Abstract
Ribonucleoprotein (RNP) granules—membraneless organelles formed through the condensation of RNA and proteins—play pivotal roles in diverse biological processes and diseases, opening new directions in molecular biology. Identifying the RNA composition of these granules is crucial for understanding their formation and functions. However, conventional approaches based on the simple immunoprecipitation of specific granule markers struggle to capture the precise nature of RNP granules. This review summarizes recent advances in granule transcriptome analysis, including the use of purification strategies, such as centrifugation and fluorescence-activated particle sorting, as well as proximity labeling techniques, which may help to increase the understanding of RNP granule biology.
Similar content being viewed by others
RNA: a central player in phase separation
Phase separation has emerged as a fundamental mechanism for cellular compartmentalization, allowing precise and efficient biochemical reactions in the crowded intracellular environment. Many phase-separated biomolecular condensates are rich in RNAs and RNA-binding proteins, which are collectively referred to as ribonucleoprotein (RNP) granules [1]. A wide variety of RNP granules have been identified across diverse cell types and organisms and are suggested to be important in dynamic physiological processes such as stress responses [2], development [3], and aging [4]. Dysfunction of RNP granules is implicated in diseases, including cancer, viral infections, and neurodegenerative disorders [5, 6], highlighting the importance of understanding these condensates from both biological and medical perspectives.
Historically, research on RNP granule formation has focused mainly on proteins containing intrinsically disordered regions. Recent findings, however, highlight RNA itself as a central player in granule assembly [1, 7]. Because RNA is a long polymer, it presents numerous binding sites for proteins or other RNAs, promoting condensate formation through multivalent interactions. The assembly of stress granules and nuclear granules, including nucleoli, is RNA dependent; they disappear or are rearranged by transcription inhibition or RNase activation [8,9,10]. Specific long noncoding RNAs termed architectural RNAs (arcRNAs), such as NEAT1 and HSATIII, are required for the assembly of paraspeckles and nuclear stress bodies, respectively [11, 12]. Moreover, purified cellular RNA mixtures, homopolymers (e.g., polyuridine), and pathological repeat RNAs can self-assemble in vitro [13, 14]. RNA-binding proteins, such as eIF4A and YB-1, also act as RNA chaperones, regulating RNA‒RNA interactions and thereby influencing stress granule formation [15, 16]. Intriguingly, depending on the secondary structure of RNAs, Whi3 droplets in the filamentous fungus Ashbya gossypii have distinct viscosities and dynamic properties [17]. These findings collectively illustrate that RNA is a key determinant of both the formation and characteristics of RNP granules.
RNP granules play important roles in RNA metabolism and gene regulation, and their link to translation has been extensively examined. Cytoplasmic RNP granules, such as stress granules and processing bodies (P-bodies), often contain untranslated, ribosome-free mRNAs [14, 18, 19]. Optogenetic condensation and in vitro reconstitution experiments have indicated that mRNA condensation can lead to translational repression [20, 21]. Conversely, phase separation can provide an active translation site [22,23,24] and even enhance the translation of stored mRNAs [25]. RNP granules also contribute to RNA transport, as neuronal RNA granules migrate along microtubules in the axon [26]. Connections to transcription have also been proposed, given that various nuclear RNP granules (e.g., nucleoli, histone locus bodies, and Cajal bodies) form at transcriptionally active gene loci [27]. Nonetheless, conflicting results are occasionally reported, and the question of whether RNP granules directly govern these processes is still under debate [28].
Understanding the RNA composition of RNP granules is, therefore, crucial for elucidating their formation mechanisms and functions. Microscopy-based methods, such as RNA fluorescence in situ hybridization (FISH) or fluorescent tagging (e.g., MS2 or PP7 stem‒loop systems), have identified granule-associated RNAs [29]. However, given that thousands of mRNA species are sequestered to varying extents in stress granules and P-bodies [30, 31], comprehensive and quantitative approaches are needed to capture the full blueprint of RNP granules.
Granule purification by differential centrifugation
Transcriptome analyses of RNP granules often rely on immunoprecipitation and deep sequencing, e.g., individual-nucleotide resolution crosslinking immunoprecipitation (iCLIP)-seq [32], which identifies RNAs bound to marker proteins. However, granular components are in equilibrium between the granule-associated and dissociated states. Microscopic observations have demonstrated that only approximately 18% of G3BP1 in mammalian stress granules [33] and approximately 30% of Dcp2 and Edc3 in yeast P-bodies [34] are sequestered in granules. Therefore, biochemical purification or labeling of RNP granules is necessary to capture the granule transcriptome more precisely.
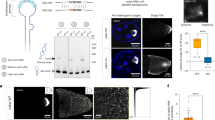
The most straightforward and widely accessible method for granule isolation is differential centrifugation (Fig. 1A) [30, 35]. Because RNP granules are relatively dense, they can be pelleted by simple benchtop centrifugation [19, 35]. Typically, cell lysates are centrifuged at low speed (~1,000 × g) to remove debris, nuclei, and other large organelles, and the supernatant is further centrifuged at 10,000–18,000 × g to isolate granules. However, the pellet fraction also includes various RNP granules and similar sedimenting membranous organelles, such as the endoplasmic reticulum (ER) and mitochondria [14, 36]. Hence, specific RNP granules should be purified by immunoprecipitation of their marker proteins.
Granule purification by centrifugation. A Purification of stress granules using differential centrifugation followed by immunoprecipitation. B Purification of granules via density gradient centrifugation and immunoprecipitation. C Adjusting the lysate density places the granules toward the bottom of the gradient to minimize cross-contamination between organelles
The differential centrifugation-based approach was originally used to analyze the proteome of stress granules in yeast and mammalian cells [35]. This approach was later applied to transcriptome analysis of stress granules, and comparison with ribosome profiling data has revealed that long and poorly translated RNAs accumulate [30]. Biases in transcript length and translation efficiency have also been observed when purifying P-bodies under stress [37, 38] and even when broadly isolating granule-enriched pellet fractions without immunoprecipitation [37, 39]. These observations suggest that RNA length and translational status may be common features of various cytoplasmic RNP granules.
Centrifugation-based purification has been widely used in a broader range of granules and organisms. For example, the application of bacterial RNP (BR) bodies has resulted in the accumulation of longer, translationally repressed mRNAs, mirroring eukaryotic granules [40, 41]. Differential centrifugation also facilitates investigations into the connections between diseases and RNP granules, e.g., stress granules in neuroblastoma cells carrying amyotrophic lateral sclerosis (ALS) mutations [42, 43], P-bodies and stress granules during T lymphocyte activation [44], and paracrine granules, which resemble stress granules yet represent distinct foci in cells neighboring virus-infected cells [45]. Moreover, granule precipitation works in vitro. The components of aggregates formed by the self-assembly of purified yeast RNAs have been successfully analyzed, revealing that the properties of RNAs assembled without proteins correlate with the features of mRNAs in the cellular stress granule [14]. Stress granules or nucleoli reconstituted by the addition of recombinant proteins to granule-free lysates (the supernatant after granule precipitation) resemble their cellular counterparts in both the proteome and transcriptome [46].
A limitation of the differential centrifugation-based method is that only stable granules or regions resistant to lysis, centrifugation, and immunoprecipitation are recovered. Stress granules isolated by centrifugation are smaller than those in cells, suggesting that the peripheral shell is diffused and that the stable core is purified [35]. Additionally, many cells are needed, as a previous study has reported culturing cells in a 500 cm2 dish for this method [47].
Granule purification by density gradient or size-exclusion chromatography
Another approach to isolating granules utilizes density gradient centrifugation, which separates RNP granules from membranous organelles. For example, RNP granules have been separated from the ER in a mouse brain lysate using a 15–30% iodixanol gradient and ultracentrifugation, with subsequent immunoprecipitation allowing the transcriptome analysis of Staufen-containing RNP granules (Fig. 1B) [48].
Modification of this approach further minimizes cross-contamination. The optimization includes adjusting the density of the lysate (dissolved in 30% iodixanol) and laying it at the bottom side of the gradient (Fig. 1C) [49]. In this setup, membranous organelles move to the top fractions, whereas cytoplasmic RNAs and proteins migrate to the bottom. The lighter fraction of the cytosol is considered granule rich because the RNAs exhibit lower ribosome associations and longer transcript lengths, corresponding to the transcriptomes of stress granules and P-bodies [30, 31]. This density gradient-based approach also allows multiple organelles to be purified from the same lysate, allowing RNA redistribution to be tracked in response to stress. Notably, upon the unfolded protein response, numerous mRNAs shift from the membrane fraction to the lighter cytosolic fraction.
Because stress granules vary in size, many smaller granules escape standard centrifugation. To address this issue, a method has been recently developed to avoid granule pelleting [50]. After large organelles are removed via brief centrifugation, ribosomes and actively translated mRNAs are separated by a sucrose density gradient. Subsequently, size-exclusion chromatography using a resin with a nominal exclusion limit of 100 MDa isolates relatively large but low-density granules that do not pellet.
Fluorescence-activated particle sorting (FAPS)
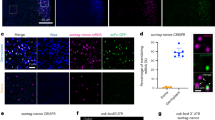
Leveraging the near-spherical morphology of RNP granules, an isolation method using a cell sorter has been developed (Fig. 2) [31]. In this method, which is called FAPS, granules are tagged with a fluorescent protein. After cell lysis, the cells are sorted on the basis of fluorescence intensity using a fluorescence-activated cell sorter. FAPS provides high-purity granules by eliminating granule-free marker proteins and other RNPs.
Fluorescence-activated particle sorting (FAPS). Cells expressing a granule marker protein tagged with a fluorescent protein are lysed and subjected to fluorescence-activated cell sorting to isolate granules on the basis of fluorescence intensity
FAPS was initially applied to P-bodies, revealing an enrichment of translationally repressed mRNAs and regulatory mRNAs related to chromatin remodeling and cell division [31]. Subsequent studies using FAPS have demonstrated that m6A modifications shape the P-body transcriptome [51] and that the RNA composition of P-bodies dynamically changes during epithelial‒mesenchymal transition [52], cell cycle progression [53], and the differentiation of human embryonic stem cells into mesodermal cells [54]. In addition to P-bodies, FAPS has been applied to TIS granules intertwined with rough ER [55], autophagic p62 bodies [56], and DACT1 condensates, promoting bone metastasis [57]. In addition, FAPS has also been used in Arabidopsis, including phytochrome B photobodies [58] and condensates of the ECT1 m6A reader [59].
The drawback of the FAPS method, similar to differential centrifugation, is that it cannot recover unstable granules that disassemble during lysis and sorting. However, stabilization by formaldehyde-mediated crosslinking expands the applications of FAPS to fragile RNP granules. For example, although stress granules are easy to disassemble by cell lysis and are thus not applicable to FAPS, fixation overcomes these issues, which has allowed RNA damage-induced subtypes containing DHX9 to be sorted [60]. The fixation approach also enabled the isolation of C. elegans oocyte P-bodies, which grow dynamically during oogenesis [61]. Moreover, in vitro reconstituted FUS droplets in lysates have been successfully sorted by crosslinking methods [62].
Another limitation is that FAPS requires a relatively large number of cells (two 15-cm dishes [31] and ~1000 C. elegans animals [61] have previously been used for FAPS). Additionally, it is time-consuming to obtain sufficient quantities of granules because the granules are sorted individually. For example, ~6–12 h are needed to sort enough P-bodies for proteomics or transcriptomics [31, 61].
Semiextractable RNA as a granule RNA
RNAs are conventionally isolated with acid guanidinium thiocyanate-phenol‒chloroform (AGPC) reagents (such as TRIzol) and separated into the aqueous phase, excluding proteins in the organic phase (Fig. 3). However, RNAs incorporated into phase-separated structures (e.g., NEAT1) form multivalent interactions with many proteins and, thus, are enriched at the interface of liquid-organic phases in the AGPC method [63]. Moreover, extensive needle shearing or heating increases the solubility of these RNAs in the aqueous phase. This characteristic, called “semiextractability,” is harnessed as a hallmark of RNAs within phase-separated granules [63, 64]. Semiextractable RNAs include nuclear lncRNAs, pre-mRNAs, antisense RNAs, and repeat RNAs, which assemble into nuclear granule-like structures.
Semiextractable RNA as a hallmark of granular RNA. RNAs within phase-separated structures (arcRNAs) remain in the protein-rich phase when extracted using acid guanidinium thiocyanate-phenol‒chloroform (AGPC) reagents (e.g., TRIzol). However, these RNAs can be recovered in the aqueous phase by needle shearing or heat treatment
Proximity labeling via peroxidases
Granule purification relies on multiple biochemical steps—i.e., cell lysis, centrifugation, and/or sorting—that can disrupt granule integrity and alter their native state. To overcome this limitation, researchers have developed in-cell labeling methods that biotinylate granule RNAs directly within living cells. One prominent strategy employs APEX2, an enzyme artificially evolved from soybean ascorbate peroxidase [65], which generates radicals from phenols upon hydrogen peroxide treatment [66]. The conjugation of biotin to phenol allows the addition of this molecular handle to proteins and RNAs. The generated radical is highly reactive but short-lived (<1 ms) [67], causing biotinylation only in close proximity (<20 nm) to the active site of the enzyme [68]. Notably, owing to the diffusion and decay of these radicals, the labeling efficiency should decline progressively with distance (like a “contour map”) rather than remain confined to a fixed radius [66].
Although APEX2 was originally designed to label proteins such as biotin ligases, it also directly biotinylates RNAs, enabling APEX-Seq of the local transcriptome (Fig. 4A) [69, 70]. Because labeling occurs in living cells, an APEX2-based approach can capture unstable granules that are not suitable for biochemical purification. Notably, RNAs close to marker proteins that are free from granules may also be labeled; hence, appropriate controls (e.g., marker mutants that do not form granules) ensure specificity. APEX-Seq has illuminated the local transcriptomes of many membranous and membraneless organelles, including nucleoli and stress granules [69, 70]. Using eIF4A1 as a stress granule marker, this APEX2-based approach has revealed that heat shock-induced stress granules accumulate longer transcripts [70], which is consistent with earlier findings obtained via biochemical purification [30]. Interestingly, when stress granules are induced by hippuristanol (an eIF4A inhibitor), no bias in transcript length is observed, suggesting that stress granule composition may vary in response to stimuli. APEX-Seq has also been applied to DDX3X- and DDX3Y-containing stress granules [71].
Engineered ascorbate peroxidase (APEX2)-based proximity labeling. A APEX-Seq. APEX2 generates biotin‒phenoxyl radicals in the presence of biotin‒phenol and hydrogen peroxide. These radicals are highly reactive and covalently bind to RNA in proximity to APEX2, but their short-lived nature limits the labeling radius. B APEX-Seq with alkyne-phenol. Alkynylated RNAs are conjugated with azide-biotin via a click reaction. C APEX2 is recruited to a target RNA using the MS2 system or inactive Cas13. D Hybridization-proximity (HyPro)-Seq. In fixed cells, digoxigenin (DIG)-labeled probes hybridize to the target RNA, and APEX2 is subsequently recruited via a DIG-binding protein
One limitation of APEX-Seq is its relatively low labeling efficiency for nucleic acids. The phenoxyl radical preferentially reacts with electron-rich guanine [69], but its efficiency is lower than that of amino acids (e.g., tyrosine). To improve RNA labeling, researchers have substituted biotin-phenol with biotin-aniline [72] or employed the metabolic incorporation of electron-rich ribonucleosides, such as 4-thiouridine or 6-thioguanosine, into cellular RNAs [73, 74]. Another drawback is the low permeability of biotin-phenol in some organisms, such as yeast. To address this issue, a smaller and more permeable alkyne-phenol mixture can be employed, followed by a click reaction with azide-biotin after RNA extraction (Fig. 4B) [75]. The alkyne-phenol-based approach has been successfully applied to bacterial RNP (BR) bodies in E. coli [76]. More recently, azide-phenol has been reported to offer high RNA-labeling efficiency and an ideal balance of hydrophilicity and lipophilicity [77].
In some cases, RNA serves as a more reliable marker than protein for defining RNP granules. APEX2 can be recruited to marker RNAs, instead of fusing to marker proteins, via the MS2 stem‒loop system or a catalytically inactive Cas13, an RNA-targeting enzyme that depends on the guide RNA (Fig. 4C) [78]. This RNA-defined labeling enables the capture of proteins close to human telomerase RNA [79]. In fixed cells, multiple digoxigenin (DIG)-labeled probes can be hybridized to the target RNA, after which a DIG-binding protein fused to APEX2 is recruited (Fig. 4D) [80]. This technology, termed hybridization-proximity (HyPro)-Seq, reveals the transcriptomes of RNP granules defined by 45S pre-rRNA (for the nucleolus), NEAT1 (for the paraspeckle), and PNCTR (for the perinucleolar compartment).
Proximity labeling without APEX2 tagging has been successfully conducted by harnessing horseradish peroxidase (HRP) [81]. The application of HRP is limited to cell surface proteins because its activity depends on disulfide bonds and Ca2+-binding sites, which are disrupted in the reducing environment of the cytoplasm. However, in fixed and permeabilized cells, HRP-conjugated secondary antibodies allow proximity labeling in a similar way to immunostaining [82]. This HRP-based approach has been adopted to identify DNA regions adjacent to granules [83,84,85] and, more recently, the transcriptomes of Cajal bodies [86] and KRAS-induced granules [87].
Light-controllable proximity labeling
While hydrogen peroxide treatment in APEX-based methods is short-term, its potential toxicity remains a concern. To circumvent this problem, a light-driven proximity labeling strategy, called chromophore-assisted proximity labeling (CAP)-Seq, has been developed (Fig. 5A) [88]. CAP-Seq uses a mini singlet oxygen generator (miniSOG), an enzyme that produces reactive oxygen species (ROS) upon blue-light illumination. Like phenoxyl radicals, ROS are highly reactive yet short-lived [89], targeting nearby guanine bases of RNA. After RNA extraction, oxidized guanine is coupled to alkyne-amine, followed by a click reaction for biotinylation. CAP-Seq has been applied to stress granule transcriptomes [90]. Although CAP-Seq requires 15–20 min of light exposure and thus offers lower temporal resolution than the APEX2-based approach (1 min of hydrogen peroxide treatment), its specificity is comparable, making it a complementary tool.
Proximity labeling by light and acylating reagents. A Chromophore-assisted proximity labeling (CAP)-Seq. A mini singlet oxygen generator (miniSOG) produces reactive oxygen species (ROS) upon blue-light illumination. ROS oxidize the guanine bases of nearby RNAs, which are then coupled to an alkyne amine and biotinylated via a click reaction. B Halo-Seq. Using the HaloTag system, an ROS generator, dibromofluorescein (DBF), is conjugated to a target protein. C Photoproximity protein interaction (PhotoPPI). The SNAP-tag system recruits a synthetic compound containing a diazirine, a biotin, and a UV-cleavable linker to a specific subcellular region. Upon UV irradiation, diazirine generates a reactive carbene that covalently labels proximal proteins. D Microenvironment mapping (µMap). Blue light exposure activates an iridium photocatalyst, which transfers energy to adjacent diazirines, generating singlet carbenes that label proximal biomolecules. E Bioorthogonal acylating agents for proximity labeling (BAP)-Seq. Acylating agents masked by a 1-methylcyclopropyl ester are activated by BS2 esterase localized in a specific subcellular region
Light-responsive small molecules, rather than enzymes, are also used for the generation of ROS for RNA oxidization. HaloTag, a protein derived from haloalkane dehalogenase, covalently binds to small compounds (HaloTag ligands) [91]. Halo-Seq employs a ligand-conjugated dibromofluorescein (DBF), which generates ROS under illumination [92, 93] and is recruited to the target region via the HaloTag-fused organelle marker protein, allowing nucleolar transcriptomes to be revealed (Fig. 5B) [94]. Because DBF results in a greater yield of ROS than MiniSOG [95], the RNA labeling efficiency is improved.
Given the need for greater precision, more refined methods have emerged, as radicals and ROS diffuse before decaying, resulting in a relatively broad labeling radius. One strategy uses singlet carbenes generated from diazirines upon UV irradiation [96]. Singlet carbenes have an extremely short half-life (~1 ns) compared with biotin-phenoxyl radicals (<1 ms) [67] and ROS (~10 µs) [97], and they instantly bind to biomolecules or are quenched by water. As a result, singlet carbenes exhibit a much shorter labeling radius (~2–4 nm) [96, 98]. In photoproximity protein interaction (PhotoPPI) (Fig. 5C) [99], a custom compound with diazirine, biotin, and a UV-cleavable linker is directed to specific cellular locations through the SNAP-tag system (orthogonal to HaloTag) [100]. After UV activation, the resulting carbene covalently labels nearby proteins.
A narrower labeling radius has been achieved using a newly developed iridium photocatalyst (Fig. 5D) [101]. Upon blue light illumination, the catalyst is excited and transfers energy to adjacent diazirines in the immediate vicinity, producing singlet carbenes. Super-resolution microscopy with catalyst-conjugating antibodies has demonstrated that the diazirine-based method achieves a low-nm range labeling radius that is significantly narrower than that of peroxidase-based labeling (up to 200-fold) [98]. Although the original catalyst was not cell permeable, which limited its use to extracellular microenvironment mapping (µMap) [101], subsequent improvements in permeability have allowed HaloTag-based recruitment to intracellular stress granules for proteome analysis [102]. More recently, further refinements of the catalyst have enabled transcriptome analyses of nuclear RNP granules, including the nucleolus and paraspeckles [103]. Notably, even with a shorter labeling radius, this method maintains high efficiency for RNA labeling. µMAP-Seq can be performed in a 10 cm dish [103], which has the same reaction scale as APEX-Seq, CAP-Seq, and Halo-Seq [69, 88, 104].
Acylation-based proximity labeling
A novel RNA proximity labeling method that exploits acylation has emerged (Fig. 5E) [105]. This method, which is called bioorthogonal acylating agents for proximity labeling (BAP)-Seq, uses specially designed acylating agents that react with RNA. To prevent off-target reactions, the agents are masked by a 1-methylcyclopropyl ester, which is resistant to hydrolysis by endogenous human or murine esterases but can be selectively cleaved by BS2 esterase from Bacillus subtilis [106]. When BS2 is locally expressed, it unmasks the acylating agent, enabling site-specific RNA labeling. After three minutes of treatment with acylating agents, the labeling efficiency is comparable to that of APEX-based methods. Notably, the probe concentration and incubation time should be optimized because higher concentrations and longer treatment increase nonspecific signals even without BS2 expression.
Proximity labeling with mutation analysis
Even with proximity labeling, the biochemical purification of granule RNAs can cause significant loss of RNA and require large amounts of starting materials. For rare samples, such as specific cell types in tissues, researchers have established strategies that mark granule RNAs through nucleotide substitutions, avoiding the purification of the marked RNAs.
The targets of RNA-binding proteins identified by editing (TRIBE) technique uses the catalytic domain of the ADAR RNA-editing enzyme (Fig. 6A) [107, 108]. Fusing the catalytic domain of ADAR to a protein of interest converts adenosine to inosine in bound RNAs. Because inosine is read as guanosine during sequencing, mutations reveal the RNAs associated with that protein. The TRIBE technique has been applied to the FXR1 stress granule protein, facilitating single-cell granule transcriptome analysis [109]. One drawback of this technique is that editing continues as long as the ADAR is expressed, limiting temporal resolution. To overcome this issue, the TRIBE-induced through dimerization (ID) technique, which temporarily recruits ADAR to the marker protein, was developed [110]. By leveraging the rapamycin-driven dimerization of FKBP and FRB [111], researchers can induce transient ADAR activity in stress granules, enabling the observation of time-dependent changes in their transcriptomes.
Proximity labeling with mutation analysis. A Targets of RNA-binding proteins identified by editing (TRIBE). The catalytic domain of the ADAR RNA-editing enzyme converts adenosine to inosine in bound RNAs. Inosine is read as guanosine during sequencing. B Oxidation-induced nucleotide conversion (OINC)-Seq. The ROS generated by DBF oxidize guanine, leading to nucleotide misincorporations or deletions during reverse transcription
A modified version of Halo-Seq that tracks mutation sites, termed oxidation-induced nucleotide conversion (OINC)-Seq, has also been introduced (Fig. 6B) [112]. DBF oxidizes guanine into 8-oxoguanosine or other variants, causing nucleotide misincorporations, such as G-to-T or G-to-C substitutions, or even deletions during reverse transcription [113]. Identifying such mutations enables Halo-Seq without biotinylation and extends its applicability to organisms such as zebrafish.
Photo-isolation chemistry (PIC)
Visually defining the target region under a microscope is often more advantageous than relying solely on marker proteins. Although laser capture microdissection has been proposed for spatial transcriptomics [114], its spatial resolution poses challenges for studying RNP granules. Photo-isolation chemistry (PIC)-Seq addresses this issue by restricting library synthesis to RNAs within a microscopically defined region (Fig. 7) [115]. In PIC-Seq, RNAs in tissue sections undergo in situ reverse transcription with primers that contain a UV-cleavable modification called 6-nitropiperonyloxymethyl deoxythymidine. This “caged” oligonucleotide blocks second-strand synthesis by DNA polymerase until it is removed by UV illumination. Only UV-irradiated regions selected under the microscope allow the formation of double-stranded cDNA after cell lysis. An in vitro transcription step is subsequently performed with T7 polymerase, but the promoter in the adapter is recognized only when it is double-stranded. Therefore, only uncaged RNAs complete library synthesis. PIC-Seq has been applied for transcriptome analysis of RNP granules, including stress granules, nuclear speckles [115], and inclusion bodies (IBs) of mumps virus [116]. Leveraging the ability to specify regions under the microscope, PIC-Seq may further elucidate the heterogeneity of the transcriptome in terms of morphological differences in the granules.
Photo-Isolation Chemistry (PIC). RNAs in tissue sections undergo in situ reverse transcription using primers containing UV-cleavable caged nucleotides. Because these modified nucleotides prevent second-strand synthesis by DNA polymerase, double-stranded cDNA is synthesized only in UV-irradiated regions. Subsequent in vitro transcription by T7 polymerase is initiated from a double-stranded promoter sequence
Future perspectives
Recent breakthroughs in transcriptome analysis by granule purification and proximity labeling have opened the door to a new era of RNP granule research. At the same time, it is important to keep in mind that data obtained through current methods may not fully represent cellular granule states due to various issues as follows: loss of granule components during purification; artifacts introduced by crosslinking; potential cellular toxicity or RNA damage from the required compounds or light exposure; biases in purification or labeling efficiency; and contamination from nongranular fractions. The development of improved methods is urgently needed to address these challenges.
Complementary approaches based on different principles are essential for validation. In particular, fluorescence microscopy techniques, such as single-molecule FISH (smFISH), confirm the localization of granule RNAs by hybridizing multiple fluorescent probes to target RNAs [117, 118]. The recent progress in smFISH toward high-throughput applications involving the use of pooled oligonucleotides and 96-well plates [119] further enhances its utility. Moreover, advanced methods, such as MERFISH [120] and SeqFISH+ [121], which rely on multiple rounds of hybridization, enable the subcellular localization of tens of thousands of RNA species. Integrating deep sequencing with these powerful imaging methods facilitates robust discoveries in RNP granules.
Combining granule purification and proximity labeling with other RNA-Seq-based methods, such as SLAM-Seq to measure RNA half-lives [122] or SHAPE-MaP to determine RNA secondary structures [123], can increase the understanding of how RNP granules contribute to RNA metabolism. Long-read sequencing platforms from Pacific Biosciences (PacBio) and Oxford Nanopore Technologies (ONT) also offer valuable opportunities to examine RNA isoforms and modified bases [124]. As these and other RNA-related technologies continue to be advanced, research on RNP granules will undoubtedly accelerate, remaining a vibrant and rapidly evolving field.
References
Ripin N, Parker R. Formation, function, and pathology of RNP granules. Cell. 2023;186:4737–56.
Marcelo A, Koppenol R, de Almeida LP, Matos CA, Nóbrega C. Stress granules, RNA-binding proteins and polyglutamine diseases: too much aggregation?. Cell Death Dis. 2021;12:592.
Thomas L, Putnam A, Folkmann A. Germ granules in development. Development. 2023;150:dev201037.
Alberti S, Hyman AA. Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat Rev Mol Cell Biol. 2021;22:196–213.
Bauer KE, de Queiroz BR, Kiebler MA, Besse F. RNA granules in neuronal plasticity and disease. Trends Neurosci. 2023;46:525–38.
An H, de Meritens CR, Shelkovnikova TA. Connecting the “dots”: RNP granule network in health and disease. Biochim Biophys Acta Mol Cell Res. 2021;1868:119058.
Wadsworth GM, Srinivasan S, Lai LB, Datta M, Gopalan V, Banerjee PR. RNA-driven phase transitions in biomolecular condensates. Mol Cell. 2024;84:3692–705.
Decker CJ, Burke JM, Mulvaney PK, Parker R. RNA is required for the integrity of multiple nuclear and cytoplasmic membrane-less RNP granules. EMBO J. 2022;41:e110137.
Bounedjah O, Desforges B, Wu T-D, Pioche-Durieu C, Marco S, Hamon L, et al. Free mRNA in excess upon polysome dissociation is a scaffold for protein multimerization to form stress granules. Nucleic Acids Res. 2014;42:8678–91.
Shav-Tal Y, Blechman J, Darzacq X, Montagna C, Dye BT, Patton JG, et al. Dynamic sorting of nuclear components into distinct nucleolar caps during transcriptional inhibition. Mol Biol Cell. 2005;16:2395–413.
Chujo T, Yamazaki T, Hirose T. Architectural RNAs (arcRNAs): A class of long noncoding RNAs that function as the scaffold of nuclear bodies. Biochim Biophys Acta. 2016;1859:139–46.
Nakagawa S, Yamazaki T, Mannen T, Hirose T. ArcRNAs and the formation of nuclear bodies. Mamm Genome. 2022;33:382–401.
Jain A, Vale RD. RNA phase transitions in repeat expansion disorders. Nature. 2017;546:243–7.
Van Treeck B, Protter DSW, Matheny T, Khong A, Link CD, Parker R. RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. Proc Natl Acad Sci USA. 2018;115:2734–9.
Tauber D, Tauber G, Khong A, Van Treeck B, Pelletier J, Parker R. Modulation of RNA condensation by the DEAD-Box protein eIF4A. Cell. 2020;180:411–26.e16.
Budkina K, El Hage K, Clément M-J, Desforges B, Bouhss A, Joshi V, et al. YB-1 unwinds mRNA secondary structures in vitro and negatively regulates stress granule assembly in HeLa cells. Nucleic Acids Res. 2021;49:10061–81.
Langdon EM, Qiu Y, Ghanbari Niaki A, McLaughlin GA, Weidmann CA, Gerbich TM, et al. mRNA structure determines specificity of a polyQ-driven phase separation. Science. 2018;360:922–7.
Kedersha N, Cho MR, Li W, Yacono PW, Chen S, Gilks N, et al. Dynamic shuttling of TIA-1 accompanies the recruitment of mRNA to mammalian stress granules. J Cell Biol. 2000;151:1257–68.
Teixeira D, Sheth U, Valencia-Sanchez MA, Brengues M, Parker R. Processing bodies require RNA for assembly and contain nontranslating mRNAs. RNA. 2005;11:371–82.
Kim TH, Tsang B, Vernon RM, Sonenberg N, Kay LE, Forman-Kay JD. Phospho-dependent phase separation of FMRP and CAPRIN1 recapitulates regulation of translation and deadenylation. Science. 2019;365:825–9.
Lee M, Moon HC, Jeong H, Kim DW, Park HY, Shin Y. Optogenetic control of mRNA condensation reveals an intimate link between condensate material properties and functions. Nat Commun. 2024;15:3216.
Chen R, Stainier W, Dufourt J, Lagha M, Lehmann R. Direct observation of translational activation by a ribonucleoprotein granule. Nat Cell Biol. 2024;26:1322–35.
Ramat A, Haidar A, Garret C, Simonelig M. Spatial organization of translation and translational repression in two phases of germ granules. Nat Commun. 2024;15:8020.
Mateju D, Eichenberger B, Voigt F, Eglinger J, Roth G, Chao JA. Single-molecule imaging reveals translation of mRNAs localized to stress granules. Cell. 2020;183:1801–12.e13.
Kang J-Y, Wen Z, Pan D, Zhang Y, Li Q, Zhong A, et al. LLPS of FXR1 drives spermiogenesis by activating translation of stored mRNAs. Science. 2022;377:eabj6647.
Broix L, Turchetto S, Nguyen L. Coordination between transport and local translation in neurons. Trends Cell Biol. 2021;31:372–86.
Sabari BR, Dall’Agnese A, Young RA. Biomolecular condensates in the nucleus. Trends Biochem Sci. 2020;45:961–77.
Putnam A, Thomas L, Seydoux G. RNA granules: functional compartments or incidental condensates?. Genes Dev. 2023;37:354–76.
Braselmann E, Rathbun C, Richards EM, Palmer AE. Illuminating RNA biology: Tools for imaging RNA in live mammalian cells. Cell Chem Biol. 2020;27:891–903.
Khong A, Matheny T, Jain S, Mitchell SF, Wheeler JR, Parker R. The stress granule transcriptome reveals principles of mRNA accumulation in stress granules. Mol Cell. 2017;68:808–20.e5.
Hubstenberger A, Courel M, Bénard M, Souquere S, Ernoult-Lange M, Chouaib R, et al. P-body purification reveals the condensation of repressed mRNA regulons. Mol Cell. 2017;68:144–57.e5.
König J, Zarnack K, Rot G, Curk T, Kayikci M, Zupan B, et al. iCLIP reveals the function of hnRNP particles in splicing at individual nucleotide resolution. Nat Struct Mol Biol. 2010;17:909–15.
Wheeler JR, Jain S, Khong A, Parker R. Isolation of yeast and mammalian stress granule cores. Methods. 2017;126:12–17.
Xing W, Muhlrad D, Parker R, Rosen MK. A quantitative inventory of yeast P body proteins reveals principles of composition and specificity. Elife. 2020;9:e56525.
Jain S, Wheeler JR, Walters RW, Agrawal A, Barsic A, Parker R. ATPase-modulated stress granules contain a diverse proteome and substructure. Cell. 2016;164:487–98.
Williamson CD, Wong DS, Bozidis P, Zhang A, Colberg-Poley AM. Isolation of endoplasmic reticulum, mitochondria, and mitochondria-associated membrane and detergent resistant membrane fractions from transfected cells and from human Cytomegalovirus-infected primary fibroblasts. Curr Protoc Cell Biol. 2015;68:3.27.1–3.27.33.
Matheny T, Rao BS, Parker R. Transcriptome-wide comparison of stress granules and P-bodies reveals that translation plays a major role in RNA partitioning. Mol Cell Biol. 2019;39:1–18.
Kershaw CJ, Nelson MG, Lui J, Bates CP, Jennings MD, Hubbard SJ, et al. Integrated multi-omics reveals common properties underlying stress granule and P-body formation. RNA Biol. 2021;18:655–73.
Namkoong S, Ho A, Woo YM, Kwak H, Lee JH. Systematic characterization of stress-induced RNA granulation. Mol Cell. 2018;70:175–87.e8.
Al-Husini N, Tomares DT, Pfaffenberger ZJ, Muthunayake NS, Samad MA, Zuo T, et al. BR-bodies provide selectively permeable condensates that stimulate mRNA decay and prevent release of decay intermediates. Mol Cell. 2020;78:670–82.e8.
Muthunayake NS, Al-Husini N, Schrader JM. Differential centrifugation to enrich bacterial ribonucleoprotein bodies (BR bodies) from Caulobacter crescentus. STAR Protoc. 2020;1:100205.
Mariani D, Setti A, Castagnetti F, Vitiello E, Stufera Mecarelli L, Di Timoteo G, et al. ALS-associated FUS mutation reshapes the RNA and protein composition of stress granules. Nucleic Acids Res. 2024;52:13269–89.
Di Timoteo G, Giuliani A, Setti A, Biagi MC, Lisi M, Santini T, et al. m6A reduction relieves FUS-associated ALS granules. Nat Commun. 2024;15:5033.
Curdy N, Lanvin O, Cerapio J-P, Pont F, Tosolini M, Sarot E, et al. The proteome and transcriptome of stress granules and P bodies during human T lymphocyte activation. Cell Rep. 2023;42:112211.
Iadevaia V, Burke JM, Eke L, Moller-Levet C, Parker R, Locker N. Novel stress granule-like structures are induced via a paracrine mechanism during viral infection. J Cell Sci. 2022;135:jcs259194.
Freibaum BD, Messing J, Yang P, Kim HJ, Taylor JP. High-fidelity reconstitution of stress granules and nucleoli in mammalian cellular lysate. J Cell Biol. 2021;220:e202009079.
Khong A, Jain S, Matheny T, Wheeler JR, Parker R. Isolation of mammalian stress granule cores for RNA-Seq analysis. Methods. 2018;137:49–54.
Heraud-Farlow JE, Sharangdhar T, Li X, Pfeifer P, Tauber S, Orozco D, et al. Staufen2 regulates neuronal target RNAs. Cell Rep. 2013;5:1511–8.
Villanueva E, Smith T, Pizzinga M, Elzek M, Queiroz RML, Harvey RF, et al. System-wide analysis of RNA and protein subcellular localization dynamics. Nat Methods. 2024;21:60–71.
Demeshkina NA & Ferré-D’Amaré AR. New purifications reveal yeast and human stress granule cores are discrete particles with complex transcriptomes and proteomes. bioRxiv. 2023. https://doi.org/10.1101/2023.10.12.562116.
Shan T, Liu F, Wen M, Chen Z, Li S, Wang Y, et al. m6A modification negatively regulates translation by switching mRNA from polysome to P-body via IGF2BP3. Mol Cell. 2023;83:4494–508.e6.
Kami D, Ishizaki T, Taya T, Katoh A, Kouji H, Gojo S. A novel mRNA decay inhibitor abolishes pathophysiological cellular transition. Cell Death Discov. 2022;8:278.
Safieddine A, Benassy M-N, Bonte T, Slimani F, Pourcelot O, Kress M, et al. Cell-cycle-dependent mRNA localization in P-bodies. Mol Cell. 2024;84:4191–208.e7.
Jiang J, Shao Q, Xie S, Xiao X, Guo R, Jin M et al. Functional characterization of RNA profiles in processing bodies of human embryonic stem cells and mesodermal cells. bioRxiv. 2023. https://doi.org/10.1101/2023.11.22.568232.
Horste EL, Fansler MM, Cai T, Chen X, Mitschka S, Zhen G, et al. Subcytoplasmic location of translation controls protein output. Mol Cell. 2023;83:4509–23.e11.
Kurusu R, Fujimoto Y, Morishita H, Noshiro D, Takada S, Yamano K, et al. Integrated proteomics identifies p62-dependent selective autophagy of the supramolecular vault complex. Dev Cell. 2023;58:1189–205.e11.
Esposito M, Fang C, Cook KC, Park N, Wei Y, Spadazzi C, et al. TGF-β-induced DACT1 biomolecular condensates repress Wnt signalling to promote bone metastasis. Nat Cell Biol. 2021;23:257–67.
Kim C, Kwon Y, Jeong J, Kang M, Lee GS, Moon JH, et al. Phytochrome B photobodies are comprised of phytochrome B and its primary and secondary interacting proteins. Nat Commun. 2023;14:1708.
Lee KP, Liu K, Kim EY, Medina-Puche L, Dong H, Di M, et al. The m6A reader ECT1 drives mRNA sequestration to dampen salicylic acid-dependent stress responses in Arabidopsis. Plant Cell. 2024;36:746–63.
Zhou Y, Panhale A, Shvedunova M, Balan M, Gomez-Auli A, Holz H, et al. RNA damage compartmentalization by DHX9 stress granules. Cell. 2024;187:1701–18.e28.
Cardona AH, Ecsedi S, Khier M, Yi Z, Bahri A, Ouertani A, et al. Self-demixing of mRNA copies buffers mRNA:mRNA and mRNA:regulator stoichiometries. Cell. 2023;186:4310–24.e23.
Reber S, Jutzi D, Lindsay H, Devoy A, Mechtersheimer J, Levone BR, et al. The phase separation-dependent FUS interactome reveals nuclear and cytoplasmic function of liquid-liquid phase separation. Nucleic Acids Res. 2021;49:7713–31.
Chujo T, Yamazaki T, Kawaguchi T, Kurosaka S, Takumi T, Nakagawa S, et al. Unusual semi-extractability as a hallmark of nuclear body-associated architectural noncoding RNAs. EMBO J. 2017;36:1447–62.
Zeng C, Chujo T, Hirose T, Hamada M. Landscape of semi-extractable RNAs across five human cell lines. Nucleic Acids Res. 2023;51:7820–31.
Lam SS, Martell JD, Kamer KJ, Deerinck TJ, Ellisman MH, Mootha VK, et al. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat Methods. 2015;12:51–54.
Hung V, Udeshi ND, Lam SS, Loh KH, Cox KJ, Pedram K, et al. Spatially resolved proteomic mapping in living cells with the engineered peroxidase APEX2. Nat Protoc. 2016;11:456–75.
Mortensen A, Skibsted LH. Importance of carotenoid structure in radical-scavenging reactions. J Agric Food Chem. 1997;45:2970–7.
Bendayan M. Worth its weight in gold. Science. 2001;291:1363–5.
Fazal FM, Han S, Parker KR, Kaewsapsak P, Xu J, Boettiger AN, et al. Atlas of subcellular RNA localization revealed by APEX-seq. Cell. 2019;178:473–90.e26.
Padrón A, Iwasaki S, Ingolia NT. Proximity RNA labeling by APEX-seq reveals the organization of translation initiation complexes and repressive RNA granules. Mol Cell. 2019;75:875–87.e5.
Shen H, Yanas A, Owens MC, Zhang C, Fritsch C, Fare CM, et al. Sexually dimorphic RNA helicases DDX3X and DDX3Y differentially regulate RNA metabolism through phase separation. Mol Cell. 2022;82:2588–603.e9.
Zhou Y, Wang G, Wang P, Li Z, Yue T, Wang J, et al. Expanding APEX2 substrates for proximity-dependent labeling of nucleic acids and proteins in living cells. Angew Chem Int Ed Engl. 2019;58:11763–7.
Huang J, Zhao R, Qin S, Yang S, Li W, Mo J, et al. 4-thiouridine-enhanced peroxidase-generated biotinylation of RNA. Chembiochem. 2021;22:212–6.
Li R, Zou Z, Wang W, Zou P. Metabolic incorporation of electron-rich ribonucleosides enhances APEX-seq for profiling spatially restricted nascent transcriptome. Cell Chem Biol. 2022;29:1218–31.e8.
Li Y, Tian C, Liu K, Zhou Y, Yang J, Zou P. A clickable APEX probe for proximity-dependent proteomic profiling in yeast. Cell Chem Biol. 2020;27:858–65.e8.
Yassine H, Schrader JM. APEX2 proximity labeling of RNA in bacteria. bioRxiv. 2024. https://doi.org/10.1101/2024.09.18.612050.
Liang J, Han J, Gao X, Jia H, Li R, Tse ECM, et al. Clickable APEX2 probes for enhanced RNA proximity labeling in live cells. Anal Chem. 2024;96:685–93.
Yang H, Patel DJ. Structures, mechanisms and applications of RNA-centric CRISPR-Cas13. Nat Chem Biol. 2024;20:673–88.
Han S, Zhao BS, Myers SA, Carr SA, He C, Ting AY. RNA-protein interaction mapping via MS2- or Cas13-based APEX targeting. Proc Natl Acad Sci USA. 2020;117:22068–79.
Yap K, Chung TH, Makeyev EV. Hybridization-proximity labeling reveals spatially ordered interactions of nuclear RNA compartments. Mol Cell. 2022;82:463–78.e11.
Kotani N, Gu J, Isaji T, Udaka K, Taniguchi N, Honke K. Biochemical visualization of cell surface molecular clustering in living cells. Proc Natl Acad Sci USA. 2008;105:7405–9.
Bar DZ, Atkatsh K, Tavarez U, Erdos MR, Gruenbaum Y, Collins FS. Biotinylation by antibody recognition-a method for proximity labeling. Nat Methods. 2018;15:127–33.
Chen Y, Zhang Y, Wang Y, Zhang L, Brinkman EK, Adam SA, et al. Mapping 3D genome organization relative to nuclear compartments using TSA-Seq as a cytological ruler. J Cell Biol. 2018;217:4025–48.
Zhang L, Zhang Y, Chen Y, Gholamalamdari O, Wang Y, Ma J, et al. TSA-seq reveals a largely conserved genome organization relative to nuclear speckles with small position changes tightly correlated with gene expression changes. Genome Res. 2021;31:251–64.
Suzuki H, Abe R, Shimada M, Hirose T, Hirose H, Noguchi K, et al. The 3’ Pol II pausing at replication-dependent histone genes is regulated by Mediator through Cajal bodies’ association with histone locus bodies. Nat Commun. 2022;13:2905.
Noguchi K, Suzuki H, Abe R, Horiuchi K, Onoguchi-Mizutani R, Akimitsu N, et al. Multi-omics analysis using antibody-based in situ biotinylation technique suggests the mechanism of Cajal body formation. Cell Rep. 2024;43:114734.
Bortoletto A, Yang Z, Cardenas M, Wang M-T, McCormick F & Parchem R. Oncogenic KRAS promotes pancreatic ductal adenocarcinoma (PDAC) through post-transcriptionally regulated KRAS-induced granules (KGs). Res Square. 2023. https://doi.org/10.21203/rs.3.rs-3064215/v1.
Wang P, Tang W, Li Z, Zou Z, Zhou Y, Li R, et al. Mapping spatial transcriptome with light-activated proximity-dependent RNA labeling. Nat Chem Biol. 2019;15:1110–9.
Moan J. On the diffusion length of singlet oxygen in cells and tissues. J Photochem Photobiol B. 1990;6:343–4.
Ren Z, Tang W, Peng L, Zou P. Profiling stress-triggered RNA condensation with photocatalytic proximity labeling. Nat Commun. 2023;14:7390.
Los GV, Encell LP, McDougall MG, Hartzell DD, Karassina N, Zimprich C, et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem Biol. 2008;3:373–82.
Li Y, Aggarwal MB, Nguyen K, Ke K, Spitale RC. Assaying RNA localization in situ with spatially restricted nucleobase oxidation. ACS Chem Biol. 2017;12:2709–14.
Li Y, Aggarwal MB, Ke K, Nguyen K, Spitale RC. Improved analysis of RNA localization by spatially restricted oxidation of RNA-protein complexes. Biochemistry. 2018;57:1577–81.
Engel KL, Lo H-YG, Goering R, Li Y, Spitale RC, Taliaferro JM. Analysis of subcellular transcriptomes by RNA proximity labeling with Halo-seq. Nucleic Acids Res. 2022;50:e24.
Ruiz-González R, Cortajarena AL, Mejias SH, Agut M, Nonell S, Flors C. Singlet oxygen generation by the genetically encoded tag miniSOG. J Am Chem Soc. 2013;135:9564–7.
Knutson SD, Buksh BF, Huth SW, Morgan DC, MacMillan DWC. Current advances in photocatalytic proximity labeling. Cell Chem Biol. 2024;31:1145–61.
Sies H. Strategies of antioxidant defense. Eur J Biochem. 1993;215:213–9.
Oakley JV, Buksh BF, Fernández DF, Oblinsky DG, Seath CP, Geri JB, et al. Radius measurement via super-resolution microscopy enables the development of a variable radii proximity labeling platform. Proc Natl Acad Sci USA. 2022;119:e2203027119.
McCutcheon DC, Lee G, Carlos A, Montgomery JE, Moellering RE. Photoproximity profiling of protein-protein interactions in cells. J Am Chem Soc. 2020;142:146–53.
Keppler A, Gendreizig S, Gronemeyer T, Pick H, Vogel H, Johnsson K. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat Biotechnol. 2003;21:86–89.
Geri JB, Oakley JV, Reyes-Robles T, Wang T, McCarver SJ, White CH, et al. Microenvironment mapping via Dexter energy transfer on immune cells. Science. 2020;367:1091–7.
Pan CR, Knutson SD, Huth SW, MacMillan DWC. µMap proximity labeling in living cells reveals stress granule disassembly mechanisms. Nat Chem Biol. 2025;21:490–500.
Knutson SD, Pan CR, Bisballe N, Bloomer BJ, Raftopolous P, Saridakis I, et al. Parallel proteomic and transcriptomic microenvironment mapping (μMap) of nuclear condensates in living cells. J Am Chem Soc. 2025;147:488-97.
Lo H-YG, Engel KL, Goering R, Li Y, Spitale RC, Taliaferro JM. Halo-seq: An RNA proximity labeling method for the isolation and analysis of subcellular RNA populations. Curr Protoc. 2022;2:e424.
Pani S, Qiu T, Kentala K, Azizi S-A, Dickinson BC. Bioorthogonal masked acylating agents for proximity-dependent RNA labelling. Nat Chem. 2024;16:717–26.
Jones KA, Kentala K, Beck MW, An W, Lippert AR, Lewis JC, et al. Development of a split esterase for protein-protein interaction-dependent small-molecule activation. ACS Cent Sci. 2019;5:1768–76.
McMahon AC, Rahman R, Jin H, Shen JL, Fieldsend A, Luo W, et al. TRIBE: Hijacking an RNA-editing enzyme to identify cell-specific targets of RNA-binding proteins. Cell. 2016;165:742–53.
Xu W, Rahman R, Rosbash M. Mechanistic implications of enhanced editing by a HyperTRIBE RNA-binding protein. RNA. 2018;24:173–82.
van Leeuwen W, VanInsberghe M, Battich N, Salmén F, van Oudenaarden A, Rabouille C. Identification of the stress granule transcriptome via RNA-editing in single cells and in vivo. Cell Rep Methods. 2022;2:100235.
Seo KW, Kleiner RE. Profiling dynamic RNA-protein interactions using small-molecule-induced RNA editing. Nat Chem Biol. 2023;19:1361–71.
Inobe T, Nukina N. Rapamycin-induced oligomer formation system of FRB-FKBP fusion proteins. J Biosci Bioeng. 2016;122:40–46.
Lo H-YG, Goering R, Kocere A, Lo J, Pockalny MC, White LK, et al. Quantification of subcellular RNA localization through direct detection of RNA oxidation. bioRxiv. 2024. https://doi.org/10.1101/2024.11.12.623278.
Alenko A, Fleming AM, Burrows CJ. Reverse transcription past products of guanine oxidation in RNA leads to insertion of A and C opposite 8-oxo-7,8-dihydroguanine and A and G opposite 5-guanidinohydantoin and spiroiminodihydantoin diastereomers. Biochemistry. 2017;56:5053–64.
Nichterwitz S, Chen G, Aguila Benitez J, Yilmaz M, Storvall H, Cao M, et al. Laser capture microscopy coupled with Smart-seq2 for precise spatial transcriptomic profiling. Nat Commun. 2016;7:12139.
Honda M, Oki S, Kimura R, Harada A, Maehara K, Tanaka K, et al. High-depth spatial transcriptome analysis by photo-isolation chemistry. Nat Commun. 2021;12:4416.
Katoh H, Kimura R, Sekizuka T, Matsuoka K, Hosogi M, Kitai Y, et al. Structural and molecular properties of mumps virus inclusion bodies. Sci Adv. 2024;10:eadr0359.
Femino AM, Fay FS, Fogarty K, Singer RH. Visualization of single RNA transcripts in situ. Science. 1998;280:585–90.
Raj A, van den Bogaard P, Rifkin SA, van Oudenaarden A, Tyagi S. Imaging individual mRNA molecules using multiple singly labeled probes. Nat Methods. 2008;5:877–9.
Safieddine A, Coleno E, Lionneton F, Traboulsi A-M, Salloum S, Lecellier C-H, et al. HT-smFISH: a cost-effective and flexible workflow for high-throughput single-molecule RNA imaging. Nat Protoc. 2023;18:157–87.
Chen KH, Boettiger AN, Moffitt JR, Wang S, Zhuang X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science. 2015;348:aaa6090.
Eng C-HL, Lawson M, Zhu Q, Dries R, Koulena N, Takei Y, et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature. 2019;568:235–9.
Herzog VA, Reichholf B, Neumann T, Rescheneder P, Bhat P, Burkard TR, et al. Thiol-linked alkylation of RNA to assess expression dynamics. Nat Methods. 2017;14:1198–204.
Siegfried NA, Busan S, Rice GM, Nelson JAE, Weeks KM. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat Methods. 2014;11:959–65.
Ament IH, DeBruyne N, Wang, F, Lin L. Long-read RNA sequencing: A transformative technology for exploring transcriptome complexity in human diseases. Mol Ther. 2025;33:883–94.
Acknowledgements
We are grateful to all the members of the Iwasaki laboratory for critically reading the manuscript. This work was supported by the Ministry of Education, Culture, Sports, Science and Technology (MEXT) (JP23H04268 to YS; JP24H02307 to SI), the Japan Agency for Medical Research and Development (AMED) (JP23gm6910005 to YS; JP20gm1410001 to SI), the Japan Society for the Promotion of Science (JSPS) (JP23K05648 to YS; JP23H02415 and JP23H00095 to SI), the Nakajima Foundation (to YS), the Exploratory Research Center on Life and Living Systems (ExCELLS) (23EX601 to YS), and RIKEN (Pioneering project “Biology of Intracellular Environments” to YS and SI; RIKEN TRIP initiative “TRIP-AGIS” to SI).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shichino, Y., Iwasaki, S. Expanding toolkit for RNP granule transcriptomics. Polym J 57, 873–883 (2025). https://doi.org/10.1038/s41428-025-01035-7
Received:
Revised:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41428-025-01035-7