Abstract
Diarrhoeal disease is responsible for 8.6% of global child mortality. Recent epidemiological studies found the protozoan parasite Cryptosporidium to be a leading cause of paediatric diarrhoea, with particularly grave impact on infants and immunocompromised individuals. There is neither a vaccine nor an effective treatment. Here we establish a drug discovery process built on scalable phenotypic assays and mouse models that take advantage of transgenic parasites. Screening a library of compounds with anti-parasitic activity, we identify pyrazolopyridines as inhibitors of Cryptosporidium parvum and Cryptosporidium hominis. Oral treatment with the pyrazolopyridine KDU731 results in a potent reduction in intestinal infection of immunocompromised mice. Treatment also leads to rapid resolution of diarrhoea and dehydration in neonatal calves, a clinical model of cryptosporidiosis that closely resembles human infection. Our results suggest that the Cryptosporidium lipid kinase PI(4)K (phosphatidylinositol-4-OH kinase) is a target for pyrazolopyridines and that KDU731 warrants further preclinical evaluation as a drug candidate for the treatment of cryptosporidiosis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Liu, L. et al. Global, regional, and national causes of under-5 mortality in 2000–15: an updated systematic analysis with implications for the Sustainable Development Goals. Lancet 388, 3027–3035 (2016)
Kotloff, K. L. et al. Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. Lancet 382, 209–222 (2013)
Platts-Mills, J. A . et al. Pathogen-specific burdens of community diarrhoea in developing countries: a multisite birth cohort study (MAL-ED). Lancet Glob. Health 3, e564–e575 (2015)
Checkley, W. et al. A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for cryptosporidium. Lancet Infect. Dis. 15, 85–94 (2015)
Amadi, B. et al. High dose prolonged treatment with nitazoxanide is not effective for cryptosporidiosis in HIV positive Zambian children: a randomised controlled trial. BMC Infect. Dis. 9, 195 (2009)
Amadi, B. et al. Effect of nitazoxanide on morbidity and mortality in Zambian children with cryptosporidiosis: a randomised controlled trial. Lancet 360, 1375–1380 (2002)
Striepen, B. Parasitic infections: time to tackle cryptosporidiosis. Nature 503, 189–191 (2013)
Vinayak, S. et al. Genetic modification of the diarrhoeal pathogen Cryptosporidium parvum. Nature 523, 477–480 (2015)
Rottmann, M. et al. Spiroindolones, a potent compound class for the treatment of malaria. Science 329, 1175–1180 (2010)
Bürstner, N. et al. Gift from nature: cyclomarin A kills mycobacteria and malaria parasites by distinct modes of action. ChemBioChem 16, 2433–2436 (2015)
Meister, S. et al. Imaging of Plasmodium liver stages to drive next-generation antimalarial drug discovery. Science 334, 1372–1377 (2011)
McNamara, C. W. et al. Targeting Plasmodium PI(4)K to eliminate malaria. Nature 504, 248–253 (2013)
Zou, B. et al. Lead optimization of imidazopyrazines: a new class of antimalarial with activity on Plasmodium liver stages. ACS Med. Chem. Lett. 5, 947–950 (2014)
Chatterjee, A. K. et al. Preparation of pyrazolopyridine compounds for the treatment of parasitic diseases. US patent WO 2014078802 A1 (2014)
Xiao, L. Molecular epidemiology of cryptosporidiosis: an update. Exp. Parasitol. 124, 80–89 (2010)
Zeeman, A. M. et al. PI4 kinase is a prophylactic but not radical curative target in Plasmodium vivax-type malaria parasites. Antimicrob. Agents Chemother. 60, 2858–2863 (2016)
Ghidelli-Disse, S. et al. Identification of Plasmodium PI4 kinase as target of MMV390048 by chemoproteomics. Malar. J. 13 (Suppl. 1), P38 (2014)
Huston, C. D. et al. A proposed target product profile and developmental cascade for new cryptosporidiosis treatments. PLoS Negl. Trop. Dis. 9, e0003987 (2015)
Manjunatha, U. H ., Chao, A. T ., Leong, F. J. & Diagana, T. T. Cryptosporidiosis drug discovery: opportunities and challenges. ACS Infect. Dis. 2, 530–537 (2016)
Branchini, B. R. et al. Red-emitting luciferases for bioluminescence reporter and imaging applications. Anal. Biochem. 396, 290–297 (2010)
Theodos, C. M., Griffiths, J. K., D’Onfro, J., Fairfield, A. & Tzipori, S. Efficacy of nitazoxanide against Cryptosporidium parvum in cell culture and in animal models. Antimicrob. Agents Chemother. 42, 1959–1965 (1998)
Zambriski, J. A. et al. Description of fecal shedding of Cryptosporidium parvum oocysts in experimentally challenged dairy calves. Parasitol. Res. 112, 1247–1254 (2013)
Zambriski, J. A. et al. Cryptosporidium parvum: determination of ID50 and the dose-response relationship in experimentally challenged dairy calves. Vet. Parasitol. 197, 104–112 (2013)
Bellosa, M. L. et al. A comparison of fecal percent dry matter and number of Cryptosporidium parvum oocysts shed to observational fecal consistency scoring in dairy calves. J. Parasitol. 97, 349–351 (2011)
Younis, Y. et al. 3,5-Diaryl-2-aminopyridines as a novel class of orally active antimalarials demonstrating single dose cure in mice and clinical candidate potential. J. Med. Chem. 55, 3479–3487 (2012)
Castellanos-Gonzalez, A. et al. Cryptosporidium infection of human intestinal epithelial cells increases expression of osteoprotegerin: a novel mechanism for evasion of host defenses. J. Infect. Dis. 197, 916–923 (2008)
Bessoff, K., Sateriale, A., Lee, K. K. & Huston, C. D. Drug repurposing screen reveals FDA-approved inhibitors of human HMG-CoA reductase and isoprenoid synthesis that block Cryptosporidium parvum growth. Antimicrob. Agents Chemother. 57, 1804–1814 (2013)
Sharling, L. et al. A screening pipeline for antiparasitic agents targeting cryptosporidium inosine monophosphate dehydrogenase. PLoS Negl. Trop. Dis. 4, e794 (2010)
Manjunatha, U. H. et al. Direct inhibitors of InhA are active against Mycobacterium tuberculosis. Sci. Transl. Med. 7, 269ra3 (2015)
Faller, B. et al. High-throughput in vitro profiling assays: lessons learnt from experiences at Novartis. Expert Opin. Drug Metab. Toxicol. 2, 823–833 (2006)
Faller, B. Artificial membrane assays to assess permeability. Curr. Drug Metab. 9, 886–892 (2008)
Ames, B. N., Lee, F. D. & Durston, W. E. An improved bacterial test system for the detection and classification of mutagens and carcinogens. Proc. Natl Acad. Sci. USA 70, 782–786 (1973)
Jothikumar, N., da Silva, A. J., Moura, I., Qvarnstrom, Y. & Hill, V. R. Detection and differentiation of Cryptosporidium hominis and Cryptosporidium parvum by dual TaqMan assays. J. Med. Microbiol. 57, 1099–1105 (2008)
Dreier, J., Störmer, M. & Kleesiek, K. Use of bacteriophage MS2 as an internal control in viral reverse transcription-PCR assays. J. Clin. Microbiol. 43, 4551–4557 (2005)
Operario, D. J., Bristol, L. S., Liotta, J., Nydam, D. V. & Houpt, E. R. Correlation between diarrhea severity and oocyst count via quantitative PCR or fluorescence microscopy in experimental cryptosporidiosis in calves. Am. J. Trop. Med. Hyg. 92, 45–49 (2015)
Acknowledgements
We thank S. Tzipori and D. Girouard for C. hominis oocysts; B. Nare for screening; B. H. Lee and J. Selva for high-content imaging data; M. Weaver for rat toxicology studies; I. Mueller for monkey pharmacokinetics; B. Yeung, O. Simon, J. Roland, V. Bollu, A. Chatterjee, A. Nagle, R. Moreau, and P. K. Mishra for compound synthesis; other Novartis Institutes for Biomedical Research (NIBR) colleagues for profiling; J. Burrows and K. Chibale for MMV390048; and M. Meissner for a plasmid carrying the red-shifted Fluc gene. This work was supported in part by the NIBR, the Wellcome Trust (Pathfinder 107678/Z/15/Z to B.S. and U.H.M.), and the National Institutes of Health (NIH R01AI112427 to B.S.). Inhibitors of the Plasmodium PI4K were discovered with the support of translational grants (WT078285 and WT096157) from the Wellcome Trust and funding from the Medicines for Malaria Venture (M.M.V.). B.S. is a Georgia Research Alliance Distinguished Investigator and A.S. is supported by NIH Fellowship F32AI124518. We thank our colleagues from Novartis Institute for Tropical Diseases, University of Georgia, Athens, Washington State University’s Office of the Campus Veterinarian, Animal Resource Unit, and the Office of Research Support and Operations and R. Anderson, 5D Dairy Farm, for their support. We are also grateful to the animal science and veterinary students at Washington State University for their participation in data collection and care of the research calves.
Author information
Authors and Affiliations
Contributions
U.H.M., S.V., J.A.Z., B.S., and T.T.D. conceived and designed the study; B.S. wrote grant applications with contributions from U.H.M.; U.H.M., A.T.C., and G.M.C.B. developed C. parvum screening assays; C.G.N. and S.H.L. developed enzyme assays; C.B. analysed P. falciparum EC50 data; U.H.M., P.G., and T.T.D. assembled the screening library; R.R.K. and B.Z. performed compound synthesis; U.H.M., B.Z., and J.W. analysed the structure–activity relationship; S.B.L. and F.B. analysed in vivo pharmacokinetics data; L.Z. optimized formulation; U.H.M., G.F., F.J.L., and T.T.D. analysed in vivo efficacy and toxicology results; S.V., A.S., and B.S. designed mouse models based on transgenic parasites; S.V., A.S., and J.T. constructed transgenic parasites; S.V., A.S., C.F.B. and G.T.H. validated mouse models; G.T.H., S.V., and C.F.B. tested compounds; J.A.Z. developed the calf model and analysed calf data; T.L.S. executed the calf model; S.N. conducted anatomic pathology reviews for efficacy and toxicity; L.B.G. developed and executed calf stool analytics; and B.S., S.V., U.H.M., and T.T.D. wrote the manuscript with contributions from J.A.Z., A.T.C., C.G.N., and S.B.L.
Corresponding authors
Ethics declarations
Competing interests
R.R.K. and B.Z. are named as inventors on a pyrazolopyridine patent application related to this work (WO 20140788002 A1). U.H.M. and T.T.D. are named as inventors on a pending cryptosporidiosis patent application related to this work. All Novartis Institute for Tropical Diseases-affiliated authors are employees of Novartis and some own shares in Novartis.
Additional information
Reviewer Information Nature thanks J. S. Doggett, N. S. Gray and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
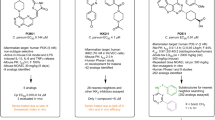
Extended Data Figure 1 Structures of the pyrazolopyridines and other known PI(4) kinase inhibitors.
Compounds described in Table 1. Important structural determinants required for anti-Cryptosporidium activity in pyrazolopyridines are shown in blue.
Extended Data Figure 2 Anti-Cryptosporidium activity does not correlate with mammalian cell toxicity.
Correlation of C. parvum cytopathic effect versus HepG2 cytotoxicity assay for selected pyrazolopyridine and imidazopyrazine analogues along with BQR695 and MMV390048. Data shown here are geometric mean EC50 values, with at least two biological replicates.
Extended Data Figure 3 Recombinant C. parvum cgd8_4500 shows phosphatidylinositide kinase activity.
C. parvum cgd8_4500 was expressed in insect cells using a Baculovirus system and recombinant enzyme was purified. A Michaelis–Menten plot of phosphatidylinositide kinase reaction with 3 nM CpPI(4)K enzyme at varying ATP concentrations is shown. Data shown here are a representative graph of two independent biological replicates.
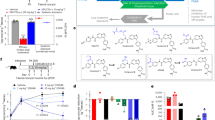
Extended Data Figure 4 KDU731 inhibits C. parvum Nluc parasites in vitro and in vivo.
a, EC50 determination of KDU731 against UGA1 Nluc transgenic parasites grown in HCT-8 cultures using luciferase activity as read out. Representative data are shown, three technical replicates. b, Mice (n = 5) were infected with 10,000 UGA1 Nluc oocysts and treated orally 3 days after infection with 1, 5, or 10 mg per kg (body weight) KDU731 for 1 week. Faecal oocyst load was determined by measuring parasite luciferase activity (b) or parasite DNA by qPCR (c) in faeces pooled from entire cage of five mice (20 mg faeces for Nluc and 100 mg for PCR assay). b, c, Means for three technical replicates are shown. Error bars, s.d. Pooled Nluc experiments for vehicle and 10 mg per kg (body weight) dose were repeated in three biological replicates and a representative result is shown.
Extended Data Figure 5 Parasite intestinal load measured by qPCR correlates with faecal shedding and tissue luminescence.
a, Mice (n = 4) were infected with 50,000 UGA2 FLuc oocysts and imaged after 1 week. Mice were killed and the small intestine was resected and imaged (representative image shown). Infection of the intestine ranged in intensity from heavy in the ileum to more moderate in the jejunum and caecum (see radiance scale bar for comparison). Intestines were cut into 12 segments and the luminescence of each segment was recorded. b, qPCR analysis of intestinal segments was performed in triplicate and plotted against the respective luminescence measurements. Regression analysis found robust correlation of tissue luminescence and PCR for parasite DNA, with r2 = 0.8. c, Mice were infected with 10,000 UGA2 FLuc oocysts, and 7 days after infection animals were treated daily for a week with vehicle or 10 mg per kg (body weight) KDU731. Whole-animal imaging during the treatment period is shown in Fig. 2g. Faecal oocyst load was determined by measuring parasite DNA by qPCR in faeces pooled from a cage of five mice. Error bars, s.d.
Extended Data Figure 6 Nitazoxanide does not reduce intestinal parasite load in IFN-γ knockout mice.
Mice (n = 5) were infected with 10,000 UGA2 FLuc oocysts, and 7 days after infection animals were treated daily for a week with 100 mg per kg (body weight) nitazoxanide or vehicle. Mice were monitored by whole-animal imaging. Radiance scale shows total flux in photons per second.
Extended Data Figure 7 Effect of KDU731 on severity of diarrhoea and dehydration in the neonatal calf model of cryptosporidiosis.
Severity of diarrhoea and dehydration in individual calves challenged with 5 × 107 C. parvum oocysts. Infected calves were treated with vehicle (n = 6) or with KDU731 (n = 7); n represents the number of calves. Every 12 h, calves were stimulated to defecate, faecal consistency was evaluated, and hydration status was assessed. Faecal consistency and hydration scores were assigned according to the study rubric (see Supplementary Information). The schematic representation shows the faecal consistency (a) and hydration scores (b) throughout the drug treatment period. Faecal consistency and hydration began to improve within 48 h of initiating treatment with KDU731.
Supplementary information
Supplementary Information (download PDF )
This file contains further details on assembly and screening of NITD parasite box and calf efficacy study and Supplementary Table 1. (PDF 336 kb)
Rights and permissions
About this article
Cite this article
Manjunatha, U., Vinayak, S., Zambriski, J. et al. A Cryptosporidium PI(4)K inhibitor is a drug candidate for cryptosporidiosis. Nature 546, 376–380 (2017). https://doi.org/10.1038/nature22337
Received:
Accepted:
Published:
Issue date:
DOI: https://doi.org/10.1038/nature22337
This article is cited by
-
A sustainable hydrotrope-assisted multicomponent synthesis of pyrazolopyridines
Research on Chemical Intermediates (2025)
-
Two pot and one pot synthesis of dipyrazolopyridines derivatives and applications
Discover Chemistry (2025)
-
In vitro molecular assessment of Cryptosporidium parvum parasitic load on human ileocecal adenocarcinoma cell culture after targeting by tavaborole (AN2690)
Journal of Parasitic Diseases (2025)
-
Identification of potent and orally efficacious phosphodiesterase inhibitors in Cryptosporidium parvum-infected immunocompromised male mice
Nature Communications (2024)
-
Cryptosporidium PI(4)K inhibitor EDI048 is a gut-restricted parasiticidal agent to treat paediatric enteric cryptosporidiosis
Nature Microbiology (2024)