Abstract
Effective precision oncology demands integration of pharmacokinetics/pharmacodynamics (PK/PD) profiling with tumor-specific genomic features. Here, we present a personalized treatment model using a patient-derived Networking Organoid Culture System (NOCS) composed of intestinal, liver, and kidney organoids differentiated from induced pluripotent stem cells (iPSCs) of an NF1-mutant breast cancer patient. This multi-organoid system enabled individualized assessment of drug absorption, distribution, metabolism, and excretion. Integrative genomic and pathway analyses uncovered therapeutic vulnerabilities, including responsiveness to a novel exon skipping therapy targeting NF1. PK/PD-guided screening on the NOCS prioritized Paxalisib, which, when combined with the exon skipping approach, demonstrated synergistic anticancer efficacy in patient-derived tumor models. These findings establish a clinically relevant framework that integrates multi-organ PK/PD modeling with genotype-driven therapeutic strategies, highlighting the potential of combining targeted gene correction with small-molecule therapy for personalized treatment. This platform offers broad applicability in precision oncology and drug development across diverse genetic contexts.
Similar content being viewed by others
Introduction
Neurofibromin (NF1) mutations define a clinically and biologically distinct subset of breast cancers (BCs) associated with aggressive behavior and limited response to targeted therapies.1 NF1 encodes a Ras GTPase-activating protein (Ras-GAP) that negatively regulates Ras signaling, and its inactivation leads to sustained activation of Ras and downstream PI3K/AKT and MAPK pathways.2,3,4 Loss-of-function mutations in NF1 promote tumor proliferation, survival, and therapeutic resistance, and have emerged as poor-prognosis genomic alterations across multiple solid tumors, including BC.5,6,7,8,9,10 Despite the availability of pathway inhibitors, patients with NF1-mutant tumors rarely achieve durable responses due to feedback reactivation and pathway redundancy. To overcome this therapeutic resistance, gene-corrective strategies such as antisense oligonucleotide (ASO)-mediated exon skipping offer a promising alternative. Exon skipping can restore the open reading frame in pre-mRNA by removing deleterious exons, allowing the production of a truncated but functional neurofibromin protein.11 Although this approach is under active investigation in neuromuscular disorders, its application to oncogenic Ras regulation remains largely unexplored.12,13,14
A major barrier to evaluating such genotype-corrective therapies lies in the lack of preclinical platforms that faithfully capture both patient-specific drug responses and systemic pharmacokinetics/pharmacodynamics (PK/PD). Organoids derived from patient-specific induced pluripotent stem cells (iPSCs) now enable the in vitro modeling of various tissues, including the intestine, liver, and kidney.15,16,17,18 However, individual organoids lack the capacity to simulate integrated absorption, distribution, metabolism, and excretion (ADME) processes observed in vivo19.
To address this, multi-organ microphysiological systems (MPSs)—including networking cell culture systems (NCCS) and organ-on-a-chip technologies—have been developed to support dynamic inter-organ crosstalk.20,21,22,23 When integrated with patient-specific iPSC-derived organoids, these platforms can model systemic drug behavior within a genetically relevant context, offering a new avenue for functional precision oncology.24,25,26
Here, we report the development of a patient-specific, iPSC-derived multi-organ PK/PD evaluation platform for NF1-mutant breast cancer. Using iPSCs derived from a patient tumor harboring an NF1 mutation, we generated intestinal, hepatic, and renal organoids and incorporated them into an NOCS-based MPS. We applied this system to evaluate antisense oligonucleotide therapy in combination with small-molecule inhibitors, demonstrating the identification of a patient-optimized treatment (Paxalisib) with synergistic efficacy when paired with NF1 exon skipping. This study establishes a mechanistically grounded, translational framework for integrating drug metabolism, genomic repair, and individualized therapy selection in genetically defined breast cancer.
Results
Establishment of BC patient-derived iPSCs harboring NF1 mutation
To establish a platform for precision oncology in BC, we developed an integrated workflow using patient-derived tissues. Genomic profiling of primary tumor cells enabled the identification of tumor-specific mutations, including a pathogenic NF1 variant. These tumor cells were used to model disease-specific phenotypes. In parallel, matched normal cells were reprogrammed into iPSCs, which were subsequently differentiated into intestine, liver, and kidney organoids. This multi-organ system was employed for in vitro PK/PD analyses, enabling personalized drug response profiling and supporting the development of individualized therapeutic strategies for NF1-mutant BC (Fig. 1a).
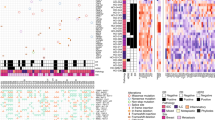
Generation of BC patient 1-derived iPSCs with NF1 mutation. a Schematic of precision medicine for BC patients. Created using BioRender.com. b BC subtypes (top) and WES analysis (bottom) of BC biospecimens from the biobank. c Dot plot of metastatic recurrence time based on NF1 mutation status. d Disease-specific survival time based on NF1 mutation status. e Volcano plot showing drug resistance related to NF1 mutation. f Mutation analysis of the NF1 allele in normal and patient-derived iPSCs. g mRNA expression of OCT4 and NANOG in BC patient-derived fibroblasts and iPSCs. Data are mean ± SEM (n 6/group). Statistical comparisons were made using a two-tailed unpaired Student’s t-test. ***P < 0.001. h Morphology and AP staining of patient-iPSCs. Scale bar, 100 μm. i Immunofluorescence of pluripotency markers in patient iPSCs. Scale bar, 100 μm. j Immunostaining for ectoderm (Tuj1, Map2), mesoderm (α-SMA, Vimentin), and endoderm (AFP, HNF-4α) markers. Fluorescence intensity of each marker was quantified from three independent experiments and shown as mean ± SEM. Scale bar, 100 μm. k Karyotype analysis of patient iPSCs. l STR analysis of BC patient fibroblasts and iPSCs
BC subtypes were identified in 14 patients and included luminal A (N = 1), luminal B (N = 5), HER2-enriched (N = 4), basal-like (N = 3), and sarcoma (N = 1). Primary cells derived from five out of 14 patients (35.7%) were successfully maintained beyond passage 5, and long-term cultures were subjected to whole-exome sequencing (WES) (Fig. 1b). Tumor tissues from patients whose cells were successfully expanded showed enhanced Ki-67 expression and reduced apoptotic activity compared to matched normal tissues, indicating a proliferative, apoptosis-resistant phenotype (Supplementary Fig. 1a, b). In normal tissues, pan-cytokeratin staining was localized to the ductal lumen, whereas it was diffusely distributed throughout the tumor epithelium, reflecting altered epithelial organization (Supplementary Fig. 1c). Flow cytometric analysis (FACS) revealed that epithelial (EpCAM) and basal (CD49f) cell populations, including the EpCAM+/CD49f+ double-positive subset, were stably maintained up to passage 5 in both normal and tumor-derived primary cultures (Supplementary Fig. 1d), suggesting that the preservation of this progenitor-like population may contribute to the sustained proliferative capacity during passaging.
Among them, Patient 1, who harbored a germline mutation in NF1, was selected for further molecular and functional investigation. Patients with NF1 mutations exhibited significantly faster metastatic recurrence in breast cancer. This effect was particularly pronounced for truncating and splice-site mutations, suggesting their role in relapse acceleration (Fig. 1c). Disease-specific survival was markedly reduced in NF1-mutated patients, with narrow survival distributions and shortened median durations compared to those in wild-type cases (Fig. 1d). NF1-mutant cells were resistant to various anticancer agents, including kinase inhibitors and cytotoxic drugs, such as Larotrectinib, Dasatinib, Tipifarnib, and Epothilone B, indicating a potential mechanism of intrinsic or acquired drug resistance (Fig. 1e).
To establish a personalized drug validation platform (NOCS) for patient 1 harboring an NF1 mutation, we generated iPSCs from somatic cells isolated from the patient’s non-tumorous (normal) tissue. Sanger sequencing revealed a germline insertion in NF1 exon 2 (c.165_166insCT), causing a frameshift and premature truncation that likely disrupted NF1 function in patient iPSCs (Fig. 1f). The patient iPSCs showed elevated OCT4 and NANOG expression when compared with the parental fibroblasts (Fig. 1g), displayed characteristic colony morphology, and stained positive for alkaline phosphatase (Fig. 1h). Immunofluorescence confirmed strong expression of the pluripotency markers OCT4, NANOG, TRA-1-60, TRA-1-81, SSEA3, and SSEA4 (Fig. 1i). Upon in-vitro differentiation, the iPSCs gave rise to derivatives of all three germ layers: ectoderm (MAP2 and Tuj-1), mesoderm (α-SMA and vimentin), and endoderm (AFP and HNF-4α), confirming their pluripotency (Fig. 1j). Karyotyping confirmed a normal chromosomal complement (46, XX) (Fig. 1k), and short tandem repeat profiling verified their genetic identity with the original fibroblasts (Fig. 1l). Patient iPSCs formed smaller, more compact colonies than healthy iPSCs derived CRL-2907 fibroblast (called normal iPSCs) (Supplementary Fig. 2a), although pluripotency gene (OCT4, NANOG, and SOX2) expression levels were comparable (Supplementary Fig. 2b). Immunostaining showed a similar distribution and intensity of OCT4, SOX2, SSEA4, and TRA-1-60 in both groups (Supplementary Fig. 2c). Embryoid-body formation or trilineage differentiation capacity did not differ significantly (Supplementary Fig. 2d, e). Transcriptome analysis revealed a strong similarity between normal and patient iPSCs, with a high correlation coefficient (R = 0.97). (Supplementary Fig. 2f). Importantly, patient iPSCs exhibited activated RhoA/ROCK signaling, leading to cytoskeletal disorganization with enhanced actin stress fibers and localized microtubule accumulation compared to the well-organized structure in normal iPSCs (Supplementary Fig. 2g, h). These findings suggest that NF1-mutant iPSCs displayed few morphological abnormalities and intact core pluripotency, supporting their utility in disease modeling and therapeutic development.
Generation and characterization of iPSC-derived intestinal epithelium (hIECs) from BC patients 1 and healthy controls for drug absorption
To model intestinal drug absorption compatible with NOCS, we first generated intestinal epithelial cells using intestinal stem cells (ISCs) derived from a healthy donor (hereafter referred to as Normal) and patient iPSCs. iPSCs were differentiated into intestinal organoids (IOs), from which proliferative ISCs were isolated following a previous report.27 The ISCs were seeded onto Trans-wells and cultured using the air-liquid interface (ALI) method, which mimics the polarized apical-basolateral environment to enhance epithelial maturation (Fig. 2a). Both normal and patient ISCs protruded apically, establishing a human intestinal villus-like 2.5-dimensional structure, yielding normal and patient-derived hIECs models, respectively (Fig. 2b). To assess cellular composition, we compared the mRNA expression of key intestinal lineage markers in organoids derived from normal and patient hIECs. The expression of the intestinal Crypt cell markers CD44 and LGR5 did not differ, whereas SOX9 expression was significantly increased in patient hIECs, suggesting enhanced crypt-like characteristics. The Paneth cell marker LYZ was significantly upregulated in patient hIECs, indicating increased Paneth cell differentiation. The expression of the differentiated epithelial cell markers VIL1 (enterocytes), MUC2, MUC13 (goblet cells), and CHGA (enteroendocrine cells) did not differ, suggesting that differentiation into mature lineages was largely preserved. Epithelial cell marker E-cadherin (ECAD) expression was similar in both groups (Fig. 2c). These findings suggested that while overall lineage differentiation was maintained, patient-derived organoids exhibited elevated crypt zone and Paneth cell signatures, potentially reflecting altered progenitor cell dynamics.
Generation of the hIECs model for drug absorption. a Schematic of the hIECs model establishment from iPSCs. b Bright-field images of normal (left) and patient-derived hIECs (right). Scale bar, 200 μm. c mRNA expression of intestinal cell type markers: CD44, SOX9, LGR5 (stem cell related), VIL1 (enterocyte), MUC2, MUC13 (goblet), LYZ (Paneth), CHGA (enteroendocrine), and ECAD (epithelial). d Immunofluorescence staining of intestinal cell-type markers and E-cadherin (top) with an enlarged view of the left panel (white box) and fluorescence intensity quantification (bottom). Scale bar, 200 μm. e mRNA expression of phase I/II drug metabolism markers. f CPR activity in normal and patient-derived hIECs. g TEER values of hIECs models and Caco-2 cells. h Immunofluorescence of CLDN1 and ZO-1. Scale bar, 200 μm. i Basolateral dextran-FITC uptake (4 kDa and 40 kDa) for passive permeability analysis. j Schematic (left) and mRNA expression (right) of intestinal influx/efflux transporters. A schematic illustration was created using Mind the Graph. k H&E staining (left) and immunofluorescence (right) of apical (VIL1, P-gp, PEPT1) and basolateral (Na⁺/K⁺ ATPase) markers. Nuclei stained with DAPI or Hoechst. Scale bar, 200 μm. Data are shown as mean ± SEM (n ≥ 4 per group). Statistical analysis for multiple groups (qPCR, IF, CPR activity, and TEER) was performed using one-way ANOVA with Tukey’s test for post hoc analysis, and data for IF was performed using a two-tailed unpaired Student’s t-test. *P < 0.05, **P < 0.01, ***P < 0.001, ns: not significant
To validate the transcriptional profiles of intestinal cell populations, we performed immunofluorescence staining of representative markers of differentiated epithelial lineages in normal and patient hIECs. Co-immunostaining of ECAD with lineage-specific markers (VIL1, MUC13, LYZ, and CHGA) was used to determine epithelial cell composition and marker localization after differentiation. The proportions of marker-positive cells were comparable between normal and patient organoids, except for goblet cells, which showed a significantly higher proportion in patient-derived organoids (Fig. 2d). These results confirmed that, despite transcriptional alterations in certain lineage-associated genes, the overall differentiation into major intestinal epithelial cell types was preserved in patient hIECs.
To evaluate the functional maturation of hIECs, we measured the expression of key phase I and II drug-metabolizing enzymes, NADPH cytochrome P450 (CYP450) oxidoreductase activity, and transepithelial electrical resistance (TEER) as an indicator of epithelial barrier integrity. mRNA levels of CYP3A4, CYP2C9, and GSTA1 were comparable between normal and patient hIECs, while CYP3A5 was significantly reduced in patient organoids, suggesting partial impairment of specific xenobiotic metabolic pathways (Fig. 2e). CYP450 reductase activity, a general indicator of enzymatic drug metabolism capacity, did not significantly differ between the groups (Fig. 2f). Patient hIECs exhibited significantly higher TEER values than normal model, although both were below the level of mature Caco-2 monolayers (Fig. 2g). Immunofluorescence staining for CLDN1 and ZO-1, markers of adherent junctions and TJs, respectively, showed intact and continuous localization at cell–cell borders in both groups (Fig. 2h).
To determine the suitability of hIECs for drug transport modeling in the NOCS, we evaluated the expression, localization, and activity of major intestinal influx and efflux transporters. Paracellular permeability assays using FITC-dextran (4 and 40 kDa) revealed significantly lower uptake in patient hIECs compared to normal hIECs, indicating enhanced TJ integrity and reduced passive diffusion, consistent with the elevated TEER values observed (Fig. 2i). Figure 2j illustrates the localization of uptake (PEPT1 and OATP1A2) and efflux (P-gp, MRP3, and BCRP) transporters across the apical and basolateral membranes and their contribution to drug movement across the intestinal epithelium. We next assessed the mRNA expression of key apical and basolateral transporters. Among influx transporters, PEPT1 expression was similar in both groups, whereas OATP1A2 expression was significantly increased in patient hIECs, suggesting an alteration in specific solute uptake capacity. As for efflux transporters, P-gp expression was unchanged, whereas MRP3 expression was significantly increased in patient hIECs, potentially affecting xenobiotic efflux dynamics (Fig. 2j). Both normal and patient-derived differentiated hIECs exhibited clear apical–basolateral polarity, as evidenced by apical localization of VIL1, P-gp, and PEPT1, basolateral localization of Na⁺/K⁺-ATPase, and lateral localization of ECAD and ZO-1, a tight junction marker. No significant differences were observed between the two models (Fig. 2k). These findings indicated that patient hIECs are suitable as an intestinal model in NOCS, retaining general epithelial traits, intestinal metabolic activity, and proper transporter expression. However, certain patient-derived organoids exhibited altered expression of specific transporters, such as OATP1A2 and MRP3, which could potentially affect drug absorption and disposition.
Generation and characterization of iPSC-derived liver organoids (LOs) from BC patient 1 and healthy controls for drug metabolism
To generate NOCS-compatible LOs from normal and patient iPSCs, we applied our protocols established based on the liver developmental process28 (Fig. 3a). Normal and patient-derived organoids were successfully generated and showed no overt morphological differences in each stage of differentiation, including the definitive endoderm, hepatic endoderm (data not shown), and expandable human liver organoids (EhLOs). However, in patient-derived differentiated human liver organoids (DhLOs), mesenchymal-like cells were observed around the epithelial layer (Fig. 3b).
Generation of the DhLOs model for drug metabolism. a Schematic of LOs generation from hiPSCs. b Representative morphology during DhLOs generation from normal and patient-derived iPSCs. Yellow arrowheads mark mesenchymal-like cells in patient-derived DhLOs, with an enlarged view provided in the right panel (yellow box). c mRNA expression of hepatocyte markers (ALB, AFP, CYP3A4) and cholangiocyte marker (KRT7) in normal and patient-derived DhLOs, compared to iPSCs. Data are mean ± SEM (n > 4 per group). d Immunofluorescence images of hepatocyte marker ALB and cholangiocyte marker KRT7 in DhLOs models. White arrowheads indicate cystic structures in patient-DhLOs. e Quantification of secreted ALB, ALT, and AST levels in EhLO and DhLOs. Data are mean ± SEM (n > 3 per group) (f) Enzyme activity of the CYP family in DhLOs with and without inducers (100 μM Omeprazole for CYP1A2 induction, 20 μM Rifampicin for other enzymes). RLU, relative luminescence unit. Data are mean ± SEM (n = 4 per group) g Fluorescence images of bile canaliculi-like structures stained with CLF in DhLOs, with an enlarged view of the left panel (white box) h mRNA expression of hepatic influx (top) and efflux (bottom) transporters in EhLO and DhLOs, compared to iPSCs. Schematic of hepatic drug transporters (lower left), created using Mind the Graph. Data are mean ± SEM (n > 4 per group). i Immunofluorescence images of influx transporter NTCP and efflux transporters (MRP2, BSEP, MRP4) in DhLOs. Data are mean ± SEM (n = 6 per group). Statistical comparisons were made using two-way ANOVA with Tukey’s test. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001
Albumin (ALB) and CYP3A4 mature hepatocyte marker expression differed minimally between normal and patient EhLOs, whereas their expression was reduced in patient DhLOs. The expression of AFP, a fetal hepatocyte and cancer marker, was increased in patient DhLOs compared to normal DhLOs. Notably, KRT7 cholangiocyte marker expression, was substantially increased in patient EhLOs and DhLOs (Fig. 3c). Importantly, in normal organoids, ALB-stained hepatocytes and KRT7-stained cholangiocytes were clearly distinguishable, whereas in patient DhLOs, cells showed strong co-staining for ALB and KRT7, and numerous small tubular structures resembling immature bile ducts were observed (Fig. 3d). In addition, patient DhLOs showed decreased expression of epithelial markers (ECAD, EPCAM, and KRT18) and increased expression of mesenchymal markers (ACTA2, COL1A1, and VIM), consistent with the presence of COL1A1-positive mesenchymal cells (Supplementary Fig. 3).
To assess patient DhLOs functionality, serum protein synthesis was measured. ALB secretion was reduced by 39% in patient DhLOs compared to normal DhLOs. Serum alanine (ALT) and aspartate aminotransferase (AST) levels were 2.45-fold and 2.64-fold higher, respectively, in patient DhLOs than in normal DhLOs (Fig. 3e). We next assessed the expression and activity of drug-metabolizing enzymes essential for liver function. Patient DhLOs exhibited lower mRNA levels of major phase I (CYP1A2, 2C8, 2C9, 2C19, and 2E1) and phase II (UGT1A1, 1A3/4/5, 2B7, and 2B15) enzymes compared to normal organoids (Supplementary Fig. 4), consistent with reduced activities of key phase I enzymes, including CYP1A2, 2C8, 2C9, 2C19, and 3A4, in patient organoids (Fig. 3f). Notably, drug-induced activation of CYP enzymes, observed in normal organoids, was largely absent in patient DhLOs, except for CYP2C9 and CYP2C19.
Transporter activity of polarized hepatocytes was assessed using cholyl-L-lysyl fluorescein (CLF) staining, revealing a significant decrease in patient DhLOs compared to normal organoids (Fig. 3g). Well-developed bile canaliculi-like structures were observed in normal DhLOs, whereas these structures were absent in patient DhLOs, which instead exhibited an abnormal dot-like pattern. These findings were further validated by CLF z-stack confocal imaging and 3D reconstruction (Supplementary Videos 1–4).
Expression levels of influx (OATP1B1, OATP1B3, OATP2B1, and OCT1) and efflux (MRP2 and MRP4) transporters were decreased in patient DhLOs compared to normal DhLOs., were decreased in patient DhLOs compared to normal DhLOs. NTCP expression did not differ between normal and patient DhLOs, whereas MDR3 expression was increased (Fig. 3h). Transporter localization was disrupted in patient DhLOs, as indicated by immunostaining (Fig. 3i). In normal DhLOs, influx (NTCP) and efflux transporters (MRP2 and BSEP in bile canaliculi; MRP4 in bloodstream) were clearly segregated, whereas they were co-localized in patient DhLOs. Collectively, these findings indicate that patient DhLOs exhibit reduced phase I/II drug-metabolizing enzyme and transporter expression and activity, accompanied by abnormal phenotypic features, compared to normal DhLOs.
Generation and comparative characterization of iPSC-derived renal tubule epithelium from BC patient 1 and healthy controls for drug reabsorption
To establish a physiologically relevant kidney reabsorption model for NOCS, tubular epithelial cells (hKORTECs) were differentiated from normal and patient iPSCs via a stepwise nephron induction protocol (Fig. 4a, b). Kidney organoids (KOs) were first generated from normal and patient iPSCs, and developed nephron-like structures, including proximal and distal tubules, collecting ducts, and podocyte clusters, by day 18 of differentiation (Supplementary Fig. 5a, b). These structures exhibited increased expression of kidney lineage markers compared to undifferentiated iPSCs, including podocyte markers (NPHS1, NPHS2), proximal tubule markers (AQP1, SLC3A1), loop of Henle (SLC12A1), distal convoluted tubule (NCC), and collecting duct (AQP2). Patient-derived hKOs showed increased NPHS2 and AQP2, but decreased SLC12A1 expression relative to normal hKOs. Immunostaining confirmed multi-lineage tubular markers, with more prominent collecting duct differentiation in patient-derived hKOs (Supplementary Fig. 5c, d). To establish a reproducible reabsorption model, cryopreservable and expandable kidney tubule organoids (hKTOs) were generated from normal and patient-derived hKOs. Both types expressed nephron segment markers (AQP1, UMOD, and AQP2) and renal transporters (OCT2, PEPT1/2, MRP2, and P-gp), with MRP2 expression notably elevated in patient-derived hKTOs (Fig. 4c, d). hKTOs were cultured under ALI conditions to generate polarized hKORTEC models. Within the NOCS platform, hKORTECs functioned as a reabsorption model, exhibiting nephron segment identity, renal transporter expression, and epithelial polarity. Immunofluorescence analysis confirmed apical–basolateral polarity and the expression of ZO-1, Na⁺/K⁺-ATPase, P-gp, OCT2, OAT3, MRP2, and PEPT1 (Fig. 4e). TEER was significantly higher in patient hKORTECs, indicating tighter barrier formation (Fig. 4f). Dextran-FITC permeability assays supported this, showing less leakage in patient hKORTECs than in normal hKORTECs (Fig. 4g). Albumin uptake via LRP2/megalin showed functional endocytosis in both groups, with slightly enhanced uptake in patient hKORTECs (Fig. 4h, i). hKORTECs were treated basolaterally with carboxy-dichlorofluorescein diacetate (CDFDA), which is converted to carboxy-dichlorofluorescein (CDF) by intracellular esterases. CDF fluorescence was detected apically, indicating active efflux via MRP2/4 transporters. Treatment with MK-571, an MRP2/4 inhibitor, blocked this efflux and led to intracellular accumulation of CDF, confirming transporter activity (Fig. 4j). To assess hKORTECs for transport-mediated nephrotoxicity, cisplatin treatment induced significant toxicity in both normal and patient-derived models. Co-treatment with cimetidine, an OCT2 inhibitor, suppressed cisplatin-induced cell death, indicating functional OCT2 activity and supporting the model’s suitability for nephrotoxicity studies. No significant differences were observed between normal and patient-derived hKORTECs in their response to cisplatin or cimetidine (Fig. 4k).
Generation of the hKORTECs model for drug reabsorption. a Schematic of the hKORTECs model generation from hiPSCs. b Representative morphology of normal and patient-derived hKORTECs. Scale bar, 100 μm. c Schematic of hKORTECs generation from hKOs (top), H&E staining, and immunofluorescence images of renal tubule markers (AQP1, UMOD, AQP2) in KTO models. Scale bar, 100 μm. A schematic illustration was created using Mind the Graph. d Schematic of renal drug transporters (lower left), created using Mind the Graph. mRNA expression of renal influx (OCT2, PEPT1, and PEPT2) and efflux (MRP2, P-gp) transporters in hKTO models. Data are mean ± SEM (n = 6/group). Statistical comparisons were made using a two-tailed unpaired Student’s t-test. **p < 0.01, ns: not significant. e IF of bidirectional markers (ZO-1, P-gp, MRP2, PEPT1, Na+/K+ ATPase, OCT2, OAT3) in hKORTECs. Scale bar, 100 μm. f, g TEER values and basolateral dextran-FITC leakage in normal and patient-hKORTECs. Mean ± SEM (n = 4 per group); unpaired two-tailed t-test. **p < 0.01. h IF of endocytic receptor megalin in hKORTECs. Scale bar, 100 μm. Mean ± SEM (n = 6/group); unpaired two-tailed t-test. *p < 0.05. i Functional analysis of megalin-mediated albumin trafficking in hKORTECs. Mean ± SEM (n = 5/group); unpaired two-tailed t-test. **p < 0.01. j Summary of transepithelial transport and MRP2/4-mediated apical efflux in normal and patient-hKORTECs (top), and apical CDF fluorescence with/without MK571. Data are mean ± SEM (n = 5/group). Statistical analysis was made using two-way ANOVA with Tukey’s test. k Summary and cytotoxicity assessment following cisplatin and cimetidine treatment via drug transporter modulation. Data are mean ± SEM (n = 5/group); two-way ANOVA with Tukey’s test. *P < 0.05 is considered significant. ns: not significant. Schematic illustrations included in (i–k) were created using Mind the Graph
These findings indicate that both normal and patient-derived hKORTECs recapitulated key in vivo kidney features, including epithelial polarity, transporter expression, and reabsorption/excretion functions. Patient-derived hKORTECs showed improved TJ integrity and albumin uptake, highlighting their potential for personalized drug screening and kidney disease modeling.
Identification of patient-specific molecular characteristics in primary cells isolated from BC patient 1 tissue
A patient-derived cancer model for personalized drug evaluation was successfully established using multicellular primary cells isolated from the tumor and matched normal BC tissues. The cells formed stable spheroids, which were embedded in a matrix and cultured for 3–5 days. The spheroids demonstrated the potential to model patient-specific drug responses, suitable for further analysis (Fig. 5a). Patient-derived 3D spheroid models of tumor and paired normal tissues were characterized using pan-cytokeratin and vimentin immunohistochemistry, which revealed preserved epithelial and stromal cell composition comparable to that in the original patient tissues (Fig. 5b). FACS analysis using EpCAM and CD49f showed that the epithelial and stromal populations remained stable up to passage 7 (Fig. 5c). To assess the expansion potential of multicellular primary cells derived from tumor tissue, we analyzed the BC stem cell population. We found that 63% of cells within tumor-derived multicellular populations expressed cancer stem cell markers at significantly higher levels than cells derived from normal tissue (Fig. 5d). This population remained stable up to passage 5. All subsequent analyses were performed using multicellular primary cells at or below passage 5.
Molecular characterization and therapeutic NF1 exon skipping in BC patient 1 tumoroids. a Schematic of 3D BC modeling using primary cells from normal and tumor tissues, created using Mind the Graph. b H&E and IHC images (pan-CK, vimentin) showing structural similarity between patient tissues (Figs. S2c) and 3D models. c FACS analysis of epithelial (EpCAM) and stromal (CD49f) populations across passages 1–7. d FACS of BC stem cell markers (CD44/CD24) at passage 3. e WES showing genetic similarity between tissues and derived cells. f Western blot of NF1/Ras pathway proteins in normal and tumor-derived primary cells. Band intensities were quantified and normalized to the normal control (set as 1). g Western blot of Her2, MEK/ERK, and PI3K/AKT signaling pathways in tissues and primary cells. Band intensities were quantified and normalized to the respective control, with values shown below each blot. h Schematic of AONs targeting NF1 exon 2. i RT-PCR of exon 2 skipping after 300 nM AON treatment in patient-derived cells. j Sanger sequencing confirming precise exon 2 skipping with AON12. k Schematic of U7snRNA lentiviral constructs (left); RT-PCR analysis of NF1 exon 2 skipping after lenti-U7 AON transduction (right). l, m RT-PCR showing time- and dose-dependent skipping efficiency using lenti-U7-AON2/12 (MOI 10). Lenti-zsGreen was used as control. n Western blot confirming NF1 exon 2 deletion after lentiviral transduction. Band intensities are indicated below each blot. All results were validated through at least three independent experiments
To evaluate the genetic similarity between patient tissues and isolated primary cells, tumor and normal tissues and their corresponding primary cells were analyzed using WES. Normal tissues and derived cells were 96.2% genetically similar, while tumor tissues and derived cells were 97.7% genetically similar. In the tumor tissue, we identified a loss-of-function mutation in NF1 (c.165_166insCT, p.I55fsX) and gain-of-function mutations in PIK3CA (c.3139 C > T, p.H1047Y) and ERBB2/HER2 (c.2711 A > T, p.D904V) (Fig. 5e). We therefore investigated alterations in the downstream NF1-RAS, PI3K-AKT, and MEK-ERK signaling pathways, and found that tumor-derived primary cells exhibited increased activation of these pathways compared to their normal counterparts (Fig. 5f–g).
Exon skipping of the NF1 mutant in BC patient 1-derived primary cells as a potential therapy
To improve drug responsiveness in patients with BC harboring NF1 mutations, we assessed exon-skipping therapy as an advanced personalized therapeutic strategy. WES identified a novel frameshift mutation in exon 2 of NF1, c.165_166insCT (p.I55fsX), which was located in a highly conserved region (Supplementary Fig. 6a). We identified exonic splicing enhancer (ESE) and silencer sites within NF1 exon 2 using the Human Splicing Finder and selected candidate antisense oligonucleotides (AONs) targeting ESE sites (Supplementary Fig. 6b). Because AON-binding sites within double-stranded RNA regions may be inaccessible, reducing exon-skipping efficacy, we analyzed the RNA secondary structure of exon 2 and designed AONs to bind to loop or single-stranded regions (Supplementary Figs. 6c and 5h). We designed 13 AONs targeting two 5′ splice sites, one 3′ splice site, and 10 ESE sites. To limit intracellular degradation, all AONs were modified with 2′-O-methyl and a phosphorothioate backbone (Supplementary Tables 8 and 9). To assess nuclear localization, which is essential for AON-mediated pre-mRNA splicing, AON12 was conjugated with a 3′-FAM dye and delivered to HEK293 cells. It localized to the nucleus and persisted for up to 72 hours (Supplementary Fig. 6d). Exon-skipping efficacy was evaluated by transfecting AONs into BC-derived primary cells, with exon 2 skipping confirmed by RT-PCR (Fig. 5i) and sequencing (Fig. 5j). As synthetic AONs are generally effective for only 2–3 days, repeated administration is necessary for a sustained effect. To overcome this limitation, we employed the U7 small nuclear RNA (U7-snRNA) system, which enables long-term AON expression. We constructed six lentiviral U7-smOPT vectors: five expressing dual AONs targeting the 3’-splice site (3’-SS; AON2) and ESE sites (AON6, AON8, AON9, AON12, and AON13), and one targeting both the 3’-SS and 5’-SS (AON7) (Supplementary Table 11). Selected AONs with high exon-skipping efficacy were evaluated as lenti-U7 constructs in BC patient-derived primary cells. The lenti-U7-AON2/12 construct exhibited the highest exon-skipping efficiency (Fig. 5k). Moreover, the lentiviral construct sustained exon skipping for up to 7 days post-transduction (Fig. 5l) and showed a dose-dependent effect (Fig. 5m). Importantly, NF1 exon skipping led to increased NF1 expression and reduced PI3K-AKT signaling (Fig. 5n).
These results demonstrated that exon skipping using a lentiviral U7-snRNA system effectively restores NF1 expression and suppresses PI3K-AKT signaling in BC-derived primary cells harboring NF1 mutations and therefore represents a promising personalized therapeutic approach to overcome drug resistance in patients with BC with NF1 mutations.
NOCS for PK/PD assessment of precision therapeutics
The NOCS was developed to closely replicate in-vivo organ connectivity and physiological fluid dynamics, providing a dynamic microenvironment that supports nutrient transport, drug metabolism, and waste elimination. By integrating multi-organ interactions in a controlled fluidic system, NOCS offers a robust platform for evaluating drug absorption, metabolism, and efficacy in a physiologically relevant context.
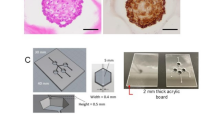
To identify optimal personalized therapeutics for a patient with BC, we generated patient iPSCs from somatic cells and differentiated them into intestinal, hepatic, and renal organoids, representing critical sites of drug absorption, metabolism, and reabsorption, respectively. A tumor model was established using patient-derived cancer cells to reflect tumor-specific characteristics. All organoids were cultured in a multi-insert dish engineered to simulate the oral drug delivery route, forming the NOCS platform (Fig. 6a). The NOCS comprised four interconnected culture chambers containing the different organoids and was continuously perfused with nutrient medium. A load cell and piezoelectric pump in each chamber maintained the medium volume around 25 mL per unit. Sensor output and applied weight were linearly correlated in all load cells, confirming measurement accuracy. Temperature-dependent variations in sensor output were observed, indicating the need for thermal calibration. The piezoelectric pumps showed a consistent nonlinear relationship between pulse count and flow rate, validating their reliability in maintaining controlled fluid delivery across channels (Supplementary Fig. 7). Fresh medium was supplied at a rate of 2 mL/min for 2 min every hour, and an equal volume of spent medium was simultaneously removed. To ensure homogeneous drug distribution, the system incorporated a motorized orbital shaker (Fig. 6b). The multi-insert dish was optimized for drug absorption and reabsorption assays, with a mesh at the bottom. A modified version, designed for metabolic and efficacy testing, included a side mesh to maximize interaction with the circulating medium (Fig. 6c). Organoid models differentiated from normal iPSCs cultured in the system exhibited structural features of their in-vivo counterparts: intestinal and kidney organoids formed epithelial and tubular architectures, while liver organoids displayed luminal structures. Over 4 days of culture, the models retained their characteristic features. (Fig. 6d).
NOCS platform for personalized PK/PD evaluation. a Schematic of NOCS for personalized therapeutic applications, integrating intestine, liver, and kidney organoids to model drug absorption, metabolism, and reabsorption. Patient-derived normal and tumor cells enable personalized medicine studies. b Experimental setup, showing the multi-insert dish within a controlled environment, connected to reservoirs for fresh medium supply and waste collection. Piezoelectric pump and load cell sensors automate fluid flow control. c Microfluidic chamber layout and system configuration, illustrating the modular design of the multi-organ system for organoid culture. d Organoid cultures of intestine, liver, and kidney derived from normal iPSCs. Representative images showing morphological changes of organoids at day 0 and day 4. e Comparative PK analysis of drug absorption, metabolism, and reabsorption in closed and semi-open systems. Drug concentration-time profiles via LC-MS/MS show PK variations across conditions. All schematic illustrations in Fig. 6 were created using BioRender
In previous studies,20 NCCS were implemented as closed systems (Supplementary Fig. 7a, left), limiting their ability to accurately replicate in vivo PK profiles, particularly for drugs cleared primarily by the kidneys. Compounds, such as Acyclovir and Chlorothiazide, characterized by minimal hepatic metabolism and predominant renal clearance, failed to exhibit physiologically relevant clearance patterns in the closed configuration. This limitation was likely due to metabolite accumulation and the absence of continuous medium exchange, which are essential for mimicking systemic drug elimination. The semi-open system, named NOCS, introduced here yielded PK profiles for these drugs that more closely mirrored in-vivo behavior, indicating improved simulation of metabolic and excretory processes. Model drugs, such as Caffeine and Acetaminophen, which follow distinct metabolic pathways, also displayed in vivo-like PK behaviors in this platform (Fig. 6e). These findings underscore the enhanced capacity of the upgraded NOCS to serve as a physiologically relevant platform for PK/PD studies.
Precise PK profiling of BC patient 1 harboring NF1-mutant using the NOCS platform
To evaluate the genomic fidelity of patient iPSCs, we compared their genome-wide variant profiles with those of normal iPSCs. Strong correlations were observed for intestinal and kidney models (r = 0.99), with slightly lower correlation in liver models (r = 0.96), indicating minimal divergence and supporting their use for downstream PK/PD studies (Supplementary Fig. 8). Next, we assessed the transcriptomic similarity of intestinal, liver, and kidney organoids derived from patient and normal iPSCs, to their corresponding native human tissues using a set of 61 drug transport-related genes critical for in vivo drug disposition, as well as within the context of NOCS platform.
In the results, organoids derived from normal and patient iPSCs showed relatively high similarity to their corresponding human tissues. Small intestinal organoids showed correlation values of 0.78–0.90 in normal and 0.79–0.87 in patient, liver organoids showed 0.78–0.82 in both groups, and kidney organoids showed 0.88–0.91 in both normal and patient (Fig. 7a). Heat map analysis of 61 drug transporter genes revealed that organoids derived from normal and patient iPSCs closely resembled their corresponding human tissues, with highly consistent expression profiles observed across two tissue samples and three organoid replicates, indicating minimal batch-to-batch variation. These results highlight the reproducibility of hIEC, DhLO, and hKORTEC organoids and support their suitability as predictive models for human drug response (Fig. 7b).
Precise PK profiling of BC patient 1 using human mimicking organoid-integrated NOCS platform. a Schematic of key drug transporters in the intestine, liver, and kidney (left), created using Mind the Graph, and a Pearson correlation matrix of expression levels of 61 drug transporter genes between iPSC-derived organoids (N = 3 for each normal (N1-3) and patient-derived (P1-3) organoid) and the corresponding human tissues (N = 2) (right). b Heatmaps comparing transporter gene expression between human tissues (SI, Liver, Kidney) and the corresponding organoids. c Schematic of PK evaluation for Bimiralisib, Paxalisib, and Gedatolisib in vivo and in the NOCS platform (left), created using Mind the Graph. Drug concentration-time profiles from mouse plasma (IV 2 mg/kg; PO 10 mg/kg; N = 3 ~ 5 mouse/group) and NOCS-based organoid models (liver: 2.5–10 nmol IV; intestine: 25 nmol PO; N = 3 independent experiments)
To identify targeted anti-cancer drugs for personalized medicine, we selected candidates based on the patient’s WES results. NF1 loss-of-function mutations activate Ras signaling, including the downstream PI3K-mTOR pathways (Supplementary Fig. 9a). Screening of targeted inhibitors in patient-derived breast tumor cells revealed that PI3K-mTOR inhibitors LY294002 and rapamycin exhibited the strongest selective cytotoxicity compared to matched normal cells (Supplementary Fig. 9b), leading us to focus on clinical-stage inhibitors Alpelisib and Everolimus. As Everolimus showed negligible absorption in the NOCS platform, we evaluated Alpelisib, which exhibited poor absorption and hepatic metabolism, with lower intracellular uptake in patient organoids than in normal organoids, likely due to differences in transporter expression or uptake mechanisms (Supplementary Fig. 9c, d). To identify effective PI3K-mTOR inhibitors with favorable PK profiles for NF1-mutant patients, we screened eight dual inhibitors using normal intestinal and liver organoids (Supplementary Fig. 9e, Supplementary Table 12). Apitolisib, Bimiralisib, Paxalisib, Samotolisib, and Voxtalisib showed moderate intestinal absorption, while Apitolisib, Samotolisib, and Voxtalisib exhibited limited hepatic metabolism, suggesting potential for enhanced systemic exposure. Gedatolisib, an FDA-approved IV drug, displayed minimal intestinal absorption and high hepatic metabolism, consistent with its clinical administration (Supplementary Fig. 9f, g). Bimiralisib, Paxalisib, and Gedatolisib were further evaluated in normal and patient-derived organoids. Bimiralisib and Gedatolisib showed increased first-pass metabolism in patient hIECs, whereas Paxalisib remained unchanged; in DhLOs, only Bimiralisib showed elevated metabolism (Supplementary Fig. 9h, i). Based on these results, we prioritized oral Bimiralisib and Paxalisib, and IV Gedatolisib, for evaluation of concordance between in vivo and in vitro PK profiles in patient-specific NOCS platforms to select the most suitable oral compound (Supplementary Table 13 and Table 14).
Initial PK profiling was performed in mice, using both IV (2 mg/kg) and oral (10 mg/kg) routes. The results, particularly for the oral route, were compared with PK data from normal NOCS platforms. Bimiralisib and Paxalisib showed strong in vivo–in vitro concordance for T_max, AUC, and half-life. Bimiralisib exhibited rapid absorption (T_max = 0.42 h) and moderate oral exposure (AUC_inf = 10,799.72 ng·h mL–1), with similar trends in the NOCS (AUC_inf = 645.095 nmol/L·h). Paxalisib showed the highest oral uptake (AUC_inf = 43,421.7 ng·h mL–1) and sustained systemic presence, which was well represented in the NOCS (AUC_inf = 1140.9 nmol/L·h). Gedatolisib demonstrated high IV uptake but poor oral absorption, with negligible AUCs in both models. PK comparisons between normal and patient NOCS revealed disease-specific alterations in drug disposition. Bimiralisib showed a higher oral AUC (1417.02 nmol/L·h) and longer half-life (72.38 h) in the patient NOCS, suggesting reduced clearance. Paxalisib exhibited increased exposure (AUC_inf = 1553.97 nmol/L·h) and prolonged T_1/2 (56.93 h), indicating altered absorption kinetics and systemic persistence under NF1-mutant conditions. Gedatolisib maintained consistent IV AUC across models but had poor oral PK, regardless of patient status. Based on patient-specific oral exposure and bioavailability, Paxalisib emerged as the most promising candidate, with the highest AUC and consistent PK across models (Fig. 7c).
These findings highlight the predictive value of the NOCS platform for human PK behavior and disease-specific drug disposition. Given its favorable in-vivo and in-vitro PK profiles, Paxalisib is a lead candidate for further preclinical and translational development for NF1-mutated BC.
Patient 1-specific PD evaluation of combination therapy in NF1-mutant BC using the NOCS platform
To identify the most effective anti-cancer drug for combination therapy, we tested three dual PI3K-mTOR inhibitors in patient-derived breast tumor and matched normal cells. Gedatolisib showed the strongest tumor-specific potency (IC50 = 6.1 μM), whereas Paxalisib and Bimiralisib were less potent (IC50 = 19.4 μM and 15.7 μM, respectively) (Supplementary Fig. 10a). When combined with NF1 exon-skipping therapy (using lenti-U7-AON2/12), these inhibitors showed enhanced cytotoxicity. Paxalisib showed the most significant sensitivity shift; its IC50 decreased from 16 μM to 1.2 μM after exon skipping (Supplementary Fig. 10b). This combination effectively inhibited tumor growth, reduced proliferation, increased apoptosis, and suppressed PI3K-mTOR signaling and tumor cell migration (Supplementary Fig. 10c–h). Thus, Paxalisib emerged as the most promising candidate, showing high selectivity and strong synergy with exon-skipping therapy. Therefore, we next evaluated its efficacy and toxicity using the NOCS platform.
The workflow of the patient-specific NOCS platform for PD analysis, incorporating patient-derived intestinal, liver, and kidney organoids, along with a primary BC spheroid model, is shown in Fig. 8a. Bright-field images of the hIECs, DhLOs, and hKORTECs confirmed stable morphology and viability throughout the assay, indicating structural integrity under NOCS conditions (Fig. 8b). To assess organ-specific responses to Paxalisib, we quantified changes in functional markers and injury-associated transcripts. In the intestinal model, Paxalisib decreased VIL1 and LYZ expression while enhancing MUC13 expression, indicating injury protection. FABP2, HO-1, and GCLC were upregulated, suggesting epithelial stress and inflammation (Fig. 8c, left panel). In the hepatic model, the liver injury marker KRT7 and CYP2E1 levels were elevated, whereas CYP3A4 expression was decreased, and oxidative stress markers (NRF2, HO-1, NQO1, and GSTA1) showed increased expression, indicating hepatocellular injury (Fig. 8c, middle panel). In the renal model, CDH1, KIM1, and LCN2 marker and oxidative stress marker levels were elevated, suggesting renal dysfunction (Fig. 8c, right panel). These results highlight that Paxalisib induced organ-specific perturbations and stress responses in the NOCS platform.
BC patient 1 specific PD evaluation of combination therapy using NOCS platform. a Schematic representation of the PD assay using patient-derived organoid models in a NOCS, created using Mind the Graph. b Representative morphology of patient-derived organoid models after Paxalisib treatment for PD assay. Scale bar, 100 μm. c mRNA expression of cell-specific injury markers after Paxalisib treatment. Mean ± SEM (n = 6/group); unpaired two-tailed t-test. *P < 0.05, **P < 0.01. d Cell viability and TEER values of patient-derived organoid models after Paxalisib treatment. Mean ± SEM (n = 8/group); unpaired two-tailed t-test. *P < 0.05, ns: not significant. e LC-MS analysis of intracellular Paxalisib concentration in patient-derived tumor models after NOCS assay. Mean ± SEM (n = 4/group); unpaired two-tailed t-test. ns: not significant. f Cell viability of patient-derived tumor models after combination treatment with lentivirus-AON exon skipping and Paxalisib in a NOCS. Data are mean ± SEM (n = 6/group). Statistical comparisons were made using two-way ANOVA with Tukey’s test. *P < 0.05, **P < 0.01. g, h Representative immunofluorescence images of Ki-67 and active-Caspase 3 staining in patient-derived tumor models after combination treatment. Scale bar, 100 μm. Lentivirus-zsGreen was used as a control. i Western blot analysis of PI3K-AKT-mTOR signaling in patient-derived tumor models after combination treatment. j Western blot analysis of apoptotic cell signaling in patient-derived tumor models after combination treatment. Band intensities are indicated below each blot. Western blot and IF were performed with 3 biological replicates
To evaluate the toxicity of Paxalisib on each organoid, we assessed the cell viability in hIECs, DhLOs, and hKORTECs, and TEER in hIECs and hKORTECs. In hIECs, paxalisib induced a modest but significant reduction in TEER, suggesting compromised barrier integrity, although cell viability remained unchanged (Fig. 8d, left panel). Paxalisib treatment did not significantly affect cell viability in DhLOs (Fig. 8d, middle panel). In the hKORTECs model, as in the hIECs model, no significant change in cell viability was observed; however, TEER values were significantly decreased, suggesting a compromised epithelial barrier in response to treatment (Fig. 8d, right panel). Following the pre-screening of the combination therapy, we evaluated its effect in the NOCS platform. Intracellular drug concentrations did not significantly between the control (lenti-zsGreen) and NF1 exon-skipping (lenti-U7-AON2/12) groups (Fig. 8e). However, both groups showed reduced tumor cell viability, with a stronger effect in the NF1 exon-skipping group, indicating synergy (Fig. 8f). Tumor spheroids in the NF1 exon-skipping group exhibited reduced Ki-67 and increased active caspase-3 expression, confirming enhanced proliferation suppression and apoptosis (Fig. 8g, h). The combination of Paxalisib and NF1 exon skipping decreased PI3K-mTOR activation and enhanced apoptosis signaling compared to those in the controls (Fig. 8i-j). NF1 exon skipping alone also suppressed PI3K-mTOR signaling, suggesting functional recovery from NF1 exon skipping.
To validate the therapeutic synergy in vivo, primary breast tumor cells from the NF1 patient were subcutaneously implanted into nude mice. Lenti-U7-AON2/12 for NF1 exon skipping was intratumorally delivered when tumors reached ~50 mm³, followed by oral Paxalisib treatment every three days (Supplementary Fig. 11a). Consistent with NOCS results, combined NF1-AON and Paxalisib treatment markedly suppressed xenograft tumor growth and weight, reduced proliferation, and increased apoptosis compared to either monotherapy, as shown by active caspase-3 staining (Supplementary Fig. 11b–e), without significant changes in body weight or histopathological abnormalities in major organs (Supplementary Fig. 11f–h). These results demonstrate that NF1 exon skipping-induced restoration of NF1 function enhances the anticancer efficacy of Paxalisib through dual PI3K/mTOR inhibition and apoptosis induction, validating the synergistic efficacy and safety of the combination in both the patient-specific in vivo model and the NOCS platform.
Discussion
We developed and validated a NOCS, a multi-organ MPS that enables dynamic inter-organ interaction and recapitulates systemic PK and PD in a patient-specific context. By incorporating iPSC-derived organoids from an NF1-mutant patient with BC into a dynamic microfluidic circuit, we successfully established NOCS models mimicking patient-relevant drug absorption, metabolism, distribution, and excretion with high physiological fidelity. Compared to conventional organ-on-a-chip systems,29 the NOCS platform supports the integration of fully matured, tissue-specific organoids across multiple compartments. Furthermore, its capacity for scalable recovery of culture media and organoid samples post-experimentation facilitates comprehensive PK/PD and molecular analyses, reinforcing its translational applicability.
Each organoid type retained hallmark functions: intestinal organoids maintained epithelial barrier integrity, kidney organoids expressed functional drug transporters, and liver organoids exhibited CYP450 activity. While no remarkable differences were observed between patient-derived and normal intestinal or kidney organoids, liver organoids from the patient displayed mesenchymal features, including increased expression of COL1A1, VIM, and ACTA2, and decreased EPCAM and ECAD expression. These alterations are consistent with epithelial-mesenchymal transition driven by Ras pathway hyper-activation due to NF1 loss-of-function and were associated with altered hepatic drug handling, including upregulation of MDR3 (ABCB4).30 To further confirm that these hepatic abnormalities were directly attributable to NF1 deficiency rather than patient-specific background, we generated an isogenic control by restoring NF1 expression in patient-derived iPSCs using a lentiviral NF1-AON strategy (Supplementary Fig. 12). When differentiated into organoids, the AON-corrected line produced intestinal and kidney organoids that were indistinguishable from those derived from normal control iPSCs (data not shown), whereas the liver organoids exhibited partial recovery of ALB and CYP450 expression along with a marked attenuation of mesenchymal features (Supplementary Fig. 13). These findings confirm that NF1 loss primarily affects hepatic differentiation and function, reinforcing the causal link between NF1 dysfunction, mesenchymal transition, and impaired drug metabolism in the liver organoid model. In patient-derived tumor models, we observed sustained Ras pathway hyper-activation, including elevated expression of MEK/ERK and PI3K/AKT signaling components. These alterations are consistent with previous findings implicating NF1 loss-of-function in Ras-driven tumorigenesis and drug resistance.5 Loss-of-function mutations in NF1 increase the risk of BC before the age of 50 years by more than five times.31 In women with BC with mutant NF1, 69.2% of tumors are HER2-positive, which is associated with poor prognosis. Furthermore, patients with BC harboring NF1 mutations show reduced overall survival compared to those without because of mutation-induced resistance to chemotherapy, endocrine therapy, and targeted therapies due to sustainably activated Ras and related signaling.32,33,34,35 To directly address the genetic basis of NF1 dysregulation, we designed AONs targeting exon 2 regions identified in the NF1 mutant to induce exon skipping of the NF1 mutation. While synthetic AONs can transiently modulate splicing events, their therapeutic window is typically limited to 2–3 days, necessitating repeated dosing. To overcome this limitation, we employed a lentiviral U7-snRNA delivery system that enables sustained nuclear AON expression and prolonged exon-skipping activity. The U7-snRNA platform, developed for inherited disorders, such as Duchenne muscular dystrophy and spinal muscular atrophy,36,37,38,39 demonstrated comparable efficacy in our cancer model. In patient-derived primary BC cells harboring an NF1 frameshift mutation, exon 2 skipping using the U7 snRNA system restored NF1 protein expression and suppressed PI3K-AKT signaling. Notably, when combined with PI3K/mTOR inhibition, lenti-AON-U7snRNA targeting NF1 further suppressed tumor growth, highlighting the therapeutic potential of combining transcript-level correction with pathway-targeted therapies.40 While AON-mediated exon skipping is traditionally used for rare genetic diseases, recent studies have expanded its potential to oncology, where it offers a gene-specific strategy to modulate aberrant splicing events or silence oncogenic transcripts.41,42 Our findings establish lentiviral U7-snRNA-mediated exon skipping as a promising, mechanism-based therapeutic approach for NF1-mutant BC and support its further development for genetically defined, treatment-refractory tumors. While this study focused on an NF1-mutant breast cancer patient, the tumor also harbored other actionable alterations, such as HER2 aberrations. The NOCS platform is flexible and could, in principle, be applied to evaluate patient-specific responses to HER2-targeted therapies, alone or in combination with NF1-AON–based strategies, highlighting its potential for broader precision oncology applications.
PK analysis of three dual PI3K/mTOR inhibitors (Bimiralisib, Paxalisib, and Gedatolisib) using the NOCS platform recapitulated in vivo mouse data, confirming its translational relevance. Among these inhibitors, Paxalisib demonstrated favorable PK properties and the most consistent systemic exposure across various organ compartments. Considering patient convenience, Paxalisib was selected based on its superior bioavailability (F% 40.55) compared to Bimiralisib, which has a bioavailability of only (F% 18.46). Its superior bioavailability, reflecting more efficient absorption and systemic distribution, rendered Paxalisib a promising candidate for personalized treatment in this study. In contrast, although Gedatolisib received FDA approval in 2022, it was formulated as a non-oral compound. This distinction was also recapitulated in the NOCS platform, where Gedatolisib exhibited poor intestinal absorption and limited systemic exposure compared to Paxalisib. These findings underscore the translational utility of the NOCS in predicting patient-relevant pharmacokinetic behaviors, including oral bioavailability. In patient-derived NOCS, hepatic clearance slightly reduced, compared to those in normal, suggesting that the observed PK profile reflects the characteristics of patient-liver organoid and intrinsic features of NF1-mutant tissue.43 Although originally developed for glioblastoma, our results suggest that Paxalisib is also effective in NF1-mutant BC, supporting its potential for broader use in Ras-driven, treatment-resistant tumors.44,45,46 NOCS represents a modular platform for patient-specific PK/PD analysis and therapeutic screening. Its ability to model organ crosstalk and capture genotype-specific drug responses underscores its potential to guide precision oncology. Nevertheless, a key limitation is the lack of validation across a larger cohort of patients harboring NF1 mutations.
Despite the strengths of this study, several limitations should be acknowledged. First, the work focused on iPSCs derived from a single breast cancer patient harboring an NF1 mutation. This patient-specific design inevitably limits broad generalizability across diverse genetic backgrounds. However, the primary aim of the study was to establish the technical feasibility and translational relevance of the NOCS platform for integrating patient-derived intestine, liver, kidney, and tumor compartments into a unified PK/PD evaluation system. Importantly, even within this single-patient setting, the platform consistently captured functionally meaningful differences in drug metabolism and therapeutic response, demonstrating its capacity to resolve patient-specific pharmacological heterogeneity. To further support the robustness and applicability of the NOCS system beyond this initial case, additional validation studies using other targeted agents are underway, and preliminary results already confirm reproducible multi-organ PK/PD dynamics. Larger-scale studies that incorporate broader patient cohorts and diverse tumor genotypes are in progress and will be essential for fully establishing the clinical generalizability of this approach. Second, the current NOCS configuration, composed of intestine, liver, and kidney organoids, lacks vascular and immune components, and thus cannot fully reproduce inter-organ or immune–tissue interactions. This limitation likely contributes to the moderate transcriptomic similarity observed between organoids and their adult tissue counterparts (intestine, 0.76–0.77; liver, 0.78–0.8; kidney, 0.70–0.72, data not shown), reflecting the absence of minor cell populations such as endothelial and immune cells and the relatively immature state typical of iPSC-derived organoids. Nonetheless, when analysis was limited to drug-response–related genes, similarity increased to 0.78–0.91, indicating that these organoids are suitable for evaluating patient-specific PK/PD and therapeutic responses (Fig. 7a, b). Nonetheless, the NOCS platform provides a versatile framework for evaluating patient-specific drug efficacy and PK/PD responses in a human-relevant context.
In conclusion, we established a NOCS, a patient-specific, multi-organ in-vitro PK/PD platform integrating tailored organoids for precision medicine. We proposed a personalized therapeutic strategy combining exon skipping with targeted inhibition and aligned with the patient’s genetic and physiological profiles. As a scalable and translationally relevant system, NOCS provides a powerful tool to guide individualized drug selection and dosing based on genotype-specific responses.
Materials and Methods
Ethical approval and consent to participate
The collection of human biospecimens and the acquisition of data derived from them were conducted with ethical approval from the Institutional Review Board (IRB, P01-202004-31-005) of the Korea Research Institute of Bioscience and Biotechnology.
All animal housing and experiments conducted were in accordance with the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Institutional Animal Care and Use Committee Guidelines (KRIBB-AEC-24058).
Development of the NOCS
NOCS, the multi-organ integrated MPS, was designed to mimic in vivo environments by integrating multiple organ components, circulating fluidics, nutrient uptake, and waste elimination. To construct each organ module, a multi-insert dish with a meshed bottom (SPL Life Sciences, #911606, Pocheon-si, Gyeonggi-do, Republic of Korea) and meshed sides (SPL Life Sciences, #911605) was used. To maintain a controlled microenvironment, a piezoelectric diaphragm pump (Takasago International Corporation, #SDMP306D, Yokohama, Kanagawa, Japan) and a weight load cell sensor (Cozy Electronics, #ZEL6J1-3kg, Ansan-si, Gyeonggi-do, Republic of Korea) precisely regulated the circulation of a common culture medium consisting of Advanced DMEM/F12 (Gibco, Thermo Fisher Scientific, #12634010, Waltham, MA, USA), N-2 supplement (Gibco, Thermo Fisher Scientific), B-27 supplement (Gibco, Thermo Fisher Scientific), 50 μg/ml Gentamycin (Gibco, Thermo Fisher Scientific, #15750060), GlutaMAX supplement (Gibco, Thermo Fisher Scientific, #35050061), and 1% penicillin/streptomycin (Gibco, Thermo Fisher Scientific, #15140122) across the system. The fluid velocity was maintained at 5 mL/min, ensuring efficient nutrient exchange and physiological shear stress, which are essential for preserving organoid function. Additionally, to prevent media stagnation and facilitate nutrient replenishment, two auxiliary pumps were programmed to activate for 2 min every hour at 2 mL/min, mimicking periodic fluid exchange observed in vivo. To further enhance physiological relevance, a brushless DC gear motor (BLDC, Xiangli, #BL3640A-24V-06P + RB35, Shanghai, China) was integrated into the main body of the system to induce orbital shaking. Detailed specifications for the equipment are provided in Supplementary Table 1.
Collection of BC biospecimens
Biospecimens were obtained from 14 women (mean age 60.2 years) who underwent breast cancer surgery and were provided by the Biobank of Chungnam University Hospital. Tissues from two patients were excluded due to yeast contamination. Of the remaining 12 patients, 10 were diagnosed with invasive ductal carcinoma, one with a malignant spindle cell tumor of the sarcoma type, and one with pleomorphic lobular carcinoma in situ. The collection of human biospecimens and the acquisition of data derived from them were conducted with ethical approval from the Institutional Review Board of the Korea Research Institute of Bioscience and Biotechnology. Informed consent was obtained from all biospecimen donors. The donor’s information of biospecimen is reported in Supplementary Tables 2.
Isolation and culture of primary cells from BC biospecimens
Tumor and adjacent normal tissue samples from breast mastectomy patients were washed with DPBS (Corning Inc., #21-031-CM) and minced into smaller pieces. For enzymatic digestion, 20–50 mg of tissue was incubated with 3 mL of 0.05% trypsin-EDTA (Gibco, #15400054) at 37 °C for 30 minutes. After digestion, 20 mL of DMEM with 5% FBS (Gibco, #16000044) was added, and the suspension was filtered through a 100 μm strainer (Corning Inc. #CLS431752). The flow-through was centrifuged, and the pellet was resuspended in MEGM (Lonza, #CC-3150) with 1 μM forskolin (Sigma-Aldrich), 1 μM Repsox (Sigma-Aldrich), 1 μM Y-27632 (Tocris Bioscience), 10 ng/mL R-SPO1 (PeproTech), and 1% P/S. The cells were seeded into T25 flasks, and after 24 h, the medium was changed every two days. Primary normal and tumor cells were maintained for up to five passages and verified by flow cytometry using EpCAM and CD49f antibodies (BD FACSVerse™).
Materials for Cell Culture
The 293 T cells were maintained in Dulbecco’s Modified Eagle’s Medium, low glucose (DMEM-low glucose, Gibco, Thermo Fisher Scientific, #11885084) supplemented with 10% FBS and 1% P/S in a humidified incubator at 37 °C with 5% CO₂. For the generation of the organoid model, DMEM/Ham’s F-12 (DMEM/F-12, #11320033), Advanced DMEM/F-12, Advanced Roswell Park Memorial Institute (RPMI) 1640 (#12633012), L-glutamine (#A2916801), 1 M HEPES (#15630080), B-27 supplement, and N-2 supplement were purchased from Thermo Fisher Scientific. RPMI 1640 medium (#10-040-LB) was purchased from Corning Inc.
Chemicals
All small molecules and selective inhibitors used in this study were purchased from Sigma-Aldrich or Selleckchem (Houston, TX, USA). Acyclovir and chlorothiazide were dissolved in dimethyl sulfoxide (DMSO, Sigma-Aldrich, #472301), while caffeine and acetaminophen were dissolved in water to the indicated stock concentrations. All selective inhibitors were dissolved in DMSO to the indicated stock concentrations. Detailed information on the chemicals, including stock and final working concentrations, is provided in Supplementary Table 3.
Establishment and maintenance of normal and BC Patient 1-derived hiPSCs
Patient 1 and healthy (normal, CRL 2097, ATCC) fibroblast-derived iPSCs were reprogrammed using the CytoTune-iPS 2.0 Sendai Reprogramming Kit (Invitrogen, Thermo Fisher Scientific). Fibroblasts (1 × 10⁵) were seeded in 6-well plates for 24 h, then transduced with Sendai virus carrying reprogramming genes (OCT4, SOX2, KLF4, L-MYC) according to the manufacturer’s recommendations. After 5 days, detached fibroblasts were re-seeded onto mitomycin C (AG scientific, San Diego, CA, USA, #M-1108)-treated mouse embryonic fibroblasts and cultured for 14 days in reprogramming medium [DMEM/F12, 1 mM MEM Non-Essential Amino Acids Solution (NEAAs, Gibco, Thermo Fisher Scientific, #11140035), 1% P/S, 0.1 mM 2-Mercaptoethanol (Gibco, Thermo Fisher Scientific, #21985023), 20% KnockOut™ Serum Replacement (KSR, Gibco, Thermo Fisher Scientific, #10828028), and 20 ng/ml basic fibroblast growth factor (bFGF, R&D Systems, #233-FB, Minneapolis, MN, USA)]. Single colonies were picked, expanded, and dissociated into clumps using 1 mg/mL collagenase type IV (Gibco, Thermo Fisher Scientific, #17104019).
Identification of the NF1 mutant allele in BC Patient 1-derived hiPSCs
To validate the genetic background of the NF1 mutation in BC patient-derived hiPSCs compared to BC tissues, genomic DNA was extracted from normal and patient-derived hiPSCs using the DNeasy Blood & Tissue Kit (QIAGEN, #69504, Hilden, Germany) according to the manufacturer’s instructions. The exon 2 region of NF1 was amplified by PCR, and the PCR products were cloned into a TA cloning vector. After TA cloning, the NF1 mutation in patient-derived hiPSCs was verified by sequencing, and the results were compared to the WES analysis of the patient tissue. WES analysis was conducted by Theragen Bio (Suwon, Republic of Korea) using standard next-generation sequencing technology.
Generation of small intestine organoid (IO) from hiPSCs and isolation of ISCs
The differentiation of human normal and BC patient 1-derived iPSCs into hIOs, and the isolation of proliferative ISCs, were performed as previously described.27 Briefly, hiPSCs were differentiated into DE by treatment with 100 ng/mL Activin A (Novus Biologicals) for 3 days in RPMI 1640 medium with increasing concentrations of fetal bovine serum (FBS). The DE cells were then cultured in RPMI 1640 medium with 2% FBS, 500 ng/mL FGF4 (PeproTech), and 500 ng/mL WNT3A (R&D Systems) to form mid-hindgut cells. These cells formed 3D spheres, which were embedded in Matrigel (Corning Inc.) and cultured in hIO medium [Advanced DMEM/F-12, 2% B-27, 100 ng/mL EGF, and 100 ng/mL Noggin].
For ISC isolation, hIOs were removed from Matrigel and gently pipetted to eliminate residual fragments. Organoids were digested in 1 mL of 0.25% trypsin-EDTA for 5 minutes at 37 °C and dissociated into small clumps. Digestion was stopped with 10 mL of ISC basal medium [Advanced DMEM/F-12 with 2mM L-glutamine, 15 mM HEPES, and 1% P/S], followed by centrifugation at 800 × g for 3 minutes. The pellet was resuspended in ISC growth medium containing 1 μM Jagged-1 (Anaspec) and 10 μM Y27632 (Tocris Bioscience), along with 2% B-27, 10 nM [Leu15]-Gastrin I, 100 ng/mL WNT3A, EGF, and Noggin, 500ng/mL R-Spondin 1, 500 nM A83-01 (Tocris Bioscience), 10 μM SB202190, 2.5 μM PGE2, 1mM N-acetyl L-cysteine, and 10 mM nicotinamide (Sigma-Aldrich), prepared in ISC basal medium. Medium was refreshed every other day, and cells were passaged at a 1:3 ratio upon reaching 80–90% confluency. Detailed information on the reagents used for hIOs generation is provided in Supplementary Table 4.
Generation of hIECs model using ALI culture
To generate the hIECs model, ISCs were seeded onto 12-well transwell plates to establish polarized cultures. Transwell membranes were pre-coated with 1% Matrigel in cold ISC basal medium and incubated for 1 hour at 37 °C. ISCs were washed with PBS, dissociated using 0.25% trypsin-EDTA for 8 minutes at 37 °C, and centrifuged. A total of 3.5 × 10⁵ cells were seeded into each insert with ISC growth medium. After reaching confluency, the apical medium was removed to initiate ALI conditions, and the basal medium was replaced every 2 days. ALI cultures were maintained for 6–10 days to promote epithelial differentiation. Detailed information on the reagents used for liver organoid generation is provided in Supplementary Table 4.
Generation of hLOs from hiPSCs
Differentiation of human normal and BC patient 1-derived iPSCs into hepatic-like organoids (hLOs) was performed as previously described.28 Briefly, iPSCs were treated with 100 ng/mL Activin A (Thermo Fisher Scientific) in RPMI 1640 with 2% B-27 for 6 days to induce DE. For HE induction, DE cells were cultured for 4 days under hypoxic conditions in RPMI 1640 with 2% B-27 minus insulin (Gibco, Thermo Fisher Scientific), 25 ng/mL BMP4, and 10 ng/mL bFGF (PeproTech). Detached HE cells were embedded in Matrigel and cultured in hepatic medium (HM) composed of advanced DMEM/F12 with 1% P/S, 1 mM GlutaMax, 1 mM HEPES, 1% N-2, 2% B-27 without vitamin A, 1% ITS, 50 ng/mL EGF, 25 ng/mL HGF, 10 ng/mL bFGF, 10 ng/mL Oncostatin M (R&D Systems), 5 μM A83-01, 10 μM forskolin (Sigma-Aldrich), 1mM N-acetyl L-cysteine, 10 nM Leu15-gastrin I, 10 mM nicotinamide, and 100 nM dexamethasone (Sigma-Aldrich). Organoids were expanded in Matrigel domes with HM refreshed every 2–3 days and passaged weekly at a 1:4–1:10 ratio.
For differentiation, one day post-passaging, HM was replaced with expansion medium (EM), composed of advanced DMEM/F12 with 1% P/S, 1 mM GlutaMax, 1 mM HEPES, 1% N-2, 2% B-27 without vitamin A, 50 ng/mL EGF, 25 ng/mL HGF, 100 ng/mL FGF10, 1mg/mL R-spondin 1, 25 ng/mL BMP7 (PeproTech), 5 μM A83-01, 10 μM forskolin, 1mM N-acetyl L-cysteine, 10 mM nicotinamide, and 10 nM Leu15-gastrin I. After 3 days in EM, the medium was replaced with differentiation medium (DM) containing advanced DMEM/F12, 1% P/S, 1 mM GlutaMax, 1 mM HEPES, 1% N-2, 2% B-27 with vitamin A, 25 ng/mL HGF, 0.5 μM A83-01, 100 ng/mL FGF19 (PeproTech), 100 ng/mL BMP7, 1mM N-acetyl L-cysteine, 10 nM Leu15-gastrin I, and 3 μM dexamethasone. Organoids were maintained in DM for 6 days with medium changes every 2 days. Reagent details are provided in Supplementary Table 4.
Generation of KOs from iPSCs
To generate KOs, human normal and BC patient 1-derived iPSCs were seeded on Matrigel-coated flasks and cultured in mTeSR1 medium (STEMCELL Technologies, #85850). When cells reached ~30–40% confluency (day 1–2), primitive streak (PS) induction was initiated using Advanced RPMI 1640 (Gibco, Thermo Fisher Scientific) supplemented with antibiotic–antimycotic (Gibco, #15240062), NEAAs, GlutaMax, B-27, and 8 μM CHIR99021 (Tocris Bioscience) for 4 days. For BC iPSCs, 1 μM Y-27632 was also included.
From days 5–7, cells were treated with 100 ng/mL FGF9 (PeproTech), 1 μg/mL heparin (Sigma-Aldrich), and 0.1 μM RA (Sigma-Aldrich, days 5–6 only) to induce intermediate mesoderm (IM). IM cells were dissociated using Accutase (STEMCELL Technologies, #07920), and 1 × 10⁵ cells/well were seeded into ultra-low attachment 96-well plates (Corning, #CLS7007), centrifuged (800 × g, 2 min), and cultured with FGF9, heparin, and 5 μM Y-27632 for 1 day. Aggregates (5–6/well) were transferred to collagen-coated transwell inserts (Corning, #3491), treated with 5 μM CHIR99021 for 1 h, then cultured in differentiation medium containing 100 ng/mL FGF9 and 100 ng/mL BMP7 (PeproTech) for 10–14 days with daily medium changes, avoiding overflow onto the apical surface.47
Generation of hKORTECs Model from hKTOs
KO-derived iPSCs were dissociated using Gentle Cell Dissociation Reagent (STEMCELL Technologies, #100-0485) for 10 min at 37 °C. Tubular segments and single cells were embedded in Matrigel and cultured in DMEM/F12 supplemented with 1.5% B-27, 1 μM N-acetyl L-cysteine, 5 μM A83-01, 100ng/mL R-spondin 1, 50 ng/mL FGF10 (PeproTech), 50 ng/mL EGF, 50 ng/mL BMP7, and 10 μM Y-27632. Medium was changed every 2 days, and tubule-like organoids appeared within 7–14 days. Organoids were passaged weekly.
For hKORTECs, ~250,000 cells from tubule organoids were seeded onto SPLInsert™ Hanging 24-well plates (SPL Life Science, #36124) and cultured in kidney tubule organoid medium. Upon confluency, the apical medium was removed to initiate ALI conditions while preventing overflow onto the membrane. Reagent details are provided in Supplementary Table 4.
Generation of BC patient 1-derived tumor spheroid model
Primary multicellular cells were isolated from the tumor and matched normal breast tissues obtained from patients with breast cancer. Cells were cultured in Mammary Epithelial Growth Medium (MEGM; Lonza, Basel, Switzerland, # CC-3150) supplemented with 1 μM forskolin, 1 μM RepSox, 1 μM Y-27632, 10 ng/mL recombinant human R-Spondin1, and 2% Matrigel (Corning Inc., #354234). Cell suspensions were placed in 15-mL SFU tubes (SPL Life Science, #911604) and subjected to rotation culture at 37 °C in 5% CO₂ incubator for 48 hours to induce multicellular aggregate formation. Following aggregation, cell clusters were embedded in a 3D extracellular matrix composed of type I collagen (Gibco, Thermo Fisher Scientific, #A10644-01) and Matrigel mixed at a 7:3 volume ratio. The embedded aggregates were transferred into ultra-low attachment 6-well plates (Corning Inc., # CLS3471) and maintained in suspension culture with MEGM for an additional 3–5 days at 37 °C under 5% CO₂. The resulting spheroids were used for subsequent functional and histological analyses.
In vitro three germ layer differentiation
To assess the differentiation potential, BC patient-derived iPSCs were detached using 1 mg/mL Collagenase Type IV and 1 mg/mL Dispase II (Gibco, Thermo Fisher Scientific, #17105041). The cell colonies were transferred to Petri dishes (SPL Life Science) and cultured in embryoid body (EB) differentiation medium containing DMEM/F-12 and 10% KSR for 5 days. Formed EBs were then re-plated onto 5% Matrigel-coated 4-well plates (Thermo Fisher Scientific, #167063) and cultured for an additional 10 days.
Cells were subsequently analyzed for germ layer-specific differentiation by immunostaining, the proportion of positive cells was quantified using ImageJ (version 1.53e), and RT-qPCR using ectoderm, mesoderm, and endoderm markers. Antibody and primer details are provided in Supplementary Tables 5 and 7.
Immunocytochemical analysis
hiPSCs and organoids were fixed with 4% paraformaldehyde at 4 °C overnight, then washed with PBS (Welgene). Samples were incubated with primary antibodies at 4 °C overnight, followed by Alexa Fluor®-conjugated secondary antibodies for 1 h at 25 °C in the dark. Organoids were washed with PBS containing 0.05% Tween-20 between steps. Nuclei were stained with Hoechst 33342 (Invitrogen) and slides mounted using Fluoromount™ Aqueous Mounting Medium (Sigma-Aldrich, #F4680). Images were captured using a confocal microscope (LSM 800, ZEISS) or EVOS FL Auto microscope (Thermo Fisher Scientific) and analyzed with ImageJ (version 1.53e).
Alkaline phosphatase (AP) staining and Immunostaining
hiPSCs were stained using the AP Detection Kit (Sigma-Aldrich) following the manufacturer’s instructions. For immunostaining, cells were washed with PBS and fixed with 4% paraformaldehyde for 15 min at 25 °C, followed by permeabilization with PBS containing 0.1% Triton X-100 for 15 min. After blocking with 4% BSA (Bovogen Biologicals, Keilor East, VIC, Australia, #BSAS-AU) for 1 h at 25 °C, cells were incubated with primary antibodies overnight at 4 °C. The next day, samples were washed with PBS containing 0.05% Tween-20 and incubated with fluorescence-conjugated secondary antibodies for 1 h at 25 °C. Nuclei were stained with DAPI (BD Biosciences, San Jose, CA, USA, #564907) and visualized using a fluorescence microscope (IX71, Olympus Corporation, Tokyo, Japan). Reagents and kits are listed in Supplementary Table 6.
RNA preparation, cDNA synthesis, and quantitative real-time PCR
Total RNA from hiPSCs and differentiated cells was extracted using the RNeasy Mini Kit (QIAGEN, #74106). cDNA was synthesized using the SuperScript™ IV First-Strand Synthesis System (Invitrogen, Thermo Fisher Scientific, #18091200). qPCR was performed using SYBR™ Green PCR Master Mix (Applied Biosystems, Thermo Fisher Scientific, #43-091-55) on a 7500 Fast Real-Time PCR System (Applied Biosystems). Gene expression levels were analyzed using the ΔΔCt method. Primer sequences are listed in Supplementary Table 7.
Short tandem repeat (STR) analysis and Karyotype analysis
Genomic DNA was extracted from breast cancer fibroblasts and hiPSCs using the DNeasy Blood & Tissue Kit. STR analysis was performed by HumanPass Inc. (Seoul, Republic of Korea), and G-banding karyotype analysis of cultured hiPSCs was conducted by GenDix Inc. (Seoul, Republic of Korea).
TEER Measurement
To assess barrier integrity and functionality of the hIEC and KORTEC models, TEER was measured using an epithelial volt/ohm meter (EVOM2, World Precision Instruments, Sarasota, FL, USA) according to the manufacturer’s instructions. TEER was assessed both to evaluate the functional properties of the models and to determine the impact of drug treatment on epithelial barrier toxicity.
Dextran uptake assay
To evaluate the paracellular permeability of the hIECs model, FITC-conjugated dextrans (4 kDa, Chondrex Inc., #4013; 40 kDa, Chondrex Inc., #4009) were used. The hIEC models were washed with PBS and treated with HBSS (Thermo Fisher Scientific, #14025092) containing 6.25 μM of each FITC-dextran in the apical compartment. After a 30-minute incubation at 37 °C in a 5% CO2 incubator, the basal solution was collected for fluorescence analysis using a multi-mode microplate reader (SpectraMax i3x, Molecular Devices).
For the hKORTECs model, FITC-labeled dextran (3 kDa, Invitrogen, Thermo Fisher Scientific, #D3306) was added to the apical compartment at a final concentration of 0.1 mg/ml. At 30, 60, 120, 240, and 360 minutes, 100 μl of medium was collected from both apical and basolateral compartments. Fluorescence intensity was measured using a multi-mode microplate reader.
Measurement of ALB, AST, and ALT
To quantify ALB, ALT, and AST, organoid culture supernatants were collected after 24 hours, and the concentrations of secreted ALB, ALT, and AST were measured using Human Albumin (Bethyl Laboratories, Montgomery, TX, USA), ALT (Abcam, Cambridge, UK), and AST (Abcam) ELISA kits according to the manufacturer’s instructions. Absorbance was measured using a microplate reader. A list of the ELISA kits used in this study is provided in Supplementary Table 6.
Uptake of CLF
To measure CLF polarity, organoids were removed from the Matrigel by washing with PBS and incubated with culture medium supplemented with 10 μg/ml CLF (Corning Inc.) and 1 μg/ml Hoechst 33342 for 30 minutes at 37 °C in a 5% CO2 incubator. After incubation, the dye was removed, and the DhLOs were gently washed twice with cold PBS containing calcium and magnesium (Gibco, Thermo Fisher Scientific, #10010023). Culture medium was then added, and the DhLOs were photographed under a confocal microscope (LSM 800, ZEISS).
Measurement of cytochrome P450 (CYP) enzyme and CYP450 reductase (CPR) activity assay
CPR activity in the hIECs was evaluated using the Cytochrome P450 Reductase Assay Kit (Abcam, #ab204704) according to the manufacturer’s instructions. Standard and reaction wells were prepared with appropriate volumes of samples, inhibitors, or controls, followed by the addition of reaction mix and 20 mM G6P solution. After 30 minutes of incubation at 25 °C, CPR activity was measured using a colorimetric microplate reader.
To assess CYP enzyme activity in the DhLOs model, specific CYP enzymes were induced with inducers: 20 μM rifampicin for CYP2C8, CYP2C9, CYP2C19, and CYP3A4, and 100 μM omeprazole for CYP1A2. CYP family activity was analyzed using a P450-Glo assay kit (Promega Corporation) and measured with a GloMax Navigator Microplate Luminometer (Promega Corporation). The results were normalized to total cell count using a TC20 Automated Cell Counter (Bio-Rad Laboratories) and expressed as enzyme activity per 1 × 10⁶ cells.
Assessment of Albumin Uptake and MRP2/4 Transport Function in the hKORTECs
For albumin uptake assessment, FITC-labeled albumin (Invitrogen, Thermo Fisher Scientific, #A23015) was added to the apical compartment at a final concentration of 0.2 mg/ml. After 6 hours, samples were collected from both the apical and basolateral compartments, and the fluorescence was measured using a multi-mode microplate reader.
For the MRP2/4 transport assay, 10 μM CDFDA (Sigma-Aldrich, #21879) ± 20 μM MK-571 (Sigma-Aldrich, #M7571) in serum-free medium was added to the basolateral compartment, and serum-free medium ± 20 μM MK-571 was added to the apical compartment. Cells were incubated at 37 °C for 2 hours, and samples were collected from the basolateral compartment and the fluorescence was measured using a multi-mode microplate reader.
Functional analysis of drug transporters in hKORTECs
For nephrotoxicity and drug transporter activity assays, hKORTECs were exposed to 20 μM cisplatin (Sigma-Aldrich) with/without 1 mM cimetidine (Sigma-Aldrich) for 24 hours and the apoptosis profile was examined in the hKORTECs. For the apoptosis assay, cells were harvested and incubated with 100 μl of Muse Annexin V & Dead Cell Reagent (Millipore, Bedford, MA, USA) for 20 minutes at room temperature. Apoptosis was determined using the Muse Cell Analyzer (Millipore), and the statistics were presented as percentages of the cells that were alive, apoptotic, or dead.
Tissue Processing
Human breast tumor and normal tissues, mouse xenografts, and major organs were fixed in 10% formalin (Sigma-Aldrich, #HT501128) for 4–6 hours at 25 °C. The tissues were then placed in cassettes, thoroughly washed with water, and stored in 70% ethanol at 4 °C overnight until further processing. Tissue processing was performed according to a standard protocol, which included sequential dehydration steps: 80% ethanol for 1 hour at 25 °C, followed by 90% ethanol for 1 hour, 95% ethanol for 1 hour, and two rounds of 100% ethanol for 1 hour each. This was followed by three xylene treatments, each for 1 hour at 25 °C. Finally, tissues were infiltrated with paraffin wax overnight at 65 °C. Paraffin-embedded tissue samples were sectioned into 5–7 μm thick slices using a rotary microtome (Epredia, Thermo Fisher Scientific, Epredia™ HM 325) and mounted onto silane-coated histological slides (MUTO CHEMICALS CO., #5116-20 F, Tokyo, Japan) to enhance tissue adhesion.
Immunohistochemical (IHC) Analysis
Paraffin-embedded tissue slides were deparaffinized in xylene and rehydrated through graded ethanol. Endogenous peroxidase activity was blocked with 0.3% H₂O₂, followed by permeabilization with PBST (1× PBS containing 0.1% Triton X-100). Antigen retrieval was performed in 10 mM sodium citrate buffer with 0.05% Tween 20 using a microwave. After blocking with 2% normal horse serum, slides were incubated with primary antibodies overnight at 4 °C. Following PBST washes, biotinylated secondary antibodies were applied, followed by ABC reagent (Vector Laboratories, Inc., Newark, CA, USA, #PK-400) and DAB substrate (Vector Laboratories, Inc., #SK-4105). Nuclei were counterstained with DAPI. Images were captured using an inverted microscope (IX71, Olympus) and analyzed with cellSense software. Antibody details are listed in Supplementary Table 5.
Flow Cytometry (FACS) analysis
Primary cells isolated from the tumor and adjacent normal tissues were washed with ice-cold PBS containing 2% FBS and counted. A total of 2 × 10⁵ cells were stained with fluorochrome-conjugated antibodies (listed in Supplementary Table 5) in PBS with 2% FBS for 30 min at 4 °C in the dark, following the manufacturer’s protocol. Secondary antibody-only samples were used as negative controls. After staining, cells were washed and analyzed using a BD FACSVerse™ Cell Analyzer (BD Biosciences). EpCAM and CD49f were used to identify epithelial and stromal populations, respectively, while CD44 and CD24 were used to detect breast cancer stem cells.
Western Blot Analysis
Tissue samples were homogenized in RIPA buffer [50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 0.5 mM EDTA, 1% NP-40, 0.1% SDS, 1 mM PMSF, and protease inhibitor cocktail (Roche, Basel, Switzerland)], and cells were directly lysed in RIPA buffer on ice for 15 min. The lysates were then clarified by centrifugation at 15,000 x g for 10 minutes at 4°C. Lysates were separated on 10–15% SDS-PAGE gels and transferred onto PVDF membranes (Merck Millipore, Burlington, MA, USA). Membranes were blocked in PBST containing 5% skim milk for 1 hour at 25 °C, incubated overnight at 4 °C with primary antibodies, and then with secondary antibodies in PBST with 0.1% skim milk for 1 hour at 25 °C. Protein bands were visualized using a chemiluminescence kit (Merck Millipore, #WBKLS0500) and quantified using Image J software. Antibodies used are listed in Supplementary Table 5.
Ras Activation Assay
Ras activity in tissues and isolated primary cells was assessed using the Active Ras Detection Kit (Cell Signaling Technology, Danvers, MA, USA) according to the manufacturer’s instructions. Briefly, homogenized tissues and cells were incubated in lysis buffer on ice, and centrifuged at 15,000 x g for 10 minutes at 4 °C to collect the supernatant. Cell lysates (500 µg of total protein) were pre-incubated with GTPγS or GDP for 15 minutes at 30 °C. The reaction was terminated by adding MgCl₂ to a final concentration of 60 mM. Lysates were incubated with 80 µg of GST-Raf-RBD pre-bound to glutathione-Sepharose in a spin column for 1 hour at 4 °C. After washing, the resin was incubated in 2 × reducing sample buffer containing 200 mM DTT for 2 minutes at 25 °C, followed by western blot analysis.
AON design for NF1 exon skipping and evaluation of exon skipping efficacy
For AON design, exon 2 of NF1 was selected as the target, and 13 different AON sequences were designed using Human Splicing Finder and Examine the Secondary Structure of mRNA software. To prevent the degradation of the newly generated transcript by nonsense-mediated decay, all AONs were chemically modified with 2′-O-methyl (2′OMe)-PS and synthesized by Bioneer (Daejeon, Republic of Korea).
For the exon skipping efficacy test, BC-derived primary cells (3 × 10⁵ cells per well) were seeded in a 6-well plate, and the most promising AON was selected by transfecting 300 nM AON using Lipofectamine™ RNAiMAX reagent (Invitrogen, Thermo Fisher Scientific, #13778075) according to the manufacturer’s instructions. At 48 hours post-transfection, cDNA was synthesized from total RNA extracted from AON-treated BC-derived primary cells. Exon skipping was assessed by conventional RT-PCR and visualized on a 3% agarose gel. Exon skipping was confirmed by Sanger sequencing, and the splicing band was quantified using ImageJ software. The sequences of AON and NF1 primers used in this study are provided in Supplementary Tables 8–10.
Lentiviral vector-mediated double-target U7 snRNA-AON constructs
The modified U7 snRNA-specific plasmids were generated using the Sf-U7snRNA-ESE-AS-NsiI vector purchased from Addgene (#190694, Watertown, MA, USA). The U7 snRNA plasmid contained a dual AON targeting exon 2, specifically the acceptor splice site and the internal ESE. The complementary sequence to the pre-mRNA was selected to promote NF1 exon skipping. AON-U7smOPT sequences were cloned into the Sf-U7snRNA vector using NsiI and SalI restriction sites by Cosmogenetech (Seoul, Republic of Korea). The AON sequences are provided in Supplementary Table 11.
Lentiviral vector production, titration, and transduction
For lentiviral vector production, 293 T cells were transfected with the packaging plasmid psPAX2, the viral envelope plasmid pMD2.G, and U7snRNA constructs using the Transporter™ 5 transfection reagent (Polysciences, Inc., Warrington, PA, USA, #26008) according to the manufacturer’s instructions. At 72 hours post-transfection, the medium containing lentiviral particles was harvested, filtered through a 0.45 µm syringe filter (Merck-Millipore), and concentrated by ultracentrifugation at 100,000 × g for 2 hours at 10 °C (Himac CP100WX, HITACHI, Tokyo, Japan). Lentivirus titers were measured using a qPCR lentivirus titer kit (Abm, BC, Canada) following the manufacturer’s protocol, and determined by transducing 293 T cells with serial dilutions in a 24-well plate. BC-derived primary cells (2 × 10⁶ cells per 10 cm dish) were transduced with infectious lentiviral particles at a multiplicity of infection (MOI) of 10 in the presence of 8 μg/ml polybrene (Santa Cruz, CA, USA, #sc-134220) for 48 hours.
Generation of Isogenic iPSCs with lentivirus-U7-AON2/12
Patient-derived iPSCs were transduced with lentiviral vectors expressing U7-AON2/12 or control (LV-zsG) at an MOI of 20 in the presence of 8 µg/mL polybrene. Single-cell clones were established and screened for successful NF1 exon 2 skipping to select stable isogenic iPSC lines. Integration of the lentiviral construct was confirmed by PCR amplification of the U7 promoter and WPRE region after 2 and 6 months of culture. Pluripotency of the established iPSC lines was confirmed by standard assays for iPSC markers and differentiation potential. The primers used in this study are provided in Supplementary Table 10.
Cell viability assay
Cell viability was assessed 72 hour after chemical treatment in the multi-organ integrated MPS. Cells were incubated with the EZ-CYTOX reagent (DoGenBio, Seoul, Republic of Korea, #EZ-1000) at 37 °C for 1 hour in a CO₂ incubator following the manufacturer’s protocol. The absorbance at 450 nm was then measured using a microplate reader.
RNA sequencing analysis
For total RNA sequencing analysis, the TrueSeq RNA Sample Preparation Kit V2 was used for RNA purification and library preparation with total RNA as input material. Sequencing was performed on the Illumina NextSeq 1000 platform (Illumina, San Diego, CA, USA) with a paired-end read length of 2 × 100 base pairs. Raw sequencing reads were trimmed using Cutadapt version 1.18 (https://cutadapt.readthedocs.io/en/stable/), with the parameters ‘-a AGATCGGAAGAGCACACGTCTGAACTCCAGTCAC -AAGATCGGAAGAGCGTCGTGTAGGGAAAGAGTGTA -m 50 -O 5’. Low-quality reads (Phred score below 20) were filtered out using Sickle v1.33 (https://github.com/najoshi/sickle), retaining sequences of at least 50 bp in length. The quality of the resulting paired-end reads was evaluated using FastQC version 0.11.4. Duplicate reads were assessed using the FastQC tool. Further quality filtering, including removal of poly-N and low-quality reads, was conducted using NGSQCToolkit version 2.3.3 (https://github.com/mjain-lab/NGSQCToolkit). Cleaned reads were aligned to the human reference genome GRCh38.97 (Accession number: GCA_000001405.27) using HISAT2 version 2.1.0 (https://daehwankimlab.github.io/hisat2/). Transcript abundance was calculated as fragments per kilobase of transcript per million mapped reads (FPKM) using StringTie version 2.2.1 (https://github.com/gpertea/stringtie), which also provided normalized expression values.
In vitro PK profiling in a NOCS
The common culture medium, consisting of Advanced DMEM/F12, 1 x N-2/B-27 supplement, 50 μg/ml Gentamycin (Gibco, Thermo Fisher Scientific, #15710064), 1x GlutaMAX, and 1% P/S, was added to the outer dish connected to the multi-insert dish containing the prepared models in the multi-organ integrated MPS. The MPS device was set with a flow rate of 5 ml per minute and a shaking speed of 0.8 revolutions per second. The input rate of fresh medium and the output rate for elimination mimicking were activated for 2 minutes every hour at 2 mL/min, respectively.
For the PK studies of chemicals in the multi-organ integrated MPS system, both oral and intravenous drug administration routes were employed. All compounds were tested at a final concentration of 10 μM or with DMSO as a control. For oral drugs, the compound was applied to the apical side of the absorption model, and samples were collected at 30 minute intervals up to 6 hours, followed by additional samples at 12, 24, 48, and 72 hours. For intravenous drugs, the compound was applied to the metabolism model, and samples were collected at the same time points as those for the oral drug. The concentration of the compounds in all samples was analyzed using LC-MS/MS, and PK parameters were obtained using PK solver version 2.0 software.
In vitro PD profiling in a NOCS
For the PD studies in the multi-organ integrated MPS system, Paxalisib was applied to the apical side of the intestinal model at a final concentration of 10 µM under the same conditions as the PK studies for 72 hours.
After the analysis, the cytotoxic effects on the intestine, liver, and kidney models were evaluated by measuring cell viability using the EZ-CYTOX reagent, and TEER measurements were performed in the intestine and kidney models to assess barrier integrity and cytotoxicity. Additionally, the intestine, liver, and kidney models were analyzed for model-specific cell types and injury markers by RT-qPCR analysis.
For the cancer model, AON-induced exon skipping was performed prior to model establishment. To evaluate the anti-cancer efficacy of the combination treatment with Paxalisib and AON-mediated exon skipping, cancer models were treated with Paxalisib and AON. PI3K-mTOR and apoptosis signaling pathways were analyzed by western blotting, and cell viability was assessed using the EZ-CYTOX reagent. Antibody and primer information is listed in Supplementary Table 5 and 7.
In vivo PK profiling using a rodent model
Animal studies
Specific pathogen-free (SPF) male ICR mice (8 weeks old) were purchased from JA BIO Co. (Suwon, Kyonggi, Republic of Korea) and maintained in an SPF environment at 23 ± 2 °C with a 12-hour light/dark cycle and 50 ± 10% relative humidity. Food and water were provided ad libitum. Mice were allowed to acclimatize for one week before the study. Mice in the oral administration group were fasted overnight and fed 4 hours after dosing. A single dose of Bimaralisib, Gedatolisib, or Paxalisib was administered i.v. (5 mg/kg) or p.o. (20 mg/kg) in a solution of dimethylacetamide/Tween 80/20% v/v 2-hydroxypropyl β-cyclodextrin in deionized water (1/1/8 v/v). Blood samples (~50 µL) were collected at predetermined time points via the saphenous vein into Sarstedt’s Microvette® capillary tubes (Sarstedt, Nümbrecht, Germany), centrifuged at 12,000 x g for 3 minutes, and stored frozen at -20°C until analysis.
Plasma sample preparation
Plasma samples (15 µL each) were transferred to PCR tubes (Axygen, Union City, USA). Four volumes of acetonitrile containing carbamazepine as the internal standard (IS) were added, vortexed for 10 minutes (Multi-Tube Vortexer, VWR International, West Chester, PA, USA), and sonicated for 30 minutes at room temperature. After centrifugation at 12,000 x g for 10 minutes, the supernatant was analyzed using a 4000 QQQ LC-MS/MS system (Applied Biosystems, Concord, Canada) in positive MRM mode. Pharmacokinetic parameters were calculated by noncompartmental analysis of plasma concentration-time profiles using Kinetica™ 4.4.1 (Thermo Fisher Scientific, Inc., Woburn, MA).
LC-MS/MS analysis
The LC-MS/MS system consisted of an Agilent 1200 series HPLC system (Agilent Technologies, Wilmington, DE) and a 4000 QQQ LC-MS/MS system (Applied Biosystems, Foster City, CA) equipped with a Turbo VTM ion spray source operated in the positive ion mode. Sample separation was performed on an Atlantis dC18 column (50 × 2.1 mm i.d., 3 µm; Waters, Milford, MA, USA) with a SecurityGuard™ C18 guard column (2.0 × 4.0 mm i.d.; Phenomenex, Torrance, CA, USA) maintained at 30 °C. The sample injection volume was 5 µL, and the flow rate was set of 0.4 mL/min. Mobile phase consisted of HPLC water (A) and acetonitrile (B), each containing 0.1% formic acid. The TurboIonSpray interface was operated in the positive ion mode at 5500 V. The detection was conducted using multiple reaction monitoring of the transitions of m/z 252 > 190 for Bimiralisib, m/z 616 > 488 for Gedatolisib, m/z 383 > 353 for Paxalisib, and m/z 237 > 194 for carbamazepine (IS). The retention times of Bimaralisib, Gedatolisib, and Paxalisib, and the IS were 3.27, 3.09, 3.28, and 3.23 min, respectively. The scan dwell time was set at 0.1 sec for every channel. Acquisition and analysis of data were performed with AnalystTM software (version 1.6.3, Applied Biosystems, Foster City, CA).
In vivo Xenograft assay
Female BALB/c nude mice (5 weeks old, n = 5 per group) were purchased from the Central Laboratory Animal Inc. (Seoul, Republic of Korea). Mice were subcutaneously injected with patient-derived tumor primary cells (passage <4, 5 × 10⁶ cells/100 µL). When tumors reached ~50 mm³ (20 days post-injection), mice received intra-tumoral injection of lentivirus (LV-zsG or LV-U7-AON2/12, 10⁹ TU/mouse), followed by oral administration of Paxalisib (10 mg/kg) every 3 days for 10 doses in a vehicle containing 10% DMSO and 10% Cremophor-EL, starting one day after viral injection. Body weight and general health were monitored throughout the 47-day study. At the endpoint, mice were sacrificed, and tumors and major organs were excised for weight measurement and further histological analyses.
Statistical analysis
Statistical analyses were performed using GraphPad Prism software (version 7.05, La Jolla, CA, USA). Data are presented as mean ± SEM. For comparisons between two groups, statistical significance was assessed using a two-tailed unpaired Student’s t-test. For experiments involving multiple factors, statistical analysis was performed using one-way or two-way ANOVA, followed by Tukey’s post hoc test for multiple comparisons. A p-value of < 0.05 was considered statistically significant.
Data availability
All data supporting the findings of this study are publicly available. Processed datasets generated during this study have been deposited in [Cho-Rok Jung] under accession number [BioProject: PRJNA1392668] to facilitate independent validation and secondary analyses. Data access complies with relevant ethical and regulatory frameworks. All other data are included in the article and its supplementary information.
References
Rubin, J. B. & Gutmann, D. H. Neurofibromatosis type 1—a model for nervous system tumour formation?. Nat. Rev. Cancer 5, 557–564 (2005).
Ratner, N. & Miller, S. J. A RASopathy gene commonly mutated in cancer: the neurofibromatosis type 1 tumour suppressor. Nat. Rev. Cancer 15, 290–301 (2015).
Giraud, J.-S., Bieche, I., Pasmant, E. & Tlemsani, C. NF1 alterations in cancers: therapeutic implications in precision medicine. Expert Opin. Investig. Drugs 32, 941–957 (2023).
Moon, A. in Translational Research in Breast Cancer 81–101 (Springer, 2021).
Pearson, A. et al. Inactivating NF1 mutations are enriched in advanced breast cancer and contribute to endocrine therapy resistance. Clin. Cancer Res. 26, 608–622 (2020).
Frayling, I. M. et al. Breast cancer risk in neurofibromatosis type 1 is a function of the type of NF1 gene mutation: a new genotype-phenotype correlation. J. Med. Genet. 56, 209–219 (2019).
Na, B., Shah, S. R. & Vasudevan, H. N. Past, present, and future therapeutic strategies for NF-1-Associated tumors. Curr. Oncol. Rep. 26, 706–713 (2024).
Berry, D. et al. The NF1 tumor suppressor regulates PD-L1 and immune evasion in melanoma. Cell Rep. 44, 115365 (2025).
House, R. R. J. et al. NF1 deficiency drives metabolic reprogramming in ER+ breast cancer. Mol. Metab. 80, 101876 (2024).
Sait, S. F. et al. Hydroxychloroquine prevents resistance and potentiates the antitumor effect of SHP2 inhibition in NF1-associated malignant peripheral nerve sheath tumors. Proc. Natl. Acad. Sci. 122, e2407745121 (2025).
Aartsma-Rus, A. & Van Ommen, G.-J. B. Antisense-mediated exon skipping: a versatile tool with therapeutic and research applications. RNA 13, 1609–1624 (2007).
Pacelli, C., Rossi, A., Milella, M., Colombo, T. & Le Pera, L. RNA-Based Strategies for Cancer Therapy: In Silico Design and Evaluation of ASOs for Targeted Exon Skipping. Int. J. Mol. Sci. 24, 14862 (2023).
Jbara, A., Siegfried, Z. & Karni, R. Splice-switching as cancer therapy. Curr. Opin. Pharmacol. 59, 140–148 (2021).
Zhou, Y. et al. Posttranslational regulation of the exon skipping machinery controls aberrant splicing in leukemia. Cancer Discov. 10, 1388–1409 (2020).
Sharma, A., Sances, S., Workman, M. J. & Svendsen, C. N. Multi-lineage human iPSC-derived platforms for disease modeling and drug discovery. Cell Stem Cell 26, 309–329 (2020).
Li, Y., Tang, P., Cai, S., Peng, J. & Hua, G. Organoid based personalized medicine: from bench to bedside. Cell Regen. 9, 1–33 (2020).
van Berlo, D. et al. Stem cells, organoids, and organ-on-a-chip models for personalized in vitro drug testing. Curr. Opin. Toxicol. 28, 7–14 (2021).
Kim, J., Koo, B.-K. & Knoblich, J. A. Human organoids: model systems for human biology and medicine. Nat. Rev. Mol. Cell Biol. 21, 571–584 (2020).
Van Ness, K. P., Cesar, F., Yeung, C. K., Himmelfarb, J. & Kelly, E. J. Microphysiological systems in absorption, distribution, metabolism, and elimination sciences. Clin. Transl. Sci. 15, 9–42 (2022).
Noh, K. H. et al. A new experimental model to study human drug responses. Biofabrication 12, 045029 (2020).
Shroff, T. et al. Studying metabolism with multi-organ chips: new tools for disease modelling, pharmacokinetics and pharmacodynamics. Open Biology 12, 210333 (2022).
Palaninathan, V. et al. Multi-organ on a chip for personalized precision medicine. MRS Commun. 8, 652–667 (2018).
Ingber, D. E. Human organs-on-chips for disease modelling, drug development and personalized medicine. Nat. Rev. Genet. 23, 467–491 (2022).
Rodrigues, R., Duarte, D. & Vale, N. Drug repurposing in cancer therapy: influence of patient’s genetic background in breast cancer treatment. Int. J. Mol. Sci. 23, 4280 (2022).
Auwerx, C., Sadler, M. C., Reymond, A. & Kutalik, Z. From pharmacogenetics to pharmaco-omics: Milestones and future directions. Hum. Genet. Genom. Adv. 3, 100100 (2022).
Fraisse, J., Dinart, D., Tosi, D., Bellera, C. & Mollevi, C. Optimal biological dose: a systematic review in cancer phase I clinical trials. BMC Cancer 21, 1–10 (2021).
Kwon, O. et al. The development of a functional human small intestinal epithelium model for drug absorption. Sci. Adv. 7, eabh1586 (2021).
Mun, S. J. et al. Generation of expandable human pluripotent stem cell-derived hepatocyte-like liver organoids. J. Hepatol. 71, 970–985 (2019).
Picollet-D’hahan, N., Zuchowska, A., Lemeunier, I. & Le Gac, S. Multiorgan-on-a-chip: a systemic approach to model and decipher inter-organ communication. Trends Biotechnol. 39, 788–810 (2021).
Zhang, H.-F. et al. The mechanisms of renin–angiotensin system in hepatocellular carcinoma: From the perspective of liver fibrosis, HCC cell proliferation, metastasis and angiogenesis, and corresponding protection measures. Biomed. Pharmacother. 141, 111868 (2021).
Yan, K., Gao, Y. & Heller, S. L. Breast cancer screening utilization and outcomes in women with neurofibromatosis Type 1. Clin. Breast Cancer 23, e200–e205 (2023).
Evans, D. G. R. et al. Breast cancer in neurofibromatosis 1: survival and risk of contralateral breast cancer in a five country cohort study. Genet. Med. 22, 398–406 (2020).
Chen, X. et al. Dual inhibition of PI3K and mTOR mitigates compensatory AKT activation and improves tamoxifen response in breast cancer. Mol. Cancer Res. 11, 1269–1278 (2013).
Kim, J. W. et al. PIK3CA mutation is associated with poor response to HER2-targeted therapy in breast cancer patients. Cancer Res. Treat. J. Korean Cancer Assoc. 55, 531–541 (2023).
Mayer, I. A. Clinical implications of mutations in the PI3K pathway in HER2+ breast cancer: prognostic or predictive?. Curr. Breast Cancer Rep. 7, 210–214 (2015).
Lesman, D., Rodriguez, Y., Rajakumar, D. & Wein, N. U7 snRNA, a small RNA with a big impact in gene therapy. Hum. Gene Ther. 32, 1317–1329 (2021).
Goyenvalle, A., Babbs, A., van Ommen, G.-J. B., Garcia, L. & Davies, K. E. Enhanced exon-skipping induced by U7 snRNA carrying a splicing silencer sequence: Promising tool for DMD therapy. Mol. Ther. 17, 1234–1240 (2009).
Gadgil, A. & Raczyńska, K. D. U7 snRNA: A tool for gene therapy. J. Gene Med. 23, e3321 (2021).
Tisdale, S. et al. SMN is essential for the biogenesis of U7 small nuclear ribonucleoprotein and 3′-end formation of histone mRNAs. Cell Rep. 5, 1187–1195 (2013).
Wallis, D. et al. Exon skipping as a therapeutic for neurofibromatosis Type I. Research Square Preprint https://doi.org/10.21203/rs.3.rs-751331/v1 (2021).
Lim, K. R. Q., Maruyama, R. & Yokota, T. Eteplirsen in the treatment of Duchenne muscular dystrophy. Drug Design Dev. Ther. 11, 533–545 (2017).
Dvinge, H., Kim, E., Abdel-Wahab, O. & Bradley, R. K. RNA splicing factors as oncoproteins and tumour suppressors. Nat. Rev. Cancer 16, 413–430 (2016).
Wu, X. et al. Recent advances in dual PI3K/mTOR inhibitors for tumour treatment. Front. Pharmacol. 13, 875372 (2022).
Salphati, L. et al. Preclinical assessment of the PI3Kα selective inhibitor inavolisib and prediction of its pharmacokinetics and efficacious dose in human. Xenobiotica 54, 808–820 (2024).
Chen, Q. et al. Paxalisib. PI3K/mTOR inhibitor, Treatment of gliomas. Drugs of the Future 47, 1–9 (2022).
Imber, B. et al. Multi-Center Phase I Study of Concurrent Paxalisib and Radiation Therapy in Patients with Solid Tumor Brain Metastases (BM) or Leptomeningeal Metastases (LM) Harboring PI3K Pathway Mutations. Int. J. Radiat. Oncol., Biol., Phys. 120, S193 (2024).
Lim, J. H. et al. Induction of nephron-ductal dual lineages via early retinoic acid signaling establishes a platform for fully patterned kidney organoids. Adv. Healthcare Mater. 14, e02044 (2025).
Acknowledgements
We would like to thank Editage (www.editage.co.kr) for English language editing. The biospecimens and data used for this study were provided by the Biobank of Chungnam University Hospital. Schematic and illustrative images in the figures were created with BioRender.com and Mind the Graph. This research was supported by the National Research Foundation of Korea (NRF), funded by the Ministry of Science and ICT (grant numbers RS-2023-00225239 and RS-2024-00340403), and by the KRIBB Research Initiative Program (KQM0042611).
Author information
Authors and Affiliations
Contributions
The Author Contributions statement has been updated to reflect the revised order of the co-first authors following substantial additional experimental work and data analysis performed during the revision process. C.R.J. conceptualized and coordinated the overall study and led the design and execution of the multi-organ microphysiological system (MPS) and PK/PD modeling. M.S. supervised the development and validation of patient-specific iPSCs and intestinal organoids. M.J.S. supervised the development and validation of liver organoids. H.C. supervised genome-wide data production and comparative analysis. J.H.L. conceptualized and performed therapeutic screening, drug response assays, and pharmacological analyses, including PK/PD. H.M.K., D.W.Y., and S.J.M. conducted experiments with organoids. S.L. and D.S.K. performed RNA-seq data analysis. J.L. and S.J.O. performed LC-MS analyses. J.L. and S.J.K. provided technical support. All authors contributed to study design, data interpretation, and manuscript preparation. All corresponding authors take joint responsibility for the integrity and accuracy of the entire work. All authors have read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare that Cho-Rok Jung is a co-inventor on Patent (US11,732,232 B2), which is related to the cell culture system used in this study. The other authors declare no competing interests.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lim, J.H., Mun, S.J., Kang, H.M. et al. Personalized pharmacokinetic–pharmacodynamic guided therapy via an induced pluripotent stem cell–derived multi-organoid platform in NF1-mutant breast cancer. Sig Transduct Target Ther 11, 79 (2026). https://doi.org/10.1038/s41392-026-02595-7
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02595-7