Abstract
Genome-wide association studies (GWAS) have identified over 800 loci associated with kidney function, yet the specific genes, variants, and pathways involved remain elusive. By integrating kidney function GWAS with human kidney expression and methylation quantitative trait analyses, we identified Ten-Eleven Translocation (TET) DNA demethylase 2 (TET2) as a novel kidney disease risk gene. Utilizing single-cell chromatin accessibility and CRISPR-based genome editing, we highlight GWAS variants that influence TET2 expression in kidney proximal tubule cells. Experiments using kidney/tubule-specific Tet2 knockout mice indicated its protective role in cisplatin-induced acute kidney injury, as well as in chronic kidney disease and fibrosis induced by unilateral ureteral obstruction or adenine diet. Single-cell gene profiling of kidneys from Tet2 knockout mice and TET2-knockdown tubule cells revealed the altered expression of DNA damage repair and chromosome segregation genes, notably including INO80, another kidney function GWAS target gene itself. Remarkably, both TET2-null and INO80-null cells exhibited an increased accumulation of micronuclei after injury, leading to the activation of cytosolic nucleotide sensor cGAS-STING. Genetic deletion of cGAS or STING in kidney tubules, or pharmacological inhibition of STING, protected TET2-null mice from disease development. In conclusion, our findings highlight TET2 and INO80 as key genes in the pathogenesis of kidney diseases, indicating the importance of DNA damage repair mechanisms.
Similar content being viewed by others
Introduction
Kidney disease is a major global health burden, affecting over 850 million people worldwide1. Kidney disease incidence is much higher in older adults2, and it is one of the fastest-growing causes of mortality3. A better understanding of the mechanisms of kidney dysfunction is crucial for developing new therapeutics to treat or cure kidney disease.
Kidney function shows strong heritability4,5,6. Large genome-wide association studies (GWAS) of kidney function have identified more than 800 loci that demonstrate statistically significant and reproducible associations with kidney function7,8. However, over 90% of variants identified by GWAS reside in the noncoding regions of the genome9, which are often correlated due to strong linkage disequilibrium (LD)10. Consequently, pinpointing the causal variants, their target genes, specific cell types involved, and underlying disease mechanisms continues to be a major challenge.
Disease-causing variants are often located in cell type-specific regulatory regions, potentially altering the binding strength of transcription factors and resulting in quantitative differences in the expression of cell type-specific target genes9. Expression quantitative trait loci (eQTL) analysis, which defines the association between genetic variants and tissue gene expression, has been widely used to identify target genes in GWAS4,11,12,13. Analyzing the effects of genotype on the epigenome, such as DNA methylation, can improve gene and cell type prioritization8. In addition, examining changes in gene expression and regulation at the single-cell level, rather than at the whole tissue bulk level, is critical9. While computational integration strategies have prioritized new kidney function risk genes based on GWAS and human kidney multi-omics datasets, these studies require careful follow-up cell and animal model experiments to substantiate the causal role of a specific gene8.
DNA methylation, the addition of a methyl group to the 5-position of cytosines, is mediated by DNA methyltransferases (DNMTs)14. The Ten-Eleven Translocation (TET) enzymes play a role in demethylation by converting 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC), 5-formylcytosine, and 5-carboxylcytosine15,16. DNA methylation is an energetically costly process which also increases the mutation rate, as 5mC is converted to thymine, which is difficult to repair17. DNA methylation is an important regulator of multiple DNA-based processes. Most studies have analyzed the role of cytosine methylation in gene expression regulation14. Cytosine methylation renders the DNA region less accessible to the transcriptional machinery by recruiting methyl binding proteins and altering transcription factor binding strength18. Cytosine methylation is critical during kidney development and our previous studies demonstrated the key role of Dnmt1 in Six2 in progenitor cells as it is required for transposable element silencing19 while Dnmt3a and Dnmt3b are essential for the silencing developmental genes20. Kidneys obtained from patients with chronic kidney disease (CKD and fibrosis) also showed important changes in cytosine methylation at more than 100 CpG sites21. Globally, more CpG sites show higher methylation level in CKD21. These methylation changes correlate and predict kidney function decline, indicating their potential role in disease development21.
DNA damage occurs in various forms, including abasic sites, adducts, DNA-protein cross-links, insertion/deletion mismatches, double-strand breaks (DSBs), and single-strand breaks (SSBs)22. Cells respond to DNA damage by initiating a series of highly coordinated events to repair DNA, known as the DNA damage response (DDR), which includes base excision repair (BER), nucleotide excision repair (NER), mismatch repair (MMR), non-homologous end joining (NHEJ), and homologous recombination (HR)22. The ataxia teleangiectasia mutated (ATM) and ATM-RAD3 related (ATR), and the DNA-dependent protein (DNA-PK) are three kinases that control DDR and orchestrate DSB22. In cases where the DNA damage exceeds the repair capacity of the cell, the DDR triggers senescence or apoptosis22. By arresting proliferation and promoting clearance of damaged cells, the DDR acts as a barrier to tumorigenesis23. Recent studies highlighted the role of ATM and ATR in kidney disease development24,25. Mutations in DNA repair pathway genes, such as ERCC1, ERCC2, ERCC6, and ERCC8, have been associated with progeria, cancer, or immunodeficiency24. The modified ERCC1 knockout mouse model, which expresses a liver-specific ERCC1 rescue transgene, developed proteinuria, segmental glomerulosclerosis, and end-stage renal failure24,25. The role of cytosine methylation in DDR is not fully understood; however, both epigenetic changes and DNA damage are strongly associated with aging26.
In this study, we identified genetic variants associated with kidney function that concurrently lower the expression of Tet2 in kidney proximal tubule cells. Kidney/tubule-specific gene knockout mice have highlighted the protective roles of Tet2 both in acute kidney injury and chronic fibrosis. Molecular studies indicated the role of TET2 in the HR pathway for DSB repair and in ensuring proper chromosome segregation in proximal tubule cells. Loss of Tet2 lead to kidney injury by leaving micronuclei in the cytosol, resulting in cGAS-STING activation and subsequent kidney disease development.
Results
Prioritization of TET2 for kidney function
Multiple genome-wide association studies have recognized common noncoding nucleotide variants on chromosome 4 associated with kidney function5,8,27,28,29. To identify the causal variants within this locus, we performed statistical fine mapping using the SuSie method. We identified two credible sets: rs6533181 (posterior inclusion probability, PIP = 9.98 × 10−8), rs115918101 (PIP = 8.82 × 10−8), and rs2214407 (PIP = 3.54 × 10−8) (Fig. 1a). The variant rs6533181 was strongly and significantly associated with kidney function in the eGFR creatinine GWAS (P = 1.41 × 10−10) (Fig. 1b). The minor allele frequency (MAF) of the variant rs6533181 was 0.3397.
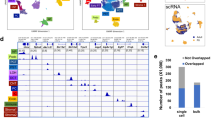
a LocusZoom plots of eGFR GWAS (genotype and eGFRcrea association, n = 1,746,932) and fine mapping for chr4: 105,934,478-106,315,964. Each dot represents a SNP, with color indicating LD association. b LocusZoom plots of eGFR GWAS (genotype and eGFRcrea association, n = 2.2M), kidney CpG cg11878490 meQTL (genotype and cg11878490 methylation association, n = 443), and kidney TET2 eQTLs (genotype and TET2 expression association, n = 686)8. The y-axis displays the -log10 (p-values) of association tests from GWAS, meQTL, and eQTLs studies. c Genotype (rs6533181, x-axis) and normalized CpG methylation (cg11878490, y-axis) in human kidneys (n = 443). The effect size (Beta) is 0.51. Center line represents the median, the box limits show 25th and 75th percentiles, and whiskers extend to 5th and 95th percentiles. P-value was derived from linear regression meQTL model. d From top: Gene browser view of the eGFR GWAS SNPs in the specified regions; single-nucleus Assay for Transposase-Accessible Chromatin using sequencing (snATAC-seq) analysis of chromatin accessibility of human kidneys, including S1, S2, and S3 segments of the proximal tubules (PT-S1, PT-S2, PT-S3), loop of Henle (LOH), distal convoluted tubule (DCT), connecting tubule (CNT), intercalated cells (IC), collecting duct principal cell types (PC), podocytes (Podo), endothelial cells (Endo), immune cells (Immune), lymphocyte (Lymph). Human kidney histone modifications by chromatin immunoprecipitation (ChIP-seq) and chromatin states are also shown. e Relative transcript levels of TET2 after CRISPR-mediated deletion of the locus, with a sample size of n = 3. GAPDH was used for normalization. Data are shown as mean ± SEM and were analyzed using one-way ANOVA followed by Tukey’s post hoc test for multigroup comparison. The diagram was created with BioRender.com. Source data are provided as a Source Data file.
To further dissect this region, we integrated the GWAS data with kidney methylation quantitative trait loci (meQTLs) and expression quantitative trait loci (eQTLs) analyses. We observed a significant association between the variant rs6533181 and CpG methylation (kidney cg11878490 meQTLs, P = 1.29 × 10−19) and TET2 expression in kidney samples (eQTL, P = 1.19 × 10−3) (Fig. 1b, c). Multi-tissue eQTL analysis from the GTEx project (Genotype-Tissue Expression) indicated that rs6533181 is associated with TET2 expression levels also in the skin (P = 1.2 × 10−10) and in fibroblasts (P = 6.1 × 10−5).
To test whether eGFR, cytosine methylation of cg11878490, and TET2 expression share causal variants at this locus, we performed a statistical colocalization analysis using Moloc. We found compelling evidence that the variants linked with kidney function, CpG methylation, and TET2 expression in the kidney tubule (allele-specific expression analysis, ASE) were shared, as indicated by a high posterior probability (ASE.PP.H4.abf = 0.87). TET2 expression in kidney tubules (613 tubule samples) was strongly genotype-dependent (rs6533181, eGFR GWAS risk allele T, ASE P = 9.7× 10−5) (Fig. S1a). The association score in the combined SNP-to-gene (cS2G) linking strategy30 for TET2 was 1, strongly suggesting that TET2 is the target gene of this genetic locus.
Considering that SNPs located in the regulatory region are more likely to be causal31, we employed our human kidney single-nuclear assay for transposase-accessible chromatin sequencing (snATAC-seq) to further prioritize risk variants. The variant rs6533181 was not located in an open chromatin region in the human kidney (Fig. S1b). However, rs1541374 and rs7661349, which are in very strong linkage disequilibrium (LD) with rs6533181 (r > 0.93 and r > 0.98, respectively), were located within an open chromatin region in the proximal tubule cell and the promoter region of TET2 (Fig. 1d). Variant rs1541374 showed the strongest association with kidney function in the eGFR creatinine GWAS (P = 8.63 × 10−11) (Fig. S2a). rs7661349 was also significantly associated with kidney function in the eGFR creatinine GWAS (P = 4.02 × 10−9) (Fig. S2b). We found a significant association between the variants rs1541374 and rs7661349 and CpG methylation (cg11878490 meQTLs, P = 2.98 × 10−18 and P = 1.58 × 10−20, respectively) (Fig. S2a–d). The MAFs of rs1541374 and rs7661349 are 0.3172 and 0.3357, respectively. We observed a significant association between the variant rs1541374 and TET2 expression in kidney tubule samples (kidney tubule eQTL, P = 5.08 × 10−4) (Fig. S2a) and a significant association between the variant rs7661349 and TET2 expression in kidney samples (kidney eQTL, P = 1.20 × 10−3) (Fig. S2b). Allele-specific expression (ASE) analysis for rs1541374 showed that TET2 expression in kidney tubules was strongly genotype-dependent (rs1541374, eGFR GWAS risk allele T, ASE P = 8.3 × 10−3) (Fig. S2e). TET2 expression in kidney tubules was also strongly genotype-dependent for rs7661349 (eGFR GWAS risk allele T, ASE P = 2.2 × 10−4) (Fig. S2f). These observations prompted us to hypothesize that rs7661349 and rs1541374 are the likely causal variants. To demonstrate that rs1541374 and rs7661349 regulate the expression of TET2 in human kidney proximal tubule cells, we performed CRISPR-based deletion of the open chromatin region containing these two variants (Figs. 1e and S3a). Deletion of rs1541374 and rs7661349, but not of the non-target regions, lowered TET2 levels, but not Inorganic pyrophosphatase 2 (PPA2) levels (Figs. 1e and S3b).
Our human kidney single-nuclear RNA sequencing (snRNA-seq) data showed that TET2 expression in the proximal tubule (S3 segment) was lower in diseased kidneys compared to healthy kidneys (Fig. S4a). Immunofluorescence staining further confirmed that the protein level of TET2 in the proximal tubule was lower in diseased kidneys compared to controls (Fig. S4b).
In summary, by integrating kidney function GWAS, single-nucleus epigenome mapping, and CRISPR-Cas9 gene editing, we identified TET2 as a risk gene for kidney disease, specifically within kidney proximal tubule cells, where lower TET2 expression was associated with an elevated risk of renal disease.
Kidney-specific Tet2 knockout mice show increased severity of acute and chronic kidney disease
To study the role of TET2 in kidney disease development, we generated mice with genetically lowered Tet2 expression in kidney cells by breeding Six2Cre mice with Tet2f/f mice32. Gene expression analysis confirmed the reduction in Tet2 expression in the proximal tubules of the kidneys in adult Six2CreTet2f/f mice compared to Six2Cre mice (Fig. S5a, b). Six2CreTet2f/f mice were born at the expected Mendelian ratio, appeared healthy at birth, and showed no significant differences in life span at 40 weeks of age32. Next, we analyzed Six2CreTet2wt/f and Six2CreTet2f/f mice in a cisplatin-induced acute kidney injury (AKI) model (Fig. 2a). Blood urea nitrogen (BUN) levels were elevated in cisplatin-injected Six2CreTet2wt/f and Six2CreTet2f/f mice compared to cisplatin-injected Six2Cre mice (Fig. 2b). Histological analysis indicated more hyaline casts, loss of brush border, and tubular lumen dilation in kidneys of cisplatin-injected Six2CreTet2wt/f and Six2CreTet2f/f mice compared to those in cisplatin-injected control mice (Fig. 2c). Expression levels of kidney injury markers, such as kidney injury molecule 1 (Kim1) and neutrophil gelatinase-associated lipocalin (Ngal), were higher in kidneys of cisplatin-injected Six2CreTet2wt/f and Six2CreTet2f/f mice when compared with cisplatin-injected control Six2Cre mice (Fig. 2d).
a Experimental design. Six2Cre, Six2creTet2wt/f, and Six2creTet2f/f mice were injected with saline or cisplatin and euthanized 3 days later. Created with BioRender.com. b BUN levels were measured in Six2Cre, Six2creTetwt/f, and Six2creTet2f/f mice injected with saline (n = 3) or cisplatin (Six2Cre n = 5, Six2creTetwt/f n = 6, Six2creTet2f/f n = 6). c Representative images of H&E-stained kidneys sections from Six2Cre and Six2creTet2f/f mice injected with saline or cisplatin. d Relative transcript level of Kim1 and Ngal in kidneys of Six2Cre, Six2creTetwt/f, and Six2creTet2f/f mice injected with saline or cisplatin. Gapdh was used for normalization. Saline-injected group: Six2Cre (n = 6), Six2creTetwt/f (n = 4), and Six2creTet2f/f (n = 6). Cisplatin-injected groups: n = 7 each. e Experimental setup. Six2Cre and Six2creTet2f/f mice underwent sham-operation or unilateral ureteral obstruction (UUO) and were euthanized 4 days post-surgery. Created with BioRender.com. f Representative images of H&E-stained kidney sections from Six2Cre and Six2creTet2f/f mice subjected to sham-operation or UUO surgery. g Representative images of Sirius Red-stained kidney sections with quantification from Six2Cre and Six2creTet2f/f mice subjected to sham-operation or UUO surgery. Sham-operated group: Six2Cre (n = 4) and Six2creTet2f/f (n = 3); UUO-operated groups: n = 4 each. h Relative transcript levels of Fibronectin1 (Fn1), Collagen1a1 (Col1a1), Collagen1a1 (Col3a1), Interleukin 6 (Il6), C-C motif chemokine ligand 2 (Ccl2), and Vascular cell adhesion molecule 1 (Vcam1) in kidneys of Six2Cre and Six2creTet2f/f mice after sham-operation or UUO surgery (n = 6 per group). i Representative Western blots and densitometric quantification of FN and alpha-smooth muscle (aSMA) in kidneys of Six2Cre and Six2creTet2f/f mice subjected to sham-operation or UUO surgery (n = 3 biological replicates). j Experimental design for adenine-induced chronic kidney disease model. Six2Cre and Six2creTet2f/f mice were placed on control or adenine diet for 4 weeks. Created with BioRender.com. k Serum BUN levels in Six2Cre and Six2creTet2f/f mice on control (n = 4) or adenine diets (n = 5). l Representative images of H&E-stained kidney sections from Six2Cre and Six2creTet2f/f mice on control or adenine diet. m Representative images of Sirius Red-stained kidney sections with quantification from Six2Cre and Six2creTet2f/f mice on control (n = 3) or adenine diet (n = 5). n Relative transcript levels of Fn1, Col1a1, and Col3a1 in kidneys of Six2Cre and Six2creTet2f/f mice on a control diet (Six2Cre: n = 5; Six2creTet2f/f: n = 4) or adenine diet (Six2Cre: n = 5; Six2creTet2f/f: n = 6). o Representative Western blots and densitometric quantification of FN and aSMA in kidneys of Six2Cre and Six2creTet2f/f mice on a control diet (n = 3 per group) or an adenine diet (n = 4 per group). Scale bar = 20 μm. Data are presented as mean ± SEM and were analyzed using one-way ANOVA followed by Tukey’s post hoc test (b, d, g-i, k, m-o). Data are representative of three independent experiments (c,f,l). Source data are provided as a Source Data file.
To understand the role of TET2 in kidney fibrosis, we subjected both control and Six2CreTet2f/f mice to unilateral ureteral obstruction (UUO) injury (Fig. 2e) and analyzed their phenotypes in an adenine diet-induced chronic kidney fibrosis model (Fig. 2j). Histological examination revealed more severe kidney fibrosis, including more dilated tubules with or without proteinaceous casts, increased immune cell infiltration, and loss of brush border in the kidneys of UUO surgery or adenine-diet-subjected Six2CreTet2f/f mice compared with those in UUO/adenine-subjected Six2Cre mice (Fig. 2f, l). Sirius Red staining indicated elevated profibrotic collagen deposition in the kidneys of Six2CreTet2f/f mice subjected to UUO surgery compared to kidneys of UUO/adenine-subjected control Six2Cre mice (Fig. 2g,m). Expression of fibrosis markers (Fibronectin 1, Collagen1a1, and Collagen3a1) was higher in the kidneys of UUO/adenine-subjected Six2CreTet2f/f mice compared to Six2Cre mice with UUO surgery (Fig. 2h, n) and inflammation markers (Interleukin 6, C-C motif chemokine ligand 2, and Vascular cell adhesion molecule 1) were as higher in kidneys of UUO-subjected Six2CreTet2f/f mice compared to kidneys of UUO-subjected Six2Cre mice (Fig. 2h). Protein expression analysis by immunoblotting further confirmed that fibronectin1 and aSMA expressions were prominently increased in kidneys of UUO/adenine-subjected Six2CreTet2f/f mice compared to UUO-subjected Six2Cre mice (Fig. 2i, o).
As Six2 is expressed during kidney development33, we generated mice with a conditional, inducible genetic deletion of Tet2 in tubule cells by crossing Pax8rtTA mice and TRECre mice with Tet2f/f mice (Fig. 3a). Gene expression analysis confirmed the reduction in Tet2 expression in the kidneys of Pax8rtTATRECreTet2f/f mice compared toTet2f/f mice (Fig. 3b). We observed that Pax8rtTATRECreTet2f/f mice were born at the expected Mendelian ratio and appeared healthy at birth, and we observed no differences in life span at 40 weeks of age. Most importantly, the kidneys of Pax8rtTATRECreTetwt/f mice or Pax8rtTATRECreTet2f/f mice injected with cisplatin showed more severe injury, including changes in kidney injury biomarkers, gene expression, and structural damage, compared to those in the control Tet2f/f mice injected with cisplatin (Fig. 3d–f).
a Experimental scheme for the generation of Pax8rtTACreTRETet2f/f mice. Created with BioRender.com. b Relative transcript levels of Tet2 in kidneys of control and Pax8rtTACreTRETet2f/f mice. Each group contains n = 4 biological replicates. c Experimental design. Tet2f/f, Pax8rtTACreTRETet2wt/f, and Pax8rtTACreTRETet2f/f mice were injected with saline or cisplatin for 3 days. Created with BioRender.com. d Representative images of H&E-stained kidney sections from Tet2f/f, Pax8rtTACreTRETet2wt/f, and Pax8rtTACreTRETet2f/f mice, injected with saline or cisplatin. Scale bar = 20 μm. Data are representative of three independent experiments. e Serum creatinine levels in Tet2f/f, Pax8rtTACreTRETet2wt/f, and Pax8rtTACreTRETet2f/f mice injected with saline or cisplatin. In the saline-injected groups: n = 3 each for each genotype. In the cisplatin-injected groups: Tet2f/f: n = 5, Pax8rtTACreTRETet2wt/f: n = 4, Pax8rtTACreTRETet2f/f: n = 5. f Relative transcript levels of Kim1 in kidneys of Tet2f/f, Pax8rtTACreTRETet2wt/f, and Pax8rtTACreTRETet2f/f mice injected with saline or cisplatin. Saline-injected groups: n = 3 for each genotype. In the cisplatin-injected groups: Tet2f/f: n = 6, Pax8rtTACreTRETet2wt/f: n = 6, Pax8rtTACreTRETet2f/f: n = 5. Data are presented as mean ± SEM and were analyzed using analyzed using a two-tailed unpaired Student’s t-test (b) or a one-way ANOVA followed by Tukey’s post hoc test for multigroup comparison (e-f). Source data are provided as a Source Data file.
In summary, our data indicate that tubule-specific Tet2 loss exacerbates kidney dysfunction in both AKI and CKD.
Tet2 deficiency in tubule cells caused chromosome mis-segregation and accumulation of micronuclei
To investigate the mechanisms by which Tet2 loss in tubule cells contributes to more severe kidney injury, we performed droplet-based snRNA-seq on kidney samples from wild-type (WT) and Six2CreTet2f/f mice at baseline and following UUO injury (Fig. 4a). Following sequencing and alignment, we identified 38,714 high-quality cells. Subsequently, we applied dimensionality reduction using uniform manifold approximation and projection (UMAP) and graph-based clustering (Figs. 4b, c, and S6a). This analysis recognized 19 distinct clusters (Figs. 4b, c, and S6a), which were annotated based on the expression of previously published gene markers (Fig. 4b)34. Tet2 level was markedly lower in Six2CreTet2f/f mice (Fig. S6c). We found that Tet2 is mainly expressed in the proximal straight tubule (PST) (Fig. S6b) which was consistent with the in situ hybridization (ISH) and immunofluorescence (IF) staining data.
a This diagram summarizes nuclear isolation and single-nucleus RNA sequencing (snRNA-seq) of kidneys from wild type (WT) control and Six2CreTet2f/f mice subjected to sham-operation or UUO surgery, using the 10X Genomics platform. Created with BioRender.com. b Bubble plots show representative marker gene expression across 19 cell clusters: Podocyte (Podo), Parietal epithelial cells (PEC), Proximal convoluted tubule (PCT), Proximal straight tubule (PST), Injured proximal tubule (iPT), Descending and ascending thin limb of loop of Henle (DTL_ATL), Thick ascending limb (TAL), Macula densa (MD), Distal convoluted tubule (DCT), Connecting tubule (CNT), Principal cells (PC), Type A and Type B intercalated cells of collecting duct (IC_A and IC_B), Endothelial cells (Endo), Proliferating cells (Prolif), Proliferating stromal (Prolif_Stromal), Myofibroblast (Myofib). The bubble size represents the percentage of positive cells, and color intensity indicates expression level. c UMAP dimension reduction of snRNA-seq from WT control and Six2CreTet2f/f mice subjected to sham-operation or UUO surgery. d Gene Ontology (GO) functional analysis of differentially expressed genes (DEGs) observed in control WT and Six2CreTet2f/f mice after UUO surgery. Enriched biological process (p < 0.05) are shown, with p-values transformed to −log10. Bubble size indicates gene count, and color shows fold change. e This diagram summarizes the RNA sequencing (RNA-seq) process for control and TET2 knockdown RPTEC (CRISPR). Created with BioRender.com. f Relative TET2 transcript levels in control and knockdown RPTEC (n = 3 biological replicates per group). g GO analysis of genes with > 30% reduced expression in TET2 knockdown RPTEC compared to control cells. Enriched terms (p < 0.05) are log-transformed. h Experimental design for primary tubule cells from control and Six2CreTet2f/f mice treated with doxorubicin. Created with BioRender.com. i,j DAPI staining shows chromosome segregation in control and Tet2-knockout primary tubule cells or TET2 knockdown RPTEC. Right panel: Quantifications of doxorubicin-treated cells with chromosome segregation defect (n = 3 biological replicates per group). k,l Representative images of γH2AX and DAPI staining, showing primary tubule cells or RPTEC with or without micronuclei. Right panel: Quantifications of the doxorubicin-treated cells with micronuclei. Each group contains n = 3 biological replicates. Scale bar = 20 μm. Data are presented as mean ± SEM, analyzed with two-tailed unpaired Student’s t-test (f,i–l). Source data are provided as a Source Data file.
We next identified 2,218 differentially expressed genes (DEGs) in PST cells in kidneys from UUO-subjected Six2CreTet2f/f mice compared to UUO-subjected control mice kidneys (with an adjusted p-value < 0.05). Pathway analysis revealed significant enrichment in pathways related to metabolic processes, cell differentiation, and chromosome organization and segregation processes (Fig. 4d).
To complement the mouse kidney analysis, we performed RNA sequencing (RNA-seq) analysis on control and TET2 knockdown (KD) human renal proximal tubule epithelial cells (RPTECs) (Fig. 4e). We generated these cells by transducing RPTECs, which stably express dCas9-ZIM3-KRAB, with either non-targeting control sgRNA or sgRNA targeting TET2 (Fig. 4e). We identified over 2000 genes, including TET2 (Figs. 4f and S6), with a reduction of more than 30% in TET2 KD RPTECs compared to control cells. Pathway analysis of these differentially expressed genes showed enrichment in processes related to nuclear division, mitotic cell cycle, and chromosome segregation (Fig. 4g). We identified 45 genes involved in the chromosome segregation process, which were dramatically lowered in TET2 KD cells, including INO80 and EME1 (Supplementary Data 1).
To confirm the role of Tet2 in chromosome segregation in tubule cells, we next conducted in vitro studies by isolating primary tubule cells from the kidneys of Six2Cre control mice and Six2CreTet2f/f mice (Fig. 4h), both in the presence and absence of doxorubicin, a DNA-intercalating agent (Fig. 4h)35. We observed various types of chromosome segregation errors in doxorubicin-treated primary tubule cells isolated from kidneys of Six2CreTet2f/f mice compared to control cells (Fig. 4i), including micronuclei formation36,37. There were more micronuclei in doxorubicin-treated tubule cells isolated from kidneys of Six2CreTet2f/f mice compared to those from kidneys of Six2Cre mice (Fig. 4k). Similar results were obtained from control and TET2 knockdown human RPTECs treated with doxorubicin (Fig. 4j, l).
In summary, our unbiased gene expression and experimental analyses underscored the critical role of TET2 in chromosome segregation and prevention of micronuclei accumulation within kidney tubule cells.
Loss of Tet2 in proximal tubule cell was associated with greater DNA damage due to the impairment in DNA damage repair
Next, we analyzed RNA-seq data from a substantial collection (n > 400) of control and diseased human kidney samples. We focused on genes whose levels strongly correlated with TET2. We identified 160 genes, all with a false discovery rate (FDR) < 0.05, that correlated with TET2 expression. Gene ontology analysis indicated strong enrichment for genes playing a role in DNA DSB repair via homologous recombination (HR) (Supplementary Data 2). DNA DSBs are highly cytotoxic and must be repaired to maintain chromosomal integrity22,38. To investigate the effects of Tet2 loss on DNA damage in tubule cells, we measured DSB marker γH2AX level in cells treated with or without doxorubicin. We observed that γH2AX level was higher in doxorubicin-treated cells isolated from kidneys of Six2CreTet2f/f mice when compared to doxorubicin-treated cells isolated from kidneys of control mice (Fig. 5a). The protein level of γH2AX was also elevated in kidneys of cisplatin-injected Six2CreTet2f/f mice or Six2CreTet2f/f mice operated with UUO surgery when compared to control cisplatin-injected mice or mice with UUO surgery (Fig. 5b).
a Representative images of immunofluorescence staining for γH2AX and DAPI in tubule cells from control and Six2CreTet2f/f mice treated with doxorubicin. n = 3 biological replicates per group. b Representative Western blots for γH2AX in kidney samples from control Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin, or subjected to sham-operation or UUO surgery. (Right panel) Densitometric quantification of γH2AX normalized to GAPDH. Saline-injected or cisplatin-injected groups: n = 3 per group. Sham-operated or UUO-operated groups: n = 4 per group. c Representative immunofluorescence staining images for p-ATM in tubule cells from control and Six2CreTet2f/f mice treated with or without doxorubicin. n = 3 biological replicates per group. d Representative Western blots for nuclear p-CHK2, p-BRCA1, and p-p53 in tubule cells from Six2Cre and Six2CreTet2f/f mice kidney treated with or without doxorubicin. Lamin B1 is the loading control. n = 3 biological replicates per group. e Double strand break (DSB) repair pathways. Created with BioRender.com. f Bubble plots showing expression of Brca2, Rad51, Xrcc2, and Eme1 in PT clusters. Circle size shows percentage of positive cells, and color indicates gene expression. g Relative transcript levels of BRCA2 DNA repair associated (Brca2), RAD51 recombinase (Rad51), X-ray repair cross complementing 2 (Xrcc2), and Essential meiotic structure-specific endonuclease 1 (Eme1) in kidneys from Six2Cre and Six2creTet2f/f mice after sham-operation or UUO surgery. Gapdh was used for normalization. Sham-operated groups: Six2Cre (n = 5), Six2creTet2f/f (n = 6); UUO-operated groups: n = 6 per group. h Relative expression of BRCA2, RAD51, XRCC2, and EME1 in control and TET2-knockdown RPTECs from RNA-seq analysis. i Relative transcript levels of BRCA2, RAD51, XRCC2, and EME1 in control and TET2-knockdown RPTECs measured by qRT-PCR. n = 3 biological replicates per group. j Representative immunofluorescence images showing γH2AX staining in tubule cells from control and Six2CreTet2f/f mice treated with or without doxorubicin in the presence or absence of PARP inhibitor Olaparib. n = 3 biological replicates per group. k Experimental design: Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin and in the presence or absence of Olaparib. Created with BioRender.com. l Survival curve for Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin and supplemented with or without Olaparib. Saline-injected groups: Six2Cre (n = 5), Six2CreTet2f/f (n = 4); Cisplatin-injected groups: Six2Cre (n = 5), Six2CreTet2f/f (n = 6); Cisplatin and Olaparib-injected groups: Six2Cre mice (n = 9) Six2CreTet2f/f mice (n = 6). Scale bar = 20 μm. Data are presented as mean ± SEM, analyzed using one-way ANOVA with Tukey’s post hoc test (b, g) or a two-tailed unpaired Student’s t-test (h, i). Source data are provided as a Source Data file.
Altered DNA damage response (DDR) during mitosis can lead to the accumulation of damaged DNA and chromosome mis-segregation35,39. We observed higher levels of phosphorylated ataxia telangiectasia mutated kinase (p-ATM), phosphorylated checkpoint kinase 2 (p-CHK2), and phosphorylated p53 (pp53), which are markers of the DDR that response to DSB, in doxorubicin-treated cells isolated from the kidneys of Six2CreTet2f/f mice when compared with those in doxorubicin-treated control cells isolated the kidneys of control mice (Fig. 5c, d). Similarly, the expression of phosphorylated ataxia telangiectasia and Rad3-related kinase (p-ATR) and phosphorylated checkpoint kinase 1 (p-CHK1), which are critical for the DNA SSBs, was elevated (Fig. S8a). Additionally, we observed higher levels of γH2AX and p-p53 in cells with rs1541374 or rs7661349 deletions compared to cells with non-target region deletions (Fig. S8b). These results are consistent with the observation of increased DNA damage and pronounced chromosome segregation errors in tubule cells lacking Tet2, underlining the critical role of Tet2 in maintaining genomic stability.
Unresolved DNA damage may cause cell cycle arrest and apoptosis22. We observed that the level of cleaved caspase 3; a marker of apoptosis, was higher in the doxorubicin-treated primary tubule cells isolated from the kidneys of Six2CreTet2f/f mice compared with those from doxorubicin-treated control (WT) cells (Fig. S8c). Additionally, we observed an increase in both cleaved caspase-3 levels and the number of TUNEL-positive cells, which are indicative of apoptosis, in the kidneys of Six2CreTet2f/f mice injected with cisplatin compared to the kidneys of Six2Cre (control) mice injected with cisplatin (Fig. S8c,d). However, no observable changes in cell cycle progression were detected between doxorubicin-treated tubule cells from the kidneys of Six2CreTet2f/f mice and those from control mice (Fig. S8e).
We hypothesized that DNA damage repair might be impaired in the absence of TET2, which would lead to damaged DNA accumulation. DNA DSBs are repaired by HR and non-homologous end joining (dominant D-NHEJ or alternative Alt-NHEJ) (Fig. 5e)40. The protein level of phosphorylated BRCA1 (p-BRCA1) was lower in the doxorubicin-treated tubule cells isolated from Six2CreTet2f/f mice kidneys compared to those cells isolated from WT kidneys (Fig. 5d). We observed that the mRNA levels of HR pathway genes, including BRCA2 DNA repair associated (Brca2), RAD51 recombinase (Rad51), X-ray repair cross complementing 2 (Xrcc2), and Essential meiotic structure-specific endonuclease 1 (Eme1), was dramatically lower in PT cells of Six2CreTet2f/f mice compared to those in control PT cells from mice with UUO surgery (Fig. 5f). Similar changes were observed in bulk gene expression data from whole kidney lysates of Six2CreTet2f/f mice with UUO surgery (Fig. 5g). In line with the snRNA-seq results, TET2 KD RPTEC showed lower expression of HR pathway genes (Fig. 5h, i). D-NHEJ and Alt-NHEJ pathway genes, including DNA activated, catalytic polypeptide (Prkdc), X-ray repair cross complementing 5 (Xrcc5), X-ray repair cross complementing 1 (Xrcc6), DNA ligase 4 (Lig4), Non-homologous end joining factor 1 (Nhej1), Poly (ADP-ribose) polymerase 1 (Parp1), and DNA ligase 3 (Lig3), were lower in PT cells of kidneys of Six2CreTet2f/f mice compared to kidneys obtained from control mice (Fig. S9a). However, we did not observe change in expression of these genes in kidneys of Six2CreTet2f/f mice following UUO injury (Fig. S9b, c) or in the TET2 KD cells (Fig. S9d–g). PARP1 plays a key role in DNA damage repair. PARP inhibitors such as Olaparib has been recently introduced to the clinic41. We observed higher γH2AX level in tubule cells from the kidneys of Six2CreTet2f/f mice treated with both doxorubicin and the PARP inhibitor Olaparib, compared to those treated with doxorubicin alone (Fig. 5j). Furthermore, Six2CreTet2f/f mice injected with both cisplatin and Olaparib exhibited severe toxicity, leading to mortality by day 2 post-cisplatin injection (Fig. 5k, l). These results are consistent with the key role of PARP in DDR and repair after injury following TET2 loss.
Amongst the genes that showed consistent changes in PT cells of Six2CreTet2f/f mice and RPTECs with TET2 loss was the INO80 complex ATPase subunit (INO80) (Fig. 6b, c, and S10). INO80 is known to play in DNA damage repair (both by HR and NHEJ pathways)42,43,44,45,46. Most importantly, we found that INO80 is a kidney disease risk gene8. Kidney function genome wide association analysis showed a strong association between rs4924532 (P = 1.31 × 10−51) and kidney function (Fig. 6a). This GWAS SNP also showed a strong association of rs4924532 with INO80 level in human kidney tubule sample (P = 6.45 × 10−6 in eQTL). Ino80 expression was higher in PT cells of kidneys of mice with UUO surgery or adenine diet (Fig. 6e, f). To further support the role of INO80, we found increased number of micronuclei in RTPEC following doxorubicin treatment with INO80 knock-down, supporting the role of INO80 in DDR (Fig. 6g, h).
a LocusZoom plot of GWAS (genotype and eGFRcrea association, n = 1,508,659 independent biological replicates). b RNA-seq of control and TET2-knockdown RPTECs showing INO80 expression in these cells. c–f Relative transcript levels of INO80 in control and TET2-knockdown RPTECs, and its levels in kidney samples from Six2Cre and Six2CreTet2f/f mice subjected to sham-operation or UUO surgery or on control or adenine diets. For the control and TET2-knockdown (KD) RPTEC groups, n = 3 biological replicates per group. In the sham-operated and UUO-operated groups, n = 6 replicates per group. In the control diet-fed groups, n = 5 replicates per group. For the adenine diet-fed group, Six2Cre mice had n = 9, and Six2creTet2f/f mice had n = 5. g Relative transcript levels of INO80 in INO80 (CRISPR) knockdown cells compared to control RPTEC, with n = 3 biological replicates per group. h Representative images of immunofluorescence staining showing micronuclei (MN) in control, INO80-knockdown, TET2- knockdown, and TET2 and INO80 double knockdown RPTECs. n = 3 biological replicates per group.Scale bar = 20 μm. Data are presented as mean ± SEM and were analyzed by a two-tailed unpaired Student’s t-test (c,g) or a one-way ANOVA followed by Tukey’s post hoc test for multigroup comparison (e,f). Source data are provided as a Source Data file.
In summary, these results indicate that Tet2 loss causes accumulation of damaged DNA due to impaired DNA damage repair, and chromosome segregation defects in kidney tubule cells.
Role of cGAS-STING activation in Tet2 loss-induced kidney disease development
Cytosolic DNA (such as cytosolic micronuclei) is a strong activator of the cytosolic nucleotide sensors (cGAS/STING)36,47. We therefore wanted to understand whether cGAS/STING activation might mediate the deleterious effect of TET2 loss. We found that cGAS-STING expression was higher in TET2 knockdown cells (Fig. 7a). Gene profiling by RNA sequencing showed that the expression of downstream STING target genes, including Radical S-adenosyl methionine domain containing 2 (RSAD2), 2’−5’-oligoadenylate synthetase 1 (OAS1), MX dynamin like GTPase 1 (MX1), ISG20, IFIT1, IFIT2, and IFIT3, was higher in the TET2 knockdown RPTECs (Fig. S11a), which was confirmed by qRT-PCR (Fig. 7b). The gene expression levels of the type I IFN response gene (Interferon k, Ifnk), interferon-stimulated genes (Isg15, Mx1, and Oas1), and the NF-kB target gene (Il6) were higher in Six2CreTet2f/f mice with UUO surgery compared to those in Six2Cre mice with UUO surgery (Figs. 2h and S11b). Protein levels of cGAS, STING, phosphorylated TBK1 (p-TBK1), phosphorylated IRF3 (p-IRF3), and phosphorylated P65 (p-P65) was higher in cisplatin-injected Six2CreTet2f/f mice compared to cisplatin-treated control Six2Cre mice (Fig. 7c and Fig. S11c). Immunofluorescence staining showed that p-STING level was higher in kidneys of Six2CreTet2f/f mice injected with cisplatin compared to those of Six2Cre mice (Fig. S11d). In situ hybridization showed higher Isg15 expression in kidneys of Six2CreTet2f/f mice injected with cisplatin when compared to those of control Six2Cre mice injected with cisplatin (Fig. 7d). Similar results were obtained in the UUO model of kidney fibrosis (Figs. 7d and S11e).
a GO functional analysis of genes upregulated (1.5-fold) in TET2-knockdown RPTECs compared to controls. Significantly biological process GO terms (p < 0.05) are presented, with p-values as negative log10-transformed. b Relative transcript levels of Isg15, Ifit1, Ifit2, Ifit3, Mx1, and Isg20 in control and TET2 knockdown RPTECs. n = 3 biological replicates per group. c Representative Western blots for cGAS, STING, TBK1, p-TBK1, IRF3, p-IRF3, P65, and phosphorylated p65 (p-p65) in kidney samples from Six2Cre and Six2CreTet2f/f mice subjected to sham operation or UUO surgery. n = 3 per group. d Representative in situ hybridization images of Isg15 (blue) and Hnf4a (red) in kidney sections from Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin, or subjected to sham-operation or UUO surgery. e Experimental design: Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin in the presence or absence of the STING inhibitor C176. Created with BioRender.com. f Serum BUN levels in Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin either in the presence or absence of C176. Saline-injected groups: n = 3 biological replicates per group; Cisplatin-injected groups: n = 4 biological replicates per group; Cisplatin and C176-injected groups: Six2Cre (n = 4) and Six2creTet2f/f (n = 5). g Representative images of H&E-stained kidney sections from Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin either in the presence or absence of C176. h Relative transcript levels of Kim1 and Ngal in kidneys from Six2Cre and Six2CreTet2f/f mice injected with saline or cisplatin, with or without C176. Saline-injected groups: n = 5 samples per group. Cisplatin-injected groups: n = 6 samples per group. Cisplatin and C176-treated groups: n = 6 samples per group. i Representative images of H&E-stained and Sirius Red-stained kidney sections from Six2Cre, Six2CreTet2f/f, Six2CreTet2f/fCgasf/f, and Six2CreTet2f/fStingf/f mice subjected to sham-operation or UUO surgery. j Relative transcript levels of Fn1, Col1a1, and Col3a1 in kidneys from Six2Cre, Six2CreTet2f/f, Six2CreTet2f/fCgasf/f, and Six2CreTet2f/fStingf/f mice after sham-operation or UUO surgery. Sham-operated groups: n = 4; UUO-operated groups: n = 5. Scale bar = 20 μm. Data are presented as mean ± SEM and were analyzed using a two-tailed unpaired Student’s t-test (b) or a one-way ANOVA followed by Tukey’s post hoc test for multigroup comparison (f, h, j). Data are representative of three independent experiments (d, g, i). Source data are provided as a Source Data file.
To prove that cGAS-STING activation mediates kidney injury induced by tubule-specific loss of Tet2, we tested the effect of the STING inhibitor C176 in cisplatin-induced AKI mice (Fig. 7e). The expression of STING target genes, such as Isg15, was lower in kidneys of Six2CreTet2f/f mice treated with C176, indicating the effectiveness of the drug (Fig. S12). BUN levels, a marker kidney function, were lower in C176-treated mice compared to those in cisplatin-treated Six2CreTet2f/f mice (Fig. 7f). Histological examinations showed less severe kidney injury (such as dilated tubules and loss of brush border) in kidneys of Six2CreTet2f/f mice treated with both cisplatin and C176, compared to those treated with cisplatin alone (Fig. 7g). Levels of kidney injury markers, Kim1 and Ngal, were lower in C176-injected animals (Fig. 7h).
To prove the role of cGAS and STING in TET2- mediated kidney injury, we used a genetic approach. We crossed Six2CreTet2f/f mice with Cgasf/f mice or Stingf/f mice (Fig. S13a) and then subjected these animals to UUO injury (Fig. S13b). Kidney protein levels of cGAS and STING confirmed the knockout efficiency of cGAS in Six2CreTet2f/fCgasf/f mice and STING in Six2CreTet2f/fStingf/f mice when compared to kidneys of UUO subjected Six2CreTet2f/f mice (Fig. S13c). Histological analyses revealed reduced injury levels in the kidneys of Six2CreTet2f/fCgasf/f and Six2CreTet2f/fStingf/f mice subjected to UUO surgery, compared to Six2CreTet2f/f mice also undergoing UUO surgery (Fig. 7i). Sirius Red staining indicated less collagen accumulation in kidneys of UUO-subjected Six2CreTet2f/fCgasf/f mice and Six2CreTet2f/fStingf/f mice compared to those in Six2CreTet2f/f mice with UUO surgery (Figs. 7i and S13d). Transcript levels of fibrosis markers, including Fibronectin1, Collagen1a1, and Collagen 3a1, were lower in kidneys of UUO-subjected Six2CreTet2f/fCgasf/f and Six2CreTet2f/fStingf/f mice compared to those in Six2CreTet2f/f mice subjected to UUO (Fig. 7j). The protein level of fibronectin was lower in kidneys of UUO-subjected Six2CreTet2f/fCgasf/f and Six2CreTet2f/fStingf/f mice compared to those in Six2CreTet2f/f mice subjected to UUO (Fig. S13c).
In summary, these findings establish the critical role of the cGAS-STING in mediating Tet2 loss-induced kidney injury and kidney disease development.
Discussion
Here, we demonstrate that TET2 is a kidney disease risk gene using a multi-omics integration of kidney function GWAS fine mapping, human kidney meQTL and eQTL data, single-cell chromatin accessibility annotation, and CRISPR/Cas9-mediated genome editing. Mice and cells with genetic deletion of TET2 in kidney tubule cells showed impaired DNA damage repair, resulting in the accumulation of cytosolic micronuclei, downstream cGAS-STING activation, and the development of kidney fibroinflammation.
TET2 is widely expressed in the body48. It is interesting to note that in the mouse and human kidneys TET2 is almost exclusively expressed in the proximal tubule S3 segment. This is likely related to the presence of a PT/S3 specific enhancer region. The kidney function GWAS variants reside at this tubule enhancer and alter tubule specific TET2 levels. Our study is the first to utilize mice with a kidney/tubule-specific deletion of Tet2, specifically exploring its role in the tubule cells, furthermore we show that the effect is not related to developmental programming as mice with conditional inducible deletion of TET2 in mature tubule cells showed the same phenotype, providing direct evidence of how germline genetic variations can alterTet2 levels in the kidney tubule. Mice with a global Tet2 deletion exhibited more severe acute renal injury following cisplatin injection49 and ischemia-reperfusion injury50. The role of TET2 in diabetic kidney disease has been proposed, based on findings from cell culture studies51.
Loss-of-function somatic mutations in TET2 within blood cell progenitors have recently gained major scientific interest, as they are commonly observed in older individuals52. Clonal hematopoiesis of indeterminate potential (CHIP) is a common aging-related phenomenon in which early blood cell progenitors contribute to the formation of a genetically distinct subpopulation of blood cells53. Somatic mutations most commonly occur in TET2 and DNMT3A53. CHIP was first reported to be associated with myeloid malignancies54, but later studies have uncovered the associations of CHIP with cardiovascular disease55, melanoma progression56, and Parkinson’s disease57. The role of CHIP and somatic TET2 mutations in kidney disease development has also been proposed58,59. Our studies suggest that germline regulatory genetic variants in TET2 predispose individuals to kidney disease development. Future studies shall determine the role of CHIP and TET2 loss in blood cell progenitors, as well as the potential interaction between tubule cells and hematopoietic stem cells in kidney disease.
Our studies indicate the important role of TET2 in DNA damage repair, likely via the HR pathway in kidney disease development. The role of TET2 in DDR in leukemia cells has already been proposed60. TET2 has also been shown to localize to DSB in HeLa cells and human embryonic stem cells and proposed to be important for maintaining genomic integrity61. Future studies shall examine the exact molecular pathways regulated by TET2. Specifically, it will be important to understand whether its role in DDR is related to a non-enzymatic function of TET2 or potentially caused by DNA methylation changes of key genes involved in DDR. Our preliminary analysis indicates that TET2 effect is linked to its enzymatic function. First, we observed important differences in methylated cytosine levels between control, KO mice and those in a disease state (Fig. S13). Our prior studies indicated that Tet2, in conjunction with Tet3, markedly alters kidney development by interfering with the methylation of WNT pathway genes. Finally, we found that vitamin C, which can enhance the enzymatic activity of TET enzymes, ameliorated kidney disease development in mice with lower TET2 levels (Fig. S15).
Our studies indicate a broader role for DDR in kidney disease development beyond TET2. Our integrated GWAS and follow-up prioritization tool identified 57 kidney disease risk genes associated with DDR, including BRCA2 and O-6-methylguanine-DNA methyltransferase (MGMT)8. We identified INO80 as another gene related to DNA repair and associated with kidney disease risk. The INO80 complex is a chromatin remodeling complex that regulates gene expression, DNA repair, and replication42,45. Aging is the strongest risk factor for kidney disease development; however, the exact mechanism of aging-associated kidney disease is poorly understood62. It is known that DNA methylation degrades with aging, and increased damage DNA accumulation and altered DDR are observed during aging24. Future studies shall establish the role of methylation changes and DDR in aging-associated kidney disease development.
Our studies indicate that the Tet2 loss results in accumulation of micronuclei in tubule cells, activating the cGAS-STING pathway. We found that damaged DNA accumulating in injured kidney tubule cells leads to alterations in genes associated with chromosome segregation. The chromosome segregation defect is the likely cause of micronuclei accumulation. Cytosolic DNA, including micronuclei, has been shown to activate the cytosolic DNA sensors such as cGAS and STING47. To demonstrate the critical role of cGAS and STING in tubule cells, we generated mice with a tubule-specific genetic deletion of cGAS and STING. These mice showed protection from TET2-mediated kidney disease, indicating the critical role of this pathway in DDR alterations and kidney disease development. Finally, we show that pharmacological inhibition of STING protected mice from severe disease, indicating that STING inhibitors might be beneficial for patients with kidney disease specifically in patients carrying DDR pathway gene variants.
In conclusion, here we identify TET2 as a kidney disease risk gene, which plays an important role in DNA damage repair, preventing altered chromosome segregation, cytosolic micronuclei accumulation, and cGAS-STING activation in kidney tubule cells.
Methods
Visualization of SNPs in GWAS, meQTL, and eQTL
LocusZoom was used to visualize genomic regions of interest on chromosome 4 for kidney function GWAS, meQTL, and eQTL of TET2 in whole kidneys or kidney tubules8. The meQTL box plot illustrates show the association between reference and alternative alleles of given genotypes and methylation level in kidney samples. The allele-specific expression (ASE) box plot was used to show the association between reference and alternative alleles of given genotypes and TET2 expression in tubule samples.
Bayesian colocalization analysis
As previously published8, we performed Bayesian coloc analysis to identify the variants where the genotype effect on kidney function, methylation, and gene expression was shared. Bayesian coloc analysis was conducted using R package moloc (v0.1.0)63 to estimate the posterior probability that an eGFR GWAS variant is associated with three traits (GWAS, meQTLs and eQTLs).
Animal studies
Mouse studies were approved by the Institutional Animal Care and Use Committee at the University of Pennsylvania. The Tet2f/f (stock #017573), Six2Cre (stock # 009606), Pax8rtTA (stock #007176), TetO-Cre (TRECre) (stock #00623), and Stingf/f (stock #031670) mice were purchased from the Jackson Laboratory. Cgas f/f mice were kindly provided by Prof. Charles Rice (Rockefeller University)64. Tet2f/f was crossed with Six2Cre mice to generate Six2CreTet2f/f mice. Pax8rtTA and TRECre mice were crossed with Tet2f/f to generate Pax8rtTECreTet2f/f mice. Cgas f/f and Sting f/f were crossed with Six2CreTet2f/f mice to generate Six2CreTet2f/fCgas f/f mice and Six2CreTet2ff/ Sting f/f mice, respectively. For the cisplatin injury model, 6-8-week-old mice were intraperitoneally injected with cisplatin (20 mg/kg; dissolved in 0.9% saline) and euthanized on day 3. For the unilateral ureteral obstruction model, 6-8-week-old mice underwent ligation of the left ureter and were euthanized on day 4. For adenine mouse disease model, 8-10-week-old mice were given a control (0.6% calcium and 0.9% phosphorus adjusted diet) or an adenine (0.2% adenine, 0.6% calcium and 0.9% phosphorus) diet and euthanized 4 weeks after starting the adenine food. In the PARP inhibition experiment, Olaparib (50 mg/kg; Tocris Bioscience™, #75-791-0) was administered intraperitoneally 30 minutes prior to cisplatin injection, with subsequent daily administrations following cisplatin injection. In the STING pharmacological inhibition experiment, C176 (750 nm per mouse, Bio Vision, #B2341) or DMSO dissolved in 85 μl corn oil was administered intraperitoneally. This administration occurred at 2 h, 24 h, and 48 h before the injection of cisplatin (20 mg/kg) or saline, and then daily after cisplatin injection. All mice were provided food and water ad libitum and monitored daily.
Single nucleus RNA sequencing
Nuclei were extracted from flash-frozen kidney tissue. Briefly, 10-30 mg of frozen kidney tissue were minced on ice into pieces in lysis buffer (Tris-HC, NaCl, MgCl2, NP40, and RNAse inhibitor), transferred into a gentleMACS C tube, and homogenized using gentleMACS homogenizer. Next, the homogenized tissue was filtered through a 40 μm strainer (Fisher Scientific, #08-771-1) and then centrifuged at 500 × g for 5 mins at 4 °C. The pellet was resuspended in wash buffer (BSA, RNAse inhibitor, 1× PBS) and then filtered through a 40 μm Flowmi cell strainer (Sigma Aldrich, #BAH136800040-50EA). After checking for nuclear quality and counting, 30,000 cells were loaded into the 10X Chromium Controller (PN-120223). The cDNA libraries were constructed using the 10X Chromium library construction kit (PN-1000190). Libraries were subjected to quality control using the Agilent Bioanalyzer High Sensitivity DNA kit (Agilent Technologies, 5067-4626). The following demultiplexing was used to sequence libraries using the Illumina Novaseq 6000 system with 2 × 150 paired-end kits: 28 bp Read1 for cell barcode and UMI, 10 bp i7 index, 10 bp i5 index, and 90 bp Read2 for transcript.
Cell culture and lentiviral production
Primary tubular epithelial cells were isolated from the kidneys of 3-4-week-old mice. Briefly, kidneys were minced into pieces and digested in RPMI 1640 medium (Gibco, #21875-034) containing collagenase I (2 mg/ml, Sigma Aldrich) for 30 min at 37 °C. Next, fetal bovine serum (FBS) was used to deactivate collagenase I activity and cells were sequentially sieved through 100, 70, and 40 μm mesh. After removing red blood cells, the tubules cells were cultured in RPMI 1640 supplemented with 10% FBS, rhEGF (20 ng/ml), 1× insulin-transferrin-selenium (ITS), and 1% penicillin-streptomycin.
Immortalized human RPTECs (ATCC; CRL-4031) were cultured in Dulbecco’s modified Eagle’s medium (DMEM)-F12, supplemented with hydrocortisone (25 ng/ml), ascorbic acid (3.5 µg/ml), sodium selenite (8.65 ng/ml), transferrin (5.0 µg/ml), insulin (5.0 µg/ml), triiodo-L-thyronine (5 pM), prostaglandin E1 (25 ng/ml), rhEGF (10 ng/ml), sodium bicarbonate (1.2 mg/ml), 1% penicillin-streptomycin, and G418 (0.1 mg/ml).
For the cytosolic micronuclei analysis experiments, the primary epithelial tubule cells or RPTECs were exposed to doxorubicin (1 μM) for 25 mins. For the DNA damage analysis experiments, the primary epithelial tubule cells or RPTECs were exposed to doxorubicin (1 μM) for 25 mins and analyzed after 8 h. For the cell cycle analysis and DDR analysis experiments, the primary epithelial tubule cells were treated with doxorubicin (1 μM) for 24 h. For the PARP inhibition experiment, cells were exposed to Olaparib (1 μM) for 1 h and then treated with or without doxorubicin (1 μM) for 8 h.
Human embryonic kidney (HEK) 293 T (HEK293T) cells (ATCC, #CRL-3216) were used for packaging lentivirus. The cells were cultured in DMEM (Mediatech, #MT10-013-CV) supplemented with 10% FBS and 1% penicillin-streptomycin and maintained in a humidified 5% CO2 atmosphere at 37 °C.
For lentivirus production, HEK293T cells were transfected with the plasmid of interest, along with the lentiviral packing and envelope plasmids, pPAX2 (Addgene, #12260, a gift from Didier Trono) and pMD2.G (Addgene, #12259, a gift from Didier Trono), using polyethylenimine (PEI, Biotechne, #7854).
Cell line generation
CRISPRi RPTEC cells were generated by stably transducing RPTECs with lentivirus expressing ZIM3 KRAB-dCas9-mCherry under the UCOE-SFFV promoter. Cells stably expressing mCherry were sorted using fluorescence-activated cell sorting. The plasmid pHR-UCOE-SFFV-dCas9-mCherry-ZIM3-KRAB was a gift from Mikko Taipale (Addgene plasmid #154473). The CRISPRi RPTEC cell line was verified by monitoring mCherry fluorescence over several generations.
CRISPR/Cas9 mediated genomic region deletion
All annealed sgRNA oligos were subcloned into a Cas9 expression plasmid (lentiCRISPR v2, Addgene, #52961, a gift from Feng Zhang) with the Bsmb1 site. All constructs were verified by Sanger sequencing.
Lentivirus was prepared as previously described. Lentivirus-containing supernatants supplemented with polybrene (Santa Cruz Biotechnology, #sc-134220) were used for lentiviral transduction for the human RPTEC. After 72 h, cells were harvested, and TET2 and PPA2 expression was determined by qRT-PCR. At the same time, DNA was isolated, and sgRNAs target region deletion were determined by Sanger sequencing. sgRNA sequences are listed in Supplementary Data 3, and PCR primer sequences are listed in Supplementary Data 4.
CRISPR interference
The sgRNA expressing plasmids were generated as previously published65. Briefly, annealed sgRNA oligos were subcloned into LRG2.1 (Addgene #108098, a gift from Christopher Vakoc) with the Bsmb1 site. All constructs were verified by Sanger sequencing. The sgRNA sequences used in this study are in Supplementary Data 3.
Lentivirus was prepared as previously described. The lentivirus-containing supernatants, supplemented with polybrene, were immediately used for lentiviral transduction of human RPTEC stable expressing dCas9-KRAB-ZIM3.
Human RPTEC RNA Sequencing
RNA was extracted using the RNeasy Mini Kit (Qiagen, Cat# 74106). After quality assessment by Agilent Bioanalyzer 2100, RNA-Seq libraries were generated using the TruSeq RNA Sample Prep Kit (Illumina, San Diego, CA). Low-quality bases were trimmed using Trim-galore, and RNA-seq reads were aligned to the human genome (hg19) using STAR (v2.6.1a) based on GENCODE v29 annotations. RSEM (v1.3.1) was used to quantify gene-level read counts and expressed as transcripts per million (TPM).
RNA extraction and qRT-PCR
RNA was isolated using Trizol reagent (Invitrogen) and was transcribed into cDNA using cDNA Reverse Transcription Kit (Applied Biosystems, Cat. #4368813). qRT-PCR was performed using the SYBR Green Master Mix (Applied Biosystems, Cat. #A25742-PEC). The data were normalized and analyzed using the 2-ΔΔCt method with indicated reference gene. Primer sequences are provided in Supplementary Data 5.
Western blot
Kidney tissue or cultured cells were homogenized in lysis buffer and were separated by SDS-PAGE and then transferred to PVDF membranes. Transferred blots were blocked for 1 h in 5% non-fat milk or 5% BSA in Tris-buffered saline. Membranes were incubated with specific primary antibodies at 4 °C overnight, followed by incubation with a horseradish peroxidase-conjugated anti-mouse antibody (1:5,000), or anti-rabbit antibody (1:5,000), or IR-conjugated anti-mouse antibody (1:20,000), and anti-rabbit (1:20,000) antibody at room temperature for 1 h. The signal was developed with Immobilon forte western HRP substrate (Millipore, #WBKLS0500) using Odyssey®Fc Imaging System (LICOR) equipment and measured with Iamge Studio Lite software. The antibodies used in the study are listed in Supplementary Data 6.
Histological analysis
Kidneys were fixed in 10% neutral formalin. Paraffin-embedded sections were stained with hematoxylin and eosin (H&E). Sirius Red staining was performed to determine the degree of fibrosis. In situ hybridization was performed using paraffin-embedded kidney tissue samples and the RNAscope 2.5 HD Duplex Detection Kit (bio-techne, Cat. #322436) following manufacturers’ original protocol. The following probes were used for the RNAscope assay: Mm-Tet2 (bio-techne, Cat. # 511591), Mm-Isg15 (bio-techne, Cat# 559271-O1), Mm-Hnf4a (bio-techne, Cat# 497651-C2). Immunofluorescence analyzes were performed on paraformaldehyde-fixed cells and formalin-fixed, paraffin-embedded kidney sections. To visualize the expression of proteins in the kidney, paraffin-embedded sections were deparaffinized, rehydrated, and incubated with indicated primary antibodies. Slides were incubated with fluorescent-conjugated secondary antibodies, counterstained and mounted with DAPI for nuclear staining. To visualize the protein expression in the paraformaldehyde-fixed cells, we permeabilized the cells with 0.2% Triton X 100, blocked with 3% BSA, then incubated with indicated primary antibodies and fluorescent conjugated secondary antibodies, and counterstained with DAPI.
BUN and creatinine level
Blood urea nitrogen levels (BUN) was measured using InfinityTM Urea Liquid Stable Reagent (Pointe Scientific, #B7552150). Serum creatinine was detected by Creatinine Enzymatic and Creatinine Standard (DIAZYME #DZ072B-KY1). Both measurements were performed following the manufacturers’ instructions.
Fluorescence-activated cell sorting analysis for cell cycle distribution
Cells were prepared for propidium iodide (Sigma) staining following standard protocols. DNA content was then analyzed using a CytoFLEX Analyzer.
Bioinformatic analysis for single-nucleus RNA sequencing data
We generated a gene-by-cell count matrix by aligning its fastq files to the mm10 reference dataset using 10X Genomics Cell Ranger ARC v2.0.2). The quality control (QC) included removing ambient mRNA contamination, eliminating low-quality cells and genes, and excluding doublet-like cells. We used “SoupX” to remove ambient mRNA. We defined low-quality cells as those with the following criteria: unique molecular identifiers (UMI) ≤ 500 or ≥ 10,000, the number of detected genes ≤300 or ≥ 3000, the percentage of mitochondrially encoded genes reads > 1%, and complex ≥ 0.25. We further removed genes that were expressed in < 10 cells. “DoubletFinder” was used to exclude doublet-like cells. To process the data, we used Seurat 4.0.4.
The data were normalized and scaled using the NormalizeData() and ScaleData() functions in Seurat, and variable features were identified with FindVariableFeatures(). Cluster identities were assigned using expression of known marker genes34. Differentially expressed genes (DEGs) in each cluster were determined using the FindAllMarkers() function with the “MAST” test for transcripts present in at least 10% of cells with an absolute log2-fold-change (log2FC) threshold of 0.25. A Bonferroni adjusted P-value (P-adj) < 0.05 was considered statistically significant.
Quantification and statistical analysis
Statistical analyses were performed using GraphPad Prism software (GraphPad Software Inc., La Jolla, CA). An unpaired, two-tailed Student’s t-test was used to compare two groups. One-way ANOVA was used to compare multiple groups, followed by Tukey’s post hoc test. p < 0.05 was considered significant. All data are shown as mean ± SEM.
Study approval
The University of Pennsylvania institutional review board (IRB) approved the human kidney sample collection. We engaged with an external honest broker who was responsible for human kidney sample collection. No personal data were acquired.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Mouse snRNA-seq data and RNA sequencing data generated in this study have been deposited in the GEO under accession code: GSE273027. The kidney meQTLs, kidney eQTLs, human kidney snATAC-seq data, and snRNA-seq data used in this study are available in the GEO under accession codes: GSE115098, GSE173343, GSE172008, GSE200547, and GSE211785 and are publicly available online at the Susztaklab Kidney Biobank (http://www.susztaklab.com/Kidney_meQTL/index.php; http://www.susztaklab.com/Kidney_eQTL/index.php). The adult human kidney histone modifications (H3K4me3 and H3K27ac) by ChIP-seq used in this study are available in the GEO under accession codes: GSM621648 and GSM1112806. Chromatin states for human adult kidneys were generated using ChromHMM (v1.17), by training a 15-state model to capture all the key interactions between the chromatin marks as previously published8. The data for the eGFRcrea GWAS and kidney tubule ASE used in this study are unpublished (Liu et al. in Revision, https://susztaklab.com/GWAS2M/GWAS.php). Additional details on protocols and special reagents used in this study are provided upon request from the corresponding author. Source data are provided with this paper.
References
Jager, K. J. et al. A single number for advocacy and communication-worldwide more than 850 million individuals have kidney diseases. Nephrol. Dial. Transpl. 34, 1803–1805 (2019).
Kovesdy, C. P. Epidemiology of chronic kidney disease: an update 2022. Kidney Int Suppl. 12, 7–11 (2022).
Collaboration, G. B. D. C. K. D. Global, regional, and national burden of chronic kidney disease, 1990-2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 395, 709–733 (2020).
Ko, Y. A. et al. Genetic-variation-driven gene-expression changes highlight genes with important functions for kidney disease. Am. J. Hum. Genet 100, 940–953 (2017).
Wuttke, M. et al. A catalog of genetic loci associated with kidney function from analyses of a million individuals. Nat. Genet 51, 957–972 (2019).
Hirohama, D. & Susztak, K. From mapping kidney function to mechanism and prediction. Nat. Rev. Nephrol. 18, 76–77 (2022).
Stanzick, K. J. et al. Discovery and prioritization of variants and genes for kidney function in >1.2 million individuals. Nat. Commun. 12, 4350 (2021).
Liu, H. et al. Epigenomic and transcriptomic analyses define core cell types, genes and targetable mechanisms for kidney disease. Nat. Genet 54, 950–962 (2022).
Sullivan, K. M. & Susztak, K. Unravelling the complex genetics of common kidney diseases: from variants to mechanisms. Nat. Rev. Nephrol. 16, 628–640 (2020).
Zhu, Q. et al. Prioritizing genetic variants for causality on the basis of preferential linkage disequilibrium. Am. J. Hum. Genet 91, 422–434 (2012).
Consortium, G. T. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Qiu, C. et al. Renal compartment-specific genetic variation analyses identify new pathways in chronic kidney disease. Nat. Med 24, 1721–1731 (2018).
Sheng, X. et al. Mapping the genetic architecture of human traits to cell types in the kidney identifies mechanisms of disease and potential treatments. Nat. Genet 53, 1322–1333 (2021).
Moore, L. D., Le, T. & Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 38, 23–38 (2013).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Sriraman, A., Debnath, T. K., Xhemalce, B. & Miller, K. M. Making it or breaking it: DNA methylation and genome integrity. Essays Biochem 64, 687–703 (2020).
Jones, P. A. & Takai, D. The role of DNA methylation in mammalian epigenetics. Science 293, 1068–1070 (2001).
Li, S. Y. et al. DNMT1 in Six2 Progenitor Cells Is Essential for Transposable Element Silencing and Kidney Development. J. Am. Soc. Nephrol. 30, 594–609 (2019).
Guan, Y. et al. Dnmt3a and Dnmt3b-decommissioned fetal enhancers are linked to kidney disease. J. Am. Soc. Nephrol. 31, 765–782 (2020).
Yan, Y. et al. Unraveling the epigenetic code: human kidney DNA methylation and chromatin dynamics in renal disease development. Nat. Commun. 15, 873 (2024).
Chatterjee, N. & Walker, G. C. Mechanisms of DNA damage, repair, and mutagenesis. Environ. Mol. Mutagen 58, 235–263 (2017).
Sun, S., Osterman, M. D. & Li, M. Tissue specificity of DNA damage response and tumorigenesis. Cancer Biol. Med 16, 396–414 (2019).
Hayashi, K., Hishikawa, A. & Itoh, H. DNA damage repair and DNA methylation in the kidney. Am. J. Nephrol. 50, 81–91 (2019).
Selfridge, J., Hsia, K. T., Redhead, N. J. & Melton, D. W. Correction of liver dysfunction in DNA repair-deficient mice with an ERCC1 transgene. Nucleic Acids Res 29, 4541–4550 (2001).
Soto-Palma, C., Niedernhofer, L. J., Faulk, C. D. & Dong, X. Epigenetics, DNA damage, and aging. J Clin Invest 132 (2022).
Morris, A. P. et al. Trans-ethnic kidney function association study reveals putative causal genes and effects on kidney-specific disease aetiologies. Nat. Commun. 10, 29 (2019).
Graham, S. E. et al. Sex-specific and pleiotropic effects underlying kidney function identified from GWAS meta-analysis. Nat. Commun. 10, 1847 (2019).
Hellwege, J. N. et al. Mapping eGFR loci to the renal transcriptome and phenome in the VA Million Veteran Program. Nat. Commun. 10, 3842 (2019).
Gazal, S. et al. Combining SNP-to-gene linking strategies to identify disease genes and assess disease omnigenicity. Nat. Genet 54, 827–836 (2022).
Gate, R. E. et al. Genetic determinants of co-accessible chromatin regions in activated T cells across humans. Nat. Genet 50, 1140–1150 (2018).
Liang, X. et al. Tet2- and Tet3- mediated cytosine hydroxymethylation in six2 progenitor cells in mice is critical for nephron progenitor differentiation and nephron endowment. J. Am. Soc. Nephrol. 34, 572–589 (2023).
Kobayashi, A. et al. Six2 defines and regulates a multipotent self-renewing nephron progenitor population throughout mammalian kidney development. Cell Stem Cell 3, 169–181 (2008).
Park, J. et al. Single-cell transcriptomics of the mouse kidney reveals potential cellular targets of kidney disease. Science 360, 758–763 (2018).
Bakhoum, S. F., Kabeche, L., Murnane, J. P., Zaki, B. I. & Compton, D. A. DNA-damage response during mitosis induces whole-chromosome missegregation. Cancer Discov. 4, 1281–1289 (2014).
Krupina, K., Goginashvili, A. & Cleveland, D. W. Causes and consequences of micronuclei. Curr. Opin. Cell Biol. 70, 91–99 (2021).
He, B. et al. Chromosomes missegregated into micronuclei contribute to chromosomal instability by missegregating at the next division. Oncotarget 10, 2660–2674 (2019).
Mehta, A. & Haber, J. E. Sources of DNA double-strand breaks and models of recombinational DNA repair. Cold Spring Harb. Perspect. Biol. 6, a016428 (2014).
Novais-Cruz, M. et al. Mitotic DNA damage promotes chromokinesin-mediated missegregation of polar chromosomes in cancer cells. Mol. Biol. Cell 34, ar47 (2023).
Shibata, A. & Jeggo, P. A. DNA double-strand break repair in a cellular context. Clin. Oncol. (R. Coll. Radio.) 26, 243–249 (2014).
Dias, M. P., Moser, S. C., Ganesan, S. & Jonkers, J. Understanding and overcoming resistance to PARP inhibitors in cancer therapy. Nat. Rev. Clin. Oncol. 18, 773–791 (2021).
Morrison, A. J. et al. INO80 and gamma-H2AX interaction links ATP-dependent chromatin remodeling to DNA damage repair. Cell 119, 767–775 (2004).
Tsukuda, T., Fleming, A. B., Nickoloff, J. A. & Osley, M. A. Chromatin remodelling at a DNA double-strand break site in Saccharomyces cerevisiae. Nature 438, 379–383 (2005).
van Attikum, H., Fritsch, O. & Gasser, S. M. Distinct roles for SWR1 and INO80 chromatin remodeling complexes at chromosomal double-strand breaks. EMBO J. 26, 4113–4125 (2007).
van Attikum, H., Fritsch, O., Hohn, B. & Gasser, S. M. Recruitment of the INO80 complex by H2A phosphorylation links ATP-dependent chromatin remodeling with DNA double-strand break repair. Cell 119, 777–788 (2004).
Gospodinov, A. et al. Mammalian Ino80 mediates double-strand break repair through its role in DNA end strand resection. Mol. Cell Biol. 31, 4735–4745 (2011).
Mackenzie, K. J. et al. cGAS surveillance of micronuclei links genome instability to innate immunity. Nature 548, 461–465 (2017).
Rasmussen, K. D. & Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 30, 733–750 (2016).
Bao, Y. et al. DNA demethylase Tet2 suppresses cisplatin-induced acute kidney injury. Cell Death Discov. 7, 167 (2021).
Yan, H. et al. Ten-eleven translocation methyl-cytosine dioxygenase 2 deficiency exacerbates renal ischemia-reperfusion injury. Clin. Epigenetics 12, 98 (2020).
Yang, L. et al. Effect of TET2 on the pathogenesis of diabetic nephropathy through activation of transforming growth factor beta1 expression via DNA demethylation. Life Sci. 207, 127–137 (2018).
Ferrone, C. K., Blydt-Hansen, M. & Rauh, M. J. Age-Associated TET2 Mutations: Common Drivers of Myeloid Dysfunction, Cancer and Cardiovascular Disease. Int J Mol Sci 21 (2020).
Kessler, M. D. et al. Common and rare variant associations with clonal haematopoiesis phenotypes. Nature 612, 301–309 (2022).
Delhommeau, F. et al. Mutation in TET2 in myeloid cancers. N. Engl. J. Med 360, 2289–2301 (2009).
Ostriker, A. C. et al. TET2 protects against vascular smooth muscle cell apoptosis and intimal thickening in transplant vasculopathy. Circulation 144, 455–470 (2021).
Pan, W. et al. The DNA methylcytosine dioxygenase Tet2 sustains immunosuppressive function of tumor-infiltrating myeloid cells to promote melanoma progression. Immunity 47, 284–297.e285 (2017).
Marshall, L. L. et al. Epigenomic analysis of Parkinson’s disease neurons identifies Tet2 loss as neuroprotective. Nat. Neurosci. 23, 1203–1214 (2020).
Vlasschaert, C. et al. Association of clonal hematopoiesis of indeterminate potential with worse kidney function and anemia in two cohorts of patients with advanced chronic kidney disease. J. Am. Soc. Nephrol. 33, 985–995 (2022).
Sano, S. et al. CRISPR-mediated gene editing to assess the roles of tet2 and dnmt3a in clonal hematopoiesis and cardiovascular disease. Circ. Res 123, 335–341 (2018).
Maifrede, S. et al. TET2 and DNMT3A mutations exert divergent effects on dna repair and sensitivity of leukemia cells to PARP inhibitors. Cancer Res 81, 5089–5101 (2021).
Kafer, G. R. et al. 5-hydroxymethylcytosine marks sites of DNA damage and promotes genome stability. Cell Rep. 14, 1283–1292 (2016).
Kazancioglu, R. Risk factors for chronic kidney disease: an update. Kidney Int Suppl. 3, 368–371 (2013).
Giambartolomei, C. et al. A Bayesian framework for multiple trait colocalization from summary association statistics. Bioinformatics 34, 2538–2545 (2018).
Schoggins, J. W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Cao, Z. et al. ZMYND8-regulated IRF8 transcription axis is an acute myeloid leukemia dependency. Mol. Cell 81, 3604–3622.e3610 (2021).
Acknowledgements
Work in the Susztak lab is supported by NIDDK grants R01DK076077, R01DK132630, R01DK105821, R01DK087635 and P50DK114786.
Author information
Authors and Affiliations
Contributions
This study was led by K.S. K.S. and X.L. designed the study, analyzed the data, and wrote the manuscript with input from other authors. X.L., A.A., and A.N. performed experiments. H.L., H.H., X.L., E.H., J.Z., and K.K. performed computational analysis.
Corresponding author
Ethics declarations
Competing interests
The Susztak lab is supported by Boehringer Ingelheim, Regeneron, Bayer, GSK, Gilead, Jnana, Maze, Novartis, ONO and Novo Nordisk for work that is unrelated to the current manuscript. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Michael Eadon, Vishal Patel and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liang, X., Liu, H., Hu, H. et al. TET2 germline variants promote kidney disease by impairing DNA repair and activating cytosolic nucleotide sensors. Nat Commun 15, 9621 (2024). https://doi.org/10.1038/s41467-024-53798-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-53798-x
This article is cited by
-
DNA damage response and cell fate decisions across the lifespan: from fetal development to age-related respiratory diseases
Cell & Bioscience (2025)
-
The Immunogenic Role of Self-DNA in T Cell Immunity
Clinical Reviews in Allergy & Immunology (2025)