Abstract
Reprogramming of lipid metabolism is a hallmark of malignant tumors, and targeting key enzymes in lipid metabolism has emerged as a critical strategy to inhibit tumor progression. Sterol regulatory element-binding protein 1 (SREBP1), a master regulator of lipid biosynthesis, drives lipid metabolic reprogramming that not only promotes malignant progression but also confers resistance to ferroptosis in tumor cells. Ferroptosis is a distinct form of regulated cell death characterized by iron accumulation and lipid peroxidation. In this study, we demonstrate that silencing SREBP1 in ovarian cancer cells leads to decreased glutathione (GSH) levels, reduced protein expression of XCT and GPX4, and increased levels of malondialdehyde (MDA) and lipid peroxidation, indicating that SREBP1 silencing induces ferroptosis in ovarian cancer cells. Further experiments, including the CCK-8, EdU, colony formation assays and flow cytometry, confirmed that SREBP1 silencing suppresses proliferation by inducing cell cycle arrest. Transwell assays, immunofluorescence (IF) staining, Nile Rad staining, and measurements of triglyceride (TG) and total cholesterol (TC) levels demonstrated that SREBP1 silencing inhibits epithelial–mesenchymal transition (EMT) and lipid synthesis in ovarian cancer cells. Notably, SREBP1 silencing downregulates the expression of Nrf2, and treatment with the Nrf2 activator THBQ reverses the effects of SREBP1 silencing on ovarian cancer cell proliferation and ferroptosis. Mechanistically, SREBP1 silencing promotes ubiquitination-mediated degradation of the Nrf2 protein, thereby suppressing the expression of XCT and GPX4, ultimately triggering ferroptosis in ovarian cancer cells. Our findings establish SREBP1 as a key mediator of ferroptosis resistance and nominates it as both a therapeutic target and a potential prognostic biomarker in ovarian cancer.

Schematic diagram illustrating the mechanism whereby silent SREBP1 mediates the Nrf2/XCT/GPX4 pathway to induce ferroptosis in ovarian cancer cells.
Similar content being viewed by others
Introduction
Ovarian cancer remains one of the most lethal gynecological malignancies, primarily due to its frequent diagnosis at advanced stages, high recurrence rate, and consequently poor prognosis [1]. The pathogenesis and progression of ovarian cancer involve the deletion or mutation of multiple genes, among which hereditary mutations in BRCA1 and BRCA2 are of particular clinical relevance [2]. Currently, platinum-based chemotherapy continues to serve as the first-line treatment for advanced disease [3], and while patients with early-stage disease often respond favorably [4], most eventually experience recurrence following surgery and chemotherapy [5]. Thus, there is an urgent need to elucidate the novel mechanisms underlying malignant progression and to identify effective therapeutic targets to improve clinical outcomes.
Emerging evidence highlights that lipid metabolic reprogramming acts as a key facilitator of ovarian cancer proliferation and metastasis because it allows the high bioenergetic and biosynthetic demands of cells to be met. Alterations in the expression of lipid metabolism-related genes frequently occur early in ovarian tumorigenesis and become more pronounced with disease progression [6]. For instance, Zhang et al. demonstrated that DLAT upregulates SREBP1 expression via the JAK2/STAT5 pathway, promoting lipid metabolic reprogramming to support malignant progression [7]. Similarly, Li et al. reported that ANGPTL4 and ESM1 interact to increase binding to lipoprotein lipase (LPL), driving lipid reprogramming and subsequent activation of the JAK2-STAT3 signaling axis [8]. These findings underscore lipid metabolism as a hallmark of ovarian cancer and position key regulators within this pathway as promising targets for therapy.
Sterol regulatory element-binding protein 1 (SREBP1) acts as a central transcription factor governing lipid biosynthesis, directly regulating the expression of genes such as fatty acid synthase (FASN), stearoyl-CoA desaturase-1 (SCD1), and acetyl-CoA carboxylase (ACC), which encode enzymes critical to lipid synthesis and turnover [9]. These metabolic enzymes disrupt lipid metabolism by influencing lipid synthesis or degradation, thereby providing raw materials and energy for the malignant progression of tumors [6]. Notably, SREBP1-driven lipid synthesis promotes resistance to ferroptosis, a form of programmed cell death, in tumor cells [10]. Lounis et al. demonstrated that SREBP1 increases monounsaturated fatty acid (MUFA) synthesis via SCD1, thereby increasing ferroptosis resistance [11]. Cao et al. reported that SREBP1 inhibition sensitizes pancreatic cancer cells to ferroptosis by downregulating GPX4 [12]. Yu et al. revealed that alkannin triggers ferroptosis in gastric cancer by suppressing the c-Fos/SREBF1 axis [13]. Moreover, within the tumor microenvironment (TME), ferroptosis modulates immune evasion via effects on T cells and macrophages [14]. Ye et al. demonstrated that asiaticoside, a triterpenoid derivative, induces ferroptosis and blocks immune escape in gastric cancer by inhibiting the Wnt/β-catenin signaling pathway [15], and Zhang et al. reported that TSPO upregulates PD-L1 expression via Nrf2/P62/Keap1 to suppress ferroptosis and promote immune evasion hepatocellular carcinoma (HCC) [16]. These insights suggest that ferroptosis inducers and immunotherapy, especially immune checkpoint blockade (ICB) therapy, may synergize to enhance antitumour efficacy. Nonetheless, the role of SREBP1 in ovarian cancer and its potential interplay with immune evasion remain unexplored.
In this study, we observed that SREBP1 expression is significantly upregulated in ovarian cancer tissues and correlates with aggressive clinicopathological features. Functional experiments revealed that SREBP1 silencing attenuates lipid metabolism and suppresses the proliferative and metastatic capacities of ovarian cancer cells. Importantly, we describe a novel mechanism whereby SREBP1 promotes the ubiquitination-mediated degradation of Nrf2, impairing its nuclear translocation and subsequently inhibiting the transcription of XCT and GPX4. This SREBP1-Nrf2-XCT/GPX4 axis represents a critical signaling node that regulates ferroptosis in ovarian cancer. Targeted disruption of this pathway concurrently restrains lipid metabolic reprogramming and induces ferroptosis and thus represents a powerful dual therapeutic strategy for ovarian cancer.
Results
SREBP1 is highly expressed in ovarian cancer
Immunohistochemical (IHC) staining was conducted to assess SREBP1 expression in ovarian cancer and adjacent normal tissues. SREBP1 protein was primarily localized in the cytoplasm (Fig. 1A). Importantly, compared with that in adjacent normal tissues, the expression level of SREBP1 in ovarian cancer tissues was significantly elevated. Specifically, the positive expression rate (92.5%, 98/106) and strong positive expression rate (62.3%, 66/106) of SREBP1 in ovarian cancer tissues were markedly higher than those in adjacent normal tissues (37.5%, 15/40 and 20.0%, 8/40, respectively; Table 1). Further analysis of the correlation between SREBP1 expression and clinicopathological characteristics of ovarian cancer patients revealed that SREBP1 expression levels were significantly associated with pathological type (P = 0.018), lymph node metastasis (P = 0.026), Ki67-positive expression rate (P = 0.023), and PD-L1-positive expression rate (P = 0.005). In contrast, no significant correlations were observed between SREBP1 expression and patient age (P = 0.432), TNM stage (P = 0.586), tumor size (P = 0.310), clinical stage (P = 0.672), distant metastasis (P = 0.072), or tumor recurrence (P = 0.761) (Table 2).
A Expression of SREBP1 was evaluated by IHC staining (Scale bar = 50 µm or 20 µm). B Kaplan–Meier analysis of the relationship between overall survival and SREBP1 protein expression in 106 patients with ovarian cancer. C The expression of SREBP1 in ovarian cells with SREBP1 silencing was detected by western blot.
To investigate the role of SREBP1 in ovarian cancer progression, overall survival (OS) was analyzed in 106 patients using the Kaplan–Meier method. Patients with high SREBP1 expression exhibited significantly shorter survival times compared to those with low SREBP1 expression (Fig. 1B). To further assess the impact of elevated SREBP1 expression on prognosis, survival analysis was conducted on ovarian cancer patients for whom complete clinical information was available. Univariate Cox regression analysis revealed that OS was significantly associated with the following factors: high SREBP1 expression (P < 0.001), age (P = 0.037), pathological type (P = 0.018), lymph node metastasis (P < 0.001), heamatogenous metastasis (P < 0.001), tumor recurrence (P = 0.014), high Ki67 expression (P < 0.001), and positive for PD-L1 (P = 0.032). Further analysis using multivariate Cox regression revealed high SREBP1 expression (P = 0.015) and high Ki67 expression (P = 0.001) as independent predictors of OS in ovarian cancer patients (Table 3). These results demonstrate that SREBP1 is significantly overexpressed in ovarian cancer tissues and is associated with poor prognosis.
SREBP1 accelerates proliferation of ovarian cancer cells
Western blot analysis was performed to assess SREBP1 protein expression in five human ovarian cancer cell lines (ES-2, A2780, OVCA429, CAOV3, SKOV3). The results showed that its expression level was significantly higher in A2780 and SKOV3 cells compared to the other cell lines, while it was significantly lower in ES-2 cells (Fig. 1C and Supplementary Fig. S2D). Based on these findings, A2780, SKOV3, and ES-2 cell lines were selected for further investigation. Using lentiviral vector-mediated shRNA, SREBP1 was stably knocked down in A2780 and SKOV3 cells, and overexpressed in ES-2 cells. Western blot analysis confirmed efficient SREBP1 knockdown, with sh-SREBP1#1 and sh-SREBP1#2 selected for subsequent experiments based on their silencing efficacy (Fig. 2A and Supplementary Fig. S1A). SREBP1 overexpression in ES-2 cells was also validated (P < 0.01; Supplementary Fig. S3A).
A The expression of SREBP1 in ovarian cancer cells with SREBP1 silencing was detected by Western blot. B The proliferation of ovarian cancer cells with SREBP1 silencing was measured using the CCK-8 assay. C, D EdU staining and Colony formation assay were performed in ovarian cancer cells with silencing SREBP1 (Scale bar = 50 µm). E Cell cycle distribution in ovarian cancer cells with SREBP1 silencing was analyzed by flow cytometry. F The expression of cell cycle-related proteins was determined by Western blot. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
We next evaluated the effect of SREBP1 on ovarian cancer cell proliferation. The effect of SREBP1 silencing on cell proliferation was assessed. CCK-8 assays revealed that cell viability was significantly lower in SREBP1-silenced A2780 and SKOV3 cells than in control cells (Fig. 2B). EdU incorporation assays indicated that DNA synthesis capacity was markedly reduced upon SREBP1 silencing (Fig. 2C). Colony formation assays demonstrated that SREBP1 silencing significantly impaired clonogenic ability (Fig. 2D and Supplementary Fig. S1B). Conversely, SREBP1 overexpression produced the opposite results (Supplementary Fig. S3B–D). Collectively, these data demonstrate that SREBP1 knockdown inhibits, while its overexpression promotes, ovarian cancer cell proliferation.
To further investigate the mechanism by which SREBP1 silencing inhibits cell proliferation, we explored its effect on the cell cycle, as it has been previously reported that inducing cell cycle arrest effectively suppresses tumor cell proliferation in various cancers [17]. Flow cytometry revealed that SREBP1 silencing induced cell cycle arrest in a cell line-specific manner, with primarily S-phase arrest being observed in A2780 cells and G0/G1-phase arrest being observed in SKOV3 cells (Fig. 2E and Supplementary Fig. S1C). Consistently, western blot analysis revealed that SREBP1 silencing downregulated the expression of cell cycle-related proteins (Fig. 2F). In summary, these results indicate that SREBP1 knockdown inhibits ovarian cancer cell proliferation and induces cell cycle arrest.
Silencing SREBP1 inhibits ovarian cancer cell metastasis by suppressing EMT
To investigate the role of SREBP1 in ovarian cancer cell metastasis, we examined its effects on cell migration, invasion, and epithelial–mesenchymal transition (EMT). Wound healing and Transwell migration assays revealed that SREBP1 silencing significantly inhibited the horizontal and vertical migration capabilities of ovarian cancer cells (Fig. 3A, B). Further mechanistic studies involving immunofluorescence (IF) staining revealed that SREBP1 silencing significantly increased the fluorescence intensity of the epithelial marker E-cadherin (Fig. 3C) and decreased that of the mesenchymal marker Vimentin (Fig. 3D), indicating suppression of EMT. Western blotting further confirmed that SREBP1 silencing upregulated E-cadherin protein expression while downregulating Vimentin and the key EMT transcription factor Snail (Fig. 3E). In summary, these findings demonstrate that targeted silencing of SREBP1 effectively inhibits ovarian cancer cell migration and invasion by reversing EMT.
A The migratory ability of ovarian cancer cells with SREBP1 silencing was assessed using a wound healing assay (Scale bar = 50 µm). B The migration and invasion abilities of ovarian cancer cells with SREBP1 silencing were evaluated by transwell assay (Scale bar = 50 µm). C, D IF staining was performed to examine the expression of EMT markers in ovarian cancer cells with SREBP1 silencing (Scale bar = 20 µm). E The expression of EMT related proteins in ovarian cancer cells with SREBP1 silencing was detected by Western blotting. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
Silencing SREBP1 inhibits lipid synthesis in ovarian cancer cells
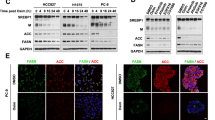
To determine whether SREBP1 regulates lipid metabolism in ovarian cancer cells, we measured intracellular lipid levels following SREBP1 knockdown. Silencing of SREBP1 led to a significant reduction in intracellular triglycerides (TG) and total cholesterol (TC), as determined by corresponding assay kits (Fig. 4A, B). The intensity of Nile Red Staining was significantly decreased in the silenced cells (Fig. 4C), indicating reduced neutral lipid content and lipid droplet formation. Western blot analysis further confirmed that the expression of lipid metabolism-related proteins was markedly downregulated in SREBP1-silenced cells (Fig. 4D). These results demonstrate that targeted silencing of SREBP1 effectively inhibits lipid synthesis in ovarian cancer cells.
A, B TG and TC levels were determined in ovarian cancer cells with silenced SREBP1 using TG or TC assay kits respectively. C Neutral lipid content in SREBP1 silenced ovarian cancer cells was detected by Nile Red Staining (Scale bar = 50 µm). D The expression of lipid metabolism-related markers in SREBP1 silenced ovarian cancer cells was analyzed by western blotting. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01 (as indicated in panels).
SREBP1 inhibits ferroptosis in ovarian cancer cells
To determine the specific mode of cell death induced by SREBP1 silencing, various cell death inhibitors including Z-VAD-FMK (apoptosis inhibitor), Necrostatin-1 (necroptosis inhibitor, Nec-1), 3-methyladenine (autophagy inhibitor, 3-MA), and Ferrostatin-1 (ferroptosis inhibitor, Fer-1), were used CCK-8 assays revealed that only Fer-1 rescued the decrease in cell proliferation caused by SREBP1 silencing (Fig. 5A), whereas Nec-1, 3-MA, and Z-VAD-FMK had no effect, suggesting that SREBP1 silencing inhibits proliferation primarily by inducing ferroptosis. Key ferroptosis indicators were subsequently assessed. Malondialdehyde (MDA) assays revealed that SREBP1 silencing significantly increased the levels of the lipid peroxidation product MDA levels (Fig. 5B). GSH assay kits revealed a significant decrease in the reduced glutathione (GSH)/oxidized glutathione (GSSG) ratio in the silenced cells (Fig. 5C). Following staining with the fluorescent probe BODIPY™ 581/591 C11, the green fluorescent intensity in the silenced cells was increased, indicating increased lipid ROS levels (Fig. 5D) and thus suggesting disrupted redox homeostasis. Following SREBP1 overexpression, a significant decrease in MDA levels (P < 0.05, Supplementary Fig. S3E) was accompanied by a significant increase in the GSH/GSSG ratio (P < 0.05 and Supplementary Fig. S3F). Furthermore, BODIPY (581/591) C11 staining revealed a significant reduction in green fluorescence, indicative of suppressed lipid peroxidation (P < 0.05, Supplementary Fig. S3G). Western blot analysis confirmed that SREBP1 silencing significantly downregulated the expression levels of key ferroptosis regulatory proteins (Fig. 5E), including the cystine/glutamate antiporter (XCT) and glutathione peroxidase 4 (GPX4) [18]. By contrast, the opposite phenotype was observed following SREBP1 overexpression (P < 0.05, Supplementary Fig. S3H). In summary, these results indicate that SREBP1 silencing induces ferroptosis in ovarian cancer cells, while its overexpression inhibits ferroptosis in ovarian cancer cells.
A Ovarian cancer cells with SREBP1 knockdown (sh-SREBP1) were treated for 24 h with either Fer-1 (10 μM), Z-VAD-FMK (10 μM), necrostatin-1 (10 μM), or 3-MA (60 μM), followed by continuous culture for 120 h. Control groups of ovarian cancer cells were maintained under normal culture conditions. Cell viability was assessed every 24 h using the CCK-8 assay. B, C MDA levels and GSH/GSSG ratios in SREBP1-silenced ovarian cancer cells were measured using commercial assay kits. D Lipid ROS in SREBP1 silenced ovarian cancer cells was detected using the BODIPY C11 fluorescent probe (Scale bar = 20 µm). E Expression of ferroptosis related markers in SREBP1-silenced ovarian cancer cells was analyzed by western blotting. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
SREBP1 knockdown promotes ferroptosis by inhibiting NRF2 in ovarian cancer cells
To further elucidate the mechanism by which SREBP1 silencing induces ferroptosis, we investigated its effect on the nuclear factor erythroid 2-related factor 2 (Nrf2) pathway, a key antioxidant signaling axis known to confer ferroptosis resistance by upregulating cytoprotective targets such as XCT and GPX4 [19, 20] To explore the role of Nrf2 in ovarian cancer, we examined the influence of SREBP1 knockdown on Nrf2 and its downstream effectors. Western blot analysis demonstrated that SREBP1 silencing markedly decreased the expression of Nrf2, as well as its target genes heme oxygenase-1 (HO-1) and NAD(P)H quinone dehydrogenase 1 (NQO1) (Fig. 6A and Supplementary Fig. S1D). By contrast, the opposite phenotype was observed following SREBP1 overexpression (P < 0.05, Supplementary Fig. S4A).
A Expression of Nrf2/HO-1/NQO1 in SREBP1 silenced ovarian cancer cells was analyzed by Western blotting. B Silent SREBP1-expressing ovarian cancer cells were treated with or without TBHQ (40 μM) for 24 h, followed by 48 h of continuous culture. Control cells were cultured under standard conditions. Cell viability was assessed every 24 h using the CCK-8 assay kit. C EdU staining was performed in sh-SREBP1 ovarian cancer cells treated with or without TBHQ (40 μM) for 24 h, with control cells cultured normally (Scale bar = 50 µm). D, E MDA levels and GSH/GSSG ratios were measured in sh-SREBP1 ovarian cancer cells following 24 h treatment with or without TBHQ (40 μM), using commercial assay kits. Control groups were cultured normally. F Detection of lipid reactive oxygen species (ROS) levels in sh-SREBP1 ovarian cancer cells treated with TBHQ (40 μM) or untreated, using the BODIPY C11 fluorescent probe (Scale bar = 20 µm). G Expression of ferroptosis-related markers was examined by western blotting in sh-SREBP1 ovarian cancer cells treated with or without TBHQ (40 μM) for 24 h. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
To assess whether SREBP1 depletion promotes ferroptosis through suppression of the Nrf2 pathway, rescue experiments were conducted using the Nrf2 activator TBHQ. CCK-8 assays revealed that TBHQ partially ameliorated the decrease in cell viability induced by SREBP1 knockdown (Fig. 6B). Similarly, colony formation assays indicated that TBHQ partly revered the impairment of clonogenicity resulting from SREBP1 silencing (Supplementary Fig. S1E). EdU incorporation assays further supported these findings, showing that TBHQ partially reversed the deficit in DNA synthesis capacity (Fig. 6C). Revaluation of ferroptosis markers revealed that TBHQ significantly attenuated lipid peroxidation, as reflected by reduced MDA levels (Fig. 6D), and increased the GSH/GSSG ratio (Fig. 6E) in SREBP1-silenced cells. Additionally, following BODIPY™ 581/591 C11 staining, the green fluorescence intensity was decreased following TBHQ treatment, indicating a reduction in lipid ROS levels (Fig. 6F). Western blot analysis confirmed that TBHQ rescued the expression of the Nrf2, XCT, and GPX4 proteins, which were downregulated upon SREBP1 silencing (Fig. 6G and Supplementary Fig. S1F). Together, these results suggest that SREBP1 knockdown induces ferroptosis in ovarian cancer cells via the suppression of Nrf2 activation.
SREBP1 knockdown promotes Nrf2 ubiquitination by upregulating Keap1
To further investigate how SREBP1 regulates Nrf2, we first assessed whether SREBP1 influences Nrf2 transcription. qPCR analysis showed that SREBP1 silencing did not affect Nrf2 mRNA levels (Fig. 7A), suggesting post-transcriptional regulation. Since Nrf2 translocates to the nucleus upon activation [21], we examined its subcellular localization. IF and nuclear-cytoplasmic fractionation assays revealed that SREBP1 knockdown reduced nuclear Nrf2 accumulation (Fig. 7B, C and Supplementary Fig. S2A), whereas SREBP1 overexpression enhanced it (Supplementary Fig. S4B, C).
A Nrf2 mRNA expression in SREBP1-silenced ovarian cancer cells was measured by qPCR. B IF staining demonstrated Nrf2 expression in SREBP1-silenced ovarian cancer cells (Scale bar = 20 µm). C Subcellular localization of Nrf2 in SREBP1-silenced ovarian cancer cells was analyzed by Western blotting. D Keap1 protein levels in SREBP1-silenced ovarian cancer cells were examined by western blotting. E Nrf2 expression was assessed by Western blotting in sh-SREBP1 ovarian cancer cells following 1 h treatment with or without MG132 (5 μM). Control cells were maintained under normal culture conditions. F In ovarian cancer cells, 10 μM chloramphenicol (CHX) was administered at 0, 15, 30, and 45 min to assess the stability of Nrf2 protein in control or SREBP1-silenced cells, validated by Western blot analysis. G Nrf2 ubiquitination was examined by immunoprecipitation using anti-Nrf2 antibody in SREBP1-silenced ovarian cancer cells, followed by Western blot detection of ubiquitin in the pulldown samples. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
We then explored whether SREBP1 influences Nrf2 stability. Kelch like ECH associated protein 1 (Keap1), a substrate adaptor for the E3 ubiquitin ligase complex, plays a key role in protein ubiquitination [22]. Under normal conditions, Keap1 binds to Nrf2 in the cytoplasm and promotes its degradation via the ubiquitin proteasome pathway, maintaining low Nrf2 expression [23]. Western blot analysis confirmed that SREBP1 silencing increased Keap1 expression (Fig. 7D). Moreover, treatment with the proteasome inhibitor MG132 significantly reversed the decrease in Nrf2 protein levels induced by SREBP1 silencing (Fig. 7E and Supplementary Fig. S2B), suggesting that the reduction in Nrf2 expression is mediated by proteasomal degradation. Subsequently, ovarian cancer cells were treated with the protein synthesis inhibitor cycloheximide (CHX). Western blot analysis revealed that compared with the control treatment, SREBP1 silencing significantly shortened the half-life of the Nrf2 protein (Fig. 7F), confirming that SREBP1 stabilizes the Nrf2 protein in ovarian cancer cells. Given the crucial role of ubiquitination in regulating protein function and stability, the effect of the ubiquitin‒proteasome pathway on Nrf2 was further investigated. Assessment of Nrf2 ubiquitination levels in SREBP1-silenced cells treated with MG132 revealed that compared with the control treatment, treatment with MG132 significantly increased Nrf2 ubiquitination (Fig. 7G).
Meanwhile, the molecular mechanism between SREBP1 and Keap1 was explored. qPCR was performed to detect the effect of SREBP1 silencing on Keap1 mRNA levels. The results showed that silencing SREBP1 increased the expression of Keap1 mRNA (P < 0.05, Supplementary Fig. S4D). Therefore, SREBP1 may bind to the Keap1 promoter, thereby regulating KEAP1 transcription and subsequently affecting protein expression levels. Subsequently, the JASPAR database was used to predict whether SREBP1 has binding sites on the Keap1 promoter (Fig. 8A, B). The results revealed two potential binding sites for SREBP1. Agarose gel electrophoresis was then conducted, and the experimental results indicated that SREBP1 has binding affinity for the two predicted binding sites in the Keap1 promoter region at the chromatin level (Fig. 8C). Finally, a chromatin immunoprecipitation (ChIP) assay was performed to confirm the binding sites, which verified that SREBP1 can bind to both site 1 and site 2 in two cell lines (Fig. 8D). In conclusion, the experimental results indicate that silencing SREBP1 promotes Keap1 transcription, thereby promoting Nrf2 degradation via the ubiquitin proteasome pathway.
A SREBP1 binding site patterns predicted by the JASPAR database. B Two potential binding regions for SREBP1 on the Keap1 promoter, as predicted by the JASPAR database. C Agarose gel electrophoresis results indicate that SREBP1 exhibits chromatin-level binding affinity for two promoter regions of Keap1 in both A2780 and SKOV3 cells. D ChIP-qPCR detection results demonstrate chromatin level binding of SREBP1 to the two Keap1 promoter regions in ovarian cancer cells. Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
SREBP1 knockdown suppresses tumor growth and modulates the immune microenvironment in vivo
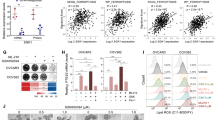
Cancer-intrinsic innate immunity can promote tumor rejection. However, cancer cells can evade these responses by overexpressing immune checkpoint regulators such as PD-L1 (CD274) [24]. Cox regression analysis of risk factors and OS in ovarian cancer patients revealed that PD-L1 positivity (P = 0.032) was closely associated with OS (Table 3), suggesting that upregulation of SREBP1 expression may inhibit immune cell infiltration in tumor tissues. Bioinformatic analysis using the TIMER 2.0 database revealed that SREBP1 expression was negatively correlated with CD4+ T-cell and CD8+ T-cell infiltration (Fig. 9A) and positively correlated with PD-L1 expression (Fig. 9B). Western blot analysis confirmed that knockdown of SREBP1 led to a decrease in PD-L1 expression (Fig. 9C). Conversely, overexpression of SREBP1 produced the opposite effect (Supplementary Fig. S4E). Consistent with these findings, IF assays showed that SREBP1 silencing reduced the PD-L1 fluorescence signals (P < 0.05, Fig. 9D), while SREBP1 overexpression enhanced the signals (Supplementary Fig. S4F).
A Correlation between SREBP1 expression and immune cell infiltration. B Correlation between SREBP1 expression and PD-L1 (CD274) expression. C PD-L1 protein levels in SREBP1-silenced ovarian cancer cells were analyzed by Western blotting. D IF staining showing PD-L1 expression in ovarian cancer cells (Scale bar = 50 µm). E The expression of SREBP1, PD-L1, and CD8+ T cells was evaluated through mIHC staining. F Subcutaneous xenograft mouse models were established by injecting nude mice with ovarian cancer cells (A2780) transfected with either sh-con or sh-SREBP1. G Representative tumor images from xenograft mouse models. H tumor weights from xenograft mouse models. I Expression of SREBP1, Ki67 (proliferation marker), 4HNE (lipid peroxidation marker), GPX4 (ferroptosis marker), Nrf2, and PD-L1 was evaluated by IHC staining (Scale bar = 50 µm or 20 µm). Data are presented as the mean ± SD of three independent experiments. p-values in panels: *p < 0.05, **p < 0.01.
Multiplex immunohistochemical (mIHC) techniques further confirmed the negative correlation between SREBP1 and the tumor immune microenvironment. In human ovarian tumor sections, high SREBP1 expression correlates with PD-L1 upregulation and reduced CD8+ T cells, whereas low SREBP1 expression shows the opposite effect (Fig. 9E). Further validation was performed in vivo. SREBP1 silencing resulted in reduced tumor volume and weight (Fig. 9F, G, H), which is consistent with the in vitro findings. IHC staining of xenograft tumor sections revealed that knockdown decreased the expression of the proliferation-related antigen Ki67, the ferroptosis-related proteins Nrf2 and GPX4, and the immune-related protein PD-L1, and increased the expression of the lipid peroxidation marker 4-HNE (Fig. 9I). In summary, these results indicate that silencing SREBP1 reduces tumor growth in mice by inducing ferroptosis and inhibiting immune evasion.
Discussion
SREBP1 is a master regulator of cellular lipid metabolism, influencing the synthesis and uptake of both cholesterol and fatty acids [25]. Previous studies have demonstrated that SREBP1 is highly expressed in various tumor tissues, including gastric [26], colorectal [27], and pancreatic cancers [12], as well as in HCC, as previously reported by our research group [28]. Zhang et al. reported that SREBP1 is associated with cell proliferation and EMT, and that its high expression promotes the progression and metastasis of breast cancer [29]. In a study by Pan et al., the small molecule GANT61 was found to upregulate SREBP1, leading to lipid droplet accumulation and enhanced proliferation in ovarian cancer cells [30]. Similarly, Lu et al. reported that silencing SREBP1 inhibits the malignant progression of nasopharyngeal carcinoma by suppressing de novo fatty acid synthesis [31]. These results suggest that cancer cells can upregulate SREBP1 expression to accelerate lipid synthesis, meeting the energy and material demands for rapid proliferation and thereby promoting malignant progression and poor prognosis [32]. Here, we confirmed that silencing SREBP1 inhibits the proliferative capacity of ovarian cancer cells by inducing cell cycle arrest. Concurrently, silencing SREBP1 attenuated the migratory and invasive abilities of ovarian cancer cells, as indicated by upregulated expression of the epithelial marker E-cadherin and downregulated expression of the mesenchymal markers Vimentin and Snail. Furthermore, SREBP1 silencing led to a significant reduction in intracellular TG and TC levels, accompanied by decreased expression of lipid metabolism-related proteins. This indicates that inhibiting SREBP1 expression can reduce the lipid synthesis capacity of ovarian cancer cells, decreasing their energy reserves. These findings suggest that targeting SREBP1 inhibits the proliferation and metastasis of ovarian cancer cells by suppressing lipid metabolic reprogramming.
Studies have shown that tumor cells undergoing lipid metabolic reprogramming and EMT alterations are more susceptible to ferroptosis, elucidating the specific conditions required to induce ferroptosis in cancer [33]. Ferroptosis is a distinct type of regulated cell death characterized by iron accumulation and lipid peroxidation damage [34]. It is now widely accepted that inducing ferroptosis can inhibit cancer progression, and this approach is considered a promising strategy for cancer therapy [35]. For instance, Zheng et al. demonstrated that a PI3Kδ inhibitor induces ferroptosis in KRAS-mutant colorectal cancer cells by inhibiting AKT/mTOR/SREBP1-mediated lipid metabolism [27]. Chen et al. showed that combination therapy with β-elemene and cetuximab induces ferroptosis in KRAS-mutant colorectal cancer cells by suppressing EMT [36]. This study experimentally confirmed that silencing SREBP1 reduces MDA levels, inhibits GSH synthesis, and downregulates the expression of the key ferroptosis-related proteins XCT and GPX4, ultimately leading to lipid peroxidation damage and the induction of ferroptosis in ovarian cancer cells. Additionally, we found that the expression of the key antioxidant transcription factor Nrf2 is significantly downregulated following SREBP1 silencing [37]. As a crucial antioxidant system, the Nrf2 pathway responds to oxidative stress and protects cells from oxidative damage [38], and its suppression by SREBP1 knockdown may directly weaken the cellular antioxidant capacity. Under normal conditions, Nrf2 undergoes ubiquitin-mediated degradation in a KEAP1-dependent manner. However, upon oxidative stress, Nrf2 is released from KEAP1, translocates to the nucleus [39], and transcriptionally promotes the expression of several antioxidant proteins and enzymes, including NQO1 and HO-1 [40]. Notably, evidence indicates that Nrf2 can protect cells from ferroptosis by modulating the system XCT/GPX4 axis. Nrf2 directly binds to the XCT promoter and transcriptionally upregulates its expression, subsequently increasing cysteine uptake and GSH synthesis. In this study, rescue experiments using the Nrf2 activator TBHQ demonstrated that the SREBP1 silencing-mediated downregulation of Nrf2 expression, reductions in XCT/GPX4 expression, an increase in MDA levels, inhibition of GSH synthesis, and induction ferroptosis in ovarian cancer cells could be reversed by TBHQ. TBHQ inhibited the ferroptotic response induced by silencing SREBP1.
To further elucidate the molecular mechanism by which SREBP1 regulates Nrf2, experimental data confirmed that although Nrf2 mRNA levels remained unchanged upon SREBP1 silencing, Nrf2 protein expression was significantly reduced. This indicates that SREBP1 depletion impairs Nrf2 protein stability rather than suppressing its transcription. Consistent with this, the expression of Keap1, the E3 ubiquitin ligase of Nrf2, was increased following SREBP1 knockdown. Subsequently, it was observed that the downregulation of Nrf2 protein induced by SREBP1 silencing could be reversed by the proteasome inhibitor MG132. Furthermore, CHX chase assays demonstrated that SREBP1 silencing shortened the half-life of Nrf2 protein, and elevated Nrf2 ubiquitination was detected in the knockdown group. As SREBP1 is a transcription factor, we hypothesized that it binds to the Keap1 promoter and represses its transcriptional activity. To investigate this, we predicted two potential SREBP1 binding sites within the Keap1 promoter using the JASPAR database. Importantly, ChIP assays confirmed the functional binding of SREBP1 to these sites. Collectively, our findings demonstrate that SREBP1 silencing enhances Keap1 transcription, which in turn promotes the ubiquitin-proteasomal degradation of Nrf2, leading to its downregulation and the subsequent induction of ferroptosis.
In addition to regulating metabolic reprogramming and redox balance, our study suggests that SREBP1 may influence immune evasion mechanisms in ovarian cancer. To date, ovarian tumors are generally considered to have low immunoreactivity, partly because tumor cells express insufficient amounts of neoantigens for effective recognition by the immune system, facilitating immune evasion [41]. The PD-1/PD-L1 pathway constitutes a crucial mechanism for the maintenance of immune homeostasis under physiological conditions [42]. The PD-1/PD-L1 interaction reduces autoimmune attacks of tissues by inhibiting T-cell activation, thereby maintaining immune balance [43]. However, in the TME, the binding of PD-1 on tumor-infiltrating lymphocytes (TILs) to PD-L1 on tumor cells inhibits lymphocyte proliferation and activation, thereby reducing immune capacity and inducing immune evasion [44, 45]. Our experimental models confirmed a positive correlation between SREBP1 and PD-L1 expression. Moreover, mIHC revealed an inverse relationship between SREBP1 levels and CD8+ T-cell infiltration, indicating an immunosuppressive microenvironment. The critical role of SREBP1 was further substantiated by in vivo xenograft experiments, wherein its silencing markedly reduced tumor growth. IHC analysis demonstrated that the knockdown group exhibited a concurrent downregulation of both PD-L1 and key ferroptosis-related proteins. These collective findings suggest that SREBP1 targeting concurrently disrupts immune evasion via PD-L1 reduction and triggers ferroptosis, providing a compelling rationale for combining ferroptosis inducers with ICIs to achieve superior antitumor efficacy [46, 47].
In conclusion, this study establishes the critical oncogenic role of SREBP1 in ovarian cancer and reveals that the induction of ferroptosis is a key mechanism through which SREBP1 silencing suppresses tumor progression. Specifically, silent SREBP1 promotes its expression by upregulating the transcriptional levels of Keap1, thereby facilitating the ubiquitin-proteasomal degradation of Nrf2. The downregulation of Nrf2 leads to reduced expression of key ferroptosis defense factors, XCT and GPX4, thereby sensitizing ovarian cancer cells to ferroptotic cell death. Moreover, this study demonstrates that SREBP1 integrates Keap1/Nrf2-mediated ferroptosis resistance with PD-L1-associated immune suppression, providing a strong mechanistic rationale for targeting SREBP1. Small molecule inhibitors targeting SREBP1 not only hold therapeutic potential on their own but may also synergistically enhance antitumor efficacy when combined with ferroptosis inducers or ICIs. Although the precise transcriptional network between SREBP1 and PD-L1 requires further elucidation, our findings consistently position SREBP1 as a promising therapeutic target and a potential prognostic biomarker in ovarian cancer. Targeted inhibition of SREBP1 is expected to trigger ferroptosis and may ultimately improve treatment responses and survival outcomes for patients.
Materials and methods
Patient specimens
An ovarian cancer tissue microarray was acquired from Shanghai Outdo Biotech Company, comprising 132 ovarian cancer and 40 normal ovarian tissue samples. Clinicopathological parameters, including age, clinical stage, tumor grade, size, lymph node metastasis, hematogenous metastasis, recurrence, Ki67 positivity, and PD-L1 positivity, were documented after securing written informed consent. All tissue specimens were sourced from partner hospitals in consultation with pathologists. Diagnoses and staging adhered to the 8th edition of the AJCC Cancer Staging Manual. The Institutional Review Board of Shanghai Outdo Biotech Company granted ethical approval for this study. (IRB Approval No.: SHYJS-CP-1804011 and SHYJS-CP-1804032).
Cell lines and reagents
Human ovarian cancer cell lines (ES-2, A2780, OVCA429, CAOV3 and SKOV3) were obtained from the American Type Culture Collection (ATCC, Rockville, MD, USA) and have undergone STR profiling and mycoplasma testing. Antibodies targeting SREBP1 (66875-1-Ig), CDK4 (11026-1-AP), CDK6 (14052-1-AP), Cyclin A2 (18202-1-AP), Cyclin D1 (26939-1-AP), Cyclin B (55004-1-AP), E-cadherin (20874-1-AP), Vimentin (10366-1-AP), Snail (12129-1-AP), GPX4 (67763-1-Ig), XCT (26864-1-AP), Nrf2 (16396-1-AP), PD-L1 (65685-1-MR), and β-tubulin (66240-1-Ig) were procured from Proteintech. Immunohistochemistry reagents, including the SP two-step detection kit, citrate buffer, PBST, DAB substrate, and hematoxylin, were sourced from Beijing Zhongshan Jinqiao Biotechnology. Cell culture supplements fetal bovine serum, DMEM, penicillin, streptomycin, trypsin, and serum-free cryopreservation medium were obtained from Gibco.
Cell culture and transfection
ES-2, A2780, OVCA429, CAOV3, and SKOV3 cells were maintained in DMEM supplemented with 10% fetal bovine serum and penicillin/streptomycin (100 U/mL). Upon reaching 80% confluence, cells were trypsinized, centrifuged, and resuspended. For transfection, cells were plated in 6-well plates at 3 × 10⁴ cells per well and transduced with lentiviral vectors according to manufacturer's protocols to generate SREBF1-knockdown (Sh-SREBF1) and control groups. Stable lines were selectively cultured using puromycin (2 μg/mL), and silencing efficiency was confirmed via western blot prior to experimental use. Lentiviral sequences are provided in Table 4.
IHC staining
IHC was conducted via the SP two-step method. Tissue sections were baked at 65 °C for 1 h, deparaffinized in xylene, and rehydrated through a graded ethanol series. Antigen retrieval was performed by microwave heating in citrate buffer for 5 min. Primary antibodies were applied at optimized dilutions and incubated overnight at 4 °C. After secondary antibody incubation, the signal was developed using DAB, and sections were counterstained with hematoxylin before mounting.
Western blot
Cells were lysed in RIPA buffer containing protease and phosphatase inhibitors. Protein concentrations were determined with a microplate reader. Samples were denatured in loading buffer, separated on 8% SDS-PAGE gels, and transferred to activated PVDF membranes. After blocking with 5% skim milk, membranes were probed with primary antibodies at 4 °C overnight. Following TBST washes, horseradish peroxidase-conjugated secondary antibodies were applied for 1 h at room temperature. Protein bands were visualized using chemiluminescence.
CCK-8 assay
Cells were seeded in 96-well plates at 2 × 10³ cells per well. After 24, 48, 72, and 96 h, medium was replaced with a CCK-8 dilution (DMEM: CCK-8 = 9:1). Following 2 h of incubation, absorbance was measured at 450 nm.
Colony formation assay
A total of 1 × 10⁴ cells were plated per well in 6-well plates and cultured for 14 days. Colonies were fixed with 4% paraformaldehyde, stained with crystal violet, and imaged after drying.
EdU assay
Cells were seeded in 96-well plates at 2 × 10⁴ cells per well. After overnight culture, EdU reagent was introduced for 2 h. Cells were fixed, permeabilized, and stained using the Apollo kit. Nuclei were counterstained with Hoechst 33342, and EdU-positive cells were visualized by fluorescence microscopy.
Cell cycle analysis by flow cytometry
Cells were harvested, fixed in ethanol, and treated with PI staining solution. DNA content was analyzed via flow cytometry to determine cell cycle distribution.
Wound healing assay
Cells were plated in 6-well plates at 5 × 10⁴ cells per well. A scratch was made using a pipette tip upon confluence. Wound closure was monitored and imaged at 0, 24, and 48 h.
Transwell assay
For migration assays, 1 × 10⁵ cells in serum-free DMEM were added to the upper chamber, with 20% FBS-DMEM below. After 24 h, migrated cells were fixed and stained. For invasion assays, Matrigel was pre-coated in the upper chamber before cell seeding.
IF assay
Place glass coverslips in a 6-well plate and seed 3 × 10⁵ cells per well. After the cells adhere, remove the plate from the incubator and aspirate the culture medium. Add 2 ml of 4% paraformaldehyde and fix the cells at room temperature for 20 min, then aspirate the fixative. Add PBST containing 0.1% Triton X-100 for permeabilization at room temperature for 10 min, then aspirate the permeabilization solution. Apply a blocking solution of 5% BSA and block at room temperature for 1 h. Dilute the primary antibody according to the manufacturer’s recommended ratio using the blocking solution, apply the diluted primary antibody to the coverslips, and incubate overnight at 4 °C. Afterwards, aspirate the primary antibody. Apply the fluorescently labeled secondary antibody diluted in PBS. Incubate at room temperature for 1 h protected from light. Aspirate the secondary antibody. Place 20 µl of an anti-fade mounting medium containing Hoechst 33342 onto a glass slide. Using forceps, carefully place the coverslip with the cell side facing down onto the mounting medium. Seal the edges of the coverslip with clear nail polish. Observe the prepared slides under a fluorescence microscope (FV3000).
MDA assay
0.37% TBA stock solution was formulated, and the MDA working solution was prepared in a ratio of TBA diluent to TBA stock to antioxidant of 150:50:3. After digestion and centrifugation, cells were maintained on ice. Then, 100 µL of cell lysate was combined with 200 µL of the MDA working solution, heated to 100 °C for 15 min, and centrifuged at 1000 × g for 10 min. Subsequently, 200 µL of supernatant was transferred to a 96-well plate for absorbance measurement at 532 nm.
GSH/GSSG assay
Working solutions for total glutathione detection, protein removal reagent M, 0.5 mg/mL NADPH, GSSG stock, diluted GSH scavenger auxiliary solution, and GSH scavenger working solution were prepared following the kit instructions. Cells were exposed to reagent M and flash-frozen in liquid nitrogen. The supernatant was collected after centrifugation. A subset of samples underwent treatment with GSH scavenger auxiliary and working solutions for 1 h. These samples, along with the total glutathione working solution, were placed in a 96-well plate, incubated for 5 min, mixed with NADPH, and further incubated for 25 min before reading absorbance at 412 nm.
BODIPY 581/591 C11 staining
Cells were plated at 3 × 10⁶ per dish and incubated for one hour with BODIPY 581/591 C11 probe diluted in DMEM at a 2:1 ratio. After gentle PBS washing, fluorescence was visualized using a fluorescence microscope.
TG assay
The TG working solution was prepared according to the manufacturer’s protocol. A total of 1 × 10⁶ cells were lysed with 100 µL of isopropanol and centrifuged at 13,000 rcf for 5 min at 4 °C. After appropriate dilution, samples were mixed with lipase in a 96-well plate and reacted for 20 min. TG working solution was then introduced and incubated for 60 min prior to measuring absorbance at 570 nm.
TC assay
TC working solution was configured as directed. Cells (1 × 10⁶) were lysed in 100 µL of TC assay buffer and centrifuged (13,000 rcf, 4 °C, 5 min). The diluted lysates were mixed with TC working solution, incubated for 30 min, and the absorbance was recorded at 570 nm.
Nile red staining
Staining solution for lipid droplets was prepared as recommended. Cells were seeded in 6-well plates at 1 × 10⁶ cells per well, fixed with 4% paraformaldehyde for 15 min, and stained in the dark with Nile Red Staining solution for 20 min. Red fluorescence was examined under a fluorescence microscope with 522 nm excitation.
Nuclear-cytoplasmic fractionation assay
Cells were harvested and lysed in cytoplasmic extraction reagent A containing PMSF (100:1), vortexed, and placed on ice for 10 min. Then, 10 µL of cytoplasmic extraction reagent B was introduced, followed by vortexing and incubation on ice for one min. After centrifugation (16,000 × g, 5 min, 4 °C), the supernatant (cytoplasmic fraction) was collected. The pellet was resuspended in nuclear extraction reagent with PMSF (100:1), vortexed, and kept on ice for 10 min; this step was repeated three times. The final supernatant (nuclear fraction) was obtained after centrifugation.
qPCR assay
Total RNA was isolated with Trizol (Life Technologies), and cDNA was synthesized using SuperScript III First-Strand Synthesis SuperMix (Invitrogen). Real-time PCR was carried out with Sybergreen Master Mix. Relative mRNA levels were determined via the 2−ΔΔCT method, using β-tubulin for normalization.
CHX chase assay
At 80% confluency, cells were exposed to cycloheximide (CHX, 10 µg/mL) and collected at 0, 15, 30, and 45 min. Cells were lysed in RIPA buffer supplemented with protease inhibitors and centrifuged (16,000 rcf, 10 min, 4 °C). Protein concentration was quantified, and degradation was evaluated by Western blot.
Protein ubiquitination assay
Cells were pre-treated with 10 µM MG132 for 6 h, lysed in IP buffer, and quantified. Lysates were immunoprecipitated with target-specific antibodies or control IgG overnight at 4 °C. Magnetic beads were blocked in 0.2% BSA and incubated with lysates for 6 h at room temperature. After washing with IP buffer, beads were resuspended in 2× SDS loading buffer, boiled for 5 min, and ubiquitination was detected via Western blot with an anti-ubiquitin antibody.
ChIP assay
When ovarian cancer cells reached 90% confluency, they were cross-linked with formaldehyde and lysed. The DNA was then fragmented into 100–500 bp fragments using sonication. Nuclear lysates were immunoprecipitated with specific antibodies or normal mouse IgG. The purified enriched sequences were detected by qPCR.
mIHC assay
Paraffin-embedded tissue sections were baked in a 60°C oven for 1 h. Deparaffinization and rehydration were performed using xylene and a graded ethanol series (100%, 95%, 85%, and 70%). Antigen retrieval was conducted using a sodium citrate antigen retrieval buffer. Endogenous peroxidase activity was blocked with a 3% hydrogen peroxide solution. The primary antibody was incubated at 4 °C overnight, followed by a secondary antibody incubation at room temperature for 1 h. The corresponding fluorophore-conjugated tyramide signal amplification (TSA) reagent was applied and incubated at room temperature for 10 min, protected from light. Quenching and antigen retrieval were performed again using the sodium citrate buffer. This “staining-quenching” cycle was repeated three times. A DAPI working solution was added and incubated at room temperature for 5–10 min, protected from light. Finally, the sections were mounted with an anti-fade mounting medium. Images were acquired using a confocal microscope (VS200).
Subcutaneous xenograft assay
Four-week-old BALB/c nude mice were supplied by Jiangsu Gem Pharmatech Co., Ltd and randomly allocated into two groups (n = 6 per group). Mice received subcutaneous injections of 200 µL PBS containing 1 × 10⁷ sh-con A2780 or sh-SREBP1#2 A2780 cells. Four weeks post-injection, all mice were euthanized, and tumors were harvested and weighed for subsequent analysis. Tumor volume calculation formula as follows: 0.5 × [long diameter] × [short diameter]2. All animal procedures were approved by the Animal Ethics Committee of Yanbian University (Approval Nos.: YD20250710017, SCXK (JI) 2020-0009) and conducted in accordance with ARRIVE guidelines.
Statistical analysis
Data analysis was performed using SPSS 27.0. Associations between clinicopathological variables were evaluated with χ² or Fisher’s exact tests. Statistical analysis of the research results was performed using GraphPad Prism 10.0 software. T-tests were used for comparisons between two groups, while one-way ANOVA analyses were employed for comparisons between multiple groups. The data are presented as mean ± standard deviation. Statistical significance was considered at p < 0.05.
Data availability
All data supporting the findings of this study are included in the article and its supplementary information files.
References
Sun G, Liu Y. Tertiary lymphoid structures in ovarian cancer. Front Immunol. 2024;15:1465516.
Chen Z, Xu L, Shi W, Zeng F, Zhuo R, Hao X, et al. Trends of female and male breast cancer incidence at the global, regional, and national levels, 1990-2017. Breast Cancer Res Treat. 2020;180:481–90.
Yang L, Xie HJ, Li YY, Wang X, Liu XX, Mai J. Molecular mechanisms of platinum‑based chemotherapy resistance in ovarian cancer (Review). Oncol Rep. 2022;47:82.
Konstantinopoulos PA, Matulonis UA. Clinical and translational advances in ovarian cancer therapy. Nat Cancer. 2023;4:1239–57.
Zhang M, Yin R, Li K. Advances and challenges in the origin and evolution of ovarian cancer organoids. Front Oncology. 2024;14:1429141.
Ji Z, Shen Y, Feng X, Kong Y, Shao Y, Meng J, et al. Deregulation of lipid metabolism: the critical factors in ovarian cancer. Front Oncol. 2020;10:593017.
Wang H, Luo S, Yin Y, Liu Y, Sun X, Qiu L, et al. DLAT is involved in ovarian cancer progression by modulating lipid metabolism through the JAK2/STAT5A/SREBP1 signaling pathway. Cancer Cell Int. 2025;25:25.
Li Y, Gao A, Zeng T, Liu D, Zhang Q, Ran X, et al. ANGPTL4 accelerates ovarian serous cystadenocarcinoma carcinogenesis and angiogenesis in the tumor microenvironment by activating the JAK2/STAT3 pathway and interacting with ESM1. J Transl Med. 2024;22:46.
Chen M, Zhang J, Sampieri K, Clohessy JG, Mendez L, Gonzalez-Billalabeitia E, et al. An aberrant SREBP-dependent lipogenic program promotes metastatic prostate cancer. Nat Genet. 2018;50:206–18.
Yu HC, Jin L, Bai L, Zhang YJ, Yang ZX. C12ORF49 inhibits ferroptosis in hepatocellular carcinoma cells via reprogramming SREBP1/SCD1-mediated lipid metabolism. Cell Death Discov. 2025;11:178.
Lounis MA, Bergeron KF, Burhans MS, Ntambi JM, Mounier C. Oleate activates SREBP-1 signaling activity in SCD1-deficient hepatocytes. Am J Physiol Endocrinol Metab. 2017;313:E710–E720.
Cao R, Feng Z, Mo J, Wu J, Li J, Li W, et al. Pharmacological inhibition of SREBP1 suppresses pancreatic cancer growth via inducing GPX4-mediated ferroptosis. Cell Signal. 2024;124:111381.
Yu H, Kou Q, Yuan H, Qi Y, Li Q, Li L, et al. Alkannin triggered apoptosis and ferroptosis in gastric cancer by suppressing lipid metabolism mediated by the c-Fos/SREBF1 axis. Phytomedicine. 2025;140:156604.
Khan A, Huo Y, Guo Y, Shi J, Hou Y. Ferroptosis is an effective strategy for cancer therapy. Med Oncol. 2024;41:124.
Ye C, Yao Z, Wang Y, Zhang C. Asiaticoside promoted ferroptosis and suppressed immune escape in gastric cancer cells by downregulating the Wnt/β-catenin pathway. Int Immunopharmacol. 2024;134:112175.
Zhang D, Man D, Lu J, Jiang Y, Ding B, Su R, et al. Mitochondrial TSPO promotes hepatocellular carcinoma progression through ferroptosis inhibition and immune evasion. Adv Sci. 2023;10:2206669.
Suski JM, Braun M, Strmiska V, Sicinski P. Targeting cell-cycle machinery in cancer. Cancer Cell. 2021;39:759–78.
Ye Y, Xie X, Bi Y, Liu Q, Qiu L, Zhao H, et al. Nrf2 alleviates acute ischemic stroke induced ferroptosis via regulating xCT/GPX4 pathway. Free Radic Biol Med. 2025;231:153–62.
Dodson M, Castro-Portuguez R, Zhang DD. NRF2 plays a critical role in mitigating lipid peroxidation and ferroptosis. Redox Biol. 2019;23:101107.
Qu Z, Zeng J, Zeng L, Li X, Zhang F. Esculetin triggers ferroptosis via inhibition of the Nrf2-xCT/GPx4 axis in hepatocellular carcinoma. Chin J Nat Med. 2025;23:443–56.
Rojo de la Vega M, Chapman E, Zhang DD. NRF2 and the hallmarks of cancer. Cancer Cell. 2018;34:21–43.
Han B, Zhen F, Sun Y, Sun B, Wang HY, Liu W, et al. Tumor suppressor KEAP1 promotes HSPA9 degradation, controlling mitochondrial biogenesis in breast cancer. Cell Rep. 2024;43:114507.
Yu C, Xiao JH. The Keap1-Nrf2 system: a mediator between oxidative stress and aging. Oxid Med Cell Longev. 2021;2021:6635460.
Leuzzi G, Vasciaveo A, Taglialatela A, Chen X, Firestone TM, Hickman AR, et al. SMARCAL1 is a dual regulator of innate immune signaling and PD-L1 expression that promotes tumor immune evasion. Cell. 2024;187:861–81.e32.
He Y, Qi S, Chen L, Zhu J, Liang L, Chen X, et al. The roles and mechanisms of SREBP1 in cancer development and drug response. Genes Dis. 2024;11:100987.
Chen M, Li H, Zheng S, Shen J, Chen Y, Li Y, et al. Nobiletin targets SREBP1/ACLY to induce autophagy-dependent cell death of gastric cancer cells through PI3K/Akt/mTOR signaling pathway. Phytomedicine. 2024;128:155360.
Zheng YN, Lou SY, Lu J, Zheng FL, Tang YM, Zhang EJ, et al. Selective PI3Kδ inhibitor TYM-3-98 suppresses AKT/mTOR/SREBP1-mediated lipogenesis and promotes ferroptosis in KRAS-mutant colorectal cancer. Cell Death Dis. 2024;15:474.
Wang X, Liu Y, Han A, Tang C, Xu R, Feng L, et al. The NQO1/p53/SREBP1 axis promotes hepatocellular carcinoma progression and metastasis by regulating Snail stability. Oncogene. 2022;41:5107–0.
Zhang N, Zhang H, Liu Y, Su P, Zhang J, Wang X, et al. SREBP1, targeted by miR-18a-5p, modulates epithelial-mesenchymal transition in breast cancer via forming a co-repressor complex with Snail and HDAC1/2. Cell Death Differ. 2019;26:843–59.
Pan Y, Chen L, Shen J, Hong S, Guan X, Ma X, et al. GANT61 modulates autophagy and lipid metabolism in ovarian cancer. Cell Prolif. 2025;58:e70051.
Lo AK-F, Lung RW-M, Dawson CW, Young LS, Ko C-W, Yeung WW, et al. Activation of sterol regulatory element-binding protein 1 (SREBP1)-mediated lipogenesis by the Epstein-Barr virus-encoded latent membrane protein 1 (LMP1) promotes cell proliferation and progression of nasopharyngeal carcinoma. J Pathol. 2018;246:180–90.
Wu R, Li N, Huang W, Yang Y, Zang R, Song H, et al. Melittin suppresses ovarian cancer growth by regulating SREBP1-mediated lipid metabolism. Phytomedicine. 2025;137:156367.
Zhou Q, Meng Y, Li D, Yao L, Le J, Liu Y, et al. Ferroptosis in cancer: From molecular mechanisms to therapeutic strategies. Signal Transduct Target Ther. 2024;9:55.
Iang X, Stockwell BR, Conrad M. Ferroptosis: mechanisms, biology and role in disease. Nat Rev Mol Cell Biol. 2021;22:266–82.
Mou Y, Wang J, Wu J, He D, Zhang C, Duan C, et al. Ferroptosis, a new form of cell death: opportunities and challenges in cancer. J Hematol Oncol. 2019;12:34.
Chen P, Li X, Zhang R, Liu S, Xiang Y, Zhang M, et al. Combinative treatment of β-elemene and cetuximab is sensitive to KRAS mutant colorectal cancer cells by inducing ferroptosis and inhibiting epithelial-mesenchymal transformation. Theranostics. 2020;10:5107–19.
Basak P, Sadhukhan P, Sarkar P, Sil PC. Perspectives of the Nrf-2 signaling pathway in cancer progression and therapy. Toxicol Rep. 2017;4:306–18.
Bae T, Hallis SP, Kwak M-K. Hypoxia, oxidative stress, and the interplay of HIFs and NRF2 signaling in cancer. Exp Mol Med. 2024;56:501–14.
Zhou Y, Chen Y, Shi Y, Wu L, Tan Y, Li T, et al. FAM117B promotes gastric cancer growth and drug resistance by targeting the KEAP1/NRF2 signaling pathway. J Clin Investig. 2023;133:e158705.
Guo WY, Wu QM, Zeng HF, Chen YL, Xu J, Yu ZY, et al. A sinomenine derivative alleviates bone destruction in collagen-induced arthritis mice by suppressing mitochondrial dysfunction and oxidative stress via the NRF2/HO-1/NQO1 signaling pathway. Pharm Res. 2025;215:107686.
Khatoon E, Parama D, Kumar A, Alqahtani MS, Abbas M, Girisa S, et al. Targeting PD-1/PD-L1 axis as new horizon for ovarian cancer therapy. Life Sci. 2022;306:120827.
Jiang Y, Chen M, Nie H, Yuan Y. PD-1 and PD-L1 in cancer immunotherapy: clinical implications and future considerations. Hum Vaccines Immunother. 2019;15:1111–22.
Dermani FK, Samadi P, Rahmani G, Kohlan AK, Najafi R. PD-1/PD-L1 immune checkpoint: potential target for cancer therapy. J Cell Physiol. 2019;234:1313–25.
Petrovas C, Casazza JP, Brenchley JM, Price DA, Gostick E, Adams WC, et al. PD-1 is a regulator of virus-specific CD8+ T cell survival in HIV infection. J Exp Med. 2006;203:2281–92.
Kuol N, Stojanovska L, Nurgali K, Apostolopoulos V. PD-1/PD-L1 in disease. Immunotherapy. 2018;10:149–60.
Kwon JE, Jang Y, Yun BS, Kang S, Kim YH, Kim BG, et al. MET overexpression in ovarian cancer via CD24-induced downregulation of miR-181a: a signalling for cellular quiescence-like state and chemoresistance in ovarian CSCs. Cell Prolif. 2024;57:e13582.
Yu L, Huang K, Liao Y, Wang L, Sethi G, Ma Z. Targeting novel regulated cell death: ferroptosis, pyroptosis and necroptosis in anti-PD-1/PD-L1 cancer immunotherapy. Cell Prolif. 2024;57:e13644.
Funding
This research was supported by the Projects of Science and Technology Department of Jilin Province (No.202002021JC) and The National Natural Science Foundation of China (No.82360576).
Author information
Authors and Affiliations
Contributions
Rui Nie and Hou-kun Zhou were involved in the conception and design of the study; Rui Nie and Hai-yan Quan performed in vitro experiments and drafting of the manuscript; Lin Chen and Yuan Zhang performed in vivo experiments and data analysis;Rui Nie, Hou-kun Zhou, and Zhen-hua Lin were manuscript revisions. All authors listed approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
The Shanghai Outdo Biotech Co., Ltd.’s Institutional Review Board (IRB) examined and approved the research procedure. The IRB approval number for this study is SHYJS-CP-1804011 and SHYJS-CP-1804032. Animal experiments were approved by Yanbian University’s Ethics Committee (Approval No. YD20250710017) and conducted in strict accordance with institutional and national guidelines for the care and use of laboratory animals. All methods are compliant with the ARRIVE guidelines.
Informed consent
Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nie, R., Zhou, H., Chen, L. et al. SREBP1 knockdown triggers ferroptosis by suppressing the Nrf2-XCT/GPX4 axis in ovarian cancer. Cell Death Discov. 12, 101 (2026). https://doi.org/10.1038/s41420-026-02964-9
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41420-026-02964-9