elabscience, ‘Elabscience Tyrosine Ammonia-Lyase (TAL) Assay Kit’. Accessed: Apr. 09, 2024. [Online]. Available: https://www.elabscience.com/p-tyrosine_ammonia_lyase_tal_activity_assay_kit-529622.html
Abbexa, ‘Abbexa Tyrosine Ammonia-Lyase (TAL) Assay Kit’. Accessed: Apr. 09, 2024. [Online]. Available: https://www.abbexa.com/tyrosine-ammonia-lyase-tal-assay-kit
Virklund, A., Jendresen, C. B., Nielsen, A. T. & Woodley, J. M. Characterization of tyrosine ammonia lyases from Flavobacterium johnsonian and Herpetosiphon aurantiacus. Biotechnol. J. https://doi.org/10.1002/biot.202300111 (2023).
Brack, Y., Sun, C., Yi, D. & Bornscheuer, U. T. Discovery of novel tyrosine ammonia lyases for the enzymatic synthesis of p -coumaric acid. ChemBioChem https://doi.org/10.1002/cbic.202200062 (2022).
P. W. Iversen et al., ‘HTS Assay Validation’, in Assay Guidance Manual, 2012. [Online]. Available: https://www.ncbi.nlm.nih.gov/books/
S. Amirreza-Mousavi-Majd, Flavobacterium johnsoniae Tyrosine Ammonia Lyase (FjTAL) in-silico structure prediction and molecular docking to L-Tyrosine, p-Coumaric Acid (pHCA) and Caffeic Acid, (2022), https://doi.org/10.1101/2022.02.09.479702.
Hughes, J. H. et al. Dietary restriction of tyrosine and phenylalanine lowers tyrosinemia associated with nitisinone therapy of alkaptonuria. J Inherit Metab Dis 43(2), 259–268. https://doi.org/10.1002/jimd.12172 (2020).
Schein, C. H. Production of soluble recombinant proteins in bacteria. Nat. Biotechnol. 7(11), 1141–1149. https://doi.org/10.1038/nbt1189-1141 (1989).
Baneyx, F. Recombinant protein expression in Escherichia coli. Curr. Opin. Biotechnol. 10(5), 411–421. https://doi.org/10.1016/S0958-1669(99)00003-8 (1999).
Papaneophytou, C. P. & Kontopidis, G. Statistical approaches to maximize recombinant protein expression in Escherichia coli: A general review. Protein Expr. Purif. 94, 22–32. https://doi.org/10.1016/j.pep.2013.10.016 (2014).
Shaw, M. K. & Ingraham, J. L. Synthesis of macromolecules by Escherichia coli near the minimal temperature for growth. J. Bacteriol. 94(1), 157–164. https://doi.org/10.1128/jb.94.1.157-164.1967 (1967).
Pinsach, J., de Mas, C., López-Santín, J., Striedner, G. & Bayer, K. Influence of process temperature on recombinant enzyme activity in Escherichia coli fed-batch cultures. Enzyme Microb. Technol. 43(7), 507–512. https://doi.org/10.1016/j.enzmictec.2008.08.007 (2008).
Noor, R., Islam, Z., Munshi, S. K. & Rahman, F. Influence of temperature on Escherichia coli growth in different culture media. J. Pure Appl. Microbiol. 7(2), 899–904 (2013).
Iversen, P. W., Eastwood, B. J., Sittampalam, G. S. & Cox, K. L. A comparison of assay performance measures in screening assays: Signal window, Z′ Factor, and assay variability ratio. SLAS Discov. 11(3), 247–252. https://doi.org/10.1177/1087057105285610 (2006).
Zhang, J.-H., Chung, T. D. Y. & Oldenburg, K. R. A Simple statistical parameter for use in evaluation and validation of high throughput screening assays. SLAS Discov. 4(2), 67–73. https://doi.org/10.1177/108705719900400206 (1999).
Wang, Y. et al. Directed evolution: Methodologies and applications. Chem. Rev. 121(20), 12384–12444. https://doi.org/10.1021/acs.chemrev.1c00260 (2021).
Cirino, P. C., Mayer, K. M. & Umeno, D. Generating Mutant Libraries Using Error-Prone PCR. In Directed Evolution Library Creation 3–10 (Humana Press, 2003).
Acknowledgements
The authors would like to thank the Institute of Biotechnology at RWTH Aachen for the supplies, excellent know-how and assistance. Figures were partially made using Servier Medical Art.
Funding
This study was supported by the Research Foundation Flanders (FWO) under grant numbers G041521N, 11L5124N, 11P1T24N and 11P3024N.
Author information
Authors and Affiliations
Contributions
I.N. designed and executed the experiments, collected and analyzed the data, prepared the figures and wrote the manuscript. C.A.L., N.S.S. and L.D. co-executed some of the experiments and reviewed the manuscript. S.L. and J.N. co-designed the experiments and reviewed the manuscript. U.S. co-supervised the study and reviewed the manuscript. J.D.K. co-designed the experiments, supervised the study and reviewed the manuscript. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nulmans, I., Laga, C.A., Salvi, N.S. et al. A robust high-throughput screening system to assess bacterial tyrosine ammonia lyase activity in the context of tyrosine inherited metabolic disorders. Sci Rep 14, 22175 (2024). https://doi.org/10.1038/s41598-024-72360-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-72360-9