Abstract
The prognosis for hepatocellular carcinoma remains grim. Combining radiotherapy with immune checkpoint blockade (ICB) has shown potential to enhance therapeutic outcomes, yet there is a pressing need for further advancements. Our previous research demonstrated that this combined approach suppresses ALKBH5 gene expression and increases m6A modification levels in hepatocellular carcinoma tissues. High-throughput sequencing and detailed molecular analysis revealed that inhibiting ALKBH5 amplifies CIITA m6A modifications post-therapy. This modulation triggers MHC II molecule expression in tumors, facilitating the presentation of tumor-associated antigens to CD4 + T lymphocytes and the recruitment of CD8 + T cells for an anti-tumor immune response. Building on these findings, we engineered a CIITA vector with a specific site mutation to confirm that the regulation of CIITA by the combined radiotherapy and immunotherapy is mediated through m6A methylation. Consequently, we established a comprehensive network involving ALKBH5, CIITA, MHC II, and CD4+ and CD8 + T cells. To elucidate the role and underlying molecular mechanisms of this combined therapy in reshaping the tumor immune microenvironment for hepatocellular carcinoma, we employed multi-omics approaches across in vitro, animal model, and clinical multi-dimensional studies, offering novel insights for enhancing treatment efficacy.
Similar content being viewed by others
Introduction
Liver cancer is a significant contributor to global cancer-related mortality. In 2020, global estimates anticipated over 900,000 new liver cancer diagnoses, with mortality figures exceeding 830,000, closely mirroring these incidence rates [1]. In China, liver cancer notably stands as the second most prevalent cause of death, with 2020 forecasts indicating more than 410,000 new cases and approaching 390,000 fatalities [2, 3]. Current treatment protocols for advanced liver cancer predominantly include systemic therapies, such as chemotherapy, targeted therapy, radiation therapy, and immunotherapy [4]. A pivotal challenge in liver cancer management is the immunosuppressive milieu, which compromises immune responses to tumor antigens and enables tumor cells to evade immune detection [5, 6]. This challenge is reflected in the modest success of exclusive immunotherapy, with objective remission rates ranging between 14% and 19.5% [7,8,9]. Contrastingly, the confluence of immunotherapy with antiangiogenic therapy substantially enhances remission rates, reaching up to approximately 46%. This combination has become increasingly significant in the treatment of advanced hepatocellular carcinoma, emerging as a critical research focus [10]. Furthermore, the amalgamation of radiotherapy and immunotherapy demonstrates potential in amplifying the inflammatory immune response, increasing cytotoxic immune cell infiltration in tumors, and improving both local tumor control and recurrence prevention [11, 12]. Although this combinatory approach is effective, the complexity of its mechanisms in hepatocellular carcinoma remains inadequately understood, necessitating further advancement. Consequently, rigorous research into the molecular mechanisms of immune remodeling influenced by this combined treatment in liver cancer is essential for optimizing its therapeutic efficacy.
In recent years, the study of non-mutated epigenetic factors, particularly m6A methylation modification, has gained prominence in understanding the pathogenesis of malignant tumors [13]. This area, a novel facet of RNA regulation, plays a crucial role in deciphering the complexities of tumor development, immune response dynamics, and mechanisms of drug resistance [14,15,16,17,18]. Research underscores the significance of m6A methylation in upregulating PD-L1 expression, a key factor in immune responses against PD-1 and PD-L1 in cancers such as head and neck squamous carcinoma, breast, and gastric cancers [19, 20]. The role of ALKBH5, an m6A demethylase, in altering the tumor immune environment and enhancing the effectiveness of immunotherapies, has been particularly noted [21,22,23]. PANS’ research in melanoma highlights how ALKBH5 reduction influences Mct4/Slc16a3 expression, lactate levels, and the composition of tumor-infiltrating lymphocytes, thereby impacting immunotherapy outcomes [22]. Intrahepatic cholangiocarcinoma studies reveal that ALKBH5 deficiency leads to increased m6A modification of the PD-L1 mRNA 3’UTR, hastening its degradation via YTHDF2. In vitro and in vivo analyses show that ALKBH5 in tumor cells regulates T-cell function and cytotoxicity by modulating PD-L1 expression [24]. The exploration of m6A modifications in liver cancer research has unveiled new aspects of epigenetic regulation, offering insights into hepatocarcinogenesis and therapeutic resistance [25,26,27]. However, the role of m6A modification in combination treatments, particularly in radiotherapy and immunotherapy for hepatocellular carcinoma, and its influence on the tumor microenvironment, remains an area ripe for further investigation.
CIITA, a pivotal constituent of the NLR protein family [28], is subject to regulation at both the transcriptional and post-transcriptional stages [29]. This molecule is instrumental in immunomodulatory processes, chiefly by mediating the expression of MHC class II molecules within tumor contexts, thereby potentiating the anti-tumor immune response in vivo [30, 31]. Its regulatory dynamics are deeply entwined with epigenetic factors. Recent investigations have pinpointed CIITA as a critical immune prognostic element, closely related to m6A modification. An upsurge in CIITA expression is correlated with the enhanced functionality and longevity of immune effector cells, which in turn is associated with more favorable survival rates in patients [32, 33]. However, a discernible gap in current research is the absence of studies exploring the connection between ALKBH5 and CIITA.
Recent investigations and initial experimental findings indicate that the synergistic application of radiotherapy and immunotherapy in hepatocellular carcinoma therapy markedly diminishes ALKBH5 gene activity, amplifies CIITA-m6A modification, and stimulates the expression of MHC II molecules within tumor cells [34,35,36]. This process crucially mediates the presentation of tumor-associated antigens to CD4 + T lymphocytes, thereby instigating the recruitment of CD8 + T cells, which are instrumental in mounting an effective anti-tumor immune response. To build upon these pivotal findings, our research adopted an integrated multi-omics methodology, incorporating m6A sequencing, transcriptomic analysis, and single-cell sequencing. This methodology aimed to substantiate and dissect the transformative impact of CIITA methylation modifications on the immune microenvironment in hepatocellular carcinoma, unearthing novel mechanistic insights into immune evasion and tolerance. Such insights are pivotal for identifying innovative therapeutic targets to enhance the overall efficacy of hepatocellular carcinoma interventions. Furthermore, the investigation delineated a correlation between ALKBH5/CIITA expression patterns and the clinical-pathological attributes of hepatocellular carcinoma, employing both cytological examinations and animal model studies. This study also undertook an evaluation of the prognostic significance and therapeutic effectiveness of the combined treatment modality, using ALKBH5/CIITA expression profiles, tumor cell dynamics, and immune cell distributions as fundamental biomarkers. This evaluation serves as a cornerstone for advancing towards clinical translational applications.
Material and methods
Cell lines and culture
In this research, the human normal liver cell line LO2, along with a range of liver cancer cell lines, specifically HepG-2, Hep3B, Huh7, Hepa1-6, H22, and SMMC-7721, were utilized. These cell lines, sourced from the Cell Bank of the Chinese Academy of Sciences in Shanghai with STR authentication, underwent cultivation in distinct mediums. The LO2 and H22 lines were propagated in RPMI-1640 medium (Gibco, Life Technologies, USA), in contrast to the HepG-2, Hep3B, Huh7, Hepa1-6, and SMMC-7721 lines, which were cultivated in Dulbecco’s Modified Eagle Medium (DMEM) medium (Gibco, Life Technologies, USA). Both media were supplemented with 10% v/v fetal bovine serum (FBS, Hyclone, USA) and 1% penicillin/streptomycin (Gibco, Life Technologies, USA). The cell cultures were maintained under controlled conditions at 37 °C within a humidified incubator, supplemented with a 5% CO2 atmosphere.
Liver cancer cells sorting
In this experiment, Liver cancer cells were initially incubated with FITC-conjugated anti-CD44 mouse antibodies, procured from Stem Cell Technologies, USA. After this step, the cells were intricately labeled with dextran-coated magnetic nanoparticles, a process facilitated by the employment of bispecific Tetrameric Antibody Complexes (TAC) [37]. Following this, immuno-magnetic separation was conducted, leading to the successful isolation and definitive identification of CD44-positive cells, unequivocally characterized as purified cancer stem cells (CSCs) [38].
Cell transfection and reagents
ALKBH5 and CIITA overexpression and knockdown plasmids were sourced from Gene Corporation in Suzhou, China. The full-length open reading frames (ORFs) of ALKBH5 (NCBI accession NM_001363894.1) and CIITA (NCBI accession NM_003394.4) were inserted into the pcDNA3.1 eukaryotic expression vector to develop their expression vectors. Additionally, knockdown constructs for ALKBH5 (sense sequence: 5’-GGGUCAUUACAGAGAUUAAAU-3’; antisense sequence: 5’-UUAAUCUCUGUAAUGACCCUG-3’) and CIITA (sense sequence: 5’-CGAAUAGACUAAGAUGAAAUG-3’; antisense sequence: 5’-UUUCAUCUUAGUCUAUUCGAG-3’) were constructed and cloned into the pLVX-shRNA vector. In the experimental procedures, 4 μg of these plasmids were transfected into liver cancer CSC cell cultures using Lipofectamine® 2000 (Thermo Fisher Scientific, Inc.) and were subsequently incubated for 24 h in six-well plates. For comparison, control cells were treated with an empty vector.
In vivo tumorigenicity assay
Six-week-old male C57BL/6 mice, sourced from Beijing Huafukang Biotechnology Co., Ltd. (license SCXK 2014-0004, Beijing, China), were housed in a controlled barrier facility. They were allocated to ventilated cages, with no more than four mice per cage, and provided with sterile hardwood bedding. The facility maintained a specific pathogen-free environment at a stable temperature range of 22–26 °C, complemented by a 12-h light-dark cycle. The mice had continuous access to sterile pellet food and autoclaved water. After a seven-day acclimatization period, animals were randomly grouped into four groups: control (Model) group, radiotherapy (Radio) group, anti-PD-1 (PD-1) group and Radio+PD-1 group (n = 5 per group). Hepa1-6 cells (1 × 106 cells per mouse) were subcutaneously implanted into the right axilla of each mouse. Mice in the Radio treatment groups underwent irradiation at 8 Gy per day for three consecutive days. Meanwhile, the mice allocated to the PD-1 treatment groups received tail vein injections of camrelizumab with a frequency of once every three days. Tumor volume was calculated using the formula (volume = width × width × length/2). After a period of 28 days, the mice were euthanized through cervical dislocation following anesthesia with isoflurane. The researcher responsible for outcome measurement was blinded during the trial. This study has been approved by the Animal Research Committee at Mongolia Medical University, with ethical approval number YKD202002012, and all experimental protocols were conducted in accordance with institutional guidelines. All experiments adhered to the ARRIVE guidelines. In this experiment, hepatocellular carcinoma cells were cultured in DMEM supplemented with 10% v/v inactivated calf serum. The cells were incubated at a constant temperature of 37 °C in an atmosphere containing 5% CO2 and with saturated humidity. The culture medium was replaced at 48-h intervals. During their logarithmic growth phase, cells were treated with 0.25% trypsin for one minute to facilitate detachment. After trypsin removal, the cells were resuspended in a small volume of the same medium to ensure a homogeneous mixture. The cell concentration was then determined using a hemocytometer and adjusted to 1 × 106 cells/mL. Mice were subjected to isoflurane-inhalation anesthesia. Once anesthesia was confirmed effective, the cell suspension was carefully injected into the thoracic cavity beneath the liver capsule of the mice [39].
Reverse transcription-quantitative PCR (RT‒qPCR)
Extraction of total RNA was accomplished utilizing the EasyPure RNA Kit (Cat# H30828; TransGen Biotech, Beijing, China), followed by the generation of single-stranded cDNA from the extracted RNA. This process was facilitated using oligo (dT) priming in accordance with the Thermoscript RT kit protocol (Invitrogen, Carlsbad, CA, USA) and employing the iQ™5 Real-Time PCR System (Bio-Rad Laboratories, Inc., USA). The RT-qPCR reactions were conducted in strict compliance with the manufacturer’s instructions. The thermocycling parameters comprised an initial denaturation at 95 °C for 30 s, followed by 40 sequential cycles of 95 °C for 15 s and 60 °C for 30 s each. The assessment of mRNA relative expression levels utilized the 2-ΔΔCt method (cited as reference [16]), with a normalization baseline set against GAPDH mRNA levels. To ensure reproducibility, each assay was conducted in triplicate and independently replicated three times. The quantitative analysis of the data was executed using the relative standard curve method, normalizing gene expression to GAPDH mRNA levels. The experiment incorporated primer sequences synthesized by Sangon (Shanghai, China).
RNA stability detection
In this investigation, 40,000 CSCs were cultivated in a six-well plate configuration over a period of 24 hours. Cells from the initial well were isolated and stored at −80 °C in 1 ml of TRI Reagent for subsequent analysis. The cells in the five remaining wells were exposed to actinomycin D (Act-D; concentration: 10 μg/mL; Sigma-Aldrich, USA). Following Act-D exposure, cell samples were systematically collected at intervals of 1, 2, 4, 6, and 8 hours, each preserved in 1 ml of TRI Reagent as per the established protocol. The extraction of total RNA was followed by its reverse transcription into cDNA, in accordance with documented methods [40]. RNA expression from the initial well was employed as a baseline for normalization. The primer sequences used for amplifying ALKBH5 pre-mRNA were as follows: forward 5’-TGAGCTCGGTGAGAGCAAAG-3’ and reverse 5’-TTAAACCGTGGGGAGACTGC-3’. For ALKBH5 mature-mRNA, the forward primer was 5’-GTGAGCGAGACCCCACTATG-3’, and the reverse primer was 5’-CACATAGCAGCACCAGTGGA-3’.
Total m6A measurement
Cells from each experimental group were subjected to a series of post-drug treatment procedures, beginning with the aspiration of the existing culture medium and followed by a single wash using 1× PBS. This was followed by the administration of 150 μL of reporter gene cell lysate into the wells. The cells were then lysed and centrifuged at 12,000 rpm for five minutes, a step crucial for the extraction of the supernatant for assay purposes. Subsequently, total RNA was extracted using Trizol reagent. This RNA underwent a selective enrichment process for polyA mRNA via Oligo-dT magnetic beads, isolating mRNA with polyA tails. This enriched mRNA was then fragmented into segments of 50–100 nucleotides using a specific fragmentation agent. The fragmented RNA was split into two equal portions. The first portion was subjected to an additional enrichment process for m6A-methylated mRNA fragments, employing m6A antibody-coated immunomagnetic beads. The second portion acted as a control, utilized in the direct construction of a standard transcriptome sequencing library. Following the m6A-modified mRNA fragment enrichment and recovery, a conventional sequencing library was compiled, adhering to the protocol for transcriptome library construction. Finally, both the m6A-seq and RNA-seq libraries underwent partial high-throughput sequencing on a Hiseq 2500 platform, employing the PE100 sequencing modality.
MeRIP-qPCR
To explore the methylation dynamics within RNA on a transcriptome-wide scale, the selective binding of antibodies to methylation-modified bases is employed. This technique involves the use of RNA immunoprecipitation to enrich methylation-modified fragments as the foundation, followed by high-throughput sequencing, ensuring the efficient acquisition of results. In the experimental design, cellular models closely related to the disease are utilized to establish knockdown, knockout, or overexpression cell lines of RNA modification-related factor ALKBH5, collectively referred to as the knockdown/knockout/overexpression group. Each group is compared to wild-type cells, with each sample containing 5 × 10 or more cells. Alternatively, additional samples can be utilized, each containing 5 × 106 to 107 cells. Additionally, a comparative analysis is conducted between normal human cells/tissues and diseased counterparts, constituting two distinct groups. Each group comprises a minimum of three samples, with each sample containing 5 × 106 to 107 cells or 10–20 mg of tissue.
MTT assay
Cellular viability was rigorously assessed post-treatment via the MTT assay. Cells were incubated for 30 min in an MTT solution, standardized to a 5 mg/ml concentration. This incubation facilitated the formation of dark blue formazan crystals in viable cells, which were then solubilized using ethanol (100% concentration) as a lysis agent. Spectrophotometric analysis of the resultant mixture was conducted in the 540 to 595 nm wavelength range, utilizing a microplate reader from Molecular Devices (Sunnyvale, CA, USA). Comparative data analysis was performed, with results expressed as percentages relative to a baseline established by the control group of untreated cells.
Flow cytometry
In this study, single-cell suspensions isolated from both neoplastic and adjacent benign tissues were methodically prepared as detailed in the preceding sections. These cells underwent a preliminary incubation with FcBlock (clone 93, Biolegend) for 15 min, subsequently followed by a 45-minute incubation at 4 °C with fluorophore-conjugated antibodies. Post-incubation, the cells were subjected to a thorough washing in a phosphate-buffered saline (PBS) solution augmented with 0.5% Bovine Serum Albumin (BSA) and 2 mM Ethylenediaminetetraacetic Acid (EDTA), culminating in their analysis via a BD LSRII flow cytometer. The exclusion of non-viable cells was accomplished using 7-Aminoactinomycin D (7AAD), and cell doublets were efficiently segregated and removed based on their distinctive forward and side scatter profiles.
Apoptosis analysis
Quantitative assessment of cellular apoptosis was conducted using an Annexin V-FITC/PI Apoptosis Detection Kit (Hangzhou Lianke Biotechnology Co., Ltd., China), conforming rigorously to the manufacturer’s specified protocols. For this assay, precisely 40,000 cells were suspended in a solution consisting of 5 μL Annexin V-FITC and 10 μL Propidium Iodide (PI). This suspension underwent a 15-min incubation in an environment shielded from light, at standard room temperature. The analysis, executed within a critical timeframe of 60 min post-incubation, was performed using a BD FACSCantoII™ flow cytometer (BD Biosciences), ensuring the accuracy and reliability of the apoptotic rate measurements.
Cell-cycle analysis
In this study, a precise quantity of 10,000 cells was subjected to fixation in 75% ethanol, maintained at a constant temperature of 4 °C throughout the night. Post-fixation, these cells were subjected to centrifugal separation, followed by a single PBS wash. They were then resuspended in 1 milliliter of DNA Staining Solution (sourced from Hangzhou Lianke Biotechnology Co., Ltd., China) and incubated for 30 min in an environment devoid of light. The final stage entailed the assessment of the cell-cycle distribution of the treated cells, employing a Beckman Coulter flow cytometer for this purpose.
TUNEL assay
Quantitative assessment of apoptosis in tendon tissues was conducted using a TUNEL assay (Boster, China), in strict compliance with the manufacturer’s detailed instructions. The protocol commenced with the fixation of cryopreserved tissue sections in 4% paraformaldehyde for 60 minutes. This was followed by a 10-min proteolytic treatment with proteinase K. Subsequently, the sections underwent a sequential incubation: initially with terminal deoxynucleotidyl transferase (TdT), and then with biotinylated deoxyuridine triphosphate (BIO-dUTP). Post-incubation, application of a blocking solution preceded the introduction of streptavidin-biotin complex (SABC) to facilitate fluorescent tagging. The final step involved the detailed visualization of these fluorescently labeled sections utilizing a laser confocal microscope (Olympus, Japan).
Colony formation assay
Investigating anchorage-independent growth, a suspension of 1 × 104 cells/ml was established in 2 ml of 0.3% agar. This medium was augmented with 1% N2 Supplement, 2% B27 Supplement, 20 ng/ml human platelet growth factor, 100 ng/ml epidermal growth factor, and 1% antibiotic-antimycotic, all procured from Invitrogen. The suspension was carefully layered over a 0.5% agar base in 6-well plates. After an incubation period of 21 days, the resultant colonies were stained with 2% crystal violet. Any colony exceeding 0.2 mm in diameter was meticulously counted under a microscope.
Hematoxylin-eosin (H&E) staining
Procured for detailed histopathological scrutiny, allograft tumor specimens were initially fixed in a 4% paraformaldehyde solution and subsequently encased in paraffin. Thin sections, precisely 5 μm in thickness, were methodically stained employing H&E. The intricate examination of areas harboring cancer emboli was conducted using the high-resolution capabilities of an Olympus BX51 microscope.
Immunofluorescence staining
In this experiment, we first permeabilized the tissue sections using a 0.2% Triton X-100 solution for a period of 15 min. This was followed by a blocking stage, where 3% Bovine Serum Albumin (BSA) was applied for an hour. For primary antibody incubation, the sections were maintained overnight at 4 °C with a series of antibodies: anti-CD3 (ab135372, Abcam, UK) at 1:50 dilution, anti-CD8 (53-6.7, Novusbio, USA) also at 1:100 dilution, and anti-CD4 (ab288724, Abcam, UK) at 1:50 dilution. Following this, the sections were cleansed with PBS and subsequently exposed to secondary antibodies (ab150077, ab150167, or ab150157, Abcam, UK) at a 1:500 dilution for a duration of two hours at ambient temperature. Finally, nuclear staining was executed using DAPI (Solarbio), and the resultant images were captured utilizing an Olympus BX51 fluorescence microscope (Olympus Corporation).
Clinical samples
Five pairs of liver cancer tissues and matched adjacent normal tissues were collected from patients with liver cancer at Affiliated Hospital of Inner Mongolia Medical University Hospital. The study protocol was approved by the Ethical Committee of Medical Ethics Committee of Inner Mongolia Medical University (approval number YKD202201227), and informed consent was obtained from all participants.
Immunohistochemistry staining
Tumor tissues and matched adjacent normal tissues were fixed in 4% paraformaldehyde, embedded in paraffin, and sectioned into 5 μm thick slices. After dewaxing, hydration, and antigen retrieval, the sections were blocked with normal goat serum. They were then incubated overnight at 4 °C with the following primary antibodies: anti-ALKBH5 (16837-1-AP, Proteintech, China) and anti-CIITA (PA5-21031, Thermo Fisher, USA). Following primary antibody incubation, the sections were treated with corresponding secondary antibodies for 20 min at room temperature. Finally, observation was carried out using an Olympus BX51 high-resolution optical microscope.
Western blotting assay
The analysis of protein levels in tumor tissues and cells involved lysis using RIPA buffer (Thermo Fisher Scientific, Inc., USA), enhanced with 1% protease inhibitors (Pierce). Protein concentrations were subsequently measured using a BCA protein assay kit (Thermo Fisher Scientific, MA, USA). For each sample, 30 mg of protein was loaded onto PVDF membranes. These membranes were first incubated with primary antibodies at 4 °C overnight, then with HRP-conjugated secondary antibodies at room temperature for 2 h. Detailed antibody specifications are provided in the Supplementary Material (Table S1). Protein signal detection utilized the enhanced chemiluminescence (ECL) reagent (Millipore, Bedford, MA, USA) and analysis was conducted using NIH ImageJ software (Scion, Frederick, MD), with data normalization against glyceraldehyde-3-phosphate dehydrogenase (GAPDH).
Bioinformatics assay
This research leveraged publicly accessible datasets, specifically miRNA and mRNA expression profiles from the Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA). The acquisition of these datasets was facilitated through their respective official websites, including GEO, TCGA, and StarBase. Analysis of the collected data was meticulously conducted using the GEO2R tool [41]. In the realm of survival analysis, we utilized methodologies provided by the GEPIA website and Kaplan-Meier Plotter [42, 43]. Furthermore, the study engaged in Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis, deploying the advanced capabilities of GSEA software.
Statistical analysis
Data compilation employed a Microsoft Excel worksheet, while SPSS version 23.0 underpinned the statistical analysis. Categorical data were delineated as rates or composition ratios. Statistical comparisons hinged on specific thresholds: (1) Pearson’s χ2 test for N ≥ 40 and T ≥ 5, (2) a corrected χ2 test for N ≥ 40 with 1 < T < 5, and (3) Fisher’s exact probability method for N < 40 or T in the range of 1 to 5. The nonparametric rank sum test facilitated comparison of categorical data across different groups. To discern factors influencing tumor marker positivity, an unconditional logistic regression model was deployed. The Kaplan–Meier method served for initial univariate prognosis assessment in liver cancer patients, followed by a Cox regression model for multifaceted prognostic factor analysis, adhering to an α threshold of 0.05. For in vitro and in vivo experiments, ANOVA was employed to assess statistical significance across three or more groups, while the Student’s t-test was utilized for comparisons between two groups. Data are presented as mean values ± standard deviation. Differences were deemed statistically significant when p < 0.05.
Results
Radiotherapy combined with immunotherapy effectively activates immune cells in vivo
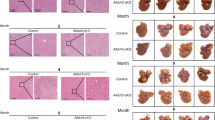
In this study, the synergistic effects of radiotherapy and immunotherapy on immune activation were assessed using C57BL/6 mice. The experimental design included four groups: a baseline control (C57BL/6), a group receiving only 8 Gy radiotherapy, one treated with PD-1 therapy alone, and a combination group subjected to both 8 Gy radiotherapy and PD-1 therapy (Fig. 1A–C). After the treatment period, mice were humanely euthanized, and tumors were collected for analysis (Fig. 1B). Tissue sections underwent H&E staining, revealing a decrease in necrotic tissue in the radiotherapy group versus control, minimal change in the PD-1 group, but a significant reduction in the combination therapy group, indicating its superior efficacy (Fig. 1D). Apoptosis levels were quantified using TUNEL staining; an increase was observed in the radiotherapy group and the PD-1 group relative to control, and a substantial elevation in the combination therapy group (Fig. 1E). Immunofluorescence assays for CD4+ and CD8 + T cell presence showed minimal difference in the radiotherapy group compared to control, a slight increase in the PD-1 group, and a marked increase in the combination group (Fig. 1F, G), suggesting that the latter therapy effectively enhances CD4+ and CD8 + T cell recruitment, potentially improving the tumor microenvironment and leading to more effective tumor cell elimination.
A Tumor formation in C57BL/6 mice. B Images of tumor tissues in mice. C Tumor volume. D H&E staining for pathological changes in different mouse groups. E TUNEL staining images of tissues from each group. F Co-immunostaining of CD3 and CD4 in tissue sections from each group. G Co-immunostaining of CD3 and CD8 in tissue sections from each group. *p < 0.05, **p < 0.01, ***p < 0.001.
To explore potential modifications in mRNA bases during treatment, we assessed the m6A methylation levels in the tissue samples from each group utilizing a colorimetric assay, as depicted in Fig. 2A. This analysis showed minor methylation increments in both the radiotherapy and PD-1 treatment groups compared to the control, with a pronounced escalation following the combined therapy (P < 0.05). Furthermore, comparative analysis of the protein and mRNA expressions of key methylation-related enzymes and proteins, including methylases (METTL3, METTL14, WTAP), demethylases (FTO, ALKBH5), and methylation regulatory proteins (YTHDC1, YTHDC2, YTHDF1, YTHDF2, YTHDF3), was conducted in all groups via Western blotting (Fig. 2B, C) and RT-qPCR (Fig. 2D). Figure 2B–D illustrate that the combined treatment resulted in a decreased expression of methylation enzymes (METTL3, METTL14, WTAP) and the demethylase ALKBH5, with FTO expression remaining constant. Similarly, a decline in YTHDC1, YTHDC2, YTHDF2, and YTHDF3 expressions was observed, whereas YTHDF1 expression remained unchanged. These findings suggest a notable reduction in methylation levels in the cancer models post-combined treatment, with a significant alteration in ALKBH5 expression being the most conspicuous.
A m6A modification status across different groups. B, C Protein expression levels of target genes in different groups. D mRNA expression levels of target genes across different groups. *p < 0.05, **p < 0.01, ***p < 0.001.
Bioinformatics analysis and explore the correlation between ALKBH5 and immune cell population and prognosis
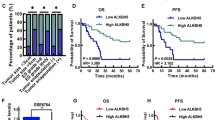
Recent studies have identified significant differential expression of the enzyme ALKBH5, a key player in demethylation, in HCC [44]. This discovery led to an investigation of ALKBH5’s expression across various cancers, utilizing the TCGA database. This analysis showed that ALKBH5 is highly expressed in 11 out of 24 cancer types, including hepatocellular carcinoma (LIHC), corroborating prior observations (Fig. 3A). Expanding on this, ALKBH5 expression in hepatocellular carcinoma was scrutinized using data from the TCGA, STARBASE, and PAN-CANCER databases. This comprehensive examination focused on the relationship between ALKBH5 expression levels and various patient survival factors, including body weight, tumor grade, ethnicity, and gender. For normal-weight individuals, ALKBH5 expression did not exhibit a consistent trend over a decade, as indicated by survival curve intersections. Conversely, among overweight and obese patients, those with lower ALKBH5 expression demonstrated higher survival rates within the first three years (Fig. S1A). A distinct inverse relationship between tumor grade and survival rate was observed; specifically, patients with lower-grade tumors and reduced ALKBH5 expression had extended survival periods — 8 years for Grades 1 and 2, 5 years for Grade 3, and 1 year for Grade 4. Notably, this trend was evident in Grade 3 and 4 patients, where lower ALKBH5 expression correlated with increased survival within respective periods of 5 and 1 year(s) (Fig. S1B). The study also revealed ethnic variations in survival rates linked to ALKBH5 expression. African American patients with lower expression had decreased survival over a 3-year period, while Asian and Caucasian patients exhibited increased survival over 10 and 5 years, respectively (Fig. S1C). Furthermore, gender-specific analysis showed that males with lower ALKBH5 expression had a higher survival rate at 3-4 years, and females at approximately 5 years (Fig. S1D). Collectively, these findings highlight a robust correlation between ALKBH5 expression and patient survival, varying significantly across different demographics.
A Prediction of ALKBH5 expression in different tumors using the TCGA database. B Correlation between ALKBH5 expression levels in liver cancer and patient survival time in TCGA-LIHC cohort. C Expression of ALKBH5 in liver cancer tissues. D Immunohistochemistry staining assessment of protein expression levels of ALKBH5 in liver cancer tissues and adjacent normal tissues. ***p < 0.001.
Furthermore, Kaplan–Meier survival analysis distinctly illustrated that lower ALKBH5 expression in hepatocellular carcinoma patients significantly correlated with improved five-year survival rates, a finding substantiated by the TCGA-LIHC database (Fig. 3B). Subsequent detailed examination disclosed a substantial elevation in ALKBH5 expression within hepatocellular carcinoma tissues (Fig. 3C). Meanwhile, immunohistochemistry staining analysis also confirmed that ALKBH5 expression was significantly elevated in HCC tissues compared to adjacent normal tissues (Fig. 3D). Advancing from previous studies on immune cells, we employed CIBERSORT analysis to probe the interplay between ALKBH5 expression and the composition of immune cells (Fig. 4). This investigation distinctly highlighted an inverse relationship between ALKBH5 expression levels and the presence of specific immune cell subsets.
Correlation scatter plots between ALKBH5 expression levels and the relative abundance of 22 immune cell subsets.
The effect of ALKBH5 on tumor phenotype was evaluated at the cellular level
In this investigation, ALKBH5’s influence on tumor cell phenotypes was meticulously examined. ALKBH5 expression across various tumor cell lines (HepG-2, Hep3B, Huh7, Hepa1-6, H22, SMMC-7721) and the normal liver cell line LO2 was assessed through RT-qPCR. This analysis revealed a consistent upregulation of ALKBH5 in tumor cells relative to LO2, with Hepa1-6 displaying the highest and HepG2 the lowest expression (P < 0.05) (Fig. 5A). Targeted manipulation of ALKBH5 expression—knockdown in Hepa1-6 and overexpression in HepG2—was verified using RT-qPCR and Western blotting, showing more than 70% reduction in Hepa1-6 and a four-to-fivefold increase in HepG2 (Fig. 5B–E). These results confirmed effective vector construction for both interference and overexpression. Subsequent EDU assays indicated a marked decrease in proliferation in ALKBH5-suppressed Hepa1-6 cells and an increase in HepG2 cells with enhanced ALKBH5 expression (P < 0.05) (Fig. 5F). Colony formation assays aligned with these findings, with Hepa1-6 exhibiting reduced colony formation post-ALKBH5 knockdown and HepG2 showing increased formation upon ALKBH5 overexpression, underscoring ALKBH5’s role in tumor cell colony proliferation (Fig. 5G). Scratch assays provided further evidence of ALKBH5’s impact on cell migration: Hepa1-6 demonstrated decreased migration following knockdown, while HepG2 showed increased migration with overexpression (Fig. 5H). Finally, flow cytometry analysis highlighted distinct cell cycle alterations: ALKBH5 knockdown led to cell cycle arrest in Hepa1-6 cells, whereas its overexpression accelerated the cell cycle in HepG2 cells, implying ALKBH5’s pivotal role in promoting hepatocellular carcinoma cell proliferation (Fig. 5I).
A RT-qPCR analysis of ALKBH5 mRNA expression levels in different cell groups. B RT-qPCR assessment of mRNA expression levels in Hepa1-6 cells following ALKBH5 knockdown. C Western blotting analysis of protein expression levels after ALKBH5 knockdown in Hepa1-6 cells. D RT-qPCR detection of mRNA expression levels in HepG-2 cells following overexpression of ALKBH5. E Western blotting analysis of protein expression levels after overexpression of ALKBH5 in HepG-2 cells. F EDU assay for evaluating cell proliferation in groups with knocked-down or overexpressed ALKBH5. G Colony formation assay to assess the proliferative capacity of cells with ALKBH5 knockdown or overexpression. H Scratch assay for examining cell migration ability following ALKBH5 knockdown or overexpression. I Flow cytometry analysis of cell cycle in groups with ALKBH5 knockdown or overexpression. *p < 0.05, **p < 0.01, ***p < 0.001.
The difference of m6A modification level and the type of modified major genes after radiotherapy combined with immunotherapy
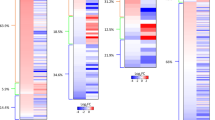
This research aimed to elucidate the variations in m6A methylation and the expression of pivotal effector genes consequent to the administration of radiotherapy combined with immunotherapy. Tissues with notable differences, identified from initial experiments, were subjected to m6A-seq and RNA-seq analyses. These comprised the model group and the group treated with radiotherapy and immunotherapy. The RNA-seq data revealed a notable downregulation of ALKBH5, a key demethylation enzyme, in the treatment group (Fig. 6A and Table S2). Comparative m6A-seq analysis showed that m6A modification was significantly reduced in this group (Fig. 6B, C). Statistical examination of the 3’UTR regions’ methylation patterns indicated a substantial increase following the treatment, depicted via pie charts and bar graphs (Fig. 6D, E).
A Heatmap analysis of m6A methylation-related genes between the model group and the radiation combined with immunotherapy group based on RNA-seq data. B Violin plot. C Overall m6A modification levels in the model group and the radiation combined with immunotherapy group. D Proportion of m6A modifications at different transcript locations in the model group and the radiation combined with immunotherapy group. E Bar graph of m6A modifications in the model group and the radiation combined with immunotherapy group. F Volcano plot of differentially expressed genes. G Volcano plot of differentially m6A modified genes. H Quadrant analysis of combined m6A-seq and RNA-seq data between the model group and the radiation combined with immunotherapy group. I KEGG pathway analysis of differentially expressed genes. J GO (Gene Ontology) analysis of differentially expressed genes.
RNA-seq elucidated a spectrum of genes with differential expression post-treatment, as visualized in volcano plots (Fig. 6F). Correspondingly, m6A-seq highlighted notable gene modifications, also presented in volcano plots (Fig. 6G). A synthesized analysis of the m6A-seq and RNA-seq data was performed, culminating in a detailed four-quadrant visualization (Fig. 6H). Further, KEGG and GO analyses of these genes intimated an immune response activation in tumor tissues following the combined treatment modality (Fig. 6I, J).
This research employed a comprehensive m6A-seq and RNA-seq data integration strategy across four quadrants, facilitating the identification of genes demonstrating simultaneous transcript level elevation and m6A modification. Detailed analysis identified 10 genes with differential expression, as depicted in Fig. 7A. Deeper analysis correlating with transcriptome data highlighted significant m6A modification within the CIITA gene. Utilizing IGV software for methylation quantification of CIITA, we observed increased methylation levels, detailed in Fig. 7B. Comparative assessment of m6A modification across various chromosomes in both control and radiotherapy-immunotherapy treated groups revealed pronounced modifications in chromosome 16, illustrated through bar graphs in Fig. 7C, D. The study culminated with a KEGG pathway enrichment analysis of the most significantly modified genes, illustrating their potential biological significance, as shown in Fig. 7E.
A Heatmap analysis of ten genes between the model group and the radiation combined with immunotherapy group based on RNA-seq data. B Integrative Genomics Viewer (IGV) image of CIITA gene. C Comparison of m6A modification levels on chromosomes in the model group versus the radiation combined with immunotherapy group. D Proportion of m6A modification levels on chromosome 16 in the model group and the radiation combined with immunotherapy group. E Analysis of the vicinity of KEGG pathways in the model group and the radiation combined with immunotherapy group.
This investigation delineated the correlation between CIITA expression and immune cell populations, employing CIBERSORT to reveal a significant inverse relationship with ALKBH5 (Fig. 8A, B). Focused analysis on CIITA within the context of hepatocellular carcinoma was then undertaken, leveraging data from the TCGA, STARBASE, and PAN-CANCER databases. This analysis aimed to delineate the association of CIITA expression with patient survival, incorporating variables such as body weight, tumor grade, ethnicity, and gender (Fig. S2A–D). In patients with a normal body weight, CIITA expression exhibited an oscillatory influence on survival over a span of ten years, lacking a distinct directional trend. Contrastingly, in cohorts categorized as overweight, obese, or severely obese, a marked enhancement in survival rates was observed in those exhibiting elevated CIITA expression within a three-year timeframe (Fig. S2A). A stratified analysis by tumor grade demonstrated a conspicuous positive correlation with survival, where heightened CIITA expression consistently correlated with increased survival durations across varying tumor grades (Fig. S2B). Ethnicity-based survival analysis elucidated divergent trends: African American patients with high CIITA levels experienced reduced survival within a three-year window, while Asian and Caucasian patients displayed augmented survival over periods of ten and five years, respectively (Fig. S2C).
A, B CIBERSORT analysis of the correlation between CIITA and immune cell populations. C Correlation between CIITA expression levels in liver cancer and patient survival time in TCGA-LIHC cohort. D Immunohistochemistry staining assessment of protein expression levels of CIITA in liver cancer tissues and adjacent normal tissues. ***p < 0.001.
In summation, these insights underscore a potential link between CIITA expression and survival in hepatocellular carcinoma. This link is further substantiated by Kaplan-Meier survival plots (Fig. 8C), underscoring a trend where lower CIITA expression aligns with poor survival over a five-year period. Immunohistochemical analysis revealed lower CIITA expression in HCC tissues compared to adjacent normal tissues (Fig. 8D).
The regulatory mechanism of the main effect gene CIITA regulated by m6A and its research value in radiotherapy combined with immunotherapy for hepatocellular carcinoma
This investigation delineates the role of CIITA as a principal gene in the confluence of radiotherapy and immunotherapy. Quantitative analysis of CIITA protein levels in four distinct experimental cohorts was conducted via Western blotting. This analysis disclosed a significant, albeit modest, augmentation of CIITA protein levels in both the standalone radiotherapy and PD-1 cohorts relative to the control group (p < 0.05). A more pronounced amplification was observed in the cohort subjected to the amalgamation of radiotherapy and PD-1 inhibition (Fig. 9A). Complementary MeRIP-RT-qPCR assays corroborated these findings, showing an escalated methylation of CIITA, with the most pronounced increase in the combined therapy cohort (Fig. 9B). Subsequent immunofluorescence assays, assessing CIITA’s co-localization with CD4+ and CD8 + T cells, revealed minimal variance in the radiotherapy-alone group compared to the control. Conversely, a slight upsurge was noted in the PD-1 group, escalating significantly in the combined therapy cohort. These observations suggest that the integration of radiotherapy and PD-1 blockade potentially enhances CIITA expression more robustly, potentially curtailing tumor proliferation through intensified immune cell engagement (Fig. 9C).
A Western blotting assay to detect CIITA protein expression levels in four groups of tumor tissues. B MeRIP-RT-qPCR assay to validate transcriptional modifications of CIITA in four groups of tumor tissues. C Immunofluorescence assay to observe co-localization of CIITA with CD4T and CD8T cells. D RT-qPCR assay to validate the overexpression and knockdown efficiency of CIITA. E Western blotting assay to confirm the overexpression and knockdown efficiency of CIITA. F EDU assay to examine the effect of overexpressing and knocking down CIITA on the proliferative capacity of liver cancer cells. *p < 0.05, **p < 0.01, ***p < 0.001.
This study rigorously examined the pivotal function of CIITA in augmenting the efficacy of radiotherapy when integrated with immunotherapy at the cellular level. We engineered various Hepa1-6 cell lines through either the suppression or augmentation of CIITA expression, yielding Hepa1-6, Hepa1-6-sh-vector, Hepa1-6-sh-CIITA, Hepa1-6-oe-vector, and Hepa1-6-oe-CIITA variants. This genetic modulation was corroborated using western blotting and RT-qPCR methodologies, with results depicted in Fig. 9D, E. Further, we orchestrated a coculture experiment, where 50,000 T cells were incubated with each Hepa1-6 variant, including those subjected to PD-1 + 8 Gy treatment, over 48 h. Post-incubation, the proliferative response of Hepa1-6 cells was meticulously quantified using an EDU assay (Fig. 9F). The data indicated a pronounced increase in Hepa1-6 cell proliferation following CIITA suppression and a significant decrease upon its overexpression. Crucially, the confluence of radiotherapy and immunotherapy was observed to substantially mitigate Hepa1-6 cell proliferation, as delineated in Fig. 9F.
The regulatory effect of ALKBH5 on the major gene CIITA
In this investigation, the ALKBH5-overexpression stabilized cell line, comprising Hepa1-6-oe-vector and Hepa1-6-oe-ALKBH5, was developed to assess changes in CIITA transcript levels. Transcriptome and m6A sequencing analyses revealed a pronounced reduction in both CIITA transcript abundance and m6A methylation levels of CIITA in the Hepa1-6-oe-ALKBH5 group relative to the vector control (Fig. 10A, B). Further, the IGV analysis indicated an decreased m6A level of CIITA modification following ALKBH5 overexpression (Fig. 10C). Furthermore, meRIP-RT-qPCR results revealed a high m6A modification level of CIITA in ALKBH5-knockdown Hepa1-6 cells (Fig. 10D). Collectively, ALKBH5 regulates both the m6A modification status and transcript levels of CIITA in Hepa1-6 cells.
A Volcano plot of differentially expressed genes between the oe-vector group and the oe-ALKBH5 group based on RNA-seq data. B Volcano plot of differentially m6A modified genes between the oe-vector group and the oe-ALKBH5 group based on m6A-seq data. C IGV image of the CIITA gene. D MeRIP-RT-qPCR assay to validate m6A modifications of CIITA between the sh-vector group and the sh-ALKBH5 group. ***p < 0.001.
The regulatory mechanism of ALKBH5 on the major gene CIITA
RT-qPCR experiments were utilized to measure the expression of pre-mRNA-CIITA and mature-mRNA-CIITA, results of which are presented in Fig. 11A. These experiments demonstrate that ALKBH5 suppression results in the upregulation of both RNA forms. Additionally, investigation into the CIITA RNA’s localization via nucleoplasm separation revealed a consistent distribution pattern before and after ALKBH5 knockout, as detailed in Fig. 11B. Furthermore, post-ALKBH5 knockout analysis indicated an increase in the stability of both pre-mRNA-CIITA and mature mRNA-CIITA, as delineated in Fig. 11C.
A RT-qPCR analysis of pre-mRNA-CIITA and mature mRNA-CIITA expression levels. B Nuclear-cytoplasmic fractionation to identify the distribution of CIITA RNA. C Half-life assay demonstrating the regulation of CIITA stability by ALKBH5. *p < 0.05, ***p < 0.001.
CIITA promote the expression of MHC II in vitro
CIITA interference and overexpression constructs were expertly engineered and subsequently transfected into Hepa1-6 cell lines. CIITA interference notably reduced MHC II expression, whereas its overexpression significantly elevated MHC II expression in Hepa1-6 cells (Fig. 12A). This was followed by a meticulous assessment of MHC II expression levels, employing both Western blotting and RT-qPCR, as delineated in Fig. 12B, C. Intriguingly, the co-expression analysis revealed a pronounced increase in MHC II levels in Hepa1-6 cells featuring CIITA overexpression in conjunction with ALKBH5 knockdown (Hepa1-6-oe-CIITA+sh-ALKBH5). Corroborative evidence from the JAPER database suggested a significant regulatory link between CIITA and MHC II, a hypothesis that was further validated through comprehensive chromatin immunoprecipitation (ChIP) assays, detailed in Fig. 12D. This confirmed the formidable regulatory influence of CIITA on MHC II.
A RT-qPCR analysis of MHC II expression levels in Hepa1-6 cells following knockdown and overexpression of CIITA. B RT-qPCR and C Western blot assays were used to detect the expression levels of MHC II under the co-regulation of CIITA and ALKBH5. D ChIP detected the binding interaction between MHC II and CIITA. ***p < 0.001.
Downregulation of CIITA or overexpression of ALKBH5 weakens the anti-tumor effects of radiotherapy combined with immunotherapy for hepatocellular carcinoma in vivo
To further investigate the roles of CIITA and ALKBH5 in the anti-tumor efficacy of radiotherapy combined with immunotherapy, we established tumor models in C57BL/6 mice by implanting Hepa1-6 cells with either CIITA knockdown (sh-CIITA) or ALKBH5 overexpression (oe-ALKBH5). As shown in Fig. 13A–C, downregulation of CIITA or overexpression of ALKBH5 significantly promoted tumor growth, as reflected by increased tumor volume and weight. In contrast, the combination of 8 Gy radiotherapy and anti-PD-1 therapy markedly suppressed tumor growth in mice (Fig. 13A–C). However, downregulation of CIITA or overexpression of ALKBH5 attenuated the anti-tumor effects of the combination therapy in vivo (Fig. 13A–C). These results indicate that tumors with reduced CIITA or elevated ALKBH5 expression exhibit diminished responsiveness to the combination therapy.
A Tumor formation in C57BL/6 mice. B Tumor volume. C Tumor weight. *p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
In this research, we expanded upon our team’s foundational work, investigating the synergistic impact of radiotherapy and immunotherapy in hepatocellular carcinoma (HCC). Our findings demonstrate that this integrative treatment modulates ALKBH5 expression, leading to increased m6A modifications in CIITA. This upregulation significantly bolsters CIITA-mRNA stability and translation, thereby activating MHC II expression in HCC cells. Crucially, this activation spurs tumor specific CD4 + T helper cells to mobilize CD8 + T cells, intensifying the immune response against HCC. Our results not only confirm previous research highlighting the importance of immune modulation in cancer therapy but also illuminate specific molecular pathways, underscoring the roles of ALKBH5 and CIITA.
Employing a robust multi-omics strategy, including m6A sequencing, transcriptomics, and single-cell sequencing, we conducted an in-depth analysis of these molecular interactions. We discovered that the therapy-induced shifts in the m6A modification landscape markedly alter the tumor microenvironment. Increased m6A modification of CIITA initiates a chain of events, culminating in a heightened immune response against HCC. These findings underscore the pivotal role of epigenetic modifications in cancer therapy and suggest innovative avenues for targeted treatment.
Nonetheless, our study has inherent limitations. Extrapolating these findings from cell and animal models to human patients presents notable challenges, given the complexity of HCC’s immune response and the diversity in human physiology. Additionally, the heterogeneity of HCC, both in tumor biology and in treatment response, emphasizes the necessity for personalized therapeutic approaches [45]. Future research should prioritize clinical trials to validate our insights and assess the therapeutic efficacy and safety of targeting the ALKBH5/CIITA-MHC II/CD4 + T cell axis in HCC patients. Investigating the potential of ALKBH5-CIITA as a biomarker for immunotherapy response also merits attention, offering prospects for revolutionizing personalized medicine and enabling more targeted treatment strategies for HCC. Additionally, Given the well-established heterogeneity of HCC [46], future studies on larger, stratified patient cohorts are necessary to precisely delineate the molecular landscape and clinicopathological heterogeneity of ALKBH5 across diverse HCC subtypes. Meanwhile, we focus only on the ALKBH5-CIITA axis in this research. It remains unclear whether and how other m6A-regulated genes contribute to this specific regulatory pathway or the broader modulation of the tumor immune microenvironment. Therefore, future studies employing broader transcriptomic analyses are warranted to construct a more comprehensive network of how m6A modification orchestrates immune responses in HCC.
In summary, this study deepens our understanding of the molecular mechanisms that drive the efficacy of combined radiotherapy and immunotherapy in HCC. By delineating the roles of ALKBH5 and CIITA, we offer novel insights into how epigenetic alterations modulate the immune response in cancer. These insights underscore the potential of integrated treatment strategies in HCC, propelling oncological research towards enhancing patient outcomes through tailored and more effective therapeutic interventions.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and supplementary material. Data supporting the conclusion of this study were included in this manuscript. Additionally, the raw MeRIP-seq data related to this study has been deposited in the OMIX database, with accession numbers OMIX013797 and OMIX013777.
References
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Sayiner M, Golabi P, Younossi ZM. Disease burden of hepatocellular carcinoma: a global perspective. Dig Dis Sci. 2019;64:910–7.
Qiu H, Cao S, Xu R. Cancer incidence, mortality, and burden in China: a time-trend analysis and comparison with the United States and United Kingdom based on the global epidemiological data released in 2020. Cancer Commun (Lond). 2021;41:1037–48.
Balogh J, Victor D 3rd, Asham EH, Burroughs SG, Boktour M Saharia A, et al. Hepatocellular carcinoma: a review. J Hepatocell Carcinoma. 2016;3:41–53.
Zhou G, Boor PPC, Bruno MJ, Sprengers D, Kwekkeboom J. Immune suppressive checkpoint interactions in the tumour microenvironment of primary liver cancers. Br J Cancer. 2022;126:10–23.
Lee JC, Green MD, Huppert LA, Chow C, Pierce RH, Daud AI. The liver-immunity nexus and cancer immunotherapy. Clin Cancer Res. 2022;28:5–12.
El-Khoueiry AB, Sangro B, Yau T, Crocenzi TS, Kudo M, Hsu C, et al. Nivolumab in patients with advanced hepatocellular carcinoma (CheckMate 040): an open-label, non-comparative, phase 1/2 dose escalation and expansion trial. Lancet. 2017;389:2492–502.
Kudo M, Finn RS, Qin S, Han KH, Ikeda K, Piscaglia F, et al. Lenvatinib versus sorafenib in first-line treatment of patients with unresectable hepatocellular carcinoma: a randomised phase 3 non-inferiority trial. Lancet. 2018;391:1163–73.
Qin S, Ren Z, Meng Z, Chen Z, Chai X, Xiong J, et al. Camrelizumab in patients with previously treated advanced hepatocellular carcinoma: a multicentre, open-label, parallel-group, randomised, phase 2 trial. Lancet Oncol. 2020;21:571–80.
Mei J, Tang YH, Wei W, Shi M, Zheng L, Li SH, et al. Hepatic arterial infusion chemotherapy combined with PD-1 inhibitors plus lenvatinib versus PD-1 inhibitors plus lenvatinib for advanced hepatocellular carcinoma. Front Oncol. 2021;11:618206.
Lee YH, Tai D, Yip C, Choo SP, Chew V. Combinational immunotherapy for hepatocellular carcinoma: radiotherapy, immune checkpoint blockade and beyond. Front Immunol. 2020;11:568759.
Huang Y, Sheng H, Xiao Y, Hu W, Zhang Z, Chen Y, et al. Wnt/β-catenin inhibitor ICG-001 enhances the antitumor efficacy of radiotherapy by increasing radiation-induced DNA damage and improving tumor immune microenvironment in hepatocellular carcinoma. Radiother Oncol. 2021;162:34–44.
Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12:31–46.
Desrosiers R, Friderici K, Rottman F. Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci USA. 1974;71:3971–5.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505:117–20.
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, et al. m(6)A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep. 2017;18:2622–34.
Zheng Q, Hou J, Zhou Y, Li Z, Cao X. The RNA helicase DDX46 inhibits innate immunity by entrapping m(6)A-demethylated antiviral transcripts in the nucleus. Nat Immunol. 2017;18:1094–103.
Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, et al. Essential role of METTL3-mediated m(6)A modification in glioma stem-like cells maintenance and radioresistance. Oncogene. 2018;37:522–33.
Li H, Wu H, Wang Q, Ning S, Xu S, Pang D. Dual effects of N(6)-methyladenosine on cancer progression and immunotherapy. Mol Ther Nucleic Acids. 2021;24:25–39.
Zhang Y, Li L, Ye Z, Zhang L, Yao N, Gai L. Identification of m6A methyltransferase-related genes predicts prognosis and immune infiltrates in head and neck squamous cell carcinoma. Ann Transl Med. 2021;9:1554.
Yan G, An Y, Xu B, Wang N, Sun X, Sun M. Potential impact of ALKBH5 and YTHDF1 on tumor immunity in colon adenocarcinoma. Front Oncol. 2021;11:670490.
Li N, Kang Y, Wang L, Huff S, Tang R, Hui H, et al. ALKBH5 regulates anti-PD-1 therapy response by modulating lactate and suppressive immune cell accumulation in tumor microenvironment. Proc Natl Acad Sci USA. 2020;117:20159–70.
Mo P, Xie S, Cai W, Ruan J, Du Q, Ye J, et al. N(6)-methyladenosine (m(6)A) RNA methylation signature as a predictor of stomach adenocarcinoma outcomes and its association with immune checkpoint molecules. J Int Med Res. 2020;48:300060520951405.
Qiu X, Yang S, Wang S, Wu J, Zheng B, Wang K, et al. M(6)A Demethylase ALKBH5 regulates PD-L1 expression and tumor immunoenvironment in intrahepatic cholangiocarcinoma. Cancer Res. 2021;81:4778–93.
Kim GW, Imam H, Khan M, Mir SA, Kim SJ, Yoon SK, et al. HBV-induced increased N6 methyladenosine modification of PTEN RNA affects innate immunity and contributes to HCC. Hepatology. 2021;73:533–47.
Chen Y, Zhao Y, Chen J, Peng C, Zhang Y, Tong R, et al. ALKBH5 suppresses malignancy of hepatocellular carcinoma via m(6)A-guided epigenetic inhibition of LYPD1. Mol Cancer. 2020;19:123.
Zhang C, Huang S, Zhuang H, Ruan S, Zhou Z, Huang K, et al. YTHDF2 promotes the liver cancer stem cell phenotype and cancer metastasis by regulating OCT4 expression via m6A RNA methylation. Oncogene. 2020;39:4507–18.
León Machado JA, Steimle V. The MHC Class II transactivator CIITA: Not (Quite) the odd-one-out anymore among NLR proteins. Int J Mol Sci. 2021;22:1074.
LeibundGut-Landmann S, Waldburger JM, Krawczyk M, Otten LA, Suter T, Fontana A, et al. Mini-review: specificity and expression of CIITA, the master regulator of MHC class II genes. Eur J Immunol. 2004;34:1513–25.
van Eggermond MC, Boom DR, Klous P, Schooten E, Marquez VE, Wierda RJ, et al. Epigenetic regulation of CIITA expression in human T-cells. Biochem Pharmacol. 2011;82:1430–7.
Oda T, Nakamura R, Kasamatsu T, Gotoh N, Okuda K, Saitoh T, et al. DNA-double strand breaks enhance the expression of major histocompatibility complex class II through the ATM-NF-κΒ-IRF1-CIITA pathway. Cancer Gene Ther. 2022;29:225–40.
Zhang L, Luo Y, Cheng T, Chen J, Yang H, Wen X, et al. Development and validation of a prognostic N6-methyladenosine-related immune gene signature for lung adenocarcinoma. Pharmgenom Pers Med. 2021;14:1549–63.
Xia T, Wu X, Cao M, Zhang P, Shi G, Zhang J, et al. The RNA m6A methyltransferase METTL3 promotes pancreatic cancer cell proliferation and invasion. Pathol Res Pract. 2019;215:152666.
Tang KC, Trzaska KA, Smirnov SV, Kotenko SV, Schwander SK, Ellner JJ, et al. Down-regulation of MHC II in mesenchymal stem cells at high IFN-gamma can be partly explained by cytoplasmic retention of CIITA. J Immunol. 2008;180:1826–33.
Forlani G, Shallak M, Celesti F, Accolla RS. Unveiling the hidden treasury: CIITA-driven MHC class II expression in tumor cells to dig up the relevant repertoire of tumor antigens for optimal stimulation of tumor specific CD4+ T helper cells. Cancers (Basel). 2020;12:3181.
Johnson AM, Bullock BL, Neuwelt AJ, Poczobutt JM, Kaspar RE, Li HY, et al. Cancer cell-intrinsic expression of MHC class II regulates the immune microenvironment and response to anti-PD-1 therapy in lung adenocarcinoma. J Immunol. 2020;204:2295–307.
Patel S, Waghela B, Shah K, Vaidya F, Mirza S, Patel S, et al. Silibinin, A natural blend in polytherapy formulation for targeting Cd44v6 expressing colon cancer stem cells. Sci Rep. 2018;8:16985.
Lai HT, Chiang CT, Tseng WK, Chao TC, Su Y. GATA6 enhances the stemness of human colon cancer cells by creating a metabolic symbiosis through upregulating LRH-1 expression. Mol Oncol. 2020;14:1327–47.
Yin H, Fu X, Gao H, Gao H, Ma Y, Chen X, et al. Hybrid peptide NTP-217 triggers ROS-mediated rapid necrosis in liver cancer cells by induction of mitochondrial leakage. Front Oncol. 2022;12:1028600.
Fang F, Lu J, Sang X, Tao YF, Wang JW, Zhang ZM, et al. Super-enhancer profiling identifies novel critical and targetable cancer survival gene LYL1 in pediatric acute myeloid leukemia. J Exp Clin Cancer Res. 2022;41:225.
Janmaat VT, Nesteruk K, Spaander MCW, Verhaar AP, Yu B, Silva RA, et al. HOXA13 in etiology and oncogenic potential of Barrett’s esophagus. Nat Commun. 2021;12:3354.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–w102.
Nagy Á, Lánczky A, Menyhárt O, Győrffy B. Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep. 2018;8:9227.
You Y, Wen D, Zeng L, Lu J, Xiao X, Chen Y, et al. ALKBH5/MAP3K8 axis regulates PD-L1+ macrophage infiltration and promotes hepatocellular carcinoma progression. Int J Biol Sci. 2022;18:5001–18.
Giannelli G, Rani B, Dituri F, Cao Y, Palasciano G. Moving towards personalised therapy in patients with hepatocellular carcinoma: the role of the microenvironment. Gut. 2014;63:1668–76.
Zhu W, Ru L, Ma Z. Identification of a novel four-gene signature correlated with the prognosis of patients with hepatocellular carcinoma: a comprehensive analysis. Front Oncol. 2021;11:626654.
Funding
This study was supported by National Natural Science Foundation of China (82260481, 81860534, 82560042); Natural Science Foundation of Inner Mongolia (2021MS08152, 2022MS08074, 2022MS08065, 2022MS08007, 2024MS08009); Program for Young Talents of Science and Technology in Universities of Inner Mongolia Autonomous Region (NJYT22004); Major Project of Inner Mongolia Medical University (YKD2022ZD002); Radiobiology system and team construction of radiotherapy for Inner Mongolia Medical University (YKD2022XK014); Key Laboratory of Radiation Physics and Biology of Inner Mongolia Medical University (PIKY2023030); Public hospital reform and high-quality development demonstration project research fund, gastrointestinal tumors (2023SGGZ114, 2023SGGZ076); Clinical Need Oriented Basic Research Project of Inner Mongolia Academy of Medical Sciences (2023GLLH0136, 2023GLLH0142, 2023GLLH0007, 2023GLLH0139; 2024GLLH0332); Key Joint Fund Project of Natural Science Foundation of Inner Mongolia Autonomous Region (2025ZDLH019); Central guiding local technology development projects (2023ZY0021, 2024ZY0152). Science and Technology Program Project of Inner Mongolia Autonomous Region (2022YFSH0112); General Project of Inner Mongolia Medical University (YKD2025MS020); Key Research and Development Science and Technology Cooperation Project of Inner Mongolia Autonomous Region (2025KJHZ0014); Affiliated Hospital of Inner Mongolia Medical University National Natural Science Foundation Cultivation Project (2023NYFYPY004).
Author information
Authors and Affiliations
Contributions
Fei Wang, Hui Hou and Hong Yang designed the experiments and provided materials and technical support. Ting Dou and Zhenxia Wang performed experiments. Hongdan Fu, Hong Li, and Wei Zhang performed experiments involved in virus infection and analyzed the data. Haiping Zhao and Hao Yang wrote the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The animal study has been approved by the Animal Research Committee at Mongolia Medical University, with ethical approval number YKD202002012, and all experimental protocols were conducted in accordance with institutional guidelines. The human study protocol was approved by the Ethical Committee of Medical Ethics Committee of Inner Mongolia Medical University (approval number YKD202201227), and informed consent was obtained from all participants.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, F., Hou, H., Yang, H. et al. Regulatory mechanisms of ALKBH5/CIITA axis in the synergistic modulation of hepatocellular carcinoma radiotherapy and immunotherapy. Genes Immun (2026). https://doi.org/10.1038/s41435-026-00382-6
Received:
Revised:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41435-026-00382-6