Abstract
Castration-resistant prostate cancer (CRPC) is one of the most prevalent cancers in men. The new generation androgen receptor (AR) inhibitor enzalutamide can improve the therapeutic effectiveness of patients with CRPC. However, these patients eventually develop acquired enzalutamide resistance (ENZR), and the mechanisms underlying resistance are not well understood. Wilms’ tumor 1-associating protein (WTAP) plays an important role in m6A modification and has been reported as an oncogene in various cancers. Here, we utilized a tissue microarray and collected tissues from prostate cancer (PCa) patients to detect WTAP expression, and found that WTAP is upregulated in PCa. Meanwhile, WTAP overexpression promotes cell proliferation and accelerates tumor growth through colony formation assays and the establishment of a subcutaneous xenograft model in vivo. These findings establish the tumor promoter role of WTAP in prostatic tumorigenesis. Furthermore, we verified that WTAP is a novel responsive gene of AR via promoter activity and chromatin immunoprecipitation (ChIP) assays. Importantly, we uncovered that WTAP is upregulated in ENZR cells, and WTAP knockdown inhibited the proliferation of ENZR cells. Mechanistically, ubiquitin-specific protease (USP7) enhanced the stability of WTAP by the ubiquitin-proteasome pathway in ENZR cells, thereby WTAP increases promote AKT signaling through an m6A-mediated way, and an AKT inhibitor can abolish the pro-resistance phenotype mediated by WTAP. Together, these findings suggest that WTAP plays a key role in ENZR development of PCa cells, and WTAP may be a potential treatment target for ENZR tumors.

In this manuscript, we utilized a tissue microarray and collected tissues from prostate cancer (PCa) patients to detect WTAP expression, and found that WTAP is upregulated in PCa. Meanwhile, WTAP overexpression promotes cell proliferation and accelerates tumor growth through colony formation assays and the establishment of a subcutaneous xenograft model in vivo. These findings establish the tumor promoter role of WTAP in prostatic tumorigenesis. Furthermore, we verified that WTAP is a novel responsive gene of AR via promoter activity and chromatin immunoprecipitation (ChIP) assays. Importantly, we uncovered that WTAP is upregulated in ENZR cells, and WTAP knockdown inhibited the proliferation of ENZR cells. Mechanistically, ubiquitin-specific protease (USP7) enhanced the stability of WTAP by the ubiquitin-proteasome pathway in ENZR cells, thereby WTAP increases promote AKT signaling through an m6A-mediated way, and an AKT inhibitor can abolish the pro-resistance phenotype mediated by WTAP. Together, these findings suggest that WTAP plays a key role in ENZR development of PCa cells, and WTAP may be a potential treatment target for ENZR tumors.
Similar content being viewed by others
Introduction
Prostate cancer (PCa) is one of the most prevalent cancers and is ranked as the second most lethal threat of malignant tumors in men [1]. Androgen receptor (AR) plays a central role in the PCa progression and serves as a decisive part in the treatment of PCa patients. Androgen deprivation therapy (ADT) as a mainstay in treating advanced PCa has been shown to be effective. However, most PCa patients will eventually experience relapse and develop into castration resistant prostate cancer (CRPC) [2]. Existing therapies for CRPC patients, including AR antagonist enzalutamide, may improve the survival of patients with CRPC at an earlier stage, but eventually acquired drug resistance is inevitable [3]. So far, various mechanisms underlying enzalutamide resistance have been reported, such as AR gene amplification, AR splice variants, and AR signaling activated by ligand-independent ways [4,5,6]. Particularly, there exist multiple AR bypass ways that escape the inhibition of enzalutamide and sustain tumor progression. Among the mechanisms skirting AR signaling, the AKT signaling pathway sparks interest in exploring AKT inhibitors as a therapeutic strategy for PCa [7]. Actually, a combination of AKT-specific inhibitors plus AR signaling inhibitors has shown promising prospects in clinical studies [8, 9]. However, further investigation is needed to explore potential biomarkers underlying this combination to treat enzalutamide resistance preferably.
Wilms’ tumor 1-associating protein (WTAP) serves as a key regulatory component of the m6A methyltransferase complex and facilitates METTL3/METTL14 to interact with target RNAs [10, 11]. WTAP plays important functions in physiological processes, such as m6A methylation and mRNA stabilization. In recent years, a large amount researches have shown that WTAP plays a crucial role in various tumors through an m6A modification-dependent or independent way. In acute myeloid leukemia (AML), WTAP is upregulated and regulates the half-life of MYC mRNA through an m6A modification-dependent manner [12]. In diffuse large B-cell (DLBCL), WTAP is overexpressed and promotes cell growth and represses apoptosis by Hsp90, which enhances the stabilization of WTAP [13]. Besides in AML and in DLBCL, the functions of WTAP have been demonstrated in colorectal cancer (CRC) [14, 15], renal cell carcinoma (RCC) [16], and breast cancer [17, 18]. However, little is known about the role of WTAP in PCa.
In our previous study, we have clarified the tumor suppressor role of FTO in PCa and found that the function of FTO may be related to AR status in PCa cells [19]. In this study, we elucidate that WTAP is regulated by AR signaling through transcriptional regulation. WTAP functions as an oncogenic factor not only in enzalutamide-sensitive PCa cells but also in enzalutamide-resistant (ENZR) PCa cells. Meanwhile, the regulatory mechanisms of WTAP expression were variable in the progression of enzalutamide treatment, and WTAP is eventually upregulated in enzalutamide-resistant cells, which was independent of transcriptional regulation of AR. Furthermore, we revealed that WTAP stability was enhanced by ubiquitin-specific protease 7 (USP7)-mediated deubiquitination in ENZR cells. Importantly, WTAP promoted enzalutamide resistance through activating AKT signaling via enhancing m6A mRNA methylation of AKT. Our study highlights the crucial role of WTAP in enzalutamide alone or in combination with AKT inhibitor treatment in enzalutamide- resistant cells, and WTAP may be a predictive biomarker for treating ENZR tumors with enzalutamide or AKT inhibitors.
Material and methods
Statistics of tissue microarrays and collection of prostate cancer samples
The tissue microarray (TAM) (HProA120Su01) containing 60 paired PCa tumor tissues and adjacent normal tissues with histological grade was used to investigate the expression of WTAP in the tumor and normal tissues, which was purchased from Outdo Biotech Company (Shanghai, China).
From the Department of Urology of Shandong Provincial Hospital Affiliated to Shandong First Medical University, we obtained paired tissue samples from 5 untreated prostate cancer patients, including tumor tissues and their corresponding normal tissues. We carried out research in conformity with the Helsinki Declaration, and the study was approved by the Ethics Committee of Shandong Provincial Hospital of Shandong First Medical University (R202406110275). All the patients participating in the research have signed the informed consent form. All collected tissue samples have been confirmed by pathological examination and first stored in liquid nitrogen for subsequent use.
Immunohistochemical staining
The section of TAM was successively stained with primary anti-WTAP antibody (ab195380, Abcam, U.K.). We quantified according to the staining intensity mentioned in previously published literature by two pathologists to evaluate the stained results independently. The staining intensity ranged from negative to strong, and was assigned a score ranging from 0 to 3, 0 signifying negative, 1 signifying weak, 2 signifying moderate, and 3 signifying strong. The proportion of the positively stained region was evaluated according to a proportion ranging from 0 to 100%. In the end, the results of the staining evaluation were multiplied by proportion scores.
Cell culture and transfection
The prostate cancer cell lines used in this research included human immortalized prostate epithelial cell lines RWPE1, C4-2B cells, PC-3 cells, LNCaP cells, 22Rv1 cells, and DU 145 cells, which were purchased from KeyGEN Biotech (Jiangsu, China). All cell lines were authenticated by STR profiling and confirmed to be free of mycoplasma contamination prior to experimental use.PC-3 cells, LNCaP cells, C4-2B cells, and 22Rv1 cells were maintained in 10% serum-containing complete RPMI-1640 medium (Cellmax, Beijing, China). RWPE1 cells were sustained in keratinocyte SFM medium containing KGS (ScienCell, California, USA). DU 145 cells were nurtured in complete MEM medium (Genscript Biotechnology, Jiangsu, China). The enzalutamide-resistant cell line (C4-2B-ENZR) was established in accordance with the research methods published previously [20]. The C4-2B-ENZR cell line was obtained by continuously treating C4-2B cells with 20 μM enzalutamide (HY-70002, MedChemExpress, USA) for more than 4 months.
Drugs used in cell experiments: Metribolone(R1881, F2220171, Aladdin, Shanghai, China); P005091(P5091, HY-15667, MedChemExpress, USA); 3-Deazaadenosine(DAA, HY-W013332, MedChemExpress, USA); STM2457, (HY-134836, MedChemExpress, USA); Perifosine (PRFS, HY-50909, MedChemExpress, USA).
The siRNAs targeting human WTAP, AR, and METTL3 were synthesized from Sangon Biotech (Shanghai, China). The siRNA sequences targeting human WTAP were 5′-GUACACAGAUCUUAACUCU-3′ and 5′-AAUGAAAGGUGAACUGGAA-3′. The siRNA sequences for silencing METTL3 were 5′-GGUAUCAUCUCUAAACCUA-3′ and 5′- GAUGAAAUCUAUGGCAUGA-3′. The siRNA sequences targeting AR were 5′-CAAGGGAGGUUACACCAAA-3′. All siRNAs were introduced into cells by using Lipofectamine® RNAiMAX Reagent (13778-150 Invitrogen, USA).
Plasmids and lentivirus transduction
WTAP (vector: pLVX-EF1α or pcDNA 3.1), AR (vector: pLVX-EF1α), and USP7 mutants (vector: PcDNA 3.1) cDNA expression was synthesized by Sangon Biotech. HA-USP7, Myc-Ubiquitin, and HA-Ubiquitin were purchased from Miaoling Plasmid (Wuhan, China). ShRNAs targeting WTAP were inserted into pLKO.1-TRC-WRPE-puro plasmid. Sequences are listed below:
shWTAP-1: GTACACAGATCTTAACTCTTTCAAGAGAAGAGTTAAGATCTGTGTACTTTT shWTAP-2: AATGAAAGGTGAACTGGAATTCAAGAGATTCCAGTTCACCTTTCATTTTTT. Lentiviruses were packaged in 293T cells by co-transfecting shRNA or pLVX-EF1α-WTAP/AR with packing vectors. The lentivirus particles were collected at 48 h and 72 h and then infected the proper cells under polybrene after being filtered. The cells were selected for 1 week using 1–2 μg/mL puromycin.
Animal experiments
The animal assays were officially permitted by the Animal Ethics Committee of Shandong Provincial Hospital of Shandong First Medical University (W202406110553). 2 × 106 cells (PC-3 cells) were administered into the right or left flank of the 4-week-old male nude mice (n = 7). Volumes of tumors (V = 0.5 × length × width2) were estimated every 6 days. After 4 weeks, the mice were devoted, and the tumors were weighed and isolated.
Co-Immunoprecipitation (Co-IP), mass spectrometry, and Western blot assay
The protocol was carried out following our previous study [19]. Cells or tissues were harvested and lysed with RIPA buffer containing protease inhibitors (HY-K0010, MedChemExpress, USA). For immunoprecipitation, Dynabeads Protein G beads (10003D, Thermo Fisher Scientific, USA) were incubated with the specified antibodies for 2 h at 4 °C. The beads were then mixed with cell lysates and rotated overnight. The obtained samples were distinguished by SDS-PAGE gels. Then, the gels were visualized with Coomassie Blue, and the specific pieces of gel were cut out for mass spectrometry (GeneChem Co., Ltd.). The primary antibodies used in Western blotting are shown in Supplementary Table 1.
CCK-8 assay
8000 cells were plated into 96-well plates. After drug treatment or siRNA (plasmids) transfection, the culture medium was replaced with proportionally diluted CCK-8 liquid (BS350A, Biosharp, China). The well-mixed culture plates were incubated in the cell incubator for one hour. The absorbance was determined by using a microplate reader (TECAN, Switzerland) at 450 nm to evaluate cell viability.
Sphere formation in Matrigel
In our previous research, we have already described the experimental protocol for the sphere formation of PCa cells [21]. Cells were digested with trypsin and resuspended at a concentration of 2000 cells/100 μL. Add 60 μL of Matrigel (BD Biosciences, Sparks, MD) to each chamber of the 8-chamber slide. After curing Matrigel in a 37 °C incubator for 1 h, 300 μL of the cell suspension in the medium was inoculated in each chamber. After incubation for 10 days, photos were taken with a microscope.
RNA extraction and quantitative RT-PCR
Total RNA was purified using the RNA Purification Kit (AC0202-B, SparkJade, Shandong, China) following the reagent supplier’s protocol. 1000 ng RNA as templates was reverse-transcribed into cDNA with reverse transcription kit (R423-01, Vazyme, Jiangsu, China). RT-qPCR analysis was implemented by using SYBR green (Q341-02, Vazyme, Jiangsu, China) in a PCR instrument. The primers used for RT-PCR are shown in Supplementary Table 2.
RNA stability assays
To test the half-life of AKT, we inoculated C4-2B cells and C4-2B-ENZR cells in a 6-well cell culture plate. Next, actinomycin D was supplied into cells at a concentration of 10 μg/mL. Cells were obtained at 0, 4, and 8 h, respectively. 22Rv1 with WTAP knocked down cells, and control cells were inoculated in a 6-well culture plate. Following that, cells were treated with actinomycin D at a level of 10 μg/mL, and cells were isolated at 0, 4, and 8 h, respectively. Total RNA was collected, and the remaining AKT was detected by RT-qPCR.
MeRIP-RT-qPCR
MeRIP measurement was carried out in accordance with the instructions of the RIP kit (GS-ET-006, Cloudseq, Shanghai, China). Digest the adherent cells with trypsin, and wash the cells with PBS. Total RNA was extracted as described. Incubate the fragmented RNA with the magnetic beads that have already bound the m6A antibody for one hour, and then purify the binding RNA. The purified RNA was analyzed by RT-qPCR. The primers used for qRT-PCR are displayed in Supplementary Table 2. Relatively enriched normalization was performed on the input as: ΔCt = CtIP-(Ctinput-Log2[Input Dilution Factor]). Calculate the proportion of input for each IP sample: %Input = 2(-ΔCt[normalized IP]).
Luciferase reporter gene assays
WTAP-wild and WTAP-mutant promoters were cloned into the pGL3 vector. The primers used in plasmid engineering are shown in Supplementary Table 3. Cells were plated into 24-well plates following by co-transfection with 0.2 μg wild-type or mutant WTAP promoter plasmids with 20 ng renilla luciferase plasmids. The luciferase activity was assessed utilizing the dual-luciferase reporter gene detection kit (E1910, Promega, USA).
Chromatin immunoprecipitation (ChIP) assay
ChIP experiment was performed according to the supplier’s protocols of Chromatin IP Kit (53008, Active Motif, MA, USA). Fix the cells cultured in 10 cm dishes with 1% formaldehyde solution for 10 min, then scrape off the adherent cells and spin at 4 °C for 10 min. Resuspend the cell precipitate with lysis buffer at ice for 30 min. The products after enzymatic digestion were incubated with AR antibodies overnight. After washing, the DNA was recovered for PCR analysis. The primers used in ChIP were shown in Supplementary Table 2.
RNA sequencing and statistical analysis
RNA-seq was conducted by Cloudseq Biotech Inc. (Shanghai, China). Libraries were generated using the GenSeq®Low Input RNA library Prep Kit (GenSeq, Inc.), and quality was evaluated using the BioAnalyzer 2100 system. Library sequencing was conducted on the Illumina HiSeq instrument, with 150 bp of paired terminal reads. The RNA sequence data has been stored in GEO (GSE301437).
All data were shown as mean ± SEM. The difference analysis between the two groups was conducted using the Student’s t-test, and analysis of ANOVA was used for more than two groups. All in vitro experiments were biological triple replicates. All data analysis in this investigation was performed using GraphPad Prism 9.
Results
WTAP is upregulated in PCa patient tissues
To determine the expression of WTAP in PCa, we performed an IHC staining assay in a tissue microarray (TMA) containing 60 paired PCa tissues and normal tissues. Indeed, IHC analysis revealed that WTAP protein was highly expressed in tumor tissues in comparison with that of the matched normal tissues (Fig. 1A, B). However, there is no significant difference in WTAP expression between different grades of PCa (Fig. 1C). Further transcriptome analyses on the TCGA database showed that mRNA levels of WTAP were higher in tumor tissues than in normal tissues (Fig. 1D). We also performed a Western blot assay to confirm that WTAP expression was higher in human PCa tissues than normal tissues from the same patient (Fig. 1E). These results suggested that WTAP may be an oncogene in PCa patient tumors.
A Representative images of WTAP staining by Immuno-histochemistry (IHC) in PCa tissues or prostate normal tissues from a tissue microarray, which contained 60 pairs of patients’ tissues. B, C WTAP staining score was increased in tumors compared to adjacent normal tissues, but had no significant change in different grades. ns, not significant. D WTAP is over-expressed in PCa tumors (n = 493) compared with prostate normal tissues (n = 52) in the TCGA dataset. *P < 0.05. E WTAP protein expression between normal and cancerous specimens was detected by Western blotting assay. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. WTAP expression was increased in Patient 1, Patient 3, and Patient 4.
WTAP promotes cell growth and xenograft progression
To further determine the functional roles of WTAP in PCa, we first detected WTAP expression in some PCa cell lines and found that WTAP had higher expression in C4-2B and 22Rv1 cells compared to PC-3 and DU 145 cells (Fig. 2A). We then performed WTAP overexpression in PC-3 cells and WTAP knockdown in C4-2B cells (Fig. 2B and H). Enhanced WTAP expression increased sphere formation in PC-3 cells (Fig. 2C). In addition, WTAP knockdown resulted in decreased sphere formation of C4-2B cells (Fig. 2I). Furthermore, we established a xenograft model in male nude mice to explore the function of WTAP overexpression in vivo. The tumor volume in the PC-3 xenograft with WTAP overexpression was higher than that in the control group (Fig. 2D and E). Accordingly, xenograft tumor weight was also heavier for the grafts with WTAP-overexpression cells (Fig. 2F). The xenograft model was also applied to determine whether WTAP knockdown inhibited PCa tumor growth. And we found that the knockdown of WTAP in 22Rv1 decreased the volume and weight of the tumor in mice (Supplementary Fig. 1). Western blot assay confirmed PCNA expression was slightly increased, which was associated with WTAP overexpression (Fig. 2G). Unexpectedly, WTAP knockdown also induced a decrease in sphere number in enzalutamide-resistant cells (Fig. 2J, K). Taken together, these results confirmed that WTAP promoted PCa cell growth whether in enzalutamide-sensitive cells or enzalutamide-resistant cells and indicated that WTAP plays an oncogene role in the progression of prostate cancer.
A WTAP levels in prostate cancer cell lines. PC-3, DU 145, C4-2B, and 22Rv1 cells were collected, and whole lysates were subjected to Western blotting. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. ***P < 0.001. B The efficiency of WTAP overexpression in PC-3 cells was detected by Western blot. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. **P < 0.01. C Colony formation was elevated after ectopically expressing WTAP in PC-3 cells. Quantitative analysis of colony formation was examined by ImageJ. n = 3. ***P < 0.001. D Photographs of tumor formation in nude mice and tumor xenografts were harvested at 4 weeks after injection of WTAP over-expressing or empty vector cells. E, F Tumor growth curves measured after inoculation at the indicated times. The tumor volume was calculated at termination. n = 7. **P < 0.01; ***P < 0.001. G Western blotting showed PCNA was increased in cells stably transfected with WTAP compared with the control vector. The relative WTAP and PCNA quantification of the Western blotting was measured by using ImageJ software. *P < 0.05; ***P < 0.001. H The knock-down efficiency of WTAP with lentiviral constructs constitutively expressing shRNA in C4-2B cells was confirmed by Western blotting. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. I Colony formation in Matrigel was decreased in the WTAP knockdown cell line. The average number of spheres with a diameter >75 µm per well. n = 3. ***P < 0.001. J C4-2B-ENZR cells were transfected with siRNA targeting WTAP. Cells were then collected and analyzed by Western blotting. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. K Colony formation in Matrigel was decreased when WTAP was silenced in C4-2B-ENZR cells. The average number of spheres with a diameter >75 µm per well. n = 3. ***P < 0.001.
AR enhanced WTAP gene transcription by directly binding to the promoter of WTAP
Initially, we analyzed WTAP expression in prostate cancer cells using Cancer Cell Line Encyclopedia (CCLE) and noted that the mRNA levels of WTAP were higher in AR-positive cells than in AR-negative cells (Fig. 3A). Consistent results showed that AR-positive C4-2B cells and LNCaP cells express higher WTAP than AR-negative DU 145 cells and PC-3 cells (Fig. 3B). To investigate the relationship between AR and WTAP, we next explored whether WTAP expression was regulated by AR signaling. We found that WTAP was increased by R1881 at protein levels in a dose-dependent manner in C4-2B and 22Rv1 cells (Fig. 3C–F). In addition, R1881 can elevate WTAP expression in a dose-dependent manner in LNCaP cells or in a time-dependent manner 22Rv1 cells (Supplementary Fig. 2A–C) and AR depleted reduced WTAP expression (Supplementary Fig. 2D). Unexpectedly, exogenous AR expression elevated WTAP expression and rescued the activated effect of R1881 in AR negative PC-3 cells, which can increase WTAP expression (Supplementary Fig. 2E). In contrast, When WTAP was depleted by siRNA or shRNA in C4-2B and 22Rv1 cells, full-length AR did not significantly change (Supplementary Fig. 3A–C). However, AR splice variant 7 (AR-V7) was reduced after WTAP was knocked down in 22Rv1.
A The mRNA expression of WTAP in AR-positive or AR-negative prostate cancer cells, analyzed by Cancer Cell Line Encyclopedia (CCLE) database. B WTAP and AR levels in prostate cancer cell lines. RWPE1, PC-3, DU 145,22Rv1, C4-2B, and LNCaP cells were collected, and whole lysates were analyzed by Western blotting. The relative WTAP and AR quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. C–F WTAP levels were upregulated by R1881 treatment in C4–2B cells (C, D) and 22Rv1 (E, F). 22Rv1 and C4–2B cells were treated with titrated doses of R1881 for 24 h. The relative WTAP and AR quantification of the Western blotting was measured by using ImageJ software. *P < 0.05; **P < 0.01; ***P < 0.001. G Dose-dependent upregulation of WTAP by R1881. C4-2B cells were treated with R1881 for 24 h at the indicated concentrations. WTAP mRNA levels were detected at the indicated time points by qRT-PCR. n = 3. ***P < 0.001. H AR motif consensus sequence binding to WTAP promoter region was predicted by hTFtarget (Database of Human Transcription Factor Targets). WTAP promoter reporter constructs, including mutant promoter and wild-type plasmids, were designed for the luciferase reporter assay. I Luciferase reporter assay to analyze the transcriptional impact of the AR signal pathway on the WTAP promoter in C4-2B cells. WTAP luciferase activity was calculated as the ratio of firefly to Renilla luciferase activities and performed in triplicate. *P < 0.05; ns not significant. J Three ChIP-PCR amplicons were shown in the red rectangles (P1, P2, and P3), and a green circle indicated the predicted sites of WTAP promoter bound by AR (up). ChIP-qPCR assays were performed using the AR antibody. The percentages of coimmunoprecipitated DNA by antibody relative to the input DNA were shown (down). ***P < 0.001.
To further investigate whether AR regulated the transcription of the WTAP gene, we detected the WTAP mRNA level and promoter activity when AR signaling was activated. We found that WTAP mRNA level was elevated by R1881 treatment in C4-2B cells (Fig. 3G). Meanwhile, WTAP mRNA level was also reduced by RNA silencing of the AR gene in C4-2B cells (Supplementary Fig. 2F). To explore whether AR affected the promoter activity of WTAP, we identified the potential binding sites of AR on WTAP using the hTFtarget database [22] and constructed the wild-type and mutant WTAP promoters (Fig. 3H). Luciferase reporter assays showed that AR can activate wild-type promoter activity of WTAP but not mutant (Fig. 3I). Furthermore, we designed three pairs of primers as shown in Fig. 3J and used chromatin immunoprecipitation (ChIP) to verify the regions of the WTAP promoter bound by AR. ChIP-qPCR assays showed that AR had stronger binding to the P2 region than the P1 and P3 regions of the WTAP promoter. These results demonstrated that AR regulated WTAP expression by directly binding to WTAP to activate WTAP transcription.
WTAP stability was enhanced by USP7 in Enza-resistant PCa cells
On the basis of the regulation of AR on WTAP expression in PCa cells and the critical role of AR in Enza-resistant, we explored whether WTAP is involved in the progression of Enza-resistant. We first examined the expression of WTAP when C4-2B cells were added with enzalutamide and found that WTAP expression decreased within 4 weeks (Fig. 4A), which indicated that WTAP expression was regulated by AR-mediated transcriptional regulation at the early stage of enzalutamide treatment. However, WTAP expression was upregulated in Enza-resistant C4-2B (C4-2B-ENZR) cells, which were treated with enzalutamide for more than 3 months (Fig. 4B). Notably, among N6-methyladenosine (m6A)-related enzymes, METTL3 or FTO had no change in C4-2B-ENZR cells compared to parental cells. Because protein expression of WTAP was decreased in C4-2B-ENZR cells but not transcripts (Fig. 4C), we hypothesized that the protein stability of WTAP may be regulated by the ubiquitin-proteasome pathway. To verify this hypothesis, we validated the stability of WTAP upon cycloheximide (CHX) treatment in C4-2B or C4-2B-ENZR cells. Results showed that WTAP degradation in C4-2B-ENZR cells was inhibited compared to C4-2B parental cells (Fig. 4D). These results suggested that elevated WTAP expression may exert important roles in PCa with ENZR.
A C4–2B cells were treated with 20 μM enzalutamide (ENZ) for the indicated weeks, and the protein levels of WTAP, AR, METTL3, and FTO were validated by Western blotting. The relative WTAP, AR, METTL3, and FTO quantification of the Western blotting was measured by using ImageJ software. *P < 0.05 **P < 0.01. B WTAP, METTL3, FTO, and AR protein levels in C4–2B and C4–2B-ENZR cells were determined by Western blotting. The relative WTAP, AR, METTL3, and FTO quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. C The mRNA levels of WTAP in C4-2B cells and C4-2B-ENZR cells were analyzed by qRT-PCR. (D) WTAP expression was detected by Western blotting. C4–2B and C4–2B-ENZR cells were treated with 40 mg/mL cycloheximide and collected at 0, 4, 8, and 12 h. The relative WTAP quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. E Identification of interacting proteins with WTAP by Co-IP and mass spectrometry. C4-2B cells and C4-2B-ENZR cells lysates were immunoprecipitated with the WTAP antibody. All associated proteins were separated by SDS-PAGE, and Coomassie blue staining was performed. F Immunoblots of Co-IP assays showing interaction between WTAP and USP7. Co-IP assays were performed using C4-2B-ENZR cell lysates. G Structural schematic diagram of the USP7 deletion mutants used for domain mapping (left panel). 293T cells expressing exogenously WTAP were co-transfected with USP7 deletion mutants. Interactions were analyzed using a co-immunoprecipitation assay (right panel). H, I Western blotting analysis of WTAP expression in 22Rv1 and C4-2B-ENZR cells treated with DMSO or P5091 with different concentrations as indicated. The relative WTAP and USP7 quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. J C4–2B-ENZR and C4-2B cells were transfected with HA-Ub. Protein lysis was collected from these cells to perform IP with WTAP, followed by Western blotting with the indicated antibodies. K 293T cells were transfected with WTAP and HA-Ub in combination with USP7 wild or mutant plasmid. Protein lysis was collected from these cells to perform IP with WTAP antibody, followed by Western blotting with the indicated antibodies.
To investigate the related proteins that mediate the degradation of WTAP, we performed mass spectrometry analysis and identified Ubiquitin-Specific Protease 7 (USP7) as a WTAP binding protein in C4-2B-ENZR (Fig. 4E). USP7, as a deubiquitinating enzyme, can protect target proteins from proteasome-mediated degradation [23, 24]. Recently, Qiu et al. [25] also found that USP7 interacted with WTAP and enhanced the protein stability of WTAP by inhibiting the ubiquitination in glioblastoma. To confirm the interaction of WTAP with USP7 in C4-2B-ENZR cells, we performed co-immunoprecipitation (co-IP) assays with cellular extracts from C4-2B-ENZR cells and confirmed that WTAP interacted with USP7 (Fig. 4F). To determine which region of USP7 interacts with WTAP, we applied co-IP assays in 293T cells and found the TRAF domain of USP7 interacted with WTAP (Fig. 4G). Importantly, P5091, an inhibitor of USP7, reduced the increase of WTAP in C4-2B-ENZR compared to C4-2B cells (Fig. 4H) and decreased WTAP protein levels in 22Rv1 with concentration increase (Fig. 4I). The ubiquitination of WTAP protein was increased in C4-2B-ENZR cells (Fig. 4J). To verify whether the deubiquitinating activity of USP7 is required for WTAP stability, we utilized a USP7 mutation C223S that has been known to be critical for its deubiquitination function [26]. We found that USP7 but not USP7 (C223S) decreased the ubiquitination of WTAP (Fig. 4K). These results suggested that USP7 promotes WTAP expression by stabilizing WTAP protein stability.
WTAP promotes AKT signaling through an m6A-mediated mechanism
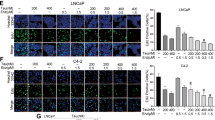
To further develop the function of WTAP in ENZR cells, we first performed RNA sequencing (RNA-seq) and found the PI3K/AKT signaling pathway is activated in C4-2B-ENZR cells compared to C4-2B parental cells (Fig. 5A). The PI3K/AKT pathway is frequently altered in the progression of resistance to androgen receptor antagonists (ARAs) [27], and some evidence has shown that AKT inhibitor combination with ARAs may be beneficial for mCRPC in clinical trials [9, 28]. However, the potential value in clinical or predictive biomarkers of this combination still needs to be studied. We verified that the increase of AKT expression in C4-2B-ENZR was reduced after silencing WTAP (Fig. 5B), and WTAP knockdown also induced the decrease of AKT in 22Rv1 cells (Fig. 5C). Considering the prevalent role of WTAP and METTL3 in m6A modification, we intended to investigate whether AKT expression was mediated by m6A modification, and we used METTL3 inhibitor STM2457 to treat PCa cells and found that the protein expression level of AKT was decreased (Fig. 5D, E). Meanwhile, we found METTL3 knockdown or 3-Deazaadenosine (DAA) treatment, a known m6A inhibitor [29], also compromised the increase of AKT expression in C4-2B-ENZR cells, which resulted in repeatable outcomes in 22Rv1 cells (Supplementary Fig. 4A–E). Moreover, Degradation of AKT mRNA was accelerated by knockdown of WTAP in 22Rv1 cells (Supplementary Fig. 4F). AKT mRNA was stabilized in C4-2B-ENZR compared to parental cells (Fig. 5F). To further verify the WTAP/METTL3-mediated m6A modification on AKT, we predicted the m6A modification site of AKT via the SRAMP prediction server [30] and found three sites (P1, P2, and P3) in the CDS of the AKT transcript had a very high combined score (Supplementary Fig. 4G). We validated that the region P1 of the AKT transcript had stronger m6A intensities in C4-2B-ENZR than in parental cells, but not P2 and P3 (Fig. 5G), and the enrichment was decreased when WTAP was depleted (Fig. 5H). More importantly, WTAP overexpression prevented the inhibition of proliferation by enzalutamide, which was abolished by AKT inhibitor (perifosine) treatment (Fig. 5I). Additionally, the enzalutamide-resistance property of C4-2B-ENZR cells can be compromised by either WTAP knockdown or perifosine treatment (Fig. 5J). WTAP silencing or AKT inhibitors enhanced the inhibitory effect of enzalutamide on 22Rv1 cells (Supplementary Fig. 4H). As previously verified, the USP7 inhibitor P5091 can reduce the expression level of WTAP. After treating C4-2B-ENZR cells and 22Rv1 cells with P5091, the protein expression of AKT was also down-regulated (Supplementary Fig. 4I and J). Furthermore, we used CCK8 assays to explore the role of USP7 in enzalutamide-resistant cells and found that, combined with the AKT inhibitor perifosine, the USP7 inhibitor P5091 can inhibit the proliferation of C4-2B-ENZR cells (Fig. 5K). These results suggested that WTAP served as a pivotal regulator in AKT signaling via an m6A-methylation manner in ENZR cells.
A Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis showing altered pathways between C4-2B and C4-2B-ENZR cells. B Immunoblots of indicated proteins in C4-2B parental and C4-2B-ENZR cells. C4-2B-ENZR cells were transfected with siRNAs targeting WTAP. The relative AKT, p-AKT, and WTAP quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. C 22Rv1 cells were transfected with siCtrl or WTAP siRNA. Western blotting was performed to detect AKT expression. The relative AKT and WTAP quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. D, E C4–2B-ENZR and 22Rv1 cells were treated with STM2457 at the indicated concentrations for 48 h. Western blotting was used to detect the level of AKT. The relative AKT quantification of the Western blotting was measured by using ImageJ software. *P < 0.05. F AKT mRNA stability was measured in C4-2B and C4-2B-ENZR cells. G MeRIP-qPCR analysis was used to validate m6A levels of AKT pre-mRNA in C4-2B and C4-2B-ENZR cells. H MeRIP-qPCR analysis was used to validate m6A levels of AKT pre-mRNA in C4-2B shCtrl and shWTAP cells. I–K Cell viability was measured by Cell Counting Kit-8 (CCK8) assay in C4-2B and C4-2B-ENZR cells under AKT inhibitor or enzalutamide treatment, or USP7 inhibitor. Cells were transiently transfected with the indicated WTAP siRNA or Flag-WTAP plasmid before treatment. *P < 0.05; **P < 0.01, ***P < 0.001.
Discussion
This study elucidated a novel function of WTAP in the development of enzalutamide-resistant (ENZR) PCa. We found that WTAP knockdown can not only inhibit PCa xenograft progression but also regain the sensitivity of ENZR cells to enzalutamide. Meanwhile, we defined WTAP as a novel responsive gene of AR signaling which enhancing the transcription of WTAP by directly binding to the WTAP promoter. Mechanistically, WTAP was associated with USP7 and was stabilized by USP7-mediated deubiquitination in ENZR cells. More importantly, WTAP can facilitate enzalutamide resistance by enhancing the AKT signaling pathway in an m6A-dependent manner. Our study can provide novel perspectives to manage PCa at different stages.
Although dysregulation of WTAP is associated with tumorigenesis in various cancers, including glioblastoma, acute myeloid leukemia, and hepatocellular carcinoma [25, 31, 32], the reports about the role of WTAP in prostate cancer are few. In this study, we further identified that WTAP functioned as an oncogenic role that WTAP was upregulated in PCa tissues, and WTAP overexpression promoted xenograft growth. Recently, Liyanage et al. [33] have reported that WTAP may play important roles in PCa progression to mCRPC, which is consistent with our findings and strengthens the credibility of our research. More importantly, we found that WTAP knockdown also inhibited cell proliferation in ENZR cells. These findings provide further exploration of WTAP at different stages of PCa from enzalutamide-sensitive to ENZR prostate tumors.
Androgen receptor (AR), a ligand-dependent transcription factor, plays important roles in prostate development and prostate cancer progression through regulating target genes’ expression [34, 35]. Here, we found that WTAP is a novel target gene of the AR signaling pathway. AR upregulated WTAP protein expression by directly binding the WTAP promoter to enhance the transcription of WTAP. Therefore, enzalutamide reduced WTAP expression in the early stage of enzalutamide treatment by repressing AR activation. Unexpectedly, WTAP was increased in ENZR cells treated with enzalutamide for more than three months. These results indicated that WTAP expression was not regulated by AR signaling in ENZR cells. Further investigations revealed that WTAP bound the TRAF region of deubiquitinating enzyme USP7, which can enhance WTAP protein stability in ENZR cells. P5091, a USP7 inhibitor, can reduce the increase of WTAP induced by ENZR in PCa cells. These findings provide a fresh perspective on whether USP7 inhibitors can modulate ENZR cell proliferation by promoting WTAP degradation.
The phosphoinositide 3-kinase (PI3K)/AKT pathway plays important roles in the pathogenesis of various cancers. This study reports that AKT signaling is upregulated in enzalutamide-resistant cells compared to parental cells. Previous studies have demonstrated that enzalutamide treatment activated AKT signaling [36], which supported the sequencing results of our study. The AKT pathway plays important roles in cell proliferation and survival. In CRPC, AKT signaling is elevated and related to poor prognosis [37]. Some preclinical researches have demonstrated that the combination of AKT inhibitor and enzalutamide may inhibit the development of enzalutamide-resistant prostate cancer [9, 28, 38]. Recent studies have demonstrated that WTAP is involved in drug resistance in some cancers, including endometrial cancer [39], pancreatic cancer [40], and esophageal cancer [41]. Some studies have reported that m6A regulators are related with PI3K/AKT pathway in cancers [42, 43]. However, the function of WTAP in PI3K/AKT is not known. In our study, we found that WTAP plays an important role in the progression of enzalutamide-resistant prostate cancer by mediating AKT expression through an m6A methylation-dependent way. These findings provide a novel perspective that AKT activity can be mediated by WTAP.
The development of enzalutamide resistance is intricate, and the mechanisms underlying it contain epigenetic and adaptive bypass ways. We have shown the promoting role of WTAP, whether in PCa or in the progression of enzalutamide resistance. These results suggest that WTAP may be a potential biomarker for treatment in PCa or ENZR PCa. Currently, there is no inhibitor for WTAP, but we found that USP7 regulates WTAP expression by preventing the degradation of WTAP. Some USP7 inhibitors have been proven to act as an anti-tumor agent in some cancers. In gastric cancer, DHPO, identified as a potent USP7 inhibitor, inhibited cell viability and tumor migration through triggering ferroptosis via targeting Stearoyl-CoA Desaturase (SCD) through deubiquitination [44]. In liver cancer, USP7 inhibitor P5091 could suppress liver cancer growth by downregulating FGL1 expression to enhance CD8+ T cell activity [45]. These data may provide a potential treatment that USP7 inhibitor alone or in combination with enzalutamide or AKT inhibitor for enzalutamide-resistant prostate cancer. But we cannot exclude the effect of the USP7 inhibitor on other m6A regulators, which may be further explored.
In conclusion, we have demonstrated that WTAP is regulated by the AR signaling pathway through regulating the transcript level of WTAP in hormone-sensitive status. Furthermore, WTAP may be an important mediator in the progression of enzalutamide resistance by elevating AKT expression via an m6A-dependent way, as depicted in Fig. 6.
In enzalutamide-sensitive PCa cells, enzalutamide inhibited the enhancing effect of AR on WTAP promoter, inducing a decrease in WTAP expression. In enzalutamide-resistant PCa cells, the effect of enzalutamide on the WTAP promoter is abolished, and USP7 elevates WTAP expression through mediating the de-ubiquitination of WTAP to enhance WTAP stability. Figure 6 was drawn by FigDraw (Permission number: ARSWW64d43).
Data availability
All data generated and analyzed during the current study are included in this article or from the corresponding authors on reasonable request. The RNA sequence data has been stored in GEO (GSE301437).
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2021. CA Cancer J Clin. 2021;71:7–33.
Teo MY, Rathkopf DE, Kantoff P. Treatment of advanced prostate cancer. Annu Rev Med. 2019;70:479–99.
Cai MP, Song XL, Li XA, Chen MK, Guo JD, Yang DH, et al. Current therapy and drug resistance in metastatic castration-resistant prostate cancer. Drug Resist Update. 2023;68:100962.
Han D, Labaf M, Zhao YW, Owiredu J, Zhang SQ, Patel K, et al. Androgen receptor splice variants drive castrationresistant prostate cancer metastasis by activating distinct transcriptional programs. J Clin Invest. 2024;134:e168649.
Ji Y, Zhang RY, Han XL, Zhou JM. Targeting the N-terminal domain of the androgen receptor: The effective approach in therapy of CRPC. Eur J Med Chem. 2023;247:115077.
Schweizer MT, Yu EY. Persistent androgen receptor addiction in castration-resistant prostate cancer. J Hematol Oncol. 2015;8:128.
He YD, Xu WD, Xiao YT, Huang HJ, Gu D, Ren SC. Targeting signaling pathways in prostate cancer: mechanisms and clinical trials. Signal Transduct Target Ther. 2022;7:198.
Mao NH, Lee YS, Salsabeel N, Zhang ZD, Li D, Kaur H, et al. Uncoupling of Akt and mTOR signaling drives resistance to Akt inhibition in PTEN loss prostate cancers. Sci Adv. 2025;11:eadq3802.
Rescigno P, Porta N, Finneran L, Riisnaes R, Figueiredo I, Carreira S, et al. Capivasertib in combination with enzalutamide for metastatic castration resistant prostate cancer after docetaxel and abiraterone: results from the randomized phase II RE-AKT trial. Eur J Cancer. 22024;205:114103.
Zhao BXS, Roundtree IA, He C. Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Bio. 2017;18:31–42.
Jiang X, Liu B, Nie Z, Duan L, Xiong Q, Jin Z, et al. The role of m6A modification in the biological functions and diseases. Signal Transduct Target Ther. 2021;6:74.
Naren D, Yan T, Gong Y, Huang J, Zhang D, Sang L, et al. High Wilms’ tumor 1 associating protein expression predicts poor prognosis in acute myeloid leukemia and regulates m(6)A methylation of MYC mRNA. J Cancer Res Clin Oncol. 2021;147:33–47.
Kuai Y, Gong X, Ding LY, Li F, Lei LZ, Gong YQ, et al. Wilms’ tumor 1-associating protein plays an aggressive role in diffuse large B-cell lymphoma and forms a complex with BCL6 via Hsp90. Cell Commun Signal. 2018;16:50.
Zhang JW, Tsoi H, Li XX, Wang H, Gao J, Wang KN, et al. Carbonic anhydrase IV inhibits colon cancer development by inhibiting the Wnt signalling pathway through targeting the WTAP-WT1-TBL1 axis. Gut. 2016;65:1482–93.
He RZ, Jiang J, Hu XL, Lei M, Li J, Luo WH, et al. Stabilization of UCA1 by N6-methyladenosine RNA methylation modification promotes colorectal cancer progression. Cancer Cell Int. 2021;21:616.
Tang JY, Wang F, Cheng G, Si SH, Sun X, Han J, et al. Wilms’ tumor 1-associating protein promotes renal cell carcinoma proliferation by regulating CDK2 mRNA stability. J Exp Clin Canc Res. 2018;37:40.
Ou BC, Liu Y, Yang XW, Xu XJ, Yan YW, Zhang JJ. C5aR1-positive neutrophils promote breast cancer glycolysis through WTAP-dependent m6A methylation of ENO1. Cell Death Dis. 2021;12:737.
Huang T, Cao LL, Feng NN, Xu B, Dong YJ, Wang M. N6-methyladenosine (m6A)-mediated lncRNA DLGAP1-AS1enhances breast canceradriamycin resistance through miR-299-3p/WTAP feedback loop. Bioengineered. 2021;12:10935–44.
Hu QX, Yin JL, Zhao SJ, Wang YB, Shi RX, Yan KQ, et al. ZFHX3 acts as a tumor suppressor in prostate cancer by targeting FTO-mediated m6A demethylation. Cell Death Discov. 2024;10:284.
Gao L, Zhang W, Zhang J, Liu J, Sun F, Liu H, et al. KIF15-mediated stabilization of AR and AR-V7 contributes to enzalutamide resistance in prostate cancer. Cancer Res. 2021;81:1026–39.
Hu Q, Zhang B, Chen R, Fu C, A J, Fu X, et al. ZFHX3 is indispensable for ERβ to inhibit cell proliferation via MYC downregulation in prostate cancer cells. Oncogenesis. 2019;8:28.
Zhang Q, Liu W, Zhang HM, Xie GY, Miao YR, Xia M, et al. hTFtarget: a comprehensive database for regulations of human transcription factors and their targets. Genom Proteom Bioinform. 2020;18:120–8.
Nie LT, Wang C, Liu XG, Teng HQ, Li ST, Huang M, et al. USP7 substrates identified by proteomics analysis reveal the specificity of USP7. Gene Dev. 2022;36,:1016–30.
Fernández-Montalvan A, Bouwmeester T, Joberty G, Mader R, Mahnke M, Pierrat B, et al. Biochemical characterization of USP7 reveals post-translational modification sites and structural requirements for substrate processing and subcellular localization. FEBS J. 2007;274:4256–70.
Qiu JW, Zhao RR, Ma CZ, Wang QT, Li BY, Qi YH, et al. O-GlcNAcylation stabilized WTAP promotes GBM malignant progression in an N6-methyladenosine-dependent manner. Dev Oncol. 2025;27:900–15.
Huang B, Cao D, Yuan X, Xiong YX, Chen BZ, Wang YJ, et al. USP7 deubiquitinates KRAS and promotes non-small cell lung cancer. Cell Rep. 2024;43:114917.
Schmidt KT, Huitema ADR, Chau CH, Figg WD. Resistance to second-generation androgen receptor antagonists in prostate cancer. Nat Rev Urol. 2021;18:209–26.
Kolinsky MP, Rescigno P, Bianchini D, Zafeirio Z, Mehra N, Mateo J, et al. A phase I dose-escalation study of enzalutamide in combination with the AKT inhibitor AZD5363 (capivasertib) in patients with metastatic castration-resistant prostate cancer. Ann Oncol. 2020;31:619–25.
Feng ZY, Zhou FH, Tan MM, Wang TT, Chen Y, Xu WW, et al. Targeting m6A modification inhibits herpes virus 1 infection. Genes Dis. 2022;9:1114–28.
Zhou Y, Zeng P, Li YH, Zhang ZD, Cui QH. SRAMP: prediction of mammalian N6-methyladenosine (m6A) sites based on sequence-derived features. Nucleic Acids Res. 2016;44:e91.
Li YQ, Liu D, Wang LL, Shao YL, Zhou HS, Hu YL, et al. WTAP-mediated m6A methylation of PHF19 facilitates cell cycle progression by remodeling the accessible chromatin landscape in t(8;21) AML. Oncogene. 2025;44:1504–16.
Chen Y, Peng C, Chen J, Chen D, Yang B, He B, et al. WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol Cancer. 2019;18:127.
Liyanage C, Fernando A, Chamberlain A, Moradi A, Batra J. RNA m6a methylation regulator expression in castration-resistant prostate cancer progression and its genetic associations. Cancers. 2024;16:1303.
Wasmuth EV, Vanden Broeck A, LaClair JR, Hoover EA, Lawrence KE, Paknejad N, et al. Allosteric interactions prime androgen receptor dimerization and activation. Mol Cell. 2022;82:2021.
Gelmann EP. Molecular biology of the androgen receptor. J Clin Oncol. 2002;20:3001–15.
Shao HeWJ, Yu YY, Huang Y, Feng W, Li GL. JH. Exendin-4 enhances the sensitivity of prostate cancer to enzalutamide by targeting Akt activation. Prostate. 2020;80:367–75.
Bitting RL, Armstrong AJ. Targeting the PI3K/Akt/mTOR pathway in castration-resistant prostate cancer. Endocr-Relat Cancer. 2013;20:R83–R99.
Toren P, Kim S, Cordonnier T, Crafter C, Davies BR, Fazli L, et al. Combination AZD5363 with enzalutamide significantly delays enzalutamide-resistant prostate cancer in preclinical models. Eur Urol. 2015;67:986–90.
Xie WL, Liu NF, Wang XY, Wei L, Xie WY, Sheng XG. Wilms’ tumor 1-associated protein contributes to chemo-resistance to cisplatin through the Wnt/β-catenin pathway in endometrial cancer. Front Oncol. 2021;11:598344.
Li BQ, Liang ZY, Seery S, Liu QF, You L, Zhang TP, et al. WT1 associated protein promotes metastasis and chemo-resistance to gemcitabine by stabilizing Fak mRNA in pancreatic cancer. Cancer Lett. 2019;451:48–57.
Zhu ZJ, Pang Y, Jin G, Zhang HY, Wang WH, Liu JW, et al. Hypoxia induces chemoresistance of esophageal cancer cells to cisplatin through regulating the lncRNA-EMS/miR-758-3p/WTAP axis. Aging. 2021;13:17155–76.
Liu J, Gu XY, Guan ZJ, Huang D, Xing HW, Zheng L Role of m6A modification in regulating the PI3K/AKT signaling pathway in cancer. J Transl Med. 2023;21:774.
Liu J, Eckert MA, Harada BT, Liu SM, Lu ZK, Yu KK, et al. m6A mRNA methylation regulates AKT activity to promote the proliferation and tumorigenicity of endometrial cancer. Nat Cell Biol. 2018;20:1074.
Guan XQ, Wang YC, Yu WK, Wei Y, Lu Y, Dai EY, et al. Blocking ubiquitin-specific protease 7 induces ferroptosis in gastric cancer via targeting Stearoyl-CoA Desaturase. Adv Sci. 2024;11:e2307899.
Sun LL, Zhao LN, Sun J, Yuan HF, Wang YF, Hou CY, et al. Inhibition of USP7 enhances CD8+ T cell activity in liver cancer by suppressing PRDM1-mediated FGL1 upregulation. Acta Pharmacol Sin. 2024;45:1686–700.
Acknowledgements
Figure 6 was drawn by FigDraw (Permission number: ARSWW64d43) (www.figdraw.com).
Funding
This work was supported by grant 82002691 from the National Natural Science Foundation of China (NSFC) and grant ZR2020QC068 from the Natural Science Foundation of Shandong Province.
Author information
Authors and Affiliations
Contributions
QH and SH conceived the project; QH, RS, QS, XW, and SH designed all experiments and analyzed all data; QH, RS, KG, LH, QS, and XW performed experiment; QH and SH prepared the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics statement
This study was conducted in accordance with the Declaration of Helsinki, and the study was vested with the authority to the Ethics Committee of Shandong Provincial Hospital of Shandong First Medical University (R202406110275). All the patients participating in the research have signed the informed consent form. The animal assays were officially permitted to the Animal Ethics Committee of Shandong Provincial Hospital of Shandong First Medical University (W202406110553).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shi, R., Gu, K., Li, H. et al. WTAP stabilized by USP7 contributes to enzalutamide resistance in prostate cancer via mediating AKT m6A-modification. Cancer Gene Ther 33, 277–288 (2026). https://doi.org/10.1038/s41417-026-01013-y
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41417-026-01013-y