Abstract
Group 3 innate lymphoid cells (ILC3s) play crucial roles in maintaining intestinal homeostasis and defending against bacterial infections. However, the epigenetic mechanisms that regulate ILC3 responses are not well understood. In this study, we show that Trmt61a, the methyltransferase responsible for the m1A58 tRNA modification, is predominantly expressed in ILC3s. We found that specific depletion of TRMT61A in ILC3s leads to dysregulated cell cycle and a reduction in cell numbers. Notably, mice with an ILC3-specific TRMT61A deficiency exhibit dysbiosis, but antibiotic treatment can restore colonic ILC3 levels. Furthermore, these mice exhibit increased susceptibility to experimental intestinal inflammation and enteric bacterial infection. Our findings uncover a previously unrecognized role for TRMT61A mediated m1A modification in the regulation of intestinal ILC3s, essential for protecting intestinal tissue during inflammation and enhancing innate immunity against enteric pathogens.
Similar content being viewed by others
Introduction
Innate lymphoid cells (ILCs) constitute a family of innate immune cells lacking antigen-specific receptors characteristic of T cells and B cells1,2,3,4. These cells are categorized into three groups based on their master transcription factors and cytokine profiles. Group 1 ILCs (ILC1s) express T-bet and produce IFN-γ, group 2 ILCs (ILC2s) express GATA3 and secrete type 2 cytokines such as interleukin (IL)-5 and IL-13, while group 3 ILCs (ILC3s) express RORγt and generate IL-17 and IL-221,2,3,4. ILC3s are primarily found in mucosal tissues like the intestinal lamina propria. Unlike T cells, which require priming and migration to inflamed tissues5, ILC3s are tissue resident6 and are quickly activated by pro-inflammatory cytokines such as IL-23, IL-1β and TNF-like factor 1A (TL1A), secreted by mononuclear phagocytes6,7,8,9,10
ILC3s are critical in intestinal immunity, particularly during injury and inflammation8,9,11. They produce IL-22, which activates the STAT3 signaling in epithelial cells, enhancing their regeneration and proliferation11. In the initial stages of bacterial infections, ILC3s are the primary source of IL-22, which induces the expression of antimicrobial peptides such as RegIIIβ and RegIIIγ, providing protection against the infection12,13,14. This process is intricately modulated at both transcriptional levels and epigenetic levels, including DNA and histone modifications. Research has highlighted the role of DNA hydroxymethylase tet methylcytosine dioxygenase 2 (TET2) in controlling ILC3 functions through the distribution of methyl-CpG and 5- hydroxymethylcytosine (5hmC)15. Additionally, the histone H3K36 methyltransferase SET domain containing 2 (SETD2) affects genome accessibility, impacting the maintenance and function of ILC3 subsets16. Yet, the impact of RNA modifications on ILC3s is less understood.
Among the more than 100 identified post-transcriptional RNA modifications, N1- methyladenosine (m1A) methylation is one of the most conserved and abundant17,18. m1A modification, particularly m1A58 in tRNA, is crucial for maintaining tRNA stability and enhancing protein translation initiation and elongation19. The m1A methyltransferase complex, consisting of the catalytic subunit TRMT61A and the non- catalytic subunit TRMT6, catalyzes this modification, which could be reversed by the demethylase ALKBH120. While m1A modifications are linked to the progression of various cancers21,22,23, their roles in the immune system, particularly within ILC3s, remain poorly defined.
In this study, we demonstrated that Trmt61a is primarily expressed by intestinal ILC3s. Genetic deletion of ILC3-intrinsic TRMT61A led to reduced numbers of total and IL-22-producing ILC3s, alongside dysregulated cell proliferation and apoptosis. Furthermore, dysbiosis in Trmt61aΔRorc mice, which could be largely corrected with antibiotics, exacerbated these effects. These findings underscore a novel role for TRMT61A mediated tRNA m1A modification in regulating ILC3s, essential for their protective immunity in intestinal health and defense against bacterial infections.
Results
Trmt61a expression and its role in ILC3 homeostasis in the intestine
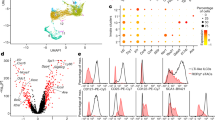
To investigate the role of tRNA m1A58 modification in the immune system, we first reanalyzed existing single-cell RNA sequencing (scRNA-seq) data of CD45+ immune cells isolated from the small intestine24. We found that Trmt61a was primarily expressed in ILC3s, dendritic cells (DCs), and plasmacytoid DCs (pDCs), with the highest expression levels observed in ILC3s (Supplementary Fig. S1a–c).
To study the function of Trmt61a and tRNA m1A modification in ILC3 biology, we crossed Trmt61afl/fl mice with Rorccre mice to specifically delete Trmt61a expression in ILC3s. The resulting RorccreTrmt61afl/fl mice, referred to as Trmt61aΔRorc mice, exhibited significantly reduced expression of Trmt61a in ILC3s from both small intestine and colon (Fig. 1a; Supplementary Fig. S2a). Notably, even under steady-state conditions, Trmt61aΔRorc mice already displayed significantly decreased percentages and numbers of ILC3s in the intestine compared to control Trmt61afl/fl mice (Fig. 1b–d; Supplementary Figs. S1d, S2b–d). There were no observed differences in the percentages and numbers of total CD4+ T cells and RORγt+CD4+ T cells (Fig. 1e–j; Supplementary Fig. S2e–j). Furthermore, Trmt61aΔRorc mice exhibited significantly decreased numbers of all three subsets of ILC3s — NKp46+ ILC3s, CCR6+ ILC3s, and NKp46–CCR6– double-negative (DN) ILC3s — compared to control Trmt61afl/fl mice (Fig. 1k–m; Supplementary Fig. S2k–m). Additionally, although the percentages remained unchanged, the numbers of IL-22-producing ILC3s were significantly lower in the intestine of Trmt61aΔRorc mice compared to controls (Fig. 1n–p; Supplementary Fig. S2n–p). These findings underscore the critical role of TRMT61A in maintaining the homeostasis of all ILC3 subsets in the intestine.
a Real-time RT-PCR analysis of Trmt61a mRNA levels in ILC3s isolated from the colonic lamina propria of Trmt61afl/fl (n = 4) and Trmt61aΔRorc mice (n = 5). b–d Flow cytometry analysis of ILC3s in colonic lamina propria lymphocytes (LPLs): representative plots (b); population frequency (c); cell counts (d). n = 5 mice per group. e–j Analysis of CD4+ and RORγt+ CD4+ T cells in colonic LPLs: e, h Representative flow cytometry plots. f, i Population frequencies. g, j Cell counts. n = 4 mice per group. k–m Counts of ILC3 subsets in colonic LPLs: k CCR6+ ILC3s. l Nkp46+ ILC3s. m DN (double-negative) ILC3s. n = 6 mice per group. n–p IL-22+ ILC3s in colonic LPLs: n Representative flow cytometry plots. o Population frequency. p Cell counts. n = 5 mice per group. Data are pooled from two independent experiments, presented as means ± SEM, *p < 0.05, **p < 0.01, ***p < 0.001, and ****p < 0.0001.
Additionally, to determine whether the decreased numbers of ILC3s in Trmt61aΔRorc mice result from developmental defects, we generated bone marrow chimeras by reconstituting lethally irradiated CD45.1+ wild-type recipients with a 1:1 mixture of CD45.2+ Trmt61aΔRorc (or Trmt61afl/fl control) and CD45.1+ wild-type bone marrow (Supplementary Fig. S3a). Eight weeks post-transplantation, the CD45.2/CD45.1 ratios within intestinal ILC3s were comparable between the two groups (Supplementary Fig. S3b, c). These results suggest that the reduction of ILC3s in Trmt61aΔRorc mice is not due to developmental defects.
Impact of TRMT61A on proliferation and apoptosis in intestinal ILC3s
To explore the underlying molecular mechanism by which TRMT61A regulates ILC3 homeostasis, we isolated ILC3s from the intestines of Trmt61aΔRorc and Trmt61afl/fl mice and performed transcriptome sequencing. The RNA-seq analysis revealed 151 genes significantly downregulated and 195 genes significantly upregulated in Trmt61a-deficient ILC3s compared to control (Fig. 2a). Notably, genes associated with the cell cycle and proliferation, such as Ccna225, Cdca326, and E2f127 were significantly downregulated in Trmt61a-deficient ILC3s (Fig. 2a). Additionally, the expression of the pro-apoptotic gene Casp628 was significantly increased, while the anti-apoptotic gene Birc529 was markedly reduced in these cells (Fig. 2a). Quantitative PCR further validated these findings, confirming reduced expressions of E2f1, Cdca3, and Birc5, and increased expression of Casp6 in Trmt61a-deficient ILC3s compared to controls (Fig. 2b).
ILC3s (Lin–CD45.2intCD90.2hi) were isolated from the colonic LPLs of Trmt61afl/fl and Trmt61aΔRorc mice. The mRNA from triplicate samples of purified ILC3s was extracted and subjected to genome-wide transcriptome analysis (RNA-seq). a Differential gene expression between Trmt61afl/fl and Trmt61aΔRorc ILC3s is displayed. Red points indicate genes with significant differences in expression. b Real-time RT-PCR analysis of specific gene expression in Trmt61afl/fl and Trmt61aΔRorc ILC3s. n = 5 mice per group. c Gene-set enrichment analysis (GSEA) illustrated that genes associated with the G2- M checkpoint pathway were expressed at higher levels in Trmt61afl/fl ILC3s compared to Trmt61aΔRorc ILC3s. d Heat map displaying the expression of transcripts associated with the G2-M checkpoint pathway in Trmt61aΔRorc ILC3s vs Trmt61afl/fl ILC3s. e Gene ontology (GO) enrichment analysis was performed on genes downregulated more than 2-fold in Trmt61aΔRorc ILC3s, highlighting representative enriched biological processes. f, g Heat maps showing lists of downregulated genes related to cell division (f) and the regulation of the apoptotic process (g) in Trmt61aΔRorc ILC3s. h, i Early apoptotic ILC3s in the colon of Trmt61afl/fl and Trmt61aΔRorc mice were assessed by Annexin V/7-AAD staining. Representative flow cytometry plots showing AnnexinV⁺/7-AAD⁻ ILC3s (h). Quantification of early apoptotic ILC3s (i). n = 6 mice per group. Data are pooled from two independent experiments, presented as means ± SEM. *p < 0.05, **p < 0.01.
Gene set enrichment analysis (GSEA) demonstrated that the G2/M checkpoint signature was enriched in control ILC3s but absent in TRMT61A-deficient cells (Fig. 2c, d). Furthermore, gene ontology (GO) analysis of downregulated genes in TRMT61A-deficient ILC3s identified pathways related to cell division and the regulation of the apoptotic process (Fig. 2e–g). Consistent with these transcriptional changes, Annexin V/7-AAD staining of intestinal ILC3s showed a significant increase in early apoptotic cells in Trmt61aΔRorc mice compared with controls (Fig. 2h, i). These findings collectively indicate that TRMT61A plays a critical role in modulating genes transcription related to proliferation and apoptosis, thereby maintaining the homeostasis of ILC3s in the intestine.
The microbiota modulates TRMT61A-mediated ILC3 homeostasis
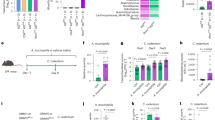
The complex interplay between ILC3s and the microbiota is crucial for maintaining intestinal homeostasis30,31,32. Disruptions to this balance, such as through blocking IL-22 or depleting ILC3s, lead to intestinal epithelial dysfunction and dysbiosis32. To determine whether dysbiosis occurred in Trmt61aΔRorc mice, we conducted 16S rRNA sequencing on fecal samples from both Trmt61aΔRorc and Trmt61afl/fl mice. The fecal microbiota of Trmt61aΔRorc mice showed significant compositional changes compared to controls (Fig. 3a), and displayed greater alpha-diversity, as indicated by the Shannon and Simpson indices33 (Fig. 3b, c). This increase in diversity in Trmt61aΔRorc mice aligns with findings in IL-22-deficient mice34.
a–d Fecal microbiota analysis of Trmt61afl/fl and Trmt61aΔRorc mice using 16S rRNA gene sequencing: principal coordinates analysis (PCA) depicting the overall microbial composition (a), Shannon-Wiener diversity index (Shannon index) of fecal samples (b), Simpson’s Diversity Index (Simpson index) of fecal samples (c), relative abundance of Muribaculaceae in fecal samples (d). n = 6 mice. e Fecal pellets were collected, and total bacterial DNA was extracted. Quantitative PCR was then performed for specific bacteria, with normalization against total bacterial DNA. n = 7 mice per group. f–h Analysis of ILC3 populations in colonic LPLs of 2- week-old and adult Trmt61afl/fl and Trmt61aΔRorc mice: representative flow cytometry plots (f), population frequencies of ILC3s (g), counts of ILC3s (h). n = 4 to 6 mice per group. i–k Effects of antibiotic treatment on ILC3 populations in colonic LPLs of Trmt61afl/fl and Trmt61aΔRorc mice, 2 months post-treatment: representative flow cytometry plots (i), population frequencies of ILC3s (j), counts of ILC3s (k). n = 5 mice per group. l–n Analysis of IL-22+ ILC3s in the colon of Trmt61afl/fl and Trmt61aΔRorc mice with or without antibiotic treatment: representative flow cytometry plots (l), population frequency of IL-22+ ILC3s (m), cell counts of IL-22+ ILC3s (n). n = 6 mice per group. o–q Analysis of ILC3 in the colon of Trmt61afl/fl mice, Trmt61aΔRorc mice, and Trmt61aΔRorc mice with intraperitoneal IL-22 administration: representative flow cytometry plots (o), population frequencies of ILC3s (p), counts of ILC3s (q). n = 4 mice per group. Data are presented as means ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001.
At the family level, there was a notable reduction in Muribaculaceae in the fecal samples of Trmt61aΔRorc mice (Fig. 3d). To validate the 16S rRNA sequencing result and also determine changes of specific Muribaculaceae strains, we performed real-time PCR and checked the relative abundance of different Muribaculaceae strains in the fecal samples from Trmt61afl/fl and Trmt61aΔRorc mice. The abundance of Muribaculum intestinale 8CFAA was significantly reduced in the fecal samples of Trmt61aΔRorc mice, and there was also a trend toward reduced levels of Muribaculum intestinale 3SKV and 6FAAM (Fig. 3e). Muribaculaceae are key utilizers of monosaccharides in the intestine, essential for producing short-chain fatty acids (SCFAs), which have been shown to significantly influence the proliferation and function of intestinal ILC3s35,36,37. This suggests that the microbiota, via SCFAs, may play a role in modulating TRMT61A activity in ILC3s.
We also examined how weaning impacts the microbiota and ILC3 populations. Weaning is a critical period for microbial diversification due to dietary changes and environmental shifts38,39,40. Upon analyzing ILC3 populations in pre-weaning 2-week-old and adult 8-week-old Trmt61aΔRorc mice, in contrast to the deficiency of ILC3s observed in adult Trmt61aΔRorc mice, we found that young Trmt61aΔRorc mice had similar ILC3 percentages and numbers as their young Trmt61afl/fl counterparts (Fig. 3f–h; Supplementary Fig. S4a–c). More interestingly, antibiotic treatment in adult mice normalized the total ILC3 and IL-22+ ILC3 counts in Trmt61aΔRorc mice to levels seen in adult Trmt61afl/fl mice (Fig. 3i–n; Supplementary Fig. S4d–i). underscoring the significant role of microbiota in regulating ILC3 homeostasis mediated by TRMT61A. Moreover, intraperitoneal administration of IL-22 partially restored ILC3 numbers in Trmt61aΔRorc mice (Fig. 3o–q; Supplementary Fig. S4j–l), further supporting a role for IL-22 in mediating ILC3 homeostasis.
To investigate the causal relationship between TRMT61A deficiency, microbiota composition, and ILC3 dysfunction, we performed fecal microbiota transfer (FMT) from Trmt61aΔRorc or Trmt61afl/fl donor mice into antibiotic-treated wild-type recipients, following established protocols41. Six weeks after transplantation, we analyzed intestinal ILC3s (Supplementary Fig. S5a). We found that recipients of Trmt61aΔRorc-derived microbiota (termed FMT-Trmt61aΔRorc) exhibited a significant reduction in both the percentage and total number of intestinal ILC3s compared with recipients of Trmt61afl/fl-derived microbiota (termed FMT-Trmt61afl/fl), along with decreased numbers of IL-22+ ILC3s (Supplementary Fig. S5b–m). These findings suggest that microbial dysbiosis in Trmt61aΔRorc mice alters the intestinal microenvironment and further exacerbates the reduction of ILC3s.
TRMT61A is essential for ILC3-mediated protective immunity
ILC3s play critical roles in maintaining intestinal epithelial homeostasis and defending against bacterial infections42,43. IL-22, secreted by ILC3s, is crucial for stimulating the production of antimicrobial peptides from intestinal epithelial cells, thereby maintaining barrier integrity40,43. To investigate the impact of ILC3-intrinsic TRMT61A on tissue repair during intestinal inflammation, we administered dextran sulfate sodium (DSS) to Trmt61aΔRorc and Trmt61afl/fl mice. Our findings revealed significantly decreased percentages and numbers of total ILC3s and IL-22+ ILC3s in the colons of Trmt61aΔRorc mice compared to controls (Fig. 4a–f). The numbers of IL-17A+ ILC3s were also significantly decreased in the colons of Trmt61aΔRorc mice (Fig. 4g–i). Noteworthily, the counts of CD4+ T cells remained similar between the two groups (Fig. 4j–l), indicative of non-involvement of CD4 counterpart. In accordance with the defects of total ILC3s and IL-22+ ILC3s, Trmt61aΔRorc mice exhibited increased weight loss, greater colonic shortening, and more severe disease pathology scores post-DSS treatment than control mice (Fig. 4m–q). Correspondingly, these mice exhibited reduced expression of IL-22 downstream genes, Reg3a and Reg3g12,13,14 in epithelial cells from Trmt61aΔRorc mice compared to controls (Fig. 4r). Together, these data underscore the essential role of TRMT61A in ILC3-mediated tissue protection during intestine inflammation.
Trmt61afl/fl and Trmt61aΔRorc mice were administered 3% dextran sulfate sodium (DSS) for 4 days, followed by regular drinking water for 2 days. Analyses were conducted on day 6 post-initiation of treatment. a–i Flow cytometry analysis: Representative flow cytometry plots showing total ILC3s (a), IL22+ ILC3s (d) and IL-17A+ ILC3s (g). Population frequencies of total ILC3s (b), IL-22+ ILC3s (e) and IL-17+ ILC3s (h). Counts of total ILC3s (c), IL-22+ ILC3s (f) and and IL-17A+ ILC3s (i) in colonic LPLs of treated mice. n = 7 mice per group. j–l CD4+ T Cells analysis in colonic LPLs: representative flow cytometry plots (j), population frequency (k), cell counts (l). n = 6 mice per group. m–q Morphological and histological assessments: body weight changes (m), representative photograph of the large intestine (n), measurements of colon length (o), representative images of colon sections (p), Scale bars represent 250 μm. Histology scores assessing tissue damage (q). n = 7 mice per group. r Gene Expression Analysis: real-time RT-PCR was used to analyze the expression of barrier function-related genes in intestinal epithelial cells. n = 4 mice per group. Data are pooled from two independent experiments, presented as means ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Given that antibiotic treatment partially rescued ILC3 defects in Trmt61aΔRorc mice under steady-state conditions (Fig. 3i–n) and in the DSS colitis model (Supplementary Fig. S6a–f), we next assessed whether modulation of the microbiota could mitigate disease severity. Trmt61aΔRorc mice were pretreated with a broad-spectrum antibiotic cocktail prior to DSS administration. Consitent with the changes observed in ILC3s, antibiotic treatment also partially ameliorated colitis in Trmt61aΔRorc mice, as reflected by reduced weight loss, improved colon length, and lower histological scores (Supplementary Fig. S6g–j). These results indicate that modulation of the intestinal microbiota can partially rescue both ILC3 defects and disease severity.
Further, to assess the function of ILC3-specific TRMT61A during bacterial infection, we infected mice with Citrobacter rodentium. Post-infection, TRMT61A expression in ILC3s significantly increased, suggesting an active role in defending against C. rodentium (Fig. 5a). Consistently, Trmt61aΔRorc mice displayed markedly reduced percentages and numbers of total ILC3s and IL-22-producing ILC3s in the colon five days post-infection, compared to controls (Fig. 5b–g), along with a decreasing trend in the numbers of IL-17A-producing ILC3s (Fig. 5h–j). These mice also showed lower percentages of Ki-67+ ILC3s, indicating compromised cellular proliferation during infection44 (Fig. 5k, l). The proportions of CD4+ T cells were unaffected during the infection (Fig. 5m, n). With diminished IL-22-producing ILC3 populations, Trmt61aΔ Rorc mice experienced more severe weight loss, colonic shortening, and heightened disease pathology, as well as increased fecal bacterial loads compared to control mice (Fig. 5o–s). These findings collectively affirm that TRMT61A is crucial for effective ILC3-mediated host defense against bacterial challenges.
a Wild-type mice were orally inoculated with 1 × 109 Citrobacter rodentium or PBS as a control. Three days post-infection, intestinal LPLs were isolated. ILC3s (Lin–CD45.2intCD90.2hi) were FACS sorted, and mRNA was extracted. Real-time RT-PCR was conducted to analyze the expression of Trmt61a in ILC3s from infected and control mice. n = 5–7 mice per group. b–n Trmt61afl/fl and Trmt61aΔRorc mice were infected with 1 × 109 Citrobacter rodentium. Flow cytometry analysis 5 days post-infection: b, e, h, k, m Representative flow cytometry plots showing various immune cell populations. Population frequencies of total ILC3s (c), IL-22+ ILC3s (f) and IL-17+ ILC3s (i). Counts of total ILC3s (d), IL-22+ ILC3s (g) and IL-17+ ILC3s (j). l Population frequency of Ki67+ ILC3s, indicating proliferation. n Population frequency of total CD4+ T cells. Data from day 5 post-infection, n = 7 mice per group. o–r Morphological and Histological Assessments: changes in body weight (o), representative image of the large intestine (p), colon length measurements (q), histology scores assessing tissue damage (r). Data from day 5 post-infection, n = 8 mice per group. s Bacterial Load Analysis: quantification of bacterial load in feces on days 3 and 5 post-infection. Data are pooled from two independent experiments, presented as means ± SEM. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Discussion
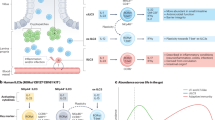
Mucosal barriers are continuously challenged by various stimuli, and the maintenance of mucosal barrier homeostasis relies crucially on tissue-resident immune cells such as ILC3s45,46. Found predominantly in lymphoid tissues and enriched in the intestine, ILC3s are pivotal in maintaining the commensal microbial community and defending against enteric bacterial infections30,43,47,48,49. Despite extensive research into how ILC3s regulate infections and facilitate tissue repair30,43,49, our understanding of their regulation at the epigenetic level, especially concerning RNA modifications, remains incomplete. In this study, we present a model where TRMT61A enhances the proliferation and survival of intestinal ILC3s, thereby protecting mice against experimental intestinal damage and enteric bacterial infections (Fig. 6).
Proposed model illustrating how tRNA-m1A58 modification regulates homeostasis and function of ILC3 in the gut.
Utilizing RNA-seq and qPCR analyses, complemented by GSEA and GO analyses, we identified significant deficits in cell proliferation and the cell cycle in Trmt61a-deficient ILC3s. Specifically, E2F1, a crucial regulator of the cell cycle transition from G1 to S phase27, was markedly downregulated, potentially leading to diminished ILC3 proliferation and numbers. Given that TRMT61A influences m1A modifications on tRNA, which are critical for protein translation initiation and elongation, the transcriptional downregulation of E2F1 suggests an indirect effect of TRMT61A activity. This highlights the necessity for further studies, such as protein mass spectrometry, to pinpoint TRMT61A’s direct targets in ILC3s. The challenges of acquiring sufficient protein from small numbers of ILC3 cells for mass spectrometry underscore the need for more advanced techniques.
Recent studies have elucidated the profound impact of bacterial colonization and its metabolites on various epigenetic modifications, including those affecting DNA methylation and histone configuration50,51,52,53,54. An investigation using Chromatin Immunoprecipitation followed by In Vitro Transcription on ILCs from antibiotic- treated mice revealed significant shifts in H3K4me2 regions55, indicating epigenetic alterations induced by microbiota depletion. Our findings of increased Trmt61a expression in ILC3s following C. rodentium colonization suggest potential alterations in tRNA m1A58 modification, which could facilitate rapid protein synthesis crucial for ILC3 activation and proliferation during infection.
Moreover, while microbiota is not essential for the initial development of ILC3s, their absence or alteration — as observed in germ-free and antibiotic-treated mice — significantly reduces NKp46+ ILC3 and IL-22-producing ILC3 numbers56. The microbial metabolites, particularly SCFAs, are known to enhance ILC3 proliferation through receptors like FFAR236. Together with the Annexin-V/7-AAD staining data, our results suggest that TRMT61A plays a intrinsic role role in maintaining ILC3 numbers. The absence of TRMT61A in ILC3s leads to reduced IL-22+ ILC3 numbers, resulting in microbial dysbiosis in Trmt61aΔRorc mice. This dysbiosis alters the intestinal microenvironment and further exacerbates the reduction of ILC3s.
This study establishes that TRMT61A expression in ILC3s is crucial for defending against intestinal infection and inflammation. The decrease in the number of IL-22-producing ILC3s in Trmt61aΔRorc mice aligns with the observations in patients with inflammatory bowel disease (IBD)57. The observed decrease in IL-22-producing ILC3s in Trmt61aΔRorc mice correlates with reduced TRMT61A expression levels, suggesting that similar mechanisms may operate in IBD, where inflammation could suppress TRMT61A expression, thereby diminishing ILC3 functionality. Future investigations into TRMT61A’s role in IBD could provide new therapeutic avenues.
In summary, our findings significantly advance the understanding of the biological roles of m1A modifications in ILC3s and underscore TRMT61A as a promising target for therapeutic intervention in IBD and related conditions.
Materials and methods
Mice
Trmt61afl/fl mice were constructed using the CRISPR-Cas9-based genome-editing system by inserting two loxP sites into the loci flanking the first exon, as previously described. Rorccre mice were kindle provided by Dr. Lei Shen, Shanghai Institute of Immunology of Shanghai Jiao Tong University, Shanghai. We crossed Trmt61afl/fl mice with Rorccre mice to obtain Trmt61afl/fl Rorccre offspring in which the expression of TRMT61A is specifically absent in ILC3s.
Mice used for in vivo studies were littermate controlled. Both male and female mice were used unless otherwise noted. All mice used in this study are on C57BL/6 background, and were maintained in specific pathogen–free facilities and used according to protocols approved by Animal Care and Use Committees of the Shanghai Jiao Tong University School of Medicine.
Isolation of intestinal lamina propria lymphocytes (LPLs) and intestinal epithelial cells
Small intestine or colon were dissected, fat tissues and peyer’s patches were removed. Intestines were cut open longitudinally and washed in PBS. Intestines were then cut into 3-cm-long pieces, washed, and shaken in PBS containing 1 mM DTT for 10 min at RT. Intestines were incubated with shaking in PBS containing 30 mM EDTA and 10 mM HEPES at 37 °C for 10 min for two cycles. Supernatant from the first round of EDTA was saved as intestinal epithelial cells. The tissues were then digested in the RPMI1640 medium (Thermo Fisher Scientific) containing DNase I (150 µg/mL, Sigma) and collagenase VIII (200 U/mL, Sigma) at 37 °C in a 5% CO2 incubator for 1.5 h. The digested tissues were homogenized by vigorous shaking and passed through 100-µm cell strainer. Mononuclear cells were then harvested from the interphase of an 80 and 40% Percoll gradient after a spin at 2500 rpm for 20 min at RT.
Flow cytometry
Anti-mouse CD16/32 antibody (Biolegend) was used to block the nonspecific binding to Fc receptors before all surface stainings. Dead cells were stained with live and dead violet viability kit (Invitrogen), and were gated out in analysis. For nuclear stainings, cells were fixed and permeabilized using a Mouse Regulatory T Cell Staining Kit (Thermo Fisher Scientific). For detection of IL-22 production in ILC3s, cells were stimulated by PMA (50 ng/mL, Sigma) and ionomycin (500 ng/mL, Sigma) for 4 h. Brefeldin A (2 µg/mL, Sigma) was added for the last 2 h before cells were harvested for analysis.
Antibodies used in this study were listed as follows:
LIVE/DEAD™ Fixable Violet Dead Cell Stain Kit, # L34964, Invitrogen, 1:1000 Zombie Aqua™ Fixable Viability Kit, # 423102, Biolegend, 1:500
CD90 BV605, # 140317, Biolegend, 1:200
CD127 FITC, # 135007, Biolegend, 1:200
NKp46 APC-ef780, # 47-3351-82, eBioscience, 1:200
CCR6 PE, # 129804, Biolegend, 1:200
GATA3 PE, # 653804, Biolegend, 1:40
RORψt APC, # 17-6981-82, eBioscience, 1:200
CD4 AF700, # 100430, Biolegend, 1:200
CD45.2 APC-CY7, # 109824, Biolegend, 1:200
CD45 PerCP-CY5.5, # 103132, Biolegend, 1:200
CD3ɛ PE-CY7, #100220, Biolegend, 1:200
CD3ɛ PerCP, # 100326, Biolegend, 1:200
CD3 BV785, # 100355, Biolegend, 1:200
TCRb PE-Cy7, # 109222, Biolegend, 1:200
CD5 PE-CY7, # 100622, Biolegend, 1:200
FcɛRI PE-CY7, # 134318, Biolegend, 1:200
F4/80 PE-CY7, # 123114, Biolegend, 1:200
CD11b PE-CY7, # 101216, Biolegend, 1:200
CD11c PE-CY7, # 117318, Biolegend, 1:200
CD11b FITC, # 101206, Biolegend, 1:200
CD11c FITC, # 117306, Biolegend, 1:200
Ki67 PE-CY7, # 652426, Biolegend, 1:200
IL-22 PE, # 51640,4 Biolegend, 1:150
Bone marrow transplantation
Bone marrow cells were isolated from the femur and tibia of Trmt61aΔRorc, Trmt61afl/fl or CD45.1 wildtype mice. Red blood cells were lysed with Red Blood Cell Lysis Buffer (Solarbio), and the remaining cells were counted. A total of 5 × 106 bone marrow cells in 200 μL PBS were intravenously injected into each lethally irradiated CD45.1 wild-type recipient. Eight weeks post-transplantation, intestinal ILC3s were analyzed by flow cytometry.
Antibiotics treatments
Pregnant mice were treated with a broad-spectrum mixture of antibiotics (ampicillin (1 g/L), vancomycin (0.5 g/L), metronidazole (1 g/L), neomycin (1 g/L), and gentamicin (1 g/L)) in their drinking water. After birth, the pubs were constantly treated with the antibiotics cocktail as describe above until 2 months.
IL-22 in vivo treatment
For IL-22 treatment, 25 μg recombinant protein (novoprotein) was injected intraperitoneally in a volume of 200 μL PBS for each Trmt61aΔRorc mice.
Fecal microbiota transfer
Fresh fecal pellets were collected from Trmt61aΔRorc or Trmt61afl/fl donor mice and suspended in sterile PBS. Recipient wild-type mice were pretreated with a broad-spectrum antibiotic cocktail to deplete endogenous microbiota. 200 μL of fecal suspension was administered to each recipient by oral gavage. Recipients were maintained for 6 weeks post-transplantation before analysis of intestinal ILC3 populations by flow cytometry.
Citrobacter rodentium infection model
Citrobacter rodentium (DBS100 strain) was cultured overnight and bacterial concentration was calculated by measuring optical density at a wavelength of 600 nm (OD600) with spectrometer. Trmt61aΔRorc and Trmt61afl/fl mice or WT mice were orally infected with 1 × 109 CFU of C. rodentium. Mice were weighed daily. At day 5 after infection, the colon was collected from the infected mice. Colon length was measured. Colon was fixed with 4% paraformaldehyde for histology or used for the isolation of colonic immune cells as describe above. For assessment of bacterial burden, feces were collected at day 3 and day 5 after infection, weighed and homogenized with PBS. The homogenates were plated on MacConkey agar plate and counted after overnight incubation at 37 °C under aerobic conditions.
DSS-induced colitis model
Trmt61aΔRorc and Trmt61afl/fl mice were treated with 3% DSS (MP Biomedicals) in the drinking water for 4 days and followed by regular drinking water for 2 days. Body weight was measured daily and mice were euthanized at day 6. Colon length was measured. Colon was fixed with 4% paraformaldehyde for histology or used for the isolation of colonic immune cells or colonic epithelial cells as describe above.
Histological analysis
Tissues from proximal colon were dissected and fixed with 4% paraformaldehyde. Tissues were then embedded in paraffin, sectioned at 5 µm, and stained with H&E. Sections were then blindly analyzed using the light microscope (Olympus), and scored according to a previously described scoring system58. The four parameters used include (i) the degree of inflammatory infiltration in the LP, range 1–3; (ii) Goblet cell loss as a marker of mucin depletion, range 0–2; (iii) mucosal erosion to frank ulcerations, range 0–2; and (iv) submucosal spread to transmural involvement, range 0–2.The severity of inflammation in sections of the colon was based on the sum of the scores in each parameter (maximum score = 9).
Microbiota sequencing
Fresh mouse fecal samples were collected and microbial DNA was isolated from stools of indicated mice by using QiAamp DNA Stool Mini Kit (QIAGEN) according to manufacturer’s instructions. DNA concentration was checked with Nanodrop (ThermoFisher Scientific). The V4 regions of the bacteria 16S rRNA gene were amplified by PCR using primers 515F-806R. 16S rRNA sequencing was performed by Oebiotech (Shanghai, China).
RNA-seq analysis
About 1 × 104 ILC3s (Lin−CD45.2intThy1.2hi) were sorted from the colon of Trmt61aΔRorc and Trmt61afl/fl mice separately and were lysed in Trizol (Invitrogen). The total RNA was extracted. Biological duplicates were generated for each group. The total RNA sample was digested by DNaseІ (NEB), and purified by oligo-dT beads (Dynabeads mRNA purification kit, Invitrogen), then poly(A) containing mRNA were fragmented into 130 bp with First-strand buffer. First-strand cDNA was generated by N6 primer, First Strand Master Mix and Super Script II reverse transcription (Invitrogen) (reaction condition: 25 °C for 10 min; 42 °C for 40 min; 70 °C for 15 min). Then add Second Strand Master Mix to synthesize the second-strand cDNA (16 °C for 1 h). Purified the cDNA with Ampure XP Beads (AGENCOURT), then combine with End Repair Mix, incubate at 20 °C for 30 min. Purified and add A-Tailing Mix, incubate at 37 °C for 30 min. Then combine the Adenylate 3′ ends DNA, Adapter and Ligation Mix, incubate the ligate reaction at 20 °C for 20 min. Several rounds of PCR amplification with PCR Primer Cocktail and PCR Master Mix were performed to enrich the cDNA fragments. Then the PCR products were purified with Ampure XP Beads (AGENCOURT). The Qualified libraries will amplify on cBot to generate the cluster on the flowcell (TruSeq PE Cluster Kit V3–cBot–HS, Illumina). The amplified flowcell will be sequenced pair end on the HiSeq 2000 System (TruSeq SBS KIT-HS V3, Illumina), read length 50. Reads were mapped to Mouse Genome Assembly GRCm38.p5 by STAR v2.5. Gene and isoform expression quantification was called by RSEM v1.2 with default parameters on GENCODE mouse M16 gene annotation file. Differential expression analysis was performed by Bioconductor package edgeR v3.18.1. Significantly changed genes were chosen according to two criteria: (1) significance level p < 0.05; (2) fold change > 2, and were used for gene ontology enrichment analysis using the website of Gene Ontology Consortium (http://www.geneontology.org). Gene set enrichment analysis was performed using GSEA software (Broad Institute). Normalized heatmap was based on the standard score (Z score) and generated with oebiotech cloud platform (https://cloud.oebiotech.com/task/). The standard score of a raw score x is Z = x−μσ, where μ is the mean of the FPKM value of each sample and σ is the standard deviation of the FPKM value of each sample.
Quantitative real-time RT-PCR
RNA was isolated with Trizol reagent (Invitrogen). cDNA was synthesized using the GoScript™ Reverse Transcription kit (Promega). Real-time PCR was performed using SYBR Green (Bio-rad). Reactions were run with the Bio-Rad CFX384. The results were displayed as relative expression values normalized to β-actin.
Target | Primer Sequence (5′-3′) |
|---|---|
Trmt61a Forward | ATGAGTTTCGTGGCATACGAG |
Trmt61a Reverse | CCTCTGCTGCAAATCACCTT |
Reg3g Forward | CAAGGTGAAGTTGCCAAGAA |
Reg3g Reverse | CCTCTGTTGGGTTCATAGCC |
Reg3b Forward | CCACTCTGGGTGCAGAAC |
Reg3b Reverse | AATTCGGGATGTTTGCTGTC |
Reg3a Forward | TCACCTGGTCCTCAACAGTATT |
Reg3a Reverse | GGAGCGATAAGCCTTGTAACC |
E2f1 Forward | CTCGACTCCTCGCAGATCG |
E2f1 Reverse | GATCCAGCCTCCGTTTCACC |
E2f1 Forward | CTGAGCGAAGTATTGGAGACAG |
E2f1 Reverse | CTGCGGATTGTTTGGCTTCC |
Birc5 Forward | GAGGCTGGCTTCATCCACTG |
Birc5 Reverse | CTTTTTGCTTGTTGTTGGTCTCC |
Casp6 Forward | GGAAGTGTTCGATCCAGCCG |
Casp6 Reverse | GGAGGGTCAGGTGCCAAAAG |
Bacterial DNA extraction and real-time PCR
Fecal pellets were collected and total bacterial DNA was extracted using the Stool DNA Kit (Omega Biotek). Quantitative PCR for detection of specific Muribaculaceae strains was performed with SYBR Green (Bio-Rad) and normalized to total bacterial DNA. Reactions were run with the Bio-Rad CFX384. Primers used in this study were shown below59.
Target | Primer Sequence (5′-3′) |
|---|---|
All bacteria | Forward CGGTGAATACGTTCCCGG |
All bacteria | Reverse TACGGCTACCTTGTTACGACTT |
Muribaculum intestinale | 3SKV Forward AACAACAACCAGCTGACAAT |
Muribaculum intestinale | 3SKV Reverse AGTAGTTCCACTGGCAGG |
Muribaculum intestinale | NM65 Forward GTAACAACAACCAGTTGACAT TG |
Muribaculum intestinale | NM65 Reverse TAATAGTTCCACTGGCATGCA |
Muribaculum intestinale | NM06 Forward CAACAACAACCAGCTGACTAT G |
Muribaculum intestinale | NM06 Reverse AATAGTTCCACTGACATGCC |
Muribaculum intestinale | 7GAM Forward AACAACAACCAGTTGACCAT G |
Muribaculum intestinale | 7GAM Reverse TAGTAGTTCCACTGGCAGG |
Muribaculum intestinale | 8CFAA Forward AACAACAACCAGCTGACAAT |
Muribaculum intestinale | 8CFAA Reverse CAACAACAACCAGCTGACA AT |
Muribaculum intestinale | 8FP Forward CAACAACAACCAGCTGACAAT |
Muribaculum intestinale | 8FP Reverse AGTAGTTCCACTGGCAAGC |
Muribaculum intestinale | 6FAAM Forward CAACAACAACCAGCTGACA AT |
Muribaculum intestinale | 6FAAM Reverse TAGTTCCACTGGCATGCC |
Muribaculum intestinale | 16COLB Forward CAACAACAACCAGCTGACA AT |
Muribaculum intestinale | 16COLB Reverse CAATAGTTCCACTGGCATGC |
Statistical analysis
Unpaired Student’s t-tests and two-way analysis of variance (ANOVA) were used to compare pairs of groups, and all data are presented as the means ± SEM. p values of < 0.05 were considered statistically significant.
Data availability
All data needed to evaluate the conclusions in this article are present in the paper or the supplementary materials (or both). The bulk RNA-seq data generated in this study are available at the Gene Expression Omnibus (GSE267755). The Trm61a flox mouse strain can be provided by H.B.L. pending scientific review and a completed materials transfer agreement. Requests for those mouse lines should be submitted to H.B.L.
References
Artis, D. & Spits, H. The biology of innate lymphoid cells. Nature 517, 293–301 (2015).
Diefenbach, A., Colonna, M. & Koyasu, S. Development, differentiation, and diversity of innate lymphoid cells. Immunity 41, 354–365 (2014).
Eberl, G., Colonna, M., Di Santo, J. P. & McKenzie, A. N. J. Innate lymphoid cells: a new paradigm in immunology. Science 348, aaa6566 (2015).
Vivier, E. et al. Innate lymphoid cells: 10 years on. Cell 174, 1054–1066 (2018).
Masopust, D. & Schenkel, J. M. The integration of T cell migration, differentiation and function. Nat. Rev. Immunol. 13, 309–320 (2013).
Gasteiger, G., Fan, X., Dikiy, S., Lee, S. Y. & Rudensky, A. Y. Tissue residency of innate lymphoid cells in lymphoid and nonlymphoid organs. Science 350, 981–985 (2015).
Forkel, M. & Mjösberg, J. Dysregulation of group 3 innate lymphoid cells in the pathogenesis of inflammatory bowel disease. Curr. Allergy Asthma Rep. 16, 73 (2016).
Zhou, W. & Sonnenberg, G. F. Activation and suppression of group 3 innate lymphoid cells in the gut. Trends Immunol. 41, 721–733 (2020).
Domingues, R. G. & Hepworth, M. R. Immunoregulatory sensory circuits in group 3 innate lymphoid cell (ILC3) function and tissue homeostasis. Front. Immunol. 11, 116 (2020).
Li, J. et al. Activation of DR3 signaling causes loss of ILC3s and exacerbates intestinal inflammation. Nat. Commun. 10, 3371 (2019).
Sonnenberg, G. F. & Artis, D. Innate lymphoid cells in the initiation, regulation and resolution of inflammation. Nat. Med. 21, 698–708 (2015).
Guo, X. et al. Innate lymphoid cells control early colonization resistance against intestinal pathogens through ID2-dependent regulation of the microbiota. Immunity 42, 731–743 (2015).
Jarade, A., Di Santo, J. P. & Serafini, N. Group 3 innate lymphoid cells mediate host defense against attaching and effacing pathogens. Curr. Opin. Microbiol. 63, 83–91 (2021).
Serafini, N. et al. Trained ILC3 responses promote intestinal defense. Science 375, 859–863 (2022).
Peng, V. et al. Whole-genome profiling of DNA methylation and hydroxymethylation identifies distinct regulatory programs among innate lymphocytes. Nat. Immunol. 23, 619–631 (2022).
Chang, J. et al. Setd2 determines distinct properties of intestinal ILC3 subsets to regulate intestinal immunity. Cell Rep. 38, 110530 (2022).
Wang, X. & He, C. Dynamic RNA modifications in posttranscriptional regulation. Mol. Cell 56, 5–12 (2014).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Liu, F. et al. ALKBH1-mediated tRNA demethylation regulates translation. Cell 167, 816–828.e16 (2016).
Zhang, C. & Jia, G. Reversible RNA modification N 1-methyladenosine (m1A) in mRNA and tRNA. Genom. Proteom. Bioinforma. 16, 155–161 (2018).
Wang, Y. et al. N1-methyladenosine methylation in tRNA drives liver tumourigenesis by regulating cholesterol metabolism. Nat. Commun. 12, 6314 (2021).
Wang, Y., Huang, Q., Deng, T., Li, B. H. & Ren, X. Q. Clinical significance of TRMT6 in hepatocellular carcinomaÿ: a bioinformatics-based study. Med. Sci. Monit. 25, 3894–3901 (2019).
Zhao, Y. et al. m1A regulated genes modulate PI3K/AKT/mTOR and ErbB pathways in gastrointestinal cancer. Transl. Oncol. 12, 1323–1333 (2019).
Xu, H. et al. Transcriptional atlas of intestinal immune cells reveals that neuropeptide α-CGRP modulates group 2 innate lymphoid cell responses. Immunity 51, 696–708.e9 (2019).
Loukil, A. Cyclin A2: At the crossroads of cell cycle and cell invasion. WJBC 6, 346 (2015).
Zhang, W. et al. CDCA3 promotes cell proliferation by activating the NF-κB/cyclin D1 signaling pathway in colorectal cancer. Biochem. Biophys. Res. Commun. 500, 196–203 (2018).
Wu, L. et al. The E2F1–3 transcription factors are essential for cellular proliferation. Nature 414, 457–462 (2001).
Zheng, M., Karki, R., Vogel, P. & Kanneganti, T. D. Caspase-6 is a key regulator of innate immunity, inflammasome activation, and host defense. Cell 181, 674–687.e13 (2020).
Mita, A. C., Mita, M. M., Nawrocki, S. T. & Giles, F. J. Survivin: key regulator of mitosis and apoptosis and novel target for cancer therapeutics. Clin. Cancer Res. 14, 5000–5005 (2008).
Sawa, S. et al. RORγt+ innate lymphoid cells regulate intestinal homeostasis by integrating negative signals from the symbiotic microbiota. Nat. Immunol. 12, 320–326 (2011).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013).
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012).
Willis, A. D. Rarefaction, alpha diversity, and statistics. Front. Microbiol. 10, 2407 (2019).
Zenewicz, L. A. et al. IL-22 deficiency alters colonic microbiota to be transmissible and colitogenic. J. Immunol. 190, 5306–5312 (2013).
Smith, B. J. et al. Changes in the gut microbiome and fermentation products concurrent with enhanced longevity in acarbose-treated mice. BMC Microbiol. 19, 130 (2019).
Chun, E. et al. Metabolite-sensing receptor Ffar2 regulates colonic group 3 innate lymphoid cells and gut immunity. Immunity 51, 871–884.e6 (2019).
Sepahi, A., Liu, Q., Friesen, L. & Kim, C. H. Dietary fiber metabolites regulate innate lymphoid cell responses. Mucosal Immunol. 14, 317–330 (2021).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat. Med. 23, 314–326 (2017).
Koenig, J. E. et al. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 108, 4578–4585 (2011).
Ganal-Vonarburg, S. C. & Duerr, C. U. The interaction of intestinal microbiota and innate lymphoid cells in health and disease throughout life. Immunology 159, 39–51 (2020).
Zhang, W. et al. Gut-innervating nociceptors regulate the intestinal microbiota to promote tissue protection. Cell 185, 4170–4189.e20 (2022).
Sanos, S. L., Vonarbourg, C., Mortha, A. & Diefenbach, A. Control of epithelial cell function by interleukin-22-producing RORγt+ innate lymphoid cells: IL-22-producing RORγt+ ILCs. Immunology 132, 453–465 (2011).
Satoh-Takayama, N. et al. Microbial flora drives interleukin 22 production in intestinal NKp46+ cells that provide innate mucosal immune defense. Immunity 29, 958–970 (2008).
Scholzen, T. & Gerdes, J. The Ki-67 protein: From the known and the unknown. J. Cell. Phys. 182, 311–322 (2000).
Kurashima, Y., Goto, Y. & Kiyono, H. Mucosal innate immune cells regulate both gut homeostasis and intestinal inflammation. Eur. J. Immunol. 43, 3108–3115 (2013).
Okumura, R. & Takeda, K. Maintenance of gut homeostasis by the mucosal immune system. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 92, 423–435 (2016).
Rankin, L. C. et al. Complementarity and redundancy of IL-22-producing innate lymphoid cells. Nat. Immunol. 17, 179–186 (2016).
Song, C. et al. Unique and redundant functions of NKp46+ ILC3s in models of intestinal inflammation. J. Exp. Med. 212, 1869–1882 (2015).
Sonnenberg, G. F., Monticelli, L. A., Elloso, M. M., Fouser, L. A. & Artis, D. CD4+ lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 34, 122–134 (2011).
Krautkramer, K. A. et al. Diet-microbiota interactions mediate global epigenetic programming in multiple host tissues. Mol. Cell 64, 982–992 (2016).
Aleksandrova, K., Romero-Mosquera, B. & Hernandez, V. Diet, gut microbiome and epigenetics: emerging links with inflammatory bowel diseases and prospects for management and prevention. Nutrients 9, 962 (2017).
Romano, K. A. et al. Metabolic, epigenetic, and transgenerational effects of gut bacterial choline consumption. Cell Host Microbe 22, 279–290.e7 (2017).
Ansari, I. et al. The microbiota programs DNA methylation to control intestinal homeostasis and inflammation. Nat. Microbiol. 5, 610–619 (2020).
Poupeau, A. et al. Genes controlling the activation of natural killer lymphocytes are epigenetically remodeled in intestinal cells from germ-free mice. FASEB J. 33, 2719–2731 (2019).
Gury-BenAri, M. et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell 166, 1231–1246.e13 (2016).
Sanos, S. L. et al. RORγt and commensal microflora are required for the differentiation of mucosal interleukin 22–producing NKp46+ cells. Nat. Immunol. 10, 83–91 (2009).
Goc, J. et al. Dysregulation of ILC3s unleashes progression and immunotherapy resistance in colon cancer. Cell 184, 5015–5030.e16 (2021).
Wang, W., Li, Y. & Guo, X. A mouse model of Citrobacter rodentium oral infection and evaluation of innate and adaptive immune responses. STAR Protoc. 1, 100218 (2020).
Ghezzi, H. et al. PUPpy: a primer design pipeline for substrain-level microbial detection and absolute quantification. mSphere 9, e0036024 (2024).
Acknowledgements
We thank M. Yang and K. Mao for technical and administrative assistance, and all members of the Li laboratories for discussions and suggestions; the imaging, flow cytometry core facilities at Shanghai Institute of Immunology and the Core Facility of College of Basic Medical Sciences, Shanghai Jiao Tong University School of Medicine for their support. This work was supported by the the Ministry of Science and Technology of China (2025YFC3408600 to C.C., 2021YFA1100800 to H.B.L.); the National Natural Science Foundation of China (82341017, 82325024, 82350112, 82030042, 32070917 and 32070917 to H.B.L., 32370953 to C.C., 32000643 to Jingyu Li); the Shanghai Municipal Health Commission (2022XD047 and 2022JC001 to H.B.L.); the Research Fund of Vanke School of Public Health (to C.C.); and the Phase III Collaborative Agreement for Frontier Medicine and Biotechnology Research (20252000931 to C.C.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
J.L. designed the research, performed experiments, analyzed the data, and drafted the manuscript; Z.T. and Y.C. performed experiment and interpreted the data; X.C. performed experiments; H.L. and C.C. contributed to discussions and writings. H.B.L. conceived the project, designed the study, supervised the research, acquired the funding, and wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Tang, Z., Chen, Y. et al. tRNA m1A modification is essential for gut homeostasis and function of group 3 innate lymphoid cells. Cell Discov 12, 1 (2026). https://doi.org/10.1038/s41421-025-00850-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41421-025-00850-9