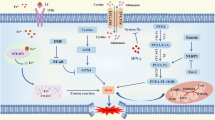
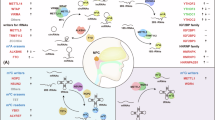
Abstract
Ferroptosis is a form of iron-dependent programmed cell death (PCD) that, once activated, inhibits tumor progression. However, the specific mechanisms influencing ferroptosis in nasopharyngeal carcinoma (NPC) are limited and warrant further investigation. This study proposes that MICAL2 not only influences the nuclear translocation of p53 but also facilitates the ubiquitination of p53 by recruiting the E3 ubiquitination ligase MDM2, leading to its degradation. This process inhibits ferroptosis in NPC cells, thereby promoting NPC progression and adversely affecting the prognosis of NPC patients. Additionally, we identified a novel long non-coding RNA (lncRNA), LINC-AC092535.5, which directly modulates the MICAL2 mRNA level through dual reinforcement at both transcriptional and post-transcriptional levels. Transcriptome sequencing and RACE experiments provided insights into the function of this new lncRNA. These findings will contribute to the expansion of the ferroptosis gene regulatory network and inform new clinical treatment strategies for NPC.

Similar content being viewed by others
Introduction
NPC is a malignancy that arises from the nasopharyngeal epithelium and is most commonly diagnosed in Southern China, Southeast Asia, and North Africa [1]. With advancements in contemporary cancer management, the survival rate of patients with non-metastatic NPC has significantly improved. However, approximately 30% of these patients experience metastasis and recurrence following standard treatment, which remains the leading cause of death among individuals with NPC [2,3,4]. These patients have developed a tolerance to conventional radiotherapy and chemotherapy, highlighting the urgent need for new treatment regimens.
Ferroptosis represents a novel form of cell death that differs from traditional mechanisms of cell death [5], which is characterized by an iron-dependent accumulation of reactive oxygen species (ROS) and primarily presents as cell shrinkage along with abnormal mitochondrial morphology, which includes reduced mitochondrial cristae, a ruptured outer mitochondrial membrane, and a condensed mitochondrial membrane [6,7,8]. Ferroptosis has been linked to cancer since the inception of the field, with numerous cancer-related genes and signaling pathways identified as modulators of ferroptosis. De-differentiated cancer cells that exhibit resistance to conventional therapies are particularly susceptible to ferroptosis inducers, especially in the cases of renal cell carcinoma, pancreatic cancer, lung cancer, and head and neck cancer [9,10,11]. These observations highlight the latent potential of inducing ferroptosis as a novel therapeutic strategy for cancer. Given that NPC cells exhibit sensitivity to ferroptosis-inducing agents [12], investigating the biological mechanisms underlying this process may reveal new therapeutic targets for the treatment of NPC.
Molecule interacting with CasL2 (MICAL2), a member of the MICAL family, is a flavin-containing monooxygenase associated with the production of ROS [13, 14]. Recent research has demonstrated that MICAL2 is highly expressed in various cancers, functioning as a novel tumor-promoting factor by participating in cell growth, vesicle trafficking, and apoptosis [15, 16]. Mechanistically, MICAL2 alters protein expression through the regulation of stability, post-translational modifications, and nuclear translocation of proteins, including components of the E-cadherin/β-catenin complex and YAP, thereby promoting epithelial-mesenchymal transition (EMT) [13,14,15,16]. So far, the expression of MICAL2, an important ROS synthase, in NPC remains unclear, as does its potential role in the occurrence of ferroptosis.
In this study, we initially employed a large clinical sample dataset in conjunction with The Cancer Genome Atlas (TCGA) database to demonstrate that MICAL2 is highly expressed in NPC and that patients with elevated MICAL2 expression exhibit a poor prognosis. Subsequently, we confirmed that MICAL2 influences ferroptosis in NPC cells by measuring the levels of ferroptosis-related ROS and Fe2+ content. We further elucidated the specific mechanism of its action through in vitro cell analyses and an in vivo BALB/c mouse tumor model. Our findings will enhance the understanding of the therapeutic potential and target population for ferritin-dependent therapy in NPC and provide new treatment strategies for patients with refractory NPC.
Materials and methods
Patients and samples
Human specimens of NPC were obtained from the Affiliated Hospital of Nantong University with approval from the Institutional Ethics Committee (Approval ID: 2021-L101). Each specimen underwent pathological confirmation as primary NPC, and written informed consent was secured from all participants. This study complies with the ethical standards outlined in the Declaration of Helsinki, the CIOMS ethical principles, the Belmont Report, and the US Common Rule. The clinical information of tissue microarrays is presented in Table S2. No treatment was received before biopsy. Tumor and normal Nasopharyngeal specimens were stored at −80 °C until preparing for RNA extraction and immunochemistry.
Cell lines and cell culture
Cell lines (NP69, CNE1, CNE2, A549, H1299, HT1080) were cultured at the Otoh Research Institute in the Department of Otorhinolaryngology Head and Neck Surgery, Affiliated Hospital of Nantong University (Jiangsu, China). All cell lines underwent validation through STR profiling and were passaged for no more than 10 generations or within 6 months of their initial revival from cryopreserved stocks. Tumor cells were cultured in RPMI 1640 (Biological Industries Israel Beit-Haemek; 01-100-1ACS) supplemented with 10% fetal bovine serum(Biological Industries Israel Beit-Haemek; 04-001-1ACS) and 100 U/mL of penicillin–streptomycin, and NP69 cells were maintained in keratinocyte-SFM (Thermo-Fisher Scientific, 17005-042), at 37 °C with 5% CO2. Before the experiment, it was confirmed that the cells were not contaminated with Mycoplasma.
Lentiviral vector transfection
Short hairpin RNAs (shRNAs) targeting human MICAL2, TP53, and LINC-AC092535.5 were constructed using lentiviral vectors, along with overexpressing lentiviruses and its negative controls. The sequences of the shRNAs targeting MICAL2 were: 5′-GCUGGGAGUUGAAAUCCAUTT-3′; The sequence of shRNA targeting TP53 was: 5′-CTAGAGACTCCAGTGGTAATCTACTTCAAGAGAGTAGATTACCACTGGAGTCTTTTTG-3′. Cells were transfected with lentiviruses for 24 h.
Western blotting
Cells were collected and lysed, and the protein concentration was measured using a BCA kit (Thermo Fisher Science; 23327). Protein samples (40 μg) were transferred to the PVDF membranes (Millipore; ISEQ00010) after the electrophoresis. ImageJ software was used to analyze the bands. Antibodies are listed in Table S1.
Real-time quantitative PCR and subcellular localization
Total RNA was collected with TRIzol reagent (Sangon Biotech; B511311). cDNA was synthesized with the Reverse Transcription Kit (Biosharp; BL699A) and qPCR was then performed in 20 μl reaction reagents using the SYBR Green PCR Master Mix (Roche; 04913914001). The following primers were used:
MICAL2: 5′-CTCACACGACACCTGGACCTA-3′(forward)
5′-CCACGCTTATCCAATTTGTACCA-3′ (reverse);
TP53:5′-TTCCTCTTCCTGCAGTACTC-3′(forward)
5′-GCCCCAGCTGCTCACCATCGCTA-3′(reverse);
GPX4:5′-GGGCTACAACGTCAAATTCG-3′(forward)
5′-TCCACTTGATGGCATTTCCC-3′ (reverse).
The 2−ΔΔCt method was used to calculate the relative expression level of mRNA and lncRNA, and was normalized to GAPDH.
Cell membrane and nuclear membrane lysis were used to achieve the separation of cytoplasmic and nuclear substances. The separated products can be used to detect the content of target RNA in the cytoplasm and nucleus through qRT-PCR method.
Immunohistochemistry (IHC)
MICAL2, GPX4, p53, MDM2 and Ki67 expression of the subcutaneous tumor tissue were analyzed by IHC. The scoring rules are as follows: staining depth is divided into four levels: 1 (negative), 2 (weak positive), 3 (medium positive), and 4 (strong positive). The staining area score is divided into four levels: 1 (0–25%), 2 (26–50%), 3 (51–75%), and 4 (>75%). The final staining score is the product (0–8 points defined as lower expression and 9–16 points defined as higher expression).In addition, Anti-Ki67 antibody (Proteintech, 27309-1-AP) was used in these studies.
Three-dimensional spheroid formation assay
Two hundred microliters of tumor cells resuspended at a density of 1 × 104cells/mL were seeded into 96-well ultra-low attachment plates. Spheroids of NPC cells formed after 48 h and then the cell culture medium was replaced with type I collagen solution. Spheroid formation was evaluated using images taken at 0 and 48 h after embedding under microscope.
Measurement the GSS/GSSG ratio in cells
Cells were inoculated in six-well plates and cultured to approximately 80% confluence and treated with Erastin (MedChemExpress; HY-15763) and Sorafenib (MedChemExpress; HY-10201) for 24 h. Cells in each group were tested using a GSH and GSSG Assay Kit (Beyotime; S0053) for GSH/GSSG ratio measurement with a Microplate reader (OD = 412 nm).
Co-immunoprecipitation (Co-IP) assay
According to the instruction, the cell lysates were prepared with the Co-IP assay kit (Thermo Fisher Scientific; 26146) and incubated with the corresponding antibody at 4 °C for 12 h. Added A/G magnetic beads to specific antibodies, cell lysate including target antibodies and incubate overnight at 4 °C. Finally, the input, IgG, and IP fractions were prepared for western blot analysis visualized using ECL chemiluminescent substrate and photographed.
BALB/c nude mice animal models
4-week-old BALB/c nude mice were randomly divided into groups. NPC cells were injected subcutaneously into the right armpit at a dose of 1 × 107/200 μl. Afterwards, Erastin (15 mg/kg), Sorafenib (20 mg/kg), and CCG-1423 (0.15 mg/kg) were intraperitoneally injected, and RG7112 (100 mg/kg) was orally administered, every 2 days until the endpoint. In the third week, the nude mice were euthanized and the tumor was completely removed. The tumor size was measured and weighed, fixed with 4% paraformaldehyde, and then embedded for subsequent IHC detection. All BALB/c nude mice were provided and approved by the Experimental Animal Center of Nantong University. All in vivo studies were conducted in accordance with the Animal Care and Use Guidelines, having received ethical approval from the Animal Experiments Committee at Nantong University (RDD number: S20210224-078).
Measurement of intracellular ROS
After collecting cells, add DCFH-DA fluorescent probes in Reactive Oxygen Species Assay Kit (Beyotime; S0033S) diluted with appropriate volume of ROS. Incubate in a 37 °C cell culture incubator for 20 min. Wash the cells three times using serum-free cell culture medium to thoroughly remove any probes that have not entered the cells. Subsequently, flow cytometry was used for detection.
Measurement of intracellular Fe2+ content
Inoculate cells on a 96 well transparent blackboard, with 1 × 104 cells per well, and incubate overnight at 37 °C in 5% CO2. Add 1 μ mol/l FerroOrange fluorescent probe working solution and place it in a 37 °C incubator for 30 min. Measure the fluorescence intensity using a multifunctional enzyme-linked immunosorbent assay.
Stability assay
1 × 107cells/2 ml were inoculated into a 6-well plate (NEST, 703001), and then 10 μM cycloheximide (CHX, Monmouth Junction, NJ, USA) was added; as well as 20 μM MG132 given at 6 h before harvest. Cell lysates were collected at 0, 2, 4, 8, and 12 h after CHX or MG132 treatment. Then, the protein expression was detected by western blotting to understand the stability of protein. Similarly, 5 g/mL actinomycin D (Act-D, Monmouth Junction, NJ, USA) was added to the above cells. The transcription level was detected by RT-qPCR to detect the RNA stability.
PI staining experiment
In the Propidium Iodide (PI) staining experiment, begin by culturing your cells until they reach the desired density. Subsequently, treat the cells as necessary to induce apoptosis or damage. Add a diluted solution of Propidium Iodide (typically 0.1–1 µg/mL) to the cells and incubate at room temperature for 15–30 min, allowing the dye to penetrate the compromised cell membranes. After incubation, wash the cells with phosphate-buffered saline (PBS) to remove any unbound dye. Finally, analyze the cells using a fluorescence microscope or flow cytometer, where live cells will appear unstained (or exhibit low fluorescence), while dead or damaged cells will emit red fluorescence due to the binding of PI to their DNA.
The dual-luciferase reporter assay
CNE2 cells were cultured under standard conditions (37 °C, 5% CO₂). For the assay, cells in the logarithmic growth phase were seeded into a 96-well plate at 0.5 × 10⁴ cells per well in 100 µL medium and incubated for 24 h. Plasmid mixtures were prepared by diluting promoter, control, and transcription factor vectors along with P3000 reagent in 5 µL OPTI-MEM, while 0.3 µL Lipofectamine™ 3000 was diluted separately in another 5 µL OPTI-MEM. After combining, the mixture was gently vortexed and incubated at room temperature for 15 min. Prior to transfection, 90 µL of fresh medium was added to each well, followed by 10 µL of the plasmid-lipid complex (total plasmid amount 50 ng/well, n = 3 replicates). Following 6 h of transfection, the medium was replaced. For detection, luciferase substrate was reconstituted with its buffer, aliquoted, and stored at −80 °C until use; Stop & Glo buffer was aliquoted and stored similarly, with its substrate added immediately before the assay. At 48 h post-transfection, the medium was aspirated, 35 µL of 1× PBS was added per well, followed by 35 µL of luciferase reagent. After 10 min of shaking, the lysate was transferred to a white 96-well plate and luminescence was measured. Subsequently, 30 µL of Stop reagent was added per well, shaken for 10 min, and the second luminescence signal was recorded.
RNA pull-down assay
The RNA pull-down assay was performed using an RNA Pull-Down Kit (Absinabs50072) according to the manufacturer’s protocol. RNA bait molecules were obtained via extraction, direct synthesis, or in vitro transcription and quantified when sequence information was available. A quarter of each RNA sample was reserved as an Input control. Streptavidin-coated magnetic beads were washed and conjugated with either biotinylated experimental or control probes (150 pmol each) through a 2-h incubation at room temperature. Following conjugation, the probe-bound beads were washed and incubated overnight at 4 °C with the bait RNA in binding buffer containing an excess of tRNA (≥30 μg) as a competitor. The captured RNA complexes were subsequently purified using Trizol-chloroform extraction and isopropanol precipitation at −80 °C. The precipitated RNA was washed multiple times with 75% ethanol, air-dried, and finally resuspended in DEPC-treated water for downstream qPCR analysis.
Sense probe primer:
A-Snse-F: TAATACGACTCACTATAGGGGGAGTCTACCAGCACGTTCC
A-Snse-R: GGGACACAGGGAAACGAGAA
A-Antisense-F: GGAGTCTACCAGCACGTTCC
A-Antisense-R: TAATACGACTCACTATAGGGGGGACACAGGGAAACGAG
AA
Statistical analysis
GraphPad Prism software (San Diego, California, USA) was applied to perform statistical analyses. The measurement results were presented as the mean ± standard deviation (SD) of repeated measurements. The Kaplan-Meier curve was used to analyze the overall survival with the log-rank test. Two-tailed Student’s t test and one-way ANOVA was used for group comparison. Image J software was applied to fluorescence intensity.
Ethical approval
This work was supported by grants from the National Natural Science Foundation of China (grant no. 82372977), Jiangsu Province excellent Youth Fund (grant no. BK20240127), Jiangsu Provincial Medical Key Discipline (Laboratory) Cultivation Unit (grant no. JSDW202244), Jiangsu Provincial Research Hospital (grant no. YJXYY202204), Postgraduate Research & Practice Innovation Program of Jiangsu Province (No. KYCX23_3429), Jiangsu Provincial Natural Science Foundation (BK20251837). Nantong Science and Technology Project (No. MS2024030), China Postdoctoral Science Foundation (No. 2025M772368).
Results
Elevated MICAL2 level in NPC alleviates ferroptosis-induced oxidative damage
From our previous study, we found that Sorafenib had poor efficacy in the treatment of NPC due to acquired ferroptosis resistance [12]. To elucidate the underlying mechanisms for drug resistance, a systematic analysis of the GEO (GSE118719 and GSE12452) database was carried out, revealing that 258 genes are significantly overexpressed in NPC tissues compared to normal nasopharyngeal tissues (Fig. S1A). Among them, MICALs, a flavin-containing monooxygenase, is linked to the generation of ROS and the enhancement of metabolic activity in cancer cells [17]. Notably, ferroptosis is characterized by the Fe2+-dependent generation of ROS in lipids. Therefore, MICAL2 was selected for subsequent investigation. Both GEO and TCGA databases indicate that MICAL2 exhibits high expression levels in NPC or head and neck squamous cell carcinoma (HNSC) (Fig. S1A, B). RT-qPCR and immunohistochemistry (IHC) revealed higher level of MICAL2 in NPC tissues than normal tissues (Fig. S1C, F). Combining these experiments, we found that MICAL2 is highly expressed in nasopharyngeal carcinoma cells, particularly in the CNE1 and CNE2 cell lines. Given that NPC is predominantly associated with Epstein-Barr virus (EBV) infection in endemic regions, such as southeastern China, we ultimately selected the CNE1, CNE2, and C666-1 cell lines for subsequent experiments (Fig. S1D, E). Given the high expression of MICAL2 in NPC cells, we proceeded to downregulate MICAL2 in these cell lines. Initially, the knockdown efficiency was verified using qRT-PCR (Fig. S2A) and Western Blot (Fig. S2B). Subsequent assays, including the CCK8 assay (Fig. S2C), colony formation assay (Fig. S2D), 3D spheroid formation assay (Fig. S2E), and EdU assay (Figure S2F), demonstrated that the knockdown of MICAL2 significantly inhibited cell proliferation. Furthermore, Transwell assays (Fig. S2G, H) and wound healing assays (Fig. S2I) indicated that cell migration and invasion were also suppressed in the shMICAL2 group. These in vitro experiments confirm that MICAL2 promotes the malignant biological behavior of nasopharyngeal carcinoma cells.
Then, to further investigate the clinical significance of MICAL2 expression and its prognostic implications, we performed IHC with tissue microarrays from 132 newly diagnosed NPC patients (Table S2). The results indicated that MICAL2 expression was significantly higher in patients with tumor node metastasis (TNM) stages III and IV compared to those with I and II (Fig. 1A). The MICAL2 staining scores from these tissue microarrays were categorized into low expression (scores of 0–9) and high expression (scores of 10–16). NPC patients exhibiting overexpression of MICAL2 demonstrated a greater malignant potential, including a higher incidence of recurrence, lymph node metastasis, and distant metastasis (Fig. 1B–E). Additionally, elevated MICAL2 expression was correlated with a poorer prognosis for patients with NPC, as demonstrated by the analysis of Overall Survival (OS), Disease-Free Survival (DFS), and Progression-Free Survival (PFS) (*P < 0.05) (Fig. 1F).
A MICAL2 staining images of NPC tissue microarray in different clinical stages (scale bar: 200 μm) and the staining scores (one-way ANOVA). B–E The relationship between MICAL2 expression and lymph node metastasis, recurrence, and metastasis in NPC patients (n = 132). F The Kaplan–Meier curves show the Overall Survival, Progression-Free Survival and Disease-Free Survival (log-rank test). G WB measured the expression of PTGS2, GPX4, FTH1 protein. H Fixable Viability Dye staining assays using sh-Ctrl or sh-MICAL2 CNE1 and CNE2 cells treated with relative inhibitors. I Flow cytometry analysis used to detect ROS content. J)Ferro-Orange probes used to detect Fe2+ content. K the ratio of intracellular GSH/GSSG content was measured in shMICAL2 cells. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
To investigate the relationship between MICAL2 expression and ferroptosis-induced oxidative damage, we examined the protein expression levels of PTGS2, GPX4, and FTH1 in NPC cells. Our study revealed that the shMICAL2 group demonstrated increased levels of ferroptosis, an effect that was further amplified when combined with the ferroptosis promoter, Erastin. (Fig. 1G). To investigate the mechanism underlying MICAL2-mediated resistance to Erastin and Sorafenib, we treated nasopharyngeal carcinoma cells with these agents in the presence or absence of regulated cell death inhibitors. The ferroptosis inhibitors ferrostatin-1 and liproxstatin-1 effectively rescued cell death induced by both Erastin and Sorafenib in shMICAL2 cells (Fig. 1H). Conversely, the apoptosis inhibitor ZVAD-FMK and the necroptosis inhibitor necrosulfonamide failed to protect against Erastin-induced death (Fig. 1H). Notably, while ZVAD-FMK partially restored viability in Sorafenib-treated cells—consistent with Sorafenib’s known ability to induce both apoptosis and ferroptosis—the ferroptosis inhibitor Fer-1 exerted a more potent protective effect. This indicates that Sorafenib primarily exerts its cytotoxic effect through ferroptosis in this context, warranting further investigation into Sorafenib-induced ferroptosis in nasopharyngeal carcinoma.
In summary, depletion of MICAL2 sensitizes human nasopharyngeal carcinoma cells to Erastin- and Sorafenib-induced cell death via the ferroptosis pathway. Furthermore, Western blot analysis of apoptosis and necroptosis marker proteins confirmed that MICAL2’s role is unrelated to these alternative cell death modalities (Fig. S6J), supporting our conclusion. Results from the fluorescent probe DCFH-DA and Ferro-Orange assays indicated that both ROS levels and intracellular Fe2+ concentrations were elevated in the shMICAL2 group, with further increases observed following the application of ferroptosis inducers (Figs. 1I, J and S2J, K). The GSH/GSSG ratio represents the proportion of reduced to oxidized glutathione within cells, serving as an indicator of cellular redox status. During ferroptosis, the levels of GSH decrease while those of GSSG increase, leading to a reduced GSH/GSSG ratio. This shift signifies an elevation in the intracellular oxidative state, thereby facilitating the occurrence of ferroptosis. The experimental results revealing that the knockdown of MICAL2 resulted in a significant reduction of this ratio (Fig. 1K). The above data indicate that, MICAL2 is highly expressed in NPC and alleviates ferroptosis-induced oxidative damage.
Knockdown of MICAL2 promotes sensitivity to ferroptosis inducers and inhibits tumor progression
In order to further verify that in addition to its own cancer-promoting role, MICAL2 can further promote the progression of NPC by inhibiting the occurrence of ferroptosis. PI cannot pass through the membrane of living cells, but it can pass through the membrane of dead cells and stain the nucleus, which is often used to quantify cell death. Our results demonstrated that the percentage of PI-positive cells increased following MICAL2 knockdown, and MICAL2 knockdown enhanced Erastin- and Sorafenib-induced PI+ dead cells (Fig. 2A, B). Additionally, the Fixable Viability Dye staining assays (Fig. 2C) and CCK8 assay (Fig. 2D) demonstrated that NPC cells lacking MICAL2 are more sensitive to ferroptosis inducers, such as Erastin and Sorafenib, resulting in increased cell mortality. Furthermore, the Fixable Viability Dye staining assays (Fig. 2C) confirmed that this effect can be reversed by the application of ferroptosis inhibitors. Based on the in vitro results of proliferation, different expression levels of MICAL2 cells were inoculated subcutaneously into nude mice. We treated the mice with Erastin or Sorafenib until the end of the experiment. After three weeks, we removed the subcutaneous tumors and measured their weight and volume. The tumors in the shMICAL2 group had a lower volume and weight than those in the control group, while the ferroptosis inducers (Erastin and Sorafenib) further inhibited tumorigenesis (Figs. 2E-G, 2I-K). Moreover, the combined treatment with the inducer and shMICAL2 significantly reduced tumor size, suggesting that the low expression of MICAL2 in vivo enhances sensitivity to ferroptosis inducers. In addition, IHC tests of the xenografts showed that lower MICAL2 expression had lower levels of proliferation markers Ki67 and ferroptosis indicators, such as GPX4 (Figs. S3, and 2H, L). Through these in vivo and in vitro experiments, we demonstrated that knocking down MICAL2 can increase the sensitivity of ferroptosis-inducing agents Erastin and Sorafenib, thereby inhibiting the progression of NPC.
A, B PI staining was used to detect cell mortality after treatment with Erastin and Sorafenib (scale: 100 μm). C, D Trypan blue staining and CCK8 cytotoxicity assay to analyze the inhibitory effects of Erastin and Sorafenib on cell growth, as well as the growth rescue effect of Fer-1. E Nude mice were subcutaneously injected with NPC cells and then intraperitoneally injected with Erastin and Sorafenib. Representative subcutaneous xenografts of CNE1 cells in the indicated group of mice (n = 6 per group). Tumor weight (F) and the volume (G) was recorded at the endpoint of experiment. H The expression of KI67, GPX4, and p53 in CNE1 cell were measured with IHC. the staining scores were observed. I Representative subcutaneous xenografts of CNE2 cells in the indicated group of mice (n = 6 per group). Tumor weight (J) and the volume (K) was recorded at the endpoint of experiment. L The expression of KI67, GPX4, and p53 in CNE2 cell were measured with IHC. Representative the staining scores were observed. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
A novel lncRNA: LINC-AC092535.5 regulates MICAL2 expression through dual reinforcement at both transcriptional and post-transcriptional levels
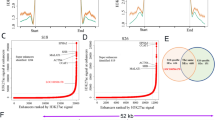
To further elucidate the regulatory mechanism of MICAL2 expression, we utilized the data in Cancer Genome Atlas (TCGA) database. Using the principle of base complementarity, we identified a novel lncRNA, LINC-AC092535.5, which exhibits the highest expression in NPC among the predicted lncRNAs (Fig. 3A, B). According to the TCGA database, LINC-AC092535.5 is highly expressed in HNSC tissues (Fig. 3C). When compared to normal nasopharyngeal epithelial cells, CNE1 and CNE2 demonstrated the highest expression levels of LINC-AC092535.5, which aligns with the observed overexpression of MICAL2 (Figs. 3D and 1D). To investigate the correlation between LINC-AC092535.5 and MICAL2 expression, LINC-AC092535.5 was knockdown in NPC cells using lentiviruses (shLINC-AC092535.5) (Fig. S6H, I), and we confirmed that MICAL2 mRNA level was significantly reduced in the shLINC-AC092535.5 group (Fig. 3G). In this study, we investigate the direct interaction between LINC-AC092535.5 and MICAL2 mRNA, as illustrated in the accompanying figure, which presents the predicted base pairing sequence between LINC-AC092535.5 and MICAL2 mRNA (Fig. 3E). Analysis of data from the TCGA database revealed a positive correlation between the expression levels of LINC-AC092535.5 and MICAL2 in tumor tissues (R: 0.2507, P < 0.001) (Fig. 3F). To elucidate the direct regulatory relationship between the identified LINC-AC092535.5 and MICAL2, we first conducted an RNA pull-down assay using in vitro transcribed, biotinylated LINC-AC092535.5. Quantitative PCR (qPCR) revealed a specific and robust enrichment of MICAL2 mRNA in the LINC-AC092535.5 captured fraction compared to the antisense control, demonstrating a physical interaction between the LINC-AC092535.5 and MICAL2 mRNA complex (Fig. 3I). Given the notably higher abundance of this interaction observed in CNE2 cells, we selected this model for functional validation. A dual-luciferase reporter assay was constructed by cloning the MICAL2 gene promoter upstream of the firefly luciferase gene. Co-transfection of this reporter with the LINC-AC092535.5 overexpression plasmid in CNE2 cells resulted in a significant increase in promoter-driven luciferase activity, indicating that the LINC-AC092535.5 functions as a transcriptional activator of MICAL2 (Fig. 3J). To determine whether this regulation extends to the post-transcriptional level, we employed Actinomycin D (Act-D) chase experiments. Upon inhibition of de novo RNA synthesis by Act-D, the decay rate of MICAL2 mRNA was markedly faster in shLINC-AC092535.5 cells compared to controls (Fig. 3H). The calculated mRNA half-life was significantly extended, confirming that the LINC-AC092535.5 enhances the transcriptional stability of MICAL2 mRNA. Consequently, we conclude that LINC-AC092535.5 directly upregulates the expression of MICAL2 in NPC. Our data delineate a dual-layer reinforcing mechanism through which this lncRNA orchestrates MICAL2 expression. Primarily, it acts as a transcriptional co-activator, significantly enhancing the promoter activity of the MICAL2 gene. Concurrently, it establishes a post-transcriptional safeguard by directly or indirectly interacting with the MICAL2 mRNA to decelerate its turnover. This model of “transcriptional and post-transcriptional reinforcement” ensures a potent and sustained upregulation of MICAL2, a master regulator of cytoskeletal dynamics and redox homeostasis. Such a robust, multi-level regulatory circuit may be crucial for amplifying the oncogenic signal output of the MICAL2 pathway, ultimately driving ferroptosis resistance and aggressive phenotypes in NPC. Thus, this lncRNA emerges as a critical upstream modulator and a potential therapeutic target within the MICAL2-centric oncogenic network. To elucidate the structure and biological properties of the novel lncRNA, we visualized the RNA sequence of LINC-AC092535.5 using rapid amplification of cDNA ends (RACE) (Fig. 3M) [18]. This sequence is slightly shorter than the conventional lncRNA sequence; however, upon re-verification, it has been confirmed that the structure of this lncRNA is complete and functional (Its expression can affect the malignant biological behavior of nasopharyngeal carcinoma, Fig. S5), enabling it to exert biological effects.
A TCGA database was used to predict the upstream molecule of MICAL2. B Heatmap of differently expression genes in NPC. C Expression level of LINC-AC092535.5 in HNSC tumor tissue based on the TCGA database. D RT-qPCR was used to detect the LINC-AC092535.5 content in NP-69 and NPC cell lines. E Schematic diagram of base complementary pairing between MICAL2 mRNA and LINC-AC092535.5. F TCGA database predicts LINC-AC092535.5 correlation with MICAL2 mRNA. G RT-qPCR result indicated knockdown efficiency of shLINC-AC092535.5 in CNE1 and CNE2. H RT-qPCR detected the stability of MICAL2 mRNA. I RNA pull-down assay detects the direct interaction between LINC-AC092535.5 and MICAL2. (J) Dual-luciferase reporter assay for the transcriptional regulation of MICAL2 by LINC-AC092535.5. K Transcript abundance of cytoplasmic and nuclear RNA fractions for LINC-AC092535.5 in CNE1 and CNE2. I The abundance of MICAL2 mRNA from RNA-seq dataset of shCtrl and shLINC-AC092535.5 group. (L) FISH to determine the intracellular localization of LINC-AC092535.5. (M) RACE technique was used to obtain the LINC-AC092535.5 transcript sequence. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
As a newly discovered lncRNA, the biological properties of LINC-AC092535.5 warrant further investigation. First, it is essential to clarify the intracellular localization of LINC-AC092535.5. Nuclear-cytoplasmic separation and fluorescence in situ hybridization (FISH) experiments revealed that LINC-AC092535.5 is predominantly located in the nucleus (Fig. 3K, L). To further validate the intracellular localization of LINC-AC092535.5 and its impact on ferroptosis, whole transcriptome RNA sequencing was conducted on shCtrl and shLINC-AC092535.5 cells. Compared to the shCtrl group, the transcription level of MICAL2 was significantly reduced in the shLINC-AC092535.5 group (Fig. 4A). Subsequent Gene Ontology (GO) analysis of the RNA sequencing results confirmed that LINC-AC092535.5 is primarily distributed in the nucleus (Fig. 4B). In summary, LINC-AC092535.5, as a new lncRNA, regulates MICAL2 mRNA stability at the post-transcriptional level.
A Transcriptome sequencing to detect the correlation between LINC-AC092535.5 and MICAL2 expression. B Results of GO enrichment analysis. (C) KEGG enrichment analysis of the pathways. D Level of GPX4 mRNA expression in CNE1 and CNE2. E PI staining suggested cell mortality rate.F, G Colony formation assay (F) and CCK8 cytotoxicity assay to analyze cell proliferation ability. (H) Flow cytometric analysis of ROS content. I Ferro-Orange fluorescence probe for the detection of intracellular Fe2+ content. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, * * * P < 0.001, ****P < 0.0001.
LINC-AC092535.5-MICAL2 axis affects ferroptosis
Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis revealed a significant correlation between LINC-AC092535.5 and ferroptosis-related signaling pathways (Fig. 4C). To elucidate the relationship between LINC-AC092535.5 and ferroptosis, results from RT-qPCR indicate that shLINC-AC092535.5 significantly inhibits the expression of the ferroptosis marker GPX4 mRNA (Fig. 4D), Knocking down LINC-AC092535.5 elevated the mortality rate of NPC cells, an effect that could be mitigated by Fer-1 (Fig. 4E). Furthermore, we observed that overexpression of MICAL2 reversed the impact of shLINC-AC092535.5 on the proliferative capacity of NPC cells, as demonstrated by CCK8 and colony formation assays (Fig. 4F, G). To investigate the effects of MICAL2 overexpression in conjunction with LINC-AC092535.5 knockdown on ferroptosis in NPC cells, we conducted measurements of intracellular Fe2+ and ROS content. The results indicated that overexpression of MICAL2 partially counteracted the beneficial effects of LINC-AC092535.5 knockdown on Fe2+ and ROS levels (Fig. 4H, I). In summary, LINC-AC092535.5 inhibits ferroptosis in NPC cells by regulating MICAL2 expression, thereby promoting cell growth and invasiveness.
Knockdown of MICAL2 or LINC-AC092535.5 upregulates and stabilizes p53
Next, we investigate the underlying cell signaling pathways associated with MICAL2-related ferroptosis resistance. Based on the TCGA database, Gene Set Variation Analysis (GSVA) functional enrichment analysis revealed that the p53 pathway was suppressed in the high MICAL2 expression group (Fig. 5A). Co-immunoprecipitation (Co-IP) experiments confirmed the direct interaction between MICAL2 and p53 (Fig. 5B). The Western blot assays clearly demonstrate that the expression of the p53 protein is negatively correlated with MICAL2 in NPC cell lines, as well as in various tumor cell lines, including human non-small cell lung cancer H1299 cells, which are naturally p53-deficient (Fig. 5C). Furthermore, the knockdown of MICAL2 leads to an increase in p53 expression in mouse tumor tissues (Fig. S3). And MICAL2 deficiency does not influence p53 transcription; but result in an increased expression of p53 protein as demonstrated by Western blot (Fig. 5D, E).
A GSVA analysis for pathways Related to MICAL2 expression (pan-cancer). B Co-IP was performed to validate protein-protein interactions between MICAL2 and p53 protein. C WB analysis of MICAL2 and p53 protein content in CNE1, CNE2, 5-8 F, 6-10B, NP69, HT1080, A549, H1299, and MCF7 cells. D RT-qPCR analysis of the p53 mRNA abundance in CNE1, CNE2 and C666-1 cells. E Western blot results demonstrated the knockdown effect of MICAL2 on cell proliferation ability. F RT-qPCR detected the stability of p53 mRNA. G Western blot analysis of p53 to analyze the stability of p53 protein treated with CHX (10 μg/mL). H Western blot detected the p53 expression in CNE1 and CNE2 cells treated with MG132 (1 μmol/ml). I Western blot showed the level of p53, p21, SLC7A11, GPX4, FTH1 expression. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
P53, often referred to as the “guardian of the genome,” primarily functions as a transcription factor that regulates essential processes including the cell cycle, DNA repair, and apoptosis. As a crucial tumor suppressor, p53 was reported to influence ferroptosis at both the transcriptional and translational levels. There are two distinct regulatory mechanisms through which wild-type p53 affects ferroptosis: one involves the abrogation of xCT inhibition, while the other entails the direct inhibition of GPX4 [19]. Previous studies have indicated that p53 can enhance ferroptosis by inhibiting the expression of Solute Carrier Family 7 A Member 11 (SLC7A11), a known cystine/glutamate antiporter. Furthermore, p53 can induce the expression of CDKN1A/p21, thereby inhibiting ferroptosis. Our western blot analysis revealed that the knockdown of MICAL2 affected the expression of downstream molecules within the p53 pathway. Specifically, p21 expression was upregulated, while SLC7A11 expression was downregulated (Fig. 5E). In order to explore the role of MICAL2 affecting p53 expression, cells transfected with shMICAL2 or shCtrl were treated with actinomycin D (Act-D) to inhibit transcriptional activity or cycloheximide (CHX) to inhibit protein translation. Results showed that the presence of CHX, but not Act-D, resulted in a time-dependent induction of p53 protein expression by MICAL2 in NPC cells (Fig. 5F-G), suggesting that MICAL2 primarily affects the post-transcriptional translation of p53 rather than its transcription. To investigate whether MICAL2 reduces p53 expression by enhancing its proteasomal degradation, the proteasome inhibitor MG132 was utilized. The results show that shMICAL2 group affects the protein level of p53 upon treatment with MG132 (Fig. 5H). Next, we explored the relationship between LINC-AC092535.5 and MICAL2 in NPC ferroptosis in vitro. Silencing LINC-AC092535.5 resulted in a reduction in the expression of GPX4 and FTH1, both of which are negatively associated with ferroptosis (Fig. 5I). Additionally, silencing LINC-AC092535.5 led to an increase in p53 expression, activation of the p53-p21 axis, and inhibition of the p53-SLC7A11 axis. Western blot analysis indicated that overexpression of MICAL2 could partially counteract the inhibitory effect of shLINC-AC092535.5 (Fig. 5I). These data indicated that knockdown of LINC-AC092535.5-MICAL2 axis upregulates p53 and stabilizes p53.
MICAL2 increases MDM2-induced ubiquitination degradation and promotes p53 nuclear export
The stabilization of p53 is closely linked to post-translational modifications, including ubiquitination and phosphorylation. Furthermore, Western blot analysis confirmed that MICAL2 did not affect the phosphorylation levels of S15 and S20 of p53, which are recognized as the most extensively studied phosphorylation sites of p53 (Fig. 6A). Therefore, exploring the specific mechanism of MICAL2 affecting p53 ubiquitination modification is our next focus. To predict ubiquitination substrates, we investigated seven known protein substrates of p53 using the UbiBrowser database in conjunction with MICAL2 expression (Fig. 6B). The MDM2 protein, which acts as an antagonist of p53, physically interacts with this tumor suppressor [20]. MDM2 functions as an E3 ligase for p53 by facilitating the binding of E2-linked ubiquitin molecules. Western blot analysis confirmed that the absence of MICAL2 significantly inhibited the ubiquitination modifications of p53 (Fig. 6C, D). Silencing MDM2 compensated for the reduction in p53 expression and inhibited the ubiquitination-mediated degradation of p53 in the oeMICAL2 group (Fig. 6E–G). Furthermore, immunofluorescence experiments demonstrated that silencing MDM2 can restore the co-localization of p53 and MDM2 in oeMICAL2 cells (Fig. 6H). In summary, the silencing of MICAL2 reduces the ubiquitination modification of p53 by interacting with MDM2, thereby decreasing the degradation of the p53 protein.
A Western blot analysis of phosphorylated S15 p53, phosphorylated S20 p53 and mutant p53 content. B Venn diagram of predicted ubiquitination substrates with the UbiBrowser database. C Co-IP experiments were conducted to verify the protein-protein interactions of MICAL2 with p53 and MDM2. D Western blot was used to detect the level of p53 ubiquitination modification. E, F Western blot was used to detect the MDM2, p53 and MICAL2 protein content. G Western blot showed the ubiquitination modification level of p53. H Immunofluorescence detection showed co-localization of MICAL2, p53 and MDM2. Representative images were captured under confocal microscopy (left, scale: 5 μm) and the average fluorescence intensity was analyzed (right). I The nuclear-cytoplasmic separation experiment verifies the intracellular location of p53 protein. J RT-qPCR showed the p53 mRNA content in the nucleus and cytoplasm. K Immunofluorescence staining (IF) showed the intracellular localization of p53 protein, and representative images were captured under confocal microscopy. (Scale: 25 μm). Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
The nuclear localization of p53 is a prerequisite for its activity; thus, investigating its nuclear-cytoplasmic localization is crucial for enhancing our understanding of its role within cells. Previous studies suggest that ubiquitination of p53 by Mdm2 could induce p53 translocation from nucleus to cytoplasm. Subsequently, we investigated whether MICAL2 influences the intracellular localization of p53. To this end, cytoplasmic and nuclear proteins were isolated from NPC cells. The separation of nuclear and cytoplasmic proteins revealed that p53 in the shMICAL2 group was predominantly localized in the nucleus, while the cytoplasmic levels of p53 were significantly reduced (Fig. 6I, J). Additionally, the results of the immunofluorescence assay corroborate these findings (Fig. 6K). These data indicate that the knockdown of MICAL2 facilitates the nuclear translocation of p53.
To further demonstrate that MICAL2 regulates ferroptosis levels in NPC and influences cancer progression through the p53 pathway, we employed a subcutaneous xenograft model in which the selective small molecule inhibitor CCG-1423 targeting MICAL2 and the MDM2 inhibitor RG7112 were administered to nude mice. Both CCG-1423 and RG7112 effectively inhibited subcutaneous tumorigenesis in these mice and exhibited a synergistic effect when combined with the ferroptosis inducer Erastin. IHC analyses revealed that the administration of MICAL2 and MDM2 inhibitors resulted in decreased levels of the proliferation marker Ki67 and the ferroptosis indicator GPX4, while the intensity of p53 staining was notably elevated. These observations were corroborated in CNE1 and CNE2 cell lines (Fig. S4). In summary, MICAL2 enhances p53 ubiquitination by recruiting MDM2 and promotes p53 nuclear export, thereby modulating the ferroptosis levels in NPC and facilitating cancer progression.
MICAL2 is vital for p53-mediated ferroptosis in pan-cancer
To assess whether p53 is a crucial protein for MICAL2’s influence on cellular ferroptosis, we conducted in vitro cell experiments to investigate this relationship from three perspectives. Initially, we established a lentiviral-mediated knockdown of MICAL2 and/or TP53 in the CNE1 and CNE2 cell lines. Western blot analysis revealed an increase in SLC7A11 expression and a decrease in MDM2 expression in the shTP53 group (Fig. 7A). PI staining indicated that shTP53 mitigated cell death induced by MICAL2 deficiency (Fig. 7B). Furthermore, we observed that silencing p53 reversed the inhibitory effect of shMICAL2 on the proliferation of NPC cells, as demonstrated by CCK8 and colony formation assays (Fig. S6A, B). Detection of ROS, Fe2+, and GSH/GSSG confirmed that silencing TP53 inhibited ferroptosis and the suppressive effect of shMICAL2 (Figs. 7C, D and S6C). Our findings suggest that MICAL2 restrains ferroptosis in NPC cells through the p53 pathway. Subsequently, we selected the natural p53-deficient cell line H1299 (a human lung adenocarcinoma cell line) along with the wild-type p53-retaining cell lines A549 (human non-small cell lung cancer cells) and HT1080 (human fibrosarcoma cells) for our experiments. As demonstrated in Fig. 5C, the expression of p53 protein in HT1080, H1299, and A549 is negatively correlated with MICAL2. We established MICAL2 knockdown cell lines using lentiviruses (Fig. S6D). The results indicate that the growth inhibitory effect observed in the shMICAL2 group is associated with ferroptosis (Fig. 7E). Additionally, the shMICAL2 group exhibits reduced proliferation ability (Fig. S6E, F). Notably, ROS levels and intracellular Fe2+ content were elevated in the shMICAL2 group, with further increases observed upon treatment with ferroptosis inducers (Fig. 7F, G). We demonstrate that p53 is a critical factor in the effect of MICAL2 on ferroptosis in various cancers. In order to further verify this conclusion, we finally selected the H1299 cell line as the research object. H1299 cells are naturally deficient in p53 and exhibit higher expression levels of MICAL2 compared to other cell lines. To precisely elucidate the molecular events regulated by MICAL2 expression, we developed a Tet-on inducible system in the lung cancer cell line [21,22,23]. We demonstrated in a unique site-directed knock-in H1299 model (H12993KR) that p53 expression is tightly regulated by doxycycline (Dox) (Fig. 7H). According to PI staining, Dox mitigated the lethality associated with the absence of MICAL2 (Fig. 7I). Furthermore, results from the colony formation assay indicated that Dox-induced p53 expression counteracted the inhibitory effect of overexpressed MICAL2 (oeMICAL2) on the proliferation of H1299 cells (Fig. S6G). The protective effects of Dox-induced p53 expression against cell ferroptosis, which was inhibited in the oeMICAL2 group, were confirmed by measuring ROS and Fe2+ content (Fig. 7J, K). Thus, the MICAL2-p53 axis was validated as playing a role in the regulation of ferroptosis in pan-cancer to a certain extent. Taken together, these data indicate that MICAL2 is required for p53-dependent ferroptosis. Furthermore, the LINC-AC092535.5-MICAL2-p53 axis was identified as a critical regulator of ferroptosis in NPC (Graphical abstract).
A The expression of SLC7A11 and MDM2 was detected through Western Blot assay. B Cell mortality was detected with PI staining in NPC cells. C Flow cytometry was used to detect the intracellular ROS content in NPC cells. D Ferro-Orange probe was used to detect the content of Fe2+ in cells in NPC cells. E PI staining suggested cell mortality rate in cells related to p53 expression. F Flow cytometry was used to detect the intracellular ROS content in cells related to p53 expression. G Ferro-Orange probe was used to detect the content of Fe2+ in cells in cells related to p53 expression. H Level of p53 expression in H1299 cells cultured in the presence of doxycycline (1 µg/ml). (I) PI staining results demonstrated the cell proliferation ability in H1299 cells. J Flow cytometry was used to detect the intracellular ROS content. (K) Ferro-Orange probe was used to detect the content of Fe2+ in cells. Data sets the mean ± SD of at least three independent experiments: *P < 0.05, **P < 0.01, * * * P < 0.001, ****P < 0.0001.
Discussion
It has been reported that over 80% of the human genome can be transcribed; however, less than 2% of these transcripts are protein-coding mRNAs, while the remainder consists of non-coding RNAs. These lncRNAs can regulate gene expression at various levels, including epigenetic, transcriptional, and post-transcriptional mechanisms, and are involved in essential biological processes such as cell growth, differentiation, apoptosis, and nuclear transport [24]. Recent developments in technology have led to the discovery of amount new lncRNAs, but their annotations are insufficient. In the present study, we identified LINC-AC092535.5, a previously uncharacterized functional lncRNA for NPC. Our research indicates that LINC-AC092535.5 is closely associated with head and neck squamous cell carcinoma, particularly overexpressed in NPC. LINC-AC092535.5 inhibits ferroptosis in NPC cells by regulating MICAL2 expression, thereby promoting cell growth and invasiveness, which offers a potential new target for the treatment of nasopharyngeal carcinoma.
LncRNAs exert their regulatory functions through four primary mechanisms [25]: (1) Signaling, where they act as molecular indicators of cellular states, such as XIST in X-chromosome inactivation; (2) Decoy, where they sequester other biomolecules (e.g., miRNAs or proteins) to inhibit their activity, exemplified by HOTAIR binding to PRC2 and the classic competitive endogenous RNA (ceRNA) interaction. Research conducted by Zi-Qi Zheng et. revealed that lncRNA FAM225A FAM225A functions as a ceRNA by binding to the response elements of miR-590-3p and miR-1275, inhibiting the regulatory effect of miR-590-3p/miR-1275 on the target gene ITGB3, consequently promoting the growth and metastasis of NPC [26]); (3) Guide, where they direct ribonucleoprotein complexes to specific genomic loci to regulate gene expression, as observed with MALAT1 in RNA splicing; and (4) Scaffold, where they serve as platforms for assembling multiple proteins into functional complexes, as illustrated by NEAT1 in paraspeckle formation. Together, these mechanisms enable lncRNAs to fine-tune gene expression, chromatin remodeling, and cellular organization. Different from the above mechanisms, our study demonstrates that LINC-AC092535.5 is localized in the nucleus and can directly bind to the mRNA of MICAL2, thereby influencing its transcription and post-transcriptional stability.
For decades, apoptosis induction has served as a cornerstone of cancer therapy. Yet, the emergence of resistance mechanisms has challenged its effectiveness. Ferroptosis, a distinct form of regulated cell death, has recently gained attention as a promising alternative to overcome apoptosis resistance. Combining ferroptosis activation with established treatments—including immunotherapy or radiation—could expand options for overcoming tumor resistance and enhancing anticancer efficacy. Notably, Ferumoxytol and C’ dots, both approved by the Food and Drug Administration (FDA), have demonstrated specificity and efficacy in inducing ferroptosis in tumor cells and show potential for application in human tumor treatment [27, 28]. Sorafenib, an FDA-approved drug that has been clinically used to treat refractory renal cell carcinoma and hepatocellular carcinoma, has recently been widely recognized for its role in regulating tumor ferroptosis [29, 30]. Our recent research also suggests that Sorafenib can induce ferroptosis in NPC cells, indicating that NPC patients may benefit from ferroptosis therapy [12]. However, ferroptosis-inducing drugs not yet been integrated into the standard treatment regimen for NPC. This may be attributed to the presence of factors that confer resistance to ferroptosis. Our research has identified that MICAL2, which is highly expressed in NPC, contributes to this resistance. Therefore, the combination of MICAL2 inhibitors with ferroptosis inducers may represent a novel therapeutic strategy for NPC. Determining whether MICAL2 functions through its enzymatic activity or as a scaffold represents an important future direction and a key challenge that we need to investigate and address in our upcoming research.
The TP53 gene is the first tumor suppressor gene to be discovered and remains the most extensively studied among such genes. As a “housekeeping gene,” p53 is influenced by various factors and exists in a low-activity state in NPC [31]. When exogenous stimulation is applied to modify p53 levels or activate p53, the cell cycle is arrested, apoptosis is increased, and both the growth and drug resistance of NPC can be managed [32, 33]. Recent studies indicate that, in addition to mediating apoptosis and autophagy, the p53 gene plays a role in regulating ferroptosis. Specifically, p53 inhibits cellular responses to cystine by downregulating SLC7A11, a crucial component of the cystine/glutamate transporter [34]. The uptake of amino acids influences ROS responses, thereby mediating ferroptosis. Jin Pinglu and colleagues demonstrated that MICAL2 influences the proliferation and apoptosis of colon cancer cells by regulating the ubiquitination of p53, thereby promoting the progression of malignant tumors [35]. This finding suggests that MICAL2 can modulate p53 expression. Additionally, we have shown that MICAL2 affects ferroptosis in NPC by regulating p53 ubiquitination levels. p53 is often maintained at a low, undetectable level in normal cells for the short half-life. Our findings demonstrate, for the first time, that MICAL2 influences protein stability by regulating the ubiquitination and degradation of P53. Interfering with MICAL2 expression is expected to enhance TP53 levels and inhibit the progression of NPC.
In conclusion, we have identified LINC-AC092535.5-MICAL2-p53 axis as a novel component for ferroptosis resistance. Inhibition of LINC-AC092535.5 or MICAL2 to enhance p53 stability offers a promising new therapeutic approach for NPC treatment.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Material or contact the corresponding author upon reasonable request.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Chen L, Zhang Y, Lai SZ, Li WF, Hu WH, Sun R, et al. 10-Year results of therapeutic ratio by intensity-modulated radiotherapy versus two-dimensional radiotherapy in patients with nasopharyngeal carcinoma. Oncologist. 2019;24:e38–e45.
Mao YP, Tang LL, Chen L, Sun Y, Qi ZY, Zhou GQ, et al. Prognostic factors and failure patterns in non-metastatic nasopharyngeal carcinoma after intensity-modulated radiotherapy. Chin J Cancer. 2016;35:103.
Blanchard P, Lee A, Marguet S, Leclercq J, Ng WT, Ma J, et al. Chemotherapy and radiotherapy in nasopharyngeal carcinoma: an update of the MAC-NPC meta-analysis. Lancet Oncol. 2015;16:645–55.
Diao J, Jia Y, Dai E, Liu J, Kang R, Tang D, et al. Ferroptotic therapy in cancer: benefits, side effects, and risks. Mol Cancer. 2024;23:89.
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE, et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell. 2012;149:1060–72.
Stockwell BR, Friedmann Angeli JP, Bayir H, Bush AI, Conrad M, Dixon SJ, et al. Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell. 2017;171:273–85.
Mou Y, Wang J, Wu J, He D, Zhang C, Duan C, et al. Ferroptosis, a new form of cell death: opportunities and challenges in cancer. J Hematol Oncol. 2019;12:34.
Xie Y, Hou W, Song X, Yu Y, Huang J, Sun X, et al. Ferroptosis: process and function. Cell Death Differ. 2016;23:369–79.
Guo J, Xu B, Han Q, Zhou H, Xia Y, Gong C, et al. Ferroptosis: a novel anti-tumor action for cisplatin. Cancer Res Treat. 2018;50:445–60.
Ma S, Dielschneider RF, Henson ES, Xiao W, Choquette TR, Blankstein AR, et al. Ferroptosis and autophagy induced cell death occur independently after siramesine and lapatinib treatment in breast cancer cells. PLoS One. 2017;12:e0182921.
Xue Z, Xie H, Shan Y, Zhang L, Cheng L, Chen W, et al. NAT10 inhibition promotes ac4C-dependent ferroptosis to counteract sorafenib resistance in nasopharyngeal carcinoma. Cancer Sci. 2024;115:3256–72.
Wang Y, Min P, Qi C, Zhao S, Yu M, Zhang Y, et al. MICAL2 facilitates gastric cancer cell migration via MRTF-A-mediated CDC42 activation. Front Mol Biosci. 2021;8:568868.
Qi C, Min P, Wang Q, Wang Y, Song Y, Zhang Y, et al. MICAL2 contributes to gastric cancer cell proliferation by promoting YAP dephosphorylation and nuclear translocation. Oxid Med Cell Longev. 2021;2021:9955717.
Cai Y, Lu J, Tang F. Overexpression of MICAL2, a novel tumor-promoting factor, accelerates tumor progression through regulating cell proliferation and EMT. J Cancer. 2018;9:521–7.
Wang Q, Qi C, Min P, Wang Y, Ye F, Xia T, et al. MICAL2 contributes to gastric cancer cell migration via Cdc42-dependent activation of E-cadherin/beta-catenin signaling pathway. Cell Commun Signal. 2022;20:136.
McDonald CA, Liu YY, Palfey BA. Actin stimulates reduction of the MICAL-2 monooxygenase domain. Biochemistry. 2013;52:6076–84.
Wu M, Harafuji N, O’Connor AK, Caldovic L, Guay-Woodford LM. Transcription factor Ap2b regulates the mouse autosomal recessive polycystic kidney disease genes, Pkhd1 and Cys1. Front Mol Biosci. 2023;9:946344.
Yang X, Wang Z, Zandkarimi F, Liu Y, Duan S, Li Z, et al. Regulation of VKORC1L1 is critical for p53-mediated tumor suppression through vitamin K metabolism. Cell metabolism. 2023;35:1474.
Yee-Lin V, Pooi-Fong W, Soo-Beng AK. Nutlin-3, A P53-Mdm2 antagonist for nasopharyngeal carcinoma treatment. Mini Rev Med Chem. 2017;17:173–83.
Das AT, Tenenbaum L, Berkhout B. Tet-on systems for doxycycline-inducible gene expression. Curr Gene Ther. 2016;16:156–167.
Karg MM, Lu YR, Refaian N, Cameron J, Hoffmann E, Hoppe C, et al. Sustained vision recovery by OSK gene therapy in a mouse model of glaucoma. Cell Reprogramm. 2023;25:288–99.
Artus J, Zenych A, Simanic I, Desterke C, Clay D, Saïm S, et al. BCR-ABL promotes hematopoietic stem and progenitor cell formation in embryonic stem cells. Exp Hematol. 2023;124:22–35.e23.
Kopp F, Mendell JT. Functional classification and experimental dissection of long noncoding RNAs. Cell. 2018;172:393–407.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43:904–14.
Zheng ZQ, Li ZX, Zhou GQ, Lin L, Zhang LL, Lv JW, et al. Long Noncoding RNA FAM225A promotes nasopharyngeal carcinoma tumorigenesis and metastasis by acting as ceRNA to sponge miR-590-3p/miR-1275 and upregulate ITGB3. Cancer Res. 2019;79:4612–26.
Zanganeh S, Hutter G, Spitler R, Lenkov O, Mahmoudi M, Shaw A, et al. Iron oxide nanoparticles inhibit tumour growth by inducing pro-inflammatory macrophage polarization in tumour tissues. Nat Nanotechnol. 2016;11:986–94.
Tarangelo A, Dixon SJ. Nanomedicine: an iron age for cancer therapy. Nat Nanotechnol. 2016;11:921–2.
Llovet JM, Ricci S, Mazzaferro V, Hilgard P, Gane E, Blanc JF, et al. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378–90.
Abou-Alfa GK, Schwartz L, Ricci S, Amadori D, Santoro A, Figer A, et al. Phase II study of sorafenib in patients with advanced hepatocellular carcinoma. J Clin Oncol. 2006;24:4293–4300.
Pan JJ, Zhang SW, Chen CB, Xiao SW, Sun Y, Liu CQ, et al. Effect of recombinant adenovirus-p53 combined with radiotherapy on long-term prognosis of advanced nasopharyngeal carcinoma. J Clin Oncol. 2009;27:799–804.
Nie X, Guo E, Wu C, Liu D, Sun W, Zhang L, et al. SALL4 induces radioresistance in nasopharyngeal carcinoma via the ATM/Chk2/p53 pathway. Cancer Med. 2019;8:1779–92.
Yuan C, Xu XH, Chen Z. Recombinant human adenovirus-p53 therapy for the treatment of nasopharyngeal carcinoma: a meta-analysis. Springerplus. 2016;5:1885.
Ma S, Sun L, Wu W, Wu J, Sun Z, Ren J. USP22 protects against myocardial ischemia-reperfusion injury via the SIRT1-p53/SLC7A11-dependent inhibition of ferroptosis-induced cardiomyocyte death. Front Physiol. 2020;11:551318.
Lu J, Li Y, Wu Y, Zhou S, Duan C, Dong Z, et al. MICAL2 mediates p53 ubiquitin degradation through oxidating p53 methionine 40 and 160 and promotes colorectal cancer malignance. Theranostics. 2018;8:5289–306.
Author information
Authors and Affiliations
Contributions
Conceptualization, QCZ, BY; Methodology, BY, SYZ, YS and WYC; Investigation, QCZ, BY, SYZ, WYC, JY, ZG, YS, ZYY, WHC and SP.; Writing, BY, SYZ, and LLB; Funding acquisition, BY, QCZ, YS; Resources, SYZ and WYC; Supervision, BY; Formal analysis, BY and SYZ; Tracking of clinical data, HOY; Final data verification, YWY; Final data verification and aesthetic formatting, YK.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, S., Chen, W., Yang, J. et al. LINC-AC092535.5 regulates MICAL2 mRNA level to inhibit p53-mediated ferroptosis in nasopharyngeal carcinoma. Oncogene 45, 1260–1274 (2026). https://doi.org/10.1038/s41388-026-03714-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41388-026-03714-y