Abstract
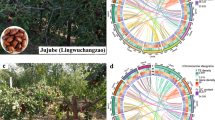
Jujube (Ziziphus jujuba Mill.), belonging to the Rhamnaceae family, is gaining increasing prominence as a perennial fruit crop with significant economic and medicinal values. Here, we conduct de novo assembly of four reference-grade genomes, encompassing one wild and three cultivated jujube accessions. We present insights into the population structure, genetic diversity, and genomic variations within a diverse collection of 1059 jujube accessions. Analyzes of the jujube pan-genome, based on our four assemblies and four previously released genomes, reveal extensive genomic variations within domestication-associated regions, potentially leading to the discovery of a candidate gene that regulates flowering and fruit ripening. By leveraging the pan-genome and a large-scale resequencing population, we identify two candidate genes involved in domestication traits, including the seed-setting rate, the bearing-shoot length and the leaf size in jujube. These genomic resources will accelerate evolutionary and functional genomics studies of jujube.
Similar content being viewed by others
Introduction
Jujube (Ziziphus jujuba Mill.), also known as Chinese date or red date, is gaining global popularity as a superfruit. Jujube is in the Rhamnaceae family and is renowned for its exceptional taste, nutritional richness (a notable source of vitamin C, cAMP, and sugar), resilience to various abiotic stresses, high economic value, and ecological friendliness1,2. A native plant of China, it originated in the middle and lower reaches of the Yellow River3. With a cultivation history spanning over 7000 years3,4, jujube has spread to nearly 50 countries across temperate to tropical regions on all five continents1,5,6.
Cultivated jujube underwent domestication from wild/sour jujube (Ziziphus acidojujuba C. Y. Cheng et M. J. Liu) through an extensive artificial selection process, which significantly altered its essential horticultural traits. These traits, including the fruit ripening period, the seed-setting rate (defined as the number of stones with plump seeds divided by the total number of detected stones), the bearing-shoot length and the leaf size, were intentionally modified during this prolonged process3,7,8,9. A recent study by Guo et al.9 highlighted that the majority of wild jujube plants exhibit earlier flowering and fruit ripening. The transition in the reproductive strategy of jujube represents a notable domestication event. Furthermore, in contrast to the prevalent seed propagation observed in wild jujube, which is characterized by one or two plump seeds within the stone, cultivated jujube predominantly employs clonal propagation. This method aligns with the propagation strategy employed by over 75% of perennial fruit trees10.
In jujube, leaves, prickles, flowers, and fruits all originate and grow on the bearing shoot, which germinates from the mother-bearing shoot in spring, is deciduous and typically drops before winter. This horticultural characteristic is rare among perennial tree plants, providing a distinctive model to understand shoot development and function1. Consequently, the bearing shoot is not only a crucial target trait for domestication, but also is a subject of scientific interest11. However, the causal genes associated with the above mentioned domestication traits remained poorly characterized, partly due to the absence of a suitable pan-genomic variation dataset.
The limitations of a single linear reference genome become evident when attempting to capture the entire spectrum of genetic diversity within a species. This approach faces challenges in identifying larger structural variants (SVs) such as presence/absence variants (PAVs), copy number variants (CNVs) and inversions, which are all known to play roles in controlling agronomical traits12,13,14,15,16,17,18,19,20. Notably, several plant pan-genomes have been constructed, including those for soybean16, rice17, tomato18,20,21, and citrus22. The utilization of graph-based pan-genome enables SV-based genome-wide association studies (GWASs) in plants, leading to the identification of numerous unreported quantitative trait loci (QTLs)17,20,23.
Jujube cultivars have traditionally been classified as fresh, dry, and dual-purpose. Genomic information on dual-purpose jujube accessions is limited. The popular elite cultivar ‘Huizao’ is a dual-purpose jujube accession with high-quality attributes and extensive cultivation24. Here, we show four reference-grade genomes, including that of ‘Huizao’ and three other accessions. Utilizing ‘Huizao’ as the reference genome, we explore the population structure and genetic diversity within a large-scale group comprising 1059 accessions. A pan-genome is constructed, encompassing our four assemblies in conjunction with four previously released genomes25,26,27. Subsequent analysis reveals a large number of genetic variations including hundreds of thousands of SVs. By integrating the pan-genomic variations and a large-scale resequencing atlas, we elucidate part of the genetic basis of domestication traits, particularly those related to the flowering and fruit ripening period, the seed-setting rate as well as the bearing-shoot length and leaf size. This research contributes a valuable genomic data resource and establishes a foundation for future basic research and improvement of jujube breeding.
Results
De novo genome assembly of jujube elite cultivar ‘Huizao’
To establish a high-quality reference genome and unravel the genomic characteristics of elite accession, we employed a comprehensive approach, utilizing Illumina sequencing, PacBio circular consensus sequencing (CCS), and high-throughput chromosome conformation capture (Hi-C) technology to generate chromosome-level genome assemblies for ‘Huizao’ (Individual code, Z95).
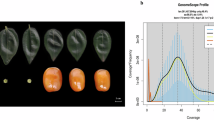
We estimated the Z95 genome size to be 411.64 Mb (GenemoScope analysis - Table 1 and Supplementary Fig. 1). Utilizing PacBio CCS technology, we generated 16.7 Gb of CCS reads, representing a sequencing depth of 42× (Supplementary Table 1). The Z95 genome was assembled using hifiasm28, employing the CCS data, resulting in a whole genome assembly of 395.06 Mb, with a contig N50 value of 20.05 Mb. Leveraging the 89× Hi-C data, we successfully anchored 96.3% of the assembled sequences to 12 chromosomes (Table 1 and Supplementary Fig. 2). The Z95 assembly exhibited a high level of intact long terminal repeats (LTRs), with an LTR assembly index (LAI) of 15.39 (Table 1), thus meeting the accepted threshold for qualification as a ‘reference’ genome (LAI > 10)23. Notably, 99.7% of the Illumina short reads were successfully mapped onto the corresponding assembled genomes. The completeness of the genome assembly was further confirmed through Benchmarking Universal Single-Copy Orthologs (BUSCO)29 evaluation, with a completeness score of 99.1% and 93.4% for complete single-copy genes (Table 1 and Supplementary Table 2).
Among the transposable elements (TEs), LTR/Gypsy repeat elements were the most abundant, accounting for 13.30%, followed by LTR/Copia at 7.97% (Supplementary Table 3). The four assembled genome was annotated using transcriptome data from different tissues, homology-based prediction, and ab initio prediction. This annotation process identified 34,061 protein-coding genes, achieving a BUSCO score of 91.6% (Table 1).
Population structure and genetic diversity of jujube
In this study, a diverse collection of 1059 jujube accessions was examined, encompassing 429 wild jujube individuals (Z. acidojujuba) and 630 jujube cultivars (Z. jujuba), and representing a broad range of jujube geographical distributions (Supplementary Fig. 3a and Supplementary Data 1). Among them, sequencing data for 562 accessions were generated in this study, while the data for the remaining 497 were sourced from previous studies1,9,11,25,27. The resequencing effort produced ~6.29 Gb of clean data per accession, achieving an average depth of 15.69× and 95.63% coverage of the Z95 reference genome (Supplementary Data 2). Upon mapping against the Z95 genome, we identified a total of 13,091,616 single nucleotide polymorphisms (SNPs), and 1,439,798 insertions/deletions (InDels) ( < 50 bp).
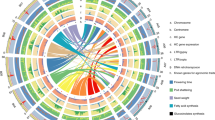
A phylogenetic analysis of the 1059 jujube accessions was conducted, utilizing 557,726 SNPs with three Indian jujube (Ziziphus mauritiana) accessions serving as the outgroup. The accessions were categorized into two major groups: wild and cultivated. Further subdivision of the cultivated group revealed five subgroups, which closely aligned with their geographical distributions (Fig. 1a, Supplementary Figs. 3b and 4). Cultivated subgroups I and III were predominantly composed of accessions from West China (west of the Taihang Mountains), while the other three cultivated subgroups consisted mainly of accessions from East China (east of the Taihang Mountains) (Supplementary Fig. 3b).
a Phylogenetic tree constructed among 1059 jujube accessions, with four de novo assembled genomes (indicated by black arrows) and four previously released genomes (red arrows) shown below the phylogenetic tree. b Population structure analysis conducted for all jujube accessions with different ancestry kinship (K = 2–6). Each vertical bar represents one accession, and the x axis displays the different groups. The y axis quantifies ancestry membership, with the orders and positions of all accessions on the x axis consistent with those in the phylogenetic tree. c Principal component analysis (PCA) plot illustrating the first two components (eigenvector 1 and 2) of all accessions. d Genome-wide decay of LD in the different groups. e Fruit morphology of four selected jujube accessions used for de novo genome assembly with bars indicating 1 cm. Abbreviation: C-sub, cultivated subgroup. Source data are provided as a Source Data file.
Subsequently, ADMIXTURE30 was employed to estimate ancestry proportions, and a principal component analysis (PCA) was conducted based on 6,185,881 SNPs. Consistent with the phylogenetic analysis, both approaches revealed a consistent pattern of six distinct clusters, including one wild and five cultivated groups (Fig. 1b, c). As expected, nucleotide diversity (π) was higher within the wild group (4.60 × 10−3) than in the five cultivated groups (average π = 3.55 × 10−3). The Dxy value (representing the mean number of nucleotide differences between samples in population X and population Y) between wild and cultivated groups was the highest compared with the other paired combinations (Supplementary Fig. 5). Additionally, we observed a rapid decay (0.25 kb) over physical distance in the wild group compared with that in the five cultivated subgroups, which ranged from 0.62 kb to 1.74 kb (Fig. 1d). These values are comparable to those reported for pear ( < 1 kb)31 and apple ( < 1 kb)32, but lower than that of peach ( ~ 35 kb for domesticated peach)33.
Characterization of a gene-based jujube pan-genome
In an effort expand the gene pool and explore the genetic diversity of jujube, we conducted a de novo assembly with three additional accessions chosen based on their phylogenetic relationships, phenotype diversity, cultivation area, and geographical distributions (Fig. 1a,e and Supplementary Data 1). Ultimately one wild accession (S21) and two cultivated accessions, namely ‘Jinsixiaozao’ (Z94) and ‘Goutouzao’ (Z203), were selected for further analysis (Fig. 1a, e and Table 1). Employing the same sequencing platform and assembly strategy used for Z95, three reference-grade genomes were assembled, exhibiting similar indicators to the Z95 genome (Table 1, Supplementary Figs. 1, 2 and Supplementary Table 2).
To create a gene-based pan-genome for jujube, we integrated data from our four de novo assemblies and four previously released genomes (‘Dongzao’26, ‘Junzao’27, and two wild accessions S202125 and S202427). The number of gene families increased significantly as the number of genomes increased from two to six, and then showed a modest increase from six to eight (Fig. 2a). Ortholog investigation assigned 241,216 (96.83%) genes from the eight jujube genomes into 32,567 gene families. Among these gene families, 11,414 (35.05%) were present in all eight genomes and were categorized as core genes, 20,707 (63.58%) were present in 2–7 genomes and were categorized as dispensable genes, and 446 (1.37%) gene families were present in only one genome, which were termed accession-specific genes (Fig. 2b, c, d). Notably, the gene-based pan-genome included 7801 gene families that are absent from the Z95 reference genome.
a Variation of gene families in the pan-genome and core genome with the increase in the number of jujube genomes. b Distribution of gene families across the eight genomes depicted in a histogram, showcasing varying frequencies. c The jujube pan-genome’s composition, detailing the proportions of core, dispensable and accession-specific gene families. d The proportion of core, dispensable, and accession-specific genes per genome. e Ka/Ks values for the dispensable and core genes illustrated through box plots. f Box plots representing FPKM values of dispensable and core genes per genome. The average FPKM value across all tissues for each gene is used here. In (a), (e), and (f), median values are denoted by center bold lines, while the box limits indicate upper and lower quartiles. Whiskers extend to data no more than 1.5× the interquartile range. In both (e) and (f), two-tailed Student’s t-tests were employed to identify significant differences. Sample size for each group is denoted in brackets. Source data are provided as a Source Data file.
Next, we computed the non-synonymous/synonymous substitution ratios (Ka/Ks) for the core and dispensable genes. The analysis revealed that the dispensable genes displayed higher Ka/Ks values compared with the core genes (Fig. 2e). These results suggest that the core genes evolved at a slower pace and are more functionally conserved. To further understand the functional significance, we conducted expression analysis using RNA-sequencing data for the four assembled accessions. The results showed much higher expression levels of the core genes compared with those of the dispensable genes (Fig. 2f), indicating that the core genes likely exert more crucial biological functions. These results, including Ka/Ks and expression analysis, are consistent with those reported for Arabidopsis and barley34.
Extensive genomic variations within the jujube pan-genome
To explore the genomic variations within the eight genomes, we performed alignments of the other seven genomes to the reference genome Z95 using MUMmer software35. In general, 2596,045–3,963,811 whole genome SNPs were identified, averaging 8.02 SNPs per kb (7.44 in cultivated accessions and 8.80 in wild accessions on average) across different jujube genomes. Among these SNPs, 95,392–148,492 were non-synonymous based on the annotation results of Z95 and 5329–8545 were predicted to be big-effect SNPs (causing changes in start codons, stop codons or splice sites) according to SnpEff software36 (Supplementary Table 4). In addition, 583,473–864,791 InDels were identified, averaging 1.86 InDels per kb (1.70 in cultivated accessions and 2.07 in wild accessions on average) across different jujube genomes. These InDels constituted a total of 1.12–1.62 Mb of sequences, with a mean of 1.33 Mb. ~20.69% of the InDels were found in genic regions, among which an average of 10,107 were predicted to be big-effect InDels (leading to frameshifts) (Supplementary Table 5).
The high-quality assemblies of these genomes present a valuable opportunity to identify SVs ( ≥ 50 bp). Comparisons of the other seven genomes to Z95 using MUMmer35, revealed a high level of collinearity (Supplementary Figs. 6 and 7) and 26,559–47,606 SVs were identified in each comparison using Syri software37 (Supplementary Table 6). Among these SVs, 1165–1947 were predicted to be big-effect SVs ( > 50% of coding region covered by SVs), which affected 2070–3839 annotated genes (Supplementary Table 6). Gene Ontology (GO) enrichment analyses revealed that the genes affected by big-effect SVs were enriched in biological processes related to peptide biosynthetic process, protein metabolic process, cellular metabolic process, and photosynthesis (Supplementary Fig. 8). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways analysis indicated that genes affected by big-effect SVs were enriched in pathways related to RNA polymerase, oxidative phosphorylation, ribosome, and metabolism (Supplementary Fig. 9). Overall, this dataset of genomic variations within the eight jujube genomes offers a rich resource for future studies of jujube trait biology and breeding practices.
Artificial selection of variations during jujube domestication
To enhance our understanding of the impact contributed by genomic variations during jujube domestication, we aggregated all cultivated sub-groups and compared the level of nucleotide diversity (π) with that of the wild group. This analysis identified 126 putative selective sweeps based on the πwild/πcultivated ratio (Supplementary Fig. 10 and Supplementary Data 3), covering 31.68 Mb (8.02%) of the reference genome and encompassing 2302 genes (Supplementary Data 4). Notably, some regions coincide with well-documented genes associated with domestication traits, such as ZjPOD1 related to reproductive system development9 (Supplementary Fig. 10).
To explore the effects of SVs during jujube domestication, we overlapped the putative swept regions with SVs obtained from the comparison between the wild accession S21 and the cultivated accession Z95. In total, we identified 4364 SVs within domestication regions, affecting 666 genes (Supplementary Data 5). Among these genes, several are potentially associated with the domestication syndrome, including increased fruit sweetness and the transition in reproductive strategy. In particular, a 1.7- kb insertion was found in the third exon of Z95_Ju00G026290 (Supplementary Fig. 11), the ortholog of AtEDA14 which regulates female gametophyte development in Arabidopsis38. This insertion resulted in an alteration in the number of exons between S21 and Z95 (Supplementary Fig. 11), leading to an alteration in the amino acid length (83 and 187 amino acid residues in Z95 and S21, respectively), thus potentially impacting the phenotype.
By exploiting the alignment results between S21 and Z95, we successfully detected 59,253 InDels within the sweep regions. Among them, 745 InDels were specifically situated in the exons of genes, impacting the protein-coding sequence of 356 annotated genes (Supplementary Data 6). Within the identified 356 genes, Z95_Ju00G026420 stood out (Fig. 3a). This gene encodes an agamous-like MADS-box protein AGL28 and features a 12- bp InDel in the sixth exon (Fig. 3b). This 12- bp insertion extends a disordered segment of ZjAGL28 (Supplementary Fig. 12), which might alter the protein’s binding ability to other molecules. Haplotype analysis centred on this InDel disclosed that the alternative allele predominated among wild accessions (73.1%), while the reference allele was prevalent in cultivated jujubes (Fig. 3c). Notably, in cultivated subgroup V, all accessions exhibited the reference allele (Fig. 3c), indicating a substantial inclination toward artificial selection for this InDel during the domestication of jujube.
a Identification of ZjAGL28 within the domestication region through π ratio analysis. b Gene structure of ZjAGL28 accompanied by genetic variation analysis in the pan-genome, revealing a 12- bp InDel located in the sixth exon. c Haplotype analysis of the InDel across all resequenced accessions, with accession numbers indicated below the pie charts. d Early flowering observed in Arabidopsis due to ectopic overexpression of ZjAGL28. Bar = 5 cm. e Statistics of days to flowering in WT and two OE lines with values expressed as mean ± SE (n = 10 plants). Differences between WT and OE lines were assessed using a two-tailed Student’s t-test. f Early ripening in Arabidopsis resulting from ZjAGL28 ectopic overexpression, demonstrated by the top shoots of WT and two OE lines. Bar = 2 cm. g Statistics of days to ripening in WT and two OE lines, with values expressed as mean ± SE (n = 10 plants). Differences between WT and OE lines were assessed using a two-tailed Student’s t-test. Abbreviations: Chr, chromosome; C-sub, cultivated subgroup. Source data are provided as a Source Data file.
To delve into the biological functions of Z95_Ju00G026420 (designated ZjAGL28), we conducted ectopic overexpression of ZjAGL28 in Arabidopsis thaliana and closely monitored any phenotypic changes throughout various developmental stages. Two overexpression (OE) lines were carefully selected and thoroughly characterized (Fig. 3d–g and Supplementary Fig. 13). During the vegetative growth stage, no significant morphological alterations were observed. However, during the reproductive growth stage, the OE lines exhibited earlier flowering compared with that of the wild-type (WT) (Fig. 3d, e). Furthermore, earlier ripening of siliques was noted in the two OE lines in comparison with that of the WT (Fig. 3f, g and Supplementary Fig. 13b). Collectively, these observations suggested that ZjAGL28 likely plays a positive regulatory role in flowering time and fruit ripening.
The pan-genome enables SV-based GWAS in jujube
To identify SVs associated with phenotypic variations, we conducted genotyping of 19,749 PAVs identified by Syri software37 across 1056 jujube accessions utilizing the Illumina short-read sequences. Subsequently, these genotyped PAVs were employed in SV-based GWAS for 16 horticultural traits, leading to the detection of 103 significantly associated SVs (Supplementary Table 7). One noteworthy finding was a 276- bp insertion on chromosome 01, which exhibited a substantial association with stone width (Fig. 4a, Supplementary Figs. 14a and 15a). Similarly, a 52- bp deletion on chromosome 10 was linked to fruit weight (Fig. 4b, Supplementary Figs. 14b and 15b). Furthermore, a 162- bp insertion on chromosome 04 (Fig. 4c, Supplementary Figs. 14c and 15c), was found to be significantly associated with the bearing-shoot length. Accessions with the alternative allele exhibited a notable decrease in the stone width, fruit weight, and bearing-shoot length (Fig. 4a, b, c). The identified SVs that are significantly associated with horticultural traits provide a foundation for further precise exploration of potential causal genes.
a–c Local Manhattan plots were generated for stone width (a), fruit weight (b), and bearing-shoot length (c). Accompanying box plots depict the distribution of these traits in accessions carrying distinct alleles. In the box plots, the upper and lower quartiles are represented by box limits, the medians are denoted by central lines, and whiskers extend to no more than 1.5× the interquartile range. Black dots indicate outliers which beyond the 1.5× the interquartile range. P-values were determined using a two-tailed Student’s t test. Accessions with the reference allele type are labelled as “Ref”, while those with alternative alleles are labelled as “Alt”. The number of accessions with the same haplotype is indicated in brackets. d Phenotypes of crushed stones from wild (left) and cultivated (right) jujubes. The seed from the wild jujube is shown on the left, with a scale bar of 1 cm. e Genome-wide Manhattan plots for the seed-setting rate; black dashed lines indicate significance thresholds ( − log10P = 4.18). The P-values for each SV were calculated using a two-sides mixed linear model implemented in the EMMAX software. f Gene structure of ZjMED12 and the associated 2.3- kb insertion in the upstream region. g Gene structure of OsMED12 and sequences at target sites in T0 plants produced using the CRISPR/Cas9 system. h Panicle morphology (upper) and plump grains per panicle (bottom) of WT rice and two CRISPR/Cas9-edited plants with bars measuring 5 cm. i Statistics of the seed-setting rate in WT rice and two CRI mutants with values expressed as mean ± SD (n = 10 panicles). Differences between WT and CRI mutants were assessed using a two-tailed Student’s t-test. Abbreviations: Del, deletion; Ins, insertion; Chr, chromosome; CRI, CRISPR/Cas9. Source data are provided as a Source Data file.
The transition from a sexual to an asexual reproductive strategy stands out as a prominent domestication event9. In contrast to wild jujube, which typically contains one or two seeds in the fruit stone, the majority of cultivated jujube varieties produce few or no seeds (Fig. 4d). Employing an SV-based GWAS on the seed-setting rate, we identified a 2.3-kb PAV located upstream ( − 10.3 kb relative to the start codon ATG) of the MED12 ortholog (ZjMED12, Z95_Ju00G334220), which was strongly associated with the seed-setting rate (Fig. 4e,f and Supplementary Fig. 16). This 2.3- kb insertion formed two haplotypes (reference and alternative alleles): accessions carrying the reference allele exhibited significantly higher seed-setting rates than did those with alternative allele (Supplementary Fig. 17a). We used two types of accessions to perform ZjMED12 expression analysis in young fruit ( ~ 5 mm). The gene displayed higher expression levels in accessions carrying the reference allele than in accessions carrying the alternative allele (Supplementary Fig. 17b). In Arabidopsis, the MED12-MED13 module of the Mediator regulates pattern formation during embryogenesis39, and loss-of-function of MED12 leads to defects in embryo development40.
To fully unravel the functions of ZjMED12, knocking it out in jujube would be essential. However, this undertaking faces several challenges, including the complexity of jujube transformation, the extended growth period of woody trees, and high genomic heterozygosity1. Consequently, we opted to employ the CRISPR/Cas9 (CRI) system to knock out OsMED12 (Os07g0648266), the ortholog of ZjMED12, in rice. For subsequent analysis, two CRI mutants were selected, featuring a 2- bp deletion and a 3- bp deletion in the sixth exon (Fig. 4g). These deletions led to a frameshift in the coding region for Osmed12-2 and the deletion of an asparagine residue from Osmed12-3. These deletions did not influence the expression levels of OsMED12 in rice (Supplementary Fig. 18). Phenotypic assessments revealed a significant decrease in the seed-setting rate for the two CRI mutants compared with that of WT rice (Fig. 4h,i). This finding suggested that MED12 functions conservatively in embryo development across both monocotyledonous and dicotyledonous plants.
ZjCDKI5 negatively regulates the bearing-shoot length and leaf size in jujube
Additionally, apart from SV-based GWAS, we conducted a SNP-based GWAS for the same 16 horticultural traits mentioned earlier. A total of 6700 SNPs were identified (Supplementary Table 7), with 2382 SNP locations overlapping (400- kb flanking region) with the results of SV-based GWAS. The remaining 4,318 SNP locations (64.45%) were exclusively detected by SNP-based GWAS. Notably, specific horticultural traits, such as the bearing-shoot length (BSL), leaf width, leaf length, and leaf area, exhibited significant increases during jujube domestication (Fig. 5a and Supplementary Fig. 19). In particular, BSL, leaf width, leaf length, and leaf area demonstrated a clear positive correlation (Supplementary Fig. 20). In the results of the SNP-based GWAS, a distinct GWAS signal on chromosome 08 was observed, simultaneously identified by the four aforementioned traits (Fig. 5b and Supplementary Fig. 21). This suggested the presence of a candidate gene with pleiotropic effects on these four domestication traits. Notably, this GWAS signal was not identified by SV-based GWAS.
a Phenotypes of bearing shoots and leaves for wild (upper) and cultivated (bottom) jujubes. b Genome-wide Manhattan plots for bearing-shoot length (BSL), leaf width, leaf length, and leaf area. The blue arrowhead indicates peak positions identified in this study. Black dashed lines represent significance thresholds ( − log10P = 5.74). The P-values for each SNP were calculated using a two-sides mixed linear model implemented in the EMMAX software. c Gene structure and sequence variations in ZjCDKI5. The number of accessions of each genotype is shown. d Box plots for BSL, leaf width, leaf length, and leaf area based on the four type accessions. In these box plots, upper and lower quartiles are represented by box limits, the mediums are denoted by central lines and whiskers extend to no more than 1.5 × the interquartile range. Gray dots indicate outliers which beyond the 1.5 × the interquartile range. The number of accessions with the same genotype is indicated in brackets. Significant differences were assessed using a two-tailed Student’s t-test. Different lowercase letters indicate significant differences among different genotypes at P < 0.05. e Relative transcript levels of ZjCDKI5 were measured by qPCR in WT rice (ZH11) and two overexpression (OE) plants in leaves. The values presented are mean ± SE of three technical repeats. f Plant morphology of WT rice and two OE plants at the heading stage. g Flag leaf morphology at the mature stage. h Statistics of flag leaf length and leaf width in WT rice and two OE plants. Values are expressed as mean ± SD (n = 10 leaves). Differences between WT and OE lines were tested using a two-tailed Student’s t-test. Abbreviations: Chr, chromosome; C-sub, cultivated subgroup. Source data are provided as a Source Data file.
Concerning BSL, the robust GWAS signal extended from 11.07 to 12.53 Mb on chromosome 08, encompassing 192 genes (Supplementary Data 7). Among them, 28 candidate genes exhibited high expression levels (FPKM > 20) in the both bearing shoot and leaf (Supplementary Fig. 22 and Supplementary Data 7). Gene description and functional annotation of orthologs in model plants directed our attention towards Z95_Ju00G226220 which encodes a cyclin-dependent kinase inhibitor (CDKI). An analysis of genetic variations in Z95_Ju00G226220 (designated ZjCDKI5) revealed one SNP in the promoter region and a 10 bp deletion in the third intron (Fig. 5c), which classified the population into four major genotype combinations (Fig. 5c). The accessions carrying genotype3 and genotype4 exhibited significantly higher phenotype indices than those with genotype1 and genotype2 (Fig. 5d). We then used the four types of accessions to perform ZjCDKI5 expression analysis in both bearing shoot and leaf. The gene displayed lower expression levels in accessions with genotype3 and genotype4 compared with those carrying genotype1 and genotype2, evident in both bearing shoot (P = 4.93 × 10−3) and leaf (P = 2.79 × 10−3) samples (Supplementary Fig. 23a). These findings, in conjunction with the gene description, suggested that ZjCDKI5 might function as a negative regulator influencing the four domestication traits.
The germination of the jujube bearing shoot occurs in spring, and it follows a deciduous pattern, typically shedding before winter. In a manner akin to annual crops, this characteristic draws a parallel with annual plant height11. To unravel the functions of ZjCDKI5, we generated ectopic overexpression rice plants, selecting two OE transgenic lines for subsequent analyzes (Fig. 5e). When assessing plant height and flag leaf size, we noted a significant reduction in plant height for the OE plants compared with that of wild-type (WT) rice, at both the seedling stage and heading stage (Fig. 5f and Supplementary Fig. 23b, c). Additionally, the flag leaf lengths and leaf widths of OE plants exhibited significant decreases compared with those of the WT rice (Fig. 5g, h). Collectively, these observations led us to the conclusion that ZjCDKI5 probably played a negative regulatory role in the increase of bearing-shoot length and leaf size during the domestication of jujube.
Discussion
We conducted de novo assembly of four high-quality, reference-grade jujube genomes by integrating PacBio CCS, Illumina short-read sequencing, and Hi-C technology. Building upon four previously released genomes25,26,27, we constructed a pan-genome using eight jujube genomes. We acknowledge that the current pan-genome, comprising only eight samples, is inadequate to encompass the full sequence diversity found within the jujube population, particularly in group C-sub5. Therefore, obtaining a more diverse reference map is essential to broaden genomic sampling in the future, which would allow better characterization of the genetic diversity of the jujube pan-genome. While our selection of eight accessions might seem limited in comparison to pan-genome analyzes in other plant species16,17,18,41, it is essential to note that these jujube accessions cover almost all phylogenetic groups, including the wild group and four cultivated subgroups (Fig. 1a and Supplementary Data 1). This diverse representation ensures the inclusion of various genetic backgrounds within the jujube population. Moreover, utilizing the pan-genome constructed from these eight representative accessions and subsequent GWAS analyzes, we successfully pinpointed several candidate genes that regulate crucial domestication traits. Noteworthy examples include ZjAGL28, ZjMED12, and ZjCDKI5, which play roles in regulating flowering and ripening time, the seed-setting rate, and the bearing-shoot length and leaf size, respectively. Furthermore, the assemblies of the four additional jujube accessions established in this study serve as a foundation for future pan-genome analysis in the field.
In this study, the biological functions of ZjAGL28 and ZjCDKI5 were validated by ectopic transformations. We cannot definitely confirm that the transgenic proteins expressed in Arabidopsis thaliana and rice are the same as in jujube. To fully unravel the functions of above two candidates, knocking out and overexpressing them in jujube would be essential. However, this work faces several challenges, including the difficulty in jujube transformation, the long growth period of woody trees, and high genomic heterozygosity1. Consequently, we opted to conduct ectopic overexpression of these two candidates in model plants. In this study, three candidate genes were identified by comparative genomics and GWAS, not using linkage mapping. For annual crops, using a segregated group to mapping QTLs is a common strategy42. However, it is substantially difficult and a time-consuming process for jujube. First, artificial pollination is difficult for jujube43. Second, under normal conditions, it is only less than 1% flower which can develop into fruits. Most of flowers fall off from bearing shoots43. Third, jujube is a perennial woody tree with a relative long juvenile period. So, to date, it is still a challenge to map candidate genes/QTLs using segregated groups in jujube.
Selective sweep analysis is a common approach to identify genes associated with domestication traits8,9,44. Herein, a total of 2302 potential domestication genes were identified within the selective sweep regions through π ratio analysis (Supplementary Data 4). Although we functionally verified some candidate genes within the selective sweeps, it is important to acknowledge that the method used in our study might have detected some false positives. In the future, more robust methodologies should be employed to more precisely identify domestication-related genes.
To unravel the genetic basis underlying crucial horticultural traits, we simultaneously performed SV-based and SNP-based GWAS for 16 traits, leading to the identification of 103 SVs and 6,700 SNPs associated with these traits (Supplementary Table 7). Statistically, 61 out of 103 SVs (59.22%) shared an overlap (within a 400- kb flanking sequence) with the SNPs detected by SNP-based GWAS. Additionally, the genomic locations of 35.55% of the identified SNPs also overlapped with SVs detected by SV-based GWAS. Notably, the remaining loci were exclusively detected by either SNPs or SVs, underscoring the technical complementarity between SNP-based and SV-based GWAS. This observation aligns with findings from previous studies, in which 17.5% of SVs showed very low linkage with nearby SNPs detected using SNP-based GWAS in rice17 and only 5.2% of loci overlapped between SV-based and SNP-based GWAS in tomato20, highlighting the importance of the simultaneous application of both approaches. This dual approach is recommended to ensure comprehensive identification of candidate genes and to prevent the oversight of key genetic loci.
In recent years, notable advances have been made in identifying candidate or causal genes associated with horticultural traits in jujube, employing both forward and reverse genetic approaches. These genes primarily belong to three functional categories: (1) morphological traits related to fruit size and shape, root growth, and flowering9,11,45,46; (2) quality and metabolism-related traits associated with fruit sweetness and acidity, as well as fruit lignin biosynthesis8,44,47,48,49; and (3) biotic and abiotic stresses related to jujube witches’ broom and salt stress50,51,52. Despite these advances, in comparison to other perennial fruit crops such as apple, peach, and pear, functional genomic research in jujube is still in its early stage. Consequently, the molecular mechanisms underlying many horticultural traits remain poorly understood. Our analyzes successfully identified candidate genes contributing to flowering and fruit ripening, the seed-setting rate, BSL and leaf size in jujube. This information enhances our understanding of the genetic basis of these horticultural traits. The wealth of genomic data, including a pan-genome and large-scale resequencing data, significantly enriches the genetic resources available for basic research and facilitates future breeding efforts in jujube.
Methods
Sample collection and agronomic evaluation
The four jujube accessions (Z95, Z94, Z203, S21) used for de novo assemblies were sampled at the Experimental Station of Luoyang Normal University (Luoyang, Henan Province, China) and the National Jujube Germplasm Repository in Shanxi Agricultural University (Taigu, Shanxi, China). The jujube cultivars of resequencing population were mainly collected at the National Jujube Germplasm Repository in Shanxi Agricultural University (Taigu, Shanxi, China) and the National Foundation for Improved Cultivar of Chinese Jujube (Cangxian County, Hebei Province, China). The remaining cultivars and wild individuals of resequencing population were sampled from wilderness. The geographical distributions of all 1059 accessions were from 25 provinces/autonomous regions/municipalities of China covering almost all jujube-planting areas, and four cultivars were gathered from South Korea (Supplementary Fig. 3a and Supplementary Data 1).
For agronomic evaluation, seven traits, including stone width, fruit weight, bearing-shoot length, seed-setting rate, leaf area, leaf length, and leaf width, were measured based on previously published jujube genetic resources evaluation criteria53. For stone width, after peeling of the flesh of mature fruits, the clean stones were prepared for measurement of stone width which was determined at the widest part of stone by using vernier caliper, and it was calculated as the average value of ten stones. Fruit weight was determined by using electronic balance as the average value of ten healthy half-red fruits sampled from different orientations of tree. For bearing-shoot length (BSL), ten healthy and strong bearing shoots were collected from different orientations of tree at maturation stage, and BSL was determined by using ruler as the average value of ten bearing shoots. The seed-setting rate equals a number of stones with plump seeds/number of all detected stones and was evaluated using around 30 healthy fruits which were sampled from different orientations of the tree. Leaf area, leaf length, and leaf width were determined by using the LA-S Leaf Area Meter (Wanshen, Hangzhou, China) as the average value of ten leaves, which were sampled from ten bearing shoots. And for each bearing shoot, one leaf in the middle was sampled for measurement. For each trait, all samples were collected from one tree. The detailed information on the other nine traits, including the fruit length, fruit width, fruit shape index, stone length, stone shape index, stone weight, number of leaves per bearing shoot, internode length of bearing shoot, and ratio of edibility, can be found in Supplementary Method 1. The numbers of wild and cultivated jujube accessions investigated for each trait were listed in Supplementary Table 8.
Illumina sequencing
Genomic DNA was extracted from young leaves using cetyltrimethylammonium bromide54. A minimum of 5 μg of genomic DNA per accession was utilized to create sequencing libraries, following the manufacturer’s guidelines (Illumina, San Diego, CA, USA). The libraries were subjected to paired-end (NGS) sequencing on the Illumina NovaSeq 6000 platform, generating 150 bp reads (Supplementary Method 2). Additionally, total RNA was extracted from bearing shoot, leaf, flower, stem, phloem and fruit tissues for library construction (Supplementary Table 9), resulting in ~6 Gb of data for each tissue during subsequent sequencing (Supplementary Method 3).
Genome sequencing and assembly
The selection of the four jujube accessions for genome assembly was based on their phylogenetic grouping. Genomic DNA was extracted from the fresh leaves of each accession. SMRTbell libraries were constructed following the standard PacBio (Pacific Biosciences, Menlo Park, CA, USA) protocol and then sequenced on the PacBio Sequel II platform to generate HiFi reads. For the creation of Hi-C libraries, DNA was extracted from fresh leaves. Chromatin underwent a 12-hour digestion with 20 units of DpnII restriction enzyme (New England Biolabs, Beijing, China) at 37 °C. The resulting mixture was subsequently incubated at 62 °C for 20 minutes to deactivate the restriction digestion. DNA fragments ranging from 300 to 500 bp were excised and purified using Ampure XP beads (Beckman Colter, Brea, CA, USA). These Hi-C libraries were sequenced on the Illumina NovaSeq 6000 platform with 2 × 150-bp reads.
The estimation of genome size and heterozygosity was performed with a k-mer-based approach using Jellyfish (v 2.2.10)55 and GenomeScope 2.056, utilizing the ~50× Illumina sequencing data. Subsequently, the genomes of the four HiFi-sequenced accessions were assembled with hifiasm (v0.13)28 (https://github.com/chhylp123/hifiasm), employing default parameters. The assembled contigs were then anchored to the chromosome level with Hi-C data through the 3D-DNA pipeline57. Hi-C reads were aligned to the polished contigs using the Juicer pipeline58. The 3D-DNA pipeline was executed with the following parameters: -i 1 -r 5. The results were refined using the Juicebox Assembly Tools59.
We evaluated the completeness of the genic region in the assemblies utilizing BUSCO (v5.2.0)29 embryophyta_odb10 database, with a set of 1440 embryophyte genes. For the assessment of intergenic region completeness, we employed the LAI with LTR_retriever (v2.9.0)60. Additionally, we assessed genome completeness by aligning high-quality Illumina short reads to the corresponding assembly using BWA (v0.7.12-r1039)61 with default parameters. The full details of genome sequencing and assembly are available in the Supplementary Method 4.
Genome annotation, GO and KEGG enrichment analysis
The detailed information of TE annotation is available in the Supplementary Method 5. Protein-coding genes were predicted for each genome assembly through the MAKER262 pipelines. RNA evidence was gathered by aligning RNA-sequencing (RNA-seq) reads to the repeat-masked assembly using HISAT2 (v.2.10.2)63, followed by assembly into transcripts using StringTie (v.1.3.0)64. TACO (v.0.7.3)65 was employed to merge stringtie gtf (–filter-splice-juncs). Ab initio gene prediction was executed using AUGUSTUS (v.3.3.3)66 and SNPA67. Protein sequences from SwissProt (Viridiplantae) (https://www.uniprot.org) and previously published jujube protein sequences were also integrated. All these proteins were utilized for homology-based prediction with BRAKER (v.2.1.4)68. Only integrated gene models with AED values < 0.5 were retained. More information of gene annotation can be found in Supplementary Method 5. The methods of GO and KEGG enrichment analysis are shown in the Supplementary Method 6.
SNPs and InDels calling of 1059 jujube accessions
To identify genetic variations, we employed the BWA-mem software (v6.0.2)61 to map the clean reads to the reference genome with default parameters. Subsequently, SAM files were converted to BAM files using SAMtools (v0.1.18) software69. Following the mapping process, the BAM file underwent sorting, and duplicates were marked using Picard tools (v1.119) (http://broadinstitute.github.io/picard/).
Variants were identified through GATK (v.4.2.3.0)70 HaplotypeCaller, and the identified SNPs and InDels underwent further filtration based on following criteria: SNPs were filtered with “QD < 2.0 | | FS > 60.0 | | MQ < 40.0 | | SOR > 3.0 || MQRankSum < −12.5 | | ReadPosRankSum < −8.0”, and InDels with “QD < 2.0 | | FS > 200.0 | | SOR > 10.0 || MQRankSum < −12.5 || ReadPosRankSum < −8.0”. To ensure the quality of SNP and InDel, these variations located within TE-regions were excluded for subsequently analysis.
Phylogenetic and population structure analysis
For the phylogenetic analysis, we first obtained all genomic variation loci and exclude those in TE regions. Then, to ensure SNP representativeness and reduce computational load, we filtered SNPs with high LD using PLINK (v1.90b3.46)71. The LD filtering command was ‘plink –file input –indep-pairwise 50 10 0.2 –out output’. After LD pruning, we selected SNPs with a minor allele frequency (MAF) ≥ 0.02 and a missing rate ≤ 0.4, resulting in 557,726 SNPs for tree construction. We used FastTree with the GTR model to construct a Maximum-Likelihood (ML) phylogenetic tree. The Newick format file was then uploaded to MEGA6.072 for visualization and optimization.
In population structure analysis, we extracted SNPs outside the TE regions and filtered for those with a minor allele frequency (MAF) ≥ 0.02 and a missing rate ≤ 0.4. This gave us 6,185,881 SNPs for analysis using ADMIXTURE (version 1.3.0)30. Taking advantage of the same data set, we also performed PCA analysis with EIGENSOFT (v6.0.1)73 and LD analysis using PopLDdecay (v3.40)74 with the command ‘PopLDdecay -InGenotype input.genotype -OutStat result.out -MAF 0.02 -Miss 0.4’. LD decay was calculated based on the r² value and the distance between SNPs.
Genomic selection signature identification
To identify potential selective sweeps, we assessed the genome-wide reduction in genetic diversity (π) using VCFtools software75. The command used for this analysis was: vcftools –gzvcf pop.vcf.gz –window-pi 100000 –window-pi-step 10000 –out result –keep target.group.list. The investigation focused on detecting selection across the genome during domestication by comparing wild and cultivated groups. Genomic regions influenced by domestication were expected to exhibit significantly lower diversity in the landrace group compared to the wild group. Windows with π < 0.001 in the wild were excluded from further analysis, and windows with the top 5% ratios of πwild/πcultivated were chosen as candidate domestication sweeps. Adjacent windows within a distance of ≤ 100 kb were merged into a single selected region.
Gene-based pan-genome construction and Ka/Ks calculation
We conducted a pan-genome analysis employing a Markov clustering approach76. All-versus-all comparisons were executed using Diamond (v0.9.25)77. Subsequently, the paired genes were clustered using OrthoFinder (v2.3.12)78. Based on their occurrence, gene families were categorized into three groups: core (present in all eight accessions), dispensable (present in two to seven accessions), and accession-specific (unique to one accession) (Supplementary Method 7). The details of Ka/Ks calculation for each gene of the pan-genome are available in the Supplementary Method 8.
Genomic variations detection
To uncover genomic variations, we aligned the other seven genomes to the Z95 reference genome using MUMmer (v4.0.0rc1)35. The alignment was conducted with the command ‘nucmer –maxmatch -c 50 -b 500 -l 20 input1.fa input2.fa’. The alignment results underwent filtration using the delta-filter program in MUMmer with parameters ‘−1 -i 90 -l 100’. The show-coords program in MUMmer was employed to extract alignment blocks from the intergenomic alignment results, and SyRI (v1.0)37 identified genomic variations in each comparison.
The extracted SNPs and InDels were annotated using the SnpEff software (v4.3t)36. Non-synonymous SNPs refer to those labeled as ‘missense_vatiant’ in the SnpEff annotation results. Variants (SNPs and InDels) with a significant impact on sequence alteration, labeled as ‘HIGH’ in the SnpEff annotation, are considered to have a high putative impact on the gene’s products and function. We further explored nine types of SVs (Supplementary Table 6) defined by Syri37. The coding regions of genes with > 50% overlap with SVs were regarded as affected by big-effect SVs. SVs containing ‘N’ sequences were excluded.
Furthermore, we investigated PAVs by selecting deletions, insertions, copy losses, and copy gains from the SVs detected by Syri37. This enabled us to genotype these PAVs using the SURVIVOR software (v.1.0.6)79 with the following parameters: ‘50 1 0 0 0 0’ and paragraph (v2.3-h8908b6f_0)80 software with the command ‘~/bin/multigrmpy.py -i merged.vcf -m mfile -r Ref.fa -o output –threads 5’ at population level.
Genome-wide association study
Leveraging a dataset comprising 19,749 PAVs and 4,844,730 filtered SNPs (excluding variations with a missing rate > 0.4 and minor allele frequency < 0.05) along with information on 16 key horticultural traits, we proceeded to conduct association tests. EMMAX (vbeta-07Mar2010)81 was employed for this analysis, where population stratification and hidden relatedness were effectively modeled using a kinship (K) matrix within the emmax-kin-intel package of EMMAX. The determination of the genome-wide significance threshold followed a uniform threshold of 1/n, with n representing the effective number of independent SVs or SNPs calculated through Genetic type 1 Error Calculator (v0.2)82. The detailed information, including SNP-based GWAS, SV detection, SV-based GWAS, and identification of candidate genes in GWAS, is available in the Supplementary Method 9–11.
qPCR
For qPCR, total RNA was extracted with a Takara MiniBEST Plant RNA Extraction Kit (TaKaRa, Dalian, China). The first-strand cDNA was synthesized using a TaKaRa PrimeScript II 1st Strand cDNA Synthesis Kit (TaKaRa). qPCR was performed in triplicate with TaKaRa SYBR Premix Ex Taq II (TaKaRa) on a Bio-Rad CFX96 machine (Bio-Rad, Hercules, CA, USA). And AtActin2 (AT3G18780), OsActin1 (Os03g0718100), and ZjActin (GenBank: KT381859) were employed as the endogenous for normalization. The relative expression levels were calculated using quantification method (2-ΔΔCT)83. Primers used for qPCR are listed in Supplementary Table 10.
Vector construction and plant transformation
For overexpression constructs, the full-length coding sequence of ZjAGL28 and ZjCDKI5 were amplified through PCR from cDNAs and the PCR products were cloned into the modified pCAMBIA-1300 vector driven by the CaMV 35S promoter and maize Ubiquitin promoter, respectively. For knocking out of OsMED12, three 23- bp gene-specific sequences (tcgcttgtttggctgggaaaggg; aatgaacgcagtcgcttgtttgg; atgttcctcatggttatcgtagg) targeting the sixth exon of OsMED12 were inserted into the sgRNA/Cas9 vector to generate the OsMED12-Cas9 construct. Primers used for vector construction are listed in Supplementary Table 10.
For Arabidopsis and rice transformations, the resulting constructs were introduced into Columbia type and Zhonghua11 (ZH11) by Agrobacterium tumefaciens-mediated transformation, respectively.
Protein structure prediction
The protein structure was predicted using online tool AlphaFold384 (https://alphafoldserver.com/), and PyMOL (3.0) software85 was used for visualizing the three-dimensional structure.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All raw PacBio CCS, transcriptome data, and resequencing data have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive database under BioProject accession number PRJNA1051535. All four assembled jujube genomes have been deposited at ENA/GenBank under the accessions: JBEONJ000000000 (GCA_041146685.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_041146685.1/]), JBEONK000000000 (GCA_041146715.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_041146715.1/]), JBEONL000000000 (GCA_041146725.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_041146725.1/]), and JBEONM000000000 (GCA_041146735.1 [https://www.ncbi.nlm.nih.gov/datasets/genome/GCA_041146735.1/]). All four assembled genomes and annotations are also accessible through Figshare [https://doi.org/10.6084/m9.figshare.24923472]. The initial version of ‘Dongzao’genome data were downloaded from Genbank under accession JREP00000000. The updated T2T version of ‘Dongzao’ genome data were downloaded from National Genomics Data Center (NGDC) under BioProject accession PRJCA016173. The genome data of ‘Junzao’ and a wild accession S2024 were downloaded from NCBI under BioProject accession PRJNA974227. The genome data of a wild accession S2021 were downloaded from Genbank under accession JAEACU000000000. The previously released jujube resequencing data which were used in this study are available at NCBI under BioProject accession PRJNA560664. Source data are provided with this paper.
Code availability
The codes used for this paper are available on GitHub [https://github.com/LianqunBio/Pan-jujube]. Codes were also archived on Zenodo [https://doi.org/10.5281/zenodo.13929708]86.
References
Liu, M. et al. The complex jujube genome provides insights into fruit tree biology. Nat. Commun. 5, 5315 (2014).
Liu, M. et al. The historical and current research progress on jujube–a superfruit for the future. Hortic. Res. 7, 119 (2020).
Qu, Z. & Wang, Y. Chinese Fruit Trees Record-Chinese Jujube (China Forestry Publishing House, Beijing, 1993).
Liu, M. & Wang, M. Germplasm Resources of Chinese Jujube (China Forestry Publishing House, Beijing, 2009).
Sapkota, D., Zhang, D., Park, S., Meinhardt, L. W. & Yao, S. Genotyping of jujube (Ziziphus spp.) germplasm in New Mexico and Southwestern Texas. Plants (Basel) 12, 2405 (2023).
Khadivi, A. Morphological characterization and interspecific variation among five species of Ziziphus genus to select superiors in Iran. BMC Plant Biol. 23, 550 (2023).
Liu, M. Chinese jujube: botany and horticulture. Hort. Revi. 32, 229–298 (2006).
Huang, J. et al. The jujube genome provides insights into genome evolution and the domestication of sweetness/acidity taste in fruit trees. PLoS Genet. 12, e1006433 (2016).
Guo, M. et al. Genomic analyses of diverse wild and cultivated accessions provide insights into the evolutionary history of jujube. Plant Biotechnol. J. 19, 517–531 (2021).
Miller, A. J. & Gross, B. L. From forest to field: perennial fruit crop domestication. Am. J. Bot. 98, 389–414 (2011).
Guo, M. et al. Comparative population genomics dissects the genetic basis of seven domestication traits in jujube. Hortic. Res. 7, 89 (2020).
Hirsch, C. N. et al. Insights into the maize pan-genome and pan-transcriptome. Plant Cell 26, 121–135 (2014).
Golicz, A. A., Batley, J. & Edwards, D. Towards plant pangenomics. Plant Biotechnol. J. 14, 1099–1105 (2016).
Hurgobin, B. et al. Homoeologous exchange is a major cause of gene presence/absence variation in the amphidiploid Brassica napus. Plant Biotechnol. J. 16, 1265–1274 (2018).
Guo, J. et al. An integrated peach genome structural variation map uncovers genes associated with fruit traits. Genome Biol. 21, 258 (2020).
Liu, Y. et al. Pan-genome of wild and cultivated soybeans. Cell 82, 162–176 (2020).
Qin, P. et al. Pan-genome analysis of 33 genetically diverse rice accessions reveals hidden genomic variations. Cell 184, 3542–3558 (2021).
Zhou, Y. et al. Graph pangenome captures missing heritability and empowers tomato breeding. Nature 606, 527–534 (2022).
Li, H. et al. Graph-based pan-genome reveals structural and sequence variations related to agronomic traits and domestication in cucumber. Nat. Commun. 13, 682 (2022).
Li, N. et al. Super-pangenome analyses highlight genomic diversity and structural variation across wild and cultivated tomato species. Nat. Genet. 55, 852–860 (2023).
Gao, L. et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 51, 1044–1051 (2019).
Huang, Y. et al. Pangenome analysis provides insight into the evolution of the orange subfamily and a key gene for citric acid accumulation in citrus fruits. Nat. Genet. 55, 1964–1975 (2023).
He, Q. et al. A graph-based genome and pan-genome variation of the model plant Setaria. Nat. Genet. 55, 1232–1242 (2023).
Muhtar, Z., Gulmira, K., Wu, Z. & Hardil, L. Comprehensive evaluation on fruit quality of different age secondary branches in ‘Huizao’ jujube. Trans. Chin. Soc. Agric. Eng. 39, 298–303 (2023).
Shen, L. Y. et al. Chromosome-scale genome assembly for Chinese sour jujube and insights into its genome evolution and domestication signature. Front. Plant Sci. 12, 773090 (2021).
Yang, M. et al. Insights into the evolution and spatial chromosome architecture of jujube from an updated gapless genome assembly. Plant Commun. 4, 100662 (2023).
Li, K. et al. Haplotype-resolved T2T reference genomes for wild and domesticated accessions shed new insights into the domestication of jujube. Hortic. Res. 11, uhae071 (2024).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Alexander, D. H. & Lange, K. Enhancements to the ADMIXTURE algorithm for individual ancestry estimation. BMC Bioinformatics 12, 246 (2011).
Wu, J. et al. Diversification and independent domestication of Asian and European pears. Genome Biol. 19, 77 (2018).
Liao, L. et al. Unraveling a genetic roadmap for improved taste in the domesticated apple. Mol. Plant 14, 1454–1471 (2021).
Li, Y. et al. Genomic analyses of an extensive collection of wild and cultivated accessions provide new insights into peach breeding history. Genome Biol. 20, 36 (2019).
Contreras-Moreira, B. et al. Analysis of plant pan-genomes and transcriptomes with GET_HOMOLOGUES-EST, a clustering solution for sequences of the same species. Front. Plant Sci. 8, 184 (2017).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944 (2018).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms. SnpEff: SNPs in the genome of Drosophila melanogaster strain w1118; iso-2; iso-3. Fly. (Austin) 6, 80–92 (2012).
Goel, M., Sun, H., Jiao, W. B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Pagnussat, G. C. et al. Genetic and molecular identification of genes required for female gametophyte development and function in Arabidopsis. Development 132, 603–614 (2005).
Gillmor, C. S. et al. The MED12-MED13 module of Mediator regulates the timing of embryo patterning in Arabidopsis. Development 137, 113–122 (2010).
Meinke, D. W. Genome-wide identification of EMBRYO-DEFECTIVE (EMB) genes required for growth and development in Arabidopsis. New Phytol. 226, 306–325 (2020).
Chen, J. et al. Pangenome analysis reveals genomic variations associated with domestication traits in broomcorn millet. Nat. Genet. 55, 2243–2254 (2023).
Bazakos, C., Hanemian, M., Trontin, C., Jiménez-Gómez, J. M. & Loudet, O. New strategies and tools in quantitative genetics: how to go from the phenotype to the genotype. Annu. Rev. Plant Biol. 68, 435–455 (2017).
Liu, M. et al. Design and practice of emasculation-free cross breeding in Chinese jujube. Acta Hortic. Sin. 41, 1495–1502 (2014).
Zhang, Z. et al. Jujube metabolome selection determined the edible properties acquired during domestication. Plant J. 109, 1116–1133 (2022).
Liu, Z. et al. Three novel adenylate cyclase genes show significant biological functions in plant. J. Agric. Food Chem. 71, 1149–1161 (2023).
Zhao, X. et al. ZjWRKY23 and ZjWRKY40 promote fruit size enlargement by targeting and downregulating cytokinin oxidase/dehydrogenase 5 expression in Chinese jujube. J. Agric. Food Chem. 71, 18046–18058 (2023).
Zhang, Q. et al. The regulation of cell wall lignification and lignin biosynthesis during pigmentation of winter jujube. Hortic. Res. 8, 238 (2021).
Zhang, C. et al. Low-acidity ALUMINUM-DEPENDENT MALATE TRANSPORTER4 genotype determines malate content in cultivated jujube. Plant Physiol. 191, 414–427 (2023).
Liu, H., Zhao, X., Bi, J., Dong, X. & Zhang, C. A natural mutation in the promoter of the aconitase gene ZjACO3 influences fruit citric acid content in jujube. Hortic. Res. 11, uhae003 (2024).
Zhou, J. et al. Jujube witches’ broom phytoplasma effectors SJP1 and SJP2 induce lateral bud outgrowth by repressing the ZjBRC1-controlled auxin efflux channel. Plant Cell Environ. 44, 3257–3272 (2021).
Chen, P. et al. Phytoplasma effector Zaofeng6 induces shoot proliferation by decreasing the expression of ZjTCP7 in Ziziphus jujuba. Hortic. Res. 9, uhab032 (2022).
Ma, Z. et al. Mycorrhizal symbiosis reprograms ion fluxes and fatty acid metabolism in wild jujube during salt stress. Plant Physiol. 189, 2481–2499 (2022).
Li, D. K. Descriptors and Data Standard for Jujube (Ziziphus jujuba Mill.) (China Agriculture Press, Beijing, 2006).
Gawel, N. J. & Jarret, R. L. A modified CTAB DNA extraction procedure for Musa and Ipomoea. Plant Mol. Biol. Rep. 3, 262–266 (1991).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 64–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11, 1432 (2020).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scafolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res. 46, e126 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Niknafs, Y. S., Pandian, B., Iyer, H. K., Chinnaiyan, A. M. & Iyer, M. K. TACO produces robust multi sample transcriptome assemblies from RNA-seq. Nat. Methods 14, 68–70 (2017).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Hoff, K. J., Lomsadze, A., Borodovsky, M. & Stanke, M. Whole-genome annotation with BRAKER. Methods Mol. Biol. 1962, 65–95 (2019).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Zhang, C., Dong, S. S., Xu, J. Y., He, W. M. & Yang, T. L. PopLDdecay: a fast and effective tool for linkage disequilibrium decay analysis based on variant call format files. Bioinformatics 35, 1786–1788 (2019).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2168 (2011).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60 (2015).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 16, 157 (2015).
Jeffares, D. C. et al. Transient structural variations have strong effects on quantitative traits and reproductive isolation in fission yeast. Nat. Commun. 8, 14061 (2017).
Chen, S. et al. Paragraph: a graph-based structural variant genotyper for short-read sequence data. Genome Biol. 20, 291 (2019).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Li, M. X., Yeung, J. M., Cherny, S. S. & Sham, P. C. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum. Genet. 131, 747–756 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408 (2001).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Yuan, S., Chan, H. C. S. & Hu, Z. Using PyMOL as a platform for computational drug design. Wiley Interdiscip. Rev. Comput. Mol. Sci. 7, e1298 (2017).
Guo, M. et al. Analyzes of pan-genome and resequencing atlas unveil the genetic basis of jujube domestication. Zenodo https://doi.org/10.5281/zenodo.13929708 (2024).
Acknowledgements
This work was supported by the National Key R&D Program of China (2023YFA0915800 and 2023ZD04076 to J.Y.); the National Natural Science Foundation of China (32271928 to M.G.); the Luoyang Key Science and Technology Innovation Program (2101016 A to X.Z.); the Tianchi Talent in Xinjiang of China (to X.Z.), the Innovation Program of Chinese Academy of Agricultural Sciences and the Elite Young Scientists Program of CAAS (to J.Y.). We thank Decang Kong (National Foundation for Improved Cultivar of Chinese Jujube, Bureau of Natural Resources and Planning of Cangxian County) and Yongkang Wang (National Jujube Germplasm Repository, Shanxi Agricultural University) for assisting for sampling.
Author information
Authors and Affiliations
Contributions
X.Z., M.G., Q.L., S.H., and J.Y. conceived and designed the research. M.G., X.Z., W.Y., W.H., Z.W., and J.Z. collected samples for DNA sequencing. M.G., W.Y., S.Z. (Suna Zhao), S.Z. (Siyuan Zhang), X.X., H.Z., and K.G. performed field experiments and phenotyping. Q.L. carried out the genome assemblies and annotations. M.G., Y.M., and Q.L. performed population structure and selective sweep analyzes. Q.L., M.G., L.C., and Z.B. carried out pan-genome related analyzes and GWAS analyzes. M.G. led in-depth analyzes of identified genomic variations associated with traits. M.G. conducted gene expression analysis and transgenic experiments. H.W. uploaded the raw sequencing data and assembled genomes and annotations in the public database. M.G. and Q.L. wrote the manuscript. J.Y. revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ludovic Duvaux, Jianquan Liu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Guo, M., Lian, Q., Mei, Y. et al. Analyzes of pan-genome and resequencing atlas unveil the genetic basis of jujube domestication. Nat Commun 15, 9320 (2024). https://doi.org/10.1038/s41467-024-53718-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53718-z
This article is cited by
-
Comparative genomics of light harvesting chlorophyll a/b binding protein (LHC) family in green jujube (Ziziphus mauritiana) reveal the effect of domestication based on haplotype-resolved genome
BMC Plant Biology (2026)
-
Haplotype-resolved genome assembly of the leading cultivar of jujube (Ziziphus jujuba Mill. ‘Huizao’)
Scientific Data (2025)