Abstract
Neddylation, a post-translational modification process involving three enzymatic steps, is crucial in regulating various cancers. However, its specific mechanisms in pancreatic cancer (PC) remain largely unexplored. This study focused on screening neddylation-related molecules in PC and identified RNF187 as a key player, demonstrating its overexpression in PC and its ability to enhance cell proliferation and invasion both in vitro and in vivo. Notably, NEDD8 could bind to RNF187, preventing its degradation via K48-linked ubiquitination. This interaction stabilized RNF187, leading to increased protein levels and subsequent stimulation of PC cell proliferation and invasion. However, this mechanism alone did not fully account for how RNF187 could exacerbate PC malignancy. Further research revealed that RNF187 upregulated IQGAP1 protein levels through modulation of K48- and K63-linked ubiquitination. This post-translational modification triggered the rearrangement of the actin cytoskeleton in PC cells by altering the transcriptional levels of MYH9, thereby promoting PC malignancy. Overall, our findings demonstrate that neddylation of RNF187 enhances PC malignancy through the IQGAP1/MYH9 axis, suggesting a new therapeutic target for PC.

Similar content being viewed by others
Introduction
Pancreatic cancer (PC), one of the most malignant digestive tumors, was the third most common cause of cancer-related death in 2023, with projections indicating it may become the second by 2025 [1, 2]. The prognosis remains extremely poor, with a 5-year survival rate of only 12%, and near-zero survival in advanced stages due to late diagnosis and the absence of reliable early biomarkers. Most patients present with non-specific symptoms, resulting in diagnosis at advanced stages [3,4,5,6]. Therefore, identifying key molecular mechanisms underlying PC progression and discovering novel therapeutic targets is an urgent priority. Recent research has increasingly focused on post-translational modifications, particularly neddylation, as critical regulators of tumor biology.
Protein neddylation, a ubiquitin-like post-translational modification, has attracted growing interest in cancer research due to its role in regulating protein function, stability, and localization [7, 8]. The neddylation cascade involves sequential enzymatic steps, where NEDD8 is activated by the NEDD8-activating enzyme E1 (NAE), transferred to E2 conjugating enzymes (UBC12/UBE2F), and finally ligated to target substrates by E3 ligases [7, 9,10,11,12]. This modification predominantly affects cullin-RING ligases (CRLs), modulating critical oncogenic processes such as cell cycle progression, DNA replication, and stress responses [13,14,15]. Dysregulation of neddylation has been implicated in multiple cancers including colorectal, hepatocellular, and breast cancer. Moreover, pharmacological inhibition of neddylation has shown promise in preclinical models, enhancing chemosensitivity and suppressing tumor growth, thereby highlighting its potential as a therapeutic target [8].
RNF187 (RING finger 187) is a RING family ubiquitin ligase also referred to as RING domain AP1 coactivator-1 [16]. It has been reported to promote epithelial-mesenchymal transition (EMT), proliferation, and metastasis in several malignancies, such as hepatocellular carcinoma, colorectal cancer, and non-small-cell lung cancer, while exerting tumor-suppressive effects in others, including esophageal squamous cell carcinoma and triple-negative breast cancer [17,18,19,20]. Mechanistically, RNF187 has been implicated in the regulation of key oncogenic pathways, such as Hippo and MAPK signaling. Given its context-dependent roles and emerging relevance in multiple tumor types, RNF187 has garnered increasing attention as a potential biomarker and therapeutic target in cancer biology. However, studies focusing on RNF187 in pancreatic cancer are still lacking, and its precise biological function in this context remains poorly understood and warrants further investigation in the context of PC. Further investigation into RNF187 in PC may uncover novel regulatory pathways and provide new opportunities for therapeutic intervention.
In this study, we demonstrated that RNF187 is overexpressed in pancreatic cancer and promotes tumor progression. We further identified a novel mechanism whereby RNF187-mediated neddylation modulates the actin cytoskeleton by regulating IQGAP1 and MYH9 expression. These findings provide new insights into the biological functions of RNF187 and uncover potential therapeutic avenues for targeting pancreatic cancer.
Methods
Cell culture
All cell lines were obtained from the American Type Culture Collection (ATCC). Human embryonic kidney (HEK293T) cells and human PC cell lines (MIA PaCa-2, PANC-1, SW-1990) were cultured in Dulbecco’s Modified Eagle Medium (DMEM) medium supplemented with 10% fetal bovine serum (FBS) and 100 units/mL penicillin/streptomycin at 37 °C in a humidified atmosphere containing 5% CO2. Human pancreatic ductal epithelial (HPDE) cells and additional human PC cell lines (AsPC-1, BxPC-3) were maintained under the identical conditions.
Transfection
Cells were transfected with small interfering RNAs (siRNAs) targeting RNF187 and NEDD8 using Lipofectamine RNAiMAX, following the manufacturer’s instructions (Invitrogen). Forty-eight hours post-transfection, the cells were utilized for subsequent research. Overexpression plasmids for RNF187, NEDD8, and a short hairpin RNA (shRNA) targeting IQGAP1 were procured from Genechem Company (Shanghai, China). Cellular transfection with the respective plasmids was conducted according to the manufacturer’s recommendations. The efficacy of transfection was subsequently assessed via quantitative polymerase chain reaction (qPCR) and western blot analysis. All oligonucleotide sequences are provided in Table S1.
RNA extraction and quantitative reverse transcription-PCR
Total RNA was isolated from PC cells using the RNeasy Mini Kit (Vazyme, China) according to the manufacturer’s instructions. cDNA was subsequently synthesized from the isolated RNA using HiScript® III RT SuperMix for qPCR (Vazyme). Quantitative PCR (qPCR) was performed to amplify target genes, with HiScript® III RT SuperMix for qPCR (Vazyme) employed for this purpose. β-Actin served as an internal control for normalization. Primer sequences for all target genes are provided in Table S1.
Western blotting
Total protein was extracted from cell lysates using RIPA buffer (Solarbio). For Co-IP experiments, RIPA buffer was replaced with NP40 buffer (Beyotime). Protein samples were denatured by boiling in SDS loading buffer for 5–10 min at 95 °C prior to separation by 10% SDS-PAGE (Epizyme). Separated proteins were transferred to PVDF membranes (Sigma-Aldrich). Membranes were blocked with 3% bovine serum albumin (BSA) for 30 min. Subsequently, membranes were incubated with primary antibodies for 12–18 h at 4 °C, followed by incubation with appropriate secondary antibodies for 1–4 h. Protein bands were visualized using chemiluminescence detection. A complete list of antibodies used in this study is provided in Table S1.
Cell Counting Kit-8 (CCK-8) assay
In accordance with the manual’s instructions, cell proliferation was evaluated using the CCK-8 assay (Beyotime). 1 × 104 PC cells were seeded in a 96-well plate, and cell viability was thereafter assessed daily for the designated duration. Absorbance was measured at 450 nm to determine cell viability.
Colony formation assay
Five hundred PC cells were plated on six-well plates and subsequently cultured for an additional 14 days. Subsequently, the cells were stained with 2% crystal violet for 30 min at room temperature before being fixed with 4% paraformaldehyde. The wells were then cleaned, allowed to dry, and photographed. ImageJ software was used to quantify the number of clones.
5-ethynyl-2-deoxyuridine (EdU) staining
EdU incorporation assays were conducted using the EdU Cell Proliferation Kit 555 (Beyotime, Shanghai, China) according to the manufacturer’s instructions. Cells were incubated with EdU (10 μM) for two hours at 37 °C. Subsequently, the plates were rinsed with PBS, fixed with 4% paraformaldehyde for 10 min, and stained with DAPI for 15 min. EdU-positive cells were visualized using fluorescence microscopy, and the percentage of EdU-positive cells was quantified with ImageJ.
Transwell assay
5 × 105 cells were seeded in the upper chamber of the non-FBS medium utilizing a transwell filter with an 8 μm pore size. 10% fetal bovine serum (FBS) was added to the lower chamber medium. Following a 24-h incubation at room temperature, the filters were fixed with 4% paraformaldehyde and subsequently stained with 2% crystal violet for 30 min. Images of the stained filters were then captured using a camera.
Wound healing assay
In a 6-well plate, cells were dislodged using a 200 μl pipette tip. Subsequently, images of the cells were acquired, and their positions within the images were recorded. After 48 h, the cells were re-imaged. The mobility of each cell group was then determined.
Co-IP and mass spectrometry
Following cell lysis of RNF187-overexpressing cells, immunoprecipitation was performed using either anti-RNF187 or anti-IgG antibodies. Co-immunoprecipitated complexes were subsequently analyzed by Western blotting and liquid chromatography-tandem mass spectrometry (LC-MS/MS).
Ubiquitination assays
The proteasome inhibitor MG132 was applied to the HA-Ub-transfected cells for 6 h. Protein A/G magnetic beads and IP buffer were used to fix the produced cell lysates before immunoprecipitation with the appropriate antibodies. Finally, the immunoprecipitated protein complexes were analyzed by Western blotting.
Immunofluorescence
Cells on slides were permeabilized with 0.1% Triton X-10 and fixed with 4% paraformaldehyde. Subsequently, cells were incubated with primary antibodies against RNF187, IQGAP1, or NEDD8 overnight at 4 °C. Finally, cells were incubated with an IgG fluorescent secondary antibody. Immunofluorescence microscopy was used to image the cells after they had been fixed and stained with DAPI.
Immunohistochemistry
The tissues were embedded in paraffin blocks after being fixed in formalin for 14 h. The tissue sections were divided into 5 μm thick sections and subjected to an overnight incubation at 4 °C with the primary antibodies, followed by a 20-min incubation at room temperature with the secondary antibodies. The total of the scores for staining intensity and the proportion of stained cells that were positive were used to determine the IHC score. The staining intensity was graded on a scale of 0–3 for absence, weakness, moderate, and strength. The percentage of cells that stained positively was graded on a scale of 1% to 100% (1% = 0, 1–25% = 1, 26–50% = 3, 51–75% = 4, 76–100% = 5).
Phalloidin staining
Cells were fixed with 4% paraformaldehyde for 20 min, and permeabilized with 0.1% TritonX-100 for 30 min. Actin cytoskeleton was stained using Actin-Tracker Red-Rhodamine (Beyotime, Shanghai, China) at room temperature for 1 h. Then cells were stained with DAPI for 10 min. Images were collected with fluorescence microscopy.
Single-cell RNA sequencing and spatial transcriptomic analysis
The single-cell RNA sequencing data were obtained from the Genome Sequence Archive under project PRJCA001063. Quality control was performed using Seurat (version 5.1.0), excluding single cells that had fewer than 400 UMIs, more than 5000 UMIs, or greater than 20% mitochondrial-derived UMIs. Batch effects across patients were corrected using the Harmony method, based on the top 30 principal components and the top 2000 variable genes. Main cell clusters were identified using Seurat’s FindClusters function and visualized using 2D UMAP. T cells, Macrophage, and cancer cells, and other clusters were annotated based on canonical markers such as CD2, CD79A, and KRT19, respectively, for further dimensionality reduction using the FindClusters function in the Seurat package.
The Gene Expression Omnibus (GEO) database was accessed to retrieve the GSE272362 dataset, from which only human samples were used for spatial transcriptomic analysis. Gene expression co-localization was performed using the SpatialFeaturePlotBlend function in the ST analysis.
Human and animal studies
Tissues were collected from 38 patients with PC at the The First Affiliated Hospital of Harbin Medical University. Pathological analysis confirmed the presence of PC in the collected patient tissues. This study was approved by the Ethics Committee of the First Affiliated Hospital of Harbin Medical University (IRB-AF/SQ-02/04.0), and all participants signed the informed consent.
All animal procedures in this study were conducted in strict accordance with the Health Guide for the Care and Use of Laboratory Animals and were approved by the Animal Care Committee at Harbin Medical University to ensure humane treatment (2024090). The female BALB/c-nude mice were obtained from Jiangsu Huachuang Xinnuo Pharmaceutical Technology at a 5-week-old age. To construct the xenograft PC-bearing model, PANC-1 cells (5 × 106 cells/100 µl) were implanted into mice. The tumor volume was calculated using the formula tumor volume = 1/2× length × width. Tumors were removed for further research after 4 weeks. To establish the liver metastasis model, 2 × 105 cells/100 µl of PC cells from each group were injected into the spleens of nude mice. The mice were sacrificed by cervical dislocation after the injection, and the number of metastatic nodules was determined.
Integration of the Rbio model for biological inference
This study utilized the Rbio model, which is based on Qwen2.5-3B-Instruct and further fine-tuned with reinforcement learning (GRPO) and soft supervision techniques [21]. The model integrates biological knowledge sources and verifiers, including TranscriptFormer and Gene Ontology, to embed gene expression data for biological reasoning. In this study, we applied the Rbio model to predict whether the knockdown of IQGAP1 would result in differential expression of MYH9. The model predicts the probability of gene knockdown using an MLP network, and this prediction serves as soft evidence during the inference process, enhancing the accuracy of the model’s reasoning.
Statistical analysis
Statistical differences were examined using GraphPad Prism 9 software. Comparisons between groups were performed using two-tailed Student’s t-tests or one-way ANOVA, as appropriate.
Results
RNF187 expression is upregulated in pancreatic cancer and correlates with NEDD8
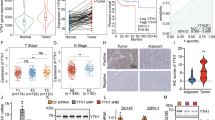
Emerging evidence suggests that inhibiting neddylation significantly reduces PC proliferation; however, the detailed mechanisms remain unclear. In this study, we explored the NEDD8 E3 ligase, which is crucial in the E3 ubiquitin ligase complex for NEDD8 molecule translocation. Our analysis cross-referenced NEDD8-associated genes with E3 ubiquitin ligases specifically implicated in PC, revealing a notable association between NEDD8 and six E3 ubiquitin ligases (RNF7, RNF25, RBX1, RNF113A, RNF181, and RNF187) in PC (Fig. 1A, B and S1A–E). The Gene Expression Profiling Interactive Analysis (GEPIA) database demonstrated elevated expression of RNF7, RNF25, RBX1, RNF181, and RNF187 in PC tissues (Figs. 1C and S1F–J). Single-cell RNA sequencing analysis identified 11 main clusters, including acinar, B cells, cancer, ductal, endocrine, endothelial, fibroblasts, macrophages, mast cells, Schwann, and T cells. Our analysis revealed that RBX1, RNF7, RNF113A, RNF181, and RNF187 were upregulated in cancer cells compared to ductal cells (Figs. 1D, E and S1K–P). While PCR assays indicated upregulation of both RNF113A and RNF187, Western blot analysis confirmed consistently elevated RNF187 protein expression across all pancreatic cancer cell lines, whereas other candidate genes exhibited more heterogeneous expression patterns (Figs. 1F, G, and S2A–J). These findings, integrating gene expression data from GEPIA, single-cell RNA sequencing, PCR, and Western blot validation, demonstrated that RNF187 exhibited the most consistent and robust upregulation across pancreatic cancer samples, making it the most promising candidate for further investigation. Further analyses were conducted to elucidate the correlation between RNF187 expression and PC. GEPIA data associated high RNF187 expression with poorer PC prognoses (Fig. 1H), and TNMplot analysis further supported RNF187 upregulation in PC (Fig. S2K). This observation was corroborated by the Human Protein Atlas (HPA), which indicated upregulated RNF187 expression in PC as determined by immunohistochemistry (IHC) (Fig. 1I). Comparative analyses of RNF187 expression in PC tissues versus adjacent non-tumorous tissues revealed significantly upregulated levels of RNF187 in PC, as evidenced by IHC staining, q-PCR, and Western blotting, with RNF187 expression being even higher in advanced stages of pancreatic cancer (Fig. 1J–M).
A Venn diagram identifying NEDD8-associated genes and overlapping E3 ligase genes. B Correlation analysis between RNF187 and NEDD8 expression. C RNF187 expression levels in PC (n = 179) and normal tissues (n = 171) from the GEPIA database. D Uniform Manifold Approximation and Projection (UMAP) plot illustrating distinct cell types. E Single-cell RNA sequencing analysis comparing RNF187 expression in cancer cells versus ductal cells. F qRT-PCR analysis of RNF187 mRNA expression in PC cells and HPDE cells. G Western blot analysis of RNF187 protein expression in PC cells and HPDE cells. H Kaplan–Meier survival curves depicting disease-free survival (DFS) based on RNF187 expression in PC patients. I IHC analysis of RNF187 expression in PC tissues from the Human Protein Atlas (HPA). J–L RNF187 mRNA and protein expression levels in PC patients, comparing tumor (T) and adjacent normal (N) tissues (Normal = 23, Tumor = 38). (M) IHC staining of RNF187 in PC and surrounding normal tissues (N = 17). All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
RNF187 enhances the malignant phenotype of pancreatic cancer cell lines
To determine RNF187’s role in PC, we utilized PANC-1 and SW1990 cell lines, characterized by high RNF187 expression. Western blot analysis confirmed the efficacy of RNF187 overexpression and knockdown in these PC cells (Fig. 2A). Subsequent functional assays, including Cell Counting Kit-8 (CCK-8), 5-ethynyl-2’-deoxyuridine (EdU) incorporation, and colony formation assays, demonstrated a significant reduction in proliferation rates of PANC-1 and SW1990 cells following RNF187 knockdown, whereas RNF187 overexpression led to increased proliferation (Figs. 2B–E and S2L–M). To assess the impact of RNF187 on cellular motility, we performed transwell and wound healing assays. Results showed slower wound closure in both cell lines with RNF187 knockdown, while overexpression enhanced wound healing (Fig. S2N, O). The transwell assay similarly revealed decreased migratory and invasive potential with suppressed RNF187, while its overexpression led to enhanced migratory and invasive properties (Fig. 2F–H). To further elucidate the role of RNF187 in pancreatic cancer progression, we established both subcutaneous and liver metastasis xenograft models in nude mice. In the subcutaneous model, tumors derived from RNF187-overexpressing PANC-1 cells exhibited significantly accelerated growth compared to controls (Fig. 2J–L). Histopathological evaluation, including Hematoxylin and eosin (HE) staining and IHC analysis, revealed a notable increase in Ki67 expression, indicative of enhanced proliferative activity (Fig. 2M). In contrast, in the RNF187 knockdown in vivo model, tumor growth was significantly slower, aligning with experimental expectations (Fig. S3A–D). In the liver metastasis model, RNF187 overexpression markedly enhanced the development of metastatic lesions and was correlated with reduced overall survival (Fig. 2M–O), whereas RNF187 knockdown resulted in diminished metastatic lesion formation and improved survival outcomes (Fig. S3E–G). These in vivo results underscore the pro-metastatic capacity of RNF187 and highlight its contribution to both tumor progression and poor prognosis in pancreatic cancer. Furthermore, Western blot analyses revealed that overexpression of RNF187 significantly upregulated the expression of several key proteins associated with cancer malignancy, such as vimentin, snail, MMP9, MMP12, N-cadherin, and PCNA, while concurrently downregulating E-cadherin expression (Fig. 2P). Conversely, RNF187 knockdown yielded the opposite effect (Fig. 2P). These findings collectively provide strong evidence that RNF187 plays a pivotal role in promoting the malignant phenotype of PC cells.
A Western blot analysis confirmed the transfection efficiency of RNF187 siRNA and RNF187 overexpression plasmids. B–E The proliferative capacity of transfected PC cells was assessed using CCK-8 and colony formation. F–H Migration and invasion abilities were evaluated through transwell assays in transfected PC cells. I Representative images of tumors from the Vector and OE-RNF187 groups (N = 5/ groups). J Tumor weights for each group. K Tumor growth curves. L HE staining and IHC for RNF187 and Ki67 in tumor sections. M Representative images of liver metastases and HE staining in the Vector and OE-RNF187 groups (N = 5/ groups). N Kaplan–-Meier survival analysis. O Quantification of liver metastasis foci. P Western blotting was employed to detect alterations in the expression of key proteins associated with PC malignancy in transfected cells. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
RNF187 interacts with NEDD8 in PC
To elucidate the relationship between RNF187 and NEDD8, we further employed Spatial Transcriptomics to identify their spatial interactions. Spatial transcriptomics analysis revealed significant spatial colocalization of RNF187 and NEDD8 within tumor epithelial cells, suggesting a potential synergistic role in tumor-associated biological processes (Fig. 3A). Single-cell analysis revealed upregulated NEDD8 expression in the RNF187 high-expression group (Fig. 3B). Analysis of the GEPIA database demonstrated a strong correlation between NEDD8 and RNF187, with upregulated expression of NEDD8 in tumor tissues (Fig. 3C, D). Similarly, single-cell analysis revealed that RNF187 expression was higher in the NEDD8 high-expression group (Fig. 3E). These findings collectively highlight a close association between RNF187 and NEDD8, reinforcing their potential cooperative roles in tumor progression. To explore the reciprocal regulatory relationship between RNF187 and NEDD8, we employed both Western blotting and RT-PCR. Interestingly, our findings revealed that the presence of RNF187 did not alter NEDD8 levels (Figs. 3F and S4A, S5A). However, Western blot analysis indicated that NEDD8 modulated RNF187 protein levels, while q-PCR results indicated that RNF187 mRNA expression was unaffected by NEDD8 (Figs. 3G and S4B, S5B). Furthermore, we utilized MLN4924, an inhibitor of the NEDD8-activating enzyme (NAE), to investigate its effect on RNF187 expression. MLN4924 treatment decreased RNF187 protein but not mRNA levels, indicating post-transcriptional regulation (Figs. 3H and S4C, S5C). Building on the spatial and single-cell analyses, we next sought to investigate the molecular mechanisms underlying the interaction and reciprocal regulation between RNF187 and NEDD8. Co-immunoprecipitation (Co-IP) assays confirmed the interaction between RNF187 and NEDD8 in HET293T, PANC-1, and SW1990 cells (Fig. 3I). Besides, imaging analyses demonstrated colocalization of RNF187 and NEDD8 in PANC-1 cells (Fig. 3J). Further Co-IP experiments indicated that the RNF187 RING domain specifically binds to NEDD8 (Fig. 3K). These results collectively indicate that the interaction between RNF187 and NEDD8 is mediated through the RNF187 RING domain, suggesting that NEDD8 may upregulate RNF187 at the post-translational level, pointing to a mechanism of protein-level regulation rather than gene expression.
A Spatial transcriptomics analysis demonstrates the co-localization of RNF187 and NEDD8. B Single-cell RNA sequencing reveals NEDD8 expression levels in pancreatic cancer tissues stratified by high and low RNF187 expression. C Analysis of NEDD8 expression in normal tissues (n = 171) and pancreatic cancer tissues (n = 179) using data from GEPIA. D Correlation analysis between RNF187 and NEDD8 expression levels from GEPIA. E Single-cell RNA sequencing analysis of RNF187 expression levels in pancreatic cancer tissues stratified by high and low NEDD8 expression. F Western blotting assay demonstrates NEDD8 protein levels in transfected PC cells. G Western blotting shows RNF187 protein levels in PC cells transfected with NEDD8 siRNA and NEDD8 overexpression plasmids. H Western blot analysis of RNF187 expression following 24 h treatment with MLN4934. I Co-immunoprecipitation (Co-IP) assays in HEK293T, PANC-1, and SW1990 cells confirm the physical interaction between RNF187 and NEDD8. J Fluorescence microscopy confirms the co-localization of RNF187 and NEDD8 in cells. K Co-IP mapping identifies the binding region between RNF187 and NEDD8. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
Neddylation stabilizes RNF187 by inhibiting K48-Linked ubiquitination and promotes PC progression
Since MLN4924 treatment reduced RNF187 protein levels without affecting its mRNA expression, we speculated that RNF187 may undergo neddylation and be stabilized at the post-translational level. To test this, we performed co-immunoprecipitation (Co-IP) assays, which revealed that NEDD8 could enhance RNF187 neddylation, while MLN4924 disrupted this process (Figs. 4A and S4D). This indicated that neddylation plays a crucial role in protecting RNF187 from degradation. We then assessed RNF187’s protein stability in PC cells using a cycloheximide (CHX) chase assay, which showed decreased RNF187 stability following NEDD8 knockdown and MLN4924 treatment (Figs. 4B and S4E). Given that protein degradation is primarily mediated by the ubiquitin-proteasome system (UPS) and the lysosomal pathway, we employed the proteasome inhibitor MG132 and the lysosomal inhibitor CQ to investigate the specific impact of NEDD8 on RNF187 degradation. The results demonstrated that neddylation markedly reduced RNF187 polyubiquitination, thereby protecting it from proteasome-mediated degradation, whereas degradation via the lysosomal pathway was not significant (Figs. 4C, D and S4F–G). Further analysis revealed that NEDD8 specifically suppressed K48-linked ubiquitination of RNF187, which is essential for its proteasomal degradation, while having no significant effect on K63-linked ubiquitination, indicating a chain-type-specific regulatory role (Figs. 4E and S4H). To further elucidate the mechanism by which NEDD8 regulates K48 ubiquitination, we generated a RING domain mutant of RNF187 and conducted functional validation. Previous evidence suggests that NEDD8 exerts its stabilizing effect through direct interaction with the RING domain of RNF187. Accordingly, we transfected NEDD8-overexpressing cells with either wild-type (WT) RNF187 or a RING domain mutant, along with a K48-specific ubiquitin plasmid. The results showed that in the presence of the RING domain mutation, NEDD8 failed to inhibit K48-linked ubiquitination of RNF187 (Figs. 4F and S4I). These findings indicate that the RING domain is essential for NEDD8 binding and that the suppressive effect of NEDD8 on K48 ubiquitination is dependent on its interaction with this domain.
A Cells were transfected with RNF187, NEDD8, and treated with MLN4934, and NEDD8 expression was assessed by immunoblot analysis of anti-RNF187 immunoprecipitates. B RNF187 expression in PC cells was evaluated by Western blot after treatment with MLN4934, siRNA, and CHX. C RNF187 expression in PC cells transfected with an overexpressed NEDD8 lentivirus and treated with DMSO, MG132, and CQ. D RNF187 ubiquitination assay in PC cells transfected with si-NEDD8 or overexpressed NEDD8. E Immunoblot analysis of K48R-Ub, K48-Ub, K63R-Ub, and K63-Ub immunoprecipitates using anti-RNF187 antibody. F Immunoblot analysis of K48-Ub immunoprecipitates using anti-RNF187 antibody. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
To investigate the functional role of neddylation in RNF187-mediated PC progression, we performed a series of in vitro and in vivo assays. In vitro, CCK-8, EdU incorporation, and colony formation assays demonstrated that MLN4924 treatment significantly attenuated the tumor-promoting effects of RNF187 overexpression by reducing cell proliferation (Fig. S4D–I). Similarly, wound healing and Transwell migration and invasion assays confirmed that MLN4924 effectively reversed the enhanced migratory and invasive capabilities induced by RNF187 overexpression (Fig. S4J, K and S6A–C). In vivo, xenograft models revealed that tumor volume and weight were significantly reduced in the RNF187 overexpression plus MLN4924 group compared to the RNF187 overexpression group (Fig. S6D–F). This inhibitory effect was further validated by HE and IHC, which showed decreased Ki67 expression, indicating reduced tumor cell proliferation (Fig. S6G). Notably, in a liver metastasis model, the number of metastatic lesions was markedly lower in the RNF187 overexpression plus MLN4924 group than in the RNF187 overexpression group, highlighting the essential role of neddylation in RNF187-driven metastasis (Fig. S6H–J). Collectively, these results demonstrate that neddylation is essential for the oncogenic activity of RNF187 in pancreatic cancer and that its inhibition by MLN4924 effectively impairs tumor growth and metastasis.
RNF187 Upregulates IQGAP1 Expression and Stabilization
To elucidate the potential mechanisms underlying the function of RNF187, we employed mass spectrometry (MS) to identify proteins interacting with RNF187 in PANC-1 cells. MS analysis indicated a strong correlation between IQGAP1 and RNF187 (Fig. 5A). Co-IP experiments conducted in PANC-1 and SW1990 cells substantiated the interaction between RNF187 and IQGAP1 (Fig. 5B). Bioinformatics analysis further revealed that IQGAP1 not only interacted with RNF187 but also exhibited a negative correlation with the prognosis of PC patients (Fig. 5C–E). Spatial transcriptomics analysis revealed significant spatial colocalization between RNF187 and NEDD8 within tumor epithelial cells, suggesting their potential synergistic roles in tumor-associated biological processes (Fig. S7A). Besides, imaging analyses confirmed the colocalization of RNF187 and IQGAP1 in PANC-1 and SW1990 cells (Fig. 5F). To investigate the relationship between RNF187 and IQGAP1, we employed Western blotting and RT-PCR techniques. Western blot results indicated a positive correlation between RNF187 and IQGAP1 protein expression, while q-PCR analysis showed no significant change in IQGAP1 mRNA levels in response to RNF187 modulation (Figs. 5G, H and S7B). These findings suggest that RNF187 may be implicated in the post-translational modification of IQGAP1. Further analysis using a CHX chase assay showed that RNF187 knockdown accelerated IQGAP1 protein degradation in PC cells (Fig. 5I, J), indicating RNF187’s role in stabilizing IQGAP1. This result supported the hypothesis that RNF187 contributes to the stabilization of IQGAP1 at the protein level. Furthermore, Co-IP experiments demonstrated that the RNF187 RING finger domain specifically interacted with the IQGAP1 protein, suggesting a critical role for this domain in the protein-protein interaction (Fig. 5K). These findings collectively underscore RNF187’s potential role in modulating PC malignancy, likely through the post-translational modification of IQGAP1.
A Mass spectrometry analysis identifying RNF187-binding proteins. B Co-IP assay demonstrating the interaction between RNF187 and IQGAP1 in PANC-1 and SW1990 cells. C Correlation analysis of RNF187 and IQGAP1 expression levels using data from GEPIA. D Comparative analysis of IQGAP1 expression in normal tissues (n = 171) and PC tissues (n = 179) from GEPIA. E Survival analysis based on IQGAP1 expression levels from GEPIA. F Fluorescence microscopy revealing colocalization of RNF187 and IQGAP1 in cells. G Western blot analysis of IQGAP1 protein levels in PC cells transfected with varying concentrations of RNF187 overexpression plasmids. H IQGAP1 protein levels in PC cells transfected with RNF187 siRNAs or overexpression plasmids. I, J Western blot analysis of IQGAP1 expression in PC cells treated with RNF187 siRNA and cycloheximide (CHX). K Reciprocal Co-IP assay of RNF187 truncation mutants and IQGAP1 in HEK293T cells. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
RNF187 inhibits IQGAP1 degradation via K48-K63-linked ubiquitination
Numerous prior studies have elucidated that IQGAP1 undergoes degradation via a ubiquitin-dependent process. To further investigate this, we employed the proteasome inhibitor MG132 or the lysosome inhibitor CQ to determine whether they could prevent the polyubiquitylation and degradation of IQGAP1 in RNF187 knockdown cells. Our results revealed that MG132 effectively inhibited the degradation of IQGAP1 protein in these RNF187-deficient cells (Fig. 6A). Subsequently, we explored the potential influence of RNF187 on the ubiquitination pattern of IQGAP1. Our ubiquitination assays indicated that overexpression of RNF187 led to an increase in IQGAP1 polyubiquitination, whereas its ubiquitination levels decreased upon RNF187 knockdown (Fig. 6B, C). Further investigations were conducted by transfecting 293 T cells with GFP-tagged RNF187 and Flag-tagged IQGAP1, along with HA-tagged ubiquitin constructs (HA-K6-Ub, HA-K11-Ub, HA-K23-Ub, HA-K26-Ub, HA-K48-Ub, HA-K63-Ub, HA-K48R, and HA-K63R) to delineate the specific ubiquitin linkage modulated by RNF187. Our findings demonstrated that the presence of K48R-Ub and K63, in contrast to K63R- and K48-Ub, significantly enhanced IQGAP1 ubiquitination. This suggested that RNF187 did not affect other forms of ubiquitination but specifically inhibited K48-linked and enhanced K63-linked ubiquitination of IQGAP1 (Fig. 6D). Moreover, knockdown of RNF187 led to an increase in K48-linked ubiquitination and a decrease in K63-linked ubiquitination of IQGAP1 (Fig. 6E). Collectively, these findings provide compelling evidence that RNF187 regulates the protein stability of IQGAP1 mainly through modulating K48- and K63-linked ubiquitination.
A RNF187 expression levels in PC cells treated with DMSO, MG-132, CQ, or transfected with RNF187 overexpression plasmids. B Ubiquitination levels of IQGAP1 in PC cells following transfection with si-RNF187. C Ubiquitination levels of IQGAP1 in PC cells transfected with RNF187 overexpression plasmids. D Co-transfection of GFP-RNF187 plasmid with Flag-IQGAP1, HA-K6-Ub, HA-K11-Ub, HA-K23-Ub, HA-K26-Ub, HA-K48-Ub, HA-K63-Ub, HA-K48R, and HA-K63R in HEK293T cells. E HEK293T cells transfected with siRNF187. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
IQGAP1 knockdown inhibited RNF187-induced proliferation, migration, and invasion in PC
IQGAP1 has been implicated in promoting tumor progression and metastasis in PC by regulating key signaling pathways. In this study, we conducted rescue experiments under conditions of RNF187 overexpression and IQGAP1 knockdown to investigate their respective roles in PC cell behaviors. We observed that knockdown of IQGAP1 in RNF187-overexpressing cells decreased expression of malignancy-associated proteins, including vimentin, snail, MMP9, MMP12, N-cadherin, and PCNA, while concomitantly increasing E-cadherin expression (Fig. 7A). Moreover, RNF187 overexpression significantly enhanced cell proliferation, which was reversed by IQGAP1 knockdown, as demonstrated by CCK-8, colony formation, and EdU assays (Figs. 7B–E and S7C, D). Furthermore, IQGAP1 knockdown mitigated the enhanced migratory and invasive capacities induced by RNF187 overexpression in PANC-1 and SW1990 cells (Figs. 7F–H and S7E, F). Employing an in vivo xenograft model, we investigated the functional impact of the RNF187/IQGAP1 axis on pancreatic cancer progression. In vivo studies showed that RNF187 knockdown significantly inhibited tumor growth in subcutaneous xenografts in nude mice, while IQGAP1 overexpression promoted this effect (Fig. 7I–K). Histological analyses, encompassing HE and IHC staining, confirmed a significant reduction in proliferation marker Ki67 and IQGAP1 expression in the RNF187 knockdown group, while IQGAP1 overexpression resulted in an increase in both Ki67 and IQGAP1 expression (Fig. 7L). Similarly, in the liver metastasis model, RNF187 knockout significantly reduced liver metastasis and improved survival, while IQGAP1 overexpression accelerated liver metastasis (Fig. 7M–O). Western blot analysis showed that NEDD8 upregulation promoted RNF187 expression, which subsequently enhanced IQGAP1 expression, suggesting a functional regulatory axis between NEDD8, RNF187, and IQGAP1 that contribute to the malignant phenotype of PC cells (Fig. 7P). Besides, spatial transcriptomics analysis provided further insights by revealing distinct co-expression patterns among RNF187, NEDD8, and IQGAP1 within specific regions of the pancreatic tumor microenvironment, indicating potential functional associations that contribute to pancreatic cancer progression (Fig. S7G). However, co-immunoprecipitation (CO-IP) experiments did not reveal a direct interaction between IQGAP1 and NEDD8, suggesting that while these proteins may be functionally linked, they do not physically associate in a complex (Fig. S7H).
A Western blot analysis demonstrated the expression levels of malignant tumor marker proteins in transfected PC cells. B–E The proliferative potential of transfected PC cells was assessed using CCK-8 and colony formation. F–H Transwell assays were conducted to evaluate the migratory and invasive capabilities of transfected PC cells. I Representative images of tumors from the NC, sh-RNF187, and sh-RNF187 + IQGAP1 groups (N = 5/ groups). J Tumor weights for each group. K Tumor growth for each group. L HE staining and IHC for RNF187, Ki67, and IQGAP1 in tumor sections. M Representative images of liver metastases and HE staining in the groups (N = 5/groups). N Kaplan–Meier survival analysis. O Quantification of liver metastasis foci. P Western blot analysis revealed the expression levels of NEDD8, RNF187, and IQGAP1 in transfected PC cells. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
To further elucidate the regulatory relationship, we performed additional experiments using constructs expressing full-length RNF187 (RNF187-FL) and a mutant version lacking the IQGAP1-binding domain (RNF187-RING). CCK-8, EdU, and colony formation assays demonstrated that sh-IQGAP1 + RNF187-FL significantly enhanced cell proliferation in IQGAP1-knockdown cells, whereas sh-IQGAP1 + RNF187-RING exhibited a markedly reduced effect (Fig. S8A–F). Similarly, migration and invasion assays showed that sh-IQGAP1 + RNF187-FL notably increased the migratory and invasive capacities of IQGAP1-knockdown cells compared to sh-IQGAP1 + RNF187-RING (Fig. S8G–K). In vivo, the subcutaneous xenograft model indicated that sh-IQGAP1 + RNF187-FL more effectively promoted tumor growth in IQGAP1-knockdown pancreatic cells than sh-IQGAP1 + RNF187-RING (Fig. S9A–D). Similarly, in the liver metastasis model, sh-IQGAP1 + RNF187-FL significantly enhanced the invasion and migration of cells relative to sh-IQGAP1 + RNF187-RING (Fig. S9E–F). Collectively, these results reveal the regulatory relationship among NEDD8, RNF187, and IQGAP1, and suggest that this axis plays an important role in the malignant phenotype of pancreatic cancer cells. Further rescue experiments with mutated RNF187 constructs (RNF187-FL and RNF187-RING) support the functional significance of the IQGAP1-binding domain in regulating tumor cell behaviors, reinforcing the mechanism proposed in our study.
IQGAP1 Regulates the Cytoskeleton by Modulating MYH9 Expression and Mediates the Malignant Phenotype of PC
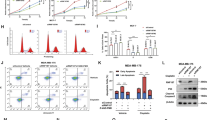
To elucidate the role of IQGAP1 in the regulation of PC, we examined differentially expressed genes involved in actin cytoskeleton regulation in IQGAP1-overexpressing PC cells using mRNA microarray analysis (Fig. S10A). This was further explored through phalloidin staining to assess the structural organization and distribution of F-actin, revealing a significant abundance of parallel F-actin fibers in cells overexpressing IQGAP1 (Fig. 8A). Moreover, we treated these IQGAP1-overexpressing PC cells with CK666, an inhibitor of actin cytoskeleton formation, and observed a consequent reduction in cellular proliferation, migration, and invasion (Figs. 8B–D and S10B–G). To further elucidate how IQGAP1 modulates the actin cytoskeleton, we integrated data from genes implicated in cytoskeletal regulation, IQGAP1-associated genes from mRNA sequencing, and related entries from the BioGRID and GeneInfo databases, identifying FN1, PTK2, and MYH9 as potential downstream targets of IQGAP1 regulation (Fig. 8E). RT-PCR and Western blot analyses confirmed that IQGAP1 knockdown resulted in decreased mRNA and protein levels of MYH9 (Figs. 8F and S11A, B), whereas IQGAP1 overexpression increased MYH9 expression (Fig. 8G). A luciferase reporter assay showed that IQGAP1 overexpression increased MYH9 promoter activity, whereas its knockdown decreased activity, confirming IQGAP1’s regulatory role in MYH9 expression (Fig. S11C). Additionally, we assessed the consistency between the Virtual Cell Model (VCM) and a post-trained Large Language Model (LLM) on a perturbation query. The VCM, part of the Rbio model, uses a two-layer MLP and ESM-2 embeddings to predict a 59.8% probability that IQGAP1 knockdown results in differential expression of MYH9 in K562 cells, suggesting a “yes” soft label. To validate this, we incorporated the soft evidence into the prompt for the LLM (Qwen-2.5-3B-Instruct, fine-tuned with LoRA using GRPO). The LLM produced a final “Yes” answer, perfectly aligning with the VCM’s prediction (Table S2). Subsequently, MYH9 knockdown in IQGAP1-overexpressing PC cells revealed that MYH9 is crucial for the formation of F-actin fibers, as evidenced by phalloidin staining (Fig. 8H–J).
A, B Visualization of F-actin using Rhodamine phalloidin staining in PC cells. C Colony formation assay to assess cell proliferation. D Transwell assay to evaluate cell migration and invasion. E Venn diagram illustrating the overlap between genes regulating the cytoskeleton, IQGAP1-related genes identified through mRNA sequencing, and IQGAP1-related genes from the Biograd and GeneInfo databases. F, G Western blot and PCR analysis demonstrating MYH9 protein levels with or without IQGAP1 overexpression. H Western blot and PCR analysis showing MYH9 protein levels with or without MYH9 knockdown. I F-actin staining using Rhodamine phalloidin in cells with or without MYH9 knockdown. J Colony formation assay following MYH9 knockdown. K Transwell assay to assess migration and invasion after MYH9 knockdown. L Western blot analysis of key proteins associated with PC malignancy in transfected PC cells. All data are expressed as the mean ± SD (three independent experiments). *p < 0.05; **p < 0.01; ***p < 0.001.
Furthermore, MYH9 knockdown impeded the proliferation, migration, and invasion of IQGAP1-overexpressing cells and was associated with the downregulation of malignancy markers such as vimentin, snail, MMP9, MMP12, N-cadherin, and PCNA, alongside the upregulation of E-cadherin (Figs. 8J–L and S11D–F). In addition, survival analysis revealed that co-expression of IQGAP1 and MYH9 was associated with poorer patient survival, suggesting their combined role in tumor aggressiveness (Fig. S11G). To further assess the clinical relevance of the RNF187/NEDD8/IQGAP1/MYH9 regulatory axis in human PC, we analyzed gene expression data from TCGA-PAAD using the GEPIA platform. All four genes were significantly upregulated in pancreatic tumor tissues compared to normal controls (P < 0.01, Figs. 1C, 3D, 5D and S11H). Co-expression analysis further revealed statistically significant positive correlations among these genes (R = 0.17–0.60, all P < 0.05; Figs. 3C, 5C and S11I–M), suggesting that they are coordinately regulated in the clinical context. Collectively, these findings suggest that the RNF187/NEDD8/IQGAP1/MYH9 signaling axis contributes to pancreatic cancer progression by regulating actin cytoskeleton dynamics and promoting malignant phenotypes.
Discussion
PC remains one of the deadliest malignancies, with limited treatment options and a poor prognosis [22, 23]. Despite the identification of significant genetic alterations, the underlying pathogenic processes that drive disease progression are not well understood at the molecular level [24]. Therefore, understanding these mechanisms is crucial for identifying potential therapeutic targets.
Neddylation impacts a multitude of biological processes by altering the stability, localization, and activity of target proteins [25, 26]. Consistent with these observations, NEDD8-activating enzyme (NAE) has been identified as a potent and druggable target in oncology. Notably, aberrant neddylation has been implicated in the pathogenesis and progression of various cancers, including PC. Pevonedistat, a selective NAE inhibitor, has demonstrated notable anticancer effects in preclinical pancreatic cancer models, including enhanced radiosensitivity, cell cycle arrest, and induction of apoptosis. [27,28,29]. These findings suggest that targeting the neddylation pathway may represent a novel therapeutic strategy in PC [30]. Moreover, MLN4924 has progressed to multiple phase I and II clinical trials in hematologic and solid tumors, such as acute myeloid leukemia and melanoma, showing manageable toxicity and preliminary efficacy. Although these findings support the therapeutic potential of targeting the neddylation pathway, the clinical application of MLN4924 remains limited by several challenges, including dose-limiting toxicities, suboptimal tumor selectivity, and variable patient responses. These limitations underscore the need to identify downstream effectors or predictive biomarkers to enhance the specificity and effectiveness of NAE-targeted therapies. In this study, we sought to identify E3 ligases associated with NEDDylation in PC. Our findings revealed that RNF187 expression was significantly elevated in PC and positively correlated with NEDD8 expression levels. Furthermore, we observed a similar suppression of RNF187 expression upon treatment with MLN4934. These findings suggest that targeting NEDD8-mediated stabilization of RNF187 may offer a valuable therapeutic strategy for PC. Notably, although mutation of the RING domain disrupted the intrinsic E3 ligase activity of RNF187 and abolished its capacity for self-ubiquitination, the level of K48-linked ubiquitination unexpectedly increased rather than decreased. We speculate that this phenomenon results from the simultaneous loss of NEDD8-mediated protection following RING domain mutation, which renders RNF187 more exposed in the cytoplasm and more susceptible to recognition by other exogenous E3 ubiquitin ligases. Based on this, we propose a regulatory model: in wild-type RNF187, NEDD8 binds to the RING domain and competitively interferes with the recruitment of E2~Ub conjugates, thereby preventing the formation of K48-linked ubiquitin chains. When the RING domain is mutated, NEDD8 can no longer bind RNF187, leading to the loss of this protective effect and increased susceptibility to degradation via K48-linked polyubiquitination by alternative E3 ligases. These findings highlight the RING domain as a critical competitive interface between NEDD8 and the ubiquitination machinery, offering novel insight into the regulation of RNF187 protein stability.
RNF187, a member of the RING finger E3 ubiquitin ligase family, is regulated by a ubiquitin switch between K48- and K63-linked polyubiquitination, modulated by MEK15 and PRMT1-mediated arginine methylation [18, 31]. Although its function has been investigated in several cancer types, the biological role of RNF187 in pancreatic cancer remains poorly defined. Our study provides new insights into the molecular mechanisms underlying RNF187’s contribution to PC progression. In this study, we demonstrated that RNF187 is markedly upregulated in PC, as evidenced by both GEPIA and TNMplot transcriptomic datasets. This finding was further validated at the protein level through IHC and Western blotting, confirming RNF187 overexpression in PC tissues and cell lines. To investigate its functional role, we performed CCK8, EdU, colony formation, transwell, and wound healing assays, which consistently showed that RNF187 significantly promotes PC cell proliferation, invasion, and migration. These comprehensive analyses collectively demonstrated that RNF187 significantly promotes these malignant phenotypes in PC cells. Previous studies have shown that NEDD8 interacts with RNF187, stabilizing it and preventing its degradation. However, this does not fully account for how RNF187 exerts its oncogenic effects in PC. To elucidate downstream effectors, we conducted mass spectrometry analysis, which identified IQGAP1 as a potential RNF187-interacting protein. Expression correlation analyses further supported a positive association between RNF187 and IQGAP1. Mechanistically, RNF187 was found to enhance K63-linked ubiquitination while suppressing K48-linked ubiquitination of IQGAP1, thereby stabilizing its expression. Collectively, our studies demonstrate that RNF187 drives PC proliferation, invasion, and metastasis both in vitro and in vivo through its interaction and modulation of IQGAP1.
IQGAP1 is a large protein (190 kDa) belonging to the IQGAP protein family [32, 33]. It comprises five key domains that mediate extensive protein-protein interactions: a calponin-homology domain, a poly-proline protein-protein domain, a domain containing four IQ motifs, a Ras GAP-related domain, and a Ras GAP C-terminal domain [34]. Consequently, it participates in various biological processes, including cell-cell adhesion, motility/invasion, proliferation, and cytoskeletal dynamics [35, 36]. Previous studies have demonstrated the crucial role of IQGAP1 in diverse cancers, including PC, colorectal cancer, and liver cancer [37,38,39,40]. Our experiments demonstrated that RNF187 promotes the malignant behavior of PC cells by modulating IQGAP1 expression. Notably, RNF187 upregulates the polyubiquitination of IQGAP1 through K63-linked ubiquitination, which leads to actin cytoskeleton remodeling via MYH9 expression. Furthermore, NEDD8 interacts with RNF187, preventing its degradation, which further enhances RNF187’s ability to upregulate IQGAP1. This mechanism is further supported by our spatial transcriptomics analysis, which revealed the co-expression of NEDD8, RNF187, and IQGAP1 in distinct regions of the pancreatic tumor microenvironment.
In conclusion, we identify RNF187 as a critical oncogenic factor in pancreatic cancer that promotes tumor proliferation, invasion, and metastasis by regulating IQGAP1 stability through K63-linked ubiquitination and activating downstream MYH9 expression. This oncogenic function of RNF187 is further enhanced by NEDD8-mediated stabilization, positioning RNF187 as both a functional effector and downstream target within the neddylation axis. These findings provide mechanistic insights into the RNF187–IQGAP1/MYH9 signaling pathway and suggest that dual targeting of RNF187 and NEDD8 may offer a promising therapeutic strategy in pancreatic cancer.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Rajkomar K, Jamieson NB. Pancreatic cancer. Surgery (Oxford International Edition) 2022;40:237–48.
Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17–48.
Yin T, Xu L, Gil B, Merali N, Sokolikova MS, Gaboriau DCA, et al. Graphene sensor arrays for rapid and accurate detection of pancreatic cancer exosomes in patients’ blood plasma samples. ACS Nano. 2023;17:14619–31.
Charles, Jacob, Signorelli HK, Charles R, Richard JL, Kashuv T, et al. Identification of novel early pancreatic cancer biomarkers KIF5B and SFRP2 from “first contact” interactions in the tumor microenvironment. J Exp Clin Cancer Res CR. 2022;41:258.
Adam MG, Beyer G, Christiansen N, Kamlage B, Pilarsky C, Distler M, et al. Identification and validation of a multivariable prediction model based on blood plasma and serum metabolomics for the distinction of chronic pancreatitis subjects from non-pancreas disease control subjects. Gut. 2021;70:2150–8.
Lin K, Zhou E, Shi T, Zhang S, Zhang J, Zheng Z, et al. m6A eraser FTO impairs gemcitabine resistance in pancreatic cancer through influencing NEDD4 mRNA stability by regulating the PTEN/PI3K/AKT pathway. J Exp Clin Cancer Res CR. 2023;42:217.
Gai W, Peng Z, Liu CH, Zhang L, Jiang H. Advances in cancer treatment by targeting the neddylation pathway. Front Cell Dev Biol. 2021;9:653882.
Mamun M, Liu Y, Geng YP, Zheng YC, Gao Y, Sun JG, et al. Discovery of neddylation E2s inhibitors with therapeutic activity. Oncogenesis. 2023;12:45.
Zhu J, Chu F, Zhang M, Sun W, Zhou F. Association between neddylation and immune response. Front Cell Dev Biol. 2022;10:890121.
Jones TM, Carew JS, Bauman JE, Nawrocki ST. Targeting NEDDylation as a novel approach to improve the treatment of head and neck cancer. Cancers. 2021;13:3250.
Kamitani T, Kito K, Nguyen HP, Yeh ET. Characterization of NEDD8, a developmentally down-regulated ubiquitin-like protein. J Biol Chem. 1997;272:28557–62.
Fu DJ, Wang T. Targeting NEDD8-activating enzyme for cancer therapy: developments, clinical trials, challenges and future research directions. J Hematol Oncol. 2023;16:87.
Zhou L, Jiang Y, Luo Q, Li L, Jia L. Neddylation: a novel modulator of the tumor microenvironment. Mol Cancer. 2019;18:77.
Enchev RI, Schulman BA, Peter M. Protein neddylation: beyond cullin-RING ligases. Nat Rev Mol Cell Biol. 2015;16:30–44.
Gonzalez-Rellan MJ, Fernández U, Parracho T, Novoa E, Fondevila MF, da Silva Lima N, et al. Neddylation of phosphoenolpyruvate carboxykinase 1 controls glucose metabolism. Cell Metab. 2023;35:1630–-45.e1635.
Davies CC, Chakraborty A, Diefenbacher ME, Skehel M, Behrens A. Arginine methylation of the c-Jun coactivator RACO-1 is required for c-Jun/AP-1 activation. EMBO J. 2013;32:1556–67.
Zhang L, Chen J, Yong J, Qiao L, Xu L, Liu C. An essential role of RNF187 in Notch1 mediated metastasis of hepatocellular carcinoma. J Exp Clin Cancer Res CR. 2019;38:384.
Chakraborty A, Diefenbacher ME, Mylona A, Kassel O, Behrens A. The E3 ubiquitin ligase Trim7 mediates c-Jun/AP-1 activation by Ras signalling. Nat Commun. 2015;6:6782.
Wang Z, Kong Q, Su P, Duan M, Xue M, Li X, et al. Regulation of Hippo signaling and triple negative breast cancer progression by an ubiquitin ligase RNF187. Oncogenesis. 2020;9:36.
Pang D, Wang W, Zhou X, Lu K, Zhang J, Chen Z, et al. RACO-1 modulates Hippo signalling in oesophageal squamous cell carcinoma. J Cell Mol Med. 2020;24:11912–21.
Istrate AM, Milletari F, Castrotorres F, Tomczak J, Torkar M, Li D, et al. rbio-1 – training scientific reasoning LLMs with biological world models as soft verifiers. bioRxiv 2025 [Preprint]. Available from: https://doi.org/10.1101/2025.08.18.670981
Hung HC, Fan MH, Wang D, Miao CH, Su P, Liu CL. Effect of chimeric antigen receptor T cells against protease-activated receptor 1 for treating pancreatic cancer. BMC Med. 2023;21:338.
Naudin S, Sampson JN, Moore SC, Albanes D, Freedman ND, Weinstein SJ, et al. Lipidomics and pancreatic cancer risk in two prospective studies. Eur J Epidemiol. 2023;38:783–93.
Fard D, Giraudo E, Tamagnone L. Mind the (guidance) signals! Translational relevance of semaphorins, plexins, and neuropilins in pancreatic cancer. Trends Mol Med. 2023;29:817–29.
Guo YJ, Pang JR, Zhang Y, Li ZR, Zi XL, Liu HM, et al. Neddylation-dependent LSD1 destabilization inhibits the stemness and chemoresistance of gastric cancer. Int J Biol Macromol. 2023;254:126801.
Shin YC, Chen JH, Chang SC. The molecular determinants for distinguishing between ubiquitin and NEDD8 by USP2. Sci Rep. 2017;7:2304.
Wei D, Li H, Yu J, Sebolt JT, Zhao L, Lawrence TS, et al. Radiosensitization of human pancreatic cancer cells by MLN4924, an investigational NEDD8-activating enzyme inhibitor. Cancer Res. 2012;72:282–93.
Xu J, Li Z, Zhuo Q, Ye Z, Fan G, Gao H, et al. Pevonedistat suppresses pancreatic cancer growth via inactivation of the neddylation pathway. Front Oncol. 2022;12:822039.
Li H, Zhou W, Li L, Wu J, Liu X, Zhao L, et al. Inhibition of neddylation modification sensitizes pancreatic cancer cells to gemcitabine. Neoplasia. 2017;19:509–18.
Tian Z, Li J, Ma R, Li T, Sun Z, Huang S. Targeting neddylation as a novel approach to lung cancer treatment (Review). Int J Oncol. 2023;62:65.
Davies CC, Chakraborty A, Cipriani F, Haigh K, Haigh JJ, Behrens A. Identification of a co-activator that links growth factor signalling to c-Jun/AP-1 activation. Nat Cell Biol. 2010;12:963–72.
Abel AM, Schuldt KM, Rajasekaran K, Hwang D, Riese MJ, Rao S, et al. IQGAP1: insights into the function of a molecular puppeteer. Mol Immunol. 2015;65:336–49.
Weissbach L, Settleman J, Kalady MF, Snijders AJ, Murthy AE, Yan YX, et al. Identification of a human rasGAP-related protein containing calmodulin-binding motifs. J Biol Chem. 1994;269:20517–21.
Brown MD, Sacks DB. IQGAP1 in cellular signaling: bridging the GAP. Trends Cell Biol. 2006;16:242–9.
Smith JM, Hedman AC, Sacks DB. IQGAPs choreograph cellular signaling from the membrane to the nucleus. Trends Cell Biol. 2015;25:171–84.
Wei T, Lambert PF. Role of IQGAP1 in carcinogenesis. Cancers 2021;13:3940.
Carmon KS, Gong X, Yi J, Thomas A, Liu Q. RSPO-LGR4 functions via IQGAP1 to potentiate Wnt signaling. Proc Natl Acad Sci USA. 2014;111:E1221–1229.
Chen F, Zhu HH, Zhou LF, Wu SS, Wang J, Chen Z. IQGAP1 is overexpressed in hepatocellular carcinoma and promotes cell proliferation by Akt activation. Exp Mol Med. 2010;42:477–83.
Jin X, Pan Y, Wang L, Ma T, Zhang L, Tang AH, et al. Fructose-1,6-bisphosphatase inhibits ERK activation and bypasses gemcitabine resistance in pancreatic cancer by blocking IQGAP1-MAPK interaction. Cancer Res. 2017;77:4328–41.
Hu W, Wang Z, Zhang S, Lu X, Wu J, Yu K, et al. IQGAP1 promotes pancreatic cancer progression and epithelial-mesenchymal transition (EMT) through Wnt/β-catenin signaling. Sci Rep. 2019;9:7539.
Acknowledgements
Thank you very much to Figdraw (www.figdraw.com) for providing the drawing software.
Funding
This work was supported by grants from the National Natural Science Foundation of China (82303333), Natural Science Foundation of Heilongjiang Province (PL2024H070), the Excellent Young Medical Talent Development Program of the First Affiliated Hospital of Harbin Medical University (2021Y11), and the Fund of Scientific Research Innovation of the First Affiliated Hospital of Harbin Medical University (2021B02).
Author information
Authors and Affiliations
Contributions
CY and XL contributed equally to this work. CY and XL performed the experiments, collected data, analyzed the results, and contributed to drafting the first manuscript; WY, LW, ZW carried out the experiments, while JL, FL, and C C participated in data analysis. XB, BS and HC designed the study, analyzed the data, provided reagents, and wrote the final version of the manuscript. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, C., Liu, X., Yang, W. et al. RNF187 neddylation in pancreatic cancer activates malignancy via IQGAP1-dependent actin cytoskeleton rearrangement. Oncogene 45, 26–41 (2026). https://doi.org/10.1038/s41388-025-03639-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-025-03639-y