Abstract
Myotonic Dystrophy type 1 (DM1), a highly prevalent form of muscular dystrophy, is caused by (CTG)n repeat expansion in the DMPK gene. Much of DM1 research has focused on the effects within the muscle and neurological tissues; however, DM1 patients also suffer from various metabolic and liver dysfunctions such as increased susceptibility to metabolic dysfunction-associated fatty liver disease (MAFLD) and heightened sensitivity to certain drugs. Here, we generated a liver-specific DM1 mouse model that reproduces molecular and pathological features of the disease, including susceptibility to MAFLD and reduced capacity to metabolize specific analgesics and muscle relaxants. Expression of CUG-expanded (CUG)exp repeat RNA within hepatocytes sequestered muscleblind-like proteins and triggered widespread gene expression and RNA processing defects. Mechanistically, we demonstrate that increased expression and alternative splicing of acetyl-CoA carboxylase 1 drives excessive lipid accumulation in DM1 livers, which is exacerbated by high-fat, high-sugar diets. Together, these findings reveal that (CUG)exp RNA toxicity disrupts normal hepatic functions, predisposing DM1 livers to injury, MAFLD, and drug clearance pathologies that may jeopardize the health of affected individuals and complicate their treatment.
Similar content being viewed by others
Introduction
Myotonic dystrophy type 1 (DM1) is an autosomal dominant disease and the second most common form of muscular dystrophy, affecting more than one in three thousand adults in North America1,2,3. The cardinal symptoms of DM1 include myotonia, debilitating muscle weakness and wasting, abnormal heart function, and excessive fatigue1,2. Despite DM1’s initial characterization as a form of muscular dystrophy, the disease is genuinely multisystemic; patients report various gastrointestinal, metabolic, and neurological dysfunctions, such as excessive daytime sleepiness and insulin resistance2,4.
DM1 is caused by a (CTG)n repeat expansion in the 3’ UTR of a ubiquitously expressed gene Dystrophia Myotonica protein kinase (DMPK)5,6,7. The (CUG)n containing RNAs resulting from the transcription of the diseased DMPK gene form hairpin secondary structures and aggregate in the nucleus, forming discrete RNA foci7,8. These foci interact with and sequester the muscleblind-like (MBNL) family of splicing factors9,10. MBNL proteins affect many developmentally regulated alternative splicing and polyadenylation decisions in various tissues throughout the process of maturation towards adulthood; thus, their loss-of-activity in DM1 shifts splicing of target pre-mRNAs towards preadolescent-like patterns, inducing specific features of the disease11,12,13,14,15,16. This reversal of transcriptomic patterns from mature-to-immature state drives many disease symptoms to become more prevalent later in life, with diagnosis often occurring in the mid to late thirties12,17,18. However, patients are typically diagnosed only after the development of significant muscular or neurological symptoms, which allows the subtle and long-term consequences of the disease to go unmanaged18.
DM1 patients are abnormally sensitive to a wide range of anesthetics and muscle relaxants, resulting in prolonged anesthesia recovery, heightened pulmonary dysfunction, and, in some cases, death1,19,20. The disruption of neurological and muscular function, hallmarks of DM1, is often blamed for this sensitivity. However, within the last twenty years, several studies have demonstrated that DM1 patients have an increased susceptibility to metabolic dysfunction-associated fatty liver disease (MAFLD), metabolic syndrome, and liver damage21,22,23,24. These studies would suggest inappropriate liver function and a predisposition for liver injury in DM1 patients. A malfunctioning liver could also help explain the sensitivity to anesthetic treatment; a liver that cannot provide adequate metabolism of xenobiotic material would prolong the clearance time for many drugs and may increase their potency. Minor malfunctions in liver response to metabolic signaling and drug metabolism may frequently occur in DM1 patients; however, none have investigated this possibility.
Here, we sought to determine what effects DM1 might have on hepatic functions and overall liver health. Utilizing two previously established mouse lines, we generated a mouse model in which we induced the expression of (CUG)960 repeat-containing RNA, specifically in the hepatocytes within the liver25,26,27. Combining acute and long-term DM1 liver mice models with systematic biochemical, molecular, and high-resolution transcriptome analyses, we found that toxic (CUG)960 RNA expression triggered global gene expression and RNA processing defects in the hepatocytes. These transcriptome defects led to various physiological and cellular pathologies, including accumulation of lipids and fatty liver, increased susceptibility to insult and injury, and misregulation of xenobiotic metabolism. Specifically, we uncovered that aberrant splicing and upregulation of Acetyl-CoA Carboxylase 1 (ACC1), the rate-limiting enzyme for de novo fatty acid biosynthesis, drives the MAFLD phenotype in DM1-afflicted mice livers. Importantly, both the fatty liver and the poor drug metabolism phenotypes are intrinsic to the repeat RNA toxicity within hepatocytes, as these defects were not detected in transgenic mice, which express the repeat RNA only in the muscle tissues. Thus, our study reveals that DM1 disrupts normal hepatic functions, predisposes the liver to fatty liver disease and injury, and confirms the need for further research into the effects of DM1 in non-traditional tissues, including the liver.
Results
A hepatocyte-specific murine model of DM1 recapitulates the molecular features of the disease in the liver
The pathogenic mechanism of DM1 is comprised of three primary parts: (i) the transcription and production of a long CUG repeat-containing RNA, (ii) the accumulation of this RNA into nuclear foci, and (iii) the sequestration of MBNL proteins into such RNA foci, which results in the decrease of MBNL-directed RNA processing activities28,29,30. To study the effects of DM1 within the liver, we generated a bi-transgenic murine model by combining two existing mice models. First is the tetracycline-inducible mouse model with a DMPK transgene containing the last five exons of human DMPK and 960 interrupted CTG repeats (labeled here as CUG960i RNA), developed by Cooper and colleagues25. The second model utilizes the expression of a reverse tetracycline trans-activator (rtTA) driven by a liver-specific apolipoprotein E (ApoE) promoter, which is highly expressed in hepatocytes26. By crossing these two mice models, we generated a double homozygous bi-transgenic line that allows for conditional, doxycycline (Dox)-dependent expression of the CUG960i RNA specifically in the liver tissue, thus allowing the study of DM1 disease in the liver (Fig. 1a). From now on this bi-transgenic model is referred to as the DM1 liver model. The control mice for this model contain only the homozygous ApoE-rtTA allele.
a Schematic illustrating the bi-transgenic, hepatocyte-specific, doxycycline (Dox)-inducible model developed to express toxic CUG960i RNA in mouse livers. The (CUG)n repeat-containing transcripts sequester RNA-binding proteins, including MBNL proteins. Administration of Dox triggers the expression of toxic RNA in hepatocytes. This model is referred to as the DM1 liver model. b Experimental protocol for Dox diet feeding to induce DM1 in mouse livers, involving a 2 g/kg Dox-supplemented diet until weaning on day 21, followed by a switch to a 0.1 g/kg Dox diet or maintenance on the 2 g/kg Dox diet for six weeks. Glucose tolerance testing (GTT) occurs a week before sacrifice. Created in BioRender. Adesanya, O. (2024) BioRender.com/k51j388. c Hybrid RNA fluorescent in-situ hybridization immuno-fluorescence (RNA FISH-IF) imaging depicts toxic (CUG)n RNA (red) and Mbnl1 (green) foci in hepatocyte nuclei (blue). d Quantification of CUG960i/Mbnl1 foci in hepatocyte nuclei via RNA FISH-IF. DM1 liver mice on 0.1 g/kg Dox diet (n = 7) are compared to the No Dox diet (n = 5) and 2-week recovery mice (n = 7). e Distribution of CUG960i/Mbnl1 foci per hepatocyte nucleus in mice fed 0.1 g/kg Dox diet for 1-month post-weaning (n = 7). f Quantitative-PCR (qPCR) analysis of CUG960i RNA in whole liver of DM1 liver mice and controls (ApoE-rtTA: n = 11, DM1 liver mice: n = 27, No-Dox mice: n = 12, 2-week recovery mice: n = 7). Box plots display the first to third quartile, a median line, upper and lower whiskers representing the minima and maxima respectively. **P < 0.01, ****P < 0.0001, by two-tailed unpaired T tests (d, f). Scale bars represent 20 µm. Source data are provided as a Source Data file.
To mimic the DM1 conditions seen in human patients, we induced the disease in newborn pups by feeding the mother a diet supplemented with 2 g of Dox per kg of chow. Once weaned, mice then continued a Dox diet at a lower dose of 0.1 g/kg until they reached adulthood at nine weeks (Fig. 1b). At eight weeks of age, mice were fasted for 20–22 hours and administered a glucose tolerance test (GTT).
To confirm the appropriate expression of the CUG960i RNA and the formation of toxic RNA/RNA binding protein foci, we utilized fluorescent in-situ hybridization and immunofluorescence (FISH-IF) imaging with probes targeting the CUG repeat sequence within the RNA (Fig. 1c)31. Bright punctae of condensed CUG RNA were seen in the nuclei of most hepatocytes. The signal from these RNA foci overlapped with the immunofluorescent signal when fluorescent MBNL1 or MBNL2 antibodies were used, indicating that the toxic CUG RNAs have successfully sequestered the MBNL proteins. The MBNL-containing RNA foci only occurred in the DM1 mice livers and not in the ApoE-rtTA control mice livers.
Quantification of the CUG960i/MBNL foci indicated that over 80% of hepatocyte nuclei in the DM1 mice contain at least one RNA focus, ensuring the uniformly distributed expression of both transgenes (Fig. 1d). The appearance of RNA foci was Dox-dependent—when mice with both the ApoE-rtTA and CUG960i alleles were not fed Dox or if the Dox diet was withdrawn for a week or more, the RNA foci were barely detectable. The distribution of CUG960i RNA foci per nucleus followed a Poisson curve, with most hepatocytes having one to three foci and a small number of hepatocytes exceeding ten foci within a single nucleus (Fig. 1e). Subsequent RNA FISH-IF experiments using hepatocyte zonation markers E-Cadherin (E-Cad) for the periportal zone (zone I) and Glutamine synthetase (GluS) for the pericentral zone (zone III), confirmed the uniform distribution of CUG960i RNA foci across the three hepatic zones (Supplementary Fig. 1)32.
Finally, to assay the amount of toxic RNA produced, CUG960i transgene expression was quantified by isolating RNA from whole livers and conducting qPCR using primers located in the final exon of the Dmpk transcript. DM1 liver mice expressed the CUG960i transgene at levels near ten percent of β-actin transcripts within the liver, compared to ApoE-rtTA control mice, which showed no evidence of CUG960i expression (Fig. 1f). A comparison of CUG960i RNA within isolated hepatocytes versus that of the whole liver confirmed that the transgene’s expression occurs primarily within hepatocytes (Supplementary Fig. 2a, b). Livers of the bi-transgenic mice not fed Dox showed no CUG960i expression, confirming that Dox must be provided to these animals to express sufficient amounts of toxic RNA. (Fig. 1f).
Expression of CUG960i RNA induces global transcriptomic changes within hepatocytes
Upon establishing that the DM1 liver model reproduces the molecular features of the disease, we prepared total RNA from purified hepatocytes isolated from the ApoE-rtTA controls and DM1-afflicted mice fed a 2.0 g/kg Dox-supplemented diet for nine weeks. We next assessed the splicing patterns of MBNL1-regulated exons within these RNA samples using end-point reverse-transcription PCR (RT-PCR) assays, including an exon in Mbnl1 transcript, which contains an auto-regulatory element that affects the sub-cellular localization of MBNL1 protein33,34,35,36. The DM1 mouse livers consistently reproduced an alternative splicing pattern that significantly deviated from the control samples (Fig. 2a), confirming that, like the muscle and brain tissues, the expanded CUG repeat-containing RNA of DM1 also induces splicing defects in the liver.
a RT-PCR splicing analysis of select MBNL1 targets (n = 5 biological replicates). Bands indicate exon presence or absence, with (+) for inclusion and (-) for exclusion. Targets are listed to the left of the image, with the percentage “spliced in” (PSI) below. b Overlap of alternative splicing, alternative polyadenylation (APA), and expression changes upon DM1 induction in hepatocytes. c Volcano Plot illustrating mRNA abundance changes from RNA-seq as identified from DESeq2. d Violin plots displaying inclusion levels of alternative splicing events from RNA-seq data. MXE mutually exclusive, A3’SS/A5’SS alternative 3’/5’ splice sites, ASE alternative cassette exon, RI retained intron. e Change in PSI determined by RT-PCR (y-axis) vs. RNA Seq analysis (x-axis) for 30 events. f Gene Tracks of representative genes showing alternative exon inclusion in DM1 liver, Mbnl1∆E3/∆E3 (KO), or wild-type animals. g Pie chart – comparison of alternatively spliced genes regulated by DM1, MBNL1, or maturation in the liver. h Pie chart – comparison of differentially expressed genes (DEG) regulated by DM1, MBNL1, or maturation. i GO Diagram - mRNA Processing: Selected processes related to genes with alternative mRNA processing events in DM1 liver. j GO Diagram - Differential Expression: Processes related to genes undergoing differential expression in DM1 liver. b–j For the RNA-seq datasets, n = 3 for ApoE-rtTA control, n = 3 for DM1 liver mice, n = 2 for Mbnl1 WT, and n = 2 for Mbnl1 KO mice. Source data are provided as a Source Data file.
We next performed high-resolution RNA sequencing of poly(A) selected RNA from the purified hepatocyte samples to further explore the genome-wide RNA processing defects in DM1-afflicted livers. Analysis of the resulting data revealed widespread changes in the DM1 hepatocyte transcriptome, with significant changes in mRNA abundance, splicing, and alternative polyadenylation (ApA) (Fig. 2b). Focusing on gene expression, inducing DM1 within the murine liver changed the mRNA abundance of 760 transcripts at a 2-fold level or higher, with 516 upregulated and 244 downregulated compared to control livers (Fig. 2c).
As the MBNL proteins are most known for regulating alternative splicing events, it is not surprising that nearly one thousand splicing events changed upon the expression of the CUG960i RNA within the liver. Of the 928 splicing events that demonstrated a greater than 10% change in PSI (Percent Spliced In) within the DM1 liver, every type of alternative splicing event was represented, with most falling under the category of cassette exons, and 35 of those events showing a ΔPSI change of 50% or higher (Fig. 2d, Supplementary Fig. 2c). Using RT-PCR, forty-one of these alternatively spliced events were validated; the comparison showed a high consistency between the RNA-seq and RT-PCR results (Fig. 2e).
Gene ontology analysis of the transcripts experiencing dysregulation in abundance, splicing, or ApA revealed enrichments in unique functional categories. Transcripts changing in abundance were enriched in glucose, lipid, and energy-related metabolism, as well as oxo-reductase and cytochrome p450 activities (Fig. 2i, Supplementary Table 1)37. The transcripts with altered splicing patterns were enriched in mRNA processing, signal transduction, and protein phosphorylation. A substantial number of transcripts encoding proteins associated with the immune response, specifically the response to viral and bacterial infection, exhibited defects in both overall abundance and splicing (Fig. 2j, Supplementary Tables 1–3). Transcripts with misregulated ApA events, much like misregulated splicing events, were enriched in nucleotide binding, protein binding, and transport-related functions.
The proposed molecular mechanism of DM1 entails disrupting MBNL protein activities, resulting in a transcriptomic shift away from the normal state of healthy adult tissue and towards an immature state in the muscles, heart, and neurons13,29,30,38,39. To test whether this pattern holds within the liver, we isolated hepatocytes from Mbnl1ΔE3/ΔE3 (Mbnl1 knockout) mice and corresponding littermate wild-type controls at ten weeks of age40. Again, RT-PCR splice assays were performed on the poly(A) selected RNAs purified from wild-type and Mbnl1 KO hepatocyte samples, and the results were compared to the DM1 liver and ApoE control samples. Notably, the DM1 liver samples showed a shift in splicing away from controls in the same direction as the Mbnl1 KO samples (Fig. 2f; Supplementary Fig. 2d, e). However, the DM1 samples often demonstrated a more significant deviation from the controls than the Mbnl1 KO samples.
This prompted a full comparison of differentially expressed genes and alternatively spliced events between the DM1 and Mbnl1 KO hepatocyte transcriptomes via RNA-Seq. Of the 705 alternatively spliced events in DM1 hepatocytes and the 508 that occurred in the hepatocytes of Mbnl1 KO mice (difference in ΔPSI > 15%), only 175 events were common to both sets (Supplementary Fig. 2f). A similar pattern was observed for differentially expressed genes, with only a 15% overlap between the DM1 and Mbnl1 KO data sets (Supplementary Fig. 2g). This limited overlap was surprising, as MBNL1 sequestration is a crucial driver of the transcriptomic defects in DM1 heart and muscle tissues.
As MBNL proteins promote tissue maturation and function, we next compared the transcriptomic changes in hepatocytes isolated from adult livers of DM1, Mbnl1 KOs, and wild-type mice livers during postnatal maturation41. By comparing alternatively spliced transcripts that change in either the context of DM1, Mbnl1 KO, or postnatal liver maturation, we found that only about 25% of the events changing in DM1 were regulated by MBNL1 (Fig. 2g, h)41. Of note, whereas only a modest portion of mis-spliced events in DM1 were developmentally regulated, over 50% of transcripts changing in abundance in the DM1 liver were also developmentally regulated.
To explore the limited overlap between the hepatic transcriptomes of DM1 and Mbnl1 KO mice with developing livers, potential compensatory mechanisms for MBNL1 function were investigated. We found that while MBNL1 levels are wholly depleted within the Mbnl1 KO livers, MBNL2 levels were elevated 4-fold, as confirmed by western blot analysis (Supplementary Fig. 2h). The upregulation of MBNL2 might explain why there is lower-than-expected overlap between changes in the Mbnl1 KO and DM1 liver models. Additionally, the increase of MBNL2 implies a compensatory mechanism that buffers the effects of Mbnl1 loss within the liver, a mechanism demonstrated in other tissues42,43.
Hepatocyte-specific expression of CUG960i RNA induces increased lipid accumulation and liver injury
As the effects of DM1 in the liver are unstudied, and even the role of MBNL proteins in the liver is unknown, we took a generalized approach to assess the pathological consequences of DM1 within the liver. This process started before sacrifice, as blood glucose levels just before sacrifice indicate a slight difference in the blood glucose levels between male DM1 liver mice and male controls (Fig. 3a). However, this difference does not occur within the female groups (Supplementary Fig. 3a). There was also no difference between DM1 liver mice and controls during glucose tolerance testing performed in the weeks before sacrifice (Fig. 3b; Supplementary Fig. 3b). Median mouse weight between female control and DM1 mice also showed no significant difference; however, DM1 male mice were 10–17% larger than ApoE-rtTA control males (Supplementary Fig. 3c). This increase in size is likely not due to the induction of DM1 in the DM1 liver mice, as male mice of this strain were larger than the ApoE-rtTA control males regardless of whether they were consuming a Dox diet.
a Blood glucose levels were measured before sacrifice after a 4-hour fast (n = 15 for ApoE-rtTA mice and 16 for DM1 mice). b Glucose tolerance testing (GTT) curves depicting blood glucose levels post-intraperitoneal glucose injection. GTT was performed a week before harvest following a 24-hour fast (n = 23 for ApoE-rtTA mice, 27 for DM1 liver mice, and 18 for HSA L/R mice). c Representative histological images of ApoE-rtTA control, DM1 liver, and DM1 liver + 2-week recovery mice. Hematoxylin and eosin (H&E) images showcasing inflammation and necrosis in the DM1 liver, with inflammation and necrosis circled, are at the top. Oil Red-O images indicating lipid droplets (Red) with nuclei stained in hematoxylin (blue) are at the bottom. Picro Sirius Red showed increased fibrosis in the DM1 liver mice compared with controls. Black scale bars represent 200 µm. d Hepatosomatic index for DM1 mice and ApoE-rtTA controls (n = 16 for ApoE-rtTA mice and 32 for DM1 liver mice). e Quantification of Oil Red O signal, relative to hematoxylin-stained nuclei, indicating lipid accumulation (n = 18 for ApoE-rtTA mice and 15 for DM1 liver mice). Box plots show the first to third quartile with a median line and minima and maxima as whiskers;. All other plots represent mean ± SD. *P < 0.05, ****P < 0.0001, by two-tailed unpaired T tests (a, d, e). Source data are provided as a Source Data file.
As glucose intolerance is a common symptom in DM1, we compared GTT analysis from the DM1 liver mice and control mice against the HSA L/R mice, a DM1 model commonly studied for skeletal muscle pathologies8. The HSA L/R model expresses the toxic CUG repeat-containing RNA only within the muscle tissues, allowing us to compare the direct contributions of liver and muscle tissue toward glucose intolerance in DM1. While the HSA L/R mice showed significant glucose intolerance, the DM1 liver mice showed normal glucose handling (Fig. 3b).
Histological analysis of the DM1 mice livers revealed varying degrees of morphological changes and regions with decreased sinusoidal spacing within the DM1 livers (Fig. 3c). Additionally, increased lobular inflammation, necrotic patches, and mild fibrosis were found in DM1 livers, which could not be reversed after 2 weeks of Dox withdrawal in the recovery mice (Fig. 3c). In contrast, H&E revealed that HSA L/R livers appear healthy, with no indication of increased injury (Supplementary Fig. 3d).
DM1 patients have shown an increased susceptibility to fatty liver disease21,22. Therefore, we used Oil Red O staining on frozen mice liver tissues to interrogate the lipid accumulation within the DM1 liver model. Relative to the control animals, DM1 liver mice showed a significant increase in lipid droplets, which was reduced after 2 weeks of Dox withdrawal in the recovery mice (Fig. 3c, e). While long-term Dox diet can result in modest hepatic steatosis, the DM1 mice consistently displayed higher lipid levels, nearly twice that of respective controls. Furthermore, the mouse liver-to-carcass weight ratio showed a significantly higher hepatosomatic index in DM1 liver mice than in controls (Fig. 3d), with a median increase of 36.6%. Oil Red O staining of HSA L/R livers demonstrated no lipid accumulation, revealing another pathology unique to the DM1 liver model (Supplementary Fig. 3d). Additionally, combined Oil Red O-IF staining with the periportal and pericentral zonation markers, indicated that fat accumulation in DM1 livers occurs similarly across the three hepatic zones (Supplementary Fig. 3f).
DM1 liver model mice demonstrate decreased drug metabolism
An often-reported challenge when treating DM1 patients is their increased susceptibility to anesthetics and analgesics2,19,44. These complications are most noticeable during surgical procedures wherein DM1 patients exhibit much longer recovery times from various anesthetics and muscle relaxants. In the case of a few anesthetics, the patient may require intervention to prevent death20,45. Because the liver is the primary organ involved in drug metabolism, we hypothesized that DM1 livers might be compromised in responding to and metabolizing xenobiotics, thereby decreasing DM1 patients’ ability to clear certain drugs from their system.
We first chose zoxazolamine, a muscle relaxant, to test this hypothesis. Zoxazolamine testing in mice consists of inducing muscle paralysis in the animals via zoxazolamine injection and then monitoring them until they can self-right and move around freely (Fig. 4a). Zoxazolamine metabolism shows a sex-specific response in mice; however, in both males and females, DM1 mice took at least 50% longer to recover from the drug-induced paralysis (Fig. 4b).
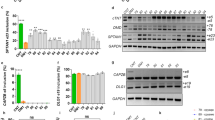
a Schematic illustrating zoxazolamine (Zox) testing procedure, measuring self-righting time post-intraperitoneal (IP) injection of 120 mg/kg Zox. Created in BioRender. Adesanya, O. (2024) BioRender.com/a13o207 b Normalized average self-righting time of Zox-injected mice (n = 9, 10, 7, 8 from left to right). Fisher’s exact test. c Diagram outlining acetaminophen (APAP) drug metabolism in hepatocytes. d Mortality rate 8 hours post 350 mg/kg APAP injection (n = 13, 19, 10, 14 from left to right). e Representative livers of male mice surviving 24 hours post-APAP injection. f H&E images of livers from male mice 24 hours post-APAP injection (n = 3 biological replicates). Black scale bars represent 500 µm. g, h Western blots showing CYP2E1 and SULT2A1 protein levels in control and DM1 mice (n = 3 biological replicates). Box plots display the first to third quartile with a median line and minima and maxima as whiskers. *P < 0.05, **P < 0.01, ***P < 0.001, by two-tailed unpaired T tests (b, d). Source data are provided as a Source Data file.
We next tested if a similar reduction in drug metabolism occurred with common, over-the-counter analgesics such as acetaminophen (APAP). APAP causes severe liver injury/damage if consumed in high concentrations as it generates toxic levels of N-acetyl-p-benzoquinone imine (NAPQI) metabolite after oxidation by cytochrome P450 2E1 (CYP2E1) in perivenous hepatocytes46 (Fig. 4c). Even a low dose of APAP can induce liver toxicity if CYP2E1 activity is high. Conversely, the liver can be insulated from APAP toxicity if CYP2E1 activity is ablated47. In most mice strains, the LD50 of APAP is between 320 and 370 mg per kg of body weight when administered intraperitoneally48. To see if DM1 changed susceptibility to APAP-induced hepatic injury, we injected 350 mg of APAP per kg of body weight into fasted DM1 and control mice. Mice were monitored for 8 hours and left to recover for 16 hours before being sacrificed.
A difference was immediately noticed between DM1 and control mice, as significantly more control mice died within the first 8 hours compared to DM1 mice (Fig. 4d). Upon collecting the liver from the surviving mice, 24 hours post-APAP injection, almost all control animals show widespread signs of liver necrosis (Fig. 4e). However, APAP-treated DM1 livers showed fewer instances of injury and necrosis. H&E staining of the APAP-treated livers also showed marked differences between DM1 and control mice, with DM1 mice still showing extensive injury and hepatocyte vacuolization but far less necrosis (Fig. 4f). Western blot analysis of CYP2E1, showed that CYP2E1 is expressed at significantly lower levels in the livers of DM1 afflicted mice (Fig. 4g). Similarly, sulfotransferase 2a1 (SULT2a1), an enzyme associated with phase 2 drug metabolism49,50, was also downregulated in the livers of the DM1 mice (Fig. 4h). Decreases in these two enzymes can lead to less conversion of APAP into the toxic NAPQI, resulting in less hepatic injury and death in the DM1 liver mice, despite being representative of a muted capacity for these mice to metabolize circulating xenobiotics46,51.
DM1 murine liver models are more susceptible to fatty liver disease and injury
As DM1 patients face dietary and mobility challenges that often require counseling and careful monitoring, we set out to test if the macronutrient composition of the patient’s diet impacts the DM1 liver’s susceptibility to MAFLD2,4,52,53. To do so, we fed DM1 liver mice and ApoE-rtTA controls standard chow supplemented with a 2 g/kg Dox diet until weaning, as previously described. Once weaned, the mice were switched to a high-fat, high-sugar, and heightened cholesterol (western) diet supplemented with 0.1 g/kg Dox for eight additional weeks (Fig. 5a)54,55. As before, we analyzed GTT and four-hour fasting glucose levels before sacrifice.
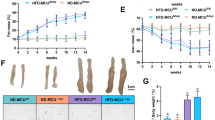
a Schematic of the feeding protocol for mice on a Western diet, starting with a 2 g/kg Dox-supplemented diet at birth and transitioning to a 0.1 g/kg Dox-supplemented high-fat, high-sugar “Western” diet at weaning, maintained for eight weeks until sacrifice. b GTT curves of male mice on the western diet performed twice with a six-day interval (n = 23 for ApoE-rtTA mice and 16 for DM1 mice). c Blood glucose levels before sacrifice after a 4-hour fast (n = 22 for ApoE-rtTA mice and 10 for DM1 liver mice). d Representative liver images from ApoE-rtTA (left) and DM1 liver mice (right) on the Western diet. e Representative histological images, including H&E (Top) images with circled inflammation and necrosis in DM1 liver and Oil Red-O (Bottom two) images staining lipid droplets (Red) with nuclei in blue. Black scale bars represent 500 µm. f Analysis of extractable triglycerides in livers of mice on basal (n = 12 for ApoE-rtTA mice and 14 for DM1 liver mice) and western diets (n = 21 for ApoE-rtTA mice and 14 for DM1 liver mice). g Hepatosomatic index for Western diet-fed DM1 mice and ApoE-rtTA mice (n = 34 for ApoE-rtTA mice and 19 for DM1 liver mice). Box plots display the first to third quartile with a median line and minima and maxima as whiskers. Other plots represent mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001, by two-tailed unpaired T test (c, f, g). Source data are provided as a Source Data file.
GTT analysis again showed no difference between DM1 mice and control animals (Fig. 5b); however, there was a slight difference in 4-hour fasted blood glucose levels between male DM1 and male control mice (Fig. 5c). In reverse of the basal diet, DM1 mice had significantly lower blood glucose.
While control livers turned pale following a high fat, high sugar diet, the DM1 livers became exceedingly lighter, with much of the usual red color replaced with off-white due to excess lipid accumulation (Fig. 5d). Both DM1 and control mice showed significant increases in micro-and macro-vesicular steatosis, inflammation, and evidence of cell death on the western diet compared to the regular chow diet (Fig. 5e). However, DM1 mice showed more macro-vesicular steatosis and patchy necrosis as well as ballooning and feathery degeneration after western diet feeding.
Oil Red O staining showed that livers of DM1 mice had a much higher density of lipid droplets and a significant increase in the number of large lipid droplets (Fig. 5e), making them challenging to quantify via image analysis. Therefore, we used an alternative method to determine the relative accumulation of lipids in the western diet-fed control and DM1 mice. A hexane/isopropanol lipid extraction protocol collected hydrophobic fatty acids from small liver portions. A colorimetric assay for triglycerides was performed on these extracts (Fig. 5f)56,57,58. This analysis revealed two features. First, the livers from DM1 mice fed the regular rodent diet accumulated as many triglycerides as those from control mice on the western diet. Second, the livers of DM1 mice fed a Western diet accumulated significantly more triglycerides than any other group.
Western diet-fed DM1 mice also had a more significant increase in the liver-to-body weight ratios than controls (Fig. 5g). The mean body weight between DM1 and control mice is invariant, suggesting that the increased accumulation of lipids in the liver is not due to a more significant increase in body weight (Supplementary Fig. 3e).
Upregulation of an alternatively spliced ACC1 isoform drives lipid accumulation in DM1 liver
To investigate how DM1 affects liver-specific lipid handling/metabolism and leads to a fatty liver, we focused on DM1-related changes in splicing/abundance of transcripts linked to lipogenesis, lipid transport, lipid metabolism, and MAFLD. Of these, acetyl-CoA carboxylase 1 (Acc1) particularly stood out (Fig. 6a). ACC1 is at the rate-limiting step for the conversion of excess citrate into free fatty acids (Fig. 6b); thus, any changes in its function or regulation could directly lead to excess lipid production and potentially explain the steatosis noted in the DM1-afflicted livers. ACC1 regulation involves multiple phosphorylation clusters and a necessary dimerization event to function59,60,61,62,63,64. Upon activation, ACC1 converts acetyl-CoA to malonyl-CoA, a necessary building block for fatty acid synthesis and lipogenesis. ACC1 activity is negatively regulated through phosphorylation by the kinases AMPK, PKA, and CDK and by excess palmitoyl-CoA levels. In contrast, it is positively regulated by insulin-induced dephosphorylation via the phosphatase PP2A and excess citrate levels (Fig. 6b)65,66,67.
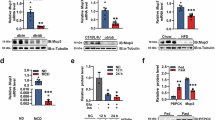
a RT-PCR-based analysis of select alternative splicing events in ApoE-rtTA, Mbnl1 KO and DM1-afflicted livers (n = 6 biological replicates). b Diagram of acetyl-CoA carboxylase 1 (ACC1) regulation and function. c Schematic of ACC1 alternative splicing in DM1 livers, displaying locations of alternatively spliced exon and notable domains. Gene names on the left. P phosphorylation cluster, BC biotin carboxylase domain, BB biotin binding/biotin carboxylase carrier protein domain, CT carboxyltransferase domain. d Percent inclusion of Acc1 exon 28 in chronic DM1 and controls and mice on a westernized diet (n = 10, 13, 24, 21 from left to right). e qPCR results showing Acc1 transcript levels in control and chronic DM1 liver mice (Control, n = 10; Chronic DM1, n = 9). f Western blot showing ACC1 protein levels in chronic DM1 and control mice (n = 6 biological replicates). g Lipid accumulation in acute and chronic DM1 mice livers via Oil Red-O (ORO) staining (n = 3 biological replicates). Black scale bars represent 50 µm. h Diagram summarizing gene abundance changes related to lipid regulation in livers of acute and chronic DM1 mice and chronic DM1 mice treated with ACC1 inhibitor CP-640186. Created in BioRender. Adesanya, O. (2024) BioRender.com/m33y833. i RT-PCR based analysis of Acc1 exon 28 splicing in acute and chronic DM1 mice livers (n = 3 biological replicates). j Changes in Acc1 exon 28 inclusion in chronic DM1 mice treated with either control antisense oligonucleotide (ASO) or Acc1 exon 28-targeting ASO (n = 3 biological replicates). Re-arranged lanes demarcated with clear borders. k ORO staining of chronic DM1 mice livers treated with Acc1 exon 28 or control ASO (n = 3 biological replicates). Black scale bars represent 200 µm. l ORO staining from chronic DM1 mice treated with ACC1 inhibitor CP-640186 or vehicle (n = 3 biological replicates). Black scale bars represent 200 µm. Box plots display the first to third quartile with a median line and minima and maxima as whiskers. ***P < 0.001, ****P < 0.0001, by two-tailed unpaired T test (d, e). Source data are provided as a Source Data file.
The exon 28 that changes in the context of DM1 is centrally situated in the protein-coding region of the Acc1 transcript; exclusion of the exon results in the loss of 8 amino acids directly N-terminal to the central phosphorylation cluster (Fig. 6c)66,68. This exon is included in Acc1 transcripts in most tissues, having only been demonstrated to be excluded in the brain, mammary tissues, ovaries, and liver68. In mammary tissues of sheep and goats, skipping of this exon results in increased phosphorylation and decreased function of ACC1, altering the lipid profiles of ovine milk68,69.
In the livers of DM1 mice, exon 28 was included in ∼30% more Acc1 transcripts than in control mice (Fig. 6a, d). Unexpectedly, MBNL1 deficiency in the liver did not affect Acc1 splicing (Fig. 6a). Furthermore, upon feeding a high-fat, high-sugar diet, the inclusion of exon 28 increased slightly in control but not in DM1 mice, which maintained similar PSI values as on standard chow (Fig. 6d). Additionally, qPCR analysis revealed a 2-fold increase in expression of Acc1 transcript levels in DM1 livers compared to controls, and western blot analysis also showed significantly higher ACC1 protein levels in the livers of DM1 mice relative to controls (Fig. 6e, f). Because reliable transfer of high molecular weight proteins such as ACC1 (250–280 kDa) can be challenging, we verified this result using the Jess capillary-based immunoassay. In the Jess system, size-based protein separation, immobilization, and target detection are performed inside individual capillaries in an automated manner, allowing sensitive and accurate quantification of target protein (Supplementary Fig. 4b). The Jess results replicated our western blot data; DM1 liver samples showed a 2-fold increase in ACC1 protein levels compared to ApoE-rtTA controls (Supplementary Fig. 4c). The size difference between the exon 28-included and excluded isoforms of ACC1 is <0.5% of the total weight of the protein and could not be resolved in either the western blot or the Jess assay. Whether increased ACC1 protein abundance in DM1 mice livers is linked to exon 28 inclusion or occurred independently through another event(s) regulating mRNA and/or protein stability/translation remains to be determined.
To discern whether the upregulation of alternatively spliced ACC1 isoform was a primary consequence of DM1 or a secondary response to steatosis, we used an acute DM1 liver model. In contrast to the chronic model, Acute DM1 mice were aged for eight weeks before being fed 0.1 g/kg Dox-containing diet for 12 or 18 days (Supplementary Fig. 4a). The acute DM1 mice did not exhibit any steatosis, which was evident in the chronic DM1 model (Fig. 6g). Next, we compared the hepatic mRNA abundance of genes that are directly linked to lipid biosynthesis, transport or metabolism-related functions in acute versus chronic DM1 mice via qPCR. In many cases, the mRNA levels were similar between acute and chronic DM1 livers. However, some transcripts in the acute model showed significant differences from the chronic model, highlighting transcripts that may have changed in response to pathological changes in the chronic DM1 liver (Fig. 6h, Supplementary Fig. 4d). Importantly, missplicing of Acc1, Pnpla6, and Sorbs1 transcripts was evident in both acute and chronic DM1 liver models (Fig. 6i; Supplementary Fig. 4e). Acute DM1 livers also demonstrated a 54% increase in ACC1 protein levels over ApoE-rtTA control mice; this increase was lower than the 2-fold increase in the chronic DM1 livers (Supplementary Fig. 4c). These data demonstrate that ACC1 misregulation in DM1 is a direct effect of repeat RNA toxicity and not a secondary response to lipid accumulation or liver injury.
To determine if the upregulation of alternatively spliced ACC1 isoform contributes to DM1-related MAFLD, chronic DM1 mice were treated with antisense oligonucleotides (ASO) targeting the 5’ss of exon 28 of Acc1 (Supplementary Fig. 5a). ASO treatment consisted of two loading doses of 12 mg/kg ASO the first two days of treatment, followed by two maintenance doses on days 6 and 10, and the mice were sacrificed on day eleven. After ASO treatment, the inclusion of exon 28 in DM1 mice livers dropped to less than 20%, an almost 60% decrease relative to untreated or control ASO-treated DM1 livers (Fig. 6j). Despite having a striking effect on Acc1 splicing, the ASO treatment did not cause a statistically significant change in Acc1 transcript or ACC1 protein abundance (Supplementary Fig. 5b, c). We next evaluated the functionality of Acc1 splicing redirection on lipid accumulation in DM1 mice livers. Chronic DM1 mice treated with ASO targeting Acc1 exon 28 showed only a modest improvement in hepatic steatosis compared to control ASO treatment (Fig. 6j, k, Supplementary Fig. 5d).
To further investigate whether ACC1 is required for the development of MAFLD phenotype in DM1-afflicted mice livers, we used the ACC1 inhibitor (CP-640186) to prevent ACC1 activity in these mice (Supplementary Fig. 6a). This inhibitor has been previously characterized for its ability to inhibit ACC1 activity, and it effectively reduces malonyl-CoA concentrations and fatty acid synthesis in cultured cells and mice livers70,71. We orally administered 25 μg/g ACC1 inhibitor twice daily to chronic DM1 mice for five days and then analyzed their liver health. Strikingly, short-term inhibition of ACC1 activity led to a significant recovery in hepatic steatosis in DM1-afflicted livers (Fig. 6k, Supplementary Fig. 6b) without affecting the expression of most lipid metabolism-related genes (Fig. 6h; Supplementary Fig. 4d). However, while the inhibition of ACC1 significantly reduced the appearance of fatty liver, markers of poor liver health associated with DM1-related MAFLD, such as increased hepatosomatic index and lobular inflammation, failed to improve after treatment (Supplementary Fig. 6c, d). This suggests that these phenotypes in chronic DM1 mice either occur independently of ACC1 dysregulation or that a longer-term ACC1 inhibition is required to reverse the pathologies developing from sustained lipid accumulation. Collectively, these data demonstrate that upregulation of the alternative ACC1 splice isoform drives the bulk of hepatic lipid accumulation in DM1 and that short-term inhibition of ACC1 can reverse the hepatic steatosis phenotype in DM1-afflicted mice livers.
Discussion
DM1 symptoms often extend beyond the musculature, wherein various tissues manifest specific pathologies that contribute to the affected individual’s overall health10,28,72. For instance, patients commonly experience gastrointestinal disturbances, metabolic dysregulation, increased sensitivity to drug injury, as well as the development of MAFLD and other liver dysfunctions20,21,22,24,73,74,75,76. However, the root cause of such dysfunctions is not fully understood. Elevated liver enzyme levels (GGT, AP) in DM1 individuals are also observed consistently but are speculated to occur secondary to the gallbladder and bile duct dysmotility, which might impair bile excretion and indirectly affect liver function24,73. In this study, we provide multiple lines of evidence that targeted expression of CUG repeat-containing RNA within hepatocytes is sufficient to alter the function of the liver, resulting in steatosis and hepatocellular injury. When combined with diet-induced metabolic stress, this predisposes the DM1 liver toward MAFLD and compromises its ability to respond to and metabolize specific analgesics and muscle relaxants. We determined that both MAFLD and poor drug metabolism phenotypes are hepatocyte-intrinsic and are direct consequences of repeat RNA toxicity in the liver, as these defects were not seen in the HSA L/R transgenic mice, which express repeat RNA only in the muscle tissues. Conversely, the HSA L/R mice suffer from significant glucose intolerance, whereas the DM1 liver mice are normal in glucose handling. These findings highlight the importance of studying the effects of DM1 within individual tissues and evaluating their respective contributions to the metabolic symptoms of this complex disease.
We further demonstrate that hepatic expression of toxic CUG960i RNA triggers global mRNA abundance and processing defects in genes enriched in ontologies that group into major functional clusters, especially lipid and drug metabolism, cell signaling and immune responses, as well as macromolecular binding and transport-related activities. These gene expression defects can directly lead to increased lipid accumulation and heighten the liver’s vulnerability to damage from toxins/dietary stress, partly explaining the high incidence of MAFLD and metabolic disorders seen in DM1 patients21,22,24. For instance, scores of genes involved in phase I and II drug metabolism were misregulated in the DM1 liver model. Notably, the levels of CYP2E1 and SULT2A1 proteins were significantly reduced in the livers of DM1 mice. A decrease in these two proteins alone may help to explain some of the poor responses to sedatives seen in DM1 patients, especially in the case of commonly inhaled anesthetics, which tend to be halogenated hydrocarbons77,78,79. Further work is needed to determine whether abnormal drug metabolism or dysregulation of drug-metabolizing enzymes is a direct consequence of repeat RNA toxicity or an aberrant response to lipid accumulation and DM1-related MAFLD.
The pathological mechanism of DM1 involves sequestration and disruption of MBNL protein activities, shifting the affected tissues’ transcriptome from an adult-to-preadolescent-like state. Also, MBNL expression is upregulated in hepatocytes as the liver matures after birth80,81. Intriguingly, when we compared the hepatic transcriptomes of DM1 and Mbnl1 KO mice, less than a third of the misregulated events occurring in the DM1 hepatocytes were detected in MBNL1 deficient hepatocytes. The dissimilarities between the DM1 liver model and Mbnl1 KOs were also evident at the tissue level, as many of the pathological consequences of expressing DM1 in the liver were not seen in the Mbnl1 KO mice, including the lack of lipid accumulation. MBNL proteins serve critical functions in maintaining tissue maturity; however, which MBNL family member is predominantly responsible for these activities is tissue-dependent11,15,81,82,83. Additionally, MBNL proteins have high sequence and structure similarity and significant overlap in binding targets; thus, substitution between the two predominant members, MBNL1 and MBNL2, allows for a tunable regulatory system that maintains appropriate splicing form for the bulk of co-regulated transcripts81,83,84. Our examination of Mbnl1 KO mice revealed that MBNL2 protein levels are increased in the KO livers, a phenomenon also seen in other tissues42,81,82,85. In the future, it would be interesting to determine whether elevated expression of MBNL2 could explain the lack of overlap in transcriptome changes between the Mbnl1 KO and DM1 livers, as well as shed light on possible compensatory mechanism(s) that buffer the effects of MBNL1 loss within the liver.
Several genes associated with insulin-regulated lipid metabolism were altered in DM1-afflicted livers. Because lipid homeostasis is regulated jointly by intra- and extra-hepatic signaling, these changes could be a primary result of repeat RNA toxicity or a secondary effect of the diseased liver. Indeed, there is some evidence of a secondary effect occurring—the expression and/or splicing of genes such as Sorbs1, Fabp1, Acox1, and Hsd17b13 were altered less in the acute DM1 liver as compared to the chronic DM1 liver, possibly indicating effects that are exacerbated with lipid accumulation. However, based on our analysis, missplicing and upregulation of ACC1 is a rapid response to the repeat RNA expression. ACC1 is a focal point for de novo lipid biosynthesis, converting excess acetyl-CoA into malonyl-CoA, which can then be used to produce palmitate86,87. Therefore, ACC1 activity must be tightly controlled in response to the nutritional state of the liver as misregulation of ACC1 stimulates excess lipogenesis and MAFLD phenotypes88,89.
Our results indicate that short-term inhibition of ACC1 activity is sufficient to reverse lipid accumulation in the DM1-afflicted mice livers, highlighting the importance and involvement of this pathway. We believe this beneficial effect is primarily due to inhibition of de novo lipogenesis; however, reduced production of malonyl-CoA would also increase the mitochondrial import and oxidation of fatty acids. Although extrapolating these findings to DM1-related MAFLD in humans is not straightforward, pharmacologic inhibition of ACC is currently viewed as one of the most promising therapeutic approaches for treating MAFLD/MASH90,91,92,93,94. It is important to note that a small proportion of MAFLD patients treated with ACC inhibitors may experience hypertriglyceridemia over time, which can increase their risk of developing cardiovascular disease95,96,97. Whether inhibiting ACC1 in the context of DM1 leads to increased serum triglycerides remains to be determined. Therefore, in future studies, it will be essential to examine the long-term effects of ACC1 inhibitors on systemic lipid trafficking and mobilization in DM1 liver mice.
In conclusion, our study provides the first characterization of the direct impact of DM1 on liver health. The findings offer valuable insights into how disrupted hepatic functions contribute to the metabolic symptoms and drug sensitivities in DM1, underscoring the idea that a malfunctioning liver can further complicate the treatment of this complex genetic disease. A current limitation of this work is that our results need to be validated in humans, and the extent of similarity between the murine DM1 liver model and actual patient livers must be systematically assessed. Therefore, enhanced screening for MAFLD/hepatocellular injury in affected individuals, along with access to liver biopsies, is urgently needed because if DM1 livers cannot provide adequate metabolism of xenobiotic material, it would prolong the clearance time for many drugs, altering their therapeutic index. Future investigations incorporating patient samples and combining clinical data with the mouse model findings will be pivotal in determining the extent of hepatic dysfunctions in DM1. This will help ensure the applicability, optimization, and effectiveness of prospective treatments being developed to treat/manage various symptoms of this debilitating disease.
Methods
Animal models
Four mouse models were utilized in this study. First, the control animals used for the DM1 experiment, the ApoE-rtTA mice, were a mixed strain C57Bl6/SJL mouse line, with a single transgene containing a reverse tetracycline transactivator (rtTA), expressed under the liver-specific ApoE promoter26. Second, the “DM1 liver” line was also a mixed strain line, combining FVB background TRE-960i mice with the ApoE-rtTA mice. The resulting FVB/C57Bl6/SJL mice contained both the ApoE-rtTA transgene as well as tetracycline response element (TRE) driven truncated DMPK gene containing only the last five exons of human DMPK25. The DMPK construct also contained an elongated CUG repeat sequence with 960 repeats. These repeats are interrupted every twenty repeats with a “ctcga” sequence to prevent the overall repeat sequence from undergoing expansion or shrinkage. The ApoE-rtTA and DM1 liver mice were maintained as homozygotes for all transgenic alleles.
HSA L/R mice were FVB mice expressing a truncated human skeletal actin (HSA) with an ∼240 CUG repeat sequence in the 3’ UTR of the transgene8. The HSA was driven by the skeletal actin promoter, allowing the exclusive expression of CUG repeats within the skeletal muscle tissue. The HSA L/R mice were maintained as homozygotes.
The final model, the Mbnl1 knock out (KO) or Mbnl1∆E3/∆E3 line, was an FVB mouse where the first coding exon of Mbnl1 was replaced with a cassette using cre-lox insertion40. This mouse line was maintained in the heterozygous state, and homozygous mutant (Mbnl1∆E3/∆E3) or homozygous wild-type (Mbnl1Wt/Wt) were generated for study or as controls when needed.
Mouse handling and care
National Institutes of Health (NIH) and University of Illinois, Urbana-Champaign (UIUC) institutional guidelines were followed in using and caring for laboratory animals. All experimental protocols were performed as approved by the UIUC Institutional Animal Care and Use Committee (IACUC). Specifically, mice were housed in a mouse facility maintain at a standard 12-hour-light/dark cycle, at 18–23 °C ambient temperature, 40–60% humidity. Mice were allowed ad libitum access to normal chow diet (2918 Envigo Teklad) or indicated experimental diet(s) and water. The study is not gender-specific, and specimens include both male and female animals; however, as attributes such as glucose regulation and body weight are sex-specific, animal sex was recorded. Euthanization was performed according to institutional guidelines with carbon dioxide inhalation or by cervical dislocation, under general anesthesia induced by isoflurane inhalation. Whole liver tissues and hepatocytes were subsequently isolated from anaesthetized mice.
Mouse diet schemes
In most experiments, DM1 liver and ApoE-rtTA mice were subjected to the disease-inducing Dox diet from birth, with mothers receiving 2.0 g/kg Dox-supplemented Teklad 2018 18% protein global rodent diet until weaning at 21 days. Subsequently, weaned mice transitioned to a 0.1 g/kg Dox diet until sacrifice at 9 weeks. This feeding protocol is referred to as the chronic DM1 liver model.
Exceptions to the protocol above include DM1 liver mice used for RNA-seq, which remained on a 2 g/kg Dox diet until sacrifice. “No-Dox” mice were ApoE-rtTA; TRE960i mice maintained on a Dox-free diet. “Recovery” mice followed the chronic DM1 protocol but were transitioned to a Dox-free diet for ten days before sacrifice. For the Western Diet model, mothers were on 2 g/kg Dox-supplemented chow until weaning. Then, mice were switched to a high-fat, high-sugar, cholesterol-supplemented “Western” diet (Teklad 88137) supplemented with 0.1 g/kg Dox for eight weeks. In the acute DM1 model, mice were Dox-free until eight weeks, followed by a 0.1 g/kg Dox diet for 12 or 18 days before sacrifice.
Sacrifice and tissue collection
Mice were fed freely until 4–6 hours before sacrifice and tissue harvest. Fasting started between the hours of 7–10 am, and tissue harvest occurred between 12–4 pm for all tissues and experiments, except for APAP-challenged mice, which occurred 24 hours post administration of APAP, typically between 8 am and 12 pm. Liver and carcass weight were taken at the time of sacrifice, as well as 600 µL of blood via retro-orbital bleeding. During sacrificing, liver tissues were collected for (1) RNA, protein, and lipid isolation, (2) paraffin-embedding, and (3) cryo-sectioning. Tissue for cryo-sectioning was collected by sectioning two pieces of liver, embedding the tissues in Optimal Cutting Temperature (OCT) compound and frozen on dry ice. Tissues for paraffin embedding were stored in neutral buffered formalin for 48 hours before being stored in 70% ethanol until paraffin embedding. The remaining tissue was flash-frozen in liquid nitrogen.
Isolation of hepatocytes
Hepatocyte isolation was performed using two-step perfusion with centrifugal separation to produce a cell population highly enriched in hepatocytes; this population was then used for RNA-seq analysis. The method for hepatocyte isolation was adapted from98. Briefly, mice were anesthetized in a chamber supplied with isoflurane and oxygen (2.5% isoflurane in oxygen, 1.5 L/min.). Mice were maintained on the anesthetic during the procedure using a nose cone. The liver was perfused via cannulation of the portal vein with 30–40 mL of a 1× HBSS (Hank’s balanced salt solution) with phenol red (without Ca2+and Mg2+), 0.5 mM EDTA solution at a flow rate between 3 and 5 mL per minute. This solution was followed by 50 mL of a 1× HBSS (with Ca2+), 5.4 mM CaCl2, 0.04 mg mL−1 soybean trypsin inhibitor, and 3000 units of collagenase type I (Worthington Chemicals). Subsequently, the liver was massaged in a Petri dish containing 1× HBSS with phenol red (without Ca2+ and Mg2+) to release cells from the liver capsule, and then the cell suspension was passed through a 70-μm filter to obtain a single-cell suspension. The cells were centrifuged at 50 × g for 5 min (4 °C) to separate live hepatocytes from non-parenchymal and dead cells. The cells were washed three times in 1× HBSS as above, then flash-frozen in liquid nitrogen and stored at −80 °C until further use.
Glucose tolerance testing (GTT)
GTT was performed on male and female mice, either seven days before sacrifice if they were maintained on the 0.1 g/kg Dox diet or 10 and 5 days before sacrifice if they were on the Western diet. GTT was performed after the mice were fasted for 24 hours for both cases. Glucose was injected through intraperitoneal injection (IP) at a 2 g/kg body weight concentration. Tails were clipped, blood was collected, and glucose was measured using a One Touch Ultra 2 glucose meter after 0, 15, 30, 60, 90, and 120 minutes.
Zoxazolamine recovery testing
Zoxazolamine (Zox) solution, prepared the day before, consisted of Zox dissolved in DMSO to achieve a final concentration of 15 µg/µL in a 95% Corn-Oil, 5% DMSO solution. After an 18–22 hour fast, mice received 120 mg/kg Zox injections. Vigorous homogenization between injections ensured a uniform solution. Post-treatment, mice freely roamed until they lost motor function, after which they were placed supine on an insulating blanket. Time was recorded until mice successfully self-righted three times99.
APAP insult testing
Acetaminophen (APAP) solutions were freshly prepared, dissolving 20 mg of APAP in 1 mL sterile 1× PBS. After heating at 55 °C for 15 minutes with periodic vortexing, the solution was maintained at 40 °C during injection, with thorough mixing between injections. Mice fasted for 18–22 hours and received a 350 mg/kg IP injection of APAP. Mice were observed for 8 hours when surviving mice returned to the animal care facility. Harvesting of serum and liver samples occurred 24 hours post-APAP injection100,101.
Acc1 exon 28 ASO treatment
Acc1 Exon 28 ASO, developed by Gene Tools, LLC, targeted the 5’ splicing site of Acc1 exon 28 (sequence: CCCTCTGTAATTAAA). A standard control in-vivo morpholino with the sequence CCTCTTACCTCAGTT served as a control. ASOs were dissolved in sterile biology-grade water. Mice designated for ASO treatment followed the chronic DM1 model until day 63, receiving IP injections of 12 mg ASO per kg body weight on days 63, 64, 68, and 72. Mice were sacrificed after a 4–6 hour fast on day 73.
ACC1 inhibitor treatment
ACC1 inhibitor CP-640186, procured from MedChemExpress (Lot#12439), was dissolved in a 5% (w/w) methylcellulose solution in molecular biology-grade water. Mice selected for ACC1 inhibitor treatment followed the chronic DM1 model until day 63. They received doses of either 25 µg inhibitor or vehicle (5% methylcellulose) per g body weight twice daily via oral gavage for five days. Sacrifice occurred on the sixth day after 4–6 hours of fasting.
Total RNA isolation and cDNA synthesis
Total RNA was isolated from the liver or perfused hepatocytes via TRIzol extraction. One milliliter of TRIzol (Invitrogen) was added to small pieces (approximately 30 mg to 50 mg) of either snap-frozen liver or hepatocyte isolate. Lysing of the cells was hastened by homogenization via bullet blending with NextAdvanced Zirconium oxide 1 mm beads. Chloroform was added, and the mixture was centrifuged for 10 minutes at 10,000xg, which caused a separation of layers. The aqueous layer containing extracted RNA was removed, and to this was added 600 µL of isopropanol. The solution was mixed and stored at −20 °C overnight (or for at least 8 hours). Afterward, the mixture was centrifuged for 40 min at 12,000xg, causing the precipitation of the RNA. The chloroform was removed, and the RNA pellet was washed with 70% ethanol. The RNA was then dissolved in water. RNA purity and concentration were assessed with a Biotek Synergy 2 UV spectrometer. cDNA synthesis was performed on 1 μg of RNA, using random hexamer primers and Maxima Reverse Transcriptase (Thermo Fisher Scientific) following manufacturer protocol. The cDNA was diluted to a total volume of 200 μL.
RNA-seq
RNA was isolated from hepatocytes using an RNeasy tissue mini-kit (Qiagen). Before library preparation, RNA quality was assessed using an Agilent Bioanalyzer by the Functional Genomics Core at the Roy J. Carver Biotechnology Center, UIUC. Poly-A selected, RNA-seq libraries preparation and 150-bp paired-end Illumina sequencing were performed on a NOVASEQ 6000 at the High Throughput Sequencing and Genotyping Unit, UIUC. RNA-seq reads were processed for quality and read length filters using Trimmomatic (version 0.39). RNA-seq reads were further aligned to the mouse genome (mm10) using STAR (version 2.5.2).
Gene expression levels were determined as TPM using count and differential expression values obtained from DESeq2 (version 1.8.2), Htseq (version 0.6.1), and Cuffdiff 2 (version 2.2.1)102,103,104. Genes were considered as having significant differential expression following imposed cutoff clearance (FDR ≤ 0.05, |log2(fold change)| ≥1). Differential splicing analysis was performed using rMATS (version 3.2.5), and significant events were identified using imposed cutoffs (FDR ≤ 0.10, junction read counts ≥10, PSI ≥ 10%, FPKM ≥ 1)105. Motif analysis for differentially spliced exons was performed using rMAPS with default parameters and putative motifs as described previously106. To perform alternative polyadenylation (APA) analysis, 3’UTR expression quantification was performed via Salmon (version 1.0.0), followed by analysis via qAPA (version 1.2.2)107,108. APA events were determined as significant if a change of 5 TPM or greater. Gene ontology analysis was performed using DAVID (version 6.8)37,109,110,111.
To perform the exon ontology analysis, exons undergoing significant changes in splicing were converted to corresponding human exons in the hg19 annotation using UCSC liftover with a minimum ratio of bases matching 0.8112. Additionally, the exons in hg19 reported by UCSC liftover were checked for gene identity corresponding to the mouse exon’s parent gene. These exons were then analyzed for ontology using the exon ontology and FasterDB packages113.
RT-PCR splice assays
Target events were amplified via PCR using the primers listed in Supplemental Table 1. The PCR cycle had a melting temperature of 95 °C, an annealing temperature of 55 °C, and an elongation temperature of 72°C. The PCR product was resolved down a 5.5% PAGE gel, stained with ethidium bromide, and imaged using a Bio-Rad Gel Doc machine.
Bio-Rad Image Lab Software (version 6.0.1) was used to measure the intensity of the bands appearing on the gel to quantify the percent splicing change. Splicing change is reported as Percent Spliced In, a ratio of the intensity of the upper band (containing the alternatively spliced exon) to the combined intensity of the upper and lower bands (the lower band does not include the alternatively spliced exon).
q-RT-PCR
Relative gene expression analysis was performed with ∼50 ng of cDNA per reaction using a SYBR® Green™ assay for Real-Time Quantitative Reverse Transcription PCR (qRT-PCR). cDNA, primer, SYBR, and water were mixed in a qPCR plate. The qPCR reaction was performed using a QuantStudio 3 Real-Time PCR System (Thermo Fisher). Standard cycling conditions for a SYBR® GreenTM based ΔΔCT assay with a reduced elongation step of 35 seconds were used. β-actin was used as a loading control unless otherwise noted. Relative gene expression is listed as relative to β-actin for quantification of the DT960i Primers used for the qPCR reactions are listed in Supplemental Table 1.
RNA-FISH and immunofluorescence (IF)
At sacrificing, pieces of liver were placed within the Tissue-Tek OCT (Optimal Cutting Temperature) Compound and frozen with dry ice. These frozen blocks were sectioned at 10 µm with a Leica CM3050 S cryostat at the Carl R. Woese Institute for Genomic Biology (UIUC).
RNA-FISH/IF was performed on these cryosections. Sections were washed in 1× PBS and then fixed with 10% NBF for 15–30 min. Slides were washed with 1× PBS, permeabilized with 0.5% Triton-X in 1× PBS for 10 minutes, washed with 1× SSC, and then washed with 30% Formamide in 2× SSC. FISH probe was then applied (the solution contained 2 μg/mL BSA, 66 μg/mL yeast tRNA, and 1 ng/μL Cy5-(CAG)10 (Integrated DNA Technologies) dissolved in 30% formamide in 2× SSC). After incubation at 37°C for two hours, sections were washed with 30% formamide in 2× SSC (for 30 minutes) and washed twice with 1× SSC. The slides were again fixed with 10% NBF (for 10 min), washed with 1× TBS, and re-permeabilized with 0.5% Triton-X in PBS (again for 10 min). Slides were washed in 1× TBS before being blocked in 10% normal goat serum with 1% BSA in 1× TBS for two hours (at room temperature). After blocking, the slides were drained with a vacuum trap and incubated in 1:500 primary antibody (anti-Mbnl1 [sc-47740] or anti-Mbnl2 [sc-136167] from Santa Cruz Biotechnology) in 1× TBS with 1% BSA at 4°C overnight.
The next day, the slides were washed in 1× TBS with 0.05% Triton-X and then washed in 1× TBS. Then, they were incubated in 1:500 secondary antibody (anti-Mouse IgG conjugated to DyLight 488 [AB_1965946] from Thermo Fisher Scientific) in 1× TBS for 1 hour at room temperature. Slides were washed with TBS with 0.05% Triton-X, washed with 1× PBS, stained with NucBlue (Invitrogen) in 1× PBS for 20 minutes, and washed with 1× PBS. Slides were imaged on a Zeiss LSM 710 microscope at the Carl R. Woese Institute for Genomic Biology at UIUC. A list of antibodies used is provided in Supplemental Table 2.
Histology: hematoxylin and eosin
To perform hematoxylin and eosin staining, paraffin-embedded tissues were sectioned into 5 µm thick sections and then deparaffinized with three xylene washes. The slides were rehydrated in ethanol solutions of decreasing concentration (100%, 95%, 80%, and 50%) before being placed in water. The slides were stained with Hematoxylin 7211 for 1.5 to 2 minutes and washed in water. Slides were blued in a 2% sodium bicarbonate and 0.2% magnesium sulfate bluing solution before being soaked in water again. After being placed in an ethanol solution, the slides were stained with eosin for 15–20 seconds, washed with ethanol, and dehydrated with xylene. They were finally coverslipped with Permount. All slides were imaged on a Hamamatsu Nanozoomer at the Carl R. Woese Institute for Genomic Biology at UIUC.
Histology: Oil Red O. staining
Oil red O. stain was performed on cryosections (reference “RNA-FISH and Immunofluorescence (IF)” for details on the preparation of cryosections). The cryosections were first fixed in 10% NBF and then hydrated in 1× PBS. After placing the slides in 60% isopropanol, the slides were stained in fresh Oil Red O. solution (consisting of Oil Red O. dye dissolved in 60% isopropanol). The slides were then washed with 60% isopropanol, counterstained briefly in Hematoxylin 7211, and washed with tap water. The slides were then mounted with CC mount. The slides were imaged on a Hamamatsu Nanozoomer at the Carl R. Woese Institute for Genomic Biology at UIUC.
To quantitatively measure the accumulation of lipids, Oil Red O. Images at ×10 zoom (∼1.75 × 1 mm), a minimum of 3 per sample, were collected and then analyzed by measuring the relative volume of red channel compared to blue (using standard RGB-based image splitting) using a pipeline on Cell Profiler114,115.
Serum alanine aminotransferase (ALT) and aspartate aminotransferase (AST) testing
Whole blood from mice was collected via retro-orbital puncture in Capiject gel/clot activator tubes, centrifuged for 3 min at 3000 × g, and then stored at −80 °C till further analysis. ALT and AST analysis is performed using commercial Thermo Scientific serum chemistry kits. Measurements were made in duplicate.
Western blot and antibodies
Approximately 50 mg of liver tissue was homogenized via bullet blending and lysed via sonication in RIPA lysis buffer (1× SDS, 1% v/v phosphatase inhibitor cocktail 3 (Sigma-Aldrich), 1× Pierce protease inhibitor mini tablets (Thermo Scientific). Samples were then separated with SDS-PAGE and transferred overnight in Towbin buffer with 0.5% SDS. Blots were then visualized by ECL after incubating with primary and secondary antibodies. All primary antibodies were diluted in AdvanBlock-Chemi Blocking solution from Advansta, and secondary antibodies were diluted in TBST (0.1% v/v Tween-20). See Supplementary table 4 for a list of antibodies and suppliers.
Jess Simple Western Immunoassay
ACC1 protein levels were quantified via Jess Automated Western Blot System (ProteinSimple Bio-techne; Cat# 004-650) according to manufacturer protocols. Briefly, protein samples were diluted to 0.04–0.08 mg/mL in Sample Buffer, added to a master mix containing dithiothreitol (DTT) and fluorescent molecular weight marker (EZ Standard Pack, 66–440 kDa; ProteinSimple), and heat-denatured at 95 °C for 5 minutes. The samples, blocking reagent (Antibody Diluent 2; ProteinSimple), ACC1-specific primary antibody (Cell Signaling Technology; Cat# 4190), anti-rabbit secondary HRP antibody, chemiluminescent substrates, as well as Total Protein Labeling Reagent (ProteinSimple; Part# DM-TP01) for total protein normalization, were added to a specialized 25-well plate. Size-based separation via capillary electrophoresis, immobilization, immunoprobing, imaging, and quantification were performed in the Jess instrument. Jess data were analyzed in the Compass for Simple Western software. For each sample, the area of ACC1 protein peak was normalized to the total protein signal in 66–440 kDa range and graphed as a ratio of ACC1 to Total Protein (ACC1/TP).
Lipid extraction and analysis
Twenty to fifty milligrams of liver tissue were weighed into locking cap 1.5 mL Eppendorf tubes. To this was added 600 µL chilled 3:2 hexanes/isopropanol solution and 6–10 NextAdvanced zirconium oxide 1 mm beads. The tubes were homogenized on a bullet blender for 30 seconds and then allowed to rest on ice for 2 minutes. This homogenization process was repeated five times before samples were spun down at 3000 × g for 10 minutes at 4°C. The liquid phase was then collected into a second Eppendorf tube and set aside. An additional 600 µL chilled 3:2 hexanes/isopropanol solution was then used to break up the pellet, and samples were rested on ice for 15 min with periodic vortexing. Again, samples were centrifuged at 3000 × g for 10 min, and the liquid phase was then combined with the liquid set aside previously. Samples were then allowed to dry in the open air overnight.
Samples were diluted in 200 µL of 1× PBS with 2% Triton X-100. Samples were vortexed and allowed to dissolve at 4°C; further dilution would occur if necessary. Once dissolved, samples were analyzed using an Infinity triglycerides colorimetric kit.
Statistical analysis and data visualization
All quantitative experiments have at least three independent biological repeats. The results were expressed with mean and standard deviation unless mentioned otherwise. Differences between groups were examined for statistical significance using unpaired T-tests when comparing directly between groups or one-way analysis of variance for more than two groups using the GraphPad Prism 9 Software. Statistical outliers were determined with the ROUT method in Prism, with Q = 5%. P-value < 0.05 or FDR < 0.10 was considered significant. RNA-seq data plots were generated in R using the ggplot2 package. In all figures, significance was set as p < 0.05, “*” indicates p < 0.05, “**” indicates p < 0.01, “***” indicates p < 0.001, and “****” indicates p < 0.0001. Data is presented as a bar graph or linear. For Box plots, the median is represented as the center line, median; box limits are the upper and lower quartiles; whiskers are set as minima and maxima.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA-Seq data that support the findings of this study have been deposited in NCBI Gene Expression Omnibus under the primary accession code GSE252827. The processed RNA-Seq data generated in this study are provided in the Supplementary Information/Source Data file. Source data are provided in this paper.
Materials availability
Requests for reagents, resources, and additional information should be directed to the corresponding author, Auinash Kalsotra (kalsotra@illinois.edu).
References
Harper, P. S. Myotonic dystrophy. (W.B. Saunders, London, New York, 2003).
Ashizawa, T. et al. Consensus-based care recommendations for adults with myotonic dystrophy type 1. Neurol. Clin. Pract. 8, 507–520 (2018).
Johnson, N. E. et al. Population-based prevalence of myotonic dystrophy type 1 using genetic analysis of statewide blood screening program. Neurology 96, e1045–e1053 (2021).
Heatwole, C. et al. Patient-reported impact of symptoms in myotonic dystrophy type 1 (PRISM-1). Neurology 79, 348–357 (2012).
Day, J. W. & Ranum, L. P. W. Genetics and molecular pathogenesis of the myotonic dystrophies. Curr. Neurol. Neurosci. Rep. 5, 55–60 (2005).
Brook, J. D. et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 68, 799–808 (1992).
Lee, J. E. & Cooper, T. A. Pathogenic mechanisms of myotonic dystrophy. Biochem. Soc. Trans. 37, 1281–1286 (2009).
Mankodi, A. Myotonic dystrophy in transgenic mice expressing an expanded CUG repeat. Science 289, 1769–1772 (2000).
Miller, J. W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J 19, 4439–4448 (2002).
Jiang, H., Mankodi, A., Swanson, M. S., Moxley, R. T. & Thornton, C. A. Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum. Mol. Genet. 13, 3079–3088 (2004).
Wang, E. T. et al. Transcriptome-wide regulation of pre-mRNA splicing and mRNA localization by muscleblind proteins. Cell 150, 710–724 (2012).
Lin, X. et al. Failure of MBNL1-dependent post-natal splicing transitions in myotonic dystrophy. Hum. Mol. Genet. 15, 2087–2097 (2006).
Chau, A. & Kalsotra, A. Developmental insights into the pathology of and therapeutic strategies for DM1: Back to the basics. Dev. Dyn. 244, 377–390 (2015).
Ho, T. H. et al. Muscleblind proteins regulate alternative splicing. EMBO J. 23, 3103–3112 (2004).
Kalsotra, A. et al. A postnatal switch of CELF and MBNL proteins reprograms alternative splicing in the developing heart. Proc. Natl. Acad. Sci. 105, 20333–20338 (2008).
Kalsotra, A. et al. The Mef2 transcription network is disrupted in myotonic dystrophy heart tissue, dramatically altering miRNA and mRNA expression. Cell Rep 6, 336–345 (2014).
Philips, A. V., Timchenko, L. T. & Cooper, T. A. Disruption of splicing regulated by a CUG-binding protein in myotonic dystrophy. Science 280, 737–741 (1998).
Yum, K., Wang, E. T. & Kalsotra, A. Myotonic dystrophy: disease repeat range, penetrance, age of onset, and relationship between repeat size and phenotypes. Curr. Opin. Genet. Dev. 44, 30–37 (2017).
Campbell, N., Brandom, B., Day, J. W. & Moxley, R. Practical suggestions for the anesthetic management of a myotonic dystrophy patient FOREWORD: 1, 1–6 (2010).
Mathieu, J. et al. Anesthetic and surgical complications in 219 cases of myotonic dystrophy. Neurology 49, 1646 LP–1650 (1997).
Bhardwaj, R. R. & Duchini, A. Non-alcoholic steatohepatitis in myotonic dystrophy: DMPK gene mutation, insulin resistance and development of steatohepatitis. Case Rep. Gastroenterol. 4, 100–103 (2010).
Shieh, K., Gilchrist, J. M. & Promrat, K. Frequency and predictors of nonalcoholic fatty liver disease in myotonic dystrophy. Muscle Nerve 41, 197–201 (2010).
John Herbick, C. J. Hepatic glycogenosis in a patient with type 1 myotonic dystrophy. J. Liver Dis. Transplant. 02, 2012–2014 (2013).
Achiron, A. et al. Abnormal liver test results in myotonic dystrophy. J. Clin. Gastroenterol. 26, 292–295 (1998).
Morriss, G. R., Rajapakshe, K., Huang, S., Coarfa, C. & Cooper, T. A. Mechanisms of skeletal muscle wasting in a mouse model for myotonic dystrophy type 1. Hum. Mol. Genet. 27, 2789–2804 (2018).
Dong, J. et al. Elucidation of a universal size-control mechanism in drosophila and mammals. Cell 130, 1120–1133 (2007).
Lee, J. et al. Intrinsically cell-penetrating multivalent and multitargeting ligands for myotonic dystrophy type 1. Proc. Natl. Acad. Sci. 116, 8709–8714 (2019).
Machuca-Tzili, L., Brook, D. & Hilton-Jones, D. Clinical and molecular aspects of the myotonic dystrophies: a review. Muscle Nerve 32, 1–18 (2005).
Wheeler, T. M. & Thornton, C. A. Myotonic dystrophy: RNA-mediated muscle disease. Curr. Opin. Neurol. 20, 572–576 (2007).
Scotti, M. M. & Swanson, M. S. RNA mis-splicing in disease. Nat. Rev. Genet. 17, 19–32 (2016).
Misra, C. et al. Aberrant expression of a non-muscle RBFOX2 isoform triggers cardiac conduction defects in myotonic dystrophy. Dev. Cell 52, 748–763.e6 (2020).
Chembazhi, U. V., Bangru, S., Hernaez, M. & Kalsotra, A. Cellular plasticity balances the metabolic and proliferation dynamics of a regenerating liver. Genome Res. 31, 576–591 (2021).
Nalavade, R., Griesche, N., Ryan, D. P., Hildebrand, S. & Krauß, S. Mechanisms of RNA-induced toxicity in CAG repeat disorders. Cell Death Amp. Dis. 4, e752–e752 (2013).
Wang, P.-Y., Chang, K.-T., Lin, Y.-M., Kuo, T.-Y. & Wang, G.-S. Ubiquitination of MBNL1 is required for its cytoplasmic localization and function in promoting neurite outgrowth. Cell Rep. 22, 2294–2306 (2018).
Gates, D. P., Coonrod, L. A. & Berglund, J. A. Autoregulated Splicing of <em>muscleblind</em>-like 1 (<em>MBNL1</em>) Pre-mRNA *. J. Biol. Chem. 286, 34224–34233 (2011).
Kino, Y. et al. Nuclear localization of MBNL1: splicing-mediated autoregulation and repression of repeat-derived aberrant proteins. Hum. Mol. Genet. 24, 740–756 (2015).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Sobczak, K. et al. Loss of MBNL leads to disruption of developmentally regulated alternative polyadenylation in RNA-mediated disease. Mol. Cell 56, 311–322 (2014).
Wang, E. T. et al. Transcriptome alterations in myotonic dystrophy skeletal muscle and heart. Hum. Mol. Genet. 28, 1312–1321 (2019).
Kanadia, R. N. et al. A muscleblind knockout model for myotonic dystrophy. Science 302, 1978–1980 (2003).
Bangru, S. et al. Alternative splicing rewires Hippo signaling pathway in hepatocytes to promote liver regeneration. Nat. Struct. Mol. Biol. 25, 928–939 (2018).
Nitschke, L., Hu, R.-C., Miller, A. N., Lucas, L. & Cooper, T. A. Alternative splicing mediates the compensatory upregulation of MBNL2 upon MBNL1 loss-of-function. Nucleic Acids Res. 51, gkac1219 (2023).
Rogalska, Z. & Sobczak, K. Sustainable recovery of MBNL activity in autoregulatory feedback loop in myotonic dystrophy. Mol. Ther. Nucleic Acids 30, 438–448 (2022).
Veyckemans, F. & Scholtes, J. L. Myotonic Dystrophies type 1 and 2: anesthetic care. Paediatr. Anaesth. 23, 794–803 (2013).
Groh, W. J. et al. Electrocardiographic abnormalities and sudden death in myotonic dystrophy type 1. N. Engl. J. Med. 358, 2688–2697 (2008).
James, L. P., Mayeux, P. R. & Hinson, J. A. Acetaminophen-induced hepatotoxicity. Drug Metab. Dispos. 31, 1499–506 (2003).
Zaher, H. et al. Protection against acetaminophen toxicity in CYP1A2 and CYP2E1 double-null mice. Toxicol. Appl. Pharmacol. 152, 193–199 (1998).
Mcclain, C. J. Potentiation of acetaminophen hepatotoxicity by alcohol. J. Am. Med. Assoc. 244, 251 (1980).
Senggunprai, L., Yoshinari, K. & Yamazoe, Y. Selective role of sulfotransferase 2A1 (SULT2A1) in the N-sulfoconjugation of quinolone drugs in humans. Drug Metab. Dispos. 37, 1711 LP–1717 (2009).
Gamage, N. et al. Human sulfotransferases and their role in chemical metabolism. Toxicol. Sci. 90, 5–22 (2006).
Bao, Y. et al. Alterations of cytochrome P450–mediated drug metabolism during liver repair and regeneration after acetaminophen-induced liver injury in mices. Drug Metab. Dispos. 50, 694–703 (2022).
Turner, C. & Hilton-Jones, D. The myotonic dystrophies: diagnosis and management. J. Neurol. Neurosurg. Psychiatry 81, 358–367 (2010).
Savkur, R. S., Philips, A. V. & Cooper, T. A. Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat. Genet. 29, 40–47 (2001).
Escolà-Gil, J. C. et al. The cholesterol content of western diets plays a major role in the paradoxical increase in high-density lipoprotein cholesterol and upregulates the macrophage reverse cholesterol transport pathway. Arterioscler. Thromb. Vasc. Biol. 31, 2493–2499 (2011).
Elmgren, A. et al. Acutely reduced locomotor activity is a major contributor to western diet-induced obesity in mice. Am. J. Physiol. Endocrinol. Metab. 294, E251–E260 (2007).
Millar, J. S., Cromley, D. A., McCoy, M. G., Rader, D. J. & Billheimer, J. T. Determining hepatic triglyceride production in mice: comparison of poloxamer 407 with Triton WR-1339. J. Lipid Res. 46, 2023–2028 (2005).
Manual, P. Serum triglyceride quantification kit. https://www.cellbiolabs.com/sites/default/files/STA-396-serum-triglyceride-quantification-kit-colorimetric.pdf (Colorimetric).
Arif, W. et al. Splicing factor SRSF1 deficiency in the liver triggers NASH-like pathology and cell death. Nat. Commun. 14, 551 (2023).
Ha, J., Daniel, S., Broyles, S. S. & Kim, K. H. Critical phosphorylation sites for acetyl-CoA carboxylase activity. J. Biol. Chem. 269, 22162–22168 (1994).
Halestrap, A. P. & Denton, R. M. Hormonal regulation of adipose-tissue acetyl-coenzyme A carboxylase by changes in the polymeric state of the enzyme. The role of long-chain fatty acyl-coenzyme A thioesters and citrate. Biochem. J. 142, 365–377 (1974).
Hunkeler, M. et al. Structural basis for regulation of human acetyl-CoA carboxylase. Nature 558, 470–474 (2018).
Munday, M. R., Campbell, D. G., Carling, D. & Hardie, D. G. Identification by amino acid sequencing of three major regulatory phosphorylation sites on rat acetyl-CoA carboxylase. Eur. J. Biochem. 175, 331–338 (1988).
Wei, J. et al. A unified molecular mechanism for the regulation of acetyl-CoA carboxylase by phosphorylation. Cell Discov. 2, 16044 (2016).
Witters, L. A., Kowaloff, E. M. & Avruch, J. Glucagon regulation of protein phosphorylation. Identification of acetyl coenzyme A carboxylase as a substrate. J. Biol. Chem. 254, 245–248 (1979).
Davies, S. P., Sim, A. T. R. & Hardie, D. G. Location and function of three sites phosphorylated on rat acetyl-CoA carboxylase by the AMP-activated protein kinase. Eur. J. Biochem. 187, 183–190 (1990).
Kong, I. S., López-Casillas, F. & Kim, K. H. Acetyl-CoA carboxylase mRNA species with or without inhibitory coding sequence for Ser-1200 phosphorylation. J. Biol. Chem. 265, 13695–13701 (1990).
Winder, W. W. et al. Phosphorylation of rat muscle acetyl-CoA carboxylase by AMP-activated protein kinase and protein kinase A. J. Appl. Physiol. 82, 219–225 (1997).
Barber, M., Pooley, L. & Travers, M. Developmental regulation of alternatively spliced acetyl-CoA carboxylase-alpha mRNAs encoding isozymes with or without an eight amino acid domain upstream of the Ser-1200 phosphorylation motif in the mammary gland. J. Mol. Endocrinol. 27, 349–356 (2001).
Badaoui, B. et al. Goat acetyl-coenzyme A carboxylase α: molecular characterization, polymorphism, and association with milk traits. J. Dairy Sci. 90, 1039–1043 (2007).
Harwood, H. J. J. et al. Isozyme-nonselective N-substituted bipiperidylcarboxamide acetyl-CoA carboxylase inhibitors reduce tissue malonyl-CoA concentrations, inhibit fatty acid synthesis, and increase fatty acid oxidation in cultured cells and in experimental animals. J. Biol. Chem. 278, 37099–37111 (2003).
Zhang, H., Tweel, B., Li, J. & Tong, L. Crystal structure of the carboxyltransferase domain of acetyl-coenzyme A carboxylase in complex with CP-640186. Structure 12, 1683–1691 (2004).
Du, H. et al. Aberrant alternative splicing and extracellular matrix gene expression in mouse models of myotonic dystrophy. Nat. Struct. Mol. Biol. 17, 187–193 (2010).
Perna, A. et al. High prevalence and gender-related differences of gastrointestinal manifestations in a cohort of DM1 patients: a perspective, cross-sectional study. Front. Neurol. 11, 394 (2020).
Miele, L. et al. Clinical characteristics of metabolic associated fatty liver disease (MAFLD) in subjects with myotonic dystrophy type 1 (DM1). Dig. Liver Dis. 53, 1451–1457 (2021).
Tanaka, N. et al. Non-alcoholic fatty liver disease later diagnosed as myotonic dystrophy. World J. Hepatol. 12, 685–692 (2020).
Hama, M. et al. Metabolic complications in myotonic dystrophy type 1: a cross-sectional survey using the National Registry of Japan. J. Neurol. Sci. 427, 117511 (2021).
Peter, R. et al. Hydroxylation of chlorzoxazone as a specific probe for human liver cytochrome P-450IIE1. Chem. Res. Toxicol. 3, 566–573 (1990).
Spracklin, D. K., Hankins, D. C., Fisher, J. M., Thummel, K. E. & Kharasch, E. D. Cytochrome P450 2E1 is the principal catalyst of human oxidative halothane metabolism in vitro. J. Pharmacol. Exp. Ther. 281, 400–411 (1997).
Gupta, N. et al. Myotonic dystrophy: an anaesthetic dilemma. Indian J. Anaesth. 53, 688–691 (2009).
Bhate, A. et al. ESRP2 controls an adult splicing programme in hepatocytes to support postnatal liver maturation. Nat. Commun. 6, 8768 (2015).
Konieczny, P., Stepniak-Konieczna, E. & Sobczak, K. MBNL proteins and their target RNAs, interaction and splicing regulation. Nucleic Acids Res. 42, 10873–10887 (2014).
Lee, K.-Y. et al. Compound loss of muscleblind-like function in myotonic dystrophy. EMBO Mol. Med. 5, 1887–1900 (2013).
Lee, S. R., Pratt, G. A., Martinez, F. J., Yeo, G. W. & Lykke-Andersen, J. Target discrimination in nonsense-mediated mRNA decay requires Upf1 ATPase activity. Mol. Cell 59, 413–425 (2015).
Sznajder, Ł. J. et al. Mechanistic determinants of MBNL activity. Nucleic Acids Res. 44, gkw915 (2016).
Konieczny, P., Stepniak-Konieczna, E. & Sobczak, K. MBNL expression in autoregulatory feedback loops. RNA Biol 15, 1–8 (2018).
Fullerton, M. D. et al. Single phosphorylation sites in Acc1 and Acc2 regulate lipid homeostasis and the insulin-sensitizing effects of metformin. Nat. Med. 19, 1649–1654 (2013).
Thampy, K. G. & Wakil, S. J. Regulation of acetyl-coenzyme A carboxylase. I. Purification and properties of two forms of acetyl-coenzyme A carboxylase from rat liver. J. Biol. Chem. 263, 6447–6453 (1988).
Softic, S. et al. Divergent effects of glucose and fructose on hepatic lipogenesis and insulin signaling. J. Clin. Invest. 127, 4059–4074 (2017).
Mao, J. et al. Liver-specific deletion of acetyl-CoA carboxylase 1 reduces hepatic triglyceride accumulation without affecting glucose homeostasis. Proc. Natl. Acad. Sci. 103, 8552–8557 (2006).
Goedeke, L. et al. Acetyl-CoA carboxylase inhibition reverses NAFLD and hepatic insulin resistance but promotes hypertriglyceridemia in rodents. Hepatology 68, 2197–2211 (2018).
Tamura, Y. O. et al. Selective acetyl-CoA carboxylase 1 inhibitor improves hepatic steatosis and hepatic fibrosis in a preclinical nonalcoholic steatohepatitis model. J. Pharmacol. Exp. Ther. 379, 280–289 (2021).
Zhang, X.-J., Cai, J. & Li, H. Targeting ACC for NASH resolution. Trends Mol. Med. 28, 5–7 (2022).
Zhang, X.-J. et al. A small molecule targeting ALOX12-ACC1 ameliorates nonalcoholic steatohepatitis in mice and macaques. Sci. Transl. Med. 13, eabg8116 (2021).
Bian, H., Liu, Y.-M. & Chen, Z.-N. New avenues for NASH therapy by targeting ACC. Cell Metab. 34, 191–193 (2022).
Kim, C.-W. et al. Acetyl CoA carboxylase inhibition reduces hepatic steatosis but elevates plasma triglycerides in mice and humans: a bedside to bench investigation. Cell Metab. 26, 576 (2017).
Calle, R. A. et al. ACC inhibitor alone or co-administered with a DGAT2 inhibitor in patients with non-alcoholic fatty liver disease: two parallel, placebo-controlled, randomized phase 2a trials. Nat. Med. 27, 1836–1848 (2021).
Loomba, R. et al. GS-0976 reduces hepatic steatosis and fibrosis markers in patients with nonalcoholic fatty liver disease. Gastroenterology 155, 1463–1473.e6 (2018).
Li, W.-C., Ralphs, K. L. & Tosh, D. Isolation and culture of adult mouse hepatocytes. In: Methods in Molecular Biology 185–196. https://doi.org/10.1007/978-1-59745-019-5_13. (Humana Press, 2010).
Fujii, K., Jaffe, H. & Epstein, S. S. Factors influencing the hexobarbital sleeping time and zoxazolamine paralysis time in mice. Toxicol. Appl. Pharmacol. 13, 431–438 (1968).
Mcgill, M. R., Williams, C. D., Xie, Y., Ramachandran, A. & Jaeschke, H. Acetaminophen-induced liver injury in rats and mice: Comparison of protein adducts, mitochondrial dysfunction, and oxidative stress in the mechanism of toxicity. Toxicol. Appl. Pharmacol. 264, 387–394 (2012).
Mossanen, J. & Tacke, F. Acetaminophen-induced acute liver injury in mice. Lab. Anim. 49, 30–6 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl. Acad. Sci. USA. 111, E5593–601 (2014).
Park, J. W., Jung, S., Rouchka, E. C., Tseng, Y.-T. & Xing, Y. rMAPS: RNA map analysis and plotting server for alternative exon regulation. Nucleic Acids Res. 44, W333–W338 (2016).
Ha, K. C. H., Blencowe, B. J. & Morris, Q. QAPA: a new method for the systematic analysis of alternative polyadenylation from RNA-seq data. Genome Biol. 19, 45 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Huang, D. W. et al. DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–W175 (2007).
Bonnot, T., Gillard, M. B. & Nagel, D. H. A simple protocol for informative visualization of enriched gene ontology terms. Bio-Protoc. 9, e3429 (2019).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800 (2011).
Kent, W. J. et al. The Human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Benoit-Pilven, C. et al. Complementarity of assembly-first and mapping-first approaches for alternative splicing annotation and differential analysis from RNAseq data. Sci. Rep. 8, 4307 (2018).
Kamentsky, L. et al. Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software. Bioinformatics 27, 1179–1180 (2011).
Mcquin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLOS Biol. 16, e2005970 (2018).
Acknowledgements
We acknowledge support from the Transgenic mouse core, High-throughput sequencing and genotyping core, and Histology-microscopy core facilities at the University of Illinois, Urbana-Champaign. We thank the members of the Kalsotra and Anakk laboratories for their valuable discussions and comments on the manuscript. Work in the Kalsotra laboratory is supported by the National Institute of Health (R01HL126845, R01AA010154), the Muscular Dystrophy Association (MDA1072487), the Chan-Zuckerberg Biohub Chicago Award, and the Beckman Fellowship from the Center for Advanced Study at the University of Illinois Urbana-Champaign. Z.D. was supported by the NIH Chemistry-Biology Interface training program (T32-GM070421); A.G. was supported by the Jenner Family Summer Research Fellowship; J.M.D. was supported by the Danica Dabich Fellowship in Biochemistry; and U.V.C. was supported by the Herbert E. Carter Fellowship in Biochemistry.
Author information
Authors and Affiliations
Contributions
Z.D. and A.K. conceived the project and designed the experiments. Z.D., O.A., H.B., J.M.D., and A.G. performed experiments and analyzed the data. Z.D., A.K., and U.V.C. facilitated mouse model development and management. Z.D. and A.K. interpreted the results and wrote the manuscript. All authors discussed the results and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
All mouse experiments were performed and reported according to the ARRIVE guidelines for reporting animal studies.
Peer review
Peer review information
Nature Communications thanks Nicholas Johnson, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dewald, Z., Adesanya, O., Bae, H. et al. Altered drug metabolism and increased susceptibility to fatty liver disease in a mouse model of myotonic dystrophy. Nat Commun 15, 9062 (2024). https://doi.org/10.1038/s41467-024-53378-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53378-z