Abstract
Epigenetic mechanisms are thought to contribute to neurodevelopmental vulnerability for psychiatric disorders, yet longitudinal evidence linking DNA methylation (DNAm) to brain maturation and psychopathology is limited. Using epigenome-wide DNAm (372,582 CpGs) and whole-brain structural MRI data from the IMAGEN cohort (n = 506, ages 14–19), we identified 18 co-regulated DNAm clusters, ten enriched for brain-expressed genes involved in neuronal development and signalling. The clusters showed consistent longitudinal change and were reproducible in independent adult samples (PPMI, n = 513; ADNI, n = 606). Multivariate analyses revealed coordinated coupling between DNAm change and cortical-subcortical maturation, such that greater DNAm reductions in brain-related clusters associated with greater cortical thinning and subcortical volume changes in the fronto-limbic-striatal axis. Increases in depressive symptoms, and frequency of cannabis use and binge drinking were linked to DNAm changes, with mediation models supporting DNAm as a mechanistic bridge between behaviour and brain change. Two clusters (C1 and C7) were associated with depressive and negative psychotic symptoms across adolescence. Associations of these clusters with depressive symptoms replicated in the PPMI dataset. Their coupling with amygdala–striatal maturation suggests that these DNAm signatures index environmentally shaped affective–motivational neurodevelopment.
Similar content being viewed by others
Introduction
Epigenetic modifications, particularly DNA methylation (DNAm) at cytosine-phosphate-guanine (CpG) dinucleotides, regulate gene expression and cellular identity across development and in response to environmental exposures [1]. Through this mechanism, genetic predispositions interact with developmental and environmental influences to shape trajectories of health and disease [2]. Although DNAm variation has been examined extensively in relation to disease states, less is known about its dynamic organisation during major neurodevelopmental transitions, such as adolescence, and how such changes relate to brain maturation and emerging mental health problems.
Blood-based DNAm has proven to be a useful proxy for investigating brain-relevant epigenetic processes. Despite tissue-specific differences, convergent studies have shown that subsets of CpG sites display correlated methylation patterns between blood and cortical brain regions [3], and that blood-based DNAm has been associated with MRI-derived brain structure in regions such as the hippocampus and cortical association areas [4,5,6]. However, studies investigating associations between DNAm and brain measures are still too scarce and lack systematic coverage, as most have been cross-sectional or region-specific, providing only limited insight into coordinated DNAm-brain associations at the whole-brain level.
Environmental exposures, including prenatal stress, early adversity, or substance use, can leave lasting DNAm signatures [7, 8], and DNAm variation has been associated with multiple psychiatric disorders, including ADHD, depression and schizophrenia [6, 9, 10]. Importantly, DNAm exhibits systematic age-related changes, especially in adolescence –a period characterised by greater susceptibility to stress, extensive brain maturation [11, 12], and heightened emergence of affective and substance-use disorders [13, 14]. Thus, adolescence may represent a critical window during which epigenetic processes influence neurodevelopmental vulnerability. Yet, the scarcity of longitudinal DNAm datasets has limited the identification of developmentally co-regulated DNAm patterns and their relevance for brain maturation and mental health.
Adolescence is also a developmental window during which depressive symptoms, psychotic-like experiences, and substance use frequently co-occur. Depression is one of the most common internalising conditions emerging during this period, and altered DNAm has been consistently linked to major depressive disorder [15]. Similarly, DNAm differences have been observed in individuals at clinical high risk for psychosis and in schizophrenia [16, 17]. Notably, depressive symptoms and negative psychosis-like symptoms show substantial phenomenological and mechanistic overlap [18, 19], which are closely tied to neural circuits involved in affect regulation and reward processing [20]. In parallel, cannabis use is a risk factor for both depressive and psychotic outcomes [21] and contributes to long-term alterations in adolescent brain maturation [22]. Together, these converging evidence justify joint examination of depression, psychosis-like symptoms, and substance use within a coherent neurodevelopmental framework grounded in DNAm plasticity and brain maturation.
Here, we address this by constructing a longitudinal DNAm atlas using IMAGEN [23], a deeply phenotyped adolescent cohort. Applying weighted gene co-expression network analysis (WGCNA) [24] on DNAm data acquired at ages 14 and 19, we identified clusters of CpG sites that show coordinated developmental patterns. We validated the stability of these clusters in two independent samples of older adults: PPMI [25] and ADNI [26]. We then examined whether longitudinal DNAm change in adolescence relates to structural brain maturation (changes in cortical thickness, surface area, and subcortical volumes) from ages 14 to 19, and whether these DNAm-brain patterns are associated with changes in depressive symptoms, psychosis symptoms, and frequency of substance use (alcohol, binge drinking, cigarette smoking, cannabis) across adolescence. An overview of the study design is provided in Fig. S1.
Results
Identification of longitudinal DNA methylation patterns
Our main analyses included data from IMAGEN, a longitudinal cohort of typically developing adolescents (n = 506, 47% males). As expected for this developmental period, the prevalence of risky behaviours (i.e., substance use) increased from ages 14 to 19 (Table S1).
To identify coordinated patterns of DNAm change across adolescence, we applied a two-step approach. First, WGCNA was applied to identify consensus modules –defined as consistently co-methylated at ages 14 and 19, which yielded 175 DNAm modules. For each module, we calculated longitudinal change in DNAm (ΔDNAm = age19 - age14) and then clustered modules based on similarity in these change patterns. This hierarchical clustering procedure identified 18 DNAm clusters (i.e., C1-C18) that showed consistent patterns of coordinated methylation change across adolescence, and spanning all 22 autosomal chromosomes (Fig. 1; Fig. S2; Table S2).
Correlation matrix of longitudinal DNAm change (ΔDNAm = DNAm19 - DNAm14) across 175 consensus co-methylation modules identified using WGCNA. Each cell reflects the Pearson correlation between longitudinal change scores of two modules. Hierarchical clustering of this matrix yielded 18 higher-order DNAm clusters (C1–C18), representing coordinated methylation trajectories across adolescence.
To evaluate whether these longitudinal DNAm patterns captured variance shared with DNAm measured at either age, we applied Mantel correlation [27] to compare similarity matrices (inter-cluster r-values) across timepoints. DNAm patterns at both age 14 and age 19 closely resembled the longitudinal covariation pattern (Mantel r = 0.94 at age 14; Mantel r = 0.81 at age 19; Fig. 2a). These results indicate that the identified clusters reflect stable, developmentally conserved DNAm organisation that is observable at each age as well as in their longitudinal change.
Similarity matrices (Pearson r) summarising inter-cluster DNAm relationships across cohorts. a IMAGEN, similarity matrices derived from ΔDNAm, DNAm14, and DNAm19 demonstrate stable cluster structure across time. b Replication in PPMI and (c) ADNI: similarity matrices computed in each cohort show significant correspondence to the IMAGEN reference pattern (Mantel correlations; 10,000 permutations; Bonferroni-corrected one-tailed P-values). SWEDD “scans without evidence of dopaminergic deficit”, MCI mild cognitive impairment.
Robustness of the DNAm pattern
Importantly, the identified DNAm clusters were technically robust, with consensus-based longitudinal clustering demonstrating significantly higher internal consistency across array platforms and batches than non-consensus approaches (see Methods and Fig. S3), supporting that the observed patterns reflect biological rather than methodological signal.
We further tested the robustness of the DNAm pattern in two independent datasets of older adults (PPMI and ADNI; Table S1). The cluster similarity structure identified in adolescence was preserved across diagnostic and healthy groups in both datasets: PPMI: Mantel r = 0.66 (Parkinson’s disease), r = 0.64 (Parkinson’s disease - SWEDD), r = 0.66 (healthy controls) (all Pperm < 0.0001; Fig. 2b); ADNI: Mantel r = 0.62 (Alzheimer’s disease), r = 0.65 (mild cognitive impairment), r = 0.62 (healthy controls) (all Pperm < 0.0001; Fig. 2c).
This cross-cohort indicates that the identified DNAm clusters represent a stable organisational architecture of the methylome that extends beyond adolescence into later life.
DNAm levels within clusters and associations with sex, pubertal development, and socioeconomic status
Table S3 summarises DNAm levels within each cluster at ages 14 and 19, the magnitude of developmental change, and associations with sex, pubertal development (PD), and socioeconomic stress (SES). Most clusters showed small-to-moderate decreases in DNAm from age 14 to 19, with the largest reductions observed in C13, C6, C7, and C11. Several clusters exhibited significant sex differences (PFDR < 0.05; |Cohen’s d| = 0.21–0.95). Notably, C1 and C2 showed higher methylation in males at both ages, with effects stronger at age 14. In contrast, C4 and C8 displayed a directional reversal across adolescence (C4: female>male at age 14, male>female at age 19; C8: male>female at age 14, female>male at age 19). Associations with SES were minimal: only C12 showed a significant association, with higher SES associated with lower DNAm (r = −0.15, PFDR = 0.0135). No DNAm cluster showed significant associations with pubertal development (Table S3).
Overall, adolescent DNAm variation was shaped primarily by sex and longitudinal developmental processes, with little evidence for systematic associations with socioeconomic factors.
Functional and tissue-specific characterisation of DNAm clusters
To infer the biological systems represented within each DNAm cluster, we performed tissue-specific gene expression enrichment (Table S4), Gene Ontology (GO), and KEGG pathway analyses (Table S5, Fig. 3). These analyses revealed marked functional differentiation across clusters, pointing to coordinated epigenetic regulation of neural, immune, and chromatin-remodelling processes during adolescence.
a–h Illustrations of the top ten significantly enriched items for KEGG pathways or GO terms in the clusters related to brain function (n = 8) (i.e., Clusters-3,4,5,6,7,8,9,11). FDR-corrected P-values were used for statistical significance. i Venn diagram displaying the numbers of brain-related DNAm clusters identified in the tissue-specific (n = 8) and functional (n = 8) enrichment analyses. A total of 10 brain-related DNAm clusters were identified, including six identified by both enrichment analyses and 4 identified by either one of the enrichment analyses. KEGG Kyoto encyclopedia of genes and genomes; GO gene ontology.
A subset of ten clusters (C1, C3, C4, C5, C6, C7, C8, C9, C11, and C12) showed convergent evidence for central nervous system relevance (Fig. 3i). Most of these clusters (C1, C3, C4, C5, C6, C7, C11, and C12) were enriched for cerebral cortex-expressed genes, suggesting that they index methylation-based coordination of neuronal structural and functional programmes. Within this group, C4, C6 and C11 were enriched for plasma membrane organisation, synaptic signalling, ion channel activity, and neuron projection, pointing to regulation of cell-cell communication, connectivity, and excitability. C5 was distinguished by strong enrichment for neurodevelopmental processes, including nervous system development, neurogenesis, and neuron differentiation, indicating a core axis of developmentally patterned methylation that aligns with brain maturation in adolescence. Other clusters within the brain-related group pointed toward cellular maintenance and metabolic regulation. C1 showed enrichment for ATP- and nucleotide-binding functions, consistent with neuronal energy mobilisation and biosynthetic regulation, whereas C3 and C9 were enriched for KEGG pathways related to neurodegeneration (Parkinson’s disease, Alzheimer’s disease, amyotrophic lateral sclerosis) and cell-cycle and senescence pathways, suggesting that these clusters may reflect molecular processes involved in long-term neuronal maintenance, vulnerability, or resilience. C8, while not enriched for cortex-specific expression, also showed enrichment for pathways related to neurodegeneration (amyotrophic lateral sclerosis) and for cell-cycle control, RNA splicing, ubiquitin-mediated turnover, and nucleocytoplasmic transport, pointing to regulation of proteostasis and nuclear-cytoplasmic communication. In contrast, C2 showed strong enrichment for peripheral immune tissues (spleen, lymph node, bone marrow) and for GO/KEGG pathways involving leukocyte activation, cytokine signalling, NF-κB and HIF-1 pathways, indicating a systemic immune signalling axis that is distinct from the brain-enriched clusters. Finally, C10 and C15 were enriched for chromatin regulation, RNA processing, and nuclear protein complex assembly, reflecting broad transcriptional regulatory machinery rather than region- or system-specific biology.
Together, these findings suggest that adolescent DNAm organisation reflects three coordinated molecular domains: (i) a neuronal and synaptic patterning axis (C1, C3, C4, C5, C6, C7, C8, C9, C11, C12), (ii) a peripheral immune regulatory axis (C2), and (iii) a global chromatin–transcriptional regulation axis (C10, C15). Because our central aim was to test how adolescent brain maturation relates to epigenetic variation, we focused subsequent analyses on the ten brain-related DNAm clusters (i.e., C1, C3, C4, C5, C6, C7, C8, C9, C11, C12) in the neuroimaging and psychopathology models presented below. These clusters were characterised by their differing overall DNAm levels and consistent patterns of change across adolescence (Table S3).
Developmental changes in cortical and subcortical morphology from ages 14 to 19
To relate the identified DNA methylation clusters to adolescent brain maturation, we first investigated changes in cortical and subcortical structure from ages 14 to 19 (i.e., Δbrain = brain19 - brain14). We observed widespread and regionally patterned changes. At the cortical level, the majority of brain regions showed significant reductions in cortical thickness (CT) over time (Table S6). The largest decreases were observed in bilateral fronto-parietal association cortices, including the rostral middle frontal, superior and inferior parietal, supramarginal, precuneus, and superior frontal regions (Cohen’s d = −0.7 to −1.0, PFDR < 0.001). Additional cortical thinning was evident in the cingulate cortex and sensorimotor regions, though with comparatively smaller effect sizes (Cohen’s d = −0.3 to −0.6). In contrast, a small subset of temporal and paralimbic regions exhibited relative preservation or increases in cortical thickness across the same period. These included the bilateral entorhinal cortex and temporal pole, as well as the left inferior temporal cortex (Cohen’s d = 0.23 to 0.39, all PFDR < 0.01). Fusiform and middle temporal regions showed no significant change. This pattern is consistent with a non-uniform maturational trajectory, with higher-order association cortices and comparatively later structural development in temporal regions [28, 29].
Subcortical grey matter volume (SubCortVol) changes were more regionally specific and of smaller magnitude. Significant reductions were observed in the bilateral caudate and putamen (all, PFDR < 0.001), and thalamus (both sides, PFDR < 0.05). A decrease was also observed in the left nucleus accumbens (PFDR = 0.0039), with a trend in the same direction on the right. In contrast, bilateral amygdala volumes increased modestly, whereas hippocampal and pallidal volumes remained stable. Bilateral lateral ventricles increased in size (both PFDR < 0.05), consistent with the observed cortical reductions [30, 31].
Taken together, these results indicate broad cortical thinning concentrated in fronto-parietal association networks, accompanied by targeted volumetric reductions in striatal and thalamic structures and selective increases in amygdala and anterior temporal cortices across adolescence. The overall pattern reflects continued structural refinement of networks supporting cognitive control, reward processing, and socio-emotional function during the transition from mid- to late adolescence [13].
Associations between changes in DNAm and brain maturation
We next investigated relationships between changes in DNAm from ages 14 to 19. All variables were entered as change scores (Δ = age19 – age14), such that more negative Δvalues reflect greater age-related reduction (e.g., cortical thinning or relative demethylation), whereas more positive values reflect relative preservation or expansion. In this framework, positive canonical loadings indicate DNAm clusters or brain regions where individuals showing relative preservation load more strongly on the CCA component, and negative loadings indicate clusters or regions where individuals showing greater age-related reduction contribute more strongly.
We ran three separate CCAs examining the association between changes in DNAm and changes in (i) cortical surface area (ΔSA), (ii) cortical thickness (ΔCT), and subcortical volumes (ΔSubCortVol) (Fig. 4a). Significant multivariate associations were found for ΔSA, ΔCT, and ΔSubCortVol (Fig. 4b–d). Statistical significances were observed for the first canonical components for the ΔCT (r = 0.56, Pperm = 0.030, Fig. 4e) and ΔSubCortVol (r = 0.41, Pperm = 0.004, Fig. 4f) analyses, but not for ΔSA (Pperm = 0.089). We therefore focused on the interpretation of the first canonical components of the ΔCT and ΔSubCortVol analyses (Tables S7–S8; Fig. 4g, h).
a Schematic overview of the canonical correlation analysis (CCA) linking ΔDNAm in the 10 brain-related clusters to longitudinal brain structural change (ΔMRI: Δcortical thickness (CT), Δcortical surface area (SA), Δsubcortical volume). b–d Permutation-derived η² distributions (10,000 permutations) for each CCA model. e, f Canonical correlations for the first significant component. g, h Brain maps displaying regional loadings (CCA weights) for Δcortical thickness and Δsubcortical volume components; FDR-corrected P < 0.05. Warmer colours indicate regions that load positively, cooler colours indicate regions that load negatively.
Cortical thickness
For ΔCT, seven DNAm clusters contributed significantly to the canonical component (|r| ≥ 0.12, PFDR < 0.05). The most influential were C12 (r = −0.51) and C6 (r = 0.44), with additional contributions from C3, C7, C9 and C11 (r ≥ 0.12), and C5 (r = −0.18). C12 -which showed a moderate decrease in methylation (Table S3)– loaded negatively, indicating that greater decreases in methylation in this cluster were associated with greater cortical thinning. In contrast, C6, which also decreased substantially in methylation, loaded positively, indicating that individuals with more preserved methylation levels in this cluster showed less cortical thinning. On the brain side, positive loadings (r ≥ 0.11) were observed in medial and lateral prefrontal (e.g., lateral orbitofrontal cortex, medial orbitofrontal cortex, pars triangularis, pars opercularis, and rostral/caudal middle frontal cortex) and cingulate regions. In contrast, negative loadings (r ≤ −0.12) were observed in temporal and parietal-association regions, including the entorhinal cortex and banks of the superior temporal sulcus.
Thus, the cortical CCA identifies two coordinated maturational profiles: (1) a profile of greater DNA methylation decrease (e.g., C12 and C5) co-occurring with greater thinning in temporal-paralimbic regions, and (2) a relative preservation profile linking less DNAm decrease (in C6, C3, C7, C9 and C11) with more stable cortical thickness in medial and lateral prefrontal regions.
Subcortical Volumes
For ΔSubCortVol, seven DNAm clusters again contributed significantly (all |r| ≥ 0.11, PFDR < 0.05). Key contributors included C1 (r = 0.44), Cluster 7 (r = −0.36), C11 (r = 0.23), and C12 (r = 0.40). Here, greater DNAm decrease in negatively loaded clusters (C7 and C6) were associated with greater volume reductions in the bilateral amygdala, left nucleus accumbens and right thalamus, hippocampus and putamen, while relative maintenance of methylation in C1, C11, C12 and C8 was associated with relative volume preservation in the bilateral caudate, and right nucleus accumbens, as well as reduced ventricular enlargement.
Longitudinal associations between DNAm change, cortical maturation, and adolescent psychopathology
The fronto-limbic-striatal axis highlighted above –encompassing prefrontal regions supporting cognitive control and valuation (lateral and medial orbitofrontal cortices), temporal-paralimbic regions involved in socio-emotional processing (entorhinal cortex, superior temporal sulcus), and striatal structures involved in reward learning and motivation– is central to affect regulation and reward sensitivity. Regions in this network have been repeatedly implicated in risk for depression and substance-use escalation, particularly in adolescence [14, 32,33,34].
Descriptive data (Table S1) show significant increases in lifetime substance use from ages 14 to 19 (all P < 0.0001). Accordingly, we next examined whether changes in depressive symptoms and lifetime frequency of substance use across adolescence were associated with the coordinated patterns of DNAm and cortical and subcortical changes identified in the first CCA components (Table S9). Significant associations were observed with the DNAm and cortical changes captured by the first ΔCT CCA component: Increased frequency in cannabis use and binge drinking were associated with greater DNAm reduction (r = −0.16) and greater cortical thinning (r < −0.10). Increases in depressive symptoms were also associated with greater DNAm reduction (r = −0.12) and greater cortical thinning (r = −0.14) (all PFDR < 0.05). In contrast, changes in alcohol use frequency and tobacco smoking showed non-significant associations. No significant associations were observed for subcortical changes identified by the ΔSubCortVol CCA. Taken together, these results suggest that adolescents who show greater increases in depressive symptoms, cannabis use, and binge drinking also exhibit greater reductions in DNAm and greater cortical thinning within a shared fronto-limbic maturation axis.
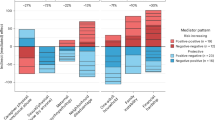
We used longitudinal mediation models to test whether DNAm change mediates the link between behavioural changes and cortical maturation, examining directional pathways among changes in DNAm, depressive symptoms, cannabis use, binge drinking and cortical thickness. Given the known effects of substances of abuse (e.g., tobacco, cannabis and alcohol) both on DNA methylation [35] and adolescent brain maturation [22, 36, 37], we tested mediation models where Δsubstance use predicted ΔCT through ΔDNAm. Both Δcannabis use and Δbinge drinking showed a significant indirect association with ΔCT via ΔDNAm (Δcannabis use: indirect = −0.04, P = 0.0002; Δbinge drinking: indirect = −0.05, P = 0.003; Fig. 5a), such that increases in cannabis use/binge drinking were associated with greater DNAm reduction, which in turn related to greater cortical thinning. The direct effects were non-significant (Δcannabis use: c′ = −0.03, P = 0.113; Δbinge drinking: c′ = −0.02, P = 0.383), indicating complete mediation.
a ΔSubstance use → ΔDNAm → ΔCortical Thickness: Indirect effects shown separately for cannabis use (left) and binge drinking (right), indicating full mediation. b Bidirectional models involving depressive symptoms: Left, ΔDepressive symptoms → ΔDNAm → ΔCortical Thickness; Right, ΔDNAm → ΔDepressive symptoms → ΔCortical Thickness. Path coefficients and P-values are shown. c, d Associations between DNAm levels in brain-related clusters and substance use at ages 14 and 19 (two-tailed PFDR < 0.05). e Significant associations of clusters C1 and C7 with depressive symptoms and negative psychotic symptom dimensions across adolescence (Pearson r; FDR-corrected two-tailed P-values). f Replication of C1 and C7 associations with depressive symptoms in PPMI (Pearson r; one-tailed P-values). g Associations of C1 and C7 with depressive symptoms and negative psychotic symptoms in IMAGEN after adjusting for depression or schizophrenia PRS, respectively (Pearson r; one-tailed P-values).
In a similar model in which Δdepressive symptoms predicted ΔCT via ΔDNAm (Fig. 5b), we again observed a significant indirect effect (indirect = −0.05, P = 0.011), such that increases in depressive symptoms predicted greater DNAm reduction (P = 0.0099), which was in turn associated with greater cortical thinning. Here, the direct effect of Δdepressive symptoms on ΔCT was attenuated and non-significant (c′ = −0.06, P = 0.062), consistent with full mediation. In the reverse model where ΔDNAm predicted ΔCT through Δdepressive symptoms, we observed a significant indirect effect (indirect = 0.01, P = 0.044), indicating that adolescents showing greater DNAm reductions tended to report increases in depressive symptoms, which in turn were associated with greater cortical thinning. The direct association between ΔDNAm and ΔCT remained strong (c′ = 0.54, P = 0.0004), suggesting partial mediation.
Together, these models suggest that DNAm changes serve as a biological pathway linking affective distress and cannabis-use or binge drinking escalation to accelerated fronto-limbic cortical maturation, consistent with frameworks in which stress- and reward-related molecular mechanisms shape neurodevelopmental risk trajectories.
Associations between cluster-defined DNAm patterns and psychopathology
Having identified coordinated DNAm patterns linking cortical maturation to psychopathology, we next examined cluster-specific DNAm signatures associated with substance use and depressive symptoms at ages 14 and 19.
Associations with substance use behaviours
As shown in Table S10 and Fig. 5c, d, associations with substance use differed across clusters and ages. At age 14, the only significant correlations (all PFDR < 0.05) were positive associations between C4 and C9 and binge drinking. By age 19, more associations emerged, specifically with C5, C6, C9, C11, and C12. Notably, C6 and C11, which showed an overall decrease in methylation from ages 14 to 19, were negatively associated with smoking and cannabis use (Table S10); associations with cannabis use, remaining significant even after controlling for smoking. C6 was also negatively associated with binge drinking. This indicates that lower methylation levels in these clusters were linked to greater substance use involvement. In contrast, C12 showed positive associations with smoking and cannabis use. None of the smoking-related associations remained significant after controlling for cannabis use, reflecting unique effects of cannabis exposure to DNAm in these clusters. C9 was the only cluster associated with all substance-related behaviours, although none of the age 19 associations remained significant after controlling for smoking or cannabis use, suggesting a distinct molecular profile related to polysubstance exposure.
Associations with mental health symptoms
Two clusters displayed significant associations with depressive symptoms (PFDR < 0.05; Fig. 5e; Table S10). C7 demonstrated a negative association with depressive symptoms at both ages, while C1 was positively associated with symptoms at age 19. Given the potential shared aetiology between depressive symptoms and negative psychotic experiences [18, 19], we also tested associations between ten brain-related DNAm clusters with dimensions of positive, negative and depressive psychotic symptoms at age 19. Significant associations were found for C7, which specifically associated with negative (r = −0.14, PFDR = 0.031), but not depressive or positive psychosis symptoms (PFDR > 0.05).
We next tested whether the associations between C1 and C7 and depressive symptoms replicated in the PPMI cohort. Both clusters again showed significant associations with depressive symptoms in this independent, older adult sample (Pone-tail < 0.005, Fig. 5f), demonstrating that these DNAm-symptom relationships are robust across cohorts, developmental stages, and clinical contexts. This cross-cohort replication suggests that C1 and C7 capture stable, developmentally persistent epigenetic variation relevant to depressive phenotype expression from adolescence through older adulthood.
Genetic and epigenetic liability to depression and psychosis
To investigate the respective contributions of epigenetic and genetic factors to the risk for depression and psychosis, we complemented the above analyses with analyses of polygenic risk score (PRS) in IMAGEN. As expected, depression PRS showed significant (PBonferroni,one-tailed < 0.05) positive associations with depressive symptoms at both ages (age 14: r = 0.09, 95% CI [0.03, +∞), Power = 0.68; age 19: r = 0.16, 95% CI [0.09, +∞), Power = 0.97). Similarly, schizophrenia PRS was significantly associated with positive (r = 0.12, 95% CI [0.05, +∞), Power = 0.86), negative (r = 0.09, 95% CI [0.01, +∞), Power = 0.61), and depressive (r = 0.18, 95% CI [0.11, +∞), Power = 0.99) psychosis symptom dimensions at age 19.
We next tested whether the associations between C1 and C7 and mental health symptoms, as identified in the section above, remained significant when controlling for genetic effects (i.e., controlling the depression PRS or schizophrenia PRS in analyses of depressive and psychosis symptoms, respectively), and did observe so (Fig. 5g). This supports a model in which epigenetics (i.e., DNAm) and genetics represent distinct risk factors for depression and psychosis.
Discussion
In this longitudinal study, we identified coordinated patterns of DNA methylation change during adolescence and demonstrated their relevance for brain maturation and mental health symptoms. Using data-driven clustering of epigenome-wide DNAm across ages 14 and 19, we derived an atlas of 18 developmentally co-regulated DNAm clusters. These clusters were robustly replicated across six independent samples of older adults, indicating that the developmental coordination captured in adolescence reflects stable organisational structure in the human methylome rather than cohort-specific effects. The atlas therefore provides a foundational resource for studying how epigenetic patterning unfolds across the lifespan.
Although prior DNAm atlas studies have been predominantly cross-sectional [38,39,40], our longitudinal design allowed us to characterise developmental trajectories of methylation change during adolescence, a neurodevelopmental window marked by pronounced cortical and subcortical reorganisation [11,12,13]. Crucially, we found that the magnitude and direction of DNAm change in specific brain-related clusters closely tracked the pace of cortical and subcortical maturation, particularly within prefrontal-limbic-striatal circuits supporting emotion regulation, reward learning, and stress sensitivity [14, 32, 34].
Several clusters enriched for genes expressed in the cortex and neurodevelopmental signalling pathways –including C3, C5, C6, C7, C9, C11 and C12– showed the strongest associations with longitudinal cortical thickness change. These clusters mapped onto two coordinated maturational profiles. First, greater DNAm reduction in clusters such as C12 and C5 (the latter enriched neuron differentiation pathways) was associated with greater cortical thinning in temporal and parietal association regions (e.g., entorhinal cortex, banks of the superior temporal sulcus). Given that DNA methylation typically represses transcription, this pattern is consistent with developmental derepression of neuronal differentiation and plasticity-related genes as paralimbic association cortices refine sensory-affective integration during adolescence [41,42,43]. Conversely, relative preservation of DNAm in clusters C6, C3, C7, C9 and C11 was linked to relative maintenance of cortical thickness in medial and lateral prefrontal regions (lateral/medial orbitofrontal cortex, pars opercularis, pars triangularis, and rostral and caudal middle frontal cortex) –regions central to valuation, cognitive control, and affective regulation [44, 45].
In addition to cortical changes, DNAm variation was also linked to subcortical maturation. Greater demethylation in clusters such as C7 and C6 was associated with more pronounced volume reductions in the amygdala, nucleus accumbens, thalamus, hippocampus and putamen, whereas relative maintenance of methylation in C1, C12, C11 and C8 was associated with relative volume preservation in the caudate and nucleus accumbens and attenuated lateral ventricular enlargement. Together, these findings indicate that the degree of methylation change in specific DNAm clusters systematically covaries with the tempo of structural brain maturation. The association between DNAm change and lateral ventricular enlargement likely reflects individual variation in the pace of normative neurodevelopmental tissue remodelling, as ventricular expansion in childhood and adolescence is a well-established marker of ongoing cortical-subcortical maturation [46], and later in life, of neurostructural vulnerability [47, 48]. Thus, the degree of DNAm change in these clusters may index individual differences in the tempo of fronto-limbic–striatal structural refinement, wherein greater demethylation corresponds to a faster maturational trajectory, while more stable methylation corresponds to relative structural preservation.
The mediation analyses further suggest a mechanistic role for DNAm in linking behavioural and affective experiences to neurodevelopment. In the case of substance use, increases in cannabis use and binge drinking predicted greater reductions in DNAm, which in turn were associated with greater cortical thinning, yielding significant indirect effects in the absence of direct behavioural-brain associations, consistent with a unidirectional, fully mediated pathway from substance use to brain maturation via DNAm change. In contrast, depressive symptoms showed evidence of bidirectional coupling with DNAm: increases in depressive symptoms predicted greater DNAm reduction, which subsequently related to greater cortical thinning, while DNAm reductions also predicted increases in depressive symptoms, partially accounting for additional variance in cortical change. Together, these findings support a model in which substance use drives neurodevelopmental change through epigenetic mechanisms, whereas affective symptom change and DNAm influence each other reciprocally, consistent with feedback regulation in stress- and reward-related molecular pathways [49].
Our cluster-specific analyses extend prior work linking DNAm to adolescent substance use [8, 50] and reveal both shared and substance-specific epigenetic signatures. C9 was associated with all substance-use behaviours at age 19, suggesting a general liability mechanism, whereas other clusters, such as C11 and C12 showed stronger specificity for cannabis or smoking. The neurobiological interpretation of these findings is strengthened by the observation that DNAm changes in these clusters relate to maturation of the prefrontal cortex, amygdala, caudate, and nucleus accumbens –regions central to reward learning and motivational salience [51,52,53].
Aligning with evidence of disrupted DNAm in schizophrenia [54,55,56] and depression [57, 58], two clusters, C1 and C7, showed consistent associations with depressive symptoms in both adolescence and older adulthood, including replication in PPMI. C7 was additionally associated with negative psychotic symptomatology, suggesting an epigenetic signature that may index transdiagnostic affective-blunting phenotypes [56] across disorders. The persistence of these associations following adjustment for polygenic risk scores supports models in which epigenetic variation constitutes an environmentally sensitive pathway, partly independent of inherited liability.
That DNA methylation variation within C1 and C7 tracked with the maturation of the amygdala and striatal nuclei is consistent with extensive evidence that these regions are highly plastic, environmentally responsive neural hubs involved in emotion regulation, stress responsivity, and reward processing [59]. Notably, C7, which loads on OFC and ACC and on amygdala/hippocampus, also shows negative associations with depressive symptoms (and with negative psychosis-like symptoms), aligning with cortical and subcortical loci repeatedly implicated in MDD [60, 61] and with OFC/ACC-centred depression subtypes [62]. The amygdala-prefrontal system is particularly sensitive to early life and social adversity, with epigenetic mechanisms proposed to mediate long-term effects on synaptic plasticity and circuit reactivity [63].
Taken together, these findings suggest that coordinated DNAm-brain maturation patterns index individual variation in the developmental tuning of affective and reward-circuitry systems. Clusters C1 and C7, in particular, capture stable, developmentally persistent epigenetic signatures that track with amygdala-striatal maturation and emerging depressive and negative-symptom phenotypes. Their reproducibility across cohorts and independence from genetic liability indicate potential utility as biomarkers capable of capturing environmentally shaped risk trajectories, informing stratification and early intervention strategies.
A key strength of this study is the integration of longitudinal DNA methylation, neuroimaging, and behavioural data within the same individuals across a developmentally critical window. The IMAGEN cohort provides a relatively large sample with repeated blood-based DNAm profiling and whole-brain MRI, alongside repeated assessment of depressive symptoms and substance use. This unique multimodal, longitudinal design enabled us to delineate coordinated trajectories of methylation change, cortical-subcortical maturation, and emerging psychopathology, offering rare insight into the dynamic coupling between molecular and neural development during adolescence. Nonetheless, several limitations warrant consideration. Although our longitudinal mediation analyses support a model in which substance-use and depressive symptoms escalation are statistically linked to DNAm change and cortical maturation, these associations cannot establish causality. The observed mediation effects suggest potential mechanistic pathways through which behavioural experiences may relate to neurodevelopmental trajectories, but the directionality of these processes may also reflect reverse causation or shared environmental influences. Effect sizes linking DNAm to behavioural symptoms were modest, in line with the polygenic and multifactorial nature of psychiatric traits. PRS-symptom associations were similarly modest in magnitude and should be interpreted as indexing broad liability rather than individual-level predictive accuracy. Replication in clinical or high-risk cohorts will be essential to determine sensitivity and predictive utility. Our metric of DNAm change reflects absolute β-value differences rather than proportional change, and while biologically interpretable, alternative representations may yield complementary information. Cannabis use was relatively uncommon at age 14; therefore early-adolescent associations should be interpreted cautiously, as non-significant findings at this age may reflect limited statistical power rather than absence of underlying effects. Finally, translation will be strengthened by future studies incorporating richer clinical phenotyping, higher temporal resolution, and experimental or interventional designs to further test mechanistic pathways.
In conclusion, this study presents the first longitudinal, whole-brain DNA methylation atlas of adolescence, revealing coordinated methylation-brain-behaviour coupling that links environmental experience to neurodevelopmental risk. By systematically integrating epigenetic, neuroimaging, and behavioural trajectories, it identifies reproducible DNAm signatures as potential biomarkers for early detection and prevention of affective and substance-use disorders in youth.
Materials and methods
Participants
IMAGEN cohort
Primary analyses were conducted in the IMAGEN study, a longitudinal population-based cohort followed from early to late adolescence [23]. The present study included 506 participants with DNAm and behavioural data available at both ages 14 and 19 (Table S1). Self-reported sex was used, reflecting biological assignment at birth. A detailed participant flow diagram for each analytical step is provided in Fig. S4.
Replication cohorts
To evaluate whether the longitudinal DNAm pattern identified in adolescence generalises across the lifespan, we examined two independent adult cohorts: (i) the Parkinson’s Progression Markers Initiative (PPMI) cohort [25] (n = 513; 32.9% female; mean age = 61.57 ± 10.04 years), consisting of individuals with Parkinson’s disease (PD, n = 329), PD and scans without evidence for dopaminergic deficit (SWEDD, n = 53), or neurologically healthy controls (n = 131) (Table S1), and (ii) the Alzheimer’s Disease Neuroimaging Initiative (ADNI) cohort (n = 606; 44.1% female; mean age = 73.16 ± 7.07 years), including participants with Alzheimer’s disease (AD, n = 72), mild cognitive impairment (MCI, n = 329), or healthy controls (n = 205) (Table S1). These datasets were used to test the robustness and reproducibility of the DNAm clustering solution.
DNA methylation preprocessing
Genome-wide DNAm was assayed in peripheral whole blood using Illumina methylation microarrays (450 K and EPIC-850K) [5, 64], according to the manufacturer’s protocols. DNAm in IMAGEN was conducted in four processing waves: ages 14 DNAm included three waves (n = 238 Wave 1; n = 207 Wave 2; n = 61 Wave 4), and ages 19 DNAm included two (n = 325 Wave 3; n = 181 Wave 4). Waves 1 & 2 were processed with the 450 K, and waves 3 & 4 with the 850 K array. Participant allocation to each wave was randomised to avoid introducing systematic bias. For a subset of individuals (n = 61), DNAm at ages 14 and 19 was processed on the same platform (850 K array) within a single wave (wave 4) to specifically evaluate the impact of technical confounders such as platform and batch effects.
Preprocessing was performed using the Bioconductor package “minfi” (version: 1.46.0) [65]. To ensure comparability across arrays, we restricted analyses to the 372,582 CpG sites present on both the 450 K and 850 K platforms. Quantile normalisation (“preprocessQuantile”) was applied to IMAGEN, and Illumina normalisation (“preprocessIllumina”) to PPMI and ADNI, to accommodate case–control global methylation differences.
For IMAGEN DNAm analyses, we regressed out technical confounders (recruitment sites and waves) before conducting the WGCNA, hierarchical clustering, and pattern similarity analyses. All other DNAm analyses in IMAGEN were adjusted for sex, recruitment site, cell-type composition (first two PCs, capturing most of the variability [5, 6]), the first four DNAm PCs (capturing major technical and biological variance) [6], the first four genotype PCs (population stratification), and wave. For PPMI, we controlled for age, sex, race, education, recruitment site, cell types, and the first four DNAm PCs in analyses of depressive symptoms.
Clustering analysis of DNAm data
A two-stage dimensionality reduction procedure was used for the DNAm data (workflow in Fig. S5).
Step 1: Consensus WGCNA modules
To minimise the impacts of technical confounders (i.e., platforms and waves) while retaining retained biological differences (i.e., changes related to age), consensus Weighted gene co-expression network analysis (WGCNA) [24] (R package “WGCNA”, version 1.72-5) was applied to DNAm matrices at ages 14 and 19 (506 ×372,582 CpGs for each timepoint), using the default and recommended settings. For adjacency matrices, soft-thresholding (β = 5) was selected using scale-free topology criteria for both age 14 and age 19 networks (Fig. S6). This resulted in the identification of 175 consensus modules, defined as sets of CpGs identified as co-methylated at both age 14 and age 19. We excluded CpGs assigned to Module 0 (i.e., unclustered CpGs) because such CpGs lack coherent co-methylation structure and therefore do not support biologically interpretable network-based inference [24, 66].
Step 2: Hierarchical clustering of longitudinal DNAm change
For each of the 175 modules, we computed the mean change in DNAm (ΔDNAm = age19 - age14). Hierarchical clustering of these ΔDNAm values was conducted using the R package “flashClust” (version: 1.01-2), using default settings. Euclidean distance [67] was used to quantify how dissimilar these ΔDNAm values were (i.e., how differently DNAm modules changed from ages 14 to 19), and average linkage to iteratively merge modules into higher-order clusters based on their mean pairwise distance, thresholding the number of clusters to 18 (i.e., 10% of 175 modules). These 18 higher-order clusters therefore capture shared developmental trajectories. Cluster-level DNAm scores (506 × 18 matrix) were computed as the mean DNAm across modules within each cluster. Batch effects (recruitment site, wave) were used as covariates during both the WGCNA and clustering steps.
Robustness of the DNAm pattern
To assess the robustness of the identified DNAm pattern, we conducted a series of internal and external validation analyses. First, we compared our consensus-based clustering approach (modules required to be present at both ages 14 and 19) with a non-consensus approach in which WGCNA was applied directly to ΔDNAm values (Fig. S7). The non-consensus approach produced 190 modules; however, when we compared the similarity matrices of longitudinal change across subsets of the IMAGEN cohort, the consensus approach demonstrated significantly greater internal stability. Specifically, correlation matrices derived from the full cohort (450 K → 850 K data) and the subset of 61 adolescents measured on the same platform and wave (850 K → 850 K) showed higher similarity for the consensus structure than for the non-consensus structure (rconsensus = 0.86 vs. rnon-consensus = 0.79; Zdiff = 2.24; Ptwo-tailed = 0.025; Fig. S3), indicating that consensus-based clustering is more robust to technical heterogeneity.
To verify that longitudinal DNAm change within each module reflects coordinated biological signal rather than noise, we calculated the variance explained by the first principal component of CpG methylation change within each module. Across 175 modules, the first principal component accounted for an average of 43% of variance (95%CI: 0.40–0.45), demonstrating a strong, systematic pattern of co-methylation within modules.
External robustness and generalizability of the 18-cluster DNAm pattern were assessed in independent datasets (PPMI and ADNI). For each cohort, we computed cluster-wise similarity matrices (cluster × cluster Pearson correlations) and compared these to the IMAGEN longitudinal similarity matrix using the Mantel test (R package vegan, v2.6-10). Statistical significance was determined using 10,000 permutations (PFDR < 0.05, FDR corrected). Significant Mantel correlations indicated that the inter-cluster co-variation structure was preserved across cohorts. Together, these analyses demonstrate that the 18-cluster DNAm architecture reflects biologically meaningful longitudinal co-variation rather than technical artefact.
Functional and tissue-specific Enrichment analyses
Gene Ontology (GO) analysis identifies biological processes and molecular functions that are statistically overrepresented within a given gene set, while Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis maps these genes to known biological pathways to infer potential functional mechanisms. Using the gometh function in the missMethyl R package (v1.34.0), we performed GO and KEGG enrichment for each DNAm cluster, accounting for probe-number bias. Genes were assigned to CpG sites based on proximity to the nearest transcription start site using the getNearestTSS function. Significance was determined using FDR correction (PFDR < 0.05).
To assess tissue relevance, we applied the teEnrichment function of the R package“TissueEnrich” (v1.20.0) to test whether genes in each cluster were disproportionately expressed in specific tissues. Clusters were defined as brain-related if they showed (i) significantly enriched expression in brain tissues (PFDR < 0.05) or (ii) major function (top ten) enriched for neurodevelopmental, synaptic, neuronal signalling, or neurodegeneration-related pathways (PFDR < 0.05). These clusters were subsequently used in brain–epigenome and psychopathology analyses.
Structural MRI processing
Structural MRI data in IMAGEN were acquired at eight sites using 3 T scanners from Siemens, Philips, General Electric, and Bruker, with harmonised acquisition protocols as previously described [23]. T1-weighted images were processed using the standard FreeSurfer pipeline (version 6.0.0), including intensity normalisation, skull stripping, surface reconstruction, and cortical parcellation [68]. Cortical thickness (CT), cortical surface area (SA), and regional grey matter volumes were derived from the Desikan–Killiany atlas (68 cortical parcels) and subcortical volumes were extracted using FreeSurfer’s aseg segmentation (16 subcortical regions) [69].
Quality control involved exclusion of regional measurements exceeding ±5 SD from the sample mean. Final analytic samples included n = 465 for SA, n = 463 for CT, and n = 443 for subcortical volume measures. Total intracranial volume (ICV), estimated via aseg, was included as a covariate in all structural analyses to account for global brain size differences.
Phenotypic assessments
Substance use behaviours
Substance use was assessed using the European School Survey Project on Alcohol and Other Drugs (ESPAD; http://espad.org), a standardised and validated questionnaire widely used in European adolescent cohorts. Four substance use indicators were examined: cigarette smoking, cannabis use, alcohol use, and binge drinking. Participants reported the number of occasions they had engaged in each behaviour over their lifetime using integer response scales. Specifically, smoking was assessed with the item “On how many occasions during your lifetime have you smoked cigarettes?”; cannabis use was assessed as the number of occasions of using “cannabis (grass, pot) or hashish”; alcohol use as the number of occasions of consuming “any alcoholic beverage”; and binge drinking as the number of occasions of “having five or more drinks in a row” during the lifetime. Prevalence rates of each behaviour at ages 14 and 19 are provided in Table S1.
Socioeconomic stress (SES)
Parents’ reports of socioeconomic stress were measured using the family stress scale, as part of the Development and Well‐Being Assessment (DAWBA) [70]. A total score, which measures the degree to which unemployment, financial difficulties, home inadequacy, and neighbourhood problems made family life stressful, was used. High scores indicate more socioeconomic stress in the family. Only data at age 14 years was available for this measure.
Pubertal development (PD)
This was assessed at age 14 in IMAGEN using the Puberty Development Scale (PDS) [71], which provides a self-report measure of physical development based on Tanner stages. Total scores were calculated separately for males and from five characteristics: growth spurt in height, pubic hair, and skin change for both boys and girls; facial hair growth and voice change in boys only; and breast development and menarche in girls only.
Psychiatric symptoms
Depressive symptoms
In IMAGEN, depressive symptoms at age 14 were assessed using the average score of self- and parents-reports from the development and well-being assessment (DAWBA) [70]. At age 19, depressive symptoms were assessed using the Adolescent Depression Rating Scale (ADRS), a 10-item self-report scale specifically developed to evaluate adolescent depression [72]. We used total scores, with higher scores reflecting higher symptom severity. To assess changes in depressive symptoms in longitudinal analyses, scores at each age were standardised (z = (X–mean)/SD), and longitudinal change was calculated as Δz (i.e., zage19−zage14).
In PPMI, depressive symptoms were assessed using the MDS-Unified Parkinson’s Disease Rating Scale (MDS-UPDRS) depressed mood item [73]. Only healthy controls were included to avoid confounding by Parkinson’s disease pathology.
Psychotic experiences
Three dimensions of psychotic-like experiences –positive, negative and depressive symptom domains– were assessed in IMAGEN at age 19 using the Community Assessment of Psychic Experience (CAPE) [55], a validated self-report questionnaire with good internal consistency [74]. Because psychotic experiences were not assessed at age 14, analyses of psychosis-related outcomes were restricted to age 19.
Polygenic risk scores of psychiatric disorders
Polygenic risk scores (PRS) for major depressive disorder and schizophrenia were derived using summary statistics from the largest available GWAS meta-analyses for depression [75] and schizophrenia [76]. PRS were computed using PRSice (v1.25) with standard clumping (r² = 0.1, 250-kb window) [77] and across 100 P-value thresholds (0 to 0.5 in increments of 0.005). For downstream analyses, we selected the PRS derived using the conventional threshold of P < 0.05.
Statistical analyses
All statistical analyses were conducted in R (version 4.3.1) or Matlab (version R2023a). False discovery rate (FDR) correction (PFDR < 0.05) was applied to all analyses involving multiple parallel tests (e.g., enrichment, CCA, and behavioural associations across clusters or symptom domains), whereas hypothesis-driven confirmatory tests (e.g., replication analyses) were evaluated without further correction.
Canonical correlation analyses (CCA)
These analyses involved the 10 DNAm clusters with evidence of central nervous system relevance (Clusters 1, 3, 4, 5, 6, 7, 8, 9, 11 and 12; totalling 276,980 CpGs) and structural MRI data from 68 cortical regions and 16 subcortical regions. We applied CCA [78, 79] (R package “CCA” version 1.2.2) to assess associations between averaged DNAm within clusters and MRI. Specifically, we investigated associations between changes in DNAm from ages 14 to 19 (i.e., △DNAm = DNAm19-DNAm14) and changes in brain structure during the same period (i.e., △brain = brain19-brain14). Three CCA were run: two investigated changes in cortical measures (i.e., surface area (SA) and average cortical thickness (CT)) within 68 cortical regions, and one, volumetric changes within 16 subcortical regions (SubCortVol). For each CCA, we first investigated the overall correlation between changes in DNAm within clusters with changes in the brain MRI, using permutation tests with 10,000 permutations to calculate exact P-values. FDR correction (PFDR < 0.05) was applied throughout, including when (i) testing the overall significance, (ii) analysing across the full set of DNAm clusters within the first canonical component, and (iii) analysing across the full set of brain ROIs within the first canonical component.
Within the first canonical components, brain regions were considered most relevant if they showed the largest absolute canonical loading ( | R | )—indicating the strongest contribution to the shared DNAm-brain covariation pattern, and the smallest associated p-value within the CCA. A similar approach was taken when considering the most relevant DNAm clusters.
Associations between DNAm, cortical maturation, and adolescent psychopathology
Longitudinal analyses
We examined whether developmental changes in substance use (binge drinking, alcohol use, cannabis use, and tobacco smoking) and mental health symptoms (depressive symptoms and psychotic experiences) were associated with the coordinated DNAm-brain maturation pattern. We focused on the first canonical variate pair from the CCAs, which captured the principal mode of covariation between DNAm change (Y₁; weighted composite of brain-related DNAm clusters) and cortical or subcortical maturation (X₁; weighted composite of cortical thickness (or subcortical volumes) changes across contributing regions). Linear regression models were estimated using change scores (Δ = age19 – age14), with ΔCCA brain or ΔCCA DNAm scores as the dependent variables.
Cross-sectional analyses
To characterise cluster-specific molecular signatures of psychopathology, we conducted linear regression analyses testing associations between average DNAm within each of the 10 brain-related clusters and behavioural measures across three domains: substance use, depressive symptoms, and psychotic symptom dimensions.
All analyses controlled for sex and recruitment site. Sensitivity models additionally controlled for pubertal development, socioeconomic status [80], and co-use of other substances to evaluate potential confounding (see Table S10). Multiple comparisons were addressed using FDR correction within each behavioural domain.
Replication in PPMI
Associations between DNAm clusters and depressive symptoms were tested in the PPMI dataset to assess reproducibility. Models controlled for age, sex, race, years of education, recruitment site, estimated blood cell composition, and the first four DNAm principal components. As the validation analysis tested a predefined directional hypothesis based on discovery results, one-tailed tests were applied to evaluate replication in the expected direction (Pone-tail < 0.05).
Mediation analyses
To further examine whether DNAm change may act as a mechanistic pathway linking behavioural and affective changes to cortical maturation, we conducted longitudinal mediation analyses. All variables were entered as change scores (Δ = age19–age14). Mediation models were implemented in MATLAB using the “MediationToolbox” [81], with non-parametric bootstrapping with 10,000 iterations to estimate indirect effects. We focused on the first canonical variate pair from the ΔCT CCA. Thus, these analyses tested whether ΔDNAm statistically mediated the associations between changes in depressive symptoms or substance-use behaviours and the shared cortical thinning pattern identified in the CCA. Conversely, we also tested whether depressive symptom change mediated the relationship between ΔDNAm and ΔCT, reflecting possible bidirectional coupling between DNAm and affect. To evaluate robustness, models were re-estimated controlling for change in cannabis use and baseline SES (age 14). The inclusion of these covariates did not alter the significance or direction of indirect effects, indicating that the mediation pathways observed were not attributable to socioeconomic or substance-use confounding.
Associations with Polygenic Risk Scores
Associations between PRS and behavioural phenotypes were tested using linear regression. One-tailed tests were used where the expected direction of effect was specified a priori based on prior literature (i.e., higher PRS predicting higher symptom severity). To quantify uncertainty, 95% confidence intervals for correlation coefficients were estimated using 10,000 bootstrap resamples to reflect the one-sided hypothesis. Achieved statistical power was estimated using G*Power (v3.1) [82]. Because only two PRS–depressive symptom tests were conducted (ages 14 and 19), Bonferroni correction was applied to control Type I error. For schizophrenia, PRS associations with three psychosis symptom dimensions, FDR correction was applied within the domain.
Ethical approval
The IMAGEN project received approval from the local ethics research committees at each research site [23]: King’s College London, University of Nottingham, Trinity College Dublin, University of Heidelberg, Technische Universität Dresden, Commissaria de l’Energie Atomique et aux Energies Alternatives and University Medical Center. Informed consent was provided by all participants or by a parent/guardian. All methods were performed in accordance with the committees’ relevant guidelines and regulations.
Data availability
The IMAGEN data is available from a dedicated database at https://imagen2.cea.fr. The PPMI data is available from the website (https://www.ppmi-info.org/). The ADNI data is available from the website (https://adni.loni.usc.edu/).
Code availability
Clustering analyses were conducted using the R package “WGCNA” (version 1.72-5). Functional enrichment analyses were performed with the R package “missMethyl” (version 1.34.0). Tissue-specific enrichment was assessed using the R package “TissueEnrich” (version 1.20.0). T1-weighted MRI images were processed with FreeSurfer (version 6.0; http://surfer.nmr.mgh.harvard.edu/). Canonical correlation analysis (CCA) was carried out using the R package “CCA” (version 1.2.2). CpGs corresponding to the 18 DNAm Clusters are available on Github (https://github.com/dichen27/DNAm_pattern/tree/main/Methylation_Running/Cluster18).
References
Moore LD, Le T, Fan GP. DNA methylation and its basic function. Neuropsychopharmacol. 2013;38:23–38.
Breton CV, Landon R, Kahn LG, Enlow MB, Peterson AK, Bastain T, et al. Exploring the evidence for epigenetic regulation of environmental influences on child health across generations. Commun Biol. 2021;4:769.
Hannon E, Lunnon K, Schalkwyk L, Mill J. Interindividual methylomic variation across blood, cortex, and cerebellum: implications for epigenetic studies of neurological and neuropsychiatric phenotypes. Epigenetics-Us. 2015;10:1024–32.
Wheater ENW, Stoye DQ, Cox SR, Wardlaw JM, Drake AJ, Bastin ME, et al. DNA methylation and brain structure and function across the life course: A systematic review. Neurosci Biobehav R. 2020;113:133–56.
Jia TY, Chu CY, Liu Y, van Dongen J, Papastergios E, Armstrong NJ, et al. Epigenome-wide meta-analysis of blood DNA methylation and its association with subcortical volumes: findings from the ENIGMA Epigenetics Working Group. Mol Psychiatr. 2021;26:3884–95.
Sun Y, Jia T, Barker ED, Chen D, Zhang Z, Xu J, et al. Associations of DNA methylation with behavioral problems, gray matter volumes, and negative life events across adolescence: Evidence from the longitudinal IMAGEN study. Biol Psychiatry. 2023;93:342–51.
Kanherkar RR, Bhatia-Dey N, Csoka AB. Epigenetics across the human lifespan. Front Cell Dev Biol. 2014;2:49.
Cecil CAM, Walton E, Smith RG, Viding E, McCrory EJ, Relton CL, et al. DNA methylation and substance-use risk: a prospective, genome-wide study spanning gestation to adolescence. Transl Psychiat. 2016;6:e976.
Alladi CG, Etain B, Bellivier F, Marie-Claire C. DNA methylation as a biomarker of treatment response variability in serious mental illnesses: A systematic review focused on bipolar disorder, schizophrenia, and major depressive disorder. Int J Mol Sci. 2018;19:3026.
Wang WJ, Li WL, Wu YL, Tian XC, Duan HP, Li SX, et al. Genome-wide DNA methylation and gene expression analyses in monozygotic twins identify potential biomarkers of depression. Transl Psychiat. 2021;11:416.
Giedd JN, Blumenthal J, Jeffries NO, Castellanos FX, Liu H, Zijdenbos A, et al. Brain development during childhood and adolescence: a longitudinal MRI study. Nat Neurosci. 1999;2:861–3.
Carrick C, Baaré WFC, Vitoratou S, Madsen KS, Fuhrmann D. Individual differences in developmental trajectories of global and subcortical brain volumes between late childhood and late adolescence: Findings from a 12-Wave neuroimaging study. Hum Brain Mapp. 2025;46:e70348.
Paus T, Keshavan M, Giedd JN. Why do many psychiatric disorders emerge during adolescence? Nat Rev Neurosci. 2008;9:947–57.
Casey BJ, Jones RM, Hare TA. The adolescent brain. Ann N Y Acad Sci. 2008;1124:111–26.
Yuan M, Yang B, Rothschild G, Mann JJ, Sanford LD, Tang X, et al. Epigenetic regulation in major depression and other stress-related disorders: molecular mechanisms, clinical relevance and therapeutic potential. Signal Transduct Target Ther. 2023;8:309.
Alameda L, Liu Z, Sham PC, Aas M, Trotta G, Rodriguez V, et al. Exploring the mediation of DNA methylation across the epigenome between childhood adversity and First Episode of Psychosis—findings from the EU-GEI study. Mol Psychiatr. 2023;28:2095–106.
Kiltschewskij DJ, Reay WR, Cairns MJ. Schizophrenia is associated with altered DNA methylation variance. Mol Psychiatry. 2025;30:1383–95.
Calderon-Mediavilla M, Vila-Badia R, Dolz M, Butjosa A, Barajas A, Del Cacho N, et al. Depressive symptoms and their relationship with negative and other psychotic symptoms in early onset psychosis. Eur Child Adoles Psy. 2021;30:1383–90.
Edwards CJ, Garety P, Hardy A. The relationship between depressive symptoms and negative symptoms in people with non-affective psychosis: a meta-analysis. Psychol Med. 2019;49:2486–98.
Shukla DK, Chiappelli JJ, Sampath H, Kochunov P, Hare SM, Wisner K, et al. Aberrant frontostriatal connectivity in negative symptoms of schizophrenia. Schizophr Bull. 2019;45:1051–9.
Jefsen OH, Erlangsen A, Nordentoft M, Hjorthøj C. Cannabis use disorder and subsequent risk of psychotic and nonpsychotic unipolar depression and bipolar disorder. Jama Psychiat. 2023;80:803–10.
Owens MM, Albaugh MD, Allgaier N, Yuan D, Robert G, Cupertino RB, et al. Bayesian causal network modeling suggests adolescent cannabis use accelerates prefrontal cortical thinning. Transl Psychiatry. 2022;12:188.
Schumann G, Loth E, Banaschewski T, Barbot A, Barker G, Buchel C, et al. The IMAGEN study: reinforcement-related behaviour in normal brain function and psychopathology. Mol Psychiatr. 2010;15:1128–39.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559.
Marek K, Jennings D, Lasch S, Siderowf A, Tanner C, Simuni T, et al. The Parkinson progression marker initiative (PPMI). Prog Neurobiol. 2011;95:629–35.
Weiner MW, Veitch DP, Aisen PS, Beckett LA, Cairns NJ, Green RC, et al. The Alzheimer’s Disease Neuroimaging Initiative: a review of papers published since its inception. Alzheimer’s Dementia. 2013;9:e111–e194.
Mantel N. The detection of disease clustering and a generalized regression approach. Cancer Res. 1967;27:209–20.
Sowell ER, Peterson BS, Thompson PM, Welcome SE, Henkenius AL, Toga AW. Mapping cortical change across the human life span. Nat Neurosci. 2003;6:309–15.
Lenroot RK, Giedd JN. Brain development in children and adolescents: insights from anatomical magnetic resonance imaging. Neurosci Biobehav Rev. 2006;30:718–29.
Zhang S, Ye X, Bai G, Fu Y, Mao C, Wu A, et al. Alterations in cortical thickness and white matter integrity in mild-to-moderate communicating hydrocephalic school-aged children measured by whole-brain cortical thickness mapping and DTI. Neural Plast. 2017;2017:5167973.
Madsen SK, Gutman BA, Joshi SH, Toga AW, Jack CR Jr, Weiner MW, et al. Mapping ventricular expansion onto cortical gray matter in older adults. Neurobiol Aging. 2015;36:S32–S41.
Drevets WC. Orbitofrontal cortex function and structure in depression. Ann N Y Acad Sci. 2007;1121:499–527.
Tottenham N, Galván A. Stress and the adolescent brain: Amygdala-prefrontal cortex circuitry and ventral striatum as developmental targets. Neurosci Biobehav Rev. 2016;70:217–27.
Volkow ND, Koob GF, McLellan AT. Neurobiologic Advances from the Brain Disease Model of Addiction. N Engl J Med. 2016;374:363–71.
Carreras-Gallo N, Dwaraka VB, Cáceres A, Smith R, Mendez TL, Went H, et al. Impact of tobacco, alcohol, and marijuana on genome-wide DNA methylation and its relationship with hypertension. Epigenetics-Us. 2023;18:2214392.
Albaugh MD, Ottino-Gonzalez J, Sidwell A, Lepage C, Juliano A, Owens MM, et al. Association of cannabis use during adolescence with neurodevelopment. Jama Psychiat. 2021;78:1–11.
Karoly HC, Kirk-Provencher KT, Schacht JP, Gowin JL. Alcohol and brain structure across the lifespan: A systematic review of large-scale neuroimaging studies. Addict Biol. 2024;29:e13439.
Loyfer N, Magenheim J, Peretz A, Cann G, Bredno J, Klochendler A, et al. A DNA methylation atlas of normal human cell types. Nature. 2023;613:355–64.
Horvath S, Zhang YF, Langfelder P, Kahn RS, Boks MPM, van Eijk K, et al. Aging effects on DNA methylation modules in human brain and blood tissue. Genome Biol. 2012;13:R97.
Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14:R115.
Arain M, Haque M, Johal L, Mathur P, Nel W, Rais A, et al. Maturation of the adolescent brain. Neuropsychiatr Dis Treat. 2013;9:449–61.
Eichenbaum H, Yonelinas AP, Ranganath C. The medial temporal lobe and recognition memory. Annu Rev Neurosci. 2007;30:123–52.
Serhan CN. Pro-resolving lipid mediators are leads for resolution physiology. Nature. 2014;510:92–101.
Friedman NP, Robbins TW. The role of prefrontal cortex in cognitive control and executive function. Neuropsychopharmacol. 2022;47:72–89.
Ahmed SP, Bittencourt-Hewitt A, Sebastian CL. Neurocognitive bases of emotion regulation development in adolescence. Dev Cogn Neurosci. 2015;15:11–25.
Giedd JN, Stockman M, Weddle C, Liverpool M, Alexander-Bloch A, Wallace GL, et al. Anatomic magnetic resonance imaging of the developing child and adolescent brain and effects of genetic variation. Neuropsychol Rev. 2010;20:349–61.
Bethlehem RAI, Seidlitz J, White SR, Vogel JW, Anderson KM, Adamson C, et al. Brain charts for the human lifespan. Nature. 2022;604:525–33.
Apostolova LG, Green AE, Babakchanian S, Hwang KS, Chou YY, Toga AW, et al. Hippocampal atrophy and ventricular enlargement in normal aging, mild cognitive impairment (MCI), and Alzheimer disease. Alz Dis Assoc Dis. 2012;26:17–27.
Russo SJ, Nestler EJ. The brain reward circuitry in mood disorders. Nat Rev Neurosci. 2013;14:609–25.
Knopik VS, Marceau K, Bidwell LC, Bolan E. Prenatal substance exposure and offspring development: Does DNA methylation play a role? Neurotoxicol Teratol. 2019;71:50–63.
Wassum KM. Amygdala-cortical collaboration in reward learning and decision making. eLife. 2022;11:e80926.
Ferri J, Eisendrath SJ, Fryer SL, Gillung E, Roach BJ, Mathalon DH. Blunted amygdala activity is associated with depression severity in treatment-resistant depression. Cogn Affect Behav Neurosci. 2017;17:1221–31.
Fahim C, Stip E, Mancini-Marïe A, Mensour B, Boulay LJ, Leroux JM, et al. Brain activity during emotionally negative pictures in schizophrenia with and without flat affect: An fMRI study. Psychiatry Res. 2005;140:1–15.
Grayson DR, Guidotti A. The dynamics of DNA methylation in schizophrenia and related psychiatric disorders. Neuropsychopharmacol. 2013;38:138–66.
Stefanis NC, Hanssen M, Smirnis NK, Avramopoulos DA, Evdokimidis IK, Stefanis CN, et al. Evidence that three dimensions of psychosis have a distribution in the general population. Psychol Med. 2002;32:347–58.
Kibel DA, Laffont I, Liddle PF. The composition of the negative syndrome of chronic-schizophrenia. Brit J Psychiat. 1993;162:744–50.
Chen DM, Meng L, Pei F, Zheng Y, Leng JY. A review of DNA methylation in depression. J Clin Neurosci. 2017;43:39–46.
Guintivano J, Arad M, Gould TD, Payne JL, Kaminsky ZA. Antenatal prediction of postpartum depression with blood DNA methylation biomarkers (vol 19, pg 560, 2013). Mol Psychiatr. 2014;19:633–633.
Fareri DS, Tottenham N. Effects of early life stress on amygdala and striatal development. Dev Cogn Neuros-Neth. 2016;19:233–47.
Schmaal L, Veltman DJ, van Erp TGM, Sämann PG, Frodl T, Jahanshad N, et al. Subcortical brain alterations in major depressive disorder: findings from the ENIGMA Major Depressive Disorder working group. Mol Psychiatr. 2016;21:806–12.
Schmaal L, Hibar DP, Sämann PG, Hall GB, Baune BT, Jahanshad N, et al. Cortical abnormalities in adults and adolescents with major depression based on brain scans from 20 cohorts worldwide in the ENIGMA Major Depressive Disorder Working Group. Mol Psychiatr. 2017;22:900–9.
Chen D, Wang X, Voon V, Jiang Y, Lo C-YZ, Wang L, et al. Neurophysiological stratification of major depressive disorder by distinct trajectories. Nature Mental Health. 2023;1:863–75.
Guadagno A, Belliveau C, Mechawar N, Walker CD. Effects of early life stress on the developing basolateral amygdala-prefrontal cortex circuit: The emerging role of local inhibition and perineuronal nets. Front Hum Neurosci. 2021;15:669120.
Sandoval J, Heyn HA, Moran S, Serra-Musach J, Pujana MA, Bibikova M, et al. Validation of a DNA methylation microarray for 450,000 CpG sites in the human genome. Epigenetics-Us. 2011;6:692–702.
Aryee MJ, Jaffe AE, Corrada-Bravo H, Ladd-Acosta C, Feinberg AP, Hansen KD, et al. Minfi: A flexible and comprehensive Bioconductor package for the analysis of Infinium DNA methylation microarrays. Bioinformatics. 2014;30:1363–9.
Jia T, Macare C, Desrivières S, Gonzalez DA, Tao C, Ji X, et al. Neural basis of reward anticipation and its genetic determinants. Proc Natl Acad Sci USA. 2016;113:3879–84.
Langfelder P, Horvath S. Fast R functions for robust correlations and hierarchical clustering. J Stat Softw. 2012;46:1–17.
Desikan RS, Segonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage. 2006;31:968–80.
Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C, et al. Whole brain segmentation: Automated labeling of neuroanatomical structures in the human brain. Neuron. 2002;33:341–55.
Goodman R, Ford T, Richards H, Gatward R, Meltzer H. The Development and Well-Being Assessment: Description and initial validation of an integrated assessment of child and adolescent psychopathology. J Child Psychol Psychiatry. 2000;41:645–55.
Petersen AC, Crockett L, Richards M, Boxer A. A self-report measure of pubertal status: Reliability, validity, and initial norms. J Youth Adolesc. 1988;17:117–33.
Revah-Levy A, Birmaher B, Gasquet I, Falissard B. The Adolescent Depression Rating Scale (ADRS): a validation study. BMC Psychiatry. 2007;7:2.
Goetz CG, Tilley BC, Shaftman SR, Stebbins GT, Fahn S, Martinez-Martin P, et al. Movement disorder society-sponsored revision of the unified Parkinson’s disease rating scale (MDS-UPDRS): Scale presentation and clinimetric testing results. Movement Disord. 2008;23:2129–70.
Brenner K, Schmitz N, Pawliuk N, Fathalli F, Joober R, Ciampi A, et al. Validation of the english and french versions of the community assessment of psychic experiences (CAPE) with a montreal community sample. Schizophr Res. 2007;95:86–95.
Wray NR, Ripke S, Mattheisen M, Trzaskowski M, Byrne EM, Abdellaoui A, et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet. 2018;50:668–81.
Trubetskoy V, Pardinas AF, Qi T, Panagiotaropoulou G, Awasthi S, Bigdeli TB, et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature. 2022;604:502–8.
Vilhjalmsson BJ, Yang J, Finucane HK, Gusev A, Lindstrom S, Ripke S, et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet. 2015;97:576–92.
Jia TY, Ing A, Quinlan EB, Tay N, Luo Q, Francesca B, et al. Neurobehavioural characterisation and stratification of reinforcement-related behaviour. Nat Hum Behav. 2020;4:544–58.
Knapp TR. Canonical correlation analysis - general parametric significance-testing system. Psychol Bull. 1978;85:410–6.
Miller AP, Baranger DAA, Paul SE, Garavan H, Mackey S, Tapert SF, et al. Neuroanatomical variability and substance use initiation in late childhood and early adolescence. Jama Netw Open. 2024;7:e2452027.
Wager TD, Davidson ML, Hughes BL, Lindquist MA, Ochsner KN. Prefrontal-subcortical pathways mediating successful emotion regulation. Neuron. 2008;59:1037–50.
Faul F, Erdfelder E, Lang AG, Buchner A. G*Power 3: A flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav Res Methods. 2007;39:175–91.
Acknowledgements
This work received support from the following sources: National Key R&D Program of China (2023YFE0199700 to T.J.); National Natural Science Foundation of China (T2122005 and 81801773 to T.J., 82150710554 to G.S.); the European Union-funded FP6 Integrated Project IMAGEN (reinforcement-related behavior in normal brain function and psychopathology; LSHM-CT- 2007-037286 to G.S.), the European Union (grant 101057429 to G.S.) and UK Research and Innovation (grant 10038599 and 10041392 to S.D.) -funded environMENTAL project, the Medical Research Council and Medical Research Foundation (MR/R00465X/, MRF-058-0004-RG-DESRI, ‘ESTRA’- Neurobiological underpinning of eating disorders: integrative biopsychosocial longitudinal analyses in adolescents; and MR/S020306/1, MRF-058-0009-RG-DESR-C0759 ‘ESTRA’-Establishing causal relationships between biopsychosocial predictors and correlates of eating disorders and their mediation by neural pathways to S.D.), the Horizon 2020-funded European Research Council Advanced Grant for STRATIFY (brain network-based stratification of reinforcement-related disorders; 695313 to G.S.), a cross- National Institute of Health (NIH) alliance that funds Big Data to Knowledge Centres of Excellence (ENIGMA; 5U54EB020403-05 and ENIGMA World Aging Center 1R56AG058854-01 grants to S.D. and P.M.T.), and the KC Wong Fellowship (to D.C.).
Other funding included: the Shanghai Pujiang Project (18PJ1400900 to T.J.), Guangdong Key Research and Development Project (2018B030335001 to J.F.), the 111 Project (B18015 to J.F.), the key project of Shanghai Science and Technology (16JC1420402 to J.F.), Shanghai Municipal Science and Technology Major Project (2018SHZDZX01 to J.F.), Zhang Jiang Lab (to J.F.), Shanghai Center for Brain Science and Brain-Inspired Technology (to J.F.), the Medical Research Council (grant MR/W002418/1: Eating Disorders: Delineating illness and recovery trajectories to inform personalized prevention and early intervention in young people (EDIFY) to S.D.), Human Brain Project (HBP SGA 2, 785907, and HBP SGA 3, 945539, to G.S.), the Medical Research Council Grant for c-VEDA (Consortium on Vulnerability to Externalising Disorders and Addictions; MR/ N000390/1 to G.S.), the National Institute of Health (NIH) (a decentralized macro and micro gene-by-environment interaction analysis of substance use behavior and its brain biomarkers; R01DA049238 to G.S.), the Bundesministeriumfür Bildung und Forschung (grants 01GS08152 and 01EV0711 to G.S.), Forschungsnetz AERIAL (01EE1406A and 01EE1406B to G.S.), Forschungsnetz IMAC-Mind (01GL1745B to G.S.), the Deutsche Forschungsgemeinschaft (SM 80/7-2, SFB 940, TRR 265 and NE 1383/14-1 to G.S.). Further support was provided by grants from the L’Agence nationale de la recherche (ANR) (ANR-12-SAMA-0004 to M.-L.P.M., and AAPG2019 – GeBra and ANR-18-NEUR00002-01 – ADORe to J.-L.M.); the Eranet Neuron (AF12-NEUR0008-01 – WM2NA to J.-L.M.); the Fondation de France (00081242 to J.-L.M.); the Fondation pour la Recherche Médicale (DPA20140629802 to J.-L.M.); the Mission Interministérielle de Lutte-contre-les-Drogues-et-les-Conduites-Addictives (MILDECA to J.-L.M.); the Assistance Publique–Hôpitaux de Paris and Institut National de la Santé et de la Recherche Médicale (interface grant to M.-L.P.M.); Paris Sud University IDEX 2012 (to J.-L.M.); the Fondation de l’Avenir (AP-RM-17-013 to M.-L.P.M.); the Fédération pour la Recherche sur le Cerveau; and the NIH, Science Foundation Ireland (16/ERCD/3797 to R.W.), USA (Axon, Testosterone and Mental Health during Adolescence; RO1 MH085772-01A1 to T.P.). This paper represents independent research, partly funded by the NIHR Maudsley Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The views expressed are those of the authors and not necessarily those of the NIHR or the Department of Health and Social Care.
Author information
Authors and Affiliations
Consortia
Contributions
TJ, SD and DC conceptualised the study and interpreted the findings. DC performed the analyses and visualised the results. XY, TB, ALWB, HF, AG, HG, PG, AH, RB, J-LM, M-LPM, EA, FN, DPO, HL, TP, LP, SH, NH, CB, MNS, NV, HW, RW, GS, and SD collected and processed the IMAGEN data. WC and WZ contributed to the analysis of the ADNI data. WC and LW contributed to the analysis of the PPMI data. SX, JF and PMT contributed to MRI analyses. DC and SD drafted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
TB served in an advisory or consultancy role for eye level, Infectopharm, Medice, Neurim Pharmaceuticals, Oberberg GmbH and Takeda. He received conference support or speaker’s fee by Janssen, Medice and Takeda. He received royalties from Hogrefe, Kohlhammer, CIP Medien and Oxford University Press. The present work is unrelated to the above grants and relationships. GJB received honoraria from General Electric Healthcare for teaching scanner programming courses. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, D., Yu, X., Cheng, W. et al. A longitudinal DNA methylation atlas and its link to brain structure and mental health. Mol Psychiatry (2026). https://doi.org/10.1038/s41380-026-03554-y
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41380-026-03554-y