Abstract
OSCA/TMEM63 channels, which have transporter-like architectures, are bona fide mechanosensitive (MS) ion channels that sense high-threshold mechanical forces in eukaryotic cells. The activation mechanism of these transporter-like channels is not fully understood. Here we report cryo-EM structures of a dimeric OSCA/TMEM63 pore mutant OSCA1.1-F516A with a sequentially extracellular dilated pore in a detergent environment. These structures suggest that the extracellular pore sequential dilation resembles a flower blooming and couples to a sequential contraction of each monomer subunit towards the dimer interface and subsequent extrusion of the dimer interface lipids. Interestingly, while OSCA1.1-F516A remains non-conducting in the native lipid environment, it can be directly activated by lyso-phosphatidylcholine (Lyso-PC) with reduced single-channel conductance. Structural analysis of OSCA1.1-F516A in lyso-PC-free and lyso-PC-containing lipid nanodiscs indicates that lyso-PC induces intracellular pore dilation by attracting the M6b to upward movement away from the intracellular side thus extending the intracellular pore. Further functional studies indicate that full activation of MS OSCA/TMEM63 dimeric channels by high-threshold mechanical force also involves the opening of both intercellular and extracellular pores. Our results provide the fundamental activation paradigm of the unique transporter-like MS OSCA/TMEM63 channels, which is likely applicable to functional branches of the TMEM63/TMEM16/TMC superfamilies.
Similar content being viewed by others
Introduction
Cells sense physical forces through dedicated receptors and respond accordingly under both physiological and pathological conditions from bacteria to humans. How cells convert forces into cellular signals remains largely unresolved at the molecular level. Encouragingly, specific membrane channels have been discovered in both prokaryotic and eukaryotic organisms, and shown to play an important role in various mechanosensory processes such as touch, hearing, and osmosensing1. Bona-fide mechanosensing or MS channels are the key to better understanding the processes of converting physical forces to cellular signals. Since the first prokaryotic MS channel, a mechanosensitive ion channel of large conductance (MscL), was identified2, only a few bona-fide eukaryotic MS channels have been discovered, such as MS K2P3, PIEZO4, and OSCA/TMEM635,6 channels. The MS K2P channels are potassium channels, while the PIEZO and OSCA/TMEM63 channels have the potential to excite the cell as they are calcium-permeable. PIEZO channels are mainly responsible for low-threshold force4, while OSCA channels are responsible for high-threshold force5. Resolving the structural basis of these channels has led to a paradigm shift in our understanding of cellular responses to physical forces.
The OSCA1 gene, originally found in Arabidopsis, mediates osmotic stress-induced calcium increases and is essential for osmotic sensing7. It was identified as a bona fide mechanosensitive ion channel using pressure clamp technology5. The mechanosensitive property of OSCA/TMEM63 family is evolutionarily involved in many processes, including insect detection of food texture8, auditory processes in mice9, and neural activities10. The first reported structure of OSCA/TMEM63 channel from plants displays eleven transmembrane (M) helices for each protomer and assembles into a dimer5. The channel forms a long and hourglass-like pore, very different from other ion channels with a central pore5. However, the dimeric architecture exhibits structural resemblance with the TMEM16 superfamily, which consists of calcium-activated anion channels and lipid scramblases11,12,13. Meanwhile, the OSCA/TMEM63 channels also exhibit structural similarity to another transmembrane channel-like subunit (TMC), which serves as the primary subunit in the auditory TMC complex14. Like the CLC family, which contains both ion channels and transporters, we designate the OSCA/TMEM63 channels, which have a transporter-like architecture, as transporter-like channels15. Interestingly, while the mammalian OSCA/TMEM63 channels possess higher pressure sensitivity, they are different from the plant dimeric OSCA/TMEM63 due to their monomeric formation16,17.
Intriguingly, while most MS channels share the concept of sensing and transducing forces from the surrounding bilayer, also known as “force-from-lipids”18,19, the architectures of MS channel families are strikingly different20,21,22. Therefore, it is likely that each MS channel family has developed distinctive molecular mechanisms to sense mechanical force. Within the bacteria, heptameric MS channels, at least three types of lipids called pore lipids, gatekeeper lipids, and pocket lipids are responsible for sensing and transducing channel force at different positions21. Similarly, in the pseudo-tetrameric MS K2P channels, there are also at least three sorts of lipids in their binding sites that regulate channel activation20,23. Notably, the pore lipids are shared by both MscS and MS K2P channels, where they seal the channel pore20,21. Thus, upon activation, the mechanical force will move the pore lipids out of the pore in these MS channels. In the trimeric PIEZO channels, membrane curvature changes are likely playing the dominant role in channel activation22, and the role of the lipids in PIEZO channels remains to be further explored. Although our previous research suggested that comparable pore lipid and dimer interface lipids in the OSCA/TMEM63 channels16, their role in channel gating is also poorly understood. Moreover, the underlying mechanism for their activation by mechanical forces remains unknown due to the lack of activation structures.
In this work, we report the cryo-EM structures of OSCA1.1-F516A that display three distinct thinner transmembrane conformations with extracellular sequentially dilated pores similar to the process of flower blooming. Interestingly, while OSCA1.1-F516A is non-conducting, it can be activated directly by lyso-PC through an upward movement of M6b that opens the intracellular pore. Further functional studies reveal that the critical residues involved in extracellular and intracellular pore opening also play dominant roles in the full activation of OSCA1.1 by mechanical force. We propose that the full activation of the transporter-like OSCA/TMEM63 channels by high-threshold mechanical force requires the opening of both extracellular and intracellular pores.
Results
Cryo-EM structures of OSCA1.1-F516A with a thinner transmembrane thickness
Activation of OSCA/TMEM63 channels requires a higher mechanical force threshold compared to PIEZO channels4,5. This makes it quite challenging to visualize the gating process of OSCA/TMEM63 channels using standard cryo-EM technology, as it is difficult to apply such a high-threshold force to the proteins. We overcame this by screening and capturing OSCA1.1 mutant structures. We have shown previously that OSCA1.1 protein is a homodimer, with each containing eleven M helices (M0-M10) to form the pore. The Q398, T515, and F516 residues are located at the constricted site of the pore, preventing the ion flow (Supplementary Fig. 1). Two mutants, OSCA1.1-Q398E and OSCA1.1-T515E, display a similar structural conformation to the wild-type, although we observed a slight extension or contraction at the dimer interface (Supplementary Fig. 2–4), as seen in wildtype OSCA1.116. On the other hand, we obtained three cryo-EM structures of pore mutant OSCA1.1-F516A at an overall resolution of 2.8 Å, 2.5 Å, and 2.7 Å, respectively (Supplementary Fig. 5 and Table S1). These three maps display very clear complexes embedded in the lipid/micelle configuration, which allowed us to calculate the thickness of transmembrane domains accurately. Surprisingly, all three OSCA1.1-F516A maps likely exhibit thinner transmembrane, ranging from 49 Å to 53 Å, compared to 58 Å of the wildtype transmembrane (Fig. 1). The reduction in the thickness observed for OSCA1.1-F516A is likely to represent structural changes responding to mechanical force.
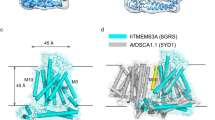
a Cryo-EM density map of non-conducting wild-type OSCA1.1 in lipid nanodiscs (8GRN [https://www.rcsb.org/structure/8GRN]) (purple) from the side view. The approximate extent of the phospholipid bilayer is shown as a thin black line. The length of the protein and the height of the TM helices are labeled. b–d Three cryo-EM density maps of OSCA1.1-F516A in non-conducting state 1 (b, brown), non-conducting state 2 (c, pink), and conducting state (d, cyan) with lipid/detergent micelle from the side view. e Side view of the wild-type OSCA1.1 dimer model in the closed state (8GRN [https://www.rcsb.org/structure/8GRN]). Coloring follows the same scheme as in the density map. The protein is shown as a cartoon and the potential lipid in the central cavity is shown as spheres. f–h Side view of OSCA1.1-F516A in non-conducting state 1 (f, brown), non-conducting state 2 (g, pink), and conducting state (h, cyan).
The three conformations present sequential extracellular pore dilation
Three accurate models were built for the three high-resolution maps of OSCA1.1-F516A, which allowed us to analyze the conformational changes between them (Supplementary Figs. 6–8). In the closed state of wildtype OSCA1.1, the long and narrow hourglass-shaped ion-permeable pores are mainly composed of M0, M3, M4, M5, and M65. The hourglass-shaped pore is separated by a restriction gate into an extracellular pore region and an intracellular pore region. Pore diameters were analyzed using the Hole2 software24, revealing that the extracellular pores of the three different conformations are stepwise extended compared to that of the closed wild-type OSCA1.1 in the lipid nanodiscs (Fig. 2). To verify that the dilated extracellular pores are not obstructed by potential lipids/detergents in the low-resolution configuration, we applied low-pass filter to the three OSCA1.1-F516A mutant density maps by ChimeraX software. The low-pass filtered density maps also display significant stepwise dilated pores and no significant visualizable lipids/detergents seal the pore (Supplementary Fig. 9). As these structures exhibit dilated extracellular pores and thinner transmembrane thickness, it suggests that they could represent the closed state to open state transition process. It is worth noting that OSCA1.1 may undergo the inactivation mode, the three extracellular pores sequentially dilated state, or may represent the open state to the inactivated state transition process. We shall refer to the three distinct states as non-conducting state 1, non-conducting state 2, and conducting state, respectively. When in the conducting state, the diameter of the narrowest constrict site is over 5 Å (Fig. 2e–h), probably enabling the permeation of some ions and even some organic compounds. This observation aligns with our previous research, which indicated that the OSCA1.1 MS channel has a lower level of ion selectivity5.
a Side view of one wildtype OSCA1.1 protomer in the closed state (purple). The calculated pore profile is shown as colored dots using Hole2 software. b–d Side views of one OSCA1.1-F516A protomer in non-conducting state 1 (b, brown), non-conducting state 2 (c, pink), and conducting state (d, cyan). e Van der Waals radii of the closed wildtype OSCA1.1 pore plotted against axial distance. The positions of the restrictive residues F516, Q398, F517, Y520, and K436 that are blocking the pore are marked by arrows. The diameter of the narrowest constriction site is less than 1 Å. f–h Van der Waals radii of the non-conducting state 1 (f brown), non-conducting state 2 (g, pink), and conducting state (h, cyan) OSCA1.1-F516A pore plotted against axial distance. The diameter of the narrowest constriction site in non-conducting state 1 (f, brown), non-conducting state 2 (g, pink), and conducting state is 3 Å, 4 Å and 5 Å, respectively. i The positions of the restrictive residues F516, Q398, F517, Y520, and K436 (colored in red) in the closed wildtype OSCA1.1 pore (purple). The protein is shown as a cartoon with a gray surface. The central pore is shown as a red star. The lipid in the pore domain is shown as spheres. j–l The positions of the hydrophobic residues F516, Q398, F517, Y520, and K436 (colored in red) in non-conducting state 1 (j, brown), non-conducting state 2 (k pink), and conducting state (l cyan).
Extracellular pore dilation couples each monomer sequential contraction and interface lipids extrusion
Sequential structural alignment of the closed state16, non-conducting state 1, non-conducting state 2, and conducting state of OSCA1.1 dimer structure indicates that the cytosolic domain (CTD), which comprises the cytosolic dimer interface and two anti-parallel transmembrane helices, remains relatively stable during channel activation. This suggests that the CTD likely acts as a stent for the channel’s main body (M0–M10) embedded in the lipid membrane (Supplementary Fig. 12a–d). Notably, the M0-M10 main body sections display sequential contraction towards the central dimer interface within the view perpendicular to the cell membrane plane and sequential anti-clockwise rotation parallel to the plane of the cell membrane during the transition from the closed to conducting state (Fig. 3). This results in a sequential reduction of the length of the dimeric OSCA1.1 protein from about 158 Å to 148 Å (Fig. 1), accompanied by an expansion of the extracellular pores. Significantly, the lipids-like molecules in the cytosolic side of the dimer interface are extruded from the gap created by M3 and M10 in the conducting state (Supplementary Fig. 10a–d and 11). Furthermore, the M5 and M6 linker (NQSPN) of each monomer creates an additional protein-protein interaction dimer interface via their main and side chains (Supplementary Fig. 10e). Our previous research into OSCA3.1 revealed a similar contraction phenomenon16. Although OSCA3.1 has a strong tendency to contract, its contracted state remains non-conductive, which is consistent with the fact that OSCA3.1 requires an additional higher mechanical force to activate the contracted state, thus explaining the failure of OSCA3.1 to operate in drought-resistant calcium response7.
a Top view of closed OSCA1.1 colored in purple is shown as a cartoon. Helices are shown as cylinders and the interface lipid in the central cavity is shown as spheres. The 11 transmembrane helices M0 to M10 are labeled. b–d Top view of OSCA1.1-F516A in non-conducting state 1 (b, brown), non-conducting state 2 (c, pink), and conducting state (d, cyan). e Superimposed top view of the transmembrane layer cross-section of wildtype OSCA1.1 in closed state (purple) and OSCA1.1-F516A in non-conducting state 1 (brown). The 11 transmembrane helices M0 to M10 are labeled and linked by a dashed curve in color following the same scheme used in transmembrane helices. The interface lipid is shown as a wavy line. The transmembrane helices exhibit a counterclockwise rotation from closed state to non-conducting state 1. M3 and M10 helices in closed state and non-conducting state 1 are linked by black lines and red lines, respectively. f Superimposed top view of the transmembrane layer cross-section of OSCA1.1-F516A in non-conducting state 1 (brown) and OSCA1.1-F516A in non-conducting state 2 (pink). M3 and M10 helices in non-conducting state 1 and non-conducting state 2 are linked by black lines and red lines, respectively. g Superimposed top view of the transmembrane layer cross-section of OSCA1.1-F516A in non-conducting state 2 (pink) and OSCA1.1-F516A in conducting state (blue). M3 and M10 helices in non-conducting state 2 and conducting state are linked by black lines and red lines, respectively. The interface lipids tended to dissociate from the central cavity in the OSCA1.1-F516A conducting state. h Superimposed top view of the transmembrane layer cross-section of wildtype OSCA1.1 in closed state (purple) and OSCA1.1-F516A in conducting state (blue). M3 and M10 helices in the closed state and conducting state are linked by black lines and red lines, respectively.
Extracellular pore dilation resembles flower blooming
The movement of M0-M10 towards the central dimer interface complicates the analysis of the single monomer pore in the dimeric OSCA1.1 channel. (Supplementary Fig. 12a–d). To simplify the analysis, we aligned every single monomer of the closed, non-conducting state 1, non-conducting state 2, and conducting state. Then, we examined the conformational changes of the monomer pore from the closed to conducting state (Supplementary Fig. 12e–h). (Supplementary Fig. 13). The results indicate that M0 experiences a 6-degree sequential rotation towards the dimer interface from the closed to conducting state. Additionally, the upper segment of M3 (M3a) kinks sequentially by approximately 75 degrees around the fulcrum of F389/P393. In coordination with the M3a kink, the upper section of M4 (M4a) sequentially kinks around 34 degrees around the fulcrum of K436/P443. Furthermore, the upper segments of M5 (M5a) and M6 (M6a) kink sequentially by ~79-degree around the fulcrum of the linker from G480 to A487 and 46-degree around the fulcrum of P511, respectively (Fig. 4). The process of M0, M3a, M4a, M5a, and M6a sequential rotations or kinks are referred to as ‘blooming-like open’, resembling a flower blooming (Supplementary Movie 1–2). It is worth noting that the location where kinking occurs is consistently at the flexible residues of proline or glycine, such as P393, P443, G480, and P511 (Supplementary Fig. 13), further suggesting that the channel OSCA1.1 in the pore region may undergo plastic deformation when exposed to high-threshold mechanical force. Previously, we have observed that the pore lipids have the potential to seal the pores, which is particularly evident in the human TMEM63A channel16. We conducted a detailed investigation of the pore lipids-like molecule in the non-conducting state 1, non-conducting state 2, and conducting state, and observed that the pore lipids appear to distance themselves from the pore regions upon activation (Supplementary Fig. 10a–d), indicating that the lipids may translocate during channel opening.
a Superimposed side view of M0 (rotate around 3°), M3a (kink around 21°), M4a (kink around 23°), M5a (kink around 75°) and M6a (kink around 33°) deformation from wildtype OSCA1.1 in closed state (purple) to OSCA1.1-F516A in non-conducting state 1 (orange). The protomer is shown as a gray surface. b Superimposed side view of M0 (rotate around 2°), M3a (kink around 68°), M4a (kink around 28°), M5a (kink around 9°) and M6a (kink around 5°) deformation from OSCA1.1-F516A in non-conducting state 1 (orange) to OSCA1.1-F516A in non-conducting state 2 (pink). The view is the same as in a, except the view of M4a, getting rotated to show the bend angle clearly. c Superimposed side view of M0 (rotate around 1°), M3a (kink around 3°), M4a (kink around 10°), M5a (kink around 34°) and M6a (kink around 20°) deformation from OSCA1.1-F516A in non-conducting state 2 (pink) to OSCA1.1-F516A in conducting state (cyan). d Superimposed side view of M0 (rotate around 6°), M3a (kink around 75°), M4a (kink around 34°), M5a (kink around 79°) and M6a (kink around 46°) deformation from wildtype OSCA1.1 in closed state (purple) to OSCA1.1-F516A in conducting state (cyan).
OSCA1.1-F516A is non-conducting and can be directly activated by lyso-PC in native membrane
It is important to note that the extracellular pore open states of OSCA1.1-F516A are obtained in the presence of detergent, which may have altered the interactions between the MS channel protein and the surrounding lipids. To test the channel activity of OSCA1.1-F516A in the native membrane environment, we performed the conventional pressure clamp. Due to the low expression level of OSCA1.1-F516A, the mechanically activated microscopic currents of OSCA1.1-F516A are lower than those of wild-type OSCA1.1 (Fig. 5a, b). Interestingly, the half-activation pressure of OSCA1.1-F516A is −124 ± 3 mmHg, which is slightly lower than wildtype OSCA1.1 of −136 ± 6 mmHg in HEK293-P1KO cells under the same pipette condition (Fig. 5c). In addition, the single-channel conductance of mechanically activated currents for both the wildtype OSCA1.1 and OSCA1.1-F516A is −10 ± 1 pA at a holding potential of −60 mV (Fig. 5d-e). Thus, the OSCA1.1-F516A exhibits a functional mechanosensitive ion channel and shows slight gain-of-function (GOF) in the native membrane. Since the perturbation of MS channel protein-lipid interactions by detergent may be too harsh to extract the protein directly from the native membrane, we may not be able to record the channel activity by the patch clamp in the detergent environment. Previously, we showed that lyso-phosphocholine (lyso-PC) can enhance the mechanosensitivity of the wildtype OSCA1.1 channel, even though it cannot directly open the OSCA1.1’s pore without pressure5. Lyso-PC may mildly interfere with the MS channel protein and lipid interaction and lead to unexpected effects on OSCA1.1-F516A. Surprisingly, when 1 mM lyso-PC was applied to the OSCA1.1-F516A transfected cell, it resulted in a unique spontaneous lyso-PC activated current, without requiring any membrane pressure. Conversely, the HEK293-P1KO cells transfected with the empty vector and wildtype OSCA1.1 were unable to achieve a significant channel current without negative pressure (Fig. 5f, g). Interestingly, the lyso-PC-activated single-channel conductance of the OSCA1.1-F516A was found to be significantly smaller than that of the wildtype OSCA1.1 channel, indicating that while lyso-PC directly activates OSCA1.1-F516A channel, the activation state is different from the full activation state by high-threshold mechanical force.
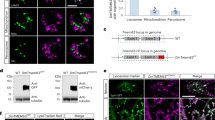
a, b Representative negative pressure activated currents from the Piezo1 knock out HEK293T cell expressing full-length wild-type OSCA1.1 (a) or OSCA1.1-F516A (b) at the holding potential of −60 mV in the cell-attached mode. Negative pressure was applied from −50 to −200 mmHg with −25 mmHg per step and shown below the current traces. c Normalized currents from a and b were fitted with the Boltzmann equation. The half-activation pressure is −124 ± 3 mmHg for OSCA1.1-F516A (red, square), and −136 ± 6 mmHg for wildtype OSCA1.1 (black, circle) in HEK293-P1KO cells under the same pipette condition of 14 MΩ. (n = 3 independent cells, data are present as mean ± SEM). d, e Negative pressure (−60 mmHg) activated single-channel currents of wildtype (d) and OSCA1.1-F516A mutant (e) at the holding potential of −60 mV in the cell-attached mode. The mechanically activated single-channel current of wildtype OSCA1.1 and OSCA1.1-F516A is ~−6 pA at a holding potential of −60 mV in HEK293-P1KO cells. f Top trace represents the 1 mM lyso-PC activated spontaneous OSCA1.1-F516A mutant current at the holding potential of −60 mV in the cell-attached mode. The two bottom traces stand for the empty vector and wildtype OSCA1.1 transfected HEK293-P1KO cell treated with the 1 mM lyso-PC, no significant single-channel current can be recorded. g, Enlargement of the red box shows lyso-PC activated single-channel OSCA1.1-F516A mutant currents. The lyso-PC-activated single-channel current of OSCA1.1-F516A is ~−6 pA at a holding potential of −60 mV in HEK293-P1KO cells.
Lyso-PC activates OSCA1.1-F516A through the upward movement of M6b and opening the intracellular pore
Motivated by the electrophysiological results that OSCA1.1-F516A is non-conducting in the native membrane, we embedded the OSCA1.1-F516A protein in the lipid nanodiscs and obtained only a 2.5 Å cryo-EM density map, indicating that OSCA1.1-F516A is stabilized in a single state in the lipid environment (Supplementary Fig. 14–17). OSCA1.1-F516A in lipid nanodiscs has a transmembrane thickness of about 57 Å, which is close to the 58 Å of wild-type OSCA1.1 in lipid nanodiscs. Meanwhile, the OSCA1.1-F516A in lipid nanodiscs presents more contracted than wild-type OSCA1.1 in lipid nanodiscs. Furthermore, the extracellular pore diameter is also slightly dilated compared to the closed state of OSCA1.1 (Supplementary Fig. 18a, b), indicating that the structure of OSCA1.1-F516A in lipid nanodiscs is non-conducting but has a tendency to be activated, which is consistent with the electrophysiological study of the sightly GOF phenotype. Since lyso-PC can directly activate OSCA1.1-F516A, we incubated 1 mM lyso-PC with the OSCA1.1-F516A nanodiscs and obtained a 2.6 Å cryo-EM map of OSCA1.1-F516A in lyso-PC-containing nanodiscs (Fig. 6a, b). Surprisingly, when compared to the lyso-PC-free structure of OSCA1.1-F516A, we observed a significant conformational change in the upward movement of about 54 degrees of M6b around the hinge of the broken helix on the intracellular side (Fig. 6f–h) and slightly conformational changes of the extracellular pore and central cavity (Supplementary Fig. 18c, d). These changes in the opening of the intracellular pore and slight reshaping of the extracellular pore (Supplementary Movie 3–4) are consistent with the fact that the single-channel conductance active with lyso-PC is smaller than the mechanically activated one (Fig. 5e–g). While OSCA/TMEM63 contains 11 transmembrane helices, in contrast to the 10 transmembrane helices typically found in the TMEM16 family, the additional M0 helix at the lateral side of the transmembrane acts as an auxiliary subunit like the TMIE auxiliary for the TMC complex14. Interestingly, we did not observe the additional M0 helix density, as well as the M0 and M1 linker, in the lyso-PC-containing nanodiscs of activated OSCA1.1-F516A, suggesting that the M0, as well as the M0 and M1 linker, may collapse under the lyso-PC activated state and play important roles in maintaining the intracellular pore closed state (Fig. 6c–e).
a, b Cryo-EM density map of OSCA1.1-F516A in lipid nanodiscs with (b, yellow) or without lyso-PC (a, green) from the side view. The approximate extent of the phospholipid bilayer is shown as a thin black line. The length of the protein and the height of the transmembrane helices are labeled. c–e Side view model of OSCA1.1-F516A in lipid nanodiscs with (d, yellow) or without (c, yellow) lyso-PC and their superimposed presentation (e). The M0 is marked as red and collapsed in the lyso-PC-containing structure. f–h The M4 and M6 of OSCA1.1-F516A in lipid nanodiscs with (g, yellow) or without (f, yellow) lyso-PC. The lyso-PC induces a slight motion of the extracellular part of M4 and M6 as well as an upward rotation/movement of M6b of about 57 degrees. The critical residues for the conformational changes of M4 and M6 are labeled.
Both extracellular and intracellular pore openings are involved in full activation by mechanical force
To further investigate whether the opening of the extracellular pore during OSCA1.1 channel activation by mechanical force also undergoes a blooming-like opening, we mutated the flexible fulcrum residues (P393, P443, G480, and P511) at the kink sites to alanine. These mutations may decrease the flexibility of the fulcrum and increase the gating energy barrier. We then performed the pressure clamp to test their mechanosensitivity. While P393A at M3 and P511A at M6 slightly reduce channel mechanosensitivity to the half-activation pressures of −174 ± 5 mmHg and −152 ± 10 mmHg, respectively, P443A at M4 and G480A at M5 largely lead to a loss-of-function phenotype as both of their half-activation pressures are beyond −200 mmHg (Fig. 7a, b). The activated state of OSCA1.1-F516A by lyso-PC suggested that when M6b moves to the extracellular side, E532 and K537 form a potential side-chain-side-chain interaction at the hinge of the broken helix. To test whether the opening of the intracellular pore is also involved in full activation of the OSCA1.1 channel by mechanical force, we mutated the potential side-chain-side-chain interaction residues E532, K537, and the flexible hinge P538 to alanine to disrupt the interaction and reduce hinge flexibility, respectively. We found that E532A and P538A displayed the reduced mechanosensitivity phenotype while K537A showed the increased mechanosensitivity phenotype (Fig. 7c, d). We reasoned that while K537A may disrupt the potential interaction between K537 and E532 in the M6b upward state, it may also disrupt the interaction between the potential lipid and K537, as predicted in OSCA1.225. Taken together, these functional results indicate that the high-threshold mechanical force applied to the lipid membrane may also result in the opening of both the extracellular pore-like flow blooming and the intracellular pore by M6b upward movement, which then fully activates the channel (Fig. 8).
a The flexible kink residues at M3a, M4a, M5a and M6a are shown as spheres. b Normalized negative pressure activated currents were fitted with the Boltzmann equation. The half-activation pressure is −136 ± 6 mmHg for wildtype OSCA1.1 (black, circle), −174 ± 5 mmHg for OSCA1.1-P393A (red, square), −152 ± 10 mmHg for OSCA1.1-P511A (blue, rhombic), and beyond −200 mmHg for both OSCA1.1-P443A (pink, square) and OSCA1.1-G480A (green, triangle) in HEK293-P1KO cells under the same pipette condition of 14 MΩ. (n = 3 independent cells, data are present as mean ± SEM). c The flexible hinge residues at M6b are shown as spheres. d Normalized negative pressure activated currents were fitted with the Boltzmann equation. The half-activation pressure is −136 ± 6 mmHg for wildtype OSCA1.1 (black, circle), −151 ± 3 mmHg for OSCA1.1-E532A (red, square), −104 ± 4 mmHg for OSCA1.1-K537A (blue, rhombic), and −145 ± 5 mmHg for OSCA1.1-P538A (pink, square) in HEK293-P1KO cells under the same pipette condition of 14 MΩ. (n = 3 independent cells, data are present as mean ± SEM).
a, b The full activation model of the OSCA channel is shown in the cartoon. Force or high-osmolality shock may cause thinning of the local membrane, which may induce transmembrane helices deformation, and then the two OSCA protomers draw close. The interface lipid dissociated from the central cavity, and pore lipids are dislocated to the local membrane (a). Consequently, each monomeric pore undergoes a blooming-like opening of the extracellular pore and opening of the intracellular pore by upward movement of M6b (b).
Discussion
The inherent MS channels embedded in the lipid membrane respond directly to physical mechanical force without changing their chemical composition. Physical mechanical force can easily plasticize the lipids, resulting in a change in membrane curvature or thickness. The interaction between the lipids and the MS channel protein may play the dominant role in sensing and transducing mechanical forces. Consequently, the mechanical force can alter the lipid and MS channel protein interaction, causing the conformational changes of the MS channel, breaking the energy barrier of the restriction gates, and then finally allowing the ion to flow through the open pore. Although it is difficult to directly visualize the mechanical force, especially the higher mechanical force, activates the MS channel at the atomic level. Some alternatives, such as directly mutating the MS channel protein or changing the lipid environment, can be used to lock its conformation from resting to activating states. In the OSCA1.1 channel, the phenyl group of F516 acts as one of the main restriction gates blocking the pore. The OSCA1.1 F516A not only removes the blocking effect of the phenyl group, but also causes the conformational changes of the extracellular pore region in the native environment and presents the relative GOF phenotype. Our results indicate that the detergent or lyso-PC could directly perturb the lipid and MS channel protein interaction, resulting in channel conformational changes, and thus OSCA1.1-F516A exhibits the pore dilated state both in the detergent environment and in lyso-PC-containing lipid nanodiscs. Interestingly, while the detergent results in dilating the extracellular pore, the lyso-PC mainly acts on M6, inducing M6b upward movement around the hinge of the broken helix, which dilates the intracellular pore. Therefore, it suggested that OSCA/TMEM63 needs to open both the intracellular pore and the extracellular pore to reach the fully activated state by the mechanical force.
Although MS channels exist in both prokaryotes and eukaryotes, they exhibit different mechanical force-responsive thresholds and structural variations. In trimeric PIEZO channels with low mechanical force thresholds, changes in membrane curvature can shift the channel to a flattened and activated state22,26,27. While in pseudo-tetrameric MS K2P and bacterial heptameric MscS channels, which possess higher thresholds of mechanical force compared to PIEZO channels, the dominant role in channel mechanosensitivity is played by pore lipids and some gating-regulated lipids located on the cytosolic side20,21. Despite the unique dimeric transporter-like architecture, our results suggest that activation of OSCA/TMEM63 channels may also involve the removal of energy barriers of the dimer interface and pore lipids. The three OSCA1.1-F516A structures in the detergent environment show thinner membrane thicknesses ranging from 49 Å to 53 Å. Among the MscS, MS K2P, and PIEZO channels, the MscS channels show significant membrane thickness changes during activation. A membrane that is too thin can cause the MscS channel to enter the non-conducting desensitized state21. In this case, OSCA/TMEM63 channels share some similar mechanisms to MscS channels.
Lyso-PC can activate most MS channels, including MscL28, MscS29, and MS K2P channels30, but there is no example of a lyso-PC-activated structure. In this study, we show the lyso-PC-activated OSCA1.1-F516A structure, suggesting that lyso-PC activates the MS channel through allosteric modulation. Interestingly, in OSCA1.1-F516A, lyso-PC causes an upward movement of M6b to the extracellular side, which is reminiscent of the calcium-induced rotation of the down segment of M6 in the TMEM16 family12. While OSCA/TMEM63 channels show structural similarity to TMEM16 and TMC families, OSCA/TMEM63 channels have an additional TM helix M0 than TMEM16 and TMC families. In the TMC complex, the TMIE subunit acts as an auxiliary for TMC, the TMIE-TMC complex can align with the full-length OSCA/TMEM6314. More interestingly, the less-PC-activated OSCA1.1-F516A structure shows that the additional M0 collapses during activation. This raises the interesting question that M0 may act as a pseudo auxiliary for the rest of OSCA/TMEM63, and the role of TMIE in the TMC complex needs to be further investigated.
Activation of OSCA/TMEM63 channels also undergoes the extracellular monomer pore blooming-like open, including the rotation of M0 and kink of M3, M4, M5, and M6. In one of the TMEM16 members mTMEM16F, which is a calcium-activated lipid scramblase or ion channel, the F518H mutant, located in the M4, presents a similar kink of M3 and M431. This similarity offers additional insight into the evolutionary divergence between the TMEM16 family and the OSCA/TMEM63 family. Upon activation of the dimeric OSCA/TMEM63 channel, each monomer sequentially contracts to the dimer interface. This motion is akin to that of the Prestin protein32,33, which senses voltage and is responsible for electromotility in the auditory system. Additionally, the TMC complex also reveals the extended or contracted motion at the dimer interface14. Based on our results, we suspected that the contraction movement at the dimer interface may be related to the activation tendency. Interestingly, OSCA/TMEM63 channels in mammals display a higher threshold for mechanical force in comparison to OSCA1.116,17. The lack of a dimer interface and monomer formation in mammalian OSCA/TMEM63 raises the question of how such high mechanical force affects its conformation, extends the pore, and ultimately allows for ion flow.
Methods
Cell lines and constructs
Sf9 cells were obtained from Thermo Fisher Scientific and cultivated in Sf-900 II SFM medium (Gibco) at a temperature of 27 °C. Expi293F suspension cells, also acquired from Thermo Fisher Scientific, were grown in HEK293 medium (SinoBiological) under 37 °C with 6% CO2. The HEK293T-P1KO cells were grown in DMEM medium supplemented with 10% FBS under 37 °C with 5% CO2 and 70% humidity. The cells were regularly screened for mycoplasma contamination and yielded negative results. The DNA fragments encoding OSCA1.1 (UniProtKB Q9XEA1) and its mutants (Q398E, T515E, and F516A) were cloned into a modified BacMam expression vector using EcoRI and XhoI restriction sites as described previously5. The DNA fragment is placed before a PreScission protease (Ppase) cleavage site, followed by a C-terminal enhanced green fluorescent protein and FLAG tag.
Protein expression and membrane pellet preparation
The baculovirus was produced using the DH10Bac bacterial strain and sf9 cells, conforming to the standard Bac-to-Bac protocol. For large-scale protein expression, Expi293F cells in suspension, grown in SinoBiological HEK293 medium at 37 °C, were infected with the baculovirus once reaching a density of 2.0 × 106 cells/mL. Following infection, 10 mM sodium butyrate was introduced 12 hours post-infection, and the temperature was reduced to 30 °C to facilitate protein expression. Cells were collected 72 hours after infection through centrifugation at 2800 × g and 4 °C for 10 minutes. The cells were then broken using sonication in cold lysis buffer containing 20 mM Tris-HCl at pH 7.4 and 150 mM NaCl. Unbroken cells and cell debris were removed by centrifugation at 5600 × g and 4 °C for 10 minutes. The supernatant was then centrifuged again, at 270,000 × g for 30 minutes in a 45Ti rotor (Beckman). Subsequently, membrane pellets were collected, frozen at −80 °C, and stored until further use.
Protein purification and EM sample preparation of detergent forms
The membrane pellets were homogenized in TBS, and then solubilized in TBS, 1% (w/v) LMNG, and 0.1% (w/v) CHS for 1 hour at 4 °C. Insoluble materials were removed through centrifugation at 270,000 × g for 30 minutes in a 45Ti rotor. The resulting supernatant was applied onto anti-FLAG resin (Genscript) by gravity flow. The resin was then washed with 10 column volumes of wash buffer (lysis buffer with 0.02% (w/v) LMNG). Protein was subsequently eluted with supplemented elution buffer (wash buffer including 230 μg/mL FLAG peptide). The recovered protein’s C-terminal GFP tag was removed via HRV3C protease cleavage at 4 °C for one hour. The protein was concentrated using a 100-kDa cutoff concentrator (Millipore) and subsequently loaded onto a Superose 6 Increase 10/300 column (GE Healthcare) in lysis buffer containing 0.01% digitonin. The peak fractions were merged and concentrated to ~10 mg/mL for cryo-EM sample preparation. The protein samples were cleared through centrifugation at 270,000 × g for 30 minutes at 4 °C before grid preparation. Holey Carbon grids (R1.2/1.3) with a mesh size of 300 and a gold thickness of 10–12 nm (Quantifoil) were glow discharged (25 s, 15 mA; Pelco easiGlow, TedPella) prior to use. A 3-μL droplet of protein solution was applied onto the grids, which were rapidly frozen by immersing them in liquid ethane cooled by nitrogen. The FEI Vitrobot Mark IV (Thermo Fisher Scientific) was set at 8 °C, 100% humidity, 3 blot force, ~5 s wait time, and 5 s blot time for cryo-embedding purposes.
Reconstitution of OSCA1.1-F516A into lipid nanodiscs and EM sample preparation of the lipid nanodiscs forms
Following the previous method16, OSCA1.1-F516A was purified in a 1% DDM environment. Lecithin solubilized in chloroform was dried under nitrogen gas and resuspended with 0.7% (w/v) DDM. OSCA1.1-F516A, MSPE3D1, and lipid mixture were mixed at a molar ratio of 1:10:500 and incubated at 4 °C for 3 h. Detergents were removed by incubation with Bio-beads SM2 (Bio-Rad) overnight at 4 °C. The protein-lipid mixture was loaded onto anti-FLAG G1 affinity resin equilibrated with TBS. The resin was further washed with 10 column volumes of TBS, and protein was eluted with an elution buffer (TBS plus 230 μg/ml FLAG peptide). The C-terminal GFP tag of eluted protein was removed by HRV3C protease cleavage for 3 h at 4 °C. The protein was incubated with TBS, further concentrated by a 100-kDa cutoff concentrator (Millipore), and loaded onto a Superose 6 increase 10/300 column (GE Healthcare) running in TBS. Peak fractions were combined and concentrated to around 2 mg/ml of OSCA1.1-F516A for cryo-EM sample preparation. For the preparation of the lyso-PC containing OSAC1.1-F516A lipid nanodiscs EM sample, the final concentration of 5 mM lyso-PC was incubated with OSAC1.1-F516A lipid nanodiscs for 3 hours before application to the grids. The rest of the procedure was the same as the conventional EM sample preparation.
Cryo-EM data acquisition, processing, and model building
Cryo-EM data were captured using a Titan Krios microscope (FEI) outfitted with a Cs-corrector and operated at 300 kV. Image stacks in super-resolution mode were automated through the EPU program operating on a Thermo Fisher Falcon4i detector. The object plane’s pixel size was 0.57 or 0.35 Å, whereas defocus ranged from −0.9 μm to −1.2 μm. In this study, a GIF Quantum energy filter with a 5 eV slit width was also employed. The total exposure dose was 40 e- Å−2. Data processing was conducted using the cryoSPARC v4 suite. The super-resolution image stacks were gain-normalized, binned with Fourier cropping, and patch-based Contrast Transfer Function parameters of the dose-weighted micrographs (0.57 or 0.35 Å per pixel) were determined by cryoSPARC and summarized in Extended Figures. The monomers’ atomic models were constructed in Coot based on an initial model (PDB: 8GRN). The models were then manually adjusted in Coot. Dimeric models were obtained through the application of a symmetry operation on the monomer. These models were refined using Phenix.real_space_refine iteratively with secondary structure restraints and Coot. FSC curves were calculated between the EM maps and final models for validation. Coot, PyMOL, and Chimera were used for figure preparation.
Electrophysiology
HEK293T-P1KO cells were transfected with AtOSCA1.1-CGFP or its mutations and were incubated for 24 to 36 hours before recording. Following the previous method, cell-attached pressure patches were performed5. At a sampling rate of 10 kHz and a filter at 2 kHz (Digidata 1440 A, Molecular Devices), the recorded currents were at −60 mV using an Axopatch 700B amplifier. Borosilicate micropipettes (OD 1.5 mm, ID 0.86 mm, Sutter) were pulled and fire-polished to 10–15 MΩ resistance to homogenize the pipette geometry. Negative pressure was applied to patch pipettes using a High-Speed Pressure Clamp-1 from ALA Scientific Instruments, Farmingdale, NY, USA. The pressure applied was recorded in millimeters of mercury (mmHg) using a piezoelectric pressure transducer from WPI, Sarasota, FL, USA. The recordings took place at room temperature (22 °C), and the data were analyzed using pClamp10.7 software. All data were acquired from at least three independent cells.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-39401 (non-conducting state 1 of OSCA1.1-F516A); EMD-39400 (non-conducting state 2 of OSCA1.1-F516A); EMD-39399 (conducting state of OSCA1.1-F516A); EMD-39403 (OSCA1.1-F516A in lipid nanodisc); and EMD-39402 (OSCA1.1-F516A in lipid nanodisc with lyso-PC). The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 8YMO (non-conducting state 1 of OSCA1.1-F516A), 8YMN (non-conducting state 2 of OSCA1.1-F516A), 8YMM (conducting state of OSCA1.1-F516A), 8YMQ (OSCA1.1-F516A in lipid nanodisc) and 8YMP (OSCA1.1-F516A in lipid nanodisc with lyso-PC). Source data are provided with this paper.
References
Kefauver, J. M., Ward, A. B. & Patapoutian, A. Discoveries in structure and physiology of mechanically activated ion channels. Nature 587, 567–576 (2020).
Sukharev, S. I., Blount, P., Martinac, B., Blattner, F. R. & Kung, C. A large-conductance mechanosensitive channel in E. coli encoded by mscL alone. Nature 368, 265–268 (1994).
Maingret, F., Fosset, M., Lesage, F., Lazdunski, M. & Honore, E. TRAAK is a mammalian neuronal mechano-gated K+ channel. J. Biol. Chem. 274, 1381–1387 (1999).
Coste, B. et al. Piezo1 and Piezo2 are essential components of distinct mechanically activated cation channels. Science 330, 55–60 (2010).
Zhang, M. et al. Structure of the mechanosensitive OSCA channels. Nat. Struct. Mol. Biol. 25, 850–858 (2018).
Murthy, S. E. et al. OSCA/TMEM63 are an evolutionarily conserved family of mechanically activated ion channels. Elife 7, e41844 (2018).
Yuan, F. et al. OSCA1 mediates osmotic-stress-evoked Ca2+ increases vital for osmosensing in arabidopsis. Nature 514, 367–371 (2014).
Li, Q. & Montell, C. Mechanism for food texture preference based on grittiness. Curr. Biol. 31, 1850–1861 e1856 (2021).
Du, H. et al. The cation channel TMEM63B is an osmosensor required for hearing. Cell Rep. 31, 107596 (2020).
Yan, H. et al. Heterozygous variants in the mechanosensitive ion channel TMEM63A result in transient hypomyelination during infancy. Am. J. Hum. Genet. 105, 996–1004 (2019).
Dang, S. et al. Cryo-EM structures of the TMEM16A calcium-activated chloride channel. Nature 552, 426–429 (2017).
Paulino, C., Kalienkova, V., Lam, A. K. M., Neldner, Y. & Dutzler, R. Activation mechanism of the calcium-activated chloride channel TMEM16A revealed by cryo-EM. Nature 552, 421–425 (2017).
Alvadia, C. et al. Cryo-EM structures and functional characterization of the murine lipid scramblase TMEM16F. Elife 8, e44365 (2019).
Jeong, H. et al. Structures of the TMC-1 complex illuminate mechanosensory transduction. Nature 610, 796–803 (2022).
Jentsch, T. J. & Pusch, M. CLC chloride channels and transporters: structure, function, physiology, and disease. Physiol. Rev. 98, 1493–1590 (2018).
Zhang, M., Shan, Y., Cox, C. D. & Pei, D. A mechanical-coupling mechanism in OSCA/TMEM63 channel mechanosensitivity. Nat. Commun. 14, 3943 (2023).
Zheng, W. et al. TMEM63 proteins function as monomeric high-threshold mechanosensitive ion channels. Neuron 111, 3195–3210.e7 (2023).
Martinac, B., Adler, J. & Kung, C. Mechanosensitive ion channels of E. coli activated by amphipaths. Nature 348, 261–263 (1990).
Anishkin, A., Loukin, S. H., Teng, J. & Kung, C. Feeling the hidden mechanical forces in lipid bilayer is an original sense. Proc. Natl Acad. Sci. USA 111, 7898–7905 (2014).
Brohawn, S. G., Campbell, E. B. & MacKinnon, R. Physical mechanism for gating and mechanosensitivity of the human TRAAK K+ channel. Nature 516, 126–130 (2014).
Zhang, Y. et al. Visualization of the mechanosensitive ion channel MscS under membrane tension. Nature 590, 509–514 (2021).
Yang, X. et al. Structure deformation and curvature sensing of PIEZO1 in lipid membranes. Nature 604, 377–383 (2022).
Schmidpeter, P. A. M. et al. Membrane phospholipids control gating of the mechanosensitive potassium leak channel TREK1. Nat. Commun. 14, 1077 (2023).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph 14, 354–360 (1996). 376.
Jojoa-Cruz, S. et al. Cryo-EM structure of the mechanically activated ion channel OSCA1.2. Elife 7, e41845 (2018).
Mulhall, E. M. et al. Direct observation of the conformational states of PIEZO1. Nature 620, 1117–1125 (2023).
Lin, Y. C. et al. Force-induced conformational changes in PIEZO1. Nature 573, 230 (2019).
Perozo, E., Kloda, A., Cortes, D. M. & Martinac, B. Physical principles underlying the transduction of bilayer deformation forces during mechanosensitive channel gating. Nat. Struct. Biol. 9, 696–703 (2002).
Nomura, T. et al. Differential effects of lipids and lyso-lipids on the mechanosensitivity of the mechanosensitive channels MscL and MscS. Proc. Natl Acad. Sci. USA 109, 8770–8775 (2012).
Maingret, F., Patel, A. J., Lesage, F., Lazdunski, M. & Honore, E. Lysophospholipids open the two-pore domain mechano-gated K(+) channels TREK-1 and TRAAK. J. Biol. Chem. 275, 10128–10133 (2000).
Arndt, M. et al. Structural basis for the activation of the lipid scramblase TMEM16F. Nat. Commun. 13, 6692 (2022).
Ge, J. et al. Molecular mechanism of prestin electromotive signal amplification. Cell 184, 4669–4679 e4613 (2021).
Bavi, N. et al. The conformational cycle of prestin underlies outer-hair cell electromotility. Nature 600, 553–558 (2021).
Acknowledgements
We would like to thank the Cryo-EM Facility of Westlake University for providing cryo-EM and the High-Performance Computing Center of Westlake University for computation support. This work was supported by National Natural Science Foundation of China (92068201) and Key R&D Program of Zhejiang (2024SSYS0031).
Author information
Authors and Affiliations
Contributions
M.F.Z. conceived the project and designed the experiments. Y.Y.S. and X.Y.G. prepared the constructs and purified the proteins. Y.Y.S., M.M.Z., and Y.L. prepared the cryo-EM sample and collected cryo-EM data. M.Y.C. performed the electrophysiological study. M.F.Z. performed image processing, built the model, analyzed data, and wrote the manuscript draft. D.Q.P. and M.F.Z. supervised the project. All authors contributed to the manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Qiong-Yao Tang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shan, Y., Zhang, M., Chen, M. et al. Activation mechanisms of dimeric mechanosensitive OSCA/TMEM63 channels. Nat Commun 15, 7504 (2024). https://doi.org/10.1038/s41467-024-51800-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-51800-0
This article is cited by
-
The double-edged sword of ion channel-targeting pesticides: a paradigm shift from health risks to precision intervention
Archives of Toxicology (2026)
-
Transmembrane protein TMEM98 as a multifunctional regulator in cancer: from signaling pathways to translational implications
Journal of Translational Medicine (2025)