Abstract
Stem cells are a hallmark of animal multicellularity. Sox and POU transcription factors are associated with stemness and were believed to be animal innovations, reported absent in their unicellular relatives. Here we describe unicellular Sox and POU factors. Choanoflagellate and filasterean Sox proteins have DNA-binding specificity similar to mammalian Sox2. Choanoflagellate—but not filasterean—Sox can replace Sox2 to reprogram mouse somatic cells into induced pluripotent stem cells (iPSCs) through interacting with the mouse POU member Oct4. In contrast, choanoflagellate POU has a distinct DNA-binding profile and cannot generate iPSCs. Ancestrally reconstructed Sox proteins indicate that iPSC formation capacity is pervasive among resurrected sequences, thus loss of Sox2-like properties fostered Sox family subfunctionalization. Our findings imply that the evolution of animal stem cells might have involved the exaptation of a pre-existing set of transcription factors, where pre-animal Sox was biochemically similar to extant Sox, whilst POU factors required evolutionary innovations.
Similar content being viewed by others
Introduction
The evolution of animal multicellularity around 700 million years ago was a key step that shaped all aspects of our planetary history. As multicellular entities, most animals including early branching sponges harbor pluripotent stem cells1. Stem cells can indefinitely produce identical copies of themselves or, upon stimulation, can form all the specialized cell types of an organism. Stem cells that can become any somatic cell type are known as pluripotent stem cells, whereas multipotent stem cells are lineage-restricted only giving rise to certain cell types. For example, neural stem cells can differentiate into neurons, astrocytes and oligodendrocytes. In vertebrates, pluripotent stem cells only transiently exist during the early stages of embryo development and are characterized by the expression of a core set of transcription factors (TFs) that induce and maintain stemness. Among these TFs, the Sry-related box 2 (Sox2) and octamer-binding transcription factor 4 (Oct4) are key pluripotency factors that need to be present and active at tightly regulated levels. Their removal or subtle perturbances to their abundance lead to the loss of self-renewal and pluripotency2,3. Sox2, Oct4, and other factors have been described as “Pioneer” TFs, capable of binding their target motifs even in closed chromatin and to DNA wrapped around a nucleosome4,5,6,7. Pioneering TFs are critical to direct cell fate transitions during embryonic development including the zygotic genome activation that precedes the formation of pluripotent stem cells8,9,10,11. To open up chromatin and regulate genes, Sox2 and all other Sox family members encode a conserved 79-amino-acid-long high mobility group (HMG) box domain that mediates sequence-specific binding to and bending at CATTGT-like DNA sequences12,13.
Previous phylogenetic reconstructions showed that the Sox clade lacked any reliable non-metazoan HMG box sequences. Therefore, Sox genes were considered a unique animal-specific sub-family within the broader HMG class14,15,16,17,18. At the onset of metazoan radiation, Sox genes expanded into five major paralogous families: SoxB-F19. As cnidarians, sponges and placozoans harbor neurogenic, pluripotent or peptidergic stem cell populations that express Sox2 orthologues (SoxB group members), likely, the association of SoxB genes with stemness evolved early in animals20,21,22,23. Given that stem cells across animals might not be homologous, it is also possible that SoxB members possess biochemical features that make them more likely to be convergently deployed in stem cell regulation across lineages. The level of functional conservation of Sox members across different animal lineages is currently not well understood.
The role of Sox2 in mammalian pluripotent stem cells is tied to its ability to form DNA-dependent heterodimers with Pit-Oct-Unc (POU) family factors such as Oct45,13,24,25,26. The degree of Sox2/Oct4 cooperativity is key for pioneer binding in naïve epiblast cells of day 4.5 mouse embryos27 and determines the quality and developmental potential of pluripotent stem cells across mammalian species28. The heterodimerization between Sox2 and Oct4 is also essential for the generation of induced pluripotent stem cells (iPSCs) during somatic cell reprogramming conventionally driven by the four-factor Yamanaka cocktail Sox2, Oct4, Klf4 and c-Myc29. Only Sox2 and other SoxB members can induce pluripotency in vertebrates as part of this cocktail, whilst Sox factors from the SoxC, D, E and F families cannot30,31,32. If the Sox2/Oct4 partnership is disrupted with mutations, iPSC generation fails25,33,34,35. POU factors are characterized by the presence of a POU-specific domain and a POU homeodomain connected by a flexible linker that jointly bind the ATGCAAAT sequence (known as the Octamer)13. POU factors form six families (POU1, 2, 3, 4, 5 and 6), where POU5/Oct4 is a vertebrate-specific duplication of a POU3 member36. POU factors were also assumed to be an animal-specific invention14,17,36,37,38,39,40,41.
In this work, we re-examine the evolutionary origins of Sox and POU transcription factors by focusing on unicellular relatives of animals. We find orthologs of these transcription factors in previously uncharted choanoflagellates and filastereans. Since these species are unicellular, they do not form stem cells. We show that choanoflagellate Sox is capable of inducing pluripotency in mouse somatic cells. In contrast, choanoflagellate POU binds DNA motifs distinct from animal homologs and cannot induce pluripotency. We propose that the Sox/POU partnership could have emerged early during animal evolution and was likely driven via molecular exaptation of the previously established Sox-POU-DNA dimerization and binding capacities. Our findings foster our understanding of the molecular evolution of Sox/POU, which we hypothesize could have been critical for the advent of stem cells and animal multicellularity.
Results
Unicellular holozoans possess Sox genes that bind DNA like Sox2
To re-evaluate the evolutionary history of Sox genes, we searched for Sox orthologs in the recently generated genomic and transcriptomic datasets from various single-celled outgroups of animals that, together with animals, are part of the clade Holozoa39,42,43,44 (Fig. 1a). We found Sox-like HMG box sequences in the genomes of the filastereans - Pigoraptor vietnamica (Pvie) and Pigoraptor chileana (Pchi) and in the transcriptomes of the choanoflagellates Mylnosiga fluctuans (Myflu) and Salpingoeca helianthica (Salhel). To validate these hits, we inferred a maximum-likelihood (ML) phylogenetic tree and found that the Pvie, Pchi, Myflu, and Salhel Sox-HMG sequences form a sister clade to animal Sox genes (Fig. 1b, Supplementary Fig. 1). Sox-like hits from other choanoflagellates present longer unstable branches, outside the Sox main clade. This suggests that Sox genes originated before the last common ancestor of multicellular animals, while many unicellular holozoans have secondarily lost their Sox or retained degenerate Sox-like sequences lacking Sox hallmarks such as Tyr72 (Fig. 1c).
a Phylogeny of holozoans and (b) Reduced phylogenetic tree of animal and unicellular Sox. c Sequence logos representing the High Mobility Group (HMG) domain of human Sox genes, Sox-like sequences found in unicellular holozoans, and human TCF/LEF genes. Residues reported to direct DNA and protein interactions are boxed in red and blue, respectively13,28,55. d Structural Models of the Sox2 DNA binding domains (DBD) superimposed with HMG Salpingoeca helianthica (Salhel), Mylnosiga fluctuans (Myflu), and Pigoraptor chileana (Pchi). e The predicted protein structure of full-length Salhel Sox-I by AlphaFold3, with model confidence color-coded. f Energy logos derived from Spec-seq using a set of sequences with one nucleotide difference to the consensus Sox motif (CATTGTT). g–i Binding of the HMG box DBD with apparent Kd shown as mean ± SD (n = independent experiments) (g) mouse Sox2 (n = 4), and Sox17 (n = 3), (h) Salhel Sox-I (n = 3) and Pchi Sox (n = 5), and (i) Sox-like sequences from Salpingoeca rosetta (Salro) (The asterisk shows the lane with 250 nM protein which is the highest concentration used for Pchi Sox) and Monosiga brevicollis (Monbr) to consensus Sox DNA (n = 3). The silhouettes of the species are sourced from PhyloPic (http://phylopic.org). Source data and statistics are provided as a Source Data file.
Importantly, all of the amino acids that are key for the sequence-specific base readout in modern Sox proteins are also conserved within Pvie, Pchi, Myflu, and Salhel Sox-HMG (Fig. 1c, d, Supplementary Fig. 2a), suggesting that unicellular Sox could bind the same DNA motifs as their mammalian counterparts. Regions outside the HMG are not evolutionary conserved, cannot be modeled with high confidence, and are therefore likely structurally disordered (Fig. 1e). To test DNA binding, we selected representative unicellular holozoan Sox sequences from a filasterean (SoxPchi) and a choanoflagellates (Sox-ISalhel) as well as two choanoflagellate sequences from the Sox-like (SoxL) branch Monsiga brevicolis (SoxLMonbr) and Salpingoeca rosetta (SoxLSalro). We purified HMG domains of holozoan Sox and performed a specificity-by-sequencing (Spec-seq45) experiment that quantitatively interrogates sequence specificity to the Sox binding motif. We found that the SoxPchi and Sox-ISalhel proteins exhibit a sequence specificity indistinguishable from mouse Sox2 (Fig. 1f, Supplementary Fig. 2b–d), which we also confirmed with electrophoretic mobility shift assays (EMSAs) (Fig. 1g, h). Unicellular holozoan Sox proteins bind DNA with nanomolar affinity, indicating that target DNA binding is highly specific (Fig. 1h, Supplementary Fig. 2e). In contrast, SoxLMonbr could not bind the Sox DNA and SoxLSalro showed a substantially reduced affinity (Fig. 1i, Supplementary Fig. 2e). Of note, these choanoflagellate Sox-like proteins could be purified with high yield and purity comparable to the Sox (Supplementary Fig. 2f). As a control, we also tested the HMG box domain from capicua (Cic) which does not belong to the Sox family, and verified that it strongly prefers binding to its cognate binding site over the Sox consensus (Supplementary Fig. 2g, h). Collectively, unicellular holozoans encode proteins with sequence characteristics and biochemical properties that are similar to mammalian Sox factors and thus represent bona fide Sox.
Choanoflagellate Sox can induce pluripotency in mouse somatic cells
Because their biochemical properties are remarkably similar to the mammalian Sox2 protein, we wondered if unicellular holozoan Sox can also recapitulate some of the functional activities of mouse Sox2. To test this hypothesis, we opted to perform induced pluripotent stem cell (iPSC) generation experiments that critically rely on the ability of Sox2 to direct cell fate transitions and regulate gene networks leading to the establishment of pluripotency and stemness32. In a typical iPSC reprogramming experiment, somatic cells such as fibroblasts are transduced with the four transgenic Yamanaka factors (Sox2, Oct4, Klf4, c-Myc, OSKM) and after culturing for 1–2 weeks in embryonic stem cell culture conditions containing critical growth factors such as LIF, clusters of iPSC gradually emerge29. Individual colonies are then serially isolated or ‘picked’ and clonal iPSC lines are established. These clonal iPSC lines are then subjected to a panel of assays to verify that they exhibit the hallmarks of pluripotency on a functional and molecular level reminiscent of embryo-derived stem cells (Fig. 2a). We used primary mouse embryonic fibroblasts with a transgenic Oct4-GFP reporter (OG2MEF) that gets activated when cells start to acquire a pluripotent state. GFP-positive cell clusters serve as an initial indicator for a pluripotent state that requires further validation (Fig. 2a).
a Schematic illustration of the procedure of mouse induced pluripotent stem cell (iPSC) reprogramming from mouse embryonic fibroblasts (MEFs) carrying an Oct4-GFP reporter (OG2MEFs) and the establishment of clonal iPSC line for pluripotency validation. b Representative microscope images show iPSC colonies generated by mSox2 and Sox factors of Choanoflagellates on reprogramming day 14. Scale bar, 80μm. Chimeric-Salhel-I (Chimeric-Salhel-Sox-I), HMG of Salhel-Sox-I fused with mSox2 NTD and CTD; Salhel-Sox-I, full-length Salhel Sox-I; Salhel-Sox-II, full-length Salhel Sox-II; Myflu-Sox-I, full-length Myflu Sox-I; Myflu-Sox-II, full-length Myflu Sox-II. c Quantification of iPSC reprogramming efficiency by Sox variants. The heatmap depicts the number of experiments with observation of GFP-positive colonies, with the red frame highlighting the ability of the variants to produce iPSCs with confirmed pluripotency through the establishment of stable clonal iPSC lines. Chim-Salhel-I, Chimeric-Salhel-Sox-I; Chim-Pchi, Chimeric-Pchi-Sox. The box plot shows the reprogramming efficiency of Sox variants normalized by the number of iPSC colonies generated by mSox2. (n = 7 technical replicates in total, 2 biological replicates each with 2 technical replicates and 1 biological replicates including 3 technical replicates). The box displays the interquartile range, with the left edge representing the lower quartile (25th percentile) and the right edge indicating the upper quartile (75th percentile). The median value is shown as a line splitting the box. The silhouettes of the species are sourced from PhyloPic (http://phylopic.org). d Representative images of iPSC colonies derived from MEFs carrying a Sox2-GFP reporter on reprogramming day 14. Scale bar, 80 μm. e Expression of pluripotency markers of clonal iPSC lines derived by choanoflagellate Sox examined by immunocytochemistry staining. Scale bar, 40μm. f Immunocytochemistry of differentiated iPSC lines stained for markers of the 3 germ layers: Class III beta-tubulin (Tuj1), Forkhead box protein A2 (FoxA2), α-smooth muscle actin (SMA). Scale bar, 40 μm. g Chimeric mice generated from full-length Salhel-Sox-I iPSC lines displaying black coat patches and eyes (indicated by arrows) representing their iPSC origin, in contrast to the wildtype mouse exhibiting a white coat and red eyes. The illustration for (a) was created in BioRender. Gao, Y. (2022) BioRender.com/z21g065. d n = 2 replicates; (e, f) n = 3 replicates. Source data and statistics are provided as a Source Data file.
We transduced fibroblasts with transgenes encoding the OSKM factor cocktail via doxycycline-inducible lentiviruses. When using mouse Sox2, dozens of GFP-positive iPSC colonies emerged in <2 weeks in a typical experiment. When Sox2 is omitted and replaced by mCherry or the non-Sox HMG box factor Cic, pluripotency reprogramming is derailed and Oct4-GFP positive colonies cannot be detected (Supplementary Fig. 3a, b). Likewise, when Sox2 is replaced with its paralog Sox17 (member of SoxF family) pluripotency reprogramming fails31,32,33 (Fig. 2b, c). To evaluate the capacity of unicellular holozoan Sox to induce pluripotency, we used four choanoflagellate Sox (Salhel-Sox-I, Salhel-Sox-II, Myflu-Sox-I, Myflu-Sox-II) and two choanoflagellate Sox-like factors (SoxLMonbr and SoxLSalro). We also generated chimeric Salhel-I and Pchi constructs, in which their respective HMG box domains were flanked by the N- and C-terminal tails of mouse Sox2 (Chimeric-Sox-ISalhel and Chimeric-SoxPchi, Supplementary Fig. 3c), as a way to test if the differences in pluripotency induction depend on regions beyond the DNA binding domain. SoxLMonbr never generated GFP-positive cell clusters (Supplementary Fig. 3d). Chimeric-SoxPchi (Fig. 2b) and SoxLSalros (Supplementary Fig. 3d) occasionally generated cell clusters with weak GFP-positive signals. However, these GFP-positive cells could not be maintained beyond two passages. Upon isolation, these cells underwent differentiation, and the GFP signal gradually faded away, indicating an incomplete reprogramming and the inability to self-renew. By contrast, all four choanoflagellate Sox and the chimeric-Sox-ISalhel reproducibly generated Oct4-GFP positive colonies (Fig. 2b, c). iPSCs could also be generated with mouse embryonic fibroblasts from another mouse genetic background where a GFP reporter is controlled by the Sox2 promoter instead of an Oct4 promoter (Fig. 2d).
Using RT-qPCR, we demonstrated that endogenous Sox2 is absent in the starting MEFs as well as during early and intermediate stages of reprogramming and only gets activated by the time Oct4-GFP positive pluripotent colonies appear (Supplementary Fig. 4a). This verifies that iPSC generation is driven by exogenously provided Sox factors. The identity of the poorly conserved and structurally disordered N/C-terminal flanks has only a minor effect on reprogramming activity as the full-length Sox-ISalhel as well as the chimeric Sox-ISalhel with Sox2 flanks, support iPSC generation (Fig. 2b). Likewise, the reciprocal chimeric constructs consisting of the mouse Sox2 HMG domain flanked by N/C-termini of Salhel-I or Salhel-II can induce pluripotency (Supplementary Fig. 4b, c). Whilst the identity of the flanks appears interchangeable despite the lack of sequence conservation and detectable structured domains, completely removing the flanks is detrimental as the isolated HMG domain of Sox2 fails to reprogram (Supplementary Fig. 4b, c)30.
To establish clonal iPSC lines for the validation of stemness and pluripotency, we serially isolated (‘picked’) cells from individual GFP-positive colonies in three successive rounds (Supplementary Fig. 5a) and verified the identity of exogenous Sox factors by genotyping (Supplementary Fig. 5b). These clonal iPSC lines displayed comparable self-renewal capacity to iPSCs reprogrammed by mouse Sox2, as they could be maintained for over 15 passages (Supplementary Fig. 5c) and express pluripotency markers such as Nanog at both the transcript (Supplementary Fig. 5d) and protein level (Fig. 2e). To verify the pluripotency of clonal iPSC lines obtained with choanoflagellate Sox in vitro, LIF was withdrawn, and cells were cultured in non-adherent conditions (Methods) initiating spontaneous differentiation into cell lineages representing all three germ layers (Fig. 2f). To verify the pluripotency of our iPSC lines in vivo, we microinjected the cells into blastocyst-stage mouse embryos which were then transplanted into pseudopregnant mice. The live offspring exhibited clear chimerism, characterized by patches of black coat and black eyes from the iPSC donor’s genetic background which highlighted the developmental potential of iPSC lines generated with choanoflagellate Sox in vivo (Fig. 2g). Altogether, choanoflagellate Sox can replace Sox2 in somatic cell reprogramming to iPSCs and induce self-renewing pluripotent stem cells. This demonstrates that choanoflagellate Sox possesses all the molecular features necessary to act as a pioneer factor and induce stemness in the context of a mammalian cell.
Choanoflagellate Sox can cooperate with mammalian Oct4 on DNA
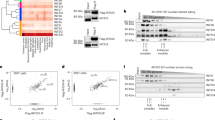
In mammalian pluripotent stem cells and during iPSC reprogramming, Sox2 partners with Oct4 on a composite binding site termed canonical SoxOct element with juxtaposed Sox (CATTGTC) and Oct (Octamer: ATGCAAAT) half sites (Fig. 3a)5,27,46,47,48. The composite SoxOct DNA element is found in the regulatory enhancer DNA of many pluripotency genes and supports the DNA-dependent heterodimerization of Sox2 and Oct425,26,46,47. In vitro, dimerization can be measured using gel shift assays that separate free DNA, Sox-bound, POU-bound, and dimerically bound DNA followed by calculation of cooperativity factors that use the relative fractional contributions of the four DNA states as input49,50 (Supplementary Fig. 6a, b). Given the intimate association of Sox2 and Oct4 in stemness induction, we wondered whether and how holozoan Sox dimerize with mouse Oct4. In control experiments, we verified that mouse Sox2/Oct4 forms dimers on canonical SoxOct DNA probes (Fig. 3a). Intriguingly, both Sox-ISalhel and SoxPchi could heterodimerize with Oct4 on the canonical SoxOct element but Sox-ISalhel does so more effectively with higher cooperativity than SoxPchi (Fig. 3b, c, Supplementary Fig. 6c).
a–f Heterodimer EMSAs with 50 or 100 nM Cy5 labeled canonical SoxOct DNA elements to monitor the heterodimer formation of POU factors (a–c) 150–190 nM mOct4 or (d–f) 50 nM mBrn2 with Sox factors - (a–d) mSox2 (a–e) Salhel Sox-I or (c–f) Pchi Sox HMG. POU factors are kept at a constant concentration indicated by + signs, triangles indicate different concentrations of Sox with the highest concentration indicated, and – sign indicates absences of either Sox or POU or both for controls. g Quantifications of heterodimer EMSAs and calculation of cooperativity factors according to (Ng et al. 2012) with the y axis depicted in log10 scale (mean ± SEM) with n = independent experiments. Oc4/Sox2 (n = 4), Oc4/Pchi Sox (n = 3), Oct4/Salhel Sox-I (n = 4) and Brn2 with Sox2, Pchi Sox and Salhel Sox-I (n = 3). Adjusted p-values are shown and were determined from a Games-Howell test with a 0.95 confidence interval after Bartlett test of homogeneity for each dataset (mOct4 - p = 2.06E-08, mBrn2 p = 0.003637) and Kruskal-Wallis test(One-way). h, i Structural models of heterodimer complexes on canonical SoxOct motifs of (f) Salhel Sox-I HMG-mOct4 POU complex or (g) Pchi Sox HMG-mOct4 POU complex highlighting differences at the heterodimer interface (i.e. positions 57, 61 and 64 previously predicted to impact dimer formation). Source data and statistics are provided as a Source Data file.
Since Oct4 is a POU5 member, a family only found in vertebrates36, we also tested dimerization of the Sox factors with the mouse POU3 factor Brn2. POU3 is the sister group to POU5, and POU3 factors are found across invertebrates including sponges. Particularly, Brn2 is expressed in mammalian multipotent neural stem cells where it can partner with Sox251,52,53. Sox-ISalhel dimerizes with mouse Brn2 on canonical SoxOct DNA with positive cooperativity indistinguishable from Sox2 (Fig. 3d–g). In contrast, SoxPchi shows weak dimerization with cooperativity < 1. Since POU3 factors are ancestral in animals, a SoxB/POU3 partnership may have evolved earlier than the prominent SoxB/POU5 dimer we see in vertebrates, and might even occur in non-bilaterian animals.
To investigate the structural basis that might underlie differences between Sox-ISalhel and SoxPchi to partner up with Oct4 and related POU proteins, we next compared structural models of Sox/Oct4 dimers for the Sox factors (Fig. 3h, i). We selected three amino acids for rational mutagenesis taking into account (i) amino acids previously found to affect dimerization and function and (ii) amino acids that differ between Sox factors that can induce pluripotency and those that cannot24,28,31,50 (Fig. 3h, i). One of the amino acids that determines Oct4 interactions and the ability to induce pluripotency is located at position 57 of the HMG box. iPSC reprogramming incompetent Sox17 encodes a glutamic acid at this position and Sox2 a lysine. A Sox2 Lys57Glu (Sox2KE) mutation disrupts the ability of Sox2 to dimerise with Oct4 and to induce pluripotency33. Likewise, mutating position 57 in Sox-ISalhel into a glutamic acid impairs the ability of mutant Sox-ISalhel to generate iPSCs (Supplementary Fig. 7a, b). Conversely, replacing two interface amino acids at positions 61 and 64 of SoxPchi with their counterparts found in mouse Sox2 re-engineers a gain-of-function Sox-IPchi double mutant that is now capable of inducing pluripotency (Supplementary Fig. 7b-h, Supplementary Fig. 8). We conclude that the cooperative dimerization of Sox-ISalhel with Oct4 on DNA sequence signatures found in pluripotency enhancers is key for its ability to induce stemness in mice. The ability to induce stemness of holozoan Sox correlates with their ability to cooperate with mammalian Oct4 on the canonical SoxOct element.
Ancestral Sox proteins can function as pluripotency inducer in mice
To understand how the amino acid positions that mediate the interaction between Sox and Oct4 and DNA evolved, we performed ancestral sequence reconstruction (ASR) which uses an alignment, a phylogenetic tree, and a probabilistic model of sequence evolution to infer the maximum a posteriori (MAP) sequences (the best estimates of the ancestral states) of ancestral proteins54. We first used the ML phylogeny, inferred the posterior probabilities and MAP sequences of ancestral Sox proteins at key evolutionary nodes, including the last common ancestor of animals and unicellular Sox sequences, as well as shallower ancestors at the base of major Sox clades (Fig. 4a, Supplementary Fig. 9a, b). All these sequences possess the signature amino acids that mediate the base-readout of the Sox DNA element (R5, F10, R18, N30, S34, W41, Y72) (Fig. 4b)55. In this set of inferred ancestors, all ancestral nodes along the trajectory from our deepest ancestor to the SoxB lineage contain K57. Residue 57 is one of the critical determinants for pluripotency induction and maintenance30,33,56,57 as well as the dimerization with Oct413,24,49,58. Only the ancestral SoxCEF node (and its descendants) encodes E57 which is detrimental to the induction and maintenance of stemness (Fig. 4b)33,57.
a Section of the maximum likelihood phylogeny of holozoan Sox HMG domains used for ancestral sequence reconstruction. Colored spheres mark reconstructed nodes. The full phylogeny is shown in Supplementary Fig. 1. b Sequence alignment showing selected key residues of the ancestral Sox HMG domains reconstructed from the phylogeny shown in (a). The arrow marks residue 57, which is critical for the selective pairing with Oct433. c Representative microscope images of iPSC colonies on day 14. Scale bar, 80 μm. d Quantification of iPSC reprogramming efficiency normalized to the colony numbers of mSox2 on day 14 (n = 5 technical replicates in total, including 2 biological replicates each with 2 technical replicates and 1 biological replicates). The box displays the interquartile range, with the left edge representing the lower quartile (25th percentile) and the right edge indicating the upper quartile (75th percentile). The median value is shown as a line splitting the box. Source data and statistics are provided as a Source Data file.
We next synthesized ancestral Sox HMG domains derived from this tree flanked by N- and C-terminal tails of Sox2 to perform iPSC generation assays (Supplementary Fig. 9c). We discovered that all ancestral nodes along the evolutionary trajectory leading to Sox2 are capable of pluripotency induction (Fig. 4c, d). Solely, the branch leading to SoxCEF families lost this capacity most likely due to the substitution of lysine 57 with glutamic acid.
In the ML phylogeny, the choanoflagellate Myflu, and Salhel Sox-HMG sequences form a monophyletic clade with the filasterean Pvie and Pchi sequences but not with other choanoflagellate sequences. This gene tree topology deviates from the expected holozoan species tree, and would imply either incomplete lineage sorting or additional gene duplications and recurrent differential losses in holozoans. We therefore also performed constrained tree (CT) searches, ensuring all Sox/Sox-like sequences followed the holozoan species tree topology. This constraint resulted in somewhat unstable tree searches, yielding 5 subtly different topologies with slightly lower likelihoods than our original ML tree. However, none of these trees were rejected by the approximately unbiased (AU) test59 (Supplementary Fig. 10), implying that our constraint is not ruled out by the data and that Sox-like sequences might be divergent Sox genes that in our original ML tree branch as sister to the rest due to accelerated evolution. We therefore also performed ASR using these alternative constrained phylogenies and compared the inferred ancestral Sox sequences (Supplementary Fig. 11). The inferred ancestral metazoan and holozoan Sox sequences were not identical, but all CT ancestors possessed the signature residues for Sox DNA element binding. This further suggests that the ability to recognize the Sox DNA motif predates the origin of animals. At position 57, four out of five CT ancestors exhibited K57, similar to the ML ancestors, while one exhibited Q57. As K57, Q57 is expected to be compatible with pluripotency reprogramming56. These results imply that regardless of the exact gene history of these proteins, the amino acid states required for DNA binding, POU partnership and iPSC generation were already present before the evolution of animals.
Choanoflagellates encode an atypical POU protein
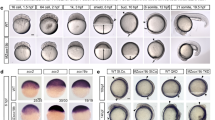
Since we found that Sox-ISalhel could interact with mammalian Oct4 on pluripotency enhancer DNA, we wondered whether POU factors and the Sox/POU partnership could have evolved before the origin of animals. In contrast to previous reports, we found high-confidence hits for Oct4/POU in the choanoflagellate transcriptomes of S. helianthica and M. fluctuans, the same species encoding Sox genes, while being absent from the rest of the 20 available choanoflagellates. Choanoflagellate POU factors similarly to their metazoan homologs, contain a bipartite domain composed of a POU-specific domain (POUS) and a POU-homeodomain (POUHD, Fig. 5a, Supplementary Fig. 12). These choanoflagellate POU homeodomains branch as a sister group to metazoan POU members in a phylogeny encompassing all homeodomains (TALE and non-TALE) across holozoans (Fig. 5a), implying that this is not a convergent acquisition of a similar domain architecture in choanoflagellate sequences. Using a focused phylogeny including the POU-specific domain alongside the homeodomain, we could recover the previously defined major POU families (1, 2, 3/5, 4, 6, Supplementary Fig. 13a). When adding few closer outgroups (LHX, SIX, ONECUT) choanoflagellate POU sequences clustered with orphan sponge POU genes nested within the POU lineage (Supplementary Fig. 13b). This confirms the allegiance of choanoflagellate POU to metazoan POU yet does not place them within any of the families present in the last common ancestor of animals (POU1, 6, and 3).
a Phylogenetic tree of holozoan homeodomains with a focus on POU branches. b Sequence logos of signature amino acids representing the bipartite POU domain of mouse and unicellular POU sequences. Residues reported to be relevant to DNA binding are boxed60. c Binding of the POU of mOct4 and Salhel to consensus Octamer DNA (n = 3 independent experiments). d Energy logos derived from Spec-seq using a set of sequences with one nucleotide difference to the consensus Octamer motif (ATGCTAAT). e Correlation scatter plots of the relative binding affinities of mOct4 versus Salhel POU for all 705 sequences tested. R = Pearson’s correlation coefficient. The color indicates the residual score from the correlation line. f Top enriched motifs from high throughput-SELEX (3rd Cycle) for mOct4 and Salhel POU. g Whole well scan and representative microscopy images of generated iPSCs with activated GFP expression on reprogramming day 14. Scale bar, 80 μm. h Quantification of iPSC reprogramming efficiency of choanoflagellate POU factors normalized to the colony numbers of mOct4 on day 14 (n = 6, 3 biological replicates each with 2 technical replicates). The box displays the interquartile range, with the left edge representing the lower quartile (25th percentile) and the right edge indicating the upper quartile (75th percentile). The median value is shown as a line splitting the box. Source data and statistics are provided as a Source Data file.
The POUSalhel and POUMyflu show deep conservation at several key positions in the POU-specific domain, in particular at helix three involved in sequence-specific DNA recognition via the major groove (i.e. Q44, T45, and R49 of the POUS) (Fig. 5b, Supplementary Fig. 12). Oct4 and other members of the POU family bind the 8 base pair ATGCAAAT octamer motif in a monomeric binding configuration whereby the POU-specific domain bind the ATGC and the POU-homeodomain the AAAT half-sites13,60.
To test the DNA binding specificity of POUSalhel, we performed EMSA titrations to the canonical Oct4 octamer motif with mouse Oct4 and POUSalhel. Interestingly, whereas Oct4 binds this DNA with a binding constant of ~17 nM and a distinct monomeric band, POUSalhel binds the octamer with lower affinity and diffuse shifts (Fig. 5c). This observation suggested that POUSalhel prefers to bind different DNA motifs. Indeed, Spec-seq and HT-SELEX analysis revealed that POUSalhel has no preference for the octamer (Fig. 5d-f). Rather, POUSalhel binds TAAT-like motifs characteristic of most other homeodomain factors61. One of the reasons for this binding mode could be the lack of C50 conserved within and characteristic for the POUHD of animal POU factors that is substituted by a Q50 in choanoflagellate POU (Fig. 5b). Q50 is directing DNA base readout and is present in many ‘non-POU’ homeodomains62. Since the interaction of Oct4 and Sox requires the presence of the composite SoxOct motif of which the octamer motif is an essential component, these findings suggest that pre-animal POU may not yet have evolved the capacity to engage with Sox proteins on enhancer signatures of pluripotency genes. We, therefore, tested the ability of POUSalhel and POUMyflu to induce stemness in mouse fibroblasts and, as expected, found that both factors are incompetent for pluripotency reprogramming (Fig. 5g, h). Collectively, the POU domain emerged earlier than previously thought before the last common ancestor of animals and choanoflagellates. Yet, unicellular POU factors, and presumably pre-animal POU factors, possess unique biochemical characteristics distinct from their animal counterparts.
Discussion
In this study, we show that Sox and POU, two of the most important TF families associated with mammalian pluripotency, emerged before the origin of animals, pre-dating stem cells in their multicellular context (Fig. 6). Sox and POU factors are therefore older than previously thought. The somatic cell reprogramming of mouse fibroblasts into iPSC requires Sox proteins to bind and open closed chromatin7,25,47. Their molecular capacity to bind, deform, and open up closed chromatin as pioneer factors might have primed Sox proteins for being deployed as regulators of diverse stem cells across animals4,5,27. It will be of interest to test with direct data from early branching animal lineages whether this pioneering activity is widespread across extant lineages. Our data show that many of the biochemical features of mammalian SoxB factors can be found in unicellular holozoans and the reconstructed sequences predating the last common ancestor of animals. We propose that these features might have facilitated Sox co-option into gene regulatory networks of stem cells. In unicellular holozoans, a regulatory role of Sox factors in driving cell fate conversions might be less critical, which could explain the pattern of rampant loss of bona fide Sox across the surveyed taxa (Fig. 6a). Yet, with the appearance of animals, Sox factors are retained across all extant lineages indicating functional essentiality. Since many unicellular holozoan species we surveyed are transcriptomes or draft genomes, higher-quality genomes from unicellular holozoans might reveal Sox or POU sequences in more species. More broadly, our study shows that no single extant species can be used as the only representative outgroup of animals63. Frequent gene loss and millions of years of independent adaptation resulted in a mosaic of ancestral and derived traits in extant unicellular holozoans, beautifully exemplified by the patchy distribution of these critical TFs. Even so, the remarkable conservation of function and the striking capacity of a unicellular Sox to direct iPSC induction in mammals shows how the complexity of multicellular gene regulation has deep roots in our unicellular ancestry.
a Phylogenetic distribution of surveyed Holozoan species marked with the presence of Sox and POU factors (green), and Sox-like and POU-like (Homeobox without POUs domain) in yellow. Asterisks indicate that available data for the species is a transcriptome, whereas the rest are genome assemblies. b Schematic for the proposed evolutionary origins of the core members of the mammalian pluripotency regulatory network (Myc, Sox, POU, Klf4 and Nanog), with molecular changes acquired along the evolutionary tree. On top, the proposed model for the evolutionary innovation from pre-metazoans to vertebrates where POU factors became binders for the new octameric POU DNA motif which made them capable of the DNA-dependent heterodimer formation Sox.
The Sox family underwent several rounds of duplications in animals19. Some newly emerging families might have lost some of their ancestral characteristics exemplified by the appearance of pluripotency incompetent ancestral sequences of SoxC/E/F classes (Fig. 4a, b). We propose that the introduction of E57 was a critical step for the gene family expansion of animal Sox as it allowed SoxC/E/F members to diversify their partners and DNA targets, allowing non-overlapping regulatory networks. SoxB factors retain K57 across animals, which we speculate might be critical for SoxB to function in stem cells in cnidarians, sponges, ctenophores, placozoans and bilaterians, including planarian neoblasts18,20,21,22,23,64. Whilst the conservation of a key roles of SoxB across animals stem cells remain to be further examined, there is cumulating evidence for a widespread use of this factors as stem cell regulators in non-bilaterian animals. Two demosponge species express SoxB in the assumed pluripotent stem cell in this lineage, the archeocytes1,65,66. The cnidarian Hydractinia symbiolongicarpus expresses a SoxB paralogue in the i-cells, the adult pluripotent stem cells in hydrozoans. However, not all SoxB expressing cnidarian stem cells are pluripotent, for example in the hydrozoan Hydractinia and the anthozoan Nematostella vectensis, SoxB factors are crucial in the specification of neural progenitors21,67. Still, in Nematostella, a SoxC factor is upstream to a SoxB (SoxB2a) in the cascade of differentiation that leads to neural progenitors, highlighting that SoxB-characteristics might be useful but not indispensable for stem cell maintenance68. Another Nematostella SoxB (SoxB(2)) is a marker of adult stem cells69. In placozoans, the peptidergic progenitor cells also express a combination of SoxB and SoxC23. In the future, the link between Sox and stem cells in non-bilaterian animals will require mechanistic approaches to determine the contribution of these TFs to stem cell maintenance. A systematic analysis of the biochemical properties of SoxB factors across extant animals and the evaluation of their capacities to act as reprogramming factors would clarify if there is a continuity and conservation of the activities we observed for Choanoflagellate Sox and ancestrally reconstructed Sox factors.
Since the emergence of Sox pre-dates the origin of stem cells, the remaining question is when and how a Sox-driven stem cell gene regulatory network evolved in animals. Our data shows that choanozoan Sox interacts with Oct4 in vitro, making POU an attractive candidate whose evolution could have aided the emergence of stem cell networks. Unicellular POUs seem to behave quite differently from their animal counterparts, as they bind different DNA motifs and consequently cannot induce mouse pluripotent stem cells, even in the presence of Sox2. Choanoflagellate POU binds to a typical Homeodomain motif, a motif shared by many other homeodomains including PRD, ANTP or LIM classes not belonging to the POU class61. Animal POU factors gained a new binding motif: the ATGCAAAT octamer. However, the association of POU with animal stem cells is less clear-cut than for Sox. Specifically, Oct4 (POU5) evolved as a vertebrate-specific paralogue of POU336,41 (Fig. 6b). Thus Oct4 is not a general animal pluripotency marker. Oct4’s ability to regulate pluripotency in vertebrates is tied to its capacity to dimerize and cooperate with Sox2. In fish, dimers of Sox2 and Oct4 orthologues drive the zygotic genome activation indicating the conservation of this partnership across vertebrates8,9. Sox2 can also interact with other POU members including more deeply conserved POU349,51,53. POU3 are already present in sponges – the earliest branching animal (Supplementary Fig. 13). Insect SoxB functionally interacts with POU factors suggesting a deep conservation of this partnership70,71. Thus, given the capacity of a choanoflagellate Sox to interact with the POU3 factor Brn2 (Fig. 3e, g), analogous partnerships between Sox and octamer binding POU members might have been important for the evolution of stem cell regulatory networks (Fig. 6b). Potentially supporting this claim, a member of the POU family plays an important role in establishing adult pluripotent stem cells in the cnidarian Hydractinia symbiolongicarpus72. But data from other non-bilaterians would be needed to determine if POU factors are frequently involved in stem cells across various lineages or if this represents a convergent deployment of POU to stem cells in this specific cnidarian clade. In the future, comprehensive analysis of POU and Sox interactions in extant non-bilaterians will be necessary to further test this hypothesis. This potential gain of complexity of TF interactions would be consistent with evolutionary trends described for other structurally distinct TF families such as bZIP. TF heterodimerization networks increase in the advent of animal multicellularity increasing regulatory complexity through cooperative binding to composite DNA motifs73. Alternatively, Sox and POU factors might have been convergently deployed in stem cells across divergent lineages, sometimes as dimers, and sometimes just Sox members.
This is reminiscent to the case of Myc, a bHLH TF also important for mammalian pluripotency (c-Myc). Myc and its heterodimer partners, including Max, originate in unicellular holozoans, where they are suggested to be involved in ribosome biogenesis17,74 (Fig. 6b). Thus, it is likely that Myc regulates an ancestral gene module important for cell proliferation (e.g. ribosome and protein production), that either was important in early stem cells or was regularly co-opted into proliferative stem cells. However, it is noteworthy that Myc has gained new functions across animal evolution, such as apoptosis regulation in vertebrates75. In vertebrates, through the innovation of Oct4 and Nanog, alongside the potentially ‘ancient’ Sox2 a conserved pluripotency regulatory network was established41,76. A precise understanding of how these ancestral regulatory networks evolved, from the hierarchy of these TFs in building stem cells to the batteries of potentially conserved downstream targets, will allow to disentangle cell type and TF factor evolution. Yet, our data clearly shows that two of the main gene families involved in vertebrate pluripotency and key developmental genes across animals were already present before the origins of multicellularity. Eventually, their biochemical capabilities were exapted to build one of the defining cell types of a complex multicellular entity.
Methods
Ethical approvals
The animal study protocols were approved by and conducted in compliance with the Committee on the Use of Live Animals in Teaching and Research (CULTAR Ref. No.: 22-276) at the University of Hong Kong and the Animals (Control of Experiments) Ordinance of Hong Kong.
Generation of chimeric mouse from choanoflagellate Sox iPSCs
OG2MEF cells were derived from day 13.5 embryos of OG2 mice (Jackson Laboratory, no. 004654), following the preparation protocol described previously in ref. 53. Clonal iPSCs derived from OG2MEF (derived from C57BL/6 J strain with black coat color) reprogrammed by full-length Salhel-Sox-I were cultured in 2i/LIF medium on gelatin-coated plates for 7–10 days before harvesting for injection, with medium change every other day. CD-1 (ICR strain, white coat color) female mice were euthanized for isolating morula or blastocysts using a 27 G blunt end needle. These blastocysts were then collected and incubated in KSOM medium (Sigma-Aldrich, #MR-101-D) until the injection of iPSCs was performed. On the day of injection, 1 million cells were prepared to be selected for microinjection into blastocysts. The injection process took place on a 6 cm plate. Using a laser objective (XYRCOS, Hamilton Throne), a slit was created on the zona pellucida, following which 8 to 10 selected iPSCs were transferred into each blastocyst using a microneedle. Following the microinjection procedure, approximately 25 blastocysts containing the integrated clonal iPSCs were implanted into pseudopregnant CD-1 mice, where they would be nurtured and allowed to develop in a conducive environment. This fostering and breeding stage is critical for the establishment of chimeric mice derived from the clonal iPSCs, ensuring the generation of genetically homogenous and robust model organisms for further experimental investigations.
Sequence search, phylogenetics and ancestral sequence reconstruction
We used the human Sox2 protein as query for a BLASTP search against the predicted proteomes of unicellular holozoans, including 22 choanoflagellates (20 transcriptomes and 2 genomes), 4 filastereans, 7 ichthyosporeans and Corallochytrium limacisporum, as well as 130 non-holozoan eukaryotes. Top blast hits had e-values below 1e-19, and bit scores above 90, which were stronger hits than the best putative hits previously identified in Monosiga brevicollis (e-value 1e-11, bitscore 70.9)77. To establish the phylogenetic framework of HMG evolution and minimize distant outgroups (e.g. non-sequence specific HMG boxes), we performed a HMMER3 search with the HMG-box domain (PF00505, e-value < 0.0001) on a select group of species (Homo sapiens, Drosophila melanogaster, Nematostella vectensis, Amphimedon queenslandica, Mnemiopsis leydi, Trichoplax adhaerens, Salpingoeca rosetta, Capsaspora owczarzaki) using the --cut_ga threshold (HMMER3 and PFAM database). We used MAFFT LINS-I for multi-sequence alignment78, trimAl with -gappyout parameter for alignment trimming79, and then used IQTREE to build a maximum likelihood phylogeny allowing for model fitting80. We selected the TCF/LEF, Maelstrom, Capicua (CIC), BobbySox (BBX) and HBP1 clades as closer outgroups of Sox, discarding more distantly related HMG-box families.
Then, using BLASTP we searched human Sox2 against the proteomes of all the unicellular holozoans, and selected all the hits with an e-value below e-10. We also included Sox hits from additional sponge genomes (Oopsacas minuta, Tethya wilhelma, Oscarella carmela, Sycon ciliatum) and an extra placozoan (Hoilungia hongkongensis) to maximize the resolution of early metazoan branching events. The substitution model selected by ModelFinder81, implemented in IQTREE, was LG + G4, with amino acid frequencies taken from the model, according to the corrected Akaike information criterion. We constructed a phylogenetic tree as described above, performing 1000 replicates to obtain SH-like approximate likelihood ratio test (SH-aLRT)82 and ultrafast bootstrap83 nodal supports. The perturbation strength (-pers option) was set to 0.2 and the number of unsuccessful iterations to stop (-nstop option) was set to 500. The tree search was repeated 10 times with different seeds, and the tree with the highest likelihood is shown in Supplementary Fig. 1. Transfer bootstrap values were computed with the Booster84 software using 500 bootstrap trees computed with RAxML-NG85. For constrained tree search, we generated a guide tree with the Sox/Sox-like genes following the holozoan species topology: (Filasteria, (Choanoflagellata, Metazoa)). Polytomies were applied to all internal branches and to those of the outgroup sequences. All branch lengths were set to 1.0. The constrained tree search was repeated 10 times using same guide tree for all tree searches and the same parameters as for the ML tree search. The ML and the five constrained trees with the highest log likelihood values were subjected to an AU-test59 (implemented into IQ-TREE) using the original multiple-sequence alignment, the substitution model LG + G4 and 10,000 replicates. Trees with p-AU values > 0.05 were included in the confidence set of plausible trees that cannot be rejected by the data, which was the case for all five tested constrained trees.
POU was searched using a similar strategy, human POU5F1 sequence was searched against the BLASTP database including unicellular holozoans and other eukaryotes, only identifying a reliable hit in Mylnosiga fluctuans (e-value 1.14e-07). Subsequent reassembly of Salpingoeca helianthica transcriptome (see below) identified another POU5F1 hit in that species. Protein domain composition containing a Homeobox (PF00046) and POU (PF00157) domains was validated using Pfam database86. To place the choanoflagellate POU within the homeobox phylogeny, we used HMMER3 (e-value < 0.0001) search to extract all homeobox domain containing proteins present in Homo sapiens, Drosophila melanogaster, Nematostella vectensis, Amphimedon queenslandica, Trichoplax adhaerens and Mnemiopsis leidyi, as well as all holozoan sequences described above. The resulting 756 sequences were aligned using MAFFT, trimmed uing trimAl, and IQTREE to build the phylogeny. Another tree was built using the same procedure, but only focusing on POU members, and using the trimAl -automated1 parameter, including new sequences from sponges, placozoans and ctenophores. This tree spanned both the homeobox and POU specific domains. Additionally, the same alignment including homeobox outgroups was tested to evaluate the topology of the focused POU phylogeny, and using ONECUT, LHX and SIX homeodomains as outgroups, following previous reports suggesting their relative proximity to POU and the presence of a structurally analogous domain N-terminal of the homeodomain.
For ancestral sequence reconstruction of Sox HMG domains, ancestral gaps were assigned using PastML87 and ancestral sequences were inferred via IQTREE using the LG + G4 substitution model and amino acid frequencies taken from the model and contain the states with the highest posterior probabilities per site.
Sequence re-annotation
Before cloning the HMG-box and POU domains, we validated the annotations using two strategies. For Pigoraptor chileana sequence, we extracted the genomic Sox locus, and used Augustus with various species models to ab initio predict the gene to validate the published genome annotation88. For choanoflagellate sequences, we downloaded the raw reads in NCBI for Salpingoeca helianthica (SRR6344974) and Mylnosiga fluctuans (SRR6344975)44, did adapter trimming using fastp89, and assembled the transcriptome using Trinity v2.8.590. TBLASTN was used to search for the transcripts encoding the POU and Sox identified above, and Open Reading Frame curation was performed using ORFfinder in NCBI.
Sequence feature analysis and visualization
The fasta files of the DNA binding domains of each protein were obtained from https://www.uniprot.org/ and combined with sequences of the unicellular Sox or POU. The M-Coffee option of the T-Coffee Multiple Sequence Alignment server (https://tcoffee.crg.eu/)91 was used to align all sequences. Then, Jalview (https://www.jalview.org/)92 was used to color using clustal colors and annotate conservation, sequence logos. The structure of Sox17 (PDB:3F27) was used to annotate the Sox alignment while the POU was annotated manually following60.
Recombinant DNA
For protein purification, pET28a-mOct4 POU, pETG20a-Sox2 and pETG20a-Sox17 from93 were used. The Salhel Sox-I HMG, Pchi Sox HMG, Salro HMG, Monbr HMG and Cic HMG domains were cloned into the plasmid pETG20a with a N-terminal His6-thioredoxin tag and tobacco etch virus (TEV) cleavage site. The Salhel POU was cloned into pET28a vector with an N-terminal His6 tag and thrombin cleavage site. For reprogramming tests, pHAGE2-TetO-Oct4 (Addgene, #136611), pHAGE2-TetO-Sox2 (Addgene, #136612), pHAGE2-TetO-Klf4 (Addgene, #136613), pHAGE2-TetO-cMyc (Addgene, #136614), pHAGE2-TetO-mSox17 (Addgene, #206367) and pHAGE2-TetO-mCherry (Addgene, #136615) were used. All sequences of other factors were synthesized by Guangzhou IGE Biotechnology and cloned into the pHAGE2-TetO vector. For EMSAs with full-length proteins, the sequences were cloned into the pLVTHM-3xflag vector. The detailed sequences were all included in the Supplementary Data 2. All plasmid reagents will be made via Addgene or by request upon publication.
Protein expression
Proteins encoded on pETG20a-Salhel-HMG,-Pchi-HMG -Salro-HMG, -Monbr-HMG and -Cic-HMG and pET28a-Salhel POU were expressed and purified as previously described for Oct4, Sox2 and Sox1793,94. Constructs were transformed into Rosetta 2(DE3) chemically competent cells (Sigma, #71397) and grown overnight (O/N) at 37 °C in 20 mL Fisher BioReagents™ Miller’s LB Broth (Fisher Scientific, #BP97235), 30 μg/mL chloramphenicol and 100 μg/ml ampicillin (Amp - proteins in pETG20a) or kanamycin (Kan- proteins in pET28a). The next day, a 10 mL subculture was grown in 1 L of LB supplemented with 0.1% glucose with 100 μg/ml of Amp or Kan for pETG20a or pET28a proteins, respectively. When the OD600 reached 0.6–0.8 (2.5–4 h), then protein expression was induced with 0.25–0.5 mM isopropyl-b-thiogalactoside (IPTG) at 18 °C for 18–20 h.
Sox protein purification
pETG20a-Salhel-HMG, -Pchi-HMG -Salro-HMG, -Monbr-HMG and -Cic-HMG were purified in a similar manner. Cells were harvested, resuspended in cold His buffer A (20 mM Tris–HCl pH 8.0, 500 mM NaCl, 30 mM imidazole) and disrupted by ultrasonication on ice (4 s on/8 s off) for 5–8 min on. The lysate was cleared by centrifugation and passed through a 0.22 μm filter. The following steps were all done at 4 °C. The His6-Thx fusion proteins were captured from the supernatant using a HisTrap HP 5 mL (Cytiva, #17524801) pre-equilibrated with buffer A and eluted using His buffer B (20 mM Tris–HCl pH 8.0, 500 mM NaCl, 300 mM imidazole). The buffer was changed to SP buffer A (20 mM Tris–HCl pH 8.0, 100 mM NaCl) using HiPrep 26/10 Desalting column (Cytiva, #17508701). The fusion tag and Sox-HMG were separated by TEV digestion using a substrate: enzyme ratio of 15:1 (w:w) at 4 °C O/N. The Sox-HMG was purified by ion-exchange chromatography using a 1 mL HiTrap SP FF (Cytiva, #17505401) pre-equilibrated with SP buffer A and eluted with a salt gradient (up to 1 M NaCl). Finally, size-exclusion chromatography was performed using a HiLoad Superdex-75 16/600 column (Cytiva, #28989333) in storage buffer (20 mM Tris–HCl pH 8.0, 250 mM NaCl). Fractions with desired protein were pooled, aliquoted, flash frozen and stored at −80 °C.
Salhel POU protein purification
Cells were harvested, resuspended in lysis buffer (100 mM HEPES pH 7.0; 500 mM NaCl; 10 mM Imidazole, 10% Glycerol + [0.5 mM TCEP, 0.4 mM PMSF, 50 U/mL Benzonase® Nuclease added fresh from stock]) and incubated for 30 min on ice. The sample was disrupted by ultrasonication on ice (4 s on/8 s off) for 5–8 min on. The lysate was cleared by centrifugation and the supernatant was discarded. The cell pellet was resuspended in denaturing His Buffer A [20 mM HEPES; 500 mM NaCl; 10 mM Imidazole; 10% Glycerol; 6 M Urea; pH 7.0] and incubated at RT with spinning O/N. All steps with 6 M Urea were done at RT. The mixture was centrifuged, and the supernatant was collected. The His-tagged proteins in the supernatant were captured using HisTrap HP 5 mL (Cytiva, #17524801) pre-equilibrated with denaturing His Buffer A and eluted using denaturing His buffer B [20 mM HEPES; 500 mM NaCl; 300 mM Imidazole; 10% Glycerol; 6 M Urea; pH7.0]. The protein was then refolded via stepwise dialysis. The elute was concentrated to 10 mL and dialyzed with Slide-A-Lyzer Dialysis Cassette (7 K MWCO, Thermo Scientific, #66710) in 1 L of storage buffer [10 mM HEPES; 100 mM NaCl,10% glycerol, 0.5 mM TCEP; pH = 7.0] with 4 M Urea at RT for 2 h. The sample was then dialyzed at 4 °C to storage buffer with 2 M Urea and twice using storage buffer with without Urea (2 h first and then O/N). The final protein was concentrated using the centrifugal units, flash-frozen, and stored at −80 °C.
Electrophoretic mobility shift assay
For purified DNA binding domains
DNA probes (Supplementary Data 1) with 5′-Cy5 or 5’- FAM dyes at the forward strand and unlabeled reverse strand were mixed, heating to 95 °C followed by gradual cooling in a thermocycler in annealing buffer (20 mM Tris/HCl, 50 mM MgCl2, 50 mM KCl, pH 8.0) to make stocks of double-stranded DNA probes. Protein samples and fluorescently labeled DNA were mixed in EMSA buffer containing (10 mM Tris/HCl pH 8.0, 0.1 mg/mL BSA, 50 µM ZnCl2, 100 mM KCl, 10% glycerol, 0.10% Igepal CA630, 2 mM b-mercaptoethanol) and incubated for 1–2 h on ice in the dark. 10 µL of binding reactions were electrophoresed using the Mini-PROTEAN Tetra cell (BioRad) for 30–40 min at 200 V in the cold room (4 °C) using 12% native PAGE mini-gels pre-run at 200 V for 30 min and 1 x TG buffer (25 mM Tris, 192 mM glycine, pH 8.0). Images were captured using an Amersham Typhoon 5 Biomolecular Imager and quantified using ImageQuantTL 7.0. Apparent dissociation constants (Kd) were calculated as described in ref. 31. Cooperativity calculations were performed using established procedures49,50,95 (Supplementary Fig. 6a, b). Binding affinity was plotted as Gibbs Free Energy and calculated through \(\Delta G^\circ={RT}{\mathrm{ln}}\left({apparent\; Kd}\right)\) were R = 0.008314 kJ and T = 277.15 K. Statistics to compare the cooperativity of Sox proteins tested were calculated with R. First, Bartlett Test of Homogeneity of Variances to evaluate whether to use parametric or non-parametric test (bartlett.test). Since there were significant differences in variances found between samples., non-parametric tests were used. Kruskal-Wallis test (kruskal.test) and Games-Howell test (games_howell_test) to calculate adjusted p-values. The functions are from stats and rstatix R packages.
For full length proteins from mammalian cell extracts
Cell extract EMSAs were performed as described in refs. 31,53. In brief, HEK293T cells were used to overexpress the full length proteins of interest from a pLVTHM-3xflag plasmid. Cells were dissociated with 0.05% trypsin-EDTA and washed twice with DPBS after 72 h of overexpression. Cell pellets were lysed in lysis buffer (20 mM Hepes-KOH pH 7.8, 150 mM NaCl, 0.2 mM EDTA pH 8.0, 25% glycerol, freshly added 1 mM DTT, cOmplete™ protease inhibitor cocktail (Roche, #11836145001) using 4x freeze-thaw cycles with liquid nitrogen). The lysate was centrifuged at 14,000 x g at 4 °C for 10 min, and the supernatant which contains the protein of interest was kept. 10 μg of total protein was subjected to SDS-PAGE and Western blot analysis using anti-Flag antibody. Protein levels were adjusted based on the quantification of bands in the blot using cell lysates without exogenous protein. Reactions with the DNA probe and protein were incubated on ice in binding buffer (25 mM Hepes-KOH pH 8.0, 50 mM NaCl, 0.5 mM EDTA, 0.07% Triton X-100, 4 mg/ml BSA, 7 mM DTT,10% glycerol). 10 µL of binding reactions were electrophoresed using the PROTEAN® II xi cell (BioRad) for 2.5 h at 300 V in the cold room (4 °C) using 6% 18.5 × 20 cm native PAGE gel pre-run at 300 V for 1.5 h in 1 x TG buffer. Images are acquired with a GE Typhoon 5 Biomolecular Imager.
Specificity by sequencing (Spec-seq)
The experiment was performed essentially as described in refs. 45,93. DNA libraries (44 bp) were designed by flanking the degenerate sequences of the Octamer motif (ATGCNNNN, ATNNNNAT, NNNNTAAT) or the Sox motif (CATNNNN and NNNNGTT) with 5’ flanking sequence of GAGTCGTCTCGTCAGCAC and 3’ flanking sequence of CCGTAGAGCACTCAGGTC for downstream processing. The resulting libraries were then made into double-stranded DNA (dsDNA) using DreamTaq Green PCR Master Mix (Thermo Scientific: K1081) with the reverse complement primer (GACCTGAGTGCTCTACGG). To get rid of single-stranded DNA (ssDNA), 1 µL of Exonuclease I (New England Biolabs:M0293S) was added to the reaction mix for 30 min. The dsDNA products were then purified using homemade Qiagen PNI binding buffer, PCR purification columns (Tiangen), and eluted in ultrapure water. The respective three Octamer and the two Sox libraries were combined in equimolar amounts. Binding reactions were prepared using different concentrations of protein and 250 nM dsDNA library, in 1x NEB Cutsmart buffer supplemented with 10% glycerol. The reactions were incubated for 1 h at 4 °C, and then EMSA was performed. After the EMSA, the gels were stained with 3x GelRed® stain (Biotium: 41003) for 15 min and visualized. Each band was excised and the DNA in the gel was extracted in 150 uL PAGE diffusion buffer [500 mM Ammonium acetate; 10 mM magnesium acetate; 1 mM EDTA; 0.1 % sodium dodecyl sulfate (SDS), pH 8.0] and purified similarly to the dsDNA libraries. 6 cycles of PCR were performed using primers containing a unique molecular identifier (UMI) to account for PCR bias. A second round of PCR (28 cycles) was performed using primers compatible with Illumina adapter and containing different indexing barcodes (Primers are in Supplementary Data 1). The resulting PCR products were combined and gel purified twice and then sent to sequencing using a Illumina NovaSeq 6000 PE150 (Novogene).
Spec-seq data analysis
R packages (QuasR, Biostrings, tidyr, data.table, stringi) were used to process the paired-end sequencing data. The reads were trimmed for adapter sequences using the preprocessReads function. A library file (pseudogenome) with all the theoretically possible sequences in each experiment was created. This file was used as an alignment template for the sequences from the Spec-seq libraries with the qAlign function (Rhisat2 as aligner) resulting in a count matrix where rows are the sequence elements and columns are samples. The relative binding energy of each sequence was calculated as described in refs. 45,93 and plotted as a scatter plot using ggpubr: ggscatter in R, where \({\#Si}\) is the number of reads per sequence, using the formula:
To plot energy logos, a subset of the sequence space was used corresponding to the consensus sequences (CATTGTT or ATGCTAAT) with all possible single base mutations (N*3 + 1 where N is the length of the sequence and ‘3’ are the three possible mutations at each of the N positions). The binding energies \(({\mathrm{ln}}\frac{{\#Si\_unbound}}{{\#Si\_bound}})\) of this subset of sequences was used to generate an energy matrix using the motif_mlr.pl script available from the Gary Stormo Lab and the logo plotted using plotEnergyLogo from TFcookbook96 (https://github.com/zeropin/TFCookbook).
High throughput systematic evolution of ligands by exponential enrichment (HT-SELEX)
The HT-SELEX experiment followed the protocol outlined in ref. 61. Selection ligands were designed with an 8 bp barcode flanking a 40 bp randomized region. In summary, 50–100 ng of barcoded DNA fragments were introduced into wells containing the target proteins along with 25 μL of binding buffer supplemented with 5 μg/ml poly-dIdC-oligonucleotide (Sigma P4929-25UN) as a competitor. The plate was gently rotated at room temperature for 30 min. Following incubation, 150 μL of binding buffer, along with 10 μL of Ni Sepharose® beads previously equilibrated with poly-dIdC, were introduced into the reaction mixture. This mixture was then incubated on a rotor at room temperature for 60 min. Unbound oligomers were subsequently removed from the bound beads using a Biotek 405TS washer. Post washing, the bound DNA was eluted in 50 μL of ddH2O. For PCR amplification, 10 μL of bead suspension was utilized. The resulting PCR products served as input oligomers for the subsequent cycle or were purified for sequencing with the NovaSeq 6000 S4 PE150. This experiment involved a total of four cycles.
Data obtained were sorted based on barcodes for each sample. After discarding low-quality reads, the remaining sequences underwent trimming to eliminate adapter sequences. The resultant 40-nt region was subjected to further analysis. PWM models were generated using initial seeds identified through Autoseed, which were subsequently refined through expert analysis, in accordance with the approach outlined in ref. 61. All motif seqlogos were generated using the R package ggseqlogo97.
Modeling of Oct4-unicellular Sox complexes
Oct4 complexed with Salhel and Pchi HMG were homology-modeled bound to the canonical and compressed SoxOct DNA motifs using MODELLER 10.498,99. The structural template for the Canonical motif is a model of ternary complex Oct4 and Sox2 on the Nanog locus (5’CAGGGTCCACCATGGACATTGTAATGCAAAAGAAGCTGTAAGGTGACCC3’)24. For the compressed motif, a model of the Oct4-Sox17 complex on an idealized sequence (5’ CGGCATTGTATGCAAATCGGCGGC3’) was used24. Motifs are in bold, and the additional base of the canonical motif is in red.
We aligned individual sequences of Salhel HMG and Pchi HMG with Sox2 or Sox17, respectively. Similarly, Salhel POU was modeled using the Sox2-Oct4-Nanog locus ternary complex as a template. Salhel POU was aligned with Oct4. For all models, the automodel function was used to generate 500 models with the DNA set as a rigid body and the model refinement level set at fast. The model with the lowest discrete optimized protein energy (DOPE) score100 was selected. ChimeraX was used to analyze and visualize clashes/contacts of the complexes modeled101,102.
Cell culture
Mouse embryonic fibroblasts (MEFs) were obtained from E13.5 embryos of OG2 mice carrying transgenic Oct4-GFP (Jackson Laboratory, no. 004654) and Sox2-GFP mice (Mutant Mouse Resource & Research Centers, no. 037525-UNC) with a GFP reported driven by endogenous Sox2, maintained at the Centre for Comparative Medicine Research (CCMR) at The University of Hong Kong (CULATR no. 4855-18). MEFs were cultured in MEF medium [DMEM (Gibco, #12100046) supplemented with 10% fetal bovine serum (FBS, Gibco, #10270106), 1x Glutamax (Gibco, #35050061), 1x nonessential amino acids (NEAA, Gibco, #11140050) and 1x penicillin/streptomycin (Gibco, #10378016)]. HEK293T cells for lentivirus packaging were cultured in DMEM supplemented with 10% FBS. Mouse ESC medium (mES medium) is composed of: DMEM with 15% FBS, 1x GlutaMax, 1x NEAA, 1 mM sodium pyruvate (Gibco, #11360070), 0.005 mM β-mercaptoethanol (Gibco, #31350010), 50 μg/mL Vitamin C (Sigma-Aldrich, #49752), 0.5x penicillin/streptomycin and 10 ng/mL leukemia inhibitory factor (LIF, produced in house). 2i/LIF medium is mES medium supplemented with 3 μM CHIR99021 (Selleck, #S2924-25mg), 1 μM PD0325901 (Selleck, #S1036-25 mg)103. Pluripotent stem cells were maintained on either feeder layers (ICR MEFs mitotically inactivated with mitomycin-C) or on 0.2% gelatin-coated plates (Sigma-Aldrich, #G1393) and cultured in mES medium or 2i/LIF medium. Medium was replaced with fresh medium every other day, and passaged at 1:10 split ratio when reaching around 70% confluency. All cells were cultured in incubators at 37 °C with 0.5% CO2 and normoxic conditions.
Lentivirus production and reprogramming
HEK 293 T cells were seeded at 8 millions of cells per 10 cm plate. On the next day, 10 μg lentiviral vector and 40 μg linear polyethyleneimine (Polysciences, #23966) dissolved in 1 mL DMEM were added. The medium was replaced after 10 ~ 15 h and virus-containing supernatants were collected at 48 h, 72 h, and 96 h post-transfection and filtered through a 0.45 µm filter (Millipore). The virus medium was supplemented with 8 μg/mL polybrene (Sigma-Aldrich, #40804ES76) before transduction. MEFs were seeded at a density of 7 × 103 cells per well of a 24-well plate one-day before transduction. The virus-containing medium was replaced after 24 h with mES medium. This day was defined as reprogramming day 0 and the medium was replaced daily on subsequent days. Whole well scans were taken using the GE Amersham Typhoon™ 5 Biomolecular Imager. To establish clonal iPSC lines, iPSC colonies were picked at day 14 using a syringe, dissociated into single-cell suspension by pipetting up and down in 30 μL of 0.05% Trypsin-EDTA (Thermo Fisher Scientific, #25300062) and incubating at 37 °C for 5 min and seeded into 48-well plate pre-coated with ICR feeders. The cells were cultured for 5 ~ 7 days until sizeable iPSC colonies developed and two more rounds of picking were conducted to obtain pure clonal lines.
Genotyping of iPSC lines
To genotype the iPSCs reprogrammed with unicellular Sox, 500,000 cells of the clonal iPSC lines were harvested for genomic DNA isolation using Quick-DNA Microprep Kit (Zymo, #D3021). The isolated genomic DNA was used to examine the integrated transgene by PCR with different specific primers (Supplementary Data 1) and verified by Sanger sequencing.
Quantitative RT-PCR analysis
Total RNA was extracted using TRIzol (Thermo Fisher Scientific, #15596026) and 2 μg was used to synthesize cDNA with ReverTra Ace® qPCR RT Master Mix (Toyobo, FSQ-201S). Quantitative PCR was performed using iTaq universal SYBR Green Supermix (Bio-Rad, #1725124) with primers listed in Supplementary Data 1. β-actin was used for normalization to calculate the relative gene expression. The R package ggplot2 was used to plot the results (https://ggplot2.tidyverse.org).
Immunocytochemistry
Cells were washed three times with PBS and fixed in 4% paraformaldehyde in PBS at room temperature for 20 min, followed by permeabilization with 0.1% Triton X-100 and blocking with 5% BSA in PBS at room temperature for 1 h. Fixed cells were washed three times with PBS, and incubated with primary antibodies Nanog (Novus Biologicals, #NB100-58842, 1:500 dilution), and Sox2 (Santa Cruz Biotechnology, #sc-365823, 1:400 dilution) at 4 °C overnight. The cells were then washed three times for 5 min with PBST (PBS with 0.1% Tween-20) and incubated with fluorescent-dye conjugated secondary antibodies (Invitrogen, Alexa Fluor 488 dye: #A21203/A21207, 1:1000 dilution) at room temperature for 1 ~ 2 h. The cells were then washed with PBST three times for 5 min. For nuclei counterstaining, NucBlue™ Fixed Cell ReadyProbes™ Reagent (DAPI) (#R37606) was used, following the instruction from the kit. Images were captured with the inverted fluorescent microscope (Olympus CKX53).
Spontaneous differentiation of iPSCs into endoderm, mesoderm, and ectoderm
Clonal iPSCs were dissociated using 0.05% Trypsin-EDTA and seeded to 96-well low attachment plates (1·103 cells/well) in mES medium without LIF for 7 days to generate embryoid bodies (EBs). EBs were then seeded on gelatin-coated 12-well plates (20 EBs per well) and cultured in the differentiation medium (DMEM/F12 + 20%FBS + 1% Glutamax) for another 10 days. The differentiation medium was exchanged every other day. After 10 day differentiation in gelatin-coated plates, spontaneously differentiated EBs were further analysed by immunocytochemistry. To evaluate tri-lineage differentiation potential, cells were stained with primary antibody against three germ layer markers respectively (FoxA2, #8186 T Cell Signaling, 1:500 dilution; TUJ1, #PA5-85639 Invitrogen, 1:500 dilution; α-SMA, #A2547 Sigma-Aldrich, 1:500 dilution) at 4 °C overnight, followed by incubation with corresponding fluorescent-dye conjugated secondary antibodies (Invitrogen, Alexa Fluor 594 dye: #A11055/A1100/A21202, 1:1000 dilution) at room temperature for 1 ~ 2 h. The steps of fixation, permeabilization, blocking, washing, DAPI staining, and imaging are the same as described above for immunocytochemistry.
Statistics & reproducibility
No statistical method was used to predetermine sample size. To ensure accurate cooperative calculations from heterodimer EMSA, lanes with a band with a fractional contribution below 0.03 were excluded. This ensures that only lanes representing all four microstates at equilibrium are included. The experiments were not randomized. For the experimental setup of reprogramming tests, technical replicates involve using the same batch of reagents and cells within an experiment, while biological replicates involve using distinct batches of MEF cells sourced from different mouse embryos. No data was excluded for quantification. All the key reagents and equipment used in the study were listed in Supplementary Data 3.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw images of gels, reassembled sequences from publicly available datasets, alignments and phylogenetic trees can be found in the public repository figshare (https://doi.org/10.6084/m9.figshare.26538691). The raw data for quantification generated in this study are provided in the Source Data file. Spec-seq and HT-SELEX sequencing data are available from the gene expression omnibus (GEO) under accession number GSE253888. Source data are provided with this paper.
References
Sogabe, S. et al. Pluripotency and the origin of animal multicellularity. Nature 570, 519–522 (2019).
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell Biol. 9, 625–635 (2007).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet 24, 372–376 (2000).
Dodonova, S. O., Zhu, F., Dienemann, C., Taipale, J. & Cramer, P. Nucleosome-bound SOX2 and SOX11 structures elucidate pioneer factor function. Nature 580, 669–672 (2020).
Michael, A. K. et al. Mechanisms of OCT4-SOX2 motif readout on nucleosomes. Science 368, 1460–1465 (2020).
Nguyen, T. et al. Chromatin sequesters pioneer transcription factor Sox2 from exerting force on DNA. Nat. Commun. 13, 3988 (2022).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Lee, M. T. et al. Nanog, Pou5f1 and SoxB1 activate zygotic gene expression during the maternal-to-zygotic transition. Nature 503, 360–364 (2013).
Leichsenring, M., Maes, J., Mossner, R., Driever, W. & Onichtchouk, D. Pou5f1 transcription factor controls zygotic gene activation in vertebrates. Science 341, 1005–1009 (2013).
Gassler, J. et al. Zygotic genome activation by the totipotency pioneer factor Nr5a2. Science 378, 1305–1315 (2022).
Ji, S. et al. OBOX regulates mouse zygotic genome activation and early development. Nature 620, 1047–1053 (2023).
Klaus, M. et al. Structure and decoy-mediated inhibition of the SOX18/Prox1-DNA interaction. Nucleic Acids Res 44, 3922–3935 (2016).
Remenyi, A. et al. Crystal structure of a POU/HMG/DNA ternary complex suggests differential assembly of Oct4 and Sox2 on two enhancers. Genes Dev. 17, 2048–2059 (2003).
de Mendoza, A. & Sebe-Pedros, A. Origin and evolution of eukaryotic transcription factors. Curr. Opin. Genet Dev. 58-59, 25–32 (2019).
de Mendoza, A. et al. Transcription factor evolution in eukaryotes and the assembly of the regulatory toolkit in multicellular lineages. Proc. Natl Acad. Sci. USA 110, E4858–E4866 (2013).
Larroux, C. et al. Genesis and expansion of metazoan transcription factor gene classes. Mol. Biol. Evol. 25, 980–996 (2008).
Sebe-Pedros, A., de Mendoza, A., Lang, B. F., Degnan, B. M. & Ruiz-Trillo, I. Unexpected repertoire of metazoan transcription factors in the unicellular holozoan Capsaspora owczarzaki. Mol. Biol. Evol. 28, 1241–1254 (2011).
Schnitzler, C. E., Simmons, D. K., Pang, K., Martindale, M. Q. & Baxevanis, A. D. Expression of multiple Sox genes through embryonic development in the ctenophore Mnemiopsis leidyi is spatially restricted to zones of cell proliferation. Evodevo 5, 15 (2014).
Bowles, J., Schepers, G. & Koopman, P. Phylogeny of the SOX family of developmental transcription factors based on sequence and structural indicators. Dev. Biol. 227, 239–255 (2000).
Sebe-Pedros, A. et al. Cnidarian cell type diversity and regulation revealed by whole-organism single-cell RNA-seq. Cell 173, 1520–1534.e1520 (2018).
Chrysostomou E. et al. A cellular and molecular analysis of SoxB-driven neurogenesis in a cnidarian. Elife 11, e78793 (2022).
Varley, A., Horkan, H. R., McMahon, E. T., Krasovec, G. & Frank, U. Pluripotent, germ cell competent adult stem cells underlie cnidarian regenerative ability and clonal growth. Curr. Biol. 33, 1883–1892.e1883 (2023).
Najle, S. R. et al. Stepwise emergence of the neuronal gene expression program in early animal evolution. Cell 186, 4676–4693.e4629 (2023).
Merino, F. et al. Structural basis for the SOX-dependent genomic redistribution of OCT4 in stem cell differentiation. Structure 22, 1274–1286 (2014).
Malik, V. et al. Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2. Nat. Commun. 10, 3477 (2019).
Rodda, D. J. et al. Transcriptional regulation of nanog by OCT4 and SOX2. J. Biol. Chem. 280, 24731–24737 (2005).
Li, L. et al. Multifaceted SOX2-chromatin interaction underpins pluripotency progression in early embryos. Science 382, eadi5516 (2023).
MacCarthy, C. M. et al. Highly cooperative chimeric super-SOX induces naive pluripotency across species. Cell Stem Cell 31, 127–147.e129 (2024).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Aksoy, I. et al. Sox transcription factors require selective interactions with Oct4 and specific transactivation functions to mediate reprogramming. Stem Cells 31, 2632–2646 (2013).
Hu H. et al. Evaluation of the determinants for improved pluripotency induction and maintenance by engineered SOX17. Nucleic Acids Res. 51, 8934–8956 (2023).
Nakagawa, M. et al. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat. Biotechnol. 26, 101–106 (2008).
Jauch, R. et al. Conversion of Sox17 into a pluripotency reprogramming factor by reengineering its association with Oct4 on DNA. Stem Cells 29, 940–951 (2011).
Jerabek, S. et al. Changing POU dimerization preferences converts Oct6 into a pluripotency inducer. EMBO Rep. 18, 319–333 (2017).
Tapia, N. et al. Dissecting the role of distinct OCT4-SOX2 heterodimer configurations in pluripotency. Sci. Rep. 5, 13533 (2015).
Gold, D. A., Gates, R. D. & Jacobs, D. K. The early expansion and evolutionary dynamics of POU class genes. Mol. Biol. Evol. 31, 3136–3147 (2014).
Bakhmet, E. I. & Tomilin, A. N. Key features of the POU transcription factor Oct4 from an evolutionary perspective. Cell Mol. Life Sci. 78, 7339–7353 (2021).
Degnan, B. M., Vervoort, M., Larroux, C. & Richards, G. S. Early evolution of metazoan transcription factors. Curr. Opin. Genet Dev. 19, 591–599 (2009).
Grau-Bove X. et al. Dynamics of genomic innovation in the unicellular ancestry of animals. Elife 6, e26036 (2017).
Paps, J. & Holland, P. W. H. Reconstruction of the ancestral metazoan genome reveals an increase in genomic novelty. Nat. Commun. 9, 1730 (2018).
Sukparangsi, W. et al. Evolutionary origin of vertebrate OCT4/POU5 functions in supporting pluripotency. Nat. Commun. 13, 5537 (2022).
Brunet, T. & King, N. The origin of animal multicellularity and cell differentiation. Dev. Cell 43, 124–140 (2017).
Ocana-Pallares, E. et al. Divergent genomic trajectories predate the origin of animals and fungi. Nature 609, 747–753 (2022).
Richter D. J., Fozouni P., Eisen M. B., King N. Gene family innovation, conservation and loss on the animal stem lineage. Elife 7, e34226 (2018).
Stormo, G. D., Zuo, Z. & Chang, Y. K. Spec-seq: determining protein-DNA-binding specificity by sequencing. Brief. Funct. Genomic. 14, 30–38 (2015).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459.e420 (2017).
Knaupp, A. S. et al. Transient and permanent reconfiguration of chromatin and transcription factor occupancy drive reprogramming. Cell Stem Cell 21, 834–845.e836 (2017).
Zviran, A. et al. Deterministic somatic cell reprogramming involves continuous transcriptional changes governed by myc and epigenetic-driven modules. Cell StemCell 24, 328–341.e329 (2019).
Chang, Y. K. et al. Quantitative profiling of selective Sox/POU pairing on hundreds of sequences in parallel by Coop-seq. Nucleic Acids Res. 45, 832–845 (2017).
Ng, C. K. et al. Deciphering the sox-oct partner code by quantitative cooperativity measurements. Nucleic Acids Res. 40, 4933–4941 (2012).
Lodato, M. A. et al. SOX2 co-occupies distal enhancer elements with distinct POU factors in ESCs and NPCs to specify cell state. PLoS Genet 9, e1003288 (2013).
Mistri, T. K. et al. Selective influence of Sox2 on POU transcription factor binding in embryonic and neural stem cells. EMBO Rep. 16, 1177–1191 (2015).
Weng, M. et al. An engineered Sox17 induces somatic to neural stem cell fate transitions independently from pluripotency reprogramming. Sci. Adv. 9, eadh2501 (2023).
Hochberg, G. K. A. & Thornton, J. W. Reconstructing ancient proteins to understand the causes of structure and function. Annu Rev. Biophys. 46, 247–269 (2017).
Hou, L., Srivastava, Y. & Jauch, R. Molecular basis for the genome engagement by Sox proteins. Semin. Cell Dev. Biol. 63, 2–12 (2017).
Veerapandian, V. et al. Directed evolution of reprogramming factors by cell selection and sequencing. Stem Cell Rep. 11, 593–606 (2018).
Niwa, H. et al. The evolutionally-conserved function of group B1 Sox family members confers the unique role of Sox2 in mouse ES cells. BMC Evol. Biol. 16, 173 (2016).
Aksoy, I. et al. Oct4 switches partnering from Sox2 to Sox17 to reinterpret the enhancer code and specify endoderm. EMBO J. 32, 938–953 (2013).
Shimodaira, H. An approximately unbiased test of phylogenetic tree selection. Syst. Biol. 51, 492–508 (2002).
Malik, V., Zimmer, D. & Jauch, R. Diversity among POU transcription factors in chromatin recognition and cell fate reprogramming. Cell Mol. Life Sci. 75, 1587–1612 (2018).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Burglin, T. R. & Affolter, M. Homeodomain proteins: an update. Chromosoma 125, 497–521 (2016).
Ruiz-Trillo, I., De Mendoza, A. Towards understanding the origin of animal development. Development 147, dev192575 (2020).
van Wolfswinkel, J. C., Wagner, D. E. & Reddien, P. W. Single-cell analysis reveals functionally distinct classes within the planarian stem cell compartment. cell stem cell 15, 326–339 (2014).
Alie, A. et al. The ancestral gene repertoire of animal stem cells. Proc. Natl Acad. Sci. USA 112, E7093–E7100 (2015).
Sebe-Pedros, A. et al. Early metazoan cell type diversity and the evolution of multicellular gene regulation. Nat. Ecol. Evol. 2, 1176–1188 (2018).
Richards, G. S. & Rentzsch, F. Regulation of Nematostella neural progenitors by SoxB, Notch and bHLH genes. Development 142, 3332–3342 (2015).
Steger, J. et al. Single-cell transcriptomics identifies conserved regulators of neuroglandular lineages. Cell Rep. 40, 111370 (2022).
Denner, A. et al. Nanos2 marks precursors of somatic lineages and is required for germline formation in the sea anemone Nematostella vectensis. Sci. Adv. 10, eado0424 (2024).
Ma, Y. et al. Functional interactions between Drosophila bHLH/PAS, Sox, and POU transcription factors regulate CNS midline expression of the slit gene. J. Neurosci. 20, 4596–4605 (2000).
Aleksic, J., Ferrero, E., Fischer, B., Shen, S. P. & Russell, S. The role of Dichaete in transcriptional regulation during Drosophila embryonic development. BMC Genomics 14, 861 (2013).
Millane, R. C. et al. Induced stem cell neoplasia in a cnidarian by ectopic expression of a POU domain transcription factor. Development 138, 2429–2439 (2011).
Reinke, A. W., Baek, J., Ashenberg, O. & Keating, A. E. Networks of bZIP protein-protein interactions diversified over a billion years of evolution. Science 340, 730–734 (2013).
Young, S. L. et al. Premetazoan ancestry of the myc-max network. Mol. Biol. Evol. 28, 2961–2971 (2011).
Cowling, V. H., Chandriani, S., Whitfield, M. L. & Cole, M. D. A conserved Myc protein domain, MBIV, regulates DNA binding, apoptosis, transformation, and G2 arrest. Mol. Cell Biol. 26, 4226–4239 (2006).
Theunissen, T. W. et al. Nanog overcomes reprogramming barriers and induces pluripotency in minimal conditions. Curr. Biol. 21, 65–71 (2011).
King, N. et al. The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 451, 783–788 (2008).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2019).
Capella-Gutierrez, S., Silla-Martinez, J. M. & Gabaldon, T. trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973 (2009).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: Improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Lemoine, F. et al. Renewing Felsenstein’s phylogenetic bootstrap in the era of big data. Nature 556, 452–456 (2018).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Mistry, J. et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 49, D412–D419 (2021).
Ishikawa, S. A., Zhukova, A., Iwasaki, W. & Gascuel, O. A fast likelihood method to reconstruct and visualize ancestral scenarios. Mol. Biol. Evol. 36, 2069–2085 (2019).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Di Tommaso, P. et al. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 39, W13–W17 (2011).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Tan, D. S. et al. Directed evolution of an enhanced POU reprogramming factor for cell fate engineering. Mol. Biol. Evol. 38, 2854–2868 (2021).
Tan, D. S. et al. The homeodomain of Oct4 is a dimeric binder of methylated CpG elements. Nucleic Acids Res. 51, 1120–1138 (2023).
Narasimhan, K. et al. DNA-mediated cooperativity facilitates the co-selection of cryptic enhancer sequences by SOX2 and PAX6 transcription factors. Nucleic Acids Res. 43, 1513–1528 (2015).
Zuo, Z. Encoding, regression, and classification of transcription factors’ specificity and methylation effects. OBM Genet. 5, 1–1 (2021).
Wagih, O. ggseqlogo: a versatile R package for drawing sequence logos. Bioinformatics 33, 3645–3647 (2017).
Fiser, A., Do, R. K. & Sali, A. Modeling of loops in protein structures. Protein Sci. 9, 1753–1773 (2000).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Shen, M. Y. & Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 15, 2507–2524 (2006).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Mulas C. et al. Defined conditions for propagation and manipulation of mouse embryonic stem cells. Development 146, dev173146 (2019).
Acknowledgements
This work used computing resources from Queen Mary University of London’s Apocrita HPC facilities by A.d.M. General Research Fund (RGC/GRF, Grant. No. 17117823, 17101120 and 17106622 to R.J.). Research Grants Council of Hong Kong Collaborative Research Fund (RGC/CRF, Grant No. C7064-22G to R.J.). Health and Medical Research Fund (08192886 to R.J.). Innovation Technology Commission Funding Scheme Health@InnoHK. European Research Council Starting Grant (950230 to A.d.M.). M.G. acknowledges support by the Peter und Traudl Engelhorn Foundation. G.K.A.H. is supported by the Max-Planck Society.
Author information
Authors and Affiliations
Contributions
R.J. and A.d.M. conceived the study with input from S.Y.H., M.G. and G.H. Y.G. (iPSC work) and D.S.T. (biochemistry) performed and analysed most wet lab experiments and prepared figures supervised by R.J. H.H. performed the whole cell extract EMSA. S.W.Y. helped with reprogramming experiments. M.G., A.d.M. and G.H. performed sequence search, phylogenetics, ancestral sequence reconstruction and analysis. X.Z. and Q.Q. performed HT-SELEX supervised by J.Y. K.S.L. generated chimeric mice. D.S.T. generated structural models with guidance from V.C. R.J., A.d.M. and M.G. drafted the initial manuscript and all authors contributed to the final version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Bernard Degnan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gao, Y., Tan, D.S., Girbig, M. et al. The emergence of Sox and POU transcription factors predates the origins of animal stem cells. Nat Commun 15, 9868 (2024). https://doi.org/10.1038/s41467-024-54152-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-54152-x
This article is cited by
-
Genome-wide identification, evolutionary diversification and developmental expression of the Sox gene family in three brachyuran crabs
BMC Genomics (2026)
-
A smartphone analogy to explore the origin of animals
The EMBO Journal (2026)
-
The interplay between driver mutation and oxidative stress in colorectal cancer: from pathogenesis to therapeutics
Journal of Translational Medicine (2025)
-
Chromatin profiling identifies putative dual roles for H3K27me3 in regulating cell type-specific genes and transposable elements in choanoflagellates
Nature Communications (2025)
-
Up-regulated microRNAs in blastocoel fluid of human implanted embryos could control circuits of pluripotency and be related to embryo competence
Journal of Assisted Reproduction and Genetics (2025)