Abstract
The spatial co-presence of aberrant long non-coding RNAs (lncRNAs) and abnormal coding genes contributes to malignancy development in various tumors. However, precise coordinated mechanisms underlying this phenomenon in tumorigenesis remains incompletely understood. Here, we show that Prohibitin 2 (PHB2) orchestrates the transcription of an oncogenic CASC15-New-Isoform 2 (CANT2) lncRNA and the coding tumor-suppressor gene CCBE1, thereby accelerating melanoma tumorigenesis. In melanoma cells, PHB2 initially accesses the open chromatin sites at the CANT2 promoter, recruiting MLL2 to augment H3K4 trimethylation and activate CANT2 transcription. Intriguingly, PHB2 further binds the activated CANT2 transcript, targeting the promoter of the tumor-suppressor gene CCBE1. This interaction recruits histone deacetylase HDAC1 to decrease H3K27 acetylation at the CCBE1 promoter and inhibit its transcription, significantly promoting tumor cell growth and metastasis both in vitro and in vivo. Our study elucidates a PHB2-mediated mechanism that orchestrates the aberrant transcription of lncRNAs and coding genes, providing an intriguing epigenetic regulatory model in tumorigenesis.
Similar content being viewed by others
Introduction
Tumor cell heterogeneity results in the coexistence of aberrant lncRNAs and abnormal coding genes within the same or different genomic loci, giving rise to diverse malignancies with various pathogenic causes. For instance, in the human genome, abnormal coding genes MYC is generated from the chr8q24 locus, a well-known genomic region implicated in the progression of various malignancies, and lncRNAs generated from this locus are recognized as contributing factors in the tumorigenesis of difference cancer types1,2,3,4,5,6. However, many critical lncRNAs and coding genes originating from distinct loci coordinately result in tumorigenesis through unique mechanisms. For example, abnormal p53 has been reported to lead to genetic instability and uncontrolled cell proliferation in gastric cancer7. Meanwhile, lncRNA GCLET increased FOXP2 expression and remarkably impacted gastric cancer phenotypes8. Additionally, our previous studies have identified that the coding gene NTS and the lncRNA ROR, derived from different loci, exerted distinct influences on the progression of uveal melanoma9,10. Therefore, it is intriguing to explore how these coding genes and lncRNAs from the same or various loci coordinately contribute to the progression of malignancies.
The eukaryotic genome is well organized and evolutionarily conserved, yet exhibits spatial plasticity with cell-specific characteristics11,12,13. Recent studies demonstrate that the intricate transcription of coding genes and lncRNAs are influenced by genetic and epigenetic mechanisms across various dimensions from one to three dimensional levels14,15,16,17,18,19. Plenty of elementary chromatin features, including DNA sequence variations, nucleosome density and position, histone methylation modification and transcription factor binding, serving as the cornerstone of transcription at coding genes and lncRNAs20,21,22,23. The activity of regulatory factors and their functional network could partially explain cell-specific regulatory mechanisms during carcinogenesis24. For instance, the ETS family members of transcription factors have clearly been shown to be a driving event in prostate tumorigenesis via chromosomal translocation events25. Furthermore, the open or closed status of specific genomic loci is closely linked to active and inactive transcription, respectively, and determines the chromatin accessibility of regulatory factors26,27,28. For example, the chromatin regulator BRD8, conjugating with H2AZ, sustained a repressive chromatin state in the p53 locus, preventing the activation of p53 and promoting the development of glioblastoma29. Typically, these multifaceted regulatory mechanisms establish an environment that permits the operation of key regulatory factors in the process of transcription.
Prohibitin-2 (PHB2) and its homologs are widely expressed scaffold proteins involved in numerous signaling pathways that regulate metabolism, mitochondrial autophagy, and cell migration and proliferation, thereby influencing diseases such as cancer and inflammation30,31. As a precursor of melanoma, melanocytes are involved in melanin biosynthesis, which is related to mitochondrial function. Previous studies have shown that the PHB family plays a crucial role in maintaining mitochondrial integrity, participating in melanin production and carcinogenic pathways, and potentially functioning as regulators of melanin production signals32.This suggests the potential of developing new drug candidates for the treatment of melanoma and other types of cancer. Furthermore, PHB2 acts as a transcriptional co-regulatory factor that can bind to histone-modifying enzymes, mediating the transcriptional regulation of tumor-related genes and thus affecting tumorigenesis33,34. Given these facts, it is plausible that unknown shuttle factors like PHB2 could coordinately orchestrate the transcription of coding genes and lncRNAs, thereby influencing tumorigenesis.
In this study, we identify that transcriptional regulator PHB2 serves as a shuttle factor which coordinately orchestrates the transcription of the oncogenic CANT2 lncRNA derived from chromosome 6p22.3 and the coding tumor-suppressor gene CCBE1 derived from chromosome 18q21.32 for accelerating tumorigenesis of melanoma. Our study provides a PHB2-mediating mechanism orchestrating aberrant transcription of lncRNA and coding gene, thus proposing an epigenetic regulatory model of tumorigenesis.
Results
Open chromatin recruits PHB2 to activate the transcription of the CANT2 lncRNA at chr6p22.3 locus
To validate our hypothesis that a shuttle factor could bridge the transcriptional connection between noncoding RNAs and coding genes in melanoma, we focused on the canonical cancer susceptibility locus chr6p22.3. This locus is known to be the origin of various noncoding transcripts, including the CASC15 lncRNA, which is implicated in the metastasis melanoma cells35,36. We initially performed rapid amplification of cDNA ends (RACE) assay at this locus to elucidate the precise transcripts present in A375 and A875 melanoma cells. A lncRNA with 1939 bp spanning 9 exons was identified at the CASC15 locus in melanoma cells (Fig. S1A, B). Specifically, exons 2–5 and 8 were consistent with the predicted exons (Fig. 1A, white box). However, exon 1 exhibited an additional 72-bp fragment at the 5’ terminus, while exon 6–7 and 9 were unique (Fig. 1A, blue box). To further strengthen the validation of this identified isoform, we employed three series of RNA-seq and TT-seq data conducted on A375 melanoma cells, available at the public GEO database (GSE22388737, GSE22388837, and GSE23237538), to examine gene density at the CASC15 locus (Fig. S2A, B). Utilizing these comprehensive datasets as a unified reference, we aligned all nine exons of the isoform with the read peaks, and confirmed that their expressions in A375 cells were consistent with our RACE findings (Fig. S2B). We then confirmed the absence of coding evidence for this transcript with both CPAT39 (Fig. S3A) and CPC2.0 (http://cpc2.gao-lab.org/index.php)40 (Fig. S3B) using the human genome as reference. Collectively, these data showed that this isoform of the CASC15 lncRNA is a non-coding transcript identified in cutaneous melanoma, and we therefore named it CASC15-New-Isoform 2 (CANT2) lncRNA (GenBank number: OR811110).
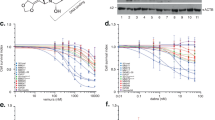
A Genomic structure of CANT2 and a schematic of the core promoter fragment deletion by CRISPR-Cas9. The rectangles indicate the exons of CANT2. The black line indicates introns. The red line indicates the core promoter of CANT2. B Chromatin accessibility status at the chr6p22.3 locus with ATAC-seq and Omni-ATAC datasets obtained from the public GEO database under accession codes GSE18839842, GSE13443243, and GSE24144544. C Real-time PCR of luciferase activity of the DNA fragments of CANT2 promoter. Ctrl, control group without any Firefly reporter plasmids; Mock, group with wild-type Firefly reporter plasmids; 1–5, groups with different 200 bp DNA fragment of CANT2 promoter contained in Firefly reporter plasmids. All groups had Renilla reporter plasmids. All data were calculated the ratio of firefly to Renilla luciferase activity (Fluc/Rluc) in dual luciferase reporter system. For comparison, the ratio of Fluc/Rluc of the mock was arbitrarily set as 1 in the calculation. All the experiments were performed in triplicate and data are presented as the mean ± SD using an unpaired two-tailed t test; *P < 0.05; **P < 0.01. D Schematic diagram of variant primer sets in FAIRE assay. a, b, c: primers in SOX4, CANT2, PRL promoter regions, respectively. E PCR of FAIRE assay at CANT2 core promoter region. Representative images from three independent experiments. F Quantification of FAIRE assay at Chr6p22.3 in melanoma cells (A375 and A875) and normal cells (PIG1). Input DNA was used as a positive control. All the experiments were performed in triplicate and data are presented as the mean ± SD using a Dunnett’s multiple comparisons test; ****P < 0.0001. G Real-time of CANT2 expression at RNA level in melanoma cells lines. All the experiments were performed in triplicate and data are presented as the mean ± SD using an unpaired two-tailed t test; ***P < 0.001; ****P < 0.0001. H Subcellular localization of CANT2 in melanoma cells. GAPDH and U6 RNA served as positive controls for the cytoplasmic (black) and nuclear (orange) fractions, respectively. All the experiments were performed in triplicate. I Survival curves of SKCM patients with a high or low PHB2 expression (cutoff = 0.5, P < 0.05) was analyzed in GEPIA using a Log-rank (Mantel-Cox) test. J, K ChIP analysis of PHB2 (J), H3K4me3 (K) and MLL2 (K) at the CANT2 core promoter. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. All the experiments were performed in triplicate and data are presented as the mean ± SD using a Dunnett’s multiple comparisons test; *P < 0.05, **P < 0.01; ***P < 0.001; ****P < 0.0001. L, M PCR of ChIP analysis of H3K4me3 (L) and MLL2 (M) at the CANT2 core promoter in melanoma and normal cells. Representative blots from three independent experiments. N Co-IP assay was performed to show the interaction between PHB2 and MLL2 in normal and melanoma cells. IgG was used as a negative control. Representative blots from three independent experiments.
Complex mechanisms such as epigenetics may play a crucial role in regulating the transcription of various isoforms at chr6p22.3 locus. Components of epigenetics, including 3D genome organization, chromatin remodeling, and histone modification, typically exert significant influence in this context. With this understanding, we investigated the chromatin topology landscape at chr6p22.3 by analyzing Hi-C data of melanoma with the 3D Genome Browser (accessible at http://3dgenome.fsm.northwestern.edu/view.php)41. However, our analysis of the Hi-C maps did not unveil any significant differences in higher-order chromatin structures such as topologically associating domain (TAD) organization between melanoma samples and normal samples (Figure S4A). Alternatively, we shifted our focus to chromatin accessibility dynamics using ATAC-seq and omni-ATAC datasets (GSE18839842, GSE13443243, and GSE24144544) performed on malignant melanoma cells and patient-derived melanoma cultures (MM lines). Comparative analysis with normal human keratinocyte cells revealed prominent peaks in the promoter region of CANT2 lncRNA, located at chr6:21,885,810–21,886,009, in melanoma cells such as COLO-823 and WM-266-4 (Fig. 1B). Similarly, significant ATAC-seq peaks were observed in the CANT2 promoter region in A375 melanoma cells used in our study (Fig. 1B). Moreover, patient-derived melanoma cultures, such as MM087 and MM099, which has undergone phenotype switching to a dedifferentiated, mesenchymal-like and therapy-resistant cell state43, exhibited higher peaks compared to the melanoma culture MM001 with a melanocyte state (Fig. 1B). To determine how CANT2 lncRNA could alter tumor behavior, the 200-bp core promoter (Fig. 1A, fragment IV) of CANT2 was identified via the dual luciferase reporter assay (Figs. S4B and 1C). We further employed formaldehyde-assisted isolation of regulatory elements (FAIRE) to examine the chromatin accessibility of CANT2 locus in melanoma (Fig. 1D), and found that an open chromatin status was exhibited in melanoma cells (Fig. 1E, lanes 2–3) compared with normal cells (Fig. 1E, lane 1). Similarly, we further confirmed the open chromatin status of CANT2 locus by using FAIRE-qPCR assay (Fig. 1F, middle). Likewise, the adjacent SOX4 and PRL genes remained in a closed chromatin status both in tumor and normal cells (Fig. 1F, left and right). Subsequently, we assessed the expression of CANT2 lncRNA in tumor cells. As anticipated, our findings revealed that CANT2 was highly expressed in both A875 and A375 cells (Fig. 1G). Additionally, we examined the expression of previous reported noncoding RNA transcripts including CANT1, CASC15 and CASC15-S within this locus. Interestingly, we observed that CANT1, CASC15 and CASC15-S were not expressed in melanoma cells used in our study (Figure S5A). Moreover, a cytoplasmic and nuclear RNA isolation assay indicated that CANT2 lncRNA predominantly localized in the nucleus (Fig. 1H). These findings unveiled that CANT2 is a nuclear lncRNA in human cutaneous melanoma cells.
We next aimed to explore the potential epigenetic regulation of CANT2 lncRNA. Intriguingly, our investigation into the gene expression profile of skin cutaneous melanoma (SKCM) revealed that PHB2, a known transcriptional co-regulator capable of binding with histone modification enzymes33,34, was highly expressed in SKCM (Fig. S5B). Additionally, high PHB2 expression correlated with inferior overall survival (Fig. 1I). We also identified higher PHB2 expression in melanoma cells compared to normal control cells (Fig. S5C). Subcellular localization studies revealed that PHB2 was presented in both the cytosol and nucleus of melanoma cells (Fig. S5D). Chromatin immunoprecipitation (ChIP) qPCR showed that PHB2 could bind to this open chromatin region in melanoma cells (Fig. 1J). Given the pivotal role of histone modification in dynamically modulating chromatin, we proceeded to investigate alterations in histone acetylation and methylation patterns at the CANT2 locus. As anticipated in the ChIP assay, all three histone markers H3K4 monomethylation (Fig. S6A), H3K27 acetylation (Fig. S6B) and H3K4 trimethylation (Fig. 1K, orange; Fig. 1L, lanes 2–3) associated with open chromatin exhibited elevated levels at the CANT2 locus in melanoma cells compared with normal PIG1 cells. Remarkably, H3K4me3 level displayed a significant increase at the CANT2 locus (Fig. 1K, orange). Consequently, we delved into the functional factors involved in the H3K4me3 modification. However, the enrichment of common histone methyltransferases, MLL1 (Fig. S6C, panel 1, lanes 2–3; Fig. S6D) and SET1A (Fig. S6C, panel 2, lanes 2–3; Fig. S6E), showed no discernible difference between melanoma cells and normal cells. It was surprising to observe the specific binding of MLL2 to the CANT2 locus in melanoma (Fig. 1K, green; Fig. 1M, lanes 2–3). Moreover, co-immunoprecipitation (co-IP) assays showed MLL2 could be pulled out by baiting PHB2 in melanoma (Fig. 1N, panels 2–3, lane 3). Taken together, these data showed that PHB2 may serve as a transcriptional regulator for recruiting MLL2 to the accessible promoter of a lncRNA CANT2, and triggering the expression of CANT2 by increased H3K4 trimethylation in melanoma.
The deletion of CANT2 diminishes tumor cell proliferation and migration in vitro
To address the role of CANT2 lncRNA in tumorigenesis, we deleted the 200-bp core promoter region in melanoma cells using the CRISPR/Cas9 method (Fig. 1A, fragment IV). As expected, CANT2 expression level was successfully knocked down to ∼20% of its original expression level in A375 and A875 cells (Figs. S7A, B; 2A, lanes 1, 2, and 4). Next, we examined cell proliferation and colony formation ability in vitro. Compared with the empty vector group (NC), tumor cell proliferation was significantly reduced after the suppression of CANT2 (KO1 and KO2) (Fig. 2B, C). Moreover, the CANT2-KO groups presented fewer and smaller colonies (Fig. S7C, lanes 2–3). By calculating the number of colonies, we found that tumor cell colony formation was significantly decreased after CANT2 deletion (Fig. S7D, E). Furthermore, wound healing assays revealed a reduction in tumor migration upon depletion of the CANT2 core promoter (Fig. 2D, E). In contrast to the 50–60% migration area observed in the NC groups, the CANT2-KO groups exhibited a diminished migration area ranging from 20–40% in A375 and A875 cells (Fig. 2F, G). Consistently, transwell assay results further corroborated this observation, indicating a decreased migration ability of tumor cells following CANT2 deletion (Fig. 2H). To confirm the role of CANT2 lncRNA in melanoma, we conducted rescue experiments by overexpressing CANT2 (CANT2-OE) in CANT2-KO melanoma cells. Transfection with CANT2 overexpression plasmids restored or exceeded the original expression level of CANT2 lncRNA (Fig. 2A, lanes 3 and 5; 2I, J). Colony formation assays indicated that the proliferation ability of CANT2-KO melanoma cells improved after CANT2 overexpression (Fig. 2K). Furthermore, transwell assays demonstrated that the CANT2-OE groups exhibited increased metastasis (Fig. S7F). Consistently, wound healing assays revealed an elevated migration area of approximately 40–60% in CANT2-OE melanoma cells compared to the NC groups, which showed only a 10–30% migration area (Fig. S7G–J). Collectively, these results suggested that CANT2 served as a oncogenic lncRNA in tumorigenesis of melanoma.
A PCR of CANT2 expression in CANT2-KO cells transfected with or without a rescue construct (CANT2-OE). GAPDH was served as a positive control. Representative images from three independent experiments. B, C CCK8 assay to measure 4-day cell growth rate in A375 (B) and A875 (C) cells after knockout of CANT2 core promoter. Triplicate experiments were performed. Data are presented as the mean ± SD using a two-tailed Pearson’s r test; **P < 0.01. D, E Representative images of the wound healing assay at 0 hours and 72 hours post-scratch in melanoma cells with or without CANT2 knockout. The dashed lines indicate the wound edge. Representative images from three independent experiments. F, G Quantification of the migration area. The migration area was calculated as the percentage of the wound area covered by cells at 72 hours compared to the initial wound area at 0 hours. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; **P < 0.01; ****P < 0.0001. H Transwell migration assay to assess the migration ability of melanoma cells after knockout of CANT2 core promoter. Representative images from three independent experiments. I, J qPCR of CANT2 expression at RNA level in melanoma cells with CANT2 knockout (CANT2-KO) and in CANT2-KO cells transfected with a rescue construct (CANT2-OE). Relative expression levels were normalized to GAPDH. Data are presented as mean ± SD from three independent experiments. Statistical significance was determined using a two-tailed t-test. *P < 0.05; **P < 0.01; ****P < 0.0001. K Colony formation assay to determine the proliferative ability of melanoma cells after CANT2 overexpression. Representative images from three independent experiments.
CANT2 enhances tumor growth and metastasis in vivo
To investigate the contribution of the CANT2 lncRNA in tumor characteristics in vivo, NC groups and CANT2-KO groups of both A375 and A875 cells were injected subcutaneously into nude mice. We also measured the size of the resultant tumors every 3 days in subcutaneous xenografts. After 12–15 days, the mice were euthanatized, and the tumors were collected for further analysis (Fig. 3A–B). In the CANT2-KO groups, we found that both tumor weight (n = 5–6; ****P < 0.0001; Figs. 3C; S8A) and volume (n = 5–6; ***P < 0.001; ****P < 0.0001; Figs. 3D; S8B) were significantly reduced compared with the NC groups. Most importantly, after suppression of CANT2 lncRNA, the survival rate of mice was significantly extended (n = 6; Fig. 3E, F). To test tumor growth and migration capability after CANT2 suppression in vivo, both A375 and A875 cells were labeled with firefly luciferase to establish metastatic models in nude mice. After tail vein injection in nude mice, tumor systemic metastasis was evaluated by bioluminescence and fluorescence imaging. In the metastatic tumor model, there were fewer metastatic loci of nude mice at 30 days in the CANT2-KO group compared with the NC groups, and luciferase signaling was significantly reduced (Fig. 3G, H). After approximately 30 days, the mice were euthanatized, and the organs were collected for further analysis. Remarkably, histological examination showed that compared with the NC groups, the absence of CANT2 dramatically suppressed metastatic colonization and reduced the number and size of macroscopic nodules observed in the lungs (Figs. 3I; S8C) and adrenal gland (Fig. 3J). These data suggested that the CANT2 lncRNA played an oncogenic regulatory role in tumor growth and metastasis of melanoma in vivo.
A Photograph of orthotopic xenograft at 15 days after the subcutaneous injection of A375 cells with or without CANT2 core promoter deletion (n = 5 mice in each group). B Photograph of orthotopic xenograft at 15 days after the subcutaneous injection of A875 cells with or without CANT2 core promoter deletion (n = 6 mice in each group). C Bar graph showed tumor weight (mg) formed by the A375 cells with or without CANT2 core promoter deletion in a subcutaneous xenograft model. Tumor weight (mg) was measured and presented as the mean ± SD (n = 5 mice) using an unpaired two-tailed t test; ****P < 0.0001. D A xenograft in vivo assay model presented tumor volume (mm3) formed by the A375 cells with or without CANT2 core promoter deletion. Tumor sizes (mm3) were calculated as the (length×width×width)/2 and presented as the mean ± SD (n = 5 mice) using a Pearson’s r test. ****, P < 0.0001. E Survival analysis of mice following intravenous injection with A375 cells with or without CANT2 core promoter deletion (n = 6 mice) were analyzed using a Log-rank (Mantel-Cox) test. F Survival analysis of mice following intravenous injection with A875 cells with or without CANT2 core promoter deletion (n = 6 mice) were analyzed using a Log-rank (Mantel-Cox) test. G Animal imaging system demonstrated the tumor migration ability of A375 cells with or without CANT2 core promoter deletion on tumor bioluminescent signals. Representative images from five independent samples. H Animal imaging system showed the tumor migration ability of A875 cells with or without CANT2 core promoter deletion on tumor bioluminescent signals. Representative images from five independent samples. I, J Representative images of the histological analysis of lung (I) and kidney (J) seeding in mice injected intravenously with CANT2-NC or CANT2-KO1 A375 cells (original magnification, 1×, scale bar: 1 mm). Representative images from three independent samples.
CCBE1 serves as a regulatory target of PHB2 in melanoma
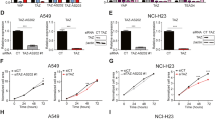
To further elucidate how CANT2 lncRNA impacted melanoma development, we concentrated on identifying and analyzing its downstream targets. We then conducted an RNA-seq (GEO accession number: GSE231936) analysis to scrutinize alterations in gene expression ensuing CANT2 suppression. Using bioinformatics analyses, we identified 249 significantly altered genes, including 174 upregulated and 75 downregulated genes in the NC group (|Fold Change | > 2, P < 0.05, Fig. S6A). The KEGG (Fig. S6B–C) and GO (Fig. S6D–E) analyses revealed significant alterations in pathways related to cancer, inflammation, and the immune system upon CANT2 suppression in tumor cells. Remarkably, within this set of differentially expressed genes, CCBE1 in chr18q21.32 locus exhibited a noteworthy and significant increase in expression following CANT2 inhibition (Fig. 4A). This observation led us to speculate that the CCBE1 is potential target gene of CANT2. As expected, we showed that the expression of CCBE1 was significantly increased in two CANT2-KO melanoma cells in RNA level (Fig. 4B, C), and the results were confirmed in protein level (Fig. 4D). We also investigated CCBE1 expression and observed the absence of CCBE1 in melanoma cells compared with normal PIG1 cells both at RNA (Fig. 4E) and protein (Fig. 4F) levels. Furthermore, the expression levels of CCBE1 in skin melanoma samples were notably reduced compared to those in corresponding normal tissues (Fig. 4G–H) as indicated by data obtained from GEPIA. Next, we investigated whether PHB2 interacts with the CCBE1 promoter (Fig. 4I). As anticipated, PHB2 was found to bind to the CCBE1 promoter (Fig. 4J). Furthermore, we examined the binding of PHB2 in CANT2-KO melanoma cells with or without CANT2 overexpression. ChIP experiments revealed that rescuing the expression of CANT2 lncRNA led to increased binding of PHB2 at the CCBE1 promoter in melanoma cells (Fig. 4K, L). These discoveries suggested that CANT2 lncRNA and its downstream target gene CCBE1 both served as regulatory candidates of PHB2 in melanoma.
A Volcano plot of differentially expressed genes in CANT2-NC compared with CANT2-KO A375 cells. Red, up-regulated genes; green, down-regulated genes; gray, unchanged genes. B, C Real-time PCR and WB of CCBE1 expression in A375 (B) and A875 (C) cells with or without CANT2 core promoter knockout. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; ***P < 0.001 and ****P < 0.0001. D Western blot of CCBE1 expression in melanoma cells with or without CANT2 suppression. Representative blots from three independent experiments. E, F CCBE1 expression at RNA (E) and protein level (F) in melanoma cells and normal PIG1 cells. GAPDH was used as negative control. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; ****P < 0.0001. G, H Gene expression profile (G) and boxplot (H) of CCBE1 in public SKCM dataset from GEPIA. T tumor samples, N normal samples. The |Log2FC| cutoff was set at 1 and the q-value cutoff at 0.01. *P < 0.05. Jitter size was 0.4, and Log2 (TPM + 1) was used for the log-scale. The box plot showed the minima, maxima, centre, bounds of box and whiskers. I Schematic diagram of the primer set d in ChIP assay. J ChIP analysis of PHB2 at the CCBE1 promoter in CANT2-NC or CANT2-KO A375 and PIG1 cells. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. Data are presented as mean ± SD from three independent experiments using a Dunnett’s multiple comparisons test; **P < 0.01; ***P < 0.001; ****P < 0.0001. K, L ChIP analysis of PHB2 at the CCBE1 promoter in CANT2-NC-NC, CANT2-KO-NC, and CANT2-KO-OE A375 and A875 cells. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. Data are presented as mean ± SD from three independent experiments using a Dunnett’s multiple comparisons test; *P < 0.05; ****P < 0.0001.
PHB2 recruits CANT2 and HDAC1 to repress CCBE1 transcription
Next, to elucidate the specific mechanism governing regulation of CCBE1 expression, we conducted chromatin isolation by RNA purification (ChIRP) using biotin-labeled oligonucleotides (Fig. S10A). We selected the CCBE1 promoter as the target site for detection. ChIRP-PCR analysis revealed the enrichment of CANT2 within the CCBE1 promoter region in melanoma cells (Figs. 5A, lane 1; 5B). Nevertheless, upon CANT2 suppression, we observed the reduced enrichment of CANT2 at the CCBE1 promoter, suggesting a direct binding capability of CANT2 to the CCBE1 promoter (Figs. 5A, lane 4; 5B). To examine whether CANT2 could interact with PHB2, we then performed ChIRP-MS, a methodology in which proteins purified through ChIRP were identified via mass spectrometry, aiming to uncover proteins that interact with CANT2 (Fig. 5C). Upon screening and analyzing peptide signals, we identified five proteins (PHB2, TKT, eIF5A, U2A’, and Nup205) enriched in the ChIRP lysate (Fig. S11A, and Table S2). As expected, PHB2 was one of the proteins identified in ChIRP-MS, confirming our above results and further prompting us to select it for a more comprehensive investigation. Following analysis via western blot assay, we successfully confirmed the interaction between PHB2 and CANT2 (Fig. 5D, lane 1 and 3). We further conducted an RNA-ChIP experiment to examine the interaction between CANT2 and PHB2, Nup205, eIF5A, U2’A and TKT. Among these proteins, CANT2 only interacted with PHB2 in A375 and A875 cells, with no significant enrichment observed for the non-specific control U2 (Fig. 5E, F). We then chose PHB2 for next investigation in A375 and A875 cells. Furthermore, given the significant increase of CCBE1 expression in CANT2-KO cells, we proceeded to investigate a representative histone modification, H3K27 trimethylation, across CCBE1 locus. Subsequent ChIP-qPCR analysis revealed that H3K27 acetylation of the CCBE1 promoter was increased in CANT2-KO melanoma cells, with markedly higher levels observed in normal PIG1 cells lacking CANT2 (Fig. 5G). Given that Class I deacetylases, such as HDAC1 and HDAC2, are pivotal in histone deacetylation, our directed our attention to their impact on CCBE1 regulation. Previous studies have illuminated both the shared characteristics and distinct regulatory roles of HDAC1 and HDAC2, illustrating their potential for collaborative or independent regulation of cellular processes45. Initially, our investigation centered on the influence of HDAC1 in the PHB2-mediated CCBE1 transcription. As anticipated, HDAC1 was found to bind to the CCBE1 promoter in the presence of CANT2 lncRNA in melanoma cells (Fig. 5H). These findings led us to hypothesize that PHB2 may facilitate the binding of HDAC1 to the CCBE1 promoter in the presence of CANT2 lncRNA. To confirm this, we initially investigated whether CANT2 directly interacted with HDAC1 through an RNA-ChIP experiment. Results showed that HDAC1 did not directly bind to CANT2 (Fig. 5I). We then examined whether PHB2 acted as the intermediary linking CANT2 and HDAC1. Co-IP assays demonstrated that the PHB2 protein could be pulled out by baiting the HDAC1 protein (Fig. 5J, top, lane 2), and reciprocally, HDAC1 could also be pulled out by baiting the PHB2 protein (Fig. 5J, bottom, lane 3), while the IgG negative control groups exhibited weak binding. Taken together, our finding shows that CANT2 interacts PHB2 and substantially PHB2 recruits HDAC1 to the CCBE1 promoter, ultimately repressing transcription of CCBE1 through H3K27 deacetylation.
A, B ChIRP assay at the CCBE1 promoter in CANT2-NC or CANT2-KO A375 cells. CANT2 oligo indicated the biotinylated antisense oligonucleotides against the CANT2 lncRNA. Negative oligo indicated the scrambled oligonucleotides. Data are presented as mean ± SD from three independent experiments using a Tukey’s multiple comparisons test; ****P < 0.0001. C Selection of ChIRP-MS for CANT2 binding proteins. D Western blot was used to verify the ChIRP-MS results. CANT2 oligo indicated the biotinylated antisense oligonucleotides against the CANT2 lncRNA. Negative oligo indicated the scrambled oligonucleotides. Representative blots from three independent experiments. E, F Real-time PCR analysis of the binding of CANT2 to PHB2, Nup205, eIF5A, U2A’, PABP and TKT using samples from the RNA-ChIP assay in A375 (E) and A875 (F) cells. IgG antibody and U2 RNA were used as negative controls. Data are presented as mean ± SD from three independent experiments using a Šídák’s multiple comparisons test; **P < 0.01 and ****P < 0.0001. G, H ChIP analysis of H3K27ac (G) and HDAC1 (H) at the CCBE1 promoter in CANT2-NC or CANT2-KO A375 and PIG1 cells. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. Data are presented as mean ± SD from three independent experiments using a Tukey’s multiple comparisons test; **P < 0.01; ***P < 0.001; ****P < 0.0001. I Real-time PCR analysis of the binding of CANT2 to HDAC1 using samples from the RNA-ChIP assay in A375 cells. IgG antibody and U2 RNA were used as negative controls. Data are presented as mean ± SD from three independent experiments using a Šídák’s multiple comparisons test; ****P < 0.0001. J Co-IP assay was performed to show the interaction between PHB2 and HDAC1 in CANT2-NC or CANT2-KO A375 cells. IgG was used as a negative control. Representative blots from three independent experiments.
CCBE1 is a tumor suppressor in tumorigenesis of melanoma
To validate the role of CCBE1 in tumorigenesis, we subsequently induced its overexpression in melanoma cells through the construction of plasmid vectors containing the coding sequence (CDS) of CCBE1 (Fig. S12A–B, and 6A). We then assessed cell proliferation and colony formation ability in vitro. Remarkably, the overexpression of CCBE1 (CCBE1-OE) led to a significant reduction in tumor cell proliferation compared with the NC groups (Fig. S12C–D). Additionally, the CCBE1-OE groups exhibited a reduced number of colonies (Fig. 6B, lane 2). Moreover, transwell assay (Fig. S12E, lane 2) and wound healing assay (Fig. 6C–E) revealed that CCBE1 overexpression resulted in a notable reduction in tumor migration compared to the NC groups. Furthermore, we conducted rescue experiments to confirm the role of CCBE1 in melanoma by knocking down CCBE1 in A375 and A875 cells using shRNAs (shCCBE1) (Fig. 6F–G). After knocking down CCBE1 in CCBE1-OE melanoma cells, we observed an increase in tumor cell proliferation (Fig. 6H). Additionally, CCBE1-OE melanoma cells with shCCBE1 (CCBE1-OE-shCCBE1 group) exhibited enhanced metastasis compared to the NC groups, as demonstrated by transwell assay (Fig. S12F) and wound healing assay (Fig. S12G–J). These findings demonstrate that CCBE1 functions as a tumor suppressor in melanoma cells.
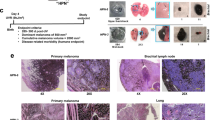
A Western blot of CCBE1 expression at protein level in melanoma cells after CCBE1 overexpression. Representative blots from three independent experiments. B Colony formation assay to determine the proliferative ability of melanoma cells after CCBE1 overexpression. Representative images from three independent experiments. C Representative images of the wound healing assay at 0 hours and 72 hours post-scratch in NC and CCBE1-OE melanoma cells. The dashed lines indicate the wound edge. Representative images from three independent experiments. D, E Quantification of the migration area. The migration area was calculated as the percentage of the wound area covered by cells at 72 hours compared to the initial wound area at 0 hours. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; **P < 0.01; ****P < 0.0001. F, G Western blot of CCBE1 expression in melanoma cells with or without CCBE1 overexpression. Representative blots from three independent experiments. H Colony formation assay to determine the proliferative ability of melanoma cells with or without CCBE1 overexpression. Representative images from three independent experiments. I A xenograft in vivo assay model presented tumor volume (mm3) formed by the A375 cells with or without CCBE1 overexpression. Tumor sizes (mm3) were calculated as the (length×width×width)/2 and presented as the mean ± SD (n = 5 mice) using a Šídák’s multiple comparisons test. ***P < 0.001. Representative images from five independent samples. J Bar graph showed tumor weight (mg) formed by the A375 cells with or without CCBE1 overexpression in a subcutaneous xenograft model. Tumor weight (mg) was measured and presented as the mean ± SD (n = 5 mice) using an unpaired two-tailed t test; *P < 0.05. K Survival analysis of mice following intravenous injection with A375 cells with or without CCBE1 overexpression (n = 5 mice) using a Log-rank (Mantel-Cox) test. Triplicate experiments were performed. L Animal imaging system demonstrated the tumor migration ability of A375 cells with or without CCBE1 overexpression on tumor bioluminescent signals. Representative images from five independent samples. M, N Representative images of the histological analysis of thigh (M) and lung (N) seeding in mice injected intravenously with CCBE1-NC or CCBE1-OE A375 cells (original magnification, 1×, scale bar: 1 mm). Representative images from three independent samples.
To further explore the impact of CCBE1 on tumor development in vivo, we subcutaneously injected CCBE1-OE and control melanoma cells into separate groups of nude mice. We also measured the size of the resultant tumors every 3 days for 12 days in subcutaneous xenografts. Tumor size was monitored every 3 days for 12 days in subcutaneous xenografts (Figs. 6I; S13A–C). Upon completion of this period, the mice were euthanized, and the tumors were collected for subsequent analysis. In the CCBE1-OE groups, we observed a significant reduction in both tumor volume (n = 5–6; **P < 0.01; ***P < 0.001; Figs. 6I; S13B) and weight (n = 5–6; *P < 0.05; ****P < 0.0001; Figs. 6J; S13C) compared with the NC groups. Significantly, the overexpression of CCBE1 led to a noteworthy extension in the survival rate of mice (n = 5–6; Figs. 6K; S13D). To assess tumor growth and migration capability following CCBE1 overexpression in vivo, A375 and A875 cells were labeled with firefly luciferase for the establishment of metastatic models in nude mice. Subsequently, tumor systemic metastasis was assessed through bioluminescence and fluorescence imaging after tail vein injection in these mice. After approximately 30 days, the CCBE1-OE groups exhibited fewer metastatic loci in nude mice compared to the NC groups, with significantly reduced luciferase signaling (Figs. 6L, bottom; S13E, bottom). Remarkably, histological examination showed that compared with the NC groups, the overexpression of CCBE1 significantly suppressed metastatic colonization and reduced the number and size of macroscopic nodules observed in the thigh bone (Figs. 6M, bottom; S13F, bottom), lung (Fig. 6N, bottom), and gastric area (Fig. S13G, bottom). These data suggested that CCBE1 functioned as a tumor-suppressor in melanoma.
PHB2 determines the transcription of CANT2 and CCBE1 during tumorigenesis
Based on the results obtained above, we identified a series of phenomena and mechanisms where PHB2 served as a shuttle factor, simultaneously regulating the transcription of the coding gene CCBE1 and the lncRNA CANT2 from distinct genomic loci. To further validate its indispensable role in these regulatory processes, we first established melanoma cells with PHB2 knockdown, which were named as shPHB2-1 and shPHB2-2 (Fig. 7A–B). Following the knockdown of PHB2, there was a significant reduction in the transcriptional level of CANT2 lncRNA (Fig. 7C). Conversely, the expression of CCBE1 exhibited a substantial upregulation at both the RNA (Fig. 7D) and protein levels (Fig. 7E). In connection with the previously elucidated mechanism governing the regulation of CANT2 and CCBE1, we subsequently investigated the regulatory landscape under the knockdown of PHB2. In the accessible promoter region of CANT2 (Fig. 7F), shPHB2 tumor cells displayed reduced binding of MLL2 (Fig. 7G) and a decreased level of H3K4 trimethylation (Fig. 7H) compared with the empty vector group. At the CCBE1 promoter (Fig. 7I), the suppression of PHB2 also led to a decrease in the binding of HDAC1 (Fig. 7J), accompanied by an increase in H3K27 acetylation level (Fig. 7K). Following analysis via western blot assay for ChIRP, we confirmed that the interaction between PHB2 and CANT2 decreased upon the suppression of PHB2 (Fig. 7L, lanes 1 and 3). Co-IP assays further demonstrated that HDAC1 could not be pulled down by baiting the PHB2 protein when PHB2 was knocked down (Fig. 7M, panel 2, lane 2). ChIRP-PCR analysis revealed that the enrichment of CANT2 at the CCBE1 promoter region was diminished in the absence of PHB2 (Fig. 7N). Furthermore, to confirm whether PHB2 and CANT2 could directly bind to the promoter of CCBE1 and subsequently regulate its expression, we applied the method enChIP46,47,48. Through constructing a dCas9/pCMV expression vector of small guide RNAs (sgRNAs) with two Flag tags, which can be pulled down by the Flag antibody to isolate specific genomic regions, we modified enChIP assay to efficiently target the CCBE1 promoter region (sgCCBE1-p groups), which allow us to directly capture the binding proteins and RNAs simultaneously (Fig. S14A). We first confirmed the efficiency of capturing the CCBE1 promoter (Fig. 7O–P). By collecting RNAs and performing reverse transcription PCR, we found that in melanoma cells, CANT2 lncRNA directly bound to the CCBE1 promoter (Fig. 7Q–R). Additionally, western blot analysis showed that PHB2 directly interacted with the promoter of CCBE1 (Fig. 7S, lane 3). These results convincingly demonstrated that PHB2 played a critical role in regulating the transcription of both the lncRNA CANT2 and the coding gene CCBE1.
A Real-time PCR of the expression of PHB2 in A375 cells with or without PHB2 suppression. Data are presented as mean ± SD from three independent experiments using a Dunnett’s multiple comparisons test; ****P < 0.0001. B Western blot of the expression of PHB2 at protein level in melanoma cells after PHB2 knockdown. Representative blots from three independent experiments. C, D Real-time PCR of the expression of CANT2 (C) and CCBE1 (D) in A375 cells with or without PHB2 suppression. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; *P < 0.05, **P < 0.01. E Western blot of the expression of CCBE1 at protein level in melanoma cells after PHB2 knockdown. Representative blots from three independent experiments. F–H ChIP analysis of PHB2, MLL2 and H3K4me3 at CANT2 core promoter in A375 cells with or without PHB2 knockdown. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. Data are presented as mean ± SD from three independent experiments using a Šídák’s multiple comparisons test; **P < 0.01, ****P < 0.0001. I–K ChIP analysis of PHB2, HDAC1 and H3K27ac at the CCBE1 promoter in A375 cells with or without PHB2 knockdown. Rabbit normal IgG served as the negative control. ChIP enrichment was presented as the percentage of bound/input signal. Data are presented as mean ± SD from three independent experiments using a Šídák’s multiple comparisons test; ****P < 0.0001. L Western blot was used to detect the interaction of PHB2 and CANT2 lncRNA after the suppression of PHB2. CANT2 oligo indicated the biotinylated antisense oligonucleotides against the CANT2 lncRNA. Negative oligo indicated the scrambled oligonucleotides. Representative blots from three independent experiments. M Co-IP assay was performed to show the interaction between PHB2 and HDAC1 after the suppression of PHB2. IgG was used as a negative control. Representative blots from three independent experiments. N ChIRP-PCR assay to dectect the binding of CANT2 lncRNA at the CCBE1 promoter after the suppression of PHB2. CANT2 oligo indicated the biotinylated antisense oligonucleotides against the CANT2 lncRNA. Negative oligo indicated the scrambled oligonucleotides. Data are presented as mean ± SD from three independent experiments using a Tukey’s multiple comparisons test; ****P < 0.0001. O–P Quality control of the enChIP assay in capturing the CCBE1 promoter via PCR. Data are presented as mean ± SD from three independent experiments using a Šídák’s multiple comparisons test; ****P < 0.0001. Q–R RT-PCR analysis of CANT2 lncRNA binding to the CCBE1 promoter using the enChIP method. Data are presented as mean ± SD from three independent experiments using an unpaired two-tailed t test; *P < 0.05; ****P < 0.0001. S Western blot analysis of PHB2 binding to the CCBE1 promoter following the enChIP assay. Representative blots from three independent experiments.
Discussion
The coexistence of aberrant lncRNAs and abnormal coding genes plays a critical role in malignancy development. Understanding how these coding genes and lncRNAs from various loci collectively contribute to malignancy progression poses an intriguing question in cancer research. In this study, we introduced a “one stone two birds” model to explain this co-presence phenomenon in tumorigenesis. We identified PHB2 as a shuttle factor that coordinately orchestrates the transcription of the oncogenic lncRNA CANT2 and the coding tumor-suppressor gene CCBE1, resulting in the accelerated tumorigenesis of melanoma (Fig. S14E).
PHB2 is a pivotal protein in cellular biology, involved in regulating various cellular functions and has potential in various diseases, including cancers35,36,49,50. It is primarily known for its role as a crucial component of the inner mitochondrial membrane and has implications for mitochondrial structure and function30. Moreover, PHB2 serves as a mitophagy receptor, mediating mitophagy and affecting cancer cell growth31,51. In addition, PHB2 has also been found to function alone or interact with other transcription factors to influence gene expression in the nucleus33,52, with emerging research suggests its significance in transcriptional regulation. In our study, however, we showed that PHB2 served as a shuttle factor (the stone) coordinately orchestrated transcription of CANT2 lncRNA (the first bird) and coding tumor-suppressor gene CCBE1 (the second bird) in the tumorigenesis of melanoma. Additionally, other proteins identified in the ChIRP-MS table, such as BRD4, HNRNPM, CTCF, YY1, and cohesin, merit further investigation to explore their interactions with lncRNAs and other functional factors. Since the spatial co-presence of lncRNAs and coding genes in various tumors and other diseases, our concept of “one stone two birds” model may provide an alternative explanation to orchestrate transcriptional regulation of lncRNA and coding gene, and proposing an interesting direction for exploring more unknown functional shuttle factors.
As a canonical disease susceptibility locus, chr6p22 hosts histone gene clusters and variant lncRNA isoforms which function in different diseases. Various mechanisms govern the distinct transcription and function of the diverse coding genes and lncRNAs within the chr6p22.3 region. The deletion on chr6p22.3–p23 harboring ATXN1, DTNBP1, JARID2, and NHLRC1 may be responsible for intellectual disability and autism spectrum disorders53. DCDC2 and KIAA0319 on chr6p22.2 act as candidate susceptibility genes of a language learning disorder developmental dyslexia54. In metastatic melanoma cells such as YDFR.SB3, WP and RKTJ-CB1, an onco-lncRNA CASC15 was transcribed in this locus, and it also promotes tumorigenesis in acute myeloid leukemia (AML), lung and gastric cancers55,56,57,58. Nevertheless, a pair of sense/antisense lncRNAs encoded by CASC15 and NBAT1 are tumor suppressors in neuroblastoma59. Surprisingly, lncRNA CASC15-S, the isoform of lncRNA CASC15, plays the similar role to inhibit neuroblastoma development60. Furthermore, lncRNA CANT1 is another variant transcribed from this locus, playing a role of tumor suppressor in eye malignancies such as uveal melanoma and retinoblastoma61,62. In our study, however, we discovered an oncogenic lncRNA CANT2 that promotes tumorigenesis of melanoma. Furthermore, in breast cancer, the major histone gene cluster at chr6p22 was subdivided into three sub-clusters of histone genes that were organized into hierarchical TADs and located at TAD boundaries, forming an active chromatin hub63. This has raised the question of whether the higher-order chromatin structure plays a role in the chr6p22 locus and initiates melanoma tumorigenesis. Unfortunately, our analysis of visualized Hi-C maps revealed that the TAD boundary remains unaltered in melanoma cells when compared to normal skin cells. In our study, however, we uncovered that the change of chromatin accessibility governing the transcription of a lncRNA variant CANT2 at chrp22.3 locus.
It should be noted that H3K4me3 is a crucial epigenetic modification that plays a central role in gene regulation. This modification is typically associated with transcriptional start sites and open chromatin structure64. H3K4me3 is primarily catalyzed by MLL proteins and SET1/COMPASS methyltransferase complexes65. H3K4me3 helps recruit transcriptional machinery, such as RNA polymerase II, and other chromatin-modifying complexes to initiate gene expression66. The sudden depletion of H3K4me3 may significantly reduce overall transcriptional output, increase RNA polymerase II pausing and slowdown elongation rather than sufficiently affecting transcriptional initiation66. In our study, we observed that PHB2 recruited MLL2 for activating transcription of CANT2 by increasing H3K4me3. It is of great interesting to explore whether this PHB2-guided H3K4me3 pattern could applied in the transcriptional regulation of other gene or lncRNA.
CCBE1 is essential to lymphangiogenesis and presents as a promising therapeutic tool for a variety of diseases involving the lymphatic system67,68. Notably, CCBE1 has emerged as a pivotal regulator of vascular endothelial growth factor-C (VEGFC) signaling69. Researchers have discovered that CCBE1 enhances VEGFC proteolysis, thereby fostering tumor lymphangiogenesis and facilitating lymphatic metastasis in colorectal cancer70. Contrastingly, CCBE1 has been observed to impede the progression of hepatocellular carcinoma by promoting mitochondrial fusion71. CCBE1 exhibits predominant expression in the ovary but is downregulated in ovarian cancer cells and primary carcinomas72. The loss of CCBE1 expression has the potential to promote ovarian carcinogenesis by augmenting cell migration and survival72. In our study, we identified CCBE1 as a tumor suppressor in melanoma. Interestingly, we demonstrated that PHB2 recruits histone deacetylase HDAC1, leading to decreased H3K27 acetylation at the CCBE1 promoter and subsequent repression of CCBE1 transcription. HDAC1, a member of the class I deacetylases, functions to remove lysine-acetyl marks from histone proteins. Previous studies have shown both similarities and differences in the regulatory roles of HDAC1 and HDAC2, highlighting how they may either collaborate or operate independently to regulate cellular processes45. Our findings specifically emphasize the crucial role of HDAC1 in this regulatory pathway. While we cannot entirely exclude the involvement of other factors in PHB2-mediated transcriptional regulation of CCBE1, our study presents a mechanism for the regulation of CCBE1 transcription. Future research should focus on identifying additional factors that may influence the regulation of CCBE1.
Methods
Cell lines
Our research complies with all relevant ethical regulations according to Tongji University. PIG1, HEK-293T cell lines, the malignant melanoma cell lines A375 and A875 cells were purchased and authenticated from American Type Culture Collection (ATCC) and China National Collection of Authenticated Cell Cultures (NCACC), and cultured in DMEM medium (GIBCO) or 1640 medium (GIBCO). All media were supplemented with 10% FBS (GIBCO), 1% penicillin/streptomycin (GIBCO). All cells were incubated at 37°C and 5% CO2. All cells were confirmed to be free of mycoplasma contamination.
Animal study
All animal studies were conducted in accordance with the guidelines provided by the Institutional Animal Care and Treatment Committee of Tongji University. The animals were treated following relevant institutional and national guidelines and regulations. The maximum allowable tumor size/burden (with a diameter of less than 1 cm) was not exceeded. Mice were housed at an ambient temperature of 24 ± 2 °C, with circulating air, constant humidity of 50 ± 10%, and a 12-hour light/dark cycle. Female BALB/c nude mice (GemPharmatech) were used, as patient gender did not have a significant impact on the observations and conclusions of this study. Mice were euthanized by cervical dislocation at the end of the experiment, after which tumors and major organs were collected using surgical scissors.
Tumor xenograft nude mice model
3 × 106 cells (CANT2-NC, CANT2-KO1 and CANT2-KO2) in 100 μl volume of DPBS were subcutaneously injected into the right anterior subcutaneous part of 5-week-old female nude mice. Tumor volume was monitored using a vernier caliper every 3 days and calculated with the formula: length (mm)×width (mm)2/2. Five mice from each group were euthanatized, and the tumors were weighed. We confirm that the maximal tumor size/burden did not exceed the limits permitted (tumor volume: 1 cm3) by our ethics committee or institutional review board.
Tumor metastasis nude mice model
Cells were transfected with a lentivirus encoded by Luc-vectors with neor screening markers. Cells were selected by incubation with 0.5 mg/ml G418 solution (InvivoGen) for 2 weeks. 1.5 × 106 cells (CANT2-NC, CANT2-KO1) in 100 μl volume of DPBS were intravenously injected through the caudal vein of 6-week-old female nude mice. 30 days later, D-Luciferin Sodium Salt (YEASEN) was injected into the right flanks of mice. Bioluminescence was detected by in vivo small animal imaging systems after anesthetization. Five mice from each group were euthanatized, and the pathological section were performed.
RNA extraction and qPCR
Total RNA was extracted from cells using TRIzol reagent (Sigma) and cDNA was synthesized using 1st Strand cDNA Synthesis SuperMix (YEASEN, 11123ES60). Real time qPCR was performed using qPCR SYBR Green Master Mix (YEASEN, 11198ES08) on Roche LightCycler 96 system. The exhibited data represents the fold change (FC) of experimental group versus control group. In brief, ΔCt was calculated as ΔCt = Ct (test gene) - Ct (Ref. gene). ΔΔCt was calculated as ΔΔCt = ΔCt (experimental group) -ΔCt (control group). The FC of a test gene in experimental group versus control group was calculated as FC = 2^(-ΔΔCt). Each gene tested in triplicates in every independent experiment, and all experiments were triplicated. Primers used are listed in Supplementary Table 1.
RACE assay
The RACE assay was performed by SMARTer RACE 5’/3’ Kit (Takara Bio) following the manufacturer’s protocol. First-strand cDNA was synthesized using a modified oligo (dT) primer and SMARTScribe Reverse Transcriptase (RT) which added several non-templated residues. The SMARTer II A Oligonucleotide annealed to the tail of the cDNA and served as an extended template for SMARTScribe RT. After having 3’- and 5’-RACE-Ready cDNA samples, RACE PCR reactions and sequencing were performed to get full-length cDNA.
FAIRE assay
The FAIRE assay was conducted following a previously described protocol73. A total of 1 × 107 cells were crosslinked with 37% formaldehyde to a final concentration of 1% in PBS. The crosslinking reaction was quenched by the addition of 2.5 M glycine. The fixed cells were collected by centrifugation at 1000 g for 5 min at 4 °C and washed three times with 10 ml of 1× PBS. The pellets were resuspended in 1 ml of cold lysis buffer and sonicated to achieve an average DNA fragment size of ∼200–500 bp. The supernatant was transferred to a fresh 1.5-ml tube and incubated with 1 μl of DNase-free RNase A for 30 min at 37°C. Subsequently, 300 μl of phenol/chloroform/isoamyl alcohol was added, and the sample was vortexed for 10 s followed by centrifugation at 12,000 g for 5 min. The aqueous layer was transferred, and the extraction process was repeated three times, pooling all the aqueous solutions. Afterwards, 1/10 volume of 3 M sodium acetate, 2 volumes of 95% ethanol, and 1 μl of 20 mg/ml glycogen were added, and the solution was incubated at −80 °C for at least 30 min. The DNA was then purified using a DNA Clean-up Kit (AxyPrep). The PCR primers used are listed in Supplementary Table 1.
Luciferase assay
Cells were collected and gDNA was achieved by Genomic DNA extraction Kit (TIANGEN, DP304) following the manufacturer’s protocol. Then the gDNA was used for PCR with Q5 High-Fidelity DNA Polymerases (NEB). Primers used are listed in Supplementary Table 1. The promoter region upstream of CANT2 TSS was dissected into 5 fragments of 200 bp and were respectively cloned into the pGL3-basic vector (Promega). HEK-293T cells in a 24-well plate reached 50% confluence within 24 hours before transfection. 900 ng pGL3-basic containing different CANT2 promoter fragments or pGL3-basic and 90 ng pRL-TK (Promega) were co-transfected into HEK-293T cells using lipofectamine 3000 (Invitrogen). After 48 hours, luciferase activity was measured by Dual Luciferase Reporter Gene Assay Kit (YEASEN, 11402ES60) following the manufacturer’s protocol. Each group was repeated with 3 technical replicates. Transfections were repeated in three independent experiments. Firefly luciferase activity was normalized to Renilla luciferase activity. P values were calculated using t test.
CRISPR/Cas9-mediated CANT2 lncRNA knockout
4 sgRNAs targeting the left and right side of the core promoter of CANT2 to be deleted were identified using Optimized CRISPR Design (http://crispor.gi.ucsc.edu) and synthesized. SgRNA oligos were cloned respectively into lentiCRISPRv2 with BsmBI (NEB). SgRNA oligos used are listed in Supplementary Table 1.
CCBE1 overexpression plasmid construction
The coding sequence of CCBE1 were generated by PCR and then cloned into the PGMLV-CMV-MCS-3xFlag-EF1-mScarlet-T2A-Puro vector with XhoI and BamHI (NEB). Primers used are listed in Supplementary Table 1.
Short hairpin RNA (shRNA) targeting PHB2 construction
3 gRNAs targeting PHB2 were synthesized. SgRNA oligos were cloned respectively into PGMLV-hU6-MCS-CMV-ZsGreen1-PGK-Puro-WPRE. SgRNA oligos used are listed in Supplementary Table 1.
Lentivirus packaging and generation of stable cell lines
The lipofectamine 3000 (Invitrogen) was incubated with Opti-MEM I Reduced Serum Medium (GIBCO) and used to transfect HEK-239T cells with 3 mg of the target plasmids, 3 mg of pMD2.D plasmids and 6 mg of PsPax plasmids. Eight hours after transfection, the medium was replaced with 10 ml of fresh medium. The supernatant containing the viruses was collected at 48 and 72 hours respectively. The virus-containing solution was filtered and concentrated. 24 hours prior to transfection, melanoma cells were seeded at 2.0 × 105 cells per well in a six-well plate. The medium was replaced with virus-containing supernatant supplemented with 10 ng/ml polybrene (Sigma). After 48 hours, the medium was replaced with fresh medium. Cells were selected by incubation with 4 mg/ml puromycin (Invitrogen) for 2 weeks. Colonies were derived from single cells tested for the loss of the targeted region, and later expanded for further analyses.
CCK8 cell viability assay
Cells were seeded into a flat-bottomed 96-well culture plate at 1000 cells per well with 100 μl culture medium, and incubated at 37°C with 5% CO2. 10 μl of CCK8 kit (YEASEN, 40203ES80) was added to each well and incubated for 2 h. Then the absorbance was measured at 450 nm with SpectraMax iD3 (Molecular Devices) for 4 consecutive days.
Colony formation assay
2000 cells were suspended in 2 ml DMEM medium and cultured in a six-well plate for 2 weeks. For quantification, the colonies grown in plates were stained with 1% crystal violet and then photographed. The number and size of the colonies were determined using ImageJ.
Wound healing assay
The migratory ability of the cells was evaluated by seeding 1 million cells into a six-well plate and cultured with DMEM medium for 24 h. Then a wound was made by manually scraping the cell monolayer with a 10 μl pipet tip and cells were incubated with FBS-free medium. Images were taken at the indicated times.
Transwell assay
The migratory ability of the cells was evaluated using a 24-well transwell plate with 8.0 µm Pore Polyester Membrane (Corning). The upper compartment contained 30,000 cells suspended in 150 μl DMEM medium containing Basement Membrane Matrix (Corning), and the lower compartment contained 600 μl DMEM medium supplemented with 10% FBS. The transwell system was stained with 0.25% crystal violet after 2 days of incubation at 37°C. The cells on the inner side of the transwell were removed by scrubbing, and the cells on the outer side were photographed.
ChIRP
A total of 1 × 108 cells were crosslinked with 3% formaldehyde at room temperature for 30 min. The reaction was quenched by adding 125 mM glycine for 5 min and then centrifuged at 13,000 g for 5 min at 4 °C. The supernatant was removed. The pellet was weighed and resuspended in 1 ml lysis buffer per 100 mg. DNA was sheared to a size of 300 bp for ChIRP-seq or 1000 bp for ChIRP-MS, respectively (Covaris M220). 1.5 μl RNA probes (100 pmol/L) targeting CANT2 exons were added to every 1 ml sample and incubated at 37 °C overnight with shaking. 50 μl MyOne Streptavidin C1 beads (Life Technologies) was pre-washed with lysis buffer for three times, and then added to the reaction and incubated at 37 °C for 2 h with shaking. Subsequently, the beads were washed 3 times with 1 ml of wash buffer for 5 min per wash. At the last wash step, 1/10 of the beads were reserved for qPCR analysis. The supernatant of the rest of beads was removed. For MS, 200 μl protein elution buffer was added to the beads and incubated at room temperature for 20 min and then at 65 °C for 10 min with shaking. The supernatant was collected and the elution was repeated for twice. Protein was purified with TCA Protein Precipitation Kit (Sangon Biotech) following the manufacturer’s instruction, and then MS was performed. For DNA sequencing, the rest of beads were suspended in 150 μl DNA elution buffer and incubated at 37 °C for 30 min with shaking. The supernatant was collected and the elution was repeated for twice. PK was added to the reaction and incubated at 65 °C for 2 h. DNA was purified and suspended in 20 μl elution buffer, and then sequencing was performed. ChIRP-MS was performed once in this study following the previous protocols74,75.
Western blot
Cells were lysed in RIPA lysis buffer (YEASEN, 20101ES60) containing 1 mM PMSF (YEASEN, 20104ES03) for 30 min, and then centrifuged at 13,000 g for 10 min at 4 °C. Protein samples were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) in 10% (w/v) polyacrylamide gels and transferred to polyvinylidene fluoride membranes (Millipore). After blocking with 5% BSA (YEASEN, 36101ES25) for 1 h at room temperature, the membrane was incubated with different antibodies in 5% BSA overnight at 4 °C. The membrane was then incubated with Peroxidase-Conjugated Goat Anti-Rabbit IgG or Peroxidase AffiniPure Goat Anti-Mouse IgG (YEASEN). The band signals were visualized and quantified using the Fully Automatic Chemiluminescence/Fluorescence Image Analysis System (Tanon). The following antibodies were used in this study: anti-PHB2 (1:1000, CST, 14085), anti-PHB2 (1:1000, Santa Cruz, sc-133094), anti-Histone H3 (1 μg/ml, Abcam, ab176842), anti-HDAC1 (1:1000, CST, 34589), anti-CCBE1 (1:1000, ImmunoWay, YN1730), anti-Nup205 (1:1000, Santa Cruz, sc-377047), anti-eIF5A (1:500, Santa Cruz, sc-390202), anti-U2A’ (1:1000, Santa Cruz, sc-393804), and anti-TKT (1:1000, Santa Cruz, sc-390179).
RNA-chromatin immunoprecipitation (RNA-ChIP)
RNA-ChIP was performed using Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore) following the manufacturer’s instructions. In brief, 107 cells were lysed with RIP lysis buffer with one freeze-thaw cycle. Cell extracts were coimmunoprecipitated with anti-PHB2, anti-Nup205, anti-eIF5A, anti-U2A’, and anti-TKT (Santa Cruz, 1:1000), and the retrieved RNA was subjected to real-time qPCR analysis. Normal mouse IgG was used as a negative control.
RNA extraction library preparation
RNA was extracted from CANT2-NC and CANT2-KO cells using the TRIzol reagent (Invitrogen). RNA purity and concentration were confirmed on a 2100 Bioanalyzer (Agilent Technologies) and a Qubit 2.0 fluorometer with the Qubit RNA Assay Kit (Life Technologies). 2 μg total RNA per sample was used as input material for the RNA sample preparation. Library preparation was completed using the VAHTS Total RNA-seq Library Prep Kit for Illumina (Vazyme) following the manufacturer’s recommendations. The libraries were sequenced by the Illumina PE150 paired-end sequencing platform.
ChIP
ChIP was conducted using an EZ-Magna ChIP A/G kit (Millipore) according to the manufacturer’s protocol. In brief, 107 cells were crosslinked and then lysed. DNA was sheared to a size of 200–500 bp (Covaris M220). DNA collected was coimmunoprecipitated with anti-PHB2 (CST, 1:100), anti-HDAC1(CST, 1:100), anti-MLL2 (1:100, Proteintech, 27266-1-AP), anti-H3K4me3 (1:500, Abclonal, A22146) and anti-H3K27ac (1:500, Abclonal, A22264). Anti-normal mouse IgG was used as a negative control. After immunoprecipitation on A/G beads, DNA was purified and sequenced or amplificated with qPCR. Primers for ChIP-qPCR are listed in Supplementary Table 1.
co-IP
107 cells were carefully washed with pre-chilled PBS for two times, and then collected with 0.5 ml of cold RIPA lysis buffer in a clean 1.5 ml Ep tube. The tube was centrifuged at 14,000 g at 4 °C for 15 min. The supernatant was transferred to a new tube immediately. The protein was diluted to 1 μg/μl. 150 μl of A/G beads (Thermo Scientific) were washed with PBS for two times. 50 μl of washed A/G beads were added to the sample and shaking on a rotator for 10 min at 4 °C. The mixed sample was centrifuged at 14,000 g at 4 °C for 15 min again. The supernatant was transferred to a new 1.5 ml Ep tube, and mixed with 100 μl washed A/G beads. Anti-PHB2 (1:50, CST) and anti-HDAC1 (1:100, CST) were added to the sample respectively, and incubated at 4 °C overnight. The beads were washed with PBS for three times and resuspended in 40 μl RIPAP buffer. Then the sample was proceeded to western blot.
Cytoplasmic and nuclear RNA or protein isolation
Cytoplasmic and nuclear RNA or protein were extracted using PARIS Kit (Thermo Fisher, AM1921) according to the manufacturer’s instructions. The localization of the CANT2 was assayed through the amplification of CANT2 by RT-qPCR. Optimal annealing U6 was used as a nuclear localization reference, and GAPDH was used as a cytoplasmic reference. The localization of the PHB2 was assayed through the western blot. Actin-β (ACTB) was used as a cytoplasmic reference, and H3 was used as a nuclear localization reference.
enChIP
We took the previous enChIP method46,47,48 as references and made some modifications. First, we constructed a dCas9/pCMV expression vector of sgRNAs with two Flag tags, which can be pulled down by the Flag antibody to isolate specific genomic regions, thereby allowing us to directly capture the binding proteins and RNAs simultaneously. After transfecting these vectors into cells, 3/6 × 106 of cells were fixed with 1% formaldehyde at 37 °C for 10 min. The cells were lysed in Cell Lysis Buffer (10 mM Tris pH 8.0, 10 mM NaCl, 0.2% IGEPAL-CA630, cOmplete EDTA-free Protease Inhibitor Cocktail, 100 mM PMSF) for 15 min, and then in Nuclei Lysis Buffer (50 mM Tris pH 8.0, 10 mM EDTA, 1% SDS, cOmplete EDTA-free Protease Inhibitor Cocktail, 100 mM PMSF). The chromatin fraction was fragmented by sonication (500 bp–1 kb). The sonicated chromatin was diluted in 1/10 Dilution Buffer (20 mM Tris pH 8.0, 2 mM EDTA, 150 mM NaCl, 1% Triton X-100, 0.01% SDS, cOmplete EDTA-free Protease Inhibitor Cocktail, 100 mM PMSF), pre-cleared with 15 μg of normal mouse IgG conjugated to 150 μl of A/G beads (Thermo Scientific), and incubated with 15 μg of anti-Flag (Proteintech, 20543-1-AP) conjugated to 150 μl of A/G beads at 4°C overnight.
The beads were washed with 1 ml of Low Salt Wash Buffer (20 mM Tris, pH 8.0, 150 mM NaCl, 2 mM EDTA, 1% TritonX-100, 0.1% SDS), High Salt Wash Buffer (20 mM Tris, pH 8.0, 500 mM NaCl, 2 mM EDTA, 1% TritonX-100, 0.1% SDS), LiCl Wash Buffer (10 mM Tris, pH 8.0, 250 mM LiCl, 1 mM EDTA, 0.5% IGEPAL-CA630, 0.5% sodium deoxycholate), and TE Buffer (10 mM Tris, pH 8.0, 1 mM EDTA). After washing, the beads were divided into three groups. In two groups, the beads were suspended in 285 μl of TE, 12 μl of 5 M NaCl, and Proteinase K, and then incubated at 65°C overnight or 8 h for reverse crosslinking, followed by DNA or RNA extraction using Phenol/chloroform or Trizol. The third group was suspended in RIPA buffer (including PMSF), incubated at 100°C for 10 min, and then subjected to western blot analysis.
Statistics and reproducibility
The analyses were performed on three biological replicates (n). All statistical analyses were performed using the GraphPad 9.0 software and Microsoft Excel. The significance was set at P < 0.05. *P < 0.05, **P < 0.01, ***P < 0.001, ns = non-significant (P > 0.05). All the values are presented as mean ± standard deviation (SD). Source data are provided as a Source Data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-seq data generated in this study are available at the Gene Expression Omnbus GSE231936, and the Sequence Read Archive (SRA) BioProject PRJNA970260. The ChIRP-MS data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD055094. The RNA-seq and ATAC-seq dataset from melanoma cells and SKCM dataset from GEPIA publicly available data used in this study are available in the GEO database under accession code GSE22388737, GSE22388837, GSE23237538, GSE18839842, GSE13443243, and GSE24144544. Source data are provided with this paper.
References
White, N. M. & Maher, C. A. The potential use of lncRNAs found in the 8q24 region as biomarkers for colon cancer. Ann. Oncol. 28, 1688–1689 (2017).
Ahmadiyeh, N. et al. 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proc. Natl Acad. Sci. USA 107, 9742–9746 (2010).
Chen, H., Liu, H. & Qing, G. Targeting oncogenic Myc as a strategy for cancer treatment. Signal Transduct. Target Ther. 3, 5 (2018).
Li, R., Wang, X., Zhu, C. & Wang, K. lncRNA PVT1: a novel oncogene in multiple cancers. Cell Mol. Biol. Lett. 27, 84 (2022).
Zhou, C. et al. LncRNA PVT1 promotes gemcitabine resistance of pancreatic cancer via activating Wnt/beta-catenin and autophagy pathway through modulating the miR-619-5p/Pygo2 and miR-619-5p/ATG14 axes. Mol. Cancer 19, 118 (2020).
Zhang, C. et al. LncRNA CCAT1 facilitates the progression of gastric cancer via PTBP1-mediated glycolysis enhancement. J. Exp. Clin. Cancer Res 42, 246 (2023).
Ohashi, M. et al. Adenovirus mediated p53 tumour suppressor gene therapy for human gastric cancer cells in vitro and in vivo. Gut 44, 366–371 (1999).
Du, M. et al. Remote modulation of lncRNA GCLET by risk variant at 16p13 underlying genetic susceptibility to gastric cancer. Sci. Adv. 6, eaay5525 (2020).
Chai, P. et al. Generation of onco-enhancer enhances chromosomal remodeling and accelerates tumorigenesis. Nucleic Acids Res 48, 12135–12150 (2020).
Fan, J. et al. A Cohesin-Mediated Intrachromosomal Loop Drives Oncogenic ROR lncRNA to Accelerate Tumorigenesis. Mol. Ther. 27, 2182–2194 (2019).
Finn, E. H. & Misteli, T. Molecular basis and biological function of variability in spatial genome organization. Science 365, 6457 eaaw9498 (2019).
Finn, E. H. et al. Extensive Heterogeneity and Intrinsic Variation in Spatial Genome Organization. Cell 176, 1502–1515 e1510 (2019).
Hansen, A. S., Pustova, I., Cattoglio, C., Tjian, R. & Darzacq X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. Elife 6, e25776 (2017).
Hogg, S. J., Beavis, P. A., Dawson, M. A. & Johnstone, R. W. Targeting the epigenetic regulation of antitumour immunity. Nat. Rev. Drug Discov. 19, 776–800 (2020).
Balsalobre, A. & Drouin, J. Pioneer factors as master regulators of the epigenome and cell fate. Nat. Rev. Mol. Cell Biol. 23, 449–464 (2022).
Ge, R. et al. Epigenetic modulations and lineage plasticity in advanced prostate cancer. Ann. Oncol. 31, 470–479 (2020).
Wen, X. et al. Interruption of aberrant chromatin looping is required for regenerating RB1 function and suppressing tumorigenesis. Commun. Biol. 5, 1036 (2022).
Wang, J. et al. Epigenomic landscape and 3D genome structure in pediatric high-grade glioma. Sci. Adv. 7, 23 eabg4126 (2021).
Xu, J. et al. Subtype-specific 3D genome alteration in acute myeloid leukaemia. Nature 611, 387–398 (2022).
Escobar, T. M. et al. Active and Repressed Chromatin Domains Exhibit Distinct Nucleosome Segregation during DNA Replication. Cell 179, 953–963 e911 (2019).
Cutter, A. R. & Hayes, J. J. A brief review of nucleosome structure. FEBS Lett. 589, 2914–2922 (2015).
Xia, W. et al. Resetting histone modifications during human parental-to-zygotic transition. Science 365, 353–360 (2019).
Batie, M. et al. Hypoxia induces rapid changes to histone methylation and reprograms chromatin. Science 363, 1222–1226 (2019).
Bushweller, J. H. Targeting transcription factors in cancer - from undruggable to reality. Nat. Rev. Cancer 19, 611–624 (2019).
Chen, Y. et al. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat. Med 19, 1023–1029 (2013).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Buitrago, D. et al. Nucleosome Dynamics: a new tool for the dynamic analysis of nucleosome positioning. Nucleic Acids Res 47, 9511–9523 (2019).
Sun, X. et al. BRD8 maintains glioblastoma by epigenetic reprogramming of the p53 network. Nature 613, 195–202 (2023).
Wei, Y., Chiang, W. C., Sumpter, R. Jr, Mishra, P. & Levine, B. Prohibitin 2 Is an Inner Mitochondrial Membrane Mitophagy Receptor. Cell 168, 224–238 e210 (2017).
Yan, C. et al. PHB2 (prohibitin 2) promotes PINK1-PRKN/Parkin-dependent mitophagy by the PARL-PGAM5-PINK1 axis. Autophagy 16, 419–434 (2020).
Djehal, A. et al. Targeting prohibitin with small molecules to promote melanogenesis and apoptosis in melanoma cells. Eur. J. Med Chem. 155, 880–888 (2018).
Bavelloni, A., Piazzi, M., Raffini, M., Faenza, I. & Blalock, W. L. Prohibitin 2: At a communications crossroads. IUBMB Life 67, 239–254 (2015).
Pabona, J. M., Velarde, M. C., Zeng, Z., Simmen, F. A. & Simmen, R. C. Nuclear receptor co-regulator Kruppel-like factor 9 and prohibitin 2 expression in estrogen-induced epithelial cell proliferation in the mouse uterus. J. Endocrinol. 200, 63–73 (2009).
Xiang, X. et al. Cellular senescence in hepatocellular carcinoma induced by a long non-coding RNA-encoded peptide PINT87aa by blocking FOXM1-mediated PHB2. Theranostics 11, 4929–4944 (2021).
Liu, M., Wang, Z., Li, S., Deng, Y. & He, N. Identification of PHB2 as a Potential Biomarker of Luminal A Breast Cancer Cells Using a Cell-Specific Aptamer. ACS Appl Mater. Interfaces 14, 51593–51601 (2022).
Insco, M. L. et al. Oncogenic CDK13 mutations impede nuclear RNA surveillance. Science 380, eabn7625 (2023).
Schlesinger, D. et al. A large-scale sORF screen identifies putative microproteins and provides insights into their interaction partners, localisation and function. bioRxiv, 2023.2006.2013.544808 (2023).
Wang, L. et al. CPAT: Coding-Potential Assessment Tool using an alignment-free logistic regression model. Nucleic Acids Res 41, e74 (2013).
Kang, Y. J. et al. CPC2: a fast and accurate coding potential calculator based on sequence intrinsic features. Nucleic Acids Res 45, W1 W12–W16 (2017).
Wang, Y. et al. The 3D Genome Browser: a web-based browser for visualizing 3D genome organization and long-range chromatin interactions. Genome Biol. 19, 151 (2018).
Donohue, L. K. H. et al. A cis-regulatory lexicon of DNA motif combinations mediating cell-type-specific gene regulation. Cell Genom 2, (2022).
Wouters, J. et al. Robust gene expression programs underlie recurrent cell states and phenotype switching in melanoma. Nat. Cell Biol. 22, 986–998 (2020).
Centore, R. C. et al. Pharmacologic inhibition of BAF chromatin remodeling complexes as a therapeutic approach to transcription factor-dependent cancers. Elife 13, RP93478 (2024).
Gonneaud, A. et al. HDAC1 and HDAC2 independently regulate common and specific intrinsic responses in murine enteroids. Sci. Rep. 9, 5363 (2019).
Campbell, A. E. et al. NuRD and CAF-1-mediated silencing of the D4Z4 array is modulated by DUX4-induced MBD3L proteins. Elife 7, e31023 (2018).
Fujita, T. & Fujii, H. Efficient isolation of specific genomic regions and identification of associated proteins by engineered DNA-binding molecule-mediated chromatin immunoprecipitation (enChIP) using CRISPR. Biochem Biophys. Res Commun. 439, 132–136 (2013).
Fujita, T. & Fujii, H. Direct identification of insulator components by insertional chromatin immunoprecipitation. PLoS One 6, e26109 (2011).
Wu, B. et al. PHB2 promotes tumorigenesis via RACK1 in non-small cell lung cancer. Theranostics 11, 3150–3166 (2021).
Ren, L. et al. PHB2 promotes colorectal cancer cell proliferation and tumorigenesis through NDUFS1-mediated oxidative phosphorylation. Cell Death Dis. 14, 44 (2023).
Jiang, M. et al. Artemisinin Alleviates Cerebral Ischemia/Reperfusion-Induced Oxidative Damage via Regulating PHB2-Mediated Autophagy in the Human Neuroblastoma SH-SY5Y Cell Line. Oxid. Med Cell Longev. 2022, 6568748 (2022).
Taniguchi, K. et al. Prohibitin-2 is a novel regulator of p21(WAF1/CIP1) induced by depletion of gamma-glutamylcyclotransferase. Biochem Biophys. Res Commun. 496, 218–224 (2018).
Celestino-Soper, P. B. et al. Deletions in chromosome 6p22.3-p24.3, including ATXN1, are associated with developmental delay and autism spectrum disorders. Mol. Cytogenet 5, 17 (2012).
Adler, W. T. et al. Position of neocortical neurons transfected at different gestational ages with shRNA targeted against candidate dyslexia susceptibility genes. PLoS One 8, e65179 (2013).
Wu, Q. et al. Long non-coding RNA CASC15 regulates gastric cancer cell proliferation, migration and epithelial mesenchymal transition by targeting CDKN1A and ZEB1. Mol. Oncol. 12, 799–813 (2018).
Sun, J. et al. Hypoxia-sensitive long noncoding RNA CASC15 promotes lung tumorigenesis by regulating the SOX4/beta-catenin axis. J. Exp. Clin. Cancer Res 40, 12 (2021).
Lessard, L. et al. The CASC15 Long Intergenic Noncoding RNA Locus Is Involved in Melanoma Progression and Phenotype Switching. J. Invest Dermatol 135, 2464–2474 (2015).
Grasedieck, S. et al. The long non-coding RNA Cancer Susceptibility 15 (CASC15) is induced by isocitrate dehydrogenase (IDH) mutations and maintains an immature phenotype in adult acute myeloid leukemia. Haematologica 105, e448–e453 (2020).
Mondal, T. et al. Sense-Antisense lncRNA Pair Encoded by Locus 6p22.3 Determines Neuroblastoma Susceptibility via the USP36-CHD7-SOX9 Regulatory Axis. Cancer Cell 33, 417–434 e417 (2018).
Russell, M. R. et al. CASC15-S Is a Tumor Suppressor lncRNA at the 6p22 Neuroblastoma Susceptibility Locus. Cancer Res 75, 3155–3166 (2015).
Xing, Y. et al. CANT1 lncRNA Triggers Efficient Therapeutic Efficacy by Correcting Aberrant lncing Cascade in Malignant Uveal Melanoma. Mol. Ther. 25, 1209–1221 (2017).
Ni, H. et al. LncRNA CANT1 suppresses retinoblastoma progression by repellinghistone methyltransferase in PI3Kgamma promoter. Cell Death Dis. 11, 306 (2020).
Fritz, A. J. et al. Intranuclear and higher-order chromatin organization of the major histone gene cluster in breast cancer. J. Cell Physiol. 233, 1278–1290 (2018).
Han, D. et al. BRWD3 promotes KDM5 degradation to maintain H3K4 methylation levels. Proc. Natl Acad. Sci. USA 120, e2305092120 (2023).
Shilatifard, A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu Rev. Biochem 81, 65–95 (2012).
Wang, H. et al. H3K4me3 regulates RNA polymerase II promoter-proximal pause-release. Nature 615, 339–348 (2023).
Jeltsch, M. et al. CCBE1 enhances lymphangiogenesis via A disintegrin and metalloprotease with thrombospondin motifs-3-mediated vascular endothelial growth factor-C activation. Circulation 129, 1962–1971 (2014).
Zou, Z. et al. The secreted lymphangiogenic factor CCBE1 is essential for fetal liver erythropoiesis. Blood 121, 3228–3236 (2013).
Roukens, M. G. et al. Functional Dissection of the CCBE1 Protein: A Crucial Requirement for the Collagen Repeat Domain. Circ. Res 116, 1660–1669 (2015).
Song, J. et al. CCBE1 promotes tumor lymphangiogenesis and is negatively regulated by TGFbeta signaling in colorectal cancer. Theranostics 10, 2327–2341 (2020).
Tian, G. A. et al. CCBE1 promotes mitochondrial fusion by inhibiting the TGFbeta-DRP1 axis to prevent the progression of hepatocellular carcinoma. Matrix Biol. 117, 31–45 (2023).
Barton, C. A. et al. Collagen and calcium-binding EGF domains 1 is frequently inactivated in ovarian cancer by aberrant promoter hypermethylation and modulates cell migration and survival. Br. J. Cancer 102, 87–96 (2010).
Gaulton, K. J. et al. A map of open chromatin in human pancreatic islets. Nat. Genet 42, 255–259 (2010).
Zhang, S. et al. Comparison of viral RNA-host protein interactomes across pathogenic RNA viruses informs rapid antiviral drug discovery for SARS-CoV-2. Cell Res 32, 9–23 (2022).
Chu, C. et al. Systematic discovery of Xist RNA binding proteins. Cell 161, 404–416 (2015).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Grant Nos. 82372705 and 31870748) to H.Z., the Shanghai Oriental Elite Project (Grant No. 2000152009) to H.Z., the National Key Research and Development Program of China (Grant No. 2017YFE0196300) to H.Z., the Shanghai Natural Science Foundation (Grant No. 22ZR1466100) to H.Z., the Fundamental Research Funds for the Central Universities (Grant No. 22120230292) to H.Z., the Key Laboratory of Organ Development and Epigenetics of Jiangxi Province (Grant No. 2024SSY07141) to H.Z., the Shuguang Project of Shanghai Municipal Education Commission and Shanghai Education Development Foundation (Grant No. 17SG19) to H.Z., the Outstanding Young Medical Scholar of Shanghai Municipal Commission of Health and Family Planning (Grant No. 2017YQ067) to H.Z., the Outstanding Yong Scholar Grant of Tongji University (Grant No. PA2019000239) to H.Z., and the Startup Funding of Frontier Science Research Center for Stem Cells & Shanghai East Hospital of Tongji University (Grant No. DFRC2019003) to H.Z.
Author information
Authors and Affiliations
Contributions
Tianyi Ding: Conceptualization, Investigation, Methodology, Visualization, Writing—original draft, Writing—review & editing. Haowen Xu: Methodology, Visualization, Writing—original draft, Writing—review & editing. Xiaoyu Zhang: Investigation, Methodology, Data curation. Fan Yang: Resources. Jixing Zhang: Methodology. Yibing Shi: Investigation. Yiran Bai: Investigation. Jiaqi Yang: Resources. Chaoqun Chen: Resources. Chengbo Zhu: Resources. He Zhang: Project administration, Funding acquisition, Conceptualization, Supervision, Writing—original draft, Writing—review & editing. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Michael Emmons, Jifan Hu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, T., Xu, H., Zhang, X. et al. Prohibitin 2 orchestrates long noncoding RNA and gene transcription to accelerate tumorigenesis. Nat Commun 15, 8385 (2024). https://doi.org/10.1038/s41467-024-52425-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-52425-z