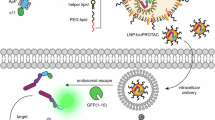
Abstract
Dysregulation of integral membrane proteins (IMPs) has been linked to a myriad of diseases, making these proteins an attractive target in drug research. Whilst PROTAC technology has had a significant impact in scientific research, its application to IMPs is still limited. Limitations of the traditional approach of immunoblotting in PROTAC research include the low throughput compared to other methods, as well as a lack of spatial information for the target. Here we compare orthogonal antibody based approaches, i.e. immunoblotting, flow cytometry and immunofluorescence, to measure PROTAC mediated degradation of two established, endogenous targets, epidermal growth factor receptor (EGFR) and hepatocyte growth-factor receptor (c-MET). We discuss advantages and limitations of each methodology for the assessment of PROTAC efficacy on IMPs. Overall, we recommend the use of immunofluorescence and flow cytometry, for an increased accuracy with both a qualitative and quantitative insight into degradation efficacy and a critical distinction between cell membrane-localized and intracellular IMP protein pools.
Similar content being viewed by others
Introduction
PROteolysis targeting chimeras (PROTACs) are heterobifunctional molecules composed of two connected ligands that link an E3 ubiquitin ligase to a target protein of interest (POI). PROTAC binding induces proximity of the E3 ligase to the target POI leading to its polyubiquitination and subsequent degradation via the ubiquitin-proteasome system (UPS)1. As an emerging class of drugs, PROTACs present novel therapeutic advantages: they remove the disease-linked protein from the cell, rather than only inhibiting it, thereby also eliminating any potential scaffolding function2,3; they act in an iterative manner inducing the degradation of the target in substoichiometric quantities, reducing the required administered dose compared to the parent inhibitor (i.e., POI-specific warhead) with tremendous impact on side effects and the therapeutic window3,4; they don’t rely on binding to an active site of target proteins but rather on availability of any accessible high affinity pocket5; they can achieve potent target degradation even with low-affinity ligands, thanks to the positive cooperative interaction between the E3 and the target4,6. These characteristics have made PROTACs potent new therapeutic agents that also open up the possibility of drugging previously undruggable targets7.
Despite their promising therapeutic potential as protein degraders, integral membrane protein (IMP) targeting PROTACs have seen slower progress compared to cytosolic or nuclear protein-targeting PROTACs due to the added complexity of location and membrane integration. Mutant and dysregulated IMPs are established driving factors for many disease indications, and their targeted degradation still remains a highly sought-after therapeutic objective (extensively reviewed in8).
Epidermal growth factor receptor (EGFR) and hepatocyte growth-factor receptor (c-MET) are membrane-integral receptor tyrosine kinases (RTKs), consisting of an N-terminal extracellular-facing ligand-binding domain, a single pass transmembrane helix and a C-terminal cytosolic domain containing a juxtamembrane domain and a kinase domain9,10. Ligand binding induces conformational changes and activation of the kinase domain, leading to autophosphorylation, effector-protein binding, and ultimately recognition by the E3 ligase Casitas B-lineage lymphoma (c-Cbl) followed by ubiquitination, internalization and degradation11,12,13.
EGFR and c-MET are implicated in the pathogenesis of various cancer types. Alterations in EGFR and aberrant activation of the receptor are commonly seen in Glioblastoma multiforme, squamous cell carcinomas in the head and neck (HNSC), lung (LUSC), and esophagus (ESCA), non-small cell lung cancer (NSCLC), bladder urothelial carcinoma and brain lower grade glioma14. Exon 19 deletion (Ex19del) is the most common activating mutation in advanced NSCLC, with a large number of molecular variants that arise from in-frame deletions, substitutions, and insertions15. Patients with Ex19del mutations are treated with EGFR tyrosine kinase inhibitors (TKIs) such as gefitinib, which competitively prevents the binding of ATP to the receptor that is required for receptor auto-phosphorylation. While gefitinib can bind WT EGFR, the affinity to Ex19del EGFR and other mutants is five to thirteen times higher16,17.
c-MET is altered in a variety of lung, kidney, ovary, thyroid, colon, gastric, liver and breast carcinomas. Oncogenic mutations can constitutively activate the receptor, prevent c-Cbl binding or generate alternatively spliced variants18. Exon skipping of exon 14, frequently ecountered in NSCLC and gastrointestinal malignancies, results in the omission of the c-Cbl-binding site within the juxtamembrane domain of c-MET leading to increased c-MET protein levels compared to cells expressing the wild type protein19,20. Exon 14 deletion (ex14del) results in tumor resistance to some c-MET inhibitors, but does not impact on Foretinib, a pan-kinase inhibitor that targets c-MET and the vascular endothelial growth factor receptor.
While PROTACs that achieve substantial degradation of either c-MET (with PROTAC SJF82402) or EGFR (e.g. with PROTAC32, or MS3921) have been described, the underlying mechanism for degradation was not defined in detail in these publications. SJF8240 (described as PROTAC7 in2) and PROTAC3 are VHL (von Hippel-Lindau)-based PROTACs with a respective c-MET inhibitor (foretinib) or EGFR inhibitor (gefitinib) as warheads. While SJF8240 induces c-MET degradation of both wildtype and mutant forms, PROTAC3 shows a selectivity towards degrading mutant EGFR, both Ex19del and L858R, while sparing wild type EGFR2.
Most of the published reports on PROTAC-mediated degradation of untagged proteins assess degradation efficacy through immunoblotting (e.g22,23,24). , as is also the case for IMP targeted protein degradation (TPD)25,26,27. This technique analyses whole cell protein lysates with no distinction between cell compartments, which requires different experimental approaches and time that complicate their use in large scale screens. Furthermore, immunoblot data are semiquantitative, and thus do not give a precise estimate of PROTAC efficacy. Here we compare different orthogonal methods of quantifying PROTAC mediated degradation of endogenous, untagged proteins of interest. Specifically, we investigate PROTAC efficacy of IMP-targeting chimeras, focussing on RTKs EGFR and c-MET. Our findings inform on the advantages and limitations of each methodology for the assessment of PROTAC efficacy on IMP TPD.
Results
While different methods are available to study target protein degradation, immunoblotting remains the primary method of choice when evaluating endogenous protein levels. We wanted to compare immunoblotting, immunofluorescence, and flow cytometry as high-throughput methods to study PROTAC-mediated degradation of integral membrane proteins c-MET and EGFR, both in their wildtype and mutant forms.
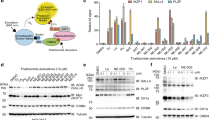
For both proteins, ligand induced endocytosis and degradation of the wildtype receptor is well established28,29. Hepatocyte growth factor (HGF) is known as the scatter factor that promotes tumor metastasis, activating c-MET and its downstream mitogenic pathways28. Similarly, the epidermal growth factor (EGF) binds and activates EGFR, stimulating cell growth and differentiation29. As positive controls of c-MET and EGFR degradation for the described experiments, we included samples treated with the aforementioned growth factors only. Their presence in the media mimics a physiological environment for the cells, and it is relevant to understand if and how they interfere with the PROTAC treatments. Another condition that we evaluated is serum starvation, which mimics the nutrient-poor tumoral environment commonly found in solid tumor30. To capture a degradation profile to compare between the different methods we treated the cells with different concentrations of PROTACs or inhibitors for 24 h. In these experiments we captured degradation profiles in full medium, either with or without pre-starvation in serum-free medium for 8 h, or with co-treatment with 100 ng/mL HGF or EGF (Fig. 1).
Degradation of WT c-MET in A549 cells
SJF8240 has previously been published as a potent c-MET degrader2. We decided to test the effect of SJF8240 on wild type c-MET in A549 cells which express wild type EGFR and c-MET, but mutant KRAS as their oncogenic driver acting downstream of EGFR and c-MET in the MAPK signalling pathway31,32. While serum starvation on its own did not have an effect on total c-MET protein levels, as detected by immunoblotting, HGF-induced endocytosis for 24 h led to 60–75% reduction in c-MET signal (Fig. 2A–D). When quantifying receptor abundance by immunofluorescence, we distinguished between whole cell (Fig. 2E, F) and plasma membrane (Fig. 2G, H) c-MET levels. The latter together with data from flow cytometric analysis (Fig. 2I, J) allows to assess alterations in the pool of active plasma membrane localized c-MET that is responsible for downstream signal transduction. Using immunofluorescence, we observed a ~ 50% reduction in total c-MET signal (Fig. 2E) after 24 h of HGF treatment, while cell-surface levels of c-MET decreased by ~ 35% (Fig. 2G). The observation that membrane occupancy of c-MET decreases less than total immunofluorescent signal in the cell suggests that HGF preferentially promotes the endocytosis of c-MET, but that endocytosed c-MET accumulates intracellularly before being degraded (Fig. 2K, S1). Flow cytometric analysis of the same treatment yielded a comparable reduction in c-MET cell-surface levels by approximately 60% (Fig. 2I, J). We did not see significant differences between the three tested methods in the HGF mediated changes of c-MET protein levels, however results for immunoblotting showed bigger variability than results for immunofluorescence or flow cytometry (Fig. S2A).
Degradation profile of wildtype c-MET in A549 cells. (A–J) Cells were pre-treated as described in (Fig. 1) with HGF as RTK ligand and either SJF8240 (A,C,E,G,I) or Foretinib (B,D,F,H,J) for 24 h. Cells were then analysed by immunoblotting (A,B) and quantification (C,D), whole cell (E,F) or plasma membrane (G,H) immunofluorescence, or by flow cytometry (I,J). Graphs show the mean and SEM of at least 3 biological replicates. The dotted line indicates a value of 0.8. Asterisks highlight p values < 0.01. All details regarding statistical analyses and p-values are provided in the (Supplementary Table 1).
In the presence of serum in the culture medium, treating A549 cells with SJF8240 resulted in a maximal reduction of c-MET signal of approximately 45% by immunoblotting, 25% by immunofluorescence (whole cell signal), and 62% with flow cytometry (Fig. 2A,C,E,I). Notably, the maximal effect in immunoblotting was observed at 100 nM SJF8240, while the other analysis methods recorded maximal effects at 1 µM. As for HGF treatment, we did not see significant differences when comparing the 1 µM treatment condition, however immunoblotting, again, yielded bigger variability than the other methods (Fig. S2A). We did not observe any synergy in promoting c-MET degradation as assessed by all the methods, when co-treating cells with HGF and SJF8240 (Fig. 2A,C,E,I). We did, however, observe a dose-dependent rescue of c-MET protein levels from HGF-induced degradation with increasing PROTAC concentrations both at the total protein level and the plasma membrane pool (Fig. 2K) with a near full rescue at 10 µM SJF8240. Foretinib binds to the c-MET ATP-binding site blocking its autophosphorylation and effector protein binding, thereby preventing RTK endocytosis and downstream signalling pathway. This indicates that the foretinib warhead in a PROTAC context can still exert its inhibitor function on c-MET. However, overall the impact of SJF8240 on c-MET inhibition and internalization was a lot less potent than with Foretinib alone (Fig. 2G,H), where full rescue was already observed at 100 nM. We observed the same effect on c-MET phosphorylation at Y1234/1235 where efficient block of phosphorylation with the PROTAC was only seen at 10 fold higher concentrations at 1 µM than with foretinib (Fig. S3A)2. Notably, ERK phosphorylation was only reduced when using Foretinib at high dose at 10 µM, in line with mutant KRAS bypassing the upstream MAPK signalling. Finally, serum starvation prior to PROTAC treatment did not influence the behaviour or maximal effects observed (Fig. 2A–J).
Degradation of mutant c-MET in Hs746T cells
Hs746T are a gastric carcinoma cell line in which the MET gene has been amplified as well as mutated resulting in exon 14 deletion, collectively leading to impaired protein turnover2. In line with the deleted c-Cbl binding site, HGF treatment did not lead to reduction of c-MET levels or endocytosis with any of the tested readout methods (Fig. 3A–J, S1A, S2B). Serum starvation did not have any effect on c-MET either (Fig. 3A–J).
Degradation profile of ex14del c-MET in HS746T cells. Cells were pre-treated as described in (Fig. 1) with HGF as RTK ligand and either SJF8240 (A,C,E,G,I) or Foretinib (B,D,F,H,J) was added for 24 h. Cells were then analysed by immunoblotting (A,B) with quantification (C,D), immunofluorescence for the whole cell (E,F) or plasma membrane only (G,H) or flow cytometry (I,J). Graphs show the mean SEM of at least 3 biological replicates. The dotted line indicates a value of 0.8. All details regarding statistical analyses and p-values are provided in the (Supplementary Table 2).
Analysis of Hs746T cells treated with PROTAC SJF8240 by immunoblotting shows c-MET degradation with a dose response and maximal degradation at 100 nM and an apparent hook effect at higher concentrations (Fig. 3A,C). More in line with the initial assessment of SJF8240 activity2, maximal c-MET signal reduction was recorded at 1 µM when using immunofluorescence and flow cytometry with maximal effects ranging from 60% signal reduction in flow cytometry and 35% in immunofluorescence (Fig. 3E,I). Intriguingly, we observed with all assays that HGF diminished the ability of SJF8240 to promote reduction of c-MET signal, even though in itself it did not do so (Fig. 3A,C,E,G,I). Additionally, serum-starvation did not change the degradation profile considerably compared to serum conditions.
The same set of experiments was also conducted using the Foretinib warhead alone. This reduced c-MET by approximately 50% by immunoblotting (Fig. 3B,D) or 20% by immunofluorescence (Fig. 3F). This was again counteracted by co-treatment with HGF or by serum-starvation. As was the case for measurements on c-MET levels in A549 cells, we did not observe any significant differences in the methods used for either HGF treatment, or 1 µM SJF8240 treatment, however, again, immunoblotting yielded high variability in both cases (Fig. S2B).
Degradation of WT EGFR in A549 cells
Similarly to c-MET, wildtype EGFR protein levels were not altered under serum-starvation in A549 cells with any of the tested methods (Fig. 4A–J). In contrast, EGF treatment on its own reduced EGFR total protein levels by approximately 75% in immunoblotting (Fig. 4A–D), 30% (whole cell) and 50% (membrane) in immunofluorescence (Fig. 4E–H, Fig. S1B) and 80% in flow cytometry (Fig. 4I,J), with immunofluorescence showing a significantly lower reduction of total protein levels (Fig. S2C) compared to immunoblotting and flow cytometry, which were not significantly different in direct comparison. Similar to what Burslem et al. show in their original publication2, we did not observe degradation of wildtype EGFR with any of the tested methods when using PROTAC3 in A549. This is in line with Gefitinib having a significantly lower affinity to wildtype EGFR compared to mutant EGFR2,17. However, we did observe very weak gefitinib warhead dependent inhibition of EGFR mediated endocytosis when using PROTAC3 at 10 µM (Fig. 4G). This is in contrast to gefitinib alone, which efficiently blocked EGF-induced degradation (Fig. 4B,D,F) and endocytosis (Fig. 4H,J) in a dose dependent manner.
Degradation profile of wildtype EGFR in A549 cells. Cells were pre-treated as described in (Fig. 1) with EGF as RTK ligand and either PROTAC3 (A,C,E,G,I) or Gefitinib (B,D,F,H,J) was added for 24 h. Cells were then analysed by immunoblotting (A,B) with quantification (C,D), immunofluorescence for the whole cell (E,F) or plasma membrane only (G,H) or flow cytometry (I,J). Graphs show the mean SEM of at least 3 biological replicates. The dotted line indicates a value of 0.8. All details regarding statistical analyses and p-values are provided in the (Supplementary Table 3).
Degradation of mutant EGFR in HCC827 cells
HCC827 cells express exon 19-deleted EGFR which is more resistant to EGF-induced downregulation, but responds to gefitinib inhibition21. In line with this, treating HCC827 cells with EGF did not change total protein levels or EGFR localization when analysed by immunofluorescence (Fig. 5E–H, Fig. S1B) or flow cytometry (Fig. 5I,J) and the two methods did not significantly differ from each other (Fig. S2D). In contrast, we observed a significant reduction of EGFR protein levels by 30–60% when treating samples under the same conditions and analysing them by immunoblotting (Fig. 5A–D, Fig. S2D). Notably, immunoblotting showed significant differences to both, immunofluorescence and flow cytometry (Fig. S2D). Similar to A549 cells, serum-starvation had no effect on EGFR protein levels regardless of the method used for analysis (Fig. 5A–J).
Degradation profile of ex19del EGFR in HCC827 cells. Cells were pre-treated as described in (Fig. 1) with EGF as RTK ligand and either PROTAC3 (A,C,E,G,I) or Gefitinib (B,D,F,H,J) was added for 24 h. Cells were then analysed by immunoblotting (A,B) with quantification (C,D), immunofluorescence for the whole cell (E,F) or plasma membrane only (G,H) or flow cytometry (I,J). Graphs show the mean SEM of at least 3 biological replicates. The dotted line indicates a value of 0.8. All details regarding statistical analyses and p-values are provided in the (Supplementary Table 4).
Treating HCC827 cells with PROTAC3 elicited the same response across all methods, with maximal signal reduction being observed at 1 µM, and signal intensity increasing again at 10 µM, reflecting a typical hook effect (Fig. 5A,C,E,G,I). Maximal measured changes in EGFR protein levels were different between immunofluorescence (Fig. 5E,G), where signal reduction was approximately 25%, and immunoblotting and flow cytometry where the signal was reduced by 70% (Fig. 5A,C,I). The measured difference in signal reduction between immunofluorescence and the other two methods was significant, while immunoblotting and flow cytometry did not differ significantly from each other (Fig. S2D). When analysing co-treatment of cells with EGF and PROTAC3 by immunoblotting or flow cytometry, there seemed to be a synergistic effect in EGFR degradation, something we did not observe with immunofluorescence. We analysed immunofluorescence images, for these conditions and observed a signal accumulation in intracellular speckles (Fig. 5K, white arrows) with an apparent reduction in plasma membrane signal, both for PROTAC3 single or EGF co-treatment. Both, treatment with PROTAC3 and Gefitinib, reduced EGFR phosphorylation at tyrosine residue 1068 with the lowest tested concentration of 100 nM (Fig. S3B). For both compounds treatment at 10 µM led to the highest effect on downstream phosphorylation of ERK.
Discussion
Traditional studies of PROTAC efficacy utilize immunoblotting as a standard method, (e.g22,23,24). providing semiquantitative data of a single time point snapshot of whole cell protein levels, without resolution of spatial information. This can be circumvented by expanding lysate preparation to subcellular fractionation that could address whether a protein a depleted from endocytic organelles, the cytosol or the nucleus. However, this strategy comes with high cost and low throughput. Depending on the protein of interest, sample preparation can also affect the result of immunoblotting, with centrifugation steps leading to potential loss of protein associated with “cellular debris”33, or heating samples leading to differential resolution upon electrophoresis34. In contrast, immunofluorescence provides the additional layer of information, where signal quantification provides the readout of target degradation, and the images themselves provide information on whether there are significant changes in target localization or aggregation. While we did not see any significant differences between the tested methods when analysing c-MET degradation, mostly due to high variability in immunoblot quantification (Fig. S2A, B), we found immunoblotting to mimic flow cytometry results far more than immunofluorescence when analysing EGFR degradation, either by PROTAC3 or by EGF (Fig. S2C,D). Overall, the trend of whether a condition induced protein level reduction was consistent between methods, however, given that flow cytometry measures protein internalization rather than degradation, the close mimic of immunoblotting is concerning, as this indicates a strong overstatement of perceived maximal degradation. One possibility for the additional removal of EGFR from the lysate could be the speckles we observed after PROTAC treatment (Fig. 5K), which might pellet during centrifugation rather than stay in the soluble fraction.
Additional downsides to immunoblotting are low and work intensive throughput, compared to other methods. This becomes especially problematic with the ever growing size of PROTAC libraries to be tested, due to different requirements in warheads and linkerology for different targets in order to achieve efficient target degradation (e.g35,36). In contrast to immunoblotting, flow cytometry and immunofluorescence enable a higher throughput, with an increased accuracy and scalability36,37,38,39. Immunofluorescence provides information about the subcellular localization of the target and in the best case allows a clear separation of membrane associated proteins from the cytosolic or intracellular protein pool resulting in a clear picture of whether a protein has been degraded or simply endocytosed (e.g. Figure 5I vs. Figure 5K at 10 µM PROTAC3 vs. DMSO).
As an example, treatment of EGFR and c-MET mutant cell lines HCC827 and HS746T with 1 µM PROTAC3 or SJF8240, respectively, showed both total protein reduction by immunoblotting and flow cytometry, indicating that the protein was either degraded (immunoblot, Figs. 3A and C and 5A and C) or endocytosed (Figs. 3I and 5I). Adding immunofluorescence reveals that more EGFR is endocytosed than degraded (Fig. 5E and G, S1), while for c-MET more protein is being degraded than endocytosed (Fig. 3E and G, S1). This indicates that EGFR degradation occurs on the endo-lysosomal (re)cycling pool, while targeted degradation of c-MET depletes the newly synthesized pool of protein. In line with this notion, we observed appearance of cytosolic EGFR speckles after PROTAC treatment, whereas c-MET results in a uniform reduction of overall signal (Fig. 5K). However, this will need to be confirmed in the future with experiments aimed specifically at answering the question of whether EGFR and c-MET are being targeted at different stages in their life cycle. Notably, for c-MET we could show that in the PROTAC context, the foretinib warhead still exerted its inhibitor function, as HGF induced endocytosis in A549 cells was dose-dependently blocked, albeit with a much lower potency than foretinib on its own (Fig. 2G, S3A). Changes of inhibitor binding affinity in a PROTAC context have been described before40. ERK phosphorylation remained unchanged, apart from when using high concentration of Foretinib, which is in line with the activating KRAS mutation in A549 cells bypassing the MAPK signalling pathway41. Similarly, PROTAC3 at 10 µM showed trends of inhibiting EGF induced endocytosis of EGFR (Fig. 4G), however given the low potency of gefitinib on wildtype EGFR (Fig. 4H)42, the result was not as clear-cut as for foretinib and c-MET. Crucially, these observations could not be made in either HS746T or HCC827 cells, as neither mutant c-MET, nor mutant EGFR were susceptible to ligand induced endocytosis (Figs. 3I and 4I).
As all the tested methods are antibody based, which can introduce different epitopes recognition and binding affinities, it is of interest to note that we observed differences in perceived maximal effects of relative signal intensities. Generally, immunoblotting seemed to yield higher differences in relative protein signals compared to immunofluorescence. However, we used the same antibody targeting the cytosolic tail of either c-MET or EGFR. As an alternative high-throughput method, flow cytometry only provides a measurement of receptor internalization from the cell surface, i.e. a smaller target pool compared to the whole cell content of target protein, which would explain the consistently increased sensitivity in measuring signal reduction compared to immunoblotting and immunofluorescence. While this generally agreed with negative data, i.e. we did not notice degradation in flow cytometry when there was no degradation in either immunoblotting or immunofluorescence, positive data have to be carefully analysed due to the aforementioned effect mixture of both protein degradation and endocytosis. Curiously, flow cytometry seemed to overall agree with immunoblotting data more than with immunofluorescence (Fig. S2).
While the PROTAC and targeted protein degradation field has gained a lot of traction in the last decade, PROTAC-mediated degradation of integral membrane proteins is still poorly understood. Published PROTACs targeting IMPs2,21,43, still leave questions on their mode of action: which pool of integral membrane proteins is being targeted by the PROTAC (i.e. newly synthesized vs. mature, membrane localized protein)? Is degradation of integral membrane proteins achieved exclusively via the lysosomal degradation pathway, or does it follow a traditional proteasomal pathway as well due to more promiscuous and potentially unconventional ubiquitination sites? In regards to the latter, it is generally assumed, and has been shown for most cytosolic and nuclear PROTAC targets, that degradation is mediated by the proteasome44,45,46. While proteasomes are generally assumed to be localized both in the nucleus and the cytosol, membrane association has been reported at both endo- and plasma membranes47,48,49,50. In this report we demonstrated how PROTAC-induced integral membrane protein degradation can be robustly characterised using immunoblotting, flow cytometry and immunofluorescence, and that the combination of these methodologies contributes to a better understanding of IMP degradation. Overall, we recommend the use of immunofluorescence rather than immunoblotting, due to the additional layer of information extracted with these types of experiments which would not be feasible by immunoblotting. Most importantly, both immunofluorescence and flow cytometry provide both a qualitative and quantitative insight into degradation efficacy and a critical distinction between cell membrane-localized and intracellular IMP protein pools. One way to enhance the resolution of the IMP degradation mechanism with PROTAC would be to use pulse-chase experiments with mass spectrometry like SILAC (stable isotope labelling by amino acids in cell culture), which delivers comparative data on both protein decay and novel protein synthesis, but such technologies are not yet ready to be delivered at scale.
Material & methods
Cell culture
Hs746T (ATCC, HTB-135) are a gastric carcinoma cell line isolated from a 74-year-old white male. A549 (ATCC, CCL-185) are a lung carcinoma cell line isolated from a 58-year-old white male. HCC-827 (ATCC, CRL-2868) are a lung adenocarcinoma cell line isolated from a white 39-year-old female. HS746T and A549 cells were cultured in DMEM + GlutaMAX (ThermoFisher, 31966-047) medium supplemented with 10% Fetal Bovine Serum (FBS; ThermoFisher, 10270-106). HCC-827 cells were cultured in RPMI-1640 (SigmaAldrich, R8758) medium supplemented with 10% FBS. For starvation experiments, cells were washed twice in PBS and grown in respective growth media without FBS for 6–8 h. Subsequently media were exchanged for full growth media and cells were treated with SJF8240 (Tocris, 7266), Foretinib (Stratech, ORB322222), PROTAC3 (Tocris, 7258) or Gefitinib (AstraZeneca). For HGF (Bio-Techne, 294-HG-025) or EGF (Bio-Techne, 236-EG-200) treatment, full growth media containing ligands at 100 ng/mL were added just before PROTAC treatment. For serum conditions full growth medium was replenished before PROTAC treatment. PROTAC treatment was conducted for 24 h.
Immunoblot
For harvesting, cells were washed in PBS and lysed in RIPA buffer (ThermoFisher, 89901) containing protease inhibitors (ThermoFisher, 87786) and phosphatase inhibitors (ThermoFisher, 78420). For efficient cell disruption, lysates were kept on ice for 20 min, with occasional vortexing, or frozen at -20⁰C and thawed twice while vortexing. Subsequently lysates were cleared by centrifugation at 16,000xg for 10 min at 4 °C. Lysates were mixed with reducing agent (ThermoFisher, NP0009) and LDS sample buffer (ThermoFisher, NP0007) at appropriate dilutions and heated to 70 °C for 10 min. Proteins were separated on 4 − 12% Bis-Tris Protein gels (ThermoFisher, NP0336BOX) in NUPAGE MOPS SDS Running buffer (ThermoFisher, NP0001) and transferred onto PVDF membranes using the iBlot2 system (ThermoFisher, IB24001) with default program P0 (20 V 1 min, 23 V 4 min and 25 V 2 min). Membranes were then blocked in Intercept (TBS) blocking buffer (Li-Cor, 927-60001) for 1 h at room temperature, followed by primary antibody incubation overnight at 4 °C. Primary antibodies were diluted in Intercept blocking buffer at 1:1,000. Membranes were washed with TBS-T buffer before incubation with IRDye secondary antibodies diluted 1:5,000 in Intercept blocking buffer for 1 h at room temperature. Membranes were washed with TBST buffer and imaged on an Odyssey DLx. Empiria Studio software was used for analysis and quantification. Data were normalised for the DMSO treated samples. The following primary and secondary antibodies were used: EGFR (C-terminus, total: CST, #4267; phospho Y1068: CST, #2236), MET (C-terminus total (main figures): CST, #8198; C-terminus total (Fig. S4): CST, #3148; phospho Y1234/1235: CST, #3077), ERK (total: CST, #9102; phospho T202/Y204: CST, #9106), IRDye® 800CW Donkey anti-Rabbit IgG (Li-Cor, 926-32213), IRDye® 680RD Donkey anti-Mouse IgG Secondary Antibody (Li-Cor, 926-68072). Immunoblots shown in figures have been cropped for visualization. Original blots can be found in supplementary file S6.
Immunofluorescence
Cells were seeded onto poly-D-lysine coated 384 well plates (Corning, 734 − 0260) and treated as described above for different treatment regimes. At observation timepoint, cells were washed in PBS and fixed in 4% paraformaldehyde (PFA) (diluted in PBS, Sigma P6148) for 10 min at room temperature. Plates were washed with PBS and permeabilized and blocked for 1 h at room temperature with 0.1% Triton + 1% BSA (Sigma X100, Sigma A7030) in PBS. Primary antibodies were diluted at 1:500 in the same blocking buffer and incubated over night at 4⁰C. The next day plates were washed with PBS and incubated with a mix of secondary antibody at 1:5,000, Hoechst33342 at 1:5,000 (Invitrogen, H3570) and Cell Mask Orange at 1:50,000–1:100,000 (ThermoFisher, C10045) for 1–2 h. Plates were again washed and then left in PBS for imaging.
Plates were imaged with Yokogawa Cell Voyager 7000. Images were analysed on Columbus (PerkinElmer). In short cells were identified by nuclear stain and cell membranes and cytosol were identified by subsequent cell mask orange stain. Cells on the image border were removed from analysis. Antibody stain intensity was quantified either for the whole cell or specifically for the membrane. Data were normalised for the DMSO treated samples. Two-tailed t-test was run for each condition with alpha 0.05 to evaluate if the difference with the DMSO treated sample is significant (P < 0.05). The following primary and secondary antibodies were used: EGFR (C-terminus; CST, #4267), MET (C-terminus; CST, #8198), Donkey anti-Rabbit IgG (H + L) Highly Cross-Adsorbed Secondary Antibody, Alexa Fluor™ 488 (ThermoFisher, A-21206).
Flow cytometry
Cells were seeded onto TC-treated 96 well plates (Corning, 3596) and treated as described above for different treatment regimes. At observation timepoint, cells were washed in PBS, harvested with TrypLE™ Express (ThermoFisher 12604013) and resuspended in cold FACS buffer – 0.5% BSA (Sigma A7030), 2 mM EDTA (ThermoFisher 15575020) in PBS. APC-conjugated antibodies were diluted at 1:200 in the same FACS buffer and incubated for 1 h at 4 °C. After 30 min, Live/Dead fixable yellow staining (Thermofisher L34967) was added at 1:1,000. Cells were washed in FACS buffer and fixed in 4% paraformaldehyde (PFA) (diluted in PBS, Sigma P6148) for 10 min at room temperature. They were washed, resuspended in FACS buffer and analysed with BD LSRFortessa with a High Throughput Sampler. 10’000 events per condition were acquired. Analysis was performed with FlowJo. Single viable cells (FSC-A vs. SSC-A, FSC-H vs. FSC-A) were gated and compensated based on the FMOs. APC signal was quantified. Data were normalised for the DMSO treated samples. Two-tailed t-test was run for each condition with alpha 0.05 to evaluate if the difference with the DMSO treated sample is significant (P < 0.05). The following APC-conjugated primary antibodies were used: EGFR (Bio-Legend 352905), MET (Bioscience 566789).
Statistical analysis
Statistical analysis was done by two-tailed t-test with alpha 0.05 to test for significant difference of treatment conditions compared to DMSO. Analysis was done on an N of at least 3. Statistical data are summarized in (Supplementary Tables 1–4).
Data availability
Data and statistical analyses are available upon request. Uncropped immunoblot images are supplied in (Supplementary Figure S4).
References
Sakamoto, K. M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl. Acad. Sci. USA 98 (15), 8554–8559 (2001).
Burslem, G. M. et al. The advantages of targeted protein degradation over inhibition: an RTK case study. Cell. Chem. Biol. 25 (1), 67–77 (2018).
Mares, A. et al. Extended pharmacodynamic responses observed upon PROTAC-mediated degradation of RIPK2. Commun. Biol. 3 (1), 1–13. (2020).
Bondeson, D. P. et al. Catalytic in vivo protein knockdown by small-molecule PROTACs. Nat. Chem. Biol. 11 (8), 611–617 (2015).
Guo, J., Liu, J. & Wei, W. Degrading proteins in animals: PROTACtion goes in vivo. Cell. Res. 29 (3), 179–180 (2019).
Han, X. et al. Discovery of ARD-69 as a highly potent proteolysis targeting chimera (PROTAC) degrader of androgen receptor (AR) for the treatment of prostate Cancer. J. Med. Chem. 62 (2), 941–964 (2019).
Bond, M. J. et al. Targeted degradation of oncogenic KRAS(G12C) by VHL-Recruiting PROTACs. ACS Cent. Sci. 6 (8), 1367–1375 (2020).
Ruffilli, C. et al. Proteolysis targeting chimeras (PROTACs): a perspective on integral membrane protein degradation. ACS Pharmacol. Transl. Sci. 2022 (2575–9108 ).
Gherardi, E. et al. Functional map and domain structure of MET, the product of the c-met protooncogene and receptor for hepatocyte growth factor/scatter factor. Proc. Natl. Acad. Sci. USA 100 (21), 12039–12044 (2003).
Lee, N. Y., Hazlett, T. L. & Koland, J. G. Structure and dynamics of the epidermal growth factor receptor C-terminal phosphorylation domain. Protein Sci. 15 (5), 1142–1152 (2006).
Peschard, P. & Park, M. Escape from Cbl-mediated downregulation. Cancer Cell. 3 (6), 519–523 (2003).
Critchley, W. R. et al. Receptor tyrosine kinase ubiquitination and de-ubiquitination in signal transduction and receptor trafficking. Cells 7 (3). (2018).
Li, N. et al. Specific Grb2-mediated interactions regulate clathrin-dependent endocytosis of the cmet-tyrosine kinase. J. Biol. Chem. 282 (23), 16764–16775 (2007).
Liu, H., Zhang, B. & Sun, Z. Spectrum of EGFR aberrations and potential clinical implications: insights from integrative pan-cancer analysis. Cancer Commun. (Lond.) 40 (1), 43–59 (2020).
Xu, C. W. et al. Molecular characteristics and clinical outcomes of EGFR exon 19 C-Helix deletion in non-small cell lung cancer and response to EGFR TKIs. Transl Oncol. 13 (9), 100791 (2020).
Mulloy, R. et al. Epidermal growth factor receptor mutants from human lung cancers exhibit enhanced catalytic activity and increased sensitivity to gefitinib. Cancer Res. 67 (5), 2325–2330 (2007).
Yun, C. H. et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl. Acad. Sci. USA 105 (6), 2070–2075 (2008).
Sierra, J. R. & Tsao, M. S. c-MET as a potential therapeutic target and biomarker in cancer. Ther. Adv. Med. Oncol. 3 (1), S21–35 (2011).
Van Der Steen, N. et al. cMET exon 14 skipping: from the structure to the clinic. J. Thorac. Oncol. 11 (9), 1423–1432 (2016).
Kim, H. J. et al. Forty-nine gastric cancer cell lines with integrative genomic profiling for development of c-MET inhibitor. Int. J. Cancer 143 (1), 151–159 (2018).
Cheng, M. et al. Discovery of potent and selective epidermal growth factor receptor (EGFR) bifunctional small-molecule degraders. J. Med. Chem. 63 (3), 1216–1232 (2020).
Khan, S. et al. A selective BCL-X(L) PROTAC degrader achieves safe and potent antitumor activity. Nat. Med. 25 (12), 1938–1947 (2019).
Nunes, J. et al. Targeting IRAK4 for degradation with PROTACs. ACS Med. Chem. Lett. 10 (7), 1081–1085 (2019).
Shiah, J. V., Grandis, J. R. & Johnson, D. E. Targeting STAT3 with proteolysis targeting chimeras and next-generation antisense oligonucleotides. Mol. Cancer Ther. 20 (2), 219–228 (2021).
Du, G. et al. Discovery of a potent degrader for fibroblast growth factor receptor 1/2. Angewandte Chemie Int. Ed. 60 (29), 15905–15911 (2021).
Chen, Y. et al. Degrading FLT3-ITD protein by proteolysis targeting chimera (PROTAC). Bioorg. Chem., 119. (2022).
Yan, G. et al. Discovery of a PROTAC targeting ALK with in vivo activity. Eur. J. Med. Chem., 212. (2021).
Birchmeier, C. et al. Met, metastasis, motility and more. Nat. Rev. Mol. Cell. Biol. 4 (12), 915–925 (2003).
Wee, P. & Wang, Z. Epidermal growth factor receptor cell proliferation signaling pathways. Cancers 9 (5), 52–52. (2017).
Kato, K. et al. Critical roles of AMP-activated protein kinase in constitutive tolerance of cancer cells to nutrient deprivation and tumor formation. Oncogene 21 (39), 6082–6090 (2002).
Korrodi-Gregorio, L. et al. From proteomic analysis to potential therapeutic targets: Functional profile of two lung cancer cell lines, A549 and SW900, widely studied in pre-clinical research. PLoS One 11 (11), e0165973 (2016).
Yoon, Y. K. et al. KRAS mutant lung cancer cells are differentially responsive to MEK inhibitor due to AKT or STAT3 activation: implication for combinatorial approach. Mol. Carcinog. 49 (4), 353–362 (2010).
Murphy, R. M., Mollica, J. P. & Lamb, G. D. Plasma membrane removal in rat skeletal muscle fibers reveals caveolin-3 hot-spots at the necks of transverse tubules. Exp. Cell. Res. 315 (6), 1015–1028 (2009).
Tsuji, Y. Transmembrane protein Western blotting: impact of sample preparation on detection of SLC11A2 (DMT1) and SLC40A1 (ferroportin). PLoS One 15 (7), e0235563 (2020).
Bashore, F. M. et al. PROTAC Linkerology leads to an optimized bivalent chemical degrader of polycomb repressive complex 2 (PRC2) components. ACS Chem. Biol. 18 (3), 494–507 (2023).
Stacey, P. et al. A phenotypic approach for the identification of new molecules for targeted protein degradation applications. SLAS Discov. 26 (7), 885–895 (2021).
Fang, X. et al. Recent advances in design of fluorescence-based assays for high-throughput screening. Anal. Chem. 91 (1), 482–504 (2019).
Yaseen, Z. et al. Validation of an accurate automated multiplex immunofluorescence method for Immuno-profiling Melanoma. Front. Mol. Biosci. 9, 810858 (2022).
Martinez-Gzegozewska, Y. et al. High-throughput cell-based immunofluorescence assays against influenza. SLAS Discov. 29 (1), 66–76 (2024).
Bondeson, D. P. et al. Lessons in PROTAC design from selective degradation with a promiscuous warhead. Cell. Chem. Biol. 25 (1), 78–87e5 (2018).
Zhao, D. et al. Clinical and molecular features of KRAS-mutated lung cancer patients treated with immune checkpoint inhibitors. Cancers (Basel) 14 (19). (2022).
Kim, B. et al. Clathrin-mediated EGFR endocytosis as a potential therapeutic strategy for overcoming primary resistance of EGFR TKI in wild-type EGFR non-small cell lung cancer. Cancer Med. 10 (1), 372–385 (2021).
Du, Y. et al. Discovery of novel EGFR-PROTACs capable of degradation of multiple EGFR-mutated proteins. Eur. J. Med. Chem. 272, 116489 (2024).
Zhang, H. et al. Discovery of potent epidermal growth factor receptor (EGFR) degraders by proteolysis targeting chimera (PROTAC). Eur. J. Med. Chem., 189. (2020).
Li, Z. et al. First small-molecule PROTACs for G protein-coupled receptors: inducing α1A-adrenergic receptor degradation. Acta Pharm. Sin. B 10 (9), 1669–1679 (2020).
Huber, M. E. et al. A Chemical biology toolbox targeting the intracellular binding site of CCR9: Fluorescent ligands, new dug leads and PROTACs. Angew. Chemie 61 (12). (2022).
Wojcik, C. & DeMartino, G. N. Intracellular localization of proteasomes. Int. J. Biochem. Cell. Biol. 35 (5), 579–589 (2003).
Guo, X. Localized proteasomal degradation: from the nucleus to cell periphery. Biomolecules 12 (2). (2022).
Bingol, B. & Sheng, M. Deconstruction for reconstruction: the role of proteolysis in neural plasticity and disease. Neuron 69 (1), 22–32 (2011).
Montenegro-Venegas, C. et al. Bassoon inhibits proteasome activity via interaction with PSMB4. Cell. Mol. Life Sci. 78 (4), 1545–1563 (2021).
Acknowledgements
CR, NZ, KM received funding from the European Union’s Horizon 2020 Research and Innovation programme under the Marie Skłodowska-Curie Grant Agreement 953489 as part of the EndoConnect training expertise.
Author information
Authors and Affiliations
Contributions
CR, SR and KM designed the project. CR and SR planned and performed the experiments and analysed the data. CR and SR drafted the manuscript. All authors discussed the results and contributed to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing financial interest(s): CR, SR and KM are employees of AstraZeneca. NZ has no competing interests to declare.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ruffilli, C., Röth, S., Zelcer, N. et al. Orthogonal validation of PROTAC mediated degradation of the integral membrane proteins EGFR and c-MET. Sci Rep 15, 504 (2025). https://doi.org/10.1038/s41598-024-84217-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-84217-2