Abstract
Epigenetic dysregulation is a fundamental cancer hallmark, and lysine demethylase 1 (LSD1) is a central target for cancer intervention. Developing novel LSD1 inhibitors with high selectivity, favorable bioavailability, and safety for acute myeloid leukemia (AML) remains challenging. We developed DC551040, a highly potent, selective irreversible LSD1 inhibitor with good tolerability in Phase I AML clinical trial (CTR20222026). DC551040-LSD1 complex crystal structure uncovered a new binding pocket, providing molecular insights for subsequent LSD1 inhibitor design. Given the significant role of LSD1 in epigenetic regulation, we performed comprehensive transcriptomic and proteomic analyses to investigate gene and protein expression dynamics following DC551040 treatment in an MV-4-11 xenograft model. These analyses revealed that multiple immune and inflammation related pathways are activated upon DC551040 treatment, including the key members STAT5, NF-κB, and AKT, suggesting the potential for adaptive resistance. Through a search of the Connectivity Map (CMAP) database, we identify homoharringtonine (HHT), an approved anti-leukemia drug, which mimics the anti-transcriptional activation of inflammatory pathways. Subsequent in vitro and in vivo experiments validated the efficacy of combining HHT with DC551040, demonstrating a synergistic antitumor effect and extended survival in MV-4-11 disseminated xenograft model mice. Together, this study not only introduces a novel LSD1 inhibitor but also delves into the molecular mechanisms underlying LSD1 inhibitors, while proposing a promising combination therapy for AML individuals in clinical trials.
Similar content being viewed by others
Introduction
Epigenetic dysregulation is a hallmark of cancer. As one of the critical epigenetic regulatory factors involved in mediating histone lysine methylation, lysine demethylase 1 (LSD1) has been established as a significant therapeutic target for cancer treatment. LSD1 exhibits multifaceted functions, serving not only as a histone modification enzyme capable of demethylating histone H3 lysine 4 (H3K4) and H3 lysine 9 (H3K9) but also as a nonhistone protein demethylase, such as the removal of a methyl group from p53K370me2, thereby inhibiting p53’s function.1 Elevated LSD1 expression facilitates tumorigenesis by modulating key biological processes, including chromatin remodeling, cell differentiation and cellular aggregation.2,3,4,5 Moreover, LSD1 is upregulated in various tumors, including those of the lung, breast, prostate, bladder, and blood,6 and is associated with poor prognosis and chemoresistance.7 As a result, LSD1 has become an increasingly significant therapeutic target for cancer treatment.
To date, several LSD1 inhibitors have been developed, including some irreversible LSD1 inhibitors discovered based on the structure of 2-phenylcyclopropan-amine (2-PCPA), which was initially identified as a monoamine oxidase (MAO) inhibitor. Promising in vitro and in vivo preclinical data have driven clinical trials of LSD1 inhibitors, including ORY-1001, GSK2879552, IMG-7289, and INCB059872, for the treatment of acute myeloid leukemia (AML).8 AML is a heterogeneous hematological malignancy characterized by the suppression of myeloid differentiation and abnormal proliferation of hematopoietic blast cells. MLL- or AML-ETO-translocated AML cells display pronounced sensitivity to LSD1 inhibitor monotherapy.9,10 ORY-1001, a representative agent, triggers blast cell differentiation in MLL-fusion AML at the molecular and morphological levels.11
LSD1 exerts a pleiotropic role in regulating gene expression through mediating epigenome establishment. Mechanistically, within the LSD1-CoREST corepressor complex, LSD1 acts as a transcriptional repressor by demethylating the active histone modifications H3K4me1/2.12 Furthermore, LSD1 is also capable of promoting gene transcription via demethylating repressive histone marks including H3K9me1/2.13,14 This suggests that LSD1 coordinates multiple cellular processes simultaneously, which may paradoxically activate oncogenic pathways and potentially weaken the anticancer effects. Therefore, developing new selective, bioavailable, and safe LSD1 inhibitors for AML therapy remains a challenging task. Furthermore, combining of LSD1 inhibitors with other agents, such as all-trans-retinoic acid and azacitidine, has shown enhanced anti-AML activity in preclinical and clinical stages.15,16 These results underscore dual priorities: advancing potent, highly selective LSD1 inhibitors toward clinical application, and investigating new drug combinations to improve the efficacy of LSD1 inhibitors in early-phase clinical trials.
In the present study, we firstly developed DC551040, an irreversible covalent LSD1 inhibitor characterized by high potency, excellent selectivity, and oral availability, which exhibits robust anti-AML effects in vitro and in vivo and is currently progressing through Phase I clinical trials (CTR20222026). The crystal structure of the DC551040-LSD1 complex revealed a novel binding pocket on the surface of LSD1, providing insights for the rational design of future LSD1 inhibitors. Secondly, recognizing the critical role of LSD1 in epigenetic regulation, we conducted comprehensive transcriptomic and proteomic dynamic analyses to systematically assess the dynamic molecular changes in an AML in vivo model in response to DC551040 treatment. The results demonstrated that prolonged DC551040 treatment induced complex immune and inflammation-related pathways in vivo. We also highlight that inflammation related pathways, including STAT5, NF-κB, and AKT, were activated in response to DC551040 treatment, shedding light on the molecular mechanisms underlying adaptive resistance to LSD1 inhibition. Moreover, through integrated proteomics and transcriptomics analyses, as well as a search of the Connectivity Map (CMAP) database, we identified the clinical AML treatment drug homoharringtonine (HHT), which mimics the anti-transcriptional activation of inflammatory pathways. Based on these findings, we hypothesized that the combination of HHT with the LSD1 inhibitor DC551040 may synergistically enhance the anticancer effects. Subsequent in vitro and in vivo experiments validated this hypothesis, demonstrating a synergistic anticancer effect of HHT in combination with DC551040 in vitro and an extended survival time in MV-4-11 disseminated xenograft model mice. Taken together, our study has not only discovered a potent and highly selective LSD1 inhibitor but has also shed light on the molecular mechanisms and utilization strategies of LSD1 inhibitors. These findings support further exploration of new combination therapies involving HHT and LSD1 inhibitors, aiming to attain effective clinical responses in the future.
Results
Discovery of a potent, selective, oral irreversible covalent LSD1 inhibitor DC551040
2-PCPA is a well-known mechanism-based irreversible MAO inhibitors. Subsequently, it was shown that 2-PCPA further exerts an inhibitory effect on LSD1 through covalent adduct formation with FAD. However, its LSD1 inhibitory activity against was relatively weak, with an IC50 value of 32 μM. Notably, 2-PCPA exhibited high inhibitory activity against both MAO-A and MAO-B, suggesting potential central nervous system (CNS) adverse effect. A comparison of co-crystal structures of LSD1 and MAO-B with the 2-PCPA-FAD adduct reveals a significant space at the amino part of the adduct in LSD1, allowing accommodation of various substituents (Fig. 1a). Based on the crystal structure of 2-PCPA, novel LSD1 inhibitors were designed by introducing fluorine-containing piperidine substituents on the amino group of 2-PCPA. Firstly, fluorine-containing piperidine substituents could occupy the S2 pocket of LSD1, enhancing LSD1 inhibitory potency and increasing selectivity for MAOs. Secondly, the incorporation of fluorine, a commonly utilized group in drug design, aim to improve the physicochemical properties, metabolic stability, and target affinity of the compounds. Compound 1 displayed nanomolar-level LSD1 inhibitory potency (IC50 = 53.67 nM), which exhibited a 596-fold improvement compared to 2-PCPA. Furthermore, compound 1 showed high selectivity for MAO-A and MAO-B, with IC50 values more than 10 μM. Analysis of the binding mode of compound 1 with LSD1 indicated that fluorine-containing piperidine substituent effectively occupied the S2 pocket of LSD1. Notably, there was still ample space at the methyl group of compound 1, allowing for the accommodation of large substituents. Motivated by this observation, a series of fluorine-containing phenylcyclopropyl derivatives was designed and synthesized as novel LSD1 inhibitors, along with detailed structure−activity relationship (SAR) studies, considering that the introduction of the hydrophobic substituent at the piperidine potentially yield selective and potent LSD1 inhibitors (Fig. 1a).
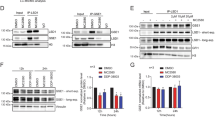
Discovery of a potent, selective, oral irreversible covalent LSD1 inhibitor DC551040. a Structure-Based Drug Design of Novel Fluorine-containing LSD1 Inhibitors. b LSD1 inhibition activity and selectivity against LSD2, MAO-A, and MAO-B of the potent LSD-1 inhibitor DC551040, GSK2879552, and 2-PCPA, data are represented as mean ± SEM, n = 4 per group. c Irreversibility of DC551040 was determined by dilution assay, and GSK2879552 was used as control, data are represented as mean ± SEM, n = 4 per group. d X-ray crystal structure of LSD1 with compound DC551040. e To characterize DC551040-mediated disruption of the LSD1–GFI1B–CoREST complex, MV-4-11 cells were treated with DMSO, 50 nmol/L, or 250 nmol/L DC551040, GSK2879552, or Bomedemstat for 24 h. Chromatin fractions were prepared, GFI1B-containing complexes were immunoprecipitated, and immunoblotting analysis was performed to detect the indicated proteins
All the synthesized compounds 1–9 were investigated in vitro for the inhibitory activity against LSD1 (Table 1). The SARs of synthesized fluorine-containing trans-2-phenylcyclopropyl-amino-methyl-piperidines were discussed as follows. GSK2879552 was treated as a postive control to evaluate the activity of synthesized compounds, which exhibited an IC50 value at 821.20 nM against LSD1 in enzymatic assays under the experimental conditions. When the methyl group of compound 1 was replaced by methylene cycloalkyl group as cyclopropylmethyl (2), cyclobutylmethyl (3), cyclopentylmethyl (4), cyclohexylmethyl (5), their IC50 values were 41.05 nM, 67.81 nM, 47.22 nM, 62.98 nM, which are similar as compound 1. In order to obtain compounds with optimal LSD1 inhibitory activity, acylation modifications were introduced to the piperidine and compound 6 was synthesized and evaluated. Firstly, an acetyl group was introduced and compound 6 (IC50 = 319.30 nM) was obtained with obviously decreased LSD1 potency. Surprisingly, when the methyl group was replaced by polar substituted group as piperidine, the resulting compound 7 showed increased LSD1 inhibitory activity with IC50 at 41.85 nM, proving that bulky polar groups may be favorable for the inhibitory activity. Then, the linker was changed with ethoxycarbonyl, and compound 8 showed excellent inhibitory potency on LSD1 with an IC50 value of 21.42 nM, indicating that bulkier substituents were favorable for S2 pocket. While the N-methyl (9) derivative displayed decreased LSD1 inhibitory potencies compared with compound 8. To investigate the influence of chirality, we conducted the enantioselective synthesis of the most potent compound 8, resulting in (1R,2S)-8 and (1S,2R)-8. The results of LSD1 inhibitory activity indicated that (1R,2S)-8 exhibited better LSD1 inhibition activity. On the basis of these results, we proposed that the hydrophobic group was required to enhance the binding affinity to LSD1, while a carboxyl group would be detrimental to binding affinity. As a result of its excellent LSD1 inhibitory activity, compound (1R,2S)-8 (DC551040) was further characterized in vitro and advanced into in vivo efficacy studies.
LSD1 falls into the FAD-dependent monoamine oxidase family, which includes MAO-A and MAO-B. Given that MAO-A and MAO-B are essential for neurotransmitter degradation in the central nervous system, selective inhibition of LSD1 is of considerable significance. In addition, another member of LSD family, LSD2 is also evaluated. Because inhibition of LSD2 was associated with toxicity in animal studies, selectivity against LSD2 was particularly important. Given the moderate to good LSD1 inhibitory activity exhibited by most synthesized LSD1 derivatives, we assessed the enzyme selectivity of DC551040 against LSD2, MAO-A, and MAO-B. (Fig. 1b). DC551040 exhibited excellent selectivity on LSD1 over LSD2 and MAO, which were better than the positive control GSK2879552 and Bomedemstat.
The irreversible mode of action of compound DC551040 on LSD1 was determined through a dilution assay (Fig. 1c). The results indicate that compound DC551040 may interact covalently with LSD1, as the activity rate of the LSD1 enzyme could not be recovered even after 3 h of dilution with DC551040. This phenomenon is similar to what was observed in the presence of Bomedemstat, a known covalent LSD1 inhibitor. Thus, it can be concluded that compound DC551040 is an irreversible and covalent LSD1 inhibitor.
To better understand the structural basis of the high LSD1 inhibitory activity, we investigated the crystal structure of LSD1 in complex with DC551040 (PDB ID: 6K3E), with a resolution of 2.87 Å (Fig. 1d). Our analysis revealed that DC551040 interacts with LSD1 in a two-step manner, involving recognition and covalent binding. First, the benzene ring of DC551040 extends into hydrophobic section S1, where it forms a π-π stacking interaction with Trp552. Meanwhile, the fluoride-substituted piperidine fragment occupies the conjunction section of S2 and FAD, heading S2 preferentially, with the fluorine forming polar interactions with Phe538. The piperidine fragment, which contains a carboethoxy group, extends deep into the S2 pocket. Second, DC551040 occupies the catalytic site of LSD1, with the 2-PCPA moiety of DC551040 located near the FAD cofactor, which results in the potential for covalent inhibition. Subsequently, the 2-PCPA moiety inactivates LSD1 covalently with FAD through a single electron transfer mechanism. Finally, the piperidine fragment of DC551040 is released through the hydrolysis of the imine intermediate. This suggests that this moiety may serve as an efficient carrier for delivering 2-PCPA into the catalytic site of LSD1. Overall, our findings demonstrate the discovery of a potent, selective, and irreversible covalent LSD1 inhibitor, DC551040, and provide insights into the structural basis underlying its inhibitory activity. Subsequently, we investigated the influence of this inhibitor on LSD1-containing protein complexes. LSD1 assembles into a complex with the corepressor proteins CoREST and GFI1B.17 As shown in Fig. 1e, the LSD1 inhibitors DC551040, GSK2879552 and Bomedemstat can all disrupt the LSD1-GFI1B association.
DC551040 has an antitumor effect in an AML cell line and a patient-derived AML cell line
DC551040 displayed significant potency in inhibiting antiproliferative activities in several AML cell lines, including MV-4-11 (IC50 = 79.51 nM), Kasumi-1 (IC50 = 25.77 nM), and HL60 (IC50 = 40.35 nM) (Fig. 2a). Further evaluation of the inhibitory activity on the other blood cancer cell lines, such as RS4:11 (acute lymphoblastic leukemia), MOLT-4 (acute lymphoblastic leukemia), Mino (mantle cell lymphoma), Z-138 (mantle cell lymphoma), MAVER-1 (mantle cell lymphoma), Raji (burkitt’s lymphoma), MM1S (multiple myeloma), which are insensitive to LSD1, revealed that DC551040 showed weak inhibitory activity against these non-AML cell lines, with an IC50 value over 20 μM. Moreover, DC551040 was evaluated in AML patient-derived cells (PDCs, Clinical information of PDCs is shown in Supplementary Tables 1, 2) and showed inhibitory effects on four AML cell lines, with IC50 values of 0.68 nM, 1.00 nM, 48.54 nM, and 4157 nM, respectively (Fig. 2b). Taken together, these results demonstrated the potential of DC551040 as an anti-AML agent through its selective inhibition of LSD1.
DC551040 has an antitumor effect in acute myeloid leukemia in vitro. a Compound DC551040 inhibits the proliferation of the AML cell lines MV-4-11, Kasumi-1, and HL-60 in vitro. This panel shows the effects of DC551040 on the growth of ten AML cell lines. Cells were treated with gradient concentrations of DC551040 or GSK2879552 for 7 days, followed by cell growth assessment. The y-axis represents the growth half-maximal inhibitory concentration (IC₅₀) value. Data are presented as mean ± SEM, n = 3 per group. b Compound DC551040 inhibits the proliferation of AML patient-derived cells (PDCs) in vitro. This panel depicts the effects of DC551040 on the growth of 24 AML PDC samples. PDCs were treated with gradient concentrations of DC551040 or GSK2879552 for 7 days prior to cell growth evaluation. The y-axis represents the growth IC₅₀ value. Data are presented as mean ± SEM, n = 3 per group. c In MV-4-11 cells, compounds DC551040 and GSK2879552 induce cell apoptosis. MV-4-11 cells were treated with each compound at concentrations of 0.5 μM, 2 μM, and 4 μM for 72 h, and apoptotic effects were detected by flow cytometry. d Summary statistics of the apoptosis rate are shown. Data are presented as mean ± SEM, n = 3 per group. e CD86 expression levels in MV-4-11 cells treated with DC551040, GSK2879552, and Bomedemstat. Data are presented as mean ± SEM, n = 3 per group. f Morphological changes of MV-4-11 cells analyzed by Wright–Giemsa staining after treatment with indicated concentrations. Compared with DMSO treatment, the morphology of MV-4-11 changed after treatment with compound DC551040 for 24 h, the purple-red cell nucleus were shrunk and appeared in a half-moon state, and the light blue cytoplasm was enlarged. Based on the above results, we speculated that MV-4-11 differentiated into monocytes/macrophages. The positive compound GSK2879552 and Bomedemstat had similar effects to DC551040. g MV-4-11 cells were incubated with vehicle and compound DC551040 at 0.1 μM, 0.5 μM and 2 μM for 1, 3, 5, and 7 days, and the dimethylation level of histone H3K4 were determined by Western blot
The effects of compound DC551040 on apoptosis were evaluated in MV-4-11 cells. The cells were treated with either vehicle alone, compound DC551040 or GSK2879552 at concentrations of 0.5 μM, 2 μM and 4 μM for 72 h, and the results were analyzed using flow cytometry. The findings indicated that, similar to GSK2879552, DC551040 induced apoptosis in a dose-dependent manner in MV-4-11 cells (Fig. 2c, d, Supplementary Fig. 1). The subsequent apoptosis analysis revealed that the proportions of MV-4-11 cells in the apoptosis phase were 7.16% at 0.5 μM, 7.78% at 2 μM and 10.76% at 4 μM DC551040, compared to 4.07% in the DMSO control group.
To evaluate the ability of compound DC551040 to induce differentiation in MV-4-11 cells, we conducted an assay to determine whether it could induce expression of the cellular biomarker CD86. The positive control GSK2879552 and Bomedemstat was also included in the assay. Our results showed that DC551040 was more effective than GSK2879552 and Bomedemstat at upregulating the expression level of CD86 (Fig. 2e). Additionally, we detected differentiation-related morphological alterations in cells after DC551040 administration, including reduced nuclear-to-cytoplasmic ratios and nuclear segmentation. These morphological changes were also observed following treatment with GSK2879552 and Bomedemstat. Therefore, we concluded that DC551040 exhibits a significant anticancer effect on cell differentiation in leukemic MV-4-11 cells (Fig. 2f).
Given that LSD1 is responsible for demethylating mono- and dimethyl histone H3 lysine 4 (H3K4), we hypothesized that DC551040 could affect the level of H3K4 dimethylation (H3K4me2) by inhibiting the enzymatic activity of LSD1. To test this hypothesis, we treated MV-4-11 cells with either vehicle or DC551040 at concentrations of 0.1 μM, 0.5 μM and 2 μM for 1, 3, 5 and 7 days. The H3K4me2 levels were detected by western blot analysis. As expected, the level of H3K4me2 in MV-4-11 cells increased in a dose-dependent manner upon treatment with DC551040 for 1, 3, 5 or 7 days (Fig. 2g, Supplementary Fig. 2).
In addition, we compared the antiproliferative effects of DC551040, GSK2879552, and Bomedemstat (two LSD1 inhibitors that have entered clinical development) on normal peripheral blood mononuclear cells (PBMCs). The results showed that the IC₅₀ values for all three compounds were greater than 10 μM in PBMCs, suggesting a favorable selectivity for malignant cells over normal hematopoietic cells (Supplementary Fig. 3).
DC551040 has an antitumor effect in acute myeloid leukemia in vivo
Further pharmacokinetic (PK) investigations indicated that DC551040 demonstrated favorable PK parameters in Institute of Cancer Research (ICR) mice (Supplementary Table 3). In ICR mice administered a 20 mg/kg oral dose of DC551040, the compound was rapidly absorbed (Tmax = 0.5 h) and well exposed (AUC0−t = 2566 h·ng/mL). Furthermore, DC551040 was observed to display high bioavailability in ICR mice (F = 74.4%). Based on the good PK profiles of compound DC551040 in mice, it was evaluated in anti-cancer efficacy studies using Kasumi-1 and MV-4-11 xenograft model systems (Fig. 3a–d).
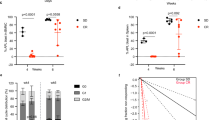
DC551040 has an antitumor effect in acute myeloid leukemia in vivo. a, b DC551040 anti-tumor effect in Kasumi-1 xenografts. The relative tumor volume (RTV) was calculated twice weekly; tumors were excised and measured at study end. Mice was treated with DC551040 (0.5/1/2 mg/kg/day), GSK2879552 (2 mg/kg/day), or vehicle (treatment duration: 17 days). Data: mean ± SEM, n = 6/group. c, d DC551040 anti-tumor effect in MV-4-11 xenografts. RTV was calculated twice weekly; tumors were excised and measured at study end. Mice was treated with DC551040 (5/10 mg/kg/day), GSK2879552 (10 mg/kg/day), or vehicle (treatment duration: 22 days). Data: mean ± SEM, n = 6/group. e, f DC551040-induced CD86 upregulation in MV-4-11 xenografts (immunohistochemistry). Mice treated with DC551040 (5/10 mg/kg/day), GSK2879552 (10 mg/kg/day), or vehicle. Data: mean ± SEM, n = 6/group. g The anti-cancer effect of DC551040 on MV-4-11 disseminated xenograft mice model. Survival for randomized mice bearing MV-4-11 tumors treated with 0.5, 2, and 4 mg/kg/day DC551040, 0.5 mg/kg/day GSK2879552, and vehicle group. Drug treatment started from recording the survival time and lasted for 21 days, n = 10 per group. h Anti-tumor effect of DC551040 ± Cytarabine (AraC) in MV-4-11 xenografts. RTV was calculated twice weekly; tumors were excised and measured at study end. Mice was treated with DC551040 (2/5 mg/kg/day), AraC (20 mg/kg/day), DC551040+AraC, or vehicle (treatment duration: 22 days). Data: mean ± SEM, n = 6/group. i The anti-cancer effect of DC551040, Cytarabine and Daunorubicin on MV-4-11 disseminated xenograft mice model. Survival for randomized mice bearing MV-4-11 tumors treated with 0.5 mg/kg/day DC551040, 7.5 mg/kg/day AraC and 0.75 mg/kg/day DAC, DC551040 + AraC + DAC, and vehicle group. Drug treatment started from recording the survival time and lasted for 21 days, n = 10 per group. The Significance of (a, b, c, d, f, h) was analyzed by equal variance two-tailed t test. The Significance of (g, i) was analyzed by logrank test. *P < 0.05, **P < 0.01, ***P < 0.001
Firstly, we evaluated its activity in a Kasumi-1 cell-derived xenograft mouse model. When administered orally once daily at doses of 0.5, 1, and 2 mg/kg, DC551040 demonstrated a dose-dependent tumor suppressive activity, with corresponding tumor growth inhibition (TGI) rates of 48.82%, 60.81%, and 86.99%, respectively. Notably, the antitumor effect of compound DC551040 (2 mg/kg) was superior to that of GSK2879552 (2 mg/kg) (Fig. 3a, b).
Then, we investigated the efficacy of compound DC551040 in a mouse model using MV-4-11 cell-derived xenografts. The mice were administered oral doses of 5 and 10 mg/kg of DC551040 once a day. Our results showed a dose-dependent tumor suppressive effect, with TGI rates of 41.08% and 61.23%, respectively. Notably, the tumor inhibitory effect of compound DC551040 (10 mg/kg) was better than that of GSK2879552 at the same dose (p < 0.05) (Fig. 3c, d). Importantly, all mice in each group tolerated the effective doses, indicating that the toxicity of compound DC551040 is manageable. To gain insights into the dose-effect relationship of compound DC551040, we evaluated the expression levels of the biomarker CD86 in mice. The results demonstrated a good dose-dependent effect of compound DC551040 on CD86 expression levels in mice (Fig. 3e, f).
To better elucidate the antitumor potential of compound DC551040, we evaluated its effects on survival in an MV-4-11 model. Administered orally once daily at doses of 0.5, 2, and 4 mg/kg, compound DC551040 demonstrated a dose-dependent survival effect (Fig. 3g). In addition, In MV-4-11 cell-derived xenograft mouse model, we found that DC551040 can significantly synergize with the commonly used AML drugs cytarabine (AraC) and daunorubicin (DAC) to exert tumor suppressive effect and prolong the survival of mice (Fig. 3h, i). Taken together, these findings suggest that compound DC551040 possesses a dose-response relationship and is able to synergize with AML chemotherapy drugs in these AML tumor models while also exhibiting no obvious toxicity to mice at the doses tested.
DC551040 displayed high oral bioavailability in rats (F = 92.0%) and dogs (F = 78.6%), and it also exhibited well exposed (AUC0−t = 5777 and 10940 h·ng/mL) (Supplementary Table 3). We further found that DC551040 showed low inhibitory activity against P450 isoforms (IC50 > 50 μM), indicating a low risk for drug-drug interactions (DDIs) (Supplementary Table 4). Meanwhile, we used the UPLC/Q-TOF MS method to identify the metabolic pathway of DC551040 in SD rats (Supplementary Fig. 4). Following a single oral administration of 0.5 mg/kg DC551040 in rats, only the parent drug (M0) was detected in plasma and feces, while a small amount of mono-oxidized dehydrogenated metabolite (M1-1, LC-MS peak area <5% of M0) was observed in urine, indicating a high metabolic stability of DC551040 in vivo. To further evaluate the safety profile of compound DC551040, we evaluated compound DC551040 in a human ether-à-go-go-related gene (hERG) assay and found that it did not inhibit hERG even at concentrations as high as 30 μM (Supplementary Table 5). This suggests that the compound DC551040 is unlikely to cause QT interval prolongation-related cardiac toxicity. Additionally, compound DC551040 was tested negative in an in vitro genotoxicity study (AMES test, with and without S9 metabolic activation). We also conducted in vivo single-dose SD rat and Beagle dog acute toxicity evaluation studies, which found that the maximum tolerated dose of DC551040 was more than 250 mg/kg and 20 mg/kg, respectively. Safety pharmacology study results indicated that no significant treatment-related effects on the central nervous system (CNS) were observed in ICR mice with DC551040 administered orally at doses of 0.5 mg/kg, 1.5 mg/kg, and 5 mg/kg. Pentobarbital-induced hypnosis tests showed no statistically significant changes in sleep latency or duration were observed in the DC551040 treatment groups, while mice demonstrated significantly shortened sleep latency (P < 0.05) and prolonged sleep duration (P < 0.05) after intravenous injection of chlorpromazine in the positive-control group (Supplementary Fig. 5). To evaluate potential CNS toxicity-related motor coordination dysfunction, the rotarod test was used—this assay was therefore employed to assess the neurotoxic effects of DC551040 at different time intervals. (Supplementary Fig. 6). DC551040 treated and vehicle groups ICR mice retained almost identical times on rotarod apparatus, which indicated non-neurotoxic nature of DC551040, while the chlorpromazine group demonstrated significant reductions in running time on the rotarod apparatus at time intervals 10, 40, 90, 120, and 180 min after treatment (P < 0.05). Overall, these results indicated that no significant treatment-related effects on the central nervous system were observed in ICR mice with DC551040 administered orally at doses of 0.5 mg/kg, 1.5 mg/kg, and 5 mg/kg. Briefly, DC551040 did not exhibit any obvious toxicity compared to the adverse effects of GSK-2879552 reported in clinical trials.
Based on its ideal drug-like properties, DC551040 has completed preclinical IND-enabling studies and is now in a Phase I clinical trials (CTR20222026). The primary objective was to assess safety of DC551040, and determine recommended phase 2 dose (RP2D); secondary objectives were to study pharmacokinetics (PK) and efficacy. Dose-escalation was performed in a modified 3 + 3 design, except for the first three dose groups which employed accelerated dose titration, dose-escalation in subsequent groups will follow the 3 + 3 design, with 3–6 patients enrolled per group. The escalation doses were 0.2, 0.4, 0.8, 1.2, 1.5, 1.8, 2.0 mg/d, and the starting dose was based on preclinical toxicology studies. Upon completion of the clinical trial unblinding process, the relevant results will be formally disclosed.
Transcriptomic and proteomic dynamic analysis of DC551040 treated AML in the MV-4-11 xenograft model
Given the epigenetic regulation of DC551040, we then examined the quantitative proteomic profile of MV-4-11 xenografts in nude mice following treatment with DC551040 at 3, 7, and 21 days (Fig. 4a). In total, 6218 proteins were successfully identified (with 1% false discovery rate at the peptide and protein levels). Among them, 233, 276, and 256 proteins changed significantly in mice treated with DC551040 compared to those treated with vehicle at 3, 7, and 21 days, respectively (|log2(fold change)| > 0.585 and p < 0.05). The proteins differentially expressed in any group were combined, and they could be clustered into four subgroups (Fig. 4b). The proteins in cluster 1 peaked at day 3 and showed a steady decrease in expression over time. This class of proteins was enriched in biosynthesis of amino acids, ribosome, lysosome, cell cycle, DNA damage checkpoint, leukocyte degranulation, myeloid leukocyte mediated immunity, and cell activation involved in immune response pathways. In contrast, the proteins in cluster 2, which were related to carbon metabolism, antigen processing and presentation, nucleotide metabolism, biosynthesis of amino acids, cytokine-mediated signaling pathway, leukocyte activation, and glucose 6-phosphate metabolic process pathways, had relatively stable expression at 3 and 7 days of administration. However, they were significantly upregulated at 21 days of administration. Unlike clusters 1 and 2, the expression of proteins in cluster 3 was significantly upregulated after 7 days and remained upregulated at 21 days. They were involved in fatty acid metabolism, biosynthesis of unsaturated fatty acids, cell cycle, autoimmune thyroid disease, chromosome localization, mitotic nuclear division, and response to lipid. In addition, cluster 4, which peaked at 7 days and dropped abruptly thereafter, encompassed proteins participating in oxidative phosphorylation, acute myeloid leukemia, neutrophil mediated immunity, leukocyte mediated immunity, immune effector process, cell activation involved in immune response, and myeloid leukocyte activation.
Drug combination exploration was conducted based on the dynamic proteomic and transcriptomic profiling of DC551040 treated MV-4-11 in vivo. a Drug combination exploration that relied on the dynamic change of transcriptome and proteome in the DC551040-treated MV-4-11 xenograft model. b Hierarchical clustering of differentially expressed proteins was performed in mice treated with 2 mg/kg DC551040 at 3, 7, and 21 days, respectively. The pathways related to the proteins in each cluster are labeled
The above results indicated that DC551040 significantly affected the metabolism and immunity of mice over time. Therefore, we further investigated the dynamic changes in the pathways involved in metabolism and immunity at 3, 7, and 21 days (Fig. 5a). Interestingly, immune-related pathways were activated at the beginning of the administration and remained active throughout administration. For example, cell activation involved in the immune response, leukocyte mediated immunity, and neutrophil mediated immunity were clearly enhanced in response to DC551040 supplementation. However, the metabolic pathways were significantly altered during administration. Oxidative phosphorylation was significantly weakened at 21 days of administration, while carbon metabolism and glycolysis/gluconeogenesis were enhanced at the same time. In addition, fatty acid metabolism was decreased remarkably at 3 days and increased at 7 and 21 days.
Dynamic proteomic changes in the pathways involved in immunity and inflammatory signaling. a Gene set enrichment analysis of the quantitative proteome in mice treated with DC551040 at 3, 7, and 21 days, respectively. Volcano plots (b) and heatmap (c) were analyzed to identify differentially expressed proteins (DEPs) enriched in IL6-JAK-STAT3/IL2-STAT5 signaling and Interferon/inflammatory signaling in mice treated with DC551040 at 3, 7, and 21 days, respectively
Given that immune and inflammatory pathways were highly enriched from the upregulated proteins in the Gene Set Enrichment Analysis (GSEA), we next explored the differentially expressed immune and inflammatory related proteins affected by DC551040 treatment at 3, 7, and 21 days (Fig. 5b, c, Supplementary Table 6). The volcano plot showed that proteins enriched in IL6-JAK-STAT3/IL2-STAT5 signaling and interferon/inflammatory pathways were obviously increased in the mice treated with DC551040 at 3, 7, and 21 days, in agreement with the previous results. Furthermore, we selected several differentially expressed proteins associated with IL6-JAK-STAT3/IL2-STAT5 and interferon/inflammatory signaling to investigate the dynamic expression changes during DC551040 treatment in mice at 3, 7, and 21 days (Fig. 5c). The heatmap showed that some proteins maintained a high expression level in mice treated with DC551040 from 3 days to 21 days, such as galactose mutarotase (GALM), which is involved in galactose metabolism (Fig. 5c). Additionally, several proteins were increased by the prolongation of treatment time with DC551040 in mice, such as stabilin-1 (STAB1), which may function in angiogenesis, lymphocyte homing, cell adhesion, or receptor scavenging, and guanine nucleotide-binding protein subunit alpha-15 (GNA15), which is involved as a modulator or transducer in various transmembrane signaling systems. In addition, we found that treatment of mice with DC551040 led to some proteins reaching peak expression levels at an early time (Day 3) and then gradually declining, such as aryl hydrocarbon receptor (AHR), which is related to development, immunity and cancer, and high affinity copper uptake protein 1 (SLC31A1), which is involved in dietary copper uptake. However, some proteins peaked at 7 days, such as T-lymphocyte activation antigen (CD86), which is involved in the costimulatory signal, and Ly6/PLAUR domain-containing protein 8 (PLAUR), which is required to prevent invasion of gram-negative bacteria. Together, these results provide insights into the potential targets and mechanisms of DC551040 for application in treating diseases on a proteome-wide level.
Screening of combination drugs for DC551040 based on transcriptomics, proteomics and CMAP database mining
We also investigated changes in the transcriptome at the same time. GSEA was performed on upregulated mRNA and protein at different time periods, and it was found that at the transcriptional level, immune-related pathways, including IL6-JAK signaling, inflammatory response, IL2-STAT5 signaling, interferon-γ response, and interferon-α response were activated (Fig. 6a). At the protein level, the results of the analysis were consistent with the previous conclusions, glycolysis, inflammatory responses, IL2-STAT5 signaling, and interferon-γ responses were also activated (Fig. 6a).
Search for a drug that can be combined with DC551040 through transcriptomic, proteomic and CMAP database analysis. a Gene set enrichment analysis (GSEA) was performed corresponding to up-regulated RNA and protein in DC551040 treatment for 3, 7, and 21 days, respectively. The inflammatory pathways, including IL2-STAT5, IL6-STAT5 and other immune-related pathways were up-regulated and enriched in DC551040 treatment. b Drug combination prediction was based on the CMAP analysis. The heatmap displayed the ability of a series of drugs to down-regulate the proteins and RNAs up-regulated in DC551040 treatment. Potential synergistic drugs for DC551040 were identified by the ability of the drugs to strongly down-regulate the proteins and RNAs, with approved drugs being marked in red. c The regulation of STAT3 phosphorylation levels by HHT and DC551040 in MV-4-11 was detected based on Western blot. d Effect of DC551040 and GSK2879552 on the transcription of IL6 in three AML cell lines based on rt-qPCR experiments. Data are represented as mean ± SEM, n = 3 per group. e Effect of DC551040, HHT and C188-9 on the transcription of IL6, CCL5, MYC, and VEGFA in MOLM-13 based on rt-qPCR experiments. Data are represented as mean ± SEM, n = 3 per group. The Significance of (d, e) was analyzed by equal variance two-tailed t test. *P < 0.05, **P < 0.01, ***P < 0.001
To enhance therapeutic outcomes and mitigate drug-associated toxicity in patients as well as non-target tissues, co-administration of one or more additional agents is a common clinical practice. Moreover, a rationally designed combination therapy has the potential to attenuate or delay the development of drug resistance. To explore the potential drugs that could reverse the upregulated gene/protein expression signatures by DC551040, we input the gene list corresponding to the proteins and mRNAs upregulated in the three time periods (six gene sets) into the CMAP database. We further searched the CMAP database for drugs that could inhibit the set of highly expressed genes and proteins (Fig. 6b). We entered the symbols corresponding to the proteins and mRNAs upregulated in the three time periods (six gene sets) into the CMAP database (Fig. 6b), selected compounds whose top 25 can significantly downregulate the expression of these genes, and investigated the research stage of the compounds, among which three kinds of compounds have been approved, including entecavir, homoharringtonine (HHT), ruxolitinib, of which ruxolitinib is a known JAK inhibitor, and HHT is a commonly used chemotherapeutic drug for AML treatment. HTT suppresses STAT3 activity by targeting the IL-6/JAK1/STAT3 signaling axis, causes cell-cycle arrest and apoptosis,18,19 and it is approved to cure a certain of tumors including AML, which is in line with the previous transcriptome and proteome functional enrichment conclusions. Furthermore, in MV-4-11, we found that DC551040 can up-regulate the phosphorylation of STAT3, while HHT can down-regulate the phosphorylation of STAT3, so these two drugs do have opposite regulatory effects on the STAT3 pathway (Fig. 6c, Supplementary Fig. 2). In addition, we performed qPCR to assess the transcriptional regulation of IL6 gene by LSD1 inhibitors (DC551040 and GSK2879552) in three AML cell lines (MOLM-13, MV-4-11, and HL-60). Both compounds significantly upregulated IL6 mRNA expression in a dose-dependent manner across all three cell lines (Fig. 6d), suggesting a conserved response to LSD1 inhibition. Since IL6 is a known upstream activator of STAT3, which in turn drives the transcription of key oncogenic and inflammation-related genes (e.g., MCL1, MYC, ICAM1, TGFB1, VEGFA, and CCL5).3 Therefore, we next examined whether DC551040 also modulates these downstream targets. qPCR analysis revealed that DC551040 significantly upregulated the mRNA expression of MYC, CCL5 and VEGFA in AML cells (Fig. 6e), suggesting activation of the IL6–STAT3 axis as a potential resistance mechanism to LSD1 inhibition,4,5,6,7,8 and we also found that HHT treatment alone could also downregulate the mRNA levels of IL6, MYC, CCL5, and VEGFA in MOLM-13 cells by qPCR experiments, and to confirm the role of STAT3 signaling, we treated MOLM-13 cells with the STAT3 inhibitor C188-9, which downregulated the mRNA expression of MYC, CCL5, and VEGFA, except for IL6, the upstream gene of STAT3 (Fig. 6e). Therefore, we examined whether DC551040 and HHT combination had the synergistic effect in AML treatment.
DC551040 in combination with HHT synergistically enhances the treatment of AML
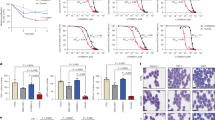
To further confirm the combined therapeutic potential of HHT and DC551040, we conducted combination experiments of HHT and DC551040 in four AML cell lines (MOLM-13, MV-4-11, HL-60, and Kasumi-1), one patient-derived cell (PDC) sample and one patient-derived tumor xenograft (PDX) model. Our comprehensive analysis using the ZIP algorithm demonstrated consistently significant synergistic effects across all tested models, with ZIP scores exceeding 10 in every case (Fig.7a–h). Notably, we also observed a clear synergistic interaction between DC551040 and HHT in the AML PDC sample (32#) (Fig. 8a, b). Therefore, supplementation with HHT significantly enhanced DC551040-induced cell death (Fig. 7a–d, Fig. 8a, b) and cell apoptosis (Fig. 8c), indicating a synergistic effect. To further evaluate the potential of DC551040 to enhance HHT sensitivity in vivo, we established a disseminated MV-4-11 AML mouse model (Fig. 8d). Neither DC551040 nor HHT as a single agent significantly improved mouse survival time; by contrast, their combination substantially extended overall survival in the MV-4-11 model (median survival time: 57.5 days vs 48.5 days for HHT monotherapy, 57.5 days vs 47.5 days for DC551040 monotherapy; P = 0.038, P = 0.013, Fig. 8d). In addition, a patient-derived tumor xenograft model was established in NDG mice using primary AML cells (sample No. AML21001-X) harboring the MLL-AF9 fusion mutation. To evaluate the residual leukemia burden, human CD33+ in mouse bone marrow will be quantified by flow cytometry after treatment completion (Fig. 8e). The results indicated that DC551040 treatment significantly reduced the proportion of human CD33+ cells in the mouse bone marrow compared to vehicle control, with a greater reduction than that observed in the GSK2879552-treated group. Notably, the DC551040 + HHT combination produced significantly reduced the CD33+ cells, demonstrating a synergistic therapeutic effect (Fig. 8f, g, Supplementary Fig. 1). These findings collectively demonstrate that DC551040 can enhance HHT sensitivity in both in vitro and in vivo models.
DC551040 combination with HHT synergistically enhances the treatment of AML. The synergistic effect of different concentrations of HHT and DC551040 was detected in MV-4-11 cells (a), MOLM-13 cells (b), Kasumi-1 cells (c) and HL60 cells (d), and the synergistic ability was calculated using the ZIP algorithm. The synergistic effect of different concentrations of C188-9 and DC551040 was detected in MV-4-11 cells (e), MOLM-13 cells (f), Kasumi-1 cells (g) and HL60 cells (h), and the synergistic ability was calculated using the ZIP algorithm. All data of this figure are represented as mean ± SEM, n = 3 per group
DC551040 combination with HHT synergistically enhances the treatment of AML in PDC and in vivo. a The synergistic effect of different concentrations of HHT and DC551040 was detected in AML PDC (32#), data are represented as mean ± SEM, n = 3 per group, and b the synergistic effect was calculated using the Bliss algorithm. c Changes of Caspase3/7 activity in MV-4-11 after DC551040 combined with HHT or alone, data are represented as mean ± SEM, n = 3 per group. d Survival curve of B-NDG mouse disseminated xenografted MV-4-11 cells treated with vehicle (NS, n = 20 mice), DC551040 (1 mg/kg, n = 10 mice), HHT (0.5 mg/kg, n = 10 mice), or the combination, n = 10 per group. e Experimental workflow for testing drug efficacy in PDX models. f Detection results and g Statistical histogram of CD33 positive cells in mouse bone marrow based on flow cytometry, data are represented as mean ± SEM, n = 4 per group. The Significance of (c, g) was analyzed by equal variance two-tailed t test. The Significance of (d) was analyzed by logrank test. *P < 0.05, **P < 0.01, ***P < 0.001
Exploration of the synergistic mechanism between HHT and LSD1 inhibitor DC551040
To investigate the mechanistic basis of the synergy between HHT and LSD1 inhibitor DC551040, particularly regarding the modulation of inflammatory signaling pathways, we conducted RNA-seq profiling on MV-4-11 following treatment with DC551040, HHT, or the combination of these two agents.
First, we observed that DC551040 primarily induced gene upregulation, whereas HHT caused a broader downregulation of gene expression (Fig. 9a, b). Subsequent GSEA results indicated that DC551040 significantly enhanced the expression of the inflammatory pathway (encompassing the IL6-JAK-STAT3 and interferon-γ pathways), while HHT induced significant downregulation of these same pathways (Fig. 9c, d).
RNA-seq data analysis and validation of MV-4-11 treated with DC551040 or HHT. a Volcano plot showing the degree of gene difference between DC551040-treated group and DMSO control group. b Volcano plot showing the degree of gene difference between HHT-treated group and DMSO control group. c Summary of significantly upregulated pathways after DC551040 treatment based on gene set enrichment analysis (GSEA). d Summary of significantly downregulated pathways after HHT treatment based on gene set enrichment analysis (GSEA). e Based on RNAseq, mRNA counts of IL6 in the DMSO group, DC551040/HHT alone group, and combination group, data are represented as mean ± SEM, n = 3 per group. f Detection of mRNA expression level of MV-4-11 after IL6 knockdown based on qPCR experiment, data are represented as mean ± SEM, n = 3 per group. g Based on the cytotoxicity experiment, the cell viability after treatment with DC551040 was detected under the condition of IL6 knockdown or not, data are represented as mean ± SEM, n = 3 per group. The Significance of (f) was analyzed by equal variance two-tailed t test. *P < 0.05, **P < 0.01, ***P < 0.001
In previous studies, HHT is a protein translation inhibitor,10 exhibits pleiotropic anti-leukemic effects beyond translation inhibition, including modulation of the epigenome, cell cycle arrest, and induction of apoptosis in AML cells.11 These mechanisms are also reflected in our RNA-seq data, where HHT treatment altered cell cycle–related pathways such as G2/M checkpoint and mitotic spindle assembly (Fig. 9d).
Importantly, HHT also is reported to regulate immune and inflammatory responses by affecting key transcription factors, such as STAT3, NF-κB, and NRF2.9,11,12 Previous studies have shown that HHT can inhibit NF-κB activation13 and reduce expression of pro-inflammatory cytokines, such as IL-1, IL-6, and TNF-α, thereby preventing NF-κB-mediated inflammatory responses and suppressing tumor growth.14 Consistently, our RNA-seq data show HHT-mediated downregulation of IL-6 and other inflammatory-associated transcripts (Fig. 9d).
Furthermore, consistent with previous results, we observed that DC551040 upregulated IL6, while HHT downregulated IL6 in RNAseq data (Fig. 9e). To further validate the role of IL-6 in the observed synergy, we used two distinct siRNAs to knock down IL6 in MV-4-11 cells (Fig. 9f). Cell viability assays (CTG), demonstrated that IL6 knockdown significantly enhanced the sensitivity of MV-4-11 cells to DC551040, with a greater effect observed in cells with higher IL6 silencing efficiency (Fig. 9g).
Collectively, these results suggest that HHT mitigates DC551040-induced inflammatory activation, particularly via suppression of IL-6 signaling, thereby contributing to the synergistic anti-leukemic activity.
Discussion
Epigenetic drugs, including LSD1 inhibitors, represent promising therapeutic options in cancer treatment due to their pleiotropic effects on different aspects of tumor biology, leading to an overall impairment of cancer cells.20 LSD1 represents the first identified histone demethylase,21 and its dual role as a transcriptional corepressor or coactivator is determined by the divergent interaction profiles in various cell types.22 Its oncoprotein role in cancer stem cell regulation, differentiation, epithelial-mesenchymal transition (EMT), and tumor immunity suppression has been summarized.23,24,25 Recently, the tumor suppressor roles of LSD1 in breast cancer have been reported by inhibiting tripartite motif-containing protein 37 (TRIM37)26 or interacting with breast cancer metastasis suppressor 1 (BRMS1).27 However, the exact tumor suppressor roles or functions in AML or other tumors remain elusive. Exploring the pleiotropic effects at the molecular level may provide the rationale for combination therapies to achieve optimal clinical results.
Lead optimization was performed based on the compound 2-PCPA to improve selectivity against LSD2, MAO-A, and MAO-B, resulting in the characterization of DC551040 as a compound with an overall profile suitable for clinical development (Figs. 1–3 and Table 1). The in vivo efficacy of DC551040 was demonstrated in the Kasumi-1 and MV-4-11 xenograft models (Fig. 3), and DC551040 displayed favorable PK profiles in multiple species and was orally available (Supplementary Tables 3, 4). During the animal experiments, the animals maintained normal body weight, and no significant adverse reactions or treatment-related toxicities were observed. Based on these results, DC551040 has been developed as a clinical candidate for the treatment of AML and is currently in Phase I clinical trials (CTR20222026).
Transcriptomic and proteomic can be used as less-biased approaches to explore compensatory signaling networks related to adaptive responses.28,29,30 Proteomics analysis has identified dynamic reprogramming of the proteome as the primary or adaptive or acquired resistance mechanism of kinase inhibitors,31,32,33 while chemo transcriptomics has discovered that PI3Kα inhibitors could stimulate a robust compensatory increase in ER-dependent transcription that limits therapeutic efficacy.34 Temporal changes in the global proteomic or transcriptomic landscape can mirror the intended pharmacological effects of a compound, adverse mechanisms, or compensatory adaptations associated with the development of resistance. Integrated dynamic proteomics and transcriptomics were used to investigate gene and protein expression dynamics upon DC551040 treatment in an MV-4-11 xenograft model (Figs. 4, 5, 6). The study revealed that multiple inflammation-related pathways, including the key numbers STAT5, NF-κB, and AKT, were activated during DC551040 treatment, indicating the potential for adaptive resistance.
LSD1 plays a role as an epigenetic regulator of the inflammatory response, with different mechanisms depending on the cell type and immune signal.35 Ablation of LSD1 in cancer cells can activate type 1 interferon,36 upregulate immune response, and cytokine-signaling pathways through GSE1-protein reduction.37 However, in hematopoietic stem cells (HSCs) and cancer cells, LSD1 acts as a negative epigenetic regulator of inflammatory responses via H3K4me2 blocking.35,38 Although intrinsic and acquired LSD1 inhibitor resistance in SCLC emerges via epigenetic reprogramming-mediated perturbations in TEAD4-driven transcriptional programs,39 further exploration is needed to understand the mechanism of immune and inflammation activation induced by DC551040 treatment.
For complex signaling, most targeted based drugs are highly ineffective. Therefore, the network pharmacology strategy provides a framework for drug discovery that can perturb dysregulated networks pharmacologically.40 Repurposing marketed drug is a powerful approach that can rapidly address therapeutic needs.41 In addition, small-molecule drugs are highly promiscuous, allowing for the search of drugs that could oppose LSD1-induced network deregulation using computational approaches. CMAP, a network pharmacology database encompassing integrated disease-gene-drug interaction profiles,42,43 has been employed to address drug repurposing in AML.
By integrating transcriptomic and proteomic analyses, as well as a search of the CMAP database (Fig. 6a). The approved anti-leukemia drug HHT was found to suppress the DC551040-induced inflammation-related gene network (Fig. 6b). HHT is a translation elongation inhibitor that binds to the 80 s ribosome and interferes with chain elongation. It can also inactivate the IL6/JAK1/STAT344,45 and inhibit the NF-κB46 and STAT547 pathways, showing pleiotropic biological effects. Consistent with these observations, HHT was found to downregulate LSD1 inhibitor-induced NF-κB, STAT5, and AKT pathways. The combination of HHT with the LSD1 inhibitor DC551040 showed potent synergistic anti-AML activity both in vitro and in vivo (Figs. 7–9). Importantly, HHT was approved by the FDA in 2012 for the treatment of CML patients and is currently in clinical trials for AML.48,49 Our study provides the first report describing the potential application of rationally designed drug combination therapies could improve treatment outcomes and delay the resistance of LSD1 inhibitors in AML clinical trials.
In conclusion, we have developed DC551040 as a novel covalent LSD1 inhibitor with high potency and selectivity. DC551040 showed better occupancy of the LSD1 binding pocket and demonstrated significant anti-AML effect in both cellular and animal studies, with promising PK properties and good tolerability. It has progressed to Phase I clinical trials in patients with AML (CTR20222026). Moreover, by integrating comprehensive transcriptomic and proteomic analyses, as well as a search of the CMAP database, we identified the clinical AML treatment drug HHT, an approved anti-leukemia drug, which mimics the anti-transcriptional activation of inflammatory pathways. The combination of DC551040 and HHT exhibited synergistic antitumor efficacy and prolonged survival in MV-4-11 disseminated xenograft model mice. These findings pave the way for a potential combination therapeutic strategy for the treatment of AML using LSD1 inhibitors.
Materials and methods
LSD1 and LSD2 inhibitory activity assay
Recombinant human LSD1 (residues 172–852) and full-length LSD2 were sourced from Sino Biological Inc. and Active Motif, respectively, for enzyme inhibition testing. These assays were carried out using the Lance Ultra LSD1 Histone H3-Lysine 4 Demethylase Assay Kit (PerkinElmer) as recommended by the manufacturer. For each reaction, LSD1/2 assay components were first diluted in Tris buffer (50 mM Tris-HCl, 50 mM NaCl, 0.01% Tween-20, 1 mM DTT, 10 μM FAD, 10% glycerol, pH 9.0) and then dispensed into a 384-well microplate, with each well containing 4 μL of enzyme (2 nM LSD1 or 172 nM LSD2), 4 μL of 2.5 μM Bio-H3K4me2 (1–24 aa) substrate, and 2 μL of inhibitor. The mixture was incubated at room temperature for 1 h before 5 μL of detection reagent (Eu-labeled H3K4me0 antibody and ULight-Streptavidin) was added. Fluorescence intensity was quantified via an Envision instrument (PerkinElmer) in TR-FRET mode, with excitation set at 320 nm and emission at 665 nm. All experiments were run in triplicate to ensure reproducibility. Curve fitting was conducted using the “sigmoidal dose-response (variable slope)” equation in GraphPad Prism 7.0 software to calculate IC50 values.
A jump dilution approach was employed to characterize biochemical reversibility. LSD1 (200 nM) was pre-incubated with 300 nM compound DC551040 or 2 μM GSK2879552 for 1 h, after which the mixture was diluted 100-fold in the reaction buffer. The activity of LSD1 in the diluted samples was monitored at three time points: 1 h, 2 h, and 4 h post-dilution.
MAO-A and MAO-B inhibition assay
We procured MAO-A and MAO-B enzymes from Sigma-Aldrich, and subsequently conducted enzyme inhibition assays using the Promega MAO-Glo™ Assay Kit as per the manufacturer’s standard protocol. Luminescent signals generated from the assays were detected by the Envision system (PerkinElmer), and each test was carried out in triplicate to ensure reproducibility. For data analysis, IC50 values were computed with GraphPad Prism 7.0 software, where the sigmoidal dose-response (variable slope) model was adopted for curve fitting.
Cell cultures
The MV-4-11, Kasumi-1, MOLM-13, HL-60, RS-4-11, MOLT-4, Mino, Z-138, Raji, MM1S cell line (National Collection of Authenticated Cell Cultures, China) was maintained in Iscove’s modified Dulbecco’s medium (IMDM, 31980030, Gibco, for MV-4-11, HL-60, Z-138) or RPMI-1640 medium (1640, A1049101, Gibco, for Kasumi-1, MOLM-13, RS-4-11, MOLT-4, Mino, Raji, MM1S) supplemented with 10–20%(v/v) fetal bovine serum (FBS, WS500T, Ausbian) and 1% (v/v) penicillin-streptomycin solution (P/S, MA0110, Meilunbio) at 37 °C in a 5% CO2 incubator. Bone marrow-derived mononuclear cells (patient-derived cells, PDC) from the patient were cultured in X-VIVO 15 medium supplemented with 10% fetal bovine serum (FBS) and a panel of cytokines50,51,52,53 at 37 °C in a humidified atmosphere containing 5% CO₂, with cytokine concentrations selected in accordance with previous studies.
Immunoprecipitation
Chromatin fractions were subjected to immunoprecipitation assays. Briefly, cells were collected and lysed in lysis buffer supplemented with protease inhibitors for 30 min; subsequent centrifugation was performed, and the resulting supernatant was designated as the chromatin fraction. For immunoprecipitation, an aliquot of the lysate was first incubated with protein G (CST, Cat. No. 2729) to eliminate nonspecific protein binding. Thereafter, antibodies targeting LSD1 (CST, Cat. No. 2184) or GFI1B (Santa Cruz, Cat. sc-28356) were supplemented into the lysates, followed by overnight incubation at 4 °C. We used PBS to wash protein A/G magnetic beads (MCE, Cat. HY-K0202) and then blocked them with BSA (5 mg/mL) for 30 min. After continuing to wash with PBS, the magnetic beads were added to the lysate and incubated at 4 °C for 4 h. We continued to wash the magnetic beads with PBS, then added the loading buffer to the magnetic beads, kept them at 100 °C for 10 min, and then performed Western bolt detection. Detection was performed using LSD1 antibody (CST, Cat. 2184), GFI1B antibody (santa cruz, Cat. sc-28356) and coREST antibody (Sigma-Aldrich Cat. 07-455).
Cell proliferation assays
For viability assays, AML cell lines (MV-4-11, Kasumi-1, MOLM-13, HL-60, RS-4-11, MOLT-4, Mino, Z-138, Raji, MM1S) and AML PDC were plated in triplicate into 96-well plates supplemented with gradient doses of drugs, followed by 3 or 7 days of culture. Cell viability was assessed via the CellTiter 96® AQueous One Solution Cell Proliferation Assay (MTS assay; G3582, Promega) for AML cell lines and the CellTiter-Glo (CTG) assay (G7573, Promega) for AML PDC. Relative cell viability was calculated by normalizing the 490 nm absorbance values (MTS) and luciferase activity counts (CTG) of drug-treated groups to those of untreated control groups, with the latter set at 100%. All experiments were performed with 3 biologically independent replicates (n = 3). Statistical significance was evaluated using an unpaired two-tailed t-test with equal variance (*P < 0.05, **P < 0.01, ***P < 0.001). The half-maximal inhibitory concentration (IC₅₀), representing the drug concentration required to reduce cell growth by 50%, was computed using GraphPad Prism software (Version 7.0).
Apoptosis assay
MV-4-11 cells were seeded into 6-well plates at a density of 3 × 10⁵ cells per well. Following 1 h of pre-incubation, the cells were treated with serial concentrations of the target drug, with an equal volume of 0.1% DMSO serving as the negative control. After 72 h of drug exposure, cells were harvested and centrifuged at 1500 rpm for 5 min, and the supernatant was removed. Cell pellets were rinsed twice with PBS (1500 rpm, 5 min per centrifugation), then resuspended in 500 μL of binding buffer. Thereafter, cells were stained with Annexin V-FITC/PI staining kit (KGA106, Keygentec, Nanjing, China) and analyzed on a CytoFLEX flow cytometer (Beckman Coulter, Inc.). Green fluorescence signals of Annexin V-FITC were captured via the FL1 channel, while red fluorescence signals of PI were detected through the FL3 channel. After the data collection was completed, the data in FSC format were exported and further analyzed by FlowJo software to calculate the percentage of apoptosis.
Caspase 3/7 activity assay
MV-4-11 cells were plated in white opaque 96-well microplates (PerkinElmer, 96-well Culture Plate) at a volume of 80 μL medium per well. Following overnight incubation, cells were treated with either 0.2% DMSO (vehicle control) or serial 1000-fold dilutions of test compounds prepared from high-concentration DMSO stock solutions (resulting in a final DMSO concentration of 0.2%) for 24 h. Subsequently, Caspase-3/7 activity was assessed using the Caspase-Glo Assay Kit (Promega) after a 0.5 h incubation period.
RT-qPCR detection of CD86 expression
MV-4-11 cells were seeded into 12-well plates at a density of 1 × 10⁶ cells per well and treated with gradient concentrations of the target drug. After 24 h of treatment, total RNA was extracted from the cells, which was subsequently reverse-transcribed into complementary DNA (cDNA). Specific primers targeting CD86 (Forward primer: 5′-CAGAAGCAGCCAAAATGGAT-3′; Reverse primer: 5′-TCAGAGGAGCAGCACCAGA-3′) and the housekeeping gene ACTB (Forward primer: 5′-AGAGCTACGAGCTGCCTGAC-3′; Reverse primer: 5′- AGCACTGTGTTGGCGTACAG-3′) were used to perform PCR amplification, with SYBR Green dye incorporated into the reaction system—this dye intercalates into double-stranded DNA to emit fluorescence, enabling real-time monitoring of the fluorescent signal generated by amplicons in each PCR cycle. A fluorescence amplification curve was generated with the accumulation of PCR products, allowing for quantitative analysis of the initial template. The relative expression level of CD86 was normalized to the geometric mean of ACTB (to minimize variability in overall expression levels) and calculated using the 2−ΔΔCt method.
Wright’s-Giemsa staining
The state of MV-4-11 cells treated with the drug was detected by the Wright-Giemsa staining method, the state of the cytoplasm and nucleus of the cells was observed, and cell differentiation was judged by the change in cell morphology. details as follows: Cells were cultured and subsequently subjected to staining using the Fast Wright’s-Giemsa Stain Kit (60529ES01, Yeaseen) per the manufacturer-recommended protocol: briefly, a small aliquot of the cell suspension was smeared evenly onto clean microscope slides, air-dried completely, and fixed with absolute methanol for 5 min. slides were then covered with working Wright-Giemsa solution for 5 min of staining, rinsed with distilled water, and air-dried at room temperature. Stained samples were observed and imaged under an Olympus IX73 microscope at 600× magnification.
Western blotting
MV-4-11 cells received two treatment protocols: DMSO or DC551040 (0.1 μM, 0.5 μM, 2 μM) for 1, 3, 5, 7 days; or DMSO, homoharringtonine (2.5 nM, 10 nM) and DC551040 (1 μM, 5 μM) for 12 h. Whole-cell lysates were prepared with 1% SDS-containing lysis buffer (SDS: A500228, Sangon Biotech), and protein concentration was determined by BCA kit (P0010, Beyotime Biotechnology). 20 μg of protein lysate per sample was separated by 15% SDS-PAGE and transferred to NC membranes (10600021, Cytiva). Membranes were blocked with 3% BSA (A600332, Sangon Biotech) in TBST (20 mM Tris pH 7.6, 150 mM NaCl, 0.1% Tween-20), incubated overnight at 4 °C with p-STAT3 (9145, CST), STAT3 (30835, CST), ACTB (4970, CST), H3K4me2 (PTM-641, PTM BIO) or histone H3 (M1309-1, Huabio) antibodies, washed thrice with TBST, then incubated with HRP-conjugated secondary antibodies (115-035-003, Jackson ImmunoResearch) for 1 h at room temperature. Target proteins were detected via ECL kit (MAD186, Meilunbio) and visualized on Tanon 4600 Imaging System.
Nu/Nu mouse subcutaneous xenograft model evaluation
All animal experimental procedures were conducted in compliance with relevant regulatory requirements and established ethical guidelines, and were reviewed and approved by the Institutional Animal Care and Use Committee (IACUC) of the Shanghai Institute of Materia Medica, Chinese Academy of Sciences (approval number: 2021-12-LJ-124). Nu/Nu mice were subcutaneously inoculated in the right flank with 5 × 10⁶ MV-4-11 or Kasumi-1 cells suspended in 0.2 mL of 1:1 PBS/Matrigel mixture. Drug treatment was initiated when the mean tumor volume reached ~100–200 mm³, with each group comprising 6 tumor-bearing mice. Mice were administered either vehicle control or the drug; tumor dimensions were measured twice weekly using a vernier caliper, and tumor volume (V, in mm³) was calculated via the formula: V = 0.5 × a × b², where a represents the longest tumor diameter and b represents the shortest tumor diameter. Mice were weighed daily. Relative tumor volume (RTV) was computed using the equation: RTV = TVt/TV0, in which TVt denotes the tumor volume of individual mice at time t and TV0 denotes the initial tumor volume at the initiation of treatment (time 0). The relative tumor growth rate (T/C, %) was calculated as follows: T/C (%) = (RTVt/RTVveh) × 100%, where RTVt is the mean RTV of drug-treated mice at time t and RTVveh is the mean RTV of vehicle-treated mice at time t. The experiment was terminated when any tumor reached a volume of 2000 mm³.
B-NDG mouse disseminated xenograft acute leukemia model
Animal experimental protocols were approved by the IACUC of Shanghai Institute of Materia Medica, Chinese Academy of Sciences (approval no. 2021-12-LJ-124). B-NDG mice received tail vein injection of 1 × 10⁷ MV-4-11 cells for intravenous engraftment. At 5 days post engraftment, mice were treated with drugs or vehicle for 4 weeks, with daily survival monitoring. Animals were humanely euthanized when showing obvious illness per IACUC ethical protocols, and the maximum permitted weight loss was 15%. Statistical significance of survival was assessed by the log-rank test.
Patient-derived tumor xenograft acute leukemia model
Animal protocols were approved by the IACUC of Shanghai Institute of Materia Medica, Chinese Academy of Sciences (IACUC approval: 2021-12-LJ-124). B-NDG mice received tail vein injection of 1 × 10⁷ primary AML cells (Sample No. AML21001-X) for intravenous engraftment. Mice were treated with drugs or vehicle for 3 weeks starting at 60 days post engraftment. To evaluate the residual leukemia burden, human CD33+ in mouse bone marrow will be quantified by flow cytometry after treatment completion.
Immunohistochemical analysis of CD86 protein in tumor tissue sections
Mice with subcutaneous transplanted tumors (MV-4-11) were treated with drugs for 22 days. The tumor tissues were isolated at the end of the experiment 2 h after administration on the last day, and then placed in 4% paraformaldehyde fixation buffer overnight, embedded in paraffin, and sectioned into sections with the largest diameter of each tumor tissue, with a section thickness of 4 μm. All sections were subjected to immunohistochemistry. The sections were first baked in a constant temperature oven at 58 °C for 30 min and then subjected to conventional HE staining and immunohistochemical analysis of CD86 protein. The Akoya vectra imaging and analysis system was used to analyze the CD86 protein positivity rate in immunohistochemical sections of tumor tissue. Statistical analyses were performed using Student’s t test, and P < 0.05 was regarded as statistically significant.
RNA-seq data analysis
After removing adaptor contaminants, polyA and polyC segments, sequencing reads were aligned to the UCSC hg38 human reference assembly using HISAT2. Aligned files were compressed, sorted, and subjected to gene expression quantification with HTSeq-count. Differentially expressed genes were identified via DESeq2 (R package) based on log2 fold change ≥1/≤−1 (two-fold change) and FDR-adjusted P < 0.05 (5% cutoff). Enrichment analysis of these DEGs was conducted on Metascape (www.metascape.org) using Fisher’s exact test, with pathway annotations derived from KEGG, GO and Reactome databases.
Sample preparation for mass spectrometry
The tissues from nude mice established with subcutaneously transplanted tumors were fully minced by a tissue grinder (Tissue lyser-24, Jingxin, Shanghai, China) and washed with ice-cold phosphate-buffered saline (PBS) twice. Then, the minced tissues were subjected to protein extraction lysis buffer (8 M urea, 50 mM NAM, 3 µM TSA, 2 mM EDTA, 5 mM dithiothreitol, and 1% Protease Inhibitor Cocktail II) and sonicated in an ultrasonic processor (XM-26A, Xiaomei Ultrasound Instrument (Kunshan) Co., Ltd.) for 15 min (25 s on and 3 s off) at 4 °C. Lysates were centrifuged at 16,000 × g for 10 min at 4 °C. The lysate supernatant was collected as the whole-tissue extract. After quantification by a BCA kit, 100 μg of mixed proteins were reduced with 10 mM dithiothreitol at 56 °C for 30 min and alkylated with 20 mM iodoacetamide at room temperature in darkness. Next, 100 mM NH4HCO3 was added to reduce the urea concentration to 2 M. Proteins were digested by incubating with trypsin (ratio 1:50) at 37 °C for 16 h. The peptide mixture was desalted by C18 tips.
LC–MS/MS analysis
Sample analysis was carried out on an EASY-nLC 1200 UHPLC system (ThermoFisher Scientific) coupled to a Q Exactive HF-X mass spectrometer (ThermoFisher Scientific). Two micrograms of peptides were separated via a capillary C18 column (75 µm ID × 25 cm length, 1.9 µm particle size, Dr. Maisch GmbH) in a 120 min gradient from 8 to 90% solvent B (A, 0.1% formic acid; B, 80% acetonitrile in 0.1% formic acid). Total scanning was acquired from m/z 350–1200 m/z at a resolution of 60,000 and the AGC target was 3E6. The resolution for the MS/MS analyses was 15,000 and the AGC number was 1e5. The 20 most intensive ions were fragmented with 28% normalized collision energy. Dynamic exclusion was set to 30.0 s.
Database search of the LC–MS/MS data
The raw data were searched against the reviewed UniProt human database (released in Jan 2020) using MaxQuant (version 1.6.15.0). Trypsin/P was selected as the digestive enzyme, and two missed cleavage sites were allowed. In all searches, cysteine carbamidomethylation was established as a fixed modification, and methionine oxidation and acetylation of the N-terminus were designated as variable modifications. Perseus software (version 1.6.2.3) was used for data analysis with LFQ values from MaxQuant results.
Bioinformatic analysis of the proteome data
First, a two-tailed Student’s t test was employed to identify differentially expressed proteins, where the inclusion criteria were an adjusted P < 0.05 and a fold change ≥1 or ≤−1 (the fold change was calculated as the log2 ratio of the median protein abundance between the two groups in each individual comparison). To facilitate intuitive interpretation, these differentially expressed proteins were visualized through volcano plots (generated by the ggplot2 package) and heatmaps (generated by the heatmap package), both of which were constructed in R. Furthermore, we performed Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis by applying the hypergeometric test within the clusterProfiler package of R. GSEA of the KEGG pathway was implemented using the clusterProfiler package in R.
Calculation of synergy scores
Drawing on prior research, the synergy scores of drug combinations were calculated using the Zero Interaction Potency (ZIP) model,54 a method that references three commonly adopted reference models: Highest Single Agent (HSA), Bliss independence, and Loewe additivity. This approach characterizes drug-drug interaction relationships by comparing shifts in the potency of dose–response curves between monotherapies and their corresponding drug combinations. This method is suitable for calculating multiple doses of large-scale drug combination experiments, with low false positives, and it is used by online software(https://synergyfinder.fimm.fi/) for analysis55 Generally, a ZIP synergy score >5 indicates synergy, and a score >10 signifies strong synergy.
Remaining Materials and Methods are described in SI Appendix.
Data availability
All data supporting the findings of this study are available in the main text and supplementary materials. RNA-sequencing data are available in the GSA repository under accession numbers CRA036092 (https://ngdc.cncb.ac.cn/gsa/browse/CRA036092). The atomic coordinates and structure factors have been deposited into the Protein Data Bank with accession code 6K3E (LSD1/CoREST in complex with DC551040).
References
Huang, J. et al. p53 is regulated by the lysine demethylase LSD1. Nature 449, 105–108 (2007).
Chen, J. et al. Knocking down LSD1 inhibits the stemness features of colorectal cancer stem cells. Braz. J. Med. Biol. Res. 53, e9230 (2020).
Ding, J. et al. The influence and mechanisms of lysine-specific demethylase 1 (LSD1) on invasion and metastasis of colon cancer cells. Zhonghua Yi Xue Za Zhi 97, 743–748 (2017).
Ferrari-Amorotti, G. et al. Inhibiting interactions of lysine demethylase LSD1 with snail/slug blocks cancer cell invasion. Cancer Res. 73, 235–245 (2013).
Lv, T. et al. Over-expression of LSD1 promotes proliferation, migration and invasion in non-small cell lung cancer. PLoS ONE 7, e35065 (2012).
Hayami, S. et al. Overexpression of LSD1 contributes to human carcinogenesis through chromatin regulation in various cancers. Int. J. Cancer 128, 574–586 (2011).
Verigos, J. et al. The histone demethylase LSD1/KappaDM1A mediates chemoresistance in breast cancer via regulation of a stem cell program. Cancers (Basel) 11, 1585 (2019).
Agboyibor, C. et al. LSD1 as a biomarker and the outcome of its inhibitors in the clinical trial: the therapy opportunity in tumor. J. Oncol. 2021, 5512524 (2021).
Harris, W. J. et al. The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21, 473–487 (2012).
Lokken, A. A. & Zeleznik-Le, N. J. Breaking the LSD1/KDM1A addiction: therapeutic targeting of the epigenetic modifier in AML. Cancer Cell 21, 451–453 (2012).
Salamero, O. et al. First-in-human phase I study of Iadademstat (ORY-1001): a first-in-class lysine-specific histone demethylase 1A inhibitor, in relapsed or refractory acute myeloid leukemia. J. Clin. Oncol. 38, 4260–4273 (2020).
Lee, M. G., Wynder, C., Cooch, N. & Shiekhattar, R. An essential role for CoREST in nucleosomal histone 3 lysine 4 demethylation. Nature 437, 432–435 (2005).
Metzger, E. et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437, 436–439 (2005).
Wang, J. et al. Opposing LSD1 complexes function in developmental gene activation and repression programmes. Nature 446, 882–887 (2007).
Wass, M. et al. A proof of concept phase I/II pilot trial of LSD1 inhibition by tranylcypromine combined with ATRA in refractory/relapsed AML patients not eligible for intensive therapy. Leukemia 35, 701–711 (2021).
Salamero, O. et al. Iadademstat in combination with azacitidine generates robust and long lasting responses in AML patients (ALICE trial). Blood 138, 3376 (2021).
Ishikawa, Y. et al. A novel LSD1 inhibitor T-3775440 disrupts GFI1B-containing complex leading to transdifferentiation and impaired growth of AML cells. Mol. Cancer Ther. 16, 273–284 (2017).
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Cao, W. et al. Homoharringtonine induces apoptosis and inhibits STAT3 via IL-6/JAK1/STAT3 signal pathway in Gefitinib-resistant lung cancer cells. Sci. Rep. 5, 8477 (2015).
Sigalotti, L. et al. Epigenetic drugs as pleiotropic agents in cancer treatment: biomolecular aspects and clinical applications. J. Cell. Physiol. 212, 330–344 (2007).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Karakaidos, P., Verigos, J. & Magklara, A. LSD1/KDM1A, a gate-keeper of cancer stemness and a promising therapeutic target. Cancers 11, 1821 (2019).
Martinez-Gamero, C., Malla, S. & Aguilo, F. LSD1: expanding functions in stem cells and differentiation. Cells 10, 3252 (2021).
Zhang, X. et al. Therapeutic potential of targeting LSD1/KDM1A in cancers. Pharmacol. Res. 175, 105958 (2022).
Kim, D., Kim, K. I. & Baek, S. H. Roles of lysine-specific demethylase 1 (LSD1) in homeostasis and diseases. J. Biomed. Sci. 28, 41 (2021).
Hu, X. et al. LSD1 suppresses invasion, migration and metastasis of luminal breast cancer cells via activation of GATA3 and repression of TRIM37 expression. Oncogene 38, 7017–7034 (2019).
Qiu, R. et al. BRMS1 coordinates with LSD1 and suppresses breast cancer cell metastasis. Am. J. Cancer Res. 8, 2030–2045 (2018).
Alaoui-Jamali, M. A., Dupre, I. & Qiang, H. Prediction of drug sensitivity and drug resistance in cancer by transcriptional and proteomic profiling. Drug Resist. Updat. 7, 245–255 (2004).
Vasan, N., Baselga, J. & Hyman, D. M. A view on drug resistance in cancer. Nature 575, 299–309 (2019).
Meissner, F., Geddes-McAlister, J., Mann, M. & Bantscheff, M. The emerging role of mass spectrometry-based proteomics in drug discovery. Nat. Rev. Drug Discov. 21, 637–654 (2022).
Sos, M. L. et al. Oncogene mimicry as a mechanism of primary resistance to BRAF inhibitors. Cell Rep. 8, 1037–1048 (2014).
Duncan, J. S. et al. Dynamic reprogramming of the kinome in response to targeted MEK inhibition in triple-negative breast cancer. Cell 149, 307–321 (2012).
Koch, H., Busto, M. E., Kramer, K., Medard, G. & Kuster, B. Chemical proteomics uncovers EPHA2 as a mechanism of acquired resistance to small molecule EGFR kinase inhibition. J. Proteome Res. 14, 2617–2625 (2015).
Toska, E. et al. PI3K pathway regulates ER-dependent transcription in breast cancer through the epigenetic regulator KMT2D. Science 355, 1324–1330 (2017).
Janzer, A. et al. Lysine-specific demethylase 1 (LSD1) and histone deacetylase 1 (HDAC1) synergistically repress proinflammatory cytokines and classical complement pathway components. Biochem. Biophys. Res. Commun. 421, 665–670 (2012).
Sheng, W. et al. LSD1 Ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563 e19 (2018).
Nicosia, L. et al. Pharmacological inhibition of LSD1 triggers myeloid differentiation by targeting GSE1 oncogenic functions in AML. Oncogene 41, 878–894 (2022).
Wang, Q. et al. Host-secreted antimicrobial peptide enforces symbiotic selectivity in Medicago truncatula. Proc. Natl. Acad. Sci. USA 114, 6854–6859 (2017).
Yan, W. et al. Intrinsic and acquired drug resistance to LSD1 inhibitors in small cell lung cancer occurs through a TEAD4-driven transcriptional state. Mol. Oncol. 16, 1309–1328 (2022).
Hopkins, A. L. Network pharmacology: the next paradigm in drug discovery. Nat. Chem. Biol. 4, 682–690 (2008).
Chong, C. R. & Sullivan, D. J. Jr. New uses for old drugs. Nature 448, 645–646 (2007).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Subramanian, A. et al. A next generation connectivity map: L1000 platform and the first 1,000,000 profiles. Cell 171, 1437–1452 e17 (2017).
Qu, M., Li, J. & Yuan, L. Uncovering the action mechanism of homoharringtonine against colorectal cancer by using network pharmacology and experimental evaluation. Bioengineered 12, 12940–12953 (2021).
Wang, F. et al. Homoharringtonine synergizes with quizartinib in FLT3-ITD acute myeloid leukemia by targeting FLT3-AKT-c-Myc pathway. Biochem. Pharmacol. 188, 114538 (2021).
Chen, X. J. et al. Homoharringtonine deregulates MYC transcriptional expression by directly binding NF-kappaB repressing factor. Proc. Natl. Acad. Sci. USA 116, 2220–2225 (2019).
Tong, H., Ren, Y., Zhang, F. & Jin, J. Homoharringtonine affects the JAK2-STAT5 signal pathway through alteration of protein tyrosine kinase phosphorylation in acute myeloid leukemia cells. Eur. J. Haematol. 81, 259–266 (2008).
Wang, F. et al. Low dose of homoharringtonine and cytarabine-based priming induction regimens for patients with de novo acute myeloid leukemia and high-risk myelodysplastic syndrome aged over 70 years. Hematology 26, 1040–1045 (2021).
Lam, S. S. et al. Homoharringtonine (omacetaxine mepesuccinate) as an adjunct for FLT3-ITD acute myeloid leukemia. Sci. Transl. Med. 8, 359ra129 (2016).
Bibikova, E. et al. TNF-mediated inflammation represses GATA1 and activates p38 MAP kinase in RPS19-deficient hematopoietic progenitors. Blood 124, 3791–3798 (2014).
Hardwick, N., Chan, L., Ingram, W., Mufti, G. & Farzaneh, F. Lytic activity against primary AML cells is stimulated in vitro by an autologous whole cell vaccine expressing IL-2 and CD80. Cancer immunol. Immunother. 59, 379–388 (2010).
Foss, B., Ulvestad, E. & Bruserud, Ø Platelet-derived growth factor (PDGF) in human acute myelogenous leukemia: PDGF receptor expression, endogenous PDGF release and responsiveness to exogenous PDGF isoforms by in vitro cultured acute myelogenous leukemia blasts. Eur. J. Haematol. 67, 267–278 (2001).
Melgar, K. et al. Overcoming adaptive therapy resistance in AML by targeting immune response pathways. Sci. Transl. Med. 11, eaaw8828 (2019).
Yadav, B., Wennerberg, K., Aittokallio, T. & Tang, J. Searching for drug synergy in complex dose–response landscapes using an interaction potency model. Comput. Struct. Biotechnol. J. 13, 504–513 (2015).
Ianevski, A., Giri, A. K. & Aittokallio, T. SynergyFinder 2.0: visual analytics of multi-drug combination synergies. Nucleic Acids Res. 48, W488–W493 (2020).
Acknowledgements
We gratefully acknowledge the National Natural Science Foundation of China (82322063 to J.W.; 82130105, 22337003, and 82121005 to H.L.; 22277125, 92253306, and 82521004 to H.H.; 82404655 to H.W.; and 92253306 to J.L.;8227366 to C.L.), National Key R&D Program of China (2022YFA1302900 to H.L.), Program of Shanghai Academic Research Leader (23XD1460300 to J.W.), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB1060000 to H.H.), Guangdong High-level new R&D Institute (2019B090904008 to J.L.). Guangdong High-level Innovative Research Institute (2021B0909050003 to J.L.), and Lingang Laboratory (LG-GG-202204-02 to J.W. and LG202103-02-06 to C.L.). the support of the Sanofi scholarship program, Shandong Laboratory Program (SYS202205 to J.L. and H.H.), Taishan Scholars Program (tstp0648 to J.L. and tstp20240845 to H.H.), Shanghai Science and Technology Development Funds (24YF2755300 to H.W), Research Funds of Hangzhou Institute for Advanced Study, UCAS (to H.H.), Innovative Drug Research and Development National Science and Technology Major Project(2025ZD1800700 to J.L.; 2025ZD1804100 to H.W.), the Institutes for Drug Discovery and Development, Chinese Academy of Sciences (CASIMM0120225007 and SIMM0120231004 to C.L.), Shanghai Institute of Materia Medica, Chinese Academy of Sciences (SIMM0120232003 to C.L.).
Author information
Authors and Affiliations
Contributions
J.W., H.W., R.D., C.L., M.S., S.W., G.L., J.S., Y.W, B.W. performed the experiments; S.W. synthesized all of the compounds with the guidance of H.L. and J.W.; H.W. and M.S. conducted the screening of compounds; R.D. and Y.Y. performed the cell assay; W.K., and X.H. performed the animal experiment; R.D. performed the in vitro activity test; F.G., W.Z., and Y.S. assisted in the synthesis of some LSD1 inhibitors; J.W., Y.Z., H.H., J.L., and H.L. provided valuable suggestions on experimental design; Y.Z. and G.L. examined the characterization data of the compounds; J.W., Y.Z., H.H., J.L., and H.L. were responsible for the study concept and design; Y.Z., H.H., J.L., and H.L. supervised the project; and J.W., H.W., R.D. Y.Z., H.H., J.L., and H.L. drafted the manuscript. All of the authors edited the manuscript and contributed intellectually. All authors have read and approved the article.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, J., Wang, H., Du, R. et al. Potent and selective LSD1 inhibitor DC551040 reveals a promising combination therapy for AML with insight into epigenetic dysregulation. Sig Transduct Target Ther 11, 108 (2026). https://doi.org/10.1038/s41392-026-02637-0
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41392-026-02637-0