Abstract
Intestinal adenomas are premalignant lesions that develop into colorectal cancer (CRC), yet the metabolic pathways underlying their malignant transformation remain poorly characterized. Using targeted metabolomics via ultra performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS), we found that serum levels of the bioactive lipid metabolite 14,15-epoxyeicosatrienoic acid (14,15-EET) were dramatically elevated in ApcMin/+ adenoma model mice as early as pre-adenoma stages, compared to C57BL/6 controls. The results were also consistent in adenomas and CRC patients. ELISA data and bioinformatics analyses revealed both elevated serum 14,15-EET levels and upregulated cytochrome P450 2J2 (CYP2J2) expression in tumor. Functional studies showed that 14,15-EET accelerates adenoma growth in vivo, and promotes proliferation, migration, and invasion in vitro by activating AKT (Ser473)/ERK1/2 signaling and inducing epithelial-mesenchymal transition (EMT). Its early elevation in premalignant lesions, and relative molecules 14,15-EET/CYP2J2 represents a novel strategy for disrupting adenoma-carcinoma transition, and offering new biomarker for CRC prevention.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most prevalent gastrointestinal malignancy and second leading cause of cancer mortality globally [1]. It develops through multistep stages: intestinal epithelial hyperplasia, adenoma formation, and ultimately evolving into CRC [2, 3], driven by oncogene activation and tumor suppressor inactivation/mutation. The adenomatous polyposis coli (APC) gene is a tumor suppressor and inactivated in approximately 80% of CRC cases at early stage [4]. Numerous studies highlight that intestinal microenvironment alterations play a key role in CRC development, including crosstalk between immunocytes, cytokines and cancer cells, as well as intestinal microbiota dysbiosis and dyslipidemia [5,6,7,8,9]. Current research focuses on how lipid metabolites influence tumorigenesis and their underlying mechanisms.
Eicosanoids are intracellular signaling molecules that play important roles in inflammation, allergic reactions, cardiovascular function, and tumorigenesis. Eicosanoids are enzymatic metabolites of arachidonic acid, synthesized by cyclooxygenase (COX), lipoxygenase (LOX) and cytochrome P450 (CYP) enzymes [10]. These enzymes are frequently overexpressed in various cancers, where they promote cancer cell proliferation and inhibit apoptosis. Previous studies have shown that increased COX-2 expression can induce APC gene instability, leading to imbalanced intestinal epithelial cell differentiation, abnormal proliferation, and an increased risk of CRC [10, 11]. Similarly, 5-LOX deficiency inhibits intestinal adenoma cell proliferation and promotes apoptosis [12], which suggests eicosanoids and their synthetic enzymes may contribute to tumorigenesis and progression.
Epoxyeicosatrienoic acids (EETs), as eicosanoid metabolites, are synthesized from arachidonic acid by CYP2Cs and CYP2Js. There are four regioisomers of EETs: 5,6-EET, 8,9-EET, 11,12-EET and 14,15-EET [13]. Previous research has shown that EETs are involved in a variety of physiological processes, including anti-inflammation, angiogenesis, proliferation, migration, neuroprotection, cardiovascular protection and glucose regulation [14,15,16,17,18]. In addition to these physiological roles, recent studies have also implicated EETs in tumorigenesis, metastasis, and drug resistance in several types of cancer [19,20,21]. Another study reported upregulation of the CYP monooxygenase pathway in colon cancer, and its inhibition via CYP inhibitors suppresses colon tumorigenesis [22]. This indicates that CYP monooxygenase and its products may contribute to the pathogenesis of colon cancer. However, relatively few studies have explored the role of CYP monooxygenase and its products, including EETs, in CRC. The impact of 14,15-EET on CRC initiation and development, and its potential mechanism, remains poorly understood.
Our study revealed elevated serum 14,15-EET levels in pre-adenoma stage ApcMin/+ mice and CRC patients. We demonstrated that 14,15-EET potently drives adenoma-carcinoma progression by activating AKT(Ser473)/ERK signaling and inducing EMT. Our findings establish it as a critical mediator of CRC, highlighting its potential as both a diagnostic biomarker for early-stage risk stratification and a therapeutic target.
Results
Serum 14,15-EET levels are elevated in Apc Min/+ mice prior to intestinal adenoma development
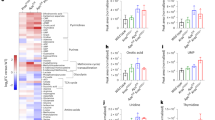
Previous studies have linked tumorigenesis to lipid metabolic reprogramming and alterations in lipid metabolites [23, 24], but whether serum lipid metabolites change prior to tumor initiation remains unclear. Here, we analyzed serum lipid profiles via UPLC-MS/MS-based targeted metabolomics in C57 mice and ApcMin/+ mice at two time points: 7-week-old (pre-adenoma stage, no intestinal glandular abnormalities or adenoma) and 24-week-old (advanced adenoma stage) (Fig. 1A). Compared with 7-week-old C57 mice, age-matched ApcMin/+ mice had 14 significantly elevated lipid subtypes (e.g., triglycerides, eicosanoids) and 9 significantly reduced (e.g., phosphatidylcholines) (Fig. 1B). Notably, eicosanoids dominated elevated lipids: serum 14,15-EET levels were 6.63-fold higher in ApcMin/+ mice (Fig. 1C). These data confirm serum lipid metabolites change in ApcMin/+ mice prior to adenoma development, with 14,15-EET showing the most prominent elevation. Elevated 14,15-EET was also observed in 24-week-old ApcMin/+ mice versus C57 controls. Additionally, triglycerides, diglycerides, eicosanoids, and acylcarnitines were significantly elevated, while ceramides, sphingomyelins, and phosphatidylcholines were reduced (Fig. 1D).
A The schematic diagram for targeted lipidomics assay. B The volcano plot and heatmap illustrating differential lipid metabolites in serum between ApcMin/+ and C57 mice (7 week). C FC of various subtypes of eicosanoid levels in serum between ApcMin/+ and C57 mice (7 week). D The volcano plot and heatmap illustrating differential lipid metabolites in serum between ApcMin/+ and C57 mice (24 week). The results are presented as mean ± SEM (n = 5 per group, P < 0.05, |FC | ≥ 1.2).
14, 15-EET is mainly produced by intestines
EETs are synthesized by CYP epoxygenases and are first identified in endothelial cells [25]. Other EET-producing cell and tissues include the small intestine, colon, liver, kidneys, and monocytic leukocytes [26,27,28]. To identify the source of elevated serum 14,15-EET, we measured mRNA expression of 10 Cyp2c and 5 Cyp2j genes (involved in 14,15-EET synthesis) in the intestine, kidney, liver, spleen, and blood cells of 7-week-old C57 and ApcMin/+ mice (Fig. 2A). Cyp2c and Cyp2j expression was detected in the intestine, kidney, and liver (Fig. 2B-D), but not in the spleen or blood cells, which indicates 14,15-EET is not derived from splenic immune cells or hemocytes. Notably, most Cyp2c and Cyp2j genes were significantly upregulated in the intestinal tissue of ApcMin/+ mice (Fig. 2B). Taken together, these results confirm that elevated serum 14,15-EET in ApcMin/+ mice primarily originates from the intestine.
A The schematic diagram for detecting Cyp2cs and Cyp2js expression in different tissues, including the (B) intestine, (C) kidney and (D) liver. (n = 5 per group, *P < 0.05, ** P < 0.01, *** P < 0.001).
14, 15-EET promotes the progression of intestinal adenomas in Apc Min/+ mice
Previous studies confirmed that 14,15-EET promotes pheochromocytoma growth and EMT in breast cancer cells [21]. To determine whether 14,15-EET has a similar role in intestinal adenomas, we treated ApcMin/+ mice with PBS or 14,15-EET (Fig. 3A). Compared with the PBS control group, 14,15-EET treatment significantly increased adenoma numbers in the jejunum and ileum of ApcMin/+ mice, but not in the duodenum or colon (Fig. 3B, C). These findings indicate that 14,15-EET may promote intestinal adenoma development in ApcMin/+ mice.
A The schematic diagram for 14,15-EET (30 μg/kg, thrice weekly, intraperitoneal injection) treated ApcMin/+ mice and observe the intestinal adenoma growth. B Methylene blue staining of intestinal adenoma. C Number of tumors in duodenum, jejunum, ileum and colon. D H&E staining and E Pathological score of duodenum, jejunum, ileum and colon in ApcMin/+ mice. F Ki67 immunohistochemical staining, G Ki67 positive cells and H Relative AOD of Ki67 staining of jejunum and ileum in ApcMin/+ mice. The results are presented as mean ± SEM (n = 4 per group; ns P å 0.05, * P < 0.05, ** P < 0.01, *** P < 0.001; 10× scale bar: 200 μm, 40× scale bar: 50 μm).
To further explore the effect of 14,15-EET on intestinal adenomas, we performed intestinal histological analysis in PBS- or 14,15-EET-treated ApcMin/+ mice. No significant differences in gland area, circularity, or their variation coefficients were observed in the duodenum between groups. However, the 14,15-EET group showed increased gland area, decreased circularity, and elevated circularity variation in the jejunum and ileum—changes indicating significant glandular disorder and structural disruption (Figs. 3D, S1A, B). Furthermore, CD45 staining revealed significantly increased immune cell infiltration in the jejunum and ileum of 14,15-EET-treated mice (Fig. S1C-E). Intestinal adenoma pathological scores were significantly higher in the 14,15-EET group than the PBS control, especially in the jejunum, ileum, and colon (Fig. 3E). Consistent with pathological scores, Ki67 staining showed increased expression in the jejunum and ileum of the 14,15-EET group versus the PBS control (Fig. 3F-H). Collectively, these findings suggest that 14,15-EET promotes intestinal adenoma progression in ApcMin/+ mice by driving glandular hyperplasia, disrupting epithelial structure, and promoting immune infiltration.
14, 15-EET contributes to proliferation of colorectal cancer in vitro
Previous reports indicated that 95% of familial adenomatous polyposis (FAP) patients with APC mutations may progress to CRC [29], prompting us to investigate 14,15-EET’s effect on intestinal adenoma and CRC cells in vitro. Due to unavailability of intestinal adenoma cells despite efforts, we used CRC cell lines (CT26 and MC38) instead. To investigate the role of 14,15-EET in CRC tumorigenesis, we evaluated CRC cell viability and colony-forming ability. 14,15-EET enhanced CRC cell viability (Fig. 4A, B) and colony-forming capacity (Fig. 4C–E) in a concentration-dependent manner, confirming its promotion of CRC cell proliferation in vitro. Similar results were observed in human CRC cells (Fig. S2A, B).
A, B CCK-8 assay was used to detect the proliferation of CT26 and MC38 cells induced by 14,15-EET. C–E Colony formation assay was used to detect the colony forming ability of CT26 and MC38 cells induced by 14,15-EET. CT26 and MC38 cells were then transfected with shNC or shCYPs (shCYP2C55, shCYP2C70, shCYP2J5 and shCYP2J13) for 48 h. F, G The proliferation ability in CRC cells was evaluated by CCK-8 assay for 6 days after shRNA transfection. H–J The colony forming ability in in CRC cells was measured by colony formation assay after 48 h shRNA transfection. The results are presented as mean ± SEM (n = 3 per group, * P < 0.05, ** P < 0.01 and *** P < 0.001). The statistical significance of two groups was determined using t-test.
CYPs (mainly CYP2C55, CYP2C70, CYP2J5, CYP2J13 in mice, and CYP2J2 in humans) are potential enzymes for 14,15-EET synthesis. We next knocked down CYPs (RT-qPCR-verified efficiency; Figs. S2C, S3A-B) and evaluated their effects on CRC cell proliferation. CCK-8 assays showed CYP knockdown significantly inhibited proliferation (P < 0.001), particularly in shCYP2J13-CT26 and shCYP2J13-MC38 (P < 0.0001; Fig. 4F, G). Consistently, CYP knockdown markedly reduced colony formation (Fig. 4H-J), suggesting CYP deficiency partially suppresses CRC proliferation.
To confirm if CYP knockdown inhibits proliferation by reducing 14,15-EET synthesis, we performed rescue experiments with shCYP2J13. Compared to PBS, exogenous 14,15-EET markedly enhanced colony formation in shNC-transfected MC38 cells (Fig. S3C, D). Conversely, CYP2J13 knockdown reduced colony numbers. Notably, this reduction was partially reversed by 14,15-EET supplementation in shCYP2J13-transfected cells. Similar effects were observed in HT-29 cells (Fig. S2D, E). These findings indicate that CYP knockdown inhibits CRC cell proliferation partly by reducing 14,15-EET biosynthesis, consistent with the pro-proliferative effect of 14,15-EET in CRC.
14, 15-EET promotes EMT and activates PI3K/AKT and MAPK/ERK signaling in vitro
To investigate 14,15-EET’s role in CRC progression, we examined its effects on migration and invasion. Wound healing assays showed 14,15-EET concentration-dependently promoted wound closure in CRC cells (Figs. 5A, B; S2F, G). Transwell assays confirmed that 14,15-EET significantly enhanced both migration and invasion (Fig. 5C, D), indicating it promotes these aggressive phenotypes in CRC cells.
A, B Wound healing assay was used to measure the migration of CT26 and MC38 cells induced by 14,15-EET for 12 h (n = 3/group). C, D Transwell assay was used to detect the migration and invasion of CT26 and MC38 cells induced by 100 nM 14,15-EET for 18 h (n = 3/group). E–G Western blot evaluation of E-cadherin, N-cadherin and Vimentin in CT26 and MC38 cells induced by 100 nM 14,15-EET for 24 and 48 h (n = 4/group). H–J Western blot evaluation of p-Akt, Akt, p-ERK1/2 and ERK1/2 in CT26 and MC38 cells treated with 100 nM 14,15-EET for 24 h (n = 4/group). The results are presented as mean ± SEM (*P < 0.05, **P < 0.01, ***P < 0.001). The statistical significance of two groups was determined using t-test.
To explore the mechanism of 14,15-EET-induced migration and invasion, we analyzed EMT-related protein levels in CRC cells treated with PBS or 14,15-EET. 14,15-EET downregulated E-cadherin and upregulated N-cadherin and Vimentin (Figs. 5E-G, S2H, I), suggesting it induces EMT in CRC cells. Given prior reports that EETs induce EMT via PI3K/AKT and MAPK/ERK pathways [30], we verified this in CRC cells. Compared with the PBS group, 14,15-EET did not alter total AKT/ERK levels but markedly increased phosphorylation of AKT (Ser473) and ERK (Thr202/Tyr204) (Figs. 5H-J, S2J, K), suggesting that activation of these pathways contributes to 14,15-EET-induced EMT.
As a lipid signaling molecule, 14,15-EET may exert additional CRC-promoting effects. RNA-seq comparing PBS- and 14,15-EET-treated CT26 cells (Fig. S4A) identified 794 upregulated and 596 downregulated differentially expressed genes (DEGs) (∣FC∣ > 2, P < 0.05; Fig. S4B, C). KEGG and GO enrichment analyses revealed that upregulated DEGs were enriched in ribosome biogenesis and function (Fig. S4D, E), whereas downregulated DEGs were associated with DNA damage repair (nucleotide excision repair, mismatch repair, base excision repair) and radiation response (Fig. S4F, G). qRT-PCR confirmed upregulated expression of ribosomal protein (RP)-related genes in 14,15-EET-treated CRC cells and ApcMin/+ mice (Fig. S4H-J), indicating 14,15-EET enhances ribosome biogenesis and suppresses DNA repair.
14, 15-EET and its synthetic enzyme CYP2J2 increased in colorectal cancer patients
Our study confirmed elevated serum 14,15-EET in ApcMin/+ mice. To extend these findings to human CRC, we measured 14,15-EET by ELISA in CRC cell lines (intracellular and supernatant levels) and serum from patients with intestinal adenomas, non-metastatic CRC (nmCRC), metastatic CRC (mCRC), and healthy controls (HC).
In vitro, CRC cells (DLD-1, LoVo, HCT116, HCT-15, HT-29) exhibited significantly higher intracellular 14,15-EET than normal intestinal epithelial NCM460 cells (Fig. 6A). Consistently, 14,15-EET secretion in CRC cell supernatants was markedly increased (Fig. 6B), recapitulating the elevation in the mouse model. In clinical samples, serum 14,15-EET differed significantly among the four groups (Fig. 6C): intestinal adenoma patients had the highest level (108.33 pg/mL), significantly higher than those in nmCRC (55.16 pg/mL), mCRC (37.85 pg/mL), and HC (3.49 pg/mL). Collectively, these results demonstrate that 14,15-EET elevation is an early event in colorectal carcinogenesis, beginning at the precancerous adenoma stage and remaining elevated throughout CRC progression.
A, B Intracellular and supernatant 14,15-EET level in CRC cell lines (DLD-1, LoVo, HCT116, HCT-15, and HT-29). C Serum 14,15-EET level in intestinal adenoma (n = 12), non-metastatic CRC (nmCRC, n = 10), metastatic CRC (mCRC, n = 22) patients and healthy donors (n = 24). The results are presented as mean ± SEM (** P < 0.01, *** P < 0.001). Human Cyp2j2 and Ephx2 gene expression in COAD and READ were analyzed (D, E) using GEPIA2 (match TCGA normal and GTEx data); F, G using TNMplot (from GEO, GTex, TCGA, and TARGET databases). Log2 (TPM + 1) was used for the log scale in GEPIA2.
CYP2J2 is the primary synthetic enzyme for 14,15-EET in humans, and sEH (EPHX2) hydrolyzes EETs to DHETs [31]. GEPIA2 analysis revealed significantly upregulated Cyp2j2 (Fig. 6D) and a downward trend in Ephx2 (Fig. 6E) in colon (COAD) and rectal (READ) adenocarcinomas versus normal tissues. TNMplot database also confirmed these findings (Fig. 6F, G).
We further analyzed associations between Cyp2j2/Ephx2 expression, pathological stage, and overall survival (OS) in CRC patients. Cyp2j2 levels were not stage-dependent in COAD/READ (Fig. S5A), but low Cyp2j2 predicted poorer OS (GEPIA2, TCGA RNA-seq; Fig. S5B). Notably, lower Ephx2 correlated with advanced stage and worse OS in COAD/READ (Fig. S5C, D). These data suggest that high Cyp2j2 and low Ephx2 may contribute to CRC progression.
Discussion
Intestinal adenomas are premalignant lesions that develop into CRC, occurring in approximately 95% of FAP patients with APC mutations [29]. Despite metabolic reprogramming occurs at the adenoma stage being critical for tumor initiation [32], the specific lipid metabolites driving adenoma progression—especially at the pre-adenoma stage—remain poorly defined. Here, we first found that serum 14,15-EET was significantly elevated in ApcMin/+ mice at pre-adenoma stages. It functionally promotes adenoma growth, proliferation, and migration by inducing EMT and activating AKT and ERK1/2 signaling. Importantly, our findings are corroborated in human clinical settings. We found that this elevation of 14,15-EET is not merely a murine phenomenon but is profoundly relevant to human carcinogenesis. Strikingly, in our clinical cohort, intestinal adenoma patients showed the highest serum levels of 14,15-EET, significantly exceeding levels in both healthy controls and patients with established cancer. We identify 14,15-EET as an early driver in the human adenoma-to-carcinoma sequence.
Previous studies reported elevated serum eicosanoids (EETs, HETEs) in individuals with enteritis or CRC versus healthy individuals [33], but the metabolic signature of intestinal adenomas—particularly at the pre-adenoma stage—has not been fully elucidated. Our study addresses this gap by demonstrating that 14,15-EET is elevated as early as the pre-adenoma stage in ApcMin/+ mice, persisting throughout adenoma development. This is consistent with findings in MMTV-PyMT breast cancer mice, where HETEs increased during established tumor growth [34].
Previous studies implicated EETs in promoting tumorigenesis—including proliferation, metastasis, angiogenesis, and drug resistance—in multiple cancers [19, 35]. However, their role in the adenoma-carcinoma transition was unclear. Our study demonstrated that 14,15-EET increases adenoma burden and accelerates pathological progression in ApcMin/+ mice, evidenced by segment-specific glandular abnormalities (e.g., increased area, decreased circularity) and enhanced inflammatory infiltration (elevated CD45-positive cells). In vitro experiments demonstrated that 14,15-EET promotes proliferation, migration, and invasion in CRC cell lines. Our study extends these observations by investigating the cellular functions of CYP2C and CYP2J, the primary synthases for 14,15-EET. Silencing either synthase in CRC cells abrogated 14,15-EET’s effects on proliferation and metastasis. Notably, a rescue experiment further validated the mechanistic link between CYP and 14,15-EET: exogenous 14,15-EET partially reversed the clonogenic defect caused by CYP2J13 knockdown in MC38 cells and CYP2J2 knockdown in HT-29 cells. This reversal supports that CYP regulates CRC cell proliferation, at least in part, via 14,15-EET synthesis. Prior research reported that deleting the key EET synthase CYP2C suppresses CRC development in the azoxymethane/dextran sodium sulfate (AOM/DSS) model—which mimics colon-specific tumorigenesis [22]. Furthermore, AOM/DSS-induced CRC models exhibit upregulated CYP2J (key EET-synthesizing synthases) expression and elevated 14,15-EET levels, with CYP2J2 contributing to tumor progression via EMT [36, 37]. Analysis of clinical databases confirms that CYP2J2, the key EET-synthesizing enzyme, is upregulated in CRC tissues. These findings confirm that the CYP2C/CYP2J-14,15-EET axis is not unique to the ApcMin/+ model but represents a mechanism in various CRC models, despite differences in tumor location.
A recent study demonstrated that upregulated CYP monooxygenases and their product 12,13-EpOME promote CRC [38]. While CYP expression is species-specific (human colorectal tissues express CYP2J2), the core CYP-mediated bioactive epoxy lipid mechanism is conserved. Our study identifies 14,15-EET as a distinct CYP-derived mediator: 12,13-EpOME promotes late-stage invasion, whereas 14,15-EET is elevated in pre-adenoma ApcMin/+ mice to drive early tumorigenesis.
The PI3K/AKT and ERK/MAPK pathways, key regulators of cell proliferation, differentiation, and survival, are commonly dysregulated in cancer. Our study shows that 14,15-EET induces EMT by upregulating mesenchymal markers (N-cadherin, Vimentin) and downregulating epithelial marker E-cadherin. We further found 14,15-EET increases AKT (Ser473) and ERK1/2 (Thr202/Tyr204) phosphorylation in CRC cells, consistent with reports that EETs/CYP2J2 activate these pathways [39]. Our findings extend previous knowledge by establishing 14,15-EET’s role in promoting both proliferative signaling and metastatic potential via EMT.
Our transcriptomic analysis identified novel functions of 14,15-EET: activating ribosome biogenesis yet suppressing DNA damage repair. As core ribosomal components, ribosomal proteins (RPs) not only mediate ribosome biogenesis and protein translation, but also participate in cell migration and tumor transformation [40]. Previous studies showed distinct regulation patterns of RPs in CRC and intestinal adenomas versus normal mucosa, with RPs dysregulation affecting key CRC processes: proliferation, stemness, differentiation, autophagy, progression, and metastasis [41,42,43]. Moreover, 14,15-EET suppresses DNA damage repair pathways (e.g., nucleotide excision repair, mismatch repair, base excision repair) and the cellular response to radiation. Such suppression may induce genomic instability and promote radiotherapy resistance—consistent with emerging evidence that genomic instability drives colorectal carcinogenesis [44, 45]. These changes suggest 14,15-EET enhances proliferation while enabling mutation accumulation. However, we have not yet determined if genetic CYP2J2 inhibition or pharmacological 14,15-EET blockade reverses ribosome biogenesis upregulation and restores DNA repair function. Additional uncertainty is whether these effects on ribosome biogenesis and DNA repair are direct (mediated by 14,15-EET itself) or indirect (driven by AKT/ERK activation). Clinically, the correlation between CYP2J2 expression and ribosome biogenesis-related signatures or DNA damage repair deficiency in CRC remains undetermined.
Although tumors in ApcMin/+ mice arise primarily in the small intestine, this model is well-validated for APC-driven tumorigenesis and recapitulates key molecular events like Wnt/β-catenin dysregulation [46]. Thus, the observed metabolic reprogramming likely reflects early events in human CRC. Notably, our study focuses on the pre-adenoma stage—when metabolic changes are driven by initiating genetic events (e.g., APC mutation) in epithelial cells, unconfounded by complex stromal interactions in advanced disease. Whether APC mutation increases 14,15-EET levels remains unclear. Future studies will explore if 14,15-EET significantly promotes intestinal adenomas and CRC using ApcMin/+ mice lacking CYP2C/J. To better recapitulate human colonic tumorigenesis, 14,15-EET’s role will be further validated in colon-specific CRC models (e.g., AOM/DSS-induced, colon-specific Apc conditional knockout). Additionally, patient-derived organoids (PDOs) will explore its clinical relevance and translational value in CRC, and the therapeutic potential of the 14,15-EET antagonist 14,15-EEZE in vitro and in vivo models will also be evaluated.
In summary, 14,15-EET as a key metabolite markedly elevated at the pre-adenoma stage. 14,15-EET drives CRC progression by activating not only AKT/ERK signaling but also ribosome biogenesis, while suppressing DNA damage repair. Our study highlights 14,15-EET’s potential as an early diagnostic biomarker and therapeutic target for intestinal adenomas and subsequent CRC progression.
Materials and methods
Mice
Although the ApcMin/+ model primarily develops small intestinal (rather than colonic) tumors, unlike human CRC, it is well-established for studying APC-driven intestinal tumorigenesis and Wnt/β-catenin signaling. Therefore, we utilized it for in vivo experiments.
Five-week-old male/female C57BL/6 J (RRID:IMSR_JAX:000664) mice were from Guangdong Medical Animal Experiment Center. Male ApcMin/+ mice mice (C57BL/6 J background) were from Cyagen Biosciences, mated with 6–8-week-old female wild-type C57 mice. Apc mutation was confirmed by PCR (primers: F: 5’-TTCTGAGAAAGACAGAAGTTA-3’, R: 5’-TTCCACTTTGGCATAAGGC-3’). All animal procedures were approved by Guangdong Pharmaceutical University’s UCUCA, with housing adhering to ethical regulations, and all methods were performed in accordance with the relevant guidelines and regulations.
UPLC-MS/MS analysis
Serum lipid profiles of C57 and ApcMin/+ mice (7 and 24 weeks, n = 5/group) were analyzed by ultra performance liquid chromatography-tandem mass spectrometry (UPLC-MS/MS). Differential lipids were identified with P < 0.05 and |fold change (FC)| ≥1.2. Volcano plots, heatmaps, and FC analyses were generated using OmicStudio tools (https://www.omicstudio.cn/tool).
EET-treatment
Female ApcMin/+ mice were randomly divided into two groups (n = 4/group). From 6 to 18 weeks of age, mice received intraperitoneal (i.p.) injections of 14,15-EET (30 μg/kg) or equal-volume normal saline, three times weekly, and were sacrificed at indicated time points.
Analysis of intestinal tumors
After sacrifice, the entire intestinal tract of ApcMin/+ mice was excised and divided into duodenum, jejunum, ileum, and colon. Each segment was longitudinally opened, tiled on filter paper, fixed in 4% paraformaldehyde (PFA), and stained with 1% methylene blue. To better compare the tumor growth in two groups, all intestinal segments from each mouse were pooled, and macroscopic tumors (blue lesions) were counted per segment.
Histological analysis
After tumor counting, intestinal tissues were fixed in 4% PFA, paraffin-embedded, and sectioned at 3 μm. For histopathology, three non-serial sections were H&E-stained and examined by light microscopy.
Morphometry of intestinal glands was performed on H&E sections using ImageJ software (RRID:SCR_003070). Five random 40× fields per section were analyzed for the mean gland area (μm2), coefficient of variation (CV) of gland area (calculated as standard deviation/mean × 100%), circularity (4π×area/perimeter2, values closer to 1.0 indicating more circular), and the CV of circularity.
Pathological grading of intestinal lesions was based on: inflammatory cell infiltration (scored 0-2), glandular hyperplasia (0-2), glandular structural disorder (0-2), and adenoma invasion (2: <50% mucosal involvement, 3: nearly entire mucosal without basement membrane penetration, 4: basement membrane penetration). The total score represented the sum of individual scores. All assessments were performed blindly by two researchers, with mean values used for statistical analysis.
Immunohistochemical assay
Tissue sections underwent high-pressure antigen retrieval with citric acid buffer (pH 6.0) for 10 min and were incubated with 3% hydrogen peroxide to block endogenous peroxidase activity. After blocking with 1% bovine serum albumin (BSA), sections were incubated with primary antibodies against Ki67 and CD45, and goat anti-rabbit secondary antibody. Immunostaining was performed using DAB (Cell Signaling Technology) with hematoxylin counterstaining, and slides were examined under a light microscope (Olympus DP27, Japan). Antibody details are provided in Supplementary Table 1.
Cell line and culture
Human intestinal adenoma cells were initially planned for in vitro experiments (consistent with the study focus on intestinal adenoma progression); however, these cells were unavailable from commercial cell banks or collaborators. Thus, CRC cell lines served as alternatives for subsequent in vitro assays.
CT26 (Cat# CRL-2638, RRID:CVCL_7256) cells were purchased from the American Type Culture Collection (ATCC). MC38, NCM460, DLD-1, LoVo, HCT116, HCT-15 and HT-29 cells were obtained from Sun Yat-sen University Cancer Center and The Sixth Affiliated Hospital of Sun Yat-sen University. Cells were cultured in RPMI 1640 or DMEM (Gibco), respectively, supplemented with 10% fetal bovine serum (FBS, ExCell Bio) and 1% penicillin-streptomycin (PS). All cells tested negative for mycoplasma contamination.
RNA interference
Short hairpin RNA (shRNA) sequences targeting CYP2C55, CYP2C70, CYP2J5 and CYP2J13 were obtained from the Genetic Perturbation Platform (GPP, broadinstitute.org). Small interfering RNA (siRNA) oligonucleotides targeted specifically for CYP2J2 downregulated EETs levels in human CRC cells. siRNA was designed and provided by Shanghai GenePharma Co., Ltd. Oligo sequences are listed in Supplementary Tables 2 and 3. Transfections were performed using Lipofectamine 3000 (Invitrogen) per manufacturer’s instructions, and efficiency was assessed by RT–qPCR 48 h post-transfection.
CCK-8 assay
CT26 and MC38 cells were cultured in at 1 × 104 cells/well. Following 12 h culture in 10% FBS medium, cells were treated with different concentrations of 14,15-EET (0, 12.5, 25, 50, 100, 200 nM) for 24 h. Subsequently, cells were cultured in 100 μL complete medium containing 10 μL CCK-8 reagent/well for 3 h at 37 °C. The absorbance values of each well were measured with a microplate spectrophotometer (Molecular Devices, USA) at 450 nm. Furthermore, cells infected with shNC or shCYPs for 48, absorbance at 450 nm was detected for 6 days. All proliferation assays were repeated as independent experiments three times.
Colony formation assay
CRC cells were cultured in 6-well plates at a density of 500 cells/well. Post-adherence, cells were refed with 5% FBS medium containing different concentrations (as previously described) of 14,15-EET for 10 days. For shNC- or shCYPs-transfected cells, cells were seeded at 1000 cells/well in 6-well plates and then incubated in 10% FBS medium for 10 days. For 14,15-EET rescue experiments in CYP-knockdown cells: cells were first transfected with shCYP2J13 or siCYP2J2. At 48 h post-transfection, cells were seeded in 6-well plates at 1000 cells/well. Cells were cultured in 10% FBS medium supplemented with 100 nM 14,15-EET or PBS. Finally, the cells were fixed with 4% PFA and stained with 0.1% crystal violet. Colony numbers were counted when colonies contained >50 cells. All experiments were performed in triplicate.
Wound healing assay
CRC cells were seeded in 6-well plates at 5 × 104 cells/well. Upon reaching 80% confluency, a sterile 200 μL pipette tip scratched the monolayer. Detached cells were removed via PBS washes. The remaining cells were treated with different concentrations (as previously described) of 14,15-EET for 12 h. Five random scratched fields were imaged at 0 and 12 h. The area of wound closure was used to assess cell migration, and images were analyzed by ImageJ software. The wound healing rate was calculated as follows: [(area of scratch at 0 h) − (area of scratch at 12 h)]/ (area of scratch at 0 h) × 100.
Transwell assay
The migration and invasion assays of CT26 and MC38 cells were conducted using Transwell chambers (8 μm; Millipore) pre-coated without or with Matrigel (BD Biosciences). Cells (5 × 104 or 1 × 105) suspended in 200 μL serum-free medium (with/without 100 nM 14,15-EET) were seeded into the upper chamber. The lower chamber contained 600 μL 10% FBS medium. After 18 h, migrated/invaded cells on the lower membrane were fixed with 4% PFA, stained with 0.1% crystal violet, and counted via microscopy (8 random fields/well).
Western blot
Protein extracts were separated by 8-10% SDS-PAGE, transferred to PVDF membranes, then incubated with primary antibodies at 4 °C overnight. Primary antibodies included: AKT, Phospho-AKT (Ser473), p44/42 MAPK (Erk1/2), Phospho-p44/42 MAPK (P-Erk1/2) (Thr202/Tyr204), E-cadherin, N-cadherin, Vimentin, and GAPDH. Peroxidase-conjugated anti-rabbit/anti-mouse antibodies were used as secondary antibodies. Antibody details are provided in Supplementary Table 1. Protein bands were visualized using an enhanced chemiluminescence. Band intensity was quantified via ImageJ software.
RNA extraction and quantitative real-time PCR
Total RNA was extracted with TRIzol reagent (Invitrogen), and cDNA synthesized with the cDNA Reverse Transcription Kit (Accurate Biotechnology). qRT-PCR was performed using 2×SYBR Green PCR Master Mix (Accurate Biotechnology) to quantify relative mRNA expression. GAPDH as the endogenous control. Specific primer sequences are provided in Supplementary Tables 4. Data were analyzed on a LightCycler 480 instrument (Roche, RRID:SCR_018626), and relative gene expression was calculated via the 2−ΔΔCt method.
RNA-seq analysis
Total RNA was isolated with TRIzol reagent from CT26 cells treated with/without 100 nM 14,15-EET for 24 h. All samples underwent RNA quality control, and RNA-seq was performed by Shanghai Yuanxin Biomedical Technology Co., Ltd. Differentially expressed genes (DEGs) were filtered by |log2 fold change (FC)| > 1 and P < 0.05. Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) enrichment analyses were ranked based on enrichment factor. Volcano plots, heatmaps, and KEGG/GO enrichment results were generated via OmicStudio tools (https://www.omicstudio.cn/tool).
14,15-EET ELISA assay
Cell culture supernatants and pellets were collected from NCM460, DLD-1, LoVo, HCT116, HCT-15, and HT-29 cells. Peripheral blood was from 12 intestinal adenoma, 10 non-metastatic CRC (nmCRC), 22 metastatic CRC (mCRC) patients (30–75 years; exclusion: no other malignancies, hyperlipidemia, hypertension) and 24 healthy donors (30–60 years). 14,15-EET levels were quantified via ELISA kit (ab175812, Abcam). Serum samples were from The Second Affiliated Hospital of Guangzhou Medical University (volunteer details: Supplementary Table 5). The study was approved by the Health Research Ethics Board with informed consent from all participants, and all related methods were performed in accordance with the relevant guidelines and regulations.
Bioinformatics analysis
The gene expression, pathological stage and overall survival (OS) of Cyp2j2 and Ephx2 in adenocarcinoma (COAD) and rectum adenocarcinoma (READ) were analyzed using Gene Expression Profiling Interactive Analysis 2 (GEPIA 2, https://GEPIA2.cancerpku.cn/) or TNMplot (https://tnmplot.com/analysis/).
Statistical analysis
SPSS 19.0 (SPSS, RRID:SCR_002865) and GraphPad Prism 8.0.2 (RRID:SCR_002798) software were used for statistical analyses. After assessing data distribution, two-tailed unpaired t-test was applied for normally distributed data; non-parametric Mann-Whitney U test was used for heteroscedastic data. All data were uniformly presented as mean ± standard error of the mean (SEM). P-value < 0.05 was considered statistically significant. All experiments were repeated at least three times with independent samples.
Data availability
All experimental data are included in this article and its supplementary information files. All raw omics data are available from the corresponding author upon reasonable request.
References
Sung H, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61:759–67.
G. B. D. Colorectal Cancer Collaborators. The global, regional, and national burden of colorectal cancer and its attributable risk factors in 195 countries and territories, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Gastroenterol Hepatol. 2019;4:913–33.
Nikolouzakis TK, et al. Improving diagnosis, prognosis and prediction by using biomarkers in CRC patients (Review). Oncol Rep. 2018;39:2455–72.
Hanus M, et al. Immune system, microbiota, and microbial metabolites: the unresolved triad in colorectal cancer microenvironment. Front Immunol. 2021;12:612826.
Schmitt M, Greten FR. The inflammatory pathogenesis of colorectal cancer. Nat Rev Immunol. 2021;21:653–67.
Wang S, et al. Tumor-associated macrophages (TAMs) depend on Shp2 for their anti-tumor roles in colorectal cancer. Am J Cancer Res. 2019;9:1957–69.
Cheng Y, Ling Z, Li L. The intestinal microbiota and colorectal cancer. Front Immunol. 2020;11:615056.
Chen K, et al. The role of dyslipidemia in colitis-associated colorectal cancer. J Oncol. 2021;2021:6640384.
Wang Y, et al. Eicosanoid signaling in carcinogenesis of colorectal cancer. Cancer Metastasis Rev. 2018;37:257–67.
Chulada PC, et al. Genetic disruption of Ptgs-1, as well as Ptgs-2, reduces intestinal tumorigenesis in Min mice. Cancer Res. 2000;60:4705–8.
Cheon EC, et al. Mast cell 5-lipoxygenase activity promotes intestinal polyposis in APCDelta468 mice. Cancer Res. 2011;71:1627–36.
Fleming I. Vascular cytochrome p450 enzymes: physiology and pathophysiology. Trends Cardiovasc Med. 2008;18:20–5.
Luo XQ, et al. Epoxyeicosatrienoic acids inhibit the activation of NLRP3 inflammasome in murine macrophages. J Cell Physiol. 2020;235:9910–21.
Hu J, Fromel T, Fleming I. Angiogenesis and vascular stability in eicosanoids and cancer. Cancer Metastasis Rev. 2018;37:425–38.
T. Aliwarga, et al., Higher epoxyeicosatrienoic acids in cardiomyocytes-specific CYP2J2 transgenic mice are associated with improved myocardial remodeling. Biomedicines. 2020;8.
H. Zhao, et al., 14,15-EET Reduced Brain Injury from Cerebral Ischemia and Reperfusion via Suppressing Neuronal Parthanatos. Int J Mol Sci 2021;22.
Fu M, et al. Epoxyeicosatrienoic acids improve glucose homeostasis by preventing NF-kappaB-mediated transcription of SGLT2 in renal tubular epithelial cells. Mol Cell Endocrinol. 2021;523:111149.
Colombero C, et al. Cytochrome 450 metabolites of arachidonic acid (20-HETE, 11,12-EET and 14,15-EET) promote pheochromocytoma cell growth and tumor associated angiogenesis. Biochimie. 2020;171-172:147–57.
Allison SE, et al. Activation of ALDH1A1 in MDA-MB-468 breast cancer cells that over-express CYP2J2 protects against paclitaxel-dependent cell death mediated by reactive oxygen species. Biochem Pharm. 2017;143:79–89.
Luo J, et al. 14, 15-EET induces breast cancer cell EMT and cisplatin resistance by up-regulating integrin alphavbeta3 and activating FAK/PI3K/AKT signaling. J Exp Clin Cancer Res. 2018;37:23.
Wang W, et al. Targeted metabolomics identifies the cytochrome P450 monooxygenase eicosanoid pathway as a novel therapeutic target of colon tumorigenesis. Cancer Res. 2019;79:1822–30.
Cheng C, Geng F, Cheng X, Guo D. Lipid metabolism reprogramming and its potential targets in cancer. Cancer Commun. 2018;38:27.
B. Faubert, A. Solmonson, and R. J. DeBerardinis, Metabolic reprogramming and cancer progression. Science, 2020;368.
Revtyak GE, Hughes MJ, Johnson AR, Campbell WB. Histamine stimulation of prostaglandin and HETE synthesis in human endothelial cells. Am J Physiol. 1988;255:C214–25.
Zeldin DC, et al. CYP2J subfamily cytochrome P450s in the gastrointestinal tract: expression, localization, and potential functional significance. Mol Pharm. 1997;51:931–43.
Snider NT, Kornilov AM, Kent UM, Hollenberg PF. Anandamide metabolism by human liver and kidney microsomal cytochrome p450 enzymes to form hydroxyeicosatetraenoic and epoxyeicosatrienoic acid ethanolamides. J Pharm Exp Ther. 2007;321:590–7.
Nakayama K, Nitto T, Inoue T, Node K. Expression of the cytochrome P450 epoxygenase CYP2J2 in human monocytic leukocytes. Life Sci. 2008;83:339–45.
Powell SM, et al. APC mutations occur early during colorectal tumorigenesis. Nature. 1992;359:235–7.
Spector AA, Norris AW. Action of epoxyeicosatrienoic acids on cellular function. Am J Physiol Cell Physiol. 2007;292:C996–1012.
Spector AA, Fang X, Snyder GD, Weintraub NL. Epoxyeicosatrienoic acids (EETs): metabolism and biochemical function. Prog Lipid Res. 2004;43:55–90.
Satoh K, et al. Global metabolic reprogramming of colorectal cancer occurs at adenoma stage and is induced by MYC. Proc Natl Acad Sci USA. 2017;114:E7697–706.
Zhang J, et al. Distinct differences in serum eicosanoids in healthy, enteritis and colorectal cancer individuals. Metabolomics. 2017;14:4.
Chen A, et al. Investigation of the content differences of arachidonic acid metabolites in a mouse model of breast cancer by using LC-MS/MS. J Pharm Biomed Anal. 2021;194:113763.
Tao P, Jiang Y, Wang H, Gao G. CYP2J2-produced epoxyeicosatrienoic acids contribute to the ferroptosis resistance of pancreatic ductal adenocarcinoma in a PPARgamma-dependent manner. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2021;46:932–41.
Chen C, et al. Selective inhibitors of CYP2J2 related to terfenadine exhibit strong activity against human cancers in vitro and in vivo. J Pharm Exp Ther. 2009;329:908–18.
Luo Y, Liu JY. Pleiotropic functions of cytochrome P450 monooxygenase-derived eicosanoids in cancer. Front Pharm. 2020;11:580897.
Kong C, et al. Fusobacterium nucleatum promotes the development of colorectal cancer by activating a cytochrome P450/Epoxyoctadecenoic acid axis via TLR4/Keap1/NRF2 Signaling. Cancer Res. 2021;81:4485–98.
Jiang JG, Shen GF, Chen C, Fu XN, Wang DW. Effects of cytochrome P450 arachidonic acid epoxygenases on the proliferation of tumor cells. Ai Zheng. 2009;28:14–9.
Xu X, Xiong X, Sun Y. The role of ribosomal proteins in the regulation of cell proliferation, tumorigenesis, and genomic integrity. Sci China Life Sci. 2016;59:656–72.
Liu X, et al. RSL1D1 promotes the progression of colorectal cancer through RAN-mediated autophagy suppression. Cell Death Dis. 2022;13:43.
S. Y. Park, et al., RPL27 contributes to colorectal cancer proliferation and stemness via PLK1 signaling. Int J Oncol. 2023;63.
Zhu J, et al. RPL21 interacts with LAMP3 to promote colorectal cancer invasion and metastasis by regulating focal adhesion formation. Cell Mol Biol Lett. 2023;28:31.
Boland CR, Goel A. Microsatellite instability in colorectal cancer. Gastroenterology. 2010;138:2073–87 e2073.
Reilly NM, Novara L, Di Nicolantonio F, Bardelli A. Exploiting DNA repair defects in colorectal cancer. Mol Oncol. 2019;13:681–700.
Su LK, et al. Multiple intestinal neoplasia caused by a mutation in the murine homolog of the APC gene. Science. 1992;256:668–70.
Acknowledgements
This work was supported by Science and Technology Planning Project of Guangdong Province, China (2023A0505050153).
Author information
Authors and Affiliations
Contributions
SH: designed the research, performed experiments, analyzed data and wrote the manuscript; SH and RZ: performed animal experiments; SH and BZ: revised the manuscript; LJ and JZ: collected serum samples; JL: funding acquisition, project administration and scientific guidance. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
He, S., Zeng, R., Zheng, B. et al. 14,15-epoxyeicosatrienoic acid drives intestinal adenoma growth and its value as an early biomarker for intestinal adenoma occurrence. Oncogenesis 15, 13 (2026). https://doi.org/10.1038/s41389-026-00604-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41389-026-00604-6