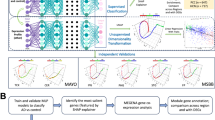
Abstract
Aging increases the risk for Alzheimer’s disease (AD), driving pathological changes like amyloid-β (Aβ) buildup, inflammation, and oxidative stress, especially in the prefrontal cortex (PFC). We present the first subcellular-resolution spatial transcriptome atlas of the human prefrontal cortex (PFC), generated with Stereo-seq from six male AD cases at varying neuropathological stages and six age-matched male controls. Our analyses revealed distinct transcriptional alterations across PFC layers, highlighted disruptions in laminar structure, and exposed AD-related shifts in layer-to-layer and cell-cell interactions. Notably, we identified genes highly upregulated in stressed neurons and nearby glial cells, where AD diminished stress-response interactions that promote Aβ clearance. Further, cell-type-specific co-expression analysis highlighted three neuronal modules linked to neuroprotection, protein dephosphorylation, and Aβ regulation, with all modules downregulated as AD progresses. We identified ZNF460 as a transcription factor regulating these modules, offering a potential therapeutic target. In summary, this spatial transcriptome atlas provides valuable insight into AD’s molecular mechanisms.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is a progressive neurodegenerative disorder primarily associated with memory deficits and cognitive decline that can eventually affect behavior, speech, visuospatial orientation and the motor system1,2,3. AD is the most common cause of dementia among the elderly, impacting an estimated 32 million individuals globally as of 20234 and its prevalence is increasing in nations with aging populations, thereby imposing significant burdens on their healthcare systems. AD brain pathology is characterized by the accumulation of extracellular amyloid-β (Aβ) plaques5 and intracellular hyperphosphorylated tau aggregates as neurofibrillary tangles (NFTs) in the gray matter6. These pathological features can trigger cytotoxic events, neuroinflammation, mitochondrial dysfunction, and contribute to the neuronal stress and degeneration, ultimately resulting in brain atrophy5,7. Interestingly, similar pathological hallmarks, can also occur in aged individuals who are not diagnosed with AD8,9. The extent of these features can vary greatly between individuals, and they may dominate or be restricted in specific brain regions8,9. However, it remains unclear what underlie these pathological features and whether they are precursors of neurodegeneration and AD, or simply the products of normal brain aging. Further, while previous studies have proposed that aging is the most profound risk factor for AD10,11, the molecular mechanisms underlying the aging-related susceptibility to AD is far from clear12. Therefore, transcriptome studies of the human brain are crucial for uncovering the molecular mechanisms that differentiate AD from normal aging (NA).
Recent advances in single-cell or single-nucleus RNA sequencing (sc/snRNA-seq), along with spatial transcriptomics, have revealed remarkable molecular diversity in the cellular landscapes of brains under NA and AD. For instance, the first snRNA-seq study on human AD conducted by Mathys et al.13 demonstrated that in human PFC, the strongest changes associated with AD manifest at the early stage of AD pathological progression and are highly specific to certain cell types. In contrast, genes that are upregulated in the later stages of AD are generally consistent across different brain cell types. Moreover, Chen et al.14 utilized the 10X Visium platform to study the human middle temporal gyrus in three AD cases and three control subjects. By aligning 10X Visium spots ( ~ 55 µm diameter) with adjacent sections stained for Aβ plaques and NFTs, the study observed upregulation of specific AD-related genes and changes in gene co-expression patterns across the bulk of cells near AD pathology, in contrast to those in more distant areas. These studies contributed to a better understanding of how gene regulatory networks drive specific transcriptional changes across various brain cell types, spatial context, and health/physiological conditions, shedding light on possible pathogenic cell subtypes underlying AD and cell-cell interactions among neuronal and non-neuronal cells. However, until now, AD-focused sc/snRNA-seq and spatial transcriptomics studies13,14,15,16 lacked the capability to capture spatial information at single-cell resolution. This limitation is particularly significant because even in brains of patients with moderate AD, neurons and non-neuronal cells with normal function may coexist with those affected by AD pathology, and the affected cells may present AD related changes at the molecular level in a cell-type AND spatial specific manner. Consequently, case-control studies that rely on sc/snRNA-seq OR spatial transcriptomics that do not incorporate single-cell resolution in spatial context may encounter biases when comparing AD-affected with NA brains. These biases can impede the accurate identification of transcriptomic markers in neurons and glial cells that are influenced by the pathological features of AD in contrast to those resulting from NA brains. Thus, comprehensive understanding of the transcriptional profiles and corresponding spatial information at single-cell resolution is essential to unveil the precise and detailed molecular mechanism of AD.
In this study, for the first time, we utilized the SpaTial Enhanced Resolution Omics-sequencing (Stereo-seq)17, a state-of-art spatial transcriptome platform offering large-field-of-view and subcellular resolution to develop a comprehensive, transcriptome-wide, and high definition atlas of the human prefrontal cortex (PFC), a vulnerable brain region in AD18, across both NA and AD-affected individuals. We successfully identify the six neocortex layers and the white matter (WM) in the PFC using supervised methods, revealing disruptions of laminar structure and detailed layer specific pathological alterations in transcriptional atlas as AD progresses. Moreover, we identified some unique cellular reactions to both physiological stress from NA and pathological stress from AD, at single-nucleus resolution. Following the construction of a cell-type specific gene co-expression network, we identified three key modules within excitatory (Ex) and inhibitory (Inh) neurons that are linked to neuronal protection and the clearance of AD-related pathological features. These modules demonstrated an inverse relationship with AD progression, pointing to a decreased ability to clear toxic substances in AD-affected samples. Furthermore, we identified ZNF460 as a potential transcription factor governing all three modules, suggesting its potential as a therapeutic target for AD.
Results
The divergence of laminar structure and transcriptional profiles between AD and NA samples
Human postmortem optimal cutting temperature (OCT) embedded samples from the prefrontal cortex (BA10 area) of six male AD patients (Braak stages IV-VI, consortium to establish a registry for AD (CERAD) scores C2-3, male, aged 70-91) and six male control samples (Braak stages II-III, CERAD scores C0-1, aged 74-95) were selected for Stereo-seq experiments (see detailed case demographics, clinical and overall neuropathological information in Supplemental Data 1). For each sample, two cryosections ( ~ 10 µm distance between each section) were captured and labeled as No.1 to No.2 from the top to the bottom sections, respectively. After the quality control and area selection on the No.1 hematoxylin and eosin (H&E) sections (Fig. S1), 10 × 10 mm area from the No.2 sections were chosen for Stereo-seq profiling. Due to the restricted sampling area on the stereo-seq chip (10 × 10 mm), the neuropathological findings related to AD across the entire brain may not accurately represent the progression of AD within the specific areas in our spatial transcriptome study. Consequently, to further identify the divergence of transcriptional profiles among different stages of AD, the third sections from each sample were captured and labeled as No.3 ( ~ 50 µm distance between the No.2 and No.3 sections). Based on immunohistochemical (IHC) stains of Aβ plaques19 on the No.3 sections (Fig. 1A; Methods), we classified the AD samples into moderate and severe AD groups. Given the minimal presence of the Aβ plaques in the control samples and the advanced age of these subjects (aged 74-95, older than AD subjects on average), we considered the individuals in control group as NA.
A The analysis pipeline of the study, with select figures created using BioRender (Created in BioRender. Gong, Y. (2024) https://BioRender.com/m56f734; Created in BioRender. Gong, Y. (2024) https://BioRender.com/l73a167; Created in BioRender. Gong, Y. (2024) https://BioRender.com/s92q855; Created in BioRender. Gong, Y. (2024) https://BioRender.com/c29s609). B The H&E staining, immunohistochemistry (IHC) staining for Aβ plaques (Aβ42; red) and nuclei (DAPI; blue), and layer clustering for NA (B01809A3), moderate (B01809C2), and severe (A02092E1) AD groups. The IHC experiment was conducted once. Scale bar for the upper panel of the IHC, 1,500 \(\mu m\), while the scale bar for the lower panel of the IHC, 100 \(\mu m\). C UMAP visualization of seven clusters across 12 samples, annotated by cortical layer I to VI (L1 to L6) and the white matter (WM). D Heatmap of the marker genes in each cortical layer and the WM. The x-axis represents six cortical layers and the WM. Colors represent the mean expression of the gene in each cortical layer and the WM. Violin and scatter plots illustrating the expression levels of previously identified (E) and unreported (F) layer marker genes in B01809A3. In the violin plot, the x-axis represents the layer specific markers, and the y-axis represents the cortical layers and the WM. The color represents the median gene expression in the cluster. For the scatter plots, six plots represent the distribution of the gene expression levels of the layer markers, respectively. The color represents the gene expression levels.
Conventionally, the human cerebral cortex is organized into six cellular layers (or laminae) and each layer exhibits unique cellular composition, intra- and interlaminar connectivity, and unique patterns of gene expression20. A previous spatial transcriptome study by Maynard et al.21 has manually delineated the laminar architecture by aligning gene expression sections with the adjacent H&E sections. They have identified the specific gene markers for each layer in healthy human dorsolateral prefrontal cortex through the 10X Visium platform. However, data-driven unsupervised approaches are crucial for uncovering and understanding the complex structures of the human brain, especially in regions lacking clear histological boundaries. These methods enable the discovery of unknown spatial domains and assist in annotating areas that are difficult to delineate manually, thus playing a key role in brain research22. Consequently, in our dataset, to annotate the specific layer based on the layer markers proposed by Maynard et al.21 and to reveal other gene markers in each cortical layer, we first converted the raw spatial expression matrix into ~55 µm x ~ 55 µm pseudo-spots (110 bins x 110 bins/spot, named bin110 resolution), each representing approximately one spot in the 10X Visium platform to facilitate comparable analyses. At the bin110 resolution, a total of 338,410 pseudo-spots were obtained across the 12 samples, comprising 170,465 pseudo-spots from NA samples, 113,324 from moderate AD, and 54,621 from severe AD samples. Each pseudo-spot captured an average of 3150 counts and 2225 genes. We then utilized the Harmony algorithm23 to remove potential batch effect of the spatial transcriptional profiles from the 12 samples and employed a newly developed unsupervised spatially constrained-clustering (scc) algorithm17,24 across all samples to identify multiple layers in human PFC. Unlike conventional clustering methods for sn/scRNA-seq data25,26, scc groups pseudo-spots not only by their gene expression patterns but also by their spatial proximity. This ensures that clusters are formed based on both transcriptional profile similarities and the close spatial arrangement of the pseudo-spots within each cluster17. In our datasets, the data-driven clustering has successfully identified distinct layers and the WM in all 12 samples based on their unique transcriptional profiles (Figs. 1B and S1A). After aligning the gene expression sections with corresponding adjacent H&E sections, we excluded the pseudo-spot without brain tissue or folded regions and plotted all the pseudo-spots on a two-dimensional panel by uniform manifold approximation and projection27 (UMAP; Figs. 1B-C and S1B). To identify the transcriptional markers of each cortical layer and the WM, we carried out differential gene expression (DGE) analysis by comparing each individual cluster against all the other clusters (Fig. 1D; Methods). We noticed that specific markers for each layer, AQP4 in Layer I, HPCAL1 in Layer II and III, PVALB in layer IV, PCP4 in layer V, KRT17 in layer VI, and MBP in WM (Fig. 1E), were consistent with the layer markers identified by Maynard et al.21, which validated our identified laminar structures of PFC. In addition, we identified multiple marker genes for each specific layers across the cortical cortex, including MT-RNR2 in layer I, TMEM59L in layer II and III, NEFM in layer IV, TUBB2A in Layer V, DIRAS2 in layer VI, and PLP1 in WM (Fig. 1F; Supplemental Data 2). Together, our results highlighted the capability of Stereo-seq platform for delineating a high-resolution, fully data-driven spatial transcriptomic atlas of the human brain with transcriptome-wide coverage.
To assess the impact of AD progression on the laminar architecture, we used a robust linear modelling-based method, Propeller28, to compare the proportion of each layer between samples with different stages of AD. The proportion of each layer is indicative of its thickness relative to the total thickness of the gray matter. Although the layer proportions appeared similar between the moderate AD and NA groups, we noticed a significant reduction in the proportions of layers II-VI in the severe AD samples (Fig. 2A, B). This finding underscores the disruption of laminar architecture and the occurrence of human PFC atrophy in later stages of AD29. The increased proportion of layer I observed in severe AD samples may be related to extensive neuronal degradation30 and the high reactivation of astrocytes (Ast) in layers II-VI, a response to the neuronal inflammation31. Since reactivated Ast in layer II-VI often express genes typically associated with Ast in layer I, such as GFAP and NEAT131 (Fig. 2C), this similarity may cause these areas being classified as layer I in our clustering analyses for layers.
A Proportion of the cortical layers in each group. The color represents cortical layers I-VI. B Bar plot of the layer proportion across NA, moderate, and severe AD groups. One-way ANOVA and two-way t-test was conducted to identify significant differences in proportion between NA, moderate, and severe AD groups for each layer. False discovery rate (FDR) adjustment was applied for the P-value adjustment. N.S. indicates non-significant (adjusted P-value > 0.05) and *** represents adjusted P-value < 0.01. The number above the N.S. and *** indicate the specific adjusted P-value for each comparison. C The expression level of the GFAP and NEAT1 in NA, moderate, and severe AD groups. Two-side t-test and FDR adjustment were applied for the adjusted P-value calculation. The adjusted P-value for the DGE analysis of GFAP between the NA and severe AD groups is 0.0, and between the moderate and severe AD groups is also 0.0. Similarly, the adjusted P-value for NEAT1 between the NA and severe AD groups is 0.0, and between the moderate and severe AD groups is 0.0. *** represents adjusted P-value < 0.01. D Number of the layer-specific up- and down-regulated genes of the pairwise DGE analysis for each layer and the WM across all three groups. E Significant layer-specific up-regulated genes in each group after the pairwise DGE analysis for the WM, layer I, II/III, and V among all three groups. Size of the spot indicates the proportion of the cells expressing the genes, and the color represents mean expression level of the genes in each group.
Furthermore, we conducted a layer-specific pairwise DGE analysis (Methods) among the NA, moderate, and severe AD groups to decipher the layer specific AD pathological changes in different AD stages (Supplemental Data 3). Notably, the number of downregulated genes in the moderate and severe AD groups significantly exceeds that of upregulated genes when compared to the NA group (Fig. 2D). In comparing NA and moderate AD groups, we observed that the WM exhibits the highest number of down-regulated genes. This suggests that significant pathological alterations in transcriptional profiles occur in the WM due to the WM hyperintensities at the early stage of AD32. Additionally, we observed that while the strongest disease-associated changes appeared early in AD progression and were highly specific to each cortical layers, the genes upregulated at later stages were common across all six layers and the WM (Figs. 2E and S2A). These late-stage genes were predominantly related to mitochondrial function, highlighting significant mitochondrial dysfunction across all layers and the WM in the later stages of AD33. In specific, genes downregulated in the WM in both moderate and severe AD groups were associated with synaptic function (e.g., SNAP25, S100B, CALM2), myelin maintenance (e.g., MBP, GFAP), and neurotransmitter synthesis (e.g., SLCO5A1, SEMA3B). This suggests impaired neurotransmission in the WM of AD samples. Furthermore, we observed that the expression level of QDPR was highest in the moderate AD group compared to the NA and severe AD groups. Quinoid dihydropteridine reductase (QDPR; encoded by QDPR) is crucial for maintaining the stability of tetrahydrobiopterin (BH4)34, which is sensitive to oxidative stress in AD samples. A previous study using a mouse model suggested that BH4 improves recognition memory in AD without altering Aβ and NFTs35. The upregulation of QDPR in the moderate AD group likely reflects a compensatory mechanism in response to BH4 deficiency in the WM during the early stage of AD. In addition to QDPR, the neuroinflammation-related gene, SPP136, and the cellular stress response-associated gene, SLC38A237, are also highly expressed in the moderate AD group but are least expressed in the severe AD group. These gene alterations suggest that significant inflammation and stress-induced changes in white matter are evident at the early stages of AD but diminish in the late stage of the disease. In contrast, across the six layers of the neocortex, unlike in white matter, multiple genes related to neuron protection (e.g., MEF2D, TXN)38,39 are downregulated in the moderate and severe AD groups compared to the NA group. Notably, QDPR and SPP1 are upregulated across all six layers in both moderate and sever AD groups in contrast to NA group, indicating that the cellular responses to stress and inflammation in the gray matter persist even in the later stages of AD (Figs. 2E and S2A).
Given the high selective vulnerability of neurons in layers II, III and V to AD40, we examined the layer specific differential expression genes (DEGs) in moderate AD group compared to NA groups within these layers. Compared to NA group, genes related to stress responses (KEAP141 and WFS142) are specifically upregulated in the layer II and III in moderate AD group. Further, one interesting gene, DAPK1, encoding Death Associated Protein Kinase 1 (DAPK1), was specifically highly expressed in the layer II and III in moderate AD group compared to the NA group. Given that the DAPK1 is a calcium/calmodulin-dependent serine/threonine kinase to regulate neuron apoptosis and autophagy43, the high expression of this gene in layer II and III may be the key factor for the layer II and III vulnerable to the early stage of AD. In layer V, we noticed that the CSRP1, a regulator for the cytoskeletal organization and cell growth44, was specifically highly expressed in the layer V in the moderate AD group compared to NA and severe AD groups. The specific upregulation of this gene in the layer V suggest the dysfunction of the neuronal development and survival under the high stress caused by the inflammation. We have also checked the specific DEGs in other cortical layers. Interestingly, the APP gene (encoding the Amyloid Precursor Protein; APP) is specifically upregulated in the layer I in the moderate AD group compared to the NA groups. The expression level of the APP is positively correlated with Aβ production, which is critical in mediating synaptic plasticity45. Since layer I neocortex is sparsely populated with neurons and heavily innervated by synapse from the neurons in lower layers, especially the pyramidal neurons in layer III and V46, the high expression of APP at the early stage of AD in the layer I suggests the synaptic dysfunction of the pyramidal neurons in layer III and V. In summary, the layer-specific DGE analysis in our study has uncovered the molecular mechanisms underlying the significant pathological alternations in the WM and the pronounced selective vulnerability of neurons in layer II, III, and V at the molecular level.
Layer-layer interaction networks reveal alteration of inter-layer communications in AD
The interactions between cortical layers are essential for integrating sensory information, supporting cognitive functions, and facilitating communication within the brain. These interactions involve the vertical transmission of signals, where neurons spanning several layers can integrate inputs and produce outputs47. In this study, we have constructed the layer-layer communication networks based on the database of interactions among ligands and receptors48 (Methods), aiming to examine the variances in the vertical connections across each cortical layer and the WM between the NA samples and the moderate/severe AD samples. We observed a reduction in the number of interactions across all layers with the progression of AD, decreasing from 3571 in the NA group to 3064 in the moderate AD group and 2244 in the severe AD group (Fig. S2B). This pattern indicated the collapse of neural connection networks due to the neurodegeneration in AD, a consequence of Aβ plaques and tau protein aggregation7. Moreover, we observed a decline in the incoming and outgoing signaling from layers II-VI and the WM, but intensified communications between layer I and the other layers, with the progression of AD (Fig. S2C), possibly due to the neurodegeneration in layers II-VI and an increased proportion of layer I in the severe AD group.
The layer-layer communication networks in each AD stage showed distinct interaction patterns (Fig. 3A). After pair-wise comparisons of the number of interactions among different layers between the NA, moderate, and severe AD groups (Methods), we observed that the interactions between the layers IV and V decreased with AD progression, along with the reduction of outgoing interactions from layers IV-V to layers II, III, and VI (Fig. 3B). As the layers II-V are crucial in cortical information collection, processing and distributing outputs to subcortical structures49,50, the breakdown of the vertical networks leads to the cognitive impairments in AD51. Further, notable alterations of communication strength related to the Glutamate and neurexins (NRXN) signaling, associated with the neuronal signal transmission52,53, were observed in the moderate AD group compared to the NA group (Fig. S2D) in all layers and the WM.
A Networks of layer-layer interactions across six cortical layers and the WM in NA, moderate, and severe AD groups. The colors of the dots and edges represent the specific layers and the outgoing signaling emanating from them. Number on the edge represents the number of the LR pairs. B Pair-wise comparison of the number of outgoing and incoming LR pairs among different cortical layers and the WM across the NA, moderate, and severe AD groups. The color of the squares in the heatmap represents the number of increased (red) or decreased (blue) signaling in the first group compared to the second one. The top color bar represents the total number of changes (increases or decreases) in incoming signaling, and the right color bar corresponds to changes in outgoing signaling, both comparing the first group to the second. C The communication strength comparison of the specific LR pairs of Glutamate (from layer V to other layers) and NRXN (from layer I to other layers) signaling pathways across control, moderate and severe AD groups. The x-axis represents the direction of the LR pairs, and the y-axis indicates the specific LR pairs. Permutation test and Bonferroni correction were used for adjusted P-value calculation. Dot size represents the adjusted P-value, and the color reflects the communication strength of the LR pairs.
Since layer V is the main output layers in the cortical structures54, our investigation delved into the alterations of the communication strength in each ligand-receptor (LR) gene pair (Methods) related to Glutamate and NRXN in layer V across NA, moderate, and severe groups, aiming to reveal the dysregulated inter-layer LR interactions that may be driving the deterioration of cognitive functions. We noticed that the SLC1A2 and GLS2—GRIA2 LR pairs, outgoing from layer V to layers II-VI, showed a considerable decline in the moderate AD group and were completely absent in the severe AD group, in contrast to the NA group (Fig. 3C). The SLC1A2, encoding the excitatory amino acid transporter 2, is a glutamate receptor gene and responsible for clearing the neurotransmitter from the synaptic cleft55. GLS2, encoding glutaminase 2, is essential for maintaining the balance between the production and recycling of glutamate for neurotransmission56. GRIA2 encodes subunits of the alpha-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid (AMPA) receptors, which are integral to fast synaptic transmission in neurons57. Additionally, we also observed the loss of interactions between SLC1A2 and GLS2 ligand genes and other AMPA receptor subunit encoding genes, including GRIA1, GRIA3, and GRIA4, from layer V to layers II-VI in the severe AD group (Fig. 3C). Further, the interaction between SLC1A2 and GLS2 ligands and the subunits of metabotropic glutamate receptors (mGluRs), including GRM5 (encoded by GRM5) and GRM7 (encoded by GRM7) also diminished in the severe AD group. These receptors play crucial roles in the central nervous system by modulating neurotransmitter release and synaptic plasticity, impacting various physiological and pathological processes58. The decreasing of the Glutamate strength in the moderate group compared to NA may be due to the downregulation of the SLC1A2 and GLS2 in layer V at early stage of AD (Fig. S2E). Interestingly, while the expression level of SLC1A2 and GRM7 were increased in the severe AD group compared to the moderate AD group, the low expression of GLS2 led to a reduction in the SLC1A2 and GLS2—GRM7 interaction in the severe AD group. In addition, from layer V to layers II-V, the interaction strength of NRXN3-CLSTN1, the ligand-receptor pairs associated with presynaptic differentiation59, decreased in the moderate and severe AD groups, compared to the NA group (Fig. 3C). The reduction of this interaction in AD groups may be the underlying mechanisms for presynaptic disruption which was previously observed in AD samples60. Further, due to the low expression of the CLSTN1 across layer II to V in the severe group compared to NA and moderate AD group, NRXN3 in layer V was compensatorily upregulated in the severe group in contrast to NA and moderate AD groups (Fig. S2E). Together, our results uncovered significant alterations in inter-layer communications within AD samples, particularly in neurotransmission processes and the upregulation of SLC1A2, GRM7, and NRXN3 in later stage of AD as a compensatory mechanism. These findings could help elucidate the key mechanisms driving cognitive decline in AD.
Identification of the location of brain segments under high stress due to NA and AD in human PFC
As the brain ages, neurons become increasingly vulnerable to oxidative stress, inflammation, and protein aggregation61, contributing to the accumulation of Aβ amyloid into plaques62 and NFTs formation63. Additionally, the presence of Aβ plaques and NFTs intensifies neuronal stress, leading to further neurodegeneration64. Furthermore, Ast and microglia (Mic) surrounding highly stressed neurons become reactivated and play a crucial role in the processes of Aβ clearance and neuroprotection65. The variations in how neurons and surrounding glial cells respond to stress in both NA and AD remain poorly understood. To investigate this at single-nucleus resolution, we first pinpointed the potential locations of the areas under the high stress in all samples. Considering the heightened vulnerability of excitatory neurons (Ex) to aging62 and AD66, coupled with the known positive correlation between cellular apoptosis and elevated mitochondrial gene ratio (number of mitochondrial genes/total detected genes) in cells64, we presumed that Ex with a high mitochondrial gene ratio may represent Ex under significant stress caused by aging or AD pathological hallmarks, and the positions of these highly stressed Ex mark the potential high stress focal point in human PFC.
To precisely identify the location of these highly stressed neurons, we refined our analytic strategy by converting the raw expression matrix into ~25 µm x ~ 25 µm pseudo-spots (50 bins × 50 bins/spot, named here bin50), which is the default resolution provided by Stereopy24. Since the mitochondrial genes are exclusively expressed within the mitochondria in the cell cytoplasm67, pseudo-spots at bin50 resolution, which capture gene expression patterns from both the cell nucleus and cytoplasm, can effectively detect mitochondrial gene expression. This capability aids in accurately identifying pseudo-spots that either fully or partially cover the neurons under high stress. After filtering out the pseudo-spots with low sequencing quality (pseudo-spots with lower than 400 genes detected), we obtained 924,780 pseudo-spots with an average of 834 (and median = 758) genes detected per pseudo-spot. To extract the pseudo-spots partially or fully covering the Ex, we first utilized Harmony23 to integrate the pseudo-spots from all the samples and then clustered them into multiple distinct subsets based on their transcriptional profiles. After the identification of the marker genes in each cluster, we annotated the clusters as Ast, endothelial cells (End), Ex, Inh, Mic, oligodendrocytes (Oli), and oligodendrocyte progenitor cells (Opc), based on the cell type specific markers (e.g., AQP4 in Ast, CLDN5 in End, SNAP25 in Ex, GAD1 in Inh, CD74 in Mic, MBP in Oli, LHFPL3 in Opc) proposed by a previous snRNA-seq study on human PFC13 (Fig. 4A, B; Supplemental Data 4). To check the accuracy of our annotation results, we aligned the annotated bin50 pseudo-spots with corresponding adjacent H&E sections. Ex was predominantly situated in the neocortex, while a high concentration of Oli was observed in the WM (Figs. 1B; 4C and S1A), which strongly support that although the pseudo-spots were not at single-nucleus resolution, our results still accurately delineated the proximate positions of the Ex. In addition, we noticed that in moderate and severe AD groups, and especially in the severe AD group, a large proportion of Ex and Oli were filtered out due to the low gene number detection (Figs. 4C and S3A), reflecting the Ex degradation30 and changes in the WM caused by Wallerian-like degeneration32 in AD.
A UMAP visualization of pseudo-spot clusters. B Expression levels of the gene markers in each cluster. The color of the violin plot reflects the median expression of the gene in each cluster. Based on the gene markers, we annotated the clusters as astrocytes (Ast), endothelial cells (End), excitatory neurons (Ex), inhibitory neurons (Inh), microglia (Mic), oligodendrocytes (Oli), and oligodendrocyte progenitor cells (Opc). C Distribution of different cell clusters on the captured area of Stereo-seq platform (B01809A3 in NA, B01809C2 in moderate, and B02008C6 in severe AD group), and the proportions of cell clusters in all samples from the NA, moderate, and severe AD groups, respectively. D The violin plot of the mitochondrial gene ratio of Ex in each sample. The x-axis represents 12 samples and the y-axis indicates the mitochondrial gene ratio. The mitochondrial gene ratio ranges are as follows: for the NA group, maxima 16.3–29.5%, 75th percentile 6.6–8.6%, median 5.5–7.4%, 25th percentile 4.4–6.5%, and minima 1.5–3.0%; for the Moderate AD group, maxima 13.3–25.7%, 75th percentile 5.5–8.7%, median 4.5–7.3%, 25th percentile 3.7–6.1%, and minima 0.9–2.2%; and for the Severe AD group, maxima 21.2–35.1%, 75th percentile 12.2–21.0%, median 10.6–18.4%, 25th percentile 9.2–16.0%, and minima 3.1–5.4%. The dashed line represents the threshold of the mitochondrial gene ratio (15%) to classify the Ex into highly stressed or normal ones. E Distribution of the highly stressed Ex in NA, moderate, and severe AD groups. The yellow dots represent the pseudo-spots covering the highly stressed Ex, and the blue dots are pseudo-spots containing the normal Ex and other types of the cells. F Proportion of the highly stressed Ex pseudo-spots among all Ex pseudo-spots in NA, moderate, and severe AD groups. G The significant up- and down-regulated genes of the highly stressed Ex pseudo-spots, compared to normal Ex pseudo-spots. Two-sided t-test and FDR adjustment were used for the adjusted P-value calculation. Number indicates the exact adjusted P-value and *** represents adjusted P-value < 0.01.
To check if the mitochondrial gene ratio is a reliable criterion to identify the highly stressed neurons, we first applied the non-parametric Kruskal-Wallis test to assess whether significant differences exist in the median mitochondrial gene ratio across all pseudo-spots covering Ex among the three groups (Fig. S3B). In Ex, our results indicated that, while the median mitochondrial gene ratio differed significantly across the three groups (Bonferroni, corrected P-value < 0.001), the medians in the NA (6.73) and moderate AD (6.47) groups were quite similar. In contrast, the median percentage of mitochondrial gene in the severe AD group (12.81) was considerably higher in the outer groups, suggesting a large proportion of Ex were under high stress at the later stage of AD. Further, we examined the differences in median of mitochondrial gene ratio in pseudo-spots containing other brain cell types among the three groups, including Ast, End, Inh, Mic, Oli, and Opc. Similar to the pattern observed in Ex, the differences in median mitochondrial gene ratio between NA and moderate AD group in each cell type, although statistically significant (Bonferroni corrected P-value < 0.001), were relatively minor (~6.3 to ~6.8). However, the median mitochondrial gene for each cell type was notably higher in the severe AD group (~12.5). These findings align with the results reported by Tonnies et al.68 that cellular stress across all cell types is moderate at the early stage of AD and increased during the later stages of AD. Therefore, the mitochondrial gene ratio is a reliable criterion to identify the highly stressed neurons from normal ones. Due to the lack of an established mitochondrial gene ratio threshold for distinguishing stressed Ex from healthy Ex in spatial transcriptome data, we set a threshold at 15% based on the distribution of mitochondrial gene ratio across the pseudo-spots covering the Ex from all the samples (Fig. 4D), that is, pseudo-spots annotated as Ex that exhibit ≥15% mitochondrial gene ratio were classified as containing neurons under high stress. In the NA group, we noticed that only 0.13% of the Ex pseudo-spots contain Ex under high stress, indicating minimal high-stress focal points in NA. In contrast, the proportion of Ex under high stress was slightly increased in the moderate AD group and was dramatically higher in the severe AD group (Fig. 4E, F), highlighting the prevalence of Ex with high stress in the later stage of AD.
We further checked the DEGs between the pseudo-spots covering highly stressed neurons and those containing the normal ones. Besides the mitochondrial genes (e.g., MT-RNR2, MT-ND4, and MT-CYB), multiple AD-related genes, including GFAP69, AQP470, and DPP1071, were also upregulated in the pseudo-spots covering highly stressed Ex, compared to those containing normal Ex (Fig. 4G). In the pseudo-spots covering highly stressed neurons, the high expression of Ast markers, GFAP and AQP4, indicated that these areas also include the transcriptional profiles of adjacent Ast. This observation reflects that when neurons are subjected to high stress, the nearby Ast become reactivated and migrate towards these neurons to support their survival72. Meanwhile, consistent with previous transcriptional studies in AD73,74,75,76, ENC1, CALM2, CCK, RGS4 were downregulated in the pseudo-spots covering highly stressed Ex (Fig. 4G). Together, these findings support our hypothesis that pseudo-spots containing Ex with elevated mitochondrial gene ratio represent brain areas under high stress affected by NA or AD pathology hallmarks.
Transcriptional divergence at single-nucleus resolution between brain cells impacted and not impacted by NA and AD pathology hallmarks
Since human brain is a complex tissue containing multiple heterogeneity cell types, including Ast, Ex, Inh, Mic, End, Oli, and Opc13, applying high resolution spatial transcriptome technology to identify the cell type specific transcriptional alternations is essential to unveil the molecular mechanism and cellular communication in AD. To further delineate the transcriptional landscape at single-nucleus resolution, we applied a deep learning model24 for the nucleus segmentation to delineate the cell nuclei based on the nuclei staining image (Methods). After filtering out the low-quality nuclei (i.e., nuclei with <150 detected genes), we analyzed 398,741 nuclei, with an average of 354 (and median=279) genes detected per nucleus. We performed clustering analysis to identify the nuclei subgroups (Fig. 5A) and conducted pairwise DGE analyses among the subgroups to identify the specific marker genes for each nucleus cluster. Based on the marker genes in each cluster, we successfully annotated nuclei as the Ast (AQP4, GLUL), End (CLDN5, IFITM2), Ex (NRGN, SYT1), Inh (GAD1, GAD2), Mic (CD74, CD14), Oli (MBP, PLP1), and Opc (VCAN)13 (Figs. 5B and S4A). The high distribution of the Oli in the WM and Ex in gray matters substantiated the accuracy of our annotation (Figs. 5C and 1B).
A UMAP visualization of clusters of nuclei. B Heatmap of the gene marker expression in each cluster of nuclei. The color represents the expression levels of the gene in each cluster of nuclei. C The nuclei type distribution on B01809A3 (NA) sample. D Distribution of nuclei in B01809C2 (moderate) across levels I, II, and III in the concentric circle analysis. The red dots represent the selected nuclei at each level, and gray dots indicate other nuclei. E The nucleus-type specific DGE analysis between the nuclei in level I and III across NA, moderate, and severe AD groups. The color represents the mean expression of the genes in each level. F The nucleus-type specific DGE analysis between the nuclei in NA, moderate, and severe AD groups at level I.
Compared to the healthy neurons, those experiencing high stress due to NA and AD pathology hallmarks exhibit distinct transcriptional profiles as part of their stress responses77,78. Additionally, the mechanisms of these stress responses can differ between neurons affected by NA and those influenced by AD pathology79. Furthermore, glial cells located near the areas under high stress are more actively involved in neuroprotective actions to aid neuronal survival, in contrast to those situated farther away72,80. Thus, to enhance our understanding of the cellular responses to the stress caused by NA and AD pathology, we extracted the nuclei within approximately 1250 µm from the location of high stress focal point (identified at bin50 resolution) across all samples, totaling 86,626 nuclei, and performed concentric circle analysis (Methods). Briefly, the extracted nuclei were classified into three groups (levels I to III) according to their proximity to the high-stress focal point (Fig. 5D). Considering that neuron somas range from approximately 4 ~ 100 µm in size, to fully include a whole stressed neuron and the nearby glial and endothelial nuclei, we categorized nuclei located within 250 µm to the stressed neurons as level I, 250-750 µm as level II, and 750-1250 µm as level III.
Given the close proximity and highly similar transcriptional profiles between nuclei in levels I and II, we focused the nuclei type specific DGE analysis between nuclei in levels I and III across NA, moderate and severe AD groups, respectively, in order to distinguish gene expression patterns between nuclei that are close to and those that are more distant from stressed neurons at different stages of AD. Our findings indicated that as with progression of AD, there was notable nucleus type specific differentiation in gene expression patterns between nuclei of level I and level III (Fig. 5E; Supplemental Data 5). For instance, in the NA group, Ex at level I significantly increased the expression of neuroprotective genes, including VGF81, LMO482, and NRN183, when compared to level III. This highlights Ex’s mechanisms in mitigating stress from NA, thereby enhancing neuron survival. However, this pattern of upregulation for VGF, LMO4, and NRN1 in Ex at level I, in contrast to level III, was not present in the moderate and severe AD groups. In the severe group, compared to level III, Ex at level I notably enhanced the expression of genes for heat-shock proteins (HSP90AA1 and HSPB1) as a reaction to neuroinflammation and neurotoxic substances84; additionally, there was a significant increase in the expression of genes associated with the clearance of Aβ peptides, such as SORT185. The long non-coding RNA (lncRNA) NEAT1, which contributes to neuron damage by downregulating microRNA-27a-3p86, was also significantly upregulated in level I Ex in the severe AD group, in contrast to Ex in level III. In the NA group’s Ast, when compared to those in level III, there was a notable increase in the expression of SLC1A3 in the level I Ast. This gene is known for its role in protecting neurons against glutamate-induced excitotoxicity by clearing excess glutamate from the synaptic gap87. Like Ex, Ast in the severe group at level I, in comparison to level III within the same group, also showed a marked increase in the expression of heat-shock protein genes (e.g., CRYAB, HSPB1, HSPA1A, HSP90AA1, HSP90AB1). Furthermore, unlike the nuclei type-specific upregulated genes observed in level I compared to level III in the moderate group, there is a notable upregulation of heat-shock protein-coding genes in Ast, Ex, Inh, and Oli in level I compared to level III in the severe group. This observation aligns with the findings proposed by Mathys et al.13, which suggest that AD-associated changes manifest early in the pathological progression and are highly cell-type specific, while genes upregulated in later stages are common across cell types and predominantly involved in the global stress response.
To further investigate the divergence in cell-type-specific transcriptional profiles during the progression of AD at equivalent distances from high-stress focal points, we conducted pairwise DGE analysis within the same level (I to III), comparing the NA, moderate, and severe AD groups (Fig. 5F; Supplemental Data 6). Interestingly, at level I, we observed that the PSAP gene was highly enriched in Ast, Ex, Mic, and Opc in the NA group, with its expression decreasing as AD progressed. Prosaposin (PSAP), encoded by the PSAP gene, plays a crucial role in maintaining neuronal survival88. The reduced expression observed in the moderate and severe AD groups compared to the NA group may suggest a deficiency in neuroprotective ability in these stages, particularly when neurons are under high stress. The upregulated of PSAP in Inh and Oli in the moderate group compared to the NA group may illustrate a compensatory mechanism for neuroprotection. Further, we have noticed the APP gene was decreased in Ex, Inh, and Oli in level I in moderate and severe AD groups, revealing the dysfunction of the APP signaling in the high stress focal point in AD. Together, our results demonstrated the intricate dynamics and differences of cellular responses to NA and AD, revealing distinct cell-specific gene expression alterations linked to the spatial position relative to stressed neurons.
Cell-cell communication networks reveal the reduction of neuron-protective LR interactions in cells near stress-affected neurons in AD
The collaborative mechanisms between neurons and glial cells play a vital role in inhibiting the accumulation of Aβ and the aggregation of tau proteins89. While prior snRNA-seq studies90,91 have uncovered distinct intercellular interactions in AD cases and NA controls, the spatial information, particularly the proximity of glial cells and End to the neurons under high stress in AD and NA, has been largely ignored. As a result, to assess whether neurons under stress and their surrounding glial cells and End employ similar or distinct mechanisms in response to the stress caused by NA and AD pathology, we conducted cell-cell communication analysis48 across nuclei within levels I and III for all three groups (NA, moderate, and severe AD groups), (Fig. 6A-B). We found that the level I nuclei showed higher number and intensity of LR pairs than the level III nuclei in the NA group (Fig. 6A), suggesting that intercellular interactions among the neurons, glial cells, and End were enhanced in cells proximal to stressed neurons during NA, potentially as a protective mechanism against the degradation of stressed neurons (Fig. 6B). However, as AD progressed, the communication networks among level I cells became compromised, indicating that the intercellular interactions close to stressed neurons are disrupted in moderate and severe AD groups91 (Fig. 6A-B).
A The total number (top) and the strength (bottom) of the LR pairs in level I and III groups across control, moderate, and severe AD groups, respectively. B Networks of cell-cell interactions show the number of LR pairs (edges) between nuclei clusters (nodes) within level III and I across NA, moderate, and severe AD groups, respectively. The colors of the dots and edges represent the specific types of nuclei and the outgoing signaling emanating from them. The number on the edge indicates the number of the outgoing signals. The size of the spot are proportional to the number of nucleus within each level of specific AD group. C The communication strength of the specific LR pairs of the nuclei in level I and III in the NA, moderate, and severe AD groups, respectively. Nuclei type in each row indicates the nuclei expressing the ligands and the column indicates the one expressing the receptors. Permutation test and Bonferroni correction were used for adjusted P-value calculation. The x-axis represents the level I and III across NA, moderate, and severe AD groups in each nucleus type. The color of the dots reflects the communication strength, and the size of the spot represents the adjusted P-value. D, E The chord diagram of the PSAP-GPR37L1 and APP-CD74 LR pairs of the nuclei in level I and III in the NA, moderate, and severe AD groups, respectively. Each segment of the circle in different color represents different types of nuclei. The chords connecting the segments represent interactions between two types of nuclei, and the thickness of each chord reflecting the strength of the interactions.
We further noticed that PSAP and APP signaling were dysregulated in the moderate and severe AD groups at level I (Fig. 6C). PSAP released by Ex, End, Mic, Oli, and Opc acts on the GPR37L1 (encoded by GPR37L1) on Ast to activate the motility of Ast and release of diffusible neuroprotective factors to shield the neurons affected by neurotoxic damage92. The APP (encoded by APP) released by Ex and Inh, interacts with the CD74 (encoded by CD74) on Mic to inhibit Aβ production93. These interactions were impaired or diminished in the nuclei proximal to the stressed neurons (level I) in moderate and severe AD groups (Fig. 6C-E). Together, our results suggest that, compared to NA, highly stressed neurons and surrounding glial cells in AD samples lose their ability for Aβ clearance and neuroprotection. This loss can contribute to the formation of Aβ plaques and exacerbate neuronal degradation, potentially explaining the high levels of Aβ deposition and brain atrophy observed in AD.
Co-expression networks at single-nucleus resolution uncover key gene markers and potential regulatory transcription factors in AD progression
Although our results uncovered nucleus-type specific changes associated with brain areas under high stress caused by NA and AD pathology, the complex relationships and patterns among co-expression gene modules, as well as the key regulators involved in the neuron protection and the clearance of AD pathological hallmarks, remain unclear. Using hdWGCNA94, we constructed the nucleus type specific weighted gene co-expression networks on the 2,000 most variable genes for each nucleus type from levels I-III across all samples. We successfully identified two co-expression gene modules in Ex and Inh, respectively (Figs. 7A and S4B-C), but not in the other nucleus types due to the low sequencing depth and limited nuclei numbers. We noticed that in these modules, several hub genes have been reported to be related to AD. For instance, in the Ex1 module, the hub gene UCHL1, encoding the ubiquitin C-terminal hydrolase L1 protein, is a major neuronal enzyme involved in the elimination of misfolded proteins95. The decreasing expression of this gene is responsible for Aβ42 accumulation95. Additionally, within the Ex1 module, we identified two hub genes, ENC1 and RTN1, which are key genes implicated in the transition from asymptomatic to symptomatic AD96. In the Inh1 module, the hub gene VAMP2 is known to be related to neurodegenerative disease where a reduction of VAMP2 expression was associated with cognitive decline97, and another hub gene, PRNP, which encodes prion protein, is involved in neuroprotection to excitotoxicity98 and has inhibitory effects on BACE1 to decrease Aβ production99. We also noticed that one hub gene in the Inh2 module, HSPA8 (encoding heat-shock protein family A member 8), can directly disassemble RHIM-amyloids to inhibit necroptosis signaling in cells and in mice100.
A The gene co-expression modules based on the Ex (Ex1) and Inh (Inh1 and Inh2) from level I to III across NA, moderate, and severe AD group. The nodes represent the hub genes, while the edge connecting two nodes indicates the co-expression of those genes. B-D GO Enrichment analysis on the top 50 hub genes in the Ex1, Inh1, and Inh2 modules. The length of the bar indicates the gene numbers enriched in the GO term and the color represents the adjusted P-values for enrichment analysis. Adjusted P-values were calculated through Fisher exact test and FDR adjustment. E Enrichment analysis to identify the Motif overlaps with Ex and Inh modules. Fisher exact test and FDR adjustment were applied for the adjusted P-value calculation. The color indicates the adjusted P-value.
To enhance our understanding of the biological functions of all four modules (Ex1, Ex2, Inh1, Inh2), we conducted GO enrichment analysis on the top 50 hub genes based on connectivity within each module. Our results indicate that all four modules are primarily associated with neuroprotective processes and inhibition of the formation of AD pathological hallmarks. For instance, the Ex1 module was significantly enriched in processes such as “Negative Regulation of Amyloid-Beta Formation”, “Negative Regulation of Protein Phosphorylation”, and “Negative Regulation of Peptidyl-Threonine Phosphorylation” (Fig. 7B), suggesting its critical role in mitigating the formation of AD pathological hallmarks. Besides the “Negative Regulation of Peptidyl-Serine Phosphorylation” GO term, the Ex2 module also supported processes like “Brain Development” and “Regulation of Nervous System Process” (Fig. S4D), which may regulate the neuronal survival. The Inh1 module was mainly enriched for processes such as “Positive Regulation of Synaptic Transmission”, “Negative Regulation of Cytokine Production”, “Response to Amyloid-Beta”, and “Regulation of DNA Repair” (Fig. 7C), while the Inh2 module was also enriched for terms related to Aβ clearance (Fig. 7D). These results underscore the involvement of these modules in neuroprotective responses against aging and AD-induced stress.
Additionally, we performed module-trait correlation analyses to assess the correlations of each module with AD progression (NA, moderate AD, and severe AD groups) and with spatial proximity to stressed brain areas (levels I-III groups). While the Ex2 module showed low correlation with AD progression (r = 0.0261, adjusted P-value < 0.01) (Supplemental Data 7) or spatial proximity to the stressed neurons (r = −0.0423, adjusted P-value < 0.01) (Supplemental Data 7), the Ex1 module exhibited negative correlations both with AD progression levels (r = −0.2241, adjusted P-value < 0.01) and spatial proximity to stressed neurons (r = −0.0921, adjusted P-value < 0.01) (Supplemental Data 7). Moreover, the Inh1 and Inh2 modules both showed significant negative correlations with AD progression (Inh1: r = −0.2193, adjusted P-value < 0.01; Inh2: r = −0.2577, adjusted P-value < 0.01) (Supplemental Data 7). This implied a diminished capacity in Ex to inhibit the formation of Aβ plaques and tau hyperphosphorylation when subjected to heightened stress in AD samples. Intriguingly, the Inh1 module also demonstrated a positive correlation with spatial proximity to high stressed neurons (r = 0.1196, adjusted P-value < 0.01; Supplemental Data 7), suggesting Inh close to the stressed brain areas may have an enhanced neuroprotective capability to support the Ex survival and maintain the Ex/Inh balance in brain101.
Given that the three modules (Ex1, Inh1, and Inh2) were negatively correlated with AD progression, the transcription factors (TFs) regulating these modules could serve as potential therapeutic targets to prevent the formation of AD pathological hallmarks and neuronal degradation in AD102. To this end, we utilized the MotifScan package103 to find the TFs significantly enriched within each module (Methods). We identified that KLF2 was significantly enriched in module Ex1 and Inh2 (Fig. 7E). Fang et al.104 have proposed that the upregulation of Kruppel-like factor 2 (encoded by KLF2) attenuates oxidative stress triggered by Aβ, improves mitochondria function, and reduce the rate of apoptosis. In addition to the neuroprotective TF, we noted that two TFs, KLF4 and KLF5 were significantly enriched in all the three modules (Fig. 7E). These two TFs are recognized for accelerating the progression of AD. Specifically, KLF4 was identified as a key mediator in promoting Aβ-induced neuroinflammation by exacerbating the release of pro-inflammatory factors105, and KLF5 can accelerate APP amyloidogenic metabolism and promote Aβ synthesis through binding to the BACE1 promoter106. Interestingly, one TF, ZNF460 showed the highest level of enrichment across Ex1, Inh1, and Inh2 gene modules (Fig. 7E), suggesting its potential role as a pivotal regulator in neuronal stress responses to NA or AD, as well as in neuron degradation processes.
Discussion
In this study, the Stereo-seq platform17, a high-resolution spatial transcriptome technology, was employed for the first time to develop the first most comprehensive, data-driven, transcriptome-wide molecular atlas of the adult human PFC ever achieved. Additionally, with the unparalleled resolution in spatial transcriptomics for human PFC to date, including six AD cases and six NA controls of the same sex and approximately matched by age with slightly older controls, our study uncovered significant molecular alterations in AD samples relative to NA controls at both cortical layer and single-nucleus levels. By analyzing high-definition landscapes of the human PFC in both AD cases and controls, we first identified six cortical layers and the WM using an unsupervised clustering algorithm (scc)17,24. Subsequently, we detected layer specific transcriptional alternations and laminar structural disruption across six cortical layers and the WM in PFC associated with AD. Furthermore, we have unveiled unique stress response mechanisms among neurons and adjacent non-neuronal cells, both during NA and throughout various stages of AD.
Here, we highlight some of our findings. First, at bin110 resolution, our study is the first one that demonstrated the increasing of the layer I proportion in neocortex at the later stage of AD. While the proportion of the cortical layers remained the same in the moderate AD group compared with the NA group, layers II-VI were diminished in the late stage of AD, which was not observed in current AD related spatial transcriptome studies on mouse models107, as well as human PFC107 and middle temporal gyrus14. An explanation for the discrepancy with previous studies is that our study included samples from more advanced stages of AD. Although multiple samples at the later stage of AD display extreme AD pathological hallmarks, there is considerable variability in pathological features that far exceed those of another, despite both being categorized at the same pathological level. For instance, in our study, besides the samples in the severe AD group (A02092E1 and B02008C6), B01809C2 sample in the moderate AD group also exhibited the highest levels of Aβ (Thal phase V and CERAD C3) and tau pathology (Braak stages VI) (Supplemental Data 1). However, after performing the immunohistochemistry for Aβ42 to assess Aβ abundance in these samples, we observed that A02092E1 was far more affected by the Aβ compared to B01809C2 (Fig. 1B). This suggests that even though these three samples (A02092E1, B02008C6, and B01809C2) have the highest level of AD pathology (Thal phase V, CERAD C3, and Braak stages VI), those two samples (A02092E1 and B02008C6) in the severe group exhibit substantially more pronounced AD pathological features than samples in other groups with similar pathological criteria. Additionally, since Ast reactivation markers, GFAP and NEAT131, were highly expressed in the severe AD group, the increase of layer I proportion reflects the atrophy of the PFC attributed to neuronal degradation108 and the reactivation of the Ast in response to toxic substance in the brain with AD pathology hallmarks109 at the later stage of AD. Besides the disruption of laminar architecture at the later stage of AD, we also observed that the most significant pathological alternations were in the WM rather than in the six cortical layers in the gray matter. The presence of WM hyperintensities in the early stages of AD32 may be a key factor contributing to this phenomenon. Furthermore, we identified an interesting gene, DAPK1, which is involved in regulating neuronal apoptosis43. DAPK1 was significantly enriched in the moderate AD group compared to the NA and severe AD groups, specifically in layers II and III. The elevated expression of DAPK1 in these layers within the moderate AD group may underlie the high selective vulnerability of layers II and III during the early stages of AD. Our study also revealed impaired interactions between cortical layers in AD samples compared to NA controls, particularly in Glutamate and NRXN signaling. These signaling, which are crucial for neurotransmission52,53, exhibited significant dysregulation in the moderate AD group compared to the NA group. This suggests that neuronal connectivity is compromised in the early stage of AD, contributing to cognitive decline.
Since the cellular responses to the stress caused by oxidative damage, inflammation, and protein aggregation are the key factors leading to Aβ plaque deposition110 and NFT formation63. We uniquely identified distinct stress response mechanisms at the single-nucleus level in neurons and adjacent glial cells, comparing the responses between NA and AD. In the NA group, our analysis revealed the motility of the nearby Ast was also activated through PSAP-GPR37L1 pairs, prompting their migration towards stressed neurons to offer protection92. We also observed the amplified APP-CD74 LR pairs between stressed Ex and adjacent Mic, which may enhance Aβ clearance93. However, these neuron protective LR pairs were found to be compromised in moderate and severe AD groups when neurons are under stress. In the moderate AD group, although Inh and End near the high-stress focal point (level I) compensatorily upregulated PSAP expression compared to the NA group, the downregulation of receptor GPR37L1 in the nearby astrocytes disrupted this protective interaction (Fig. 5F). In summary, our results suggest that impaired neuroprotection and Aβ clearance are key factors in the neuron degradation and deposition of Aβ plaques in AD brains.
To further support our findings that a decrease in neuroprotection and AD pathological hallmarks clearance contributes to the progression from NA to AD, we have conducted additional gene co-expression analysis. The nucleus-type specific gene co-expression networks revealed that in the Ex and Inh, the three gene modules, Ex1, Inh1, and Inh2, related to the Aβ plaque and NFT clearance progressively decreased during AD progression. Our findings indicate that while stressed neurons contribute to the accumulation of Aβ into plaques62 and the formation of NFTs63, in the context of the NA group, both stressed neurons and adjacent brain cells participate in the clearance of Aβ plaques and the dephosphorylation of proteins, which prevents the accumulation of Aβ plaques and tau aggregation, respectively. Conversely, in AD brains, this protective mechanism is compromised. Stressed neurons and surrounding cells lose their capacity to clear Aβ and maintain protein dephosphorylation, leading to the formation of AD pathology hallmarks and acceleration of AD progression. These observations align with previous research indicating that impaired Aβ clearance is a hallmark of AD samples111,112 and a decrease in tau phosphatase activity in AD brains contributes to an imbalance in the protein phosphorylation/dephosphorylation system, culminating in NFT formation113. Interestingly, one transcript factor, ZNF460, is a regulator with the highest enrichment in all three modules (Ex1, Inh1, and Inh2), and thus may be an important therapeutic target to promote neuron survival and AD pathological hallmarks clearance. Although the biological mechanisms linking ZNF460 to AD has yet to be established, a study by Liu et al.114 demonstrated that ZNF460 interacts with the apolipoprotein C1 (APOC1) promoter and enhances APOC1 transcription. This process is believed to contribute to the progression of gastric cancer114. Notably, APOC1, like apolipoprotein E (APOE), is involved in lipid metabolism115, and its H2 allele has been identified as a genetic risk factor for AD116. Future studies are needed to explore whether a similar interaction between ZNF460 and APOC1 occurs in the brain, and to assess its potential as a therapeutic target for AD. Overall, our study offers insights into the potential mechanisms underlying the progression of AD from NA to its late stages, and identified one key TF, ZNF460, that may regulate these mechanisms.
Despite the distinct advances and findings that this study contributes to the field, our study may also have some limitations that future studies with more advanced technologies and clinical samples may address. First, although Stereo-seq technology offers us the highest resolution among the current spatial transcriptome platforms, the median number of genes captured per nucleus ( ~ 300 median genes per nucleus) is significantly lower than that achieved by traditional snRNA-seq methods. While we successfully identified neurons, glial cells, and End at single-nucleus resolution, the reduced sequencing depth may potentially introduce bias in detecting significant alterations in certain transcriptomic molecules related to AD. Further, the low transcript capture ability limited us to identify the Aβ clearance and neuroprotection in highly stressed neurons and surrounding glial cells in AD in a layer-specific manner. Enhancements in mRNA capture efficiency could improve the sensitivity and accuracy of spatially resolved cellular taxonomy within the brain. Another limitation of our study is that we exclusively involved male samples, which might lead to bias when considering the impact of AD on females. Furthermore, as Stereo-seq represents one of the most advanced high-definition spatial transcriptome platforms, there are currently no mature software tools available to precisely identify interlayer communications using Stereo-seq data. Given that neurons primarily communicate through synapses, with most ligand release and receptor binding occurring far from the soma, cell-cell interaction analysis without spatial information may introduce bias when studying interlayer communications. Therefore, novel algorithms that incorporated spatial information for cell-cell interaction analysis on Stereo-seq data are urgently needed. Additionally, due to the inherent limitations of Stereo-seq technology, it is not feasible to perform immunohistology or HE staining alongside spatial transcriptional profiling on the same tissue section. This limitation presents a challenge in accurately aligning spatial transcriptome profiles with immunohistology staining images, potentially introducing bias when identifying cell-type-specific divergences in gene expression patterns between cells near AD pathological hallmarks and those in more distant regions. Despite these challenges, our findings offer crucial and valuable insights into the transcriptional landscape of AD on the human PFC at various resolutions, and they provide a detailed and systematic understanding of alterations in neocortical laminar architecture and specific stress responses at various stages of AD, compared to the NA.
Methods
Study subjects
A total of 12 postmortem human prefrontal cortex samples (Brodmann area 10) from age-matched male individuals were collected through the University of Kansas Alzheimer’s Disease Research Center (KU ADRC). Informed consent was approved by the KU Institutional Review Board (IRB) and obtained for all human participants. All participants in this study have signed repository consents permitting their data to be repurposed for future research. The study was approved by University of Kansas Medical Center Human Subjects Committee protocol 11132. All samples were obtained from Non-Hispanic White males age 70-95 years. We classified these samples into six AD cases and six controls based on Braak stages, Thal phases, and CERAD scores. The control group was defined based on the observation that their prefrontal cortex is not affected by NFT at Braak II-III stages117 and the low Aβ burden (CERAD score C0-1) in the six control samples. For details on human samples used in this study, please see Supplementary Data 1.
Statistics and reproducibility
No statistical method was used to predetermine sample size. No data were excluded from the analyses. The experiments were not randomized, and the investigators were not blinded to allocation during experiments and outcome assessment.
Tissue preparation, cryosection, stereo-seq library preparation, and sequencing
Stereo-seq is a state-of-art spatial transcriptome platform that captures mRNA from tissue sections using stereo chips. This technology achieves nanoscale resolution with a spot diameter of 220 nm, enabling the most detailed delineation of the transcriptome landscape currently available. Further, distinct from other spatial transcriptome platforms, the Stereo-seq chip has the capability to convert a few hundred spots into a pseudo-spot by combining the transcriptional information from the selected DNA nanoball (DNB) patterned array17, providing us the opportunity to depict transcriptional profiles at multiple resolutions.
To construct a comprehensive spatial transcriptome atlas of human PFC, the BA10 area from 12 postmortem human brains was harvested and immediately flash frozen, embedded with Tissue-Tek OCT medium (Cat # 4583, SAKURA FINETECK USA Inc.) in liquid nitrogen, and then stored at -80°C until ready for the Stereo-seq pipeline. Cryosections were cut at a thickness of 10 µm and mounted on Stereo-seq permeabilization chips (Cat # 210CP118, STOmics America Ltd) or transcriptomics chips (Cat # 210CT114, STOmics America Ltd). Tissue fixation and the following spatial transcriptomics procedures were performed according to the vendor’s manual and previous publications17,118. In brief, the tissue section on the Stereo-seq chip (10 mm x 10 mm) was incubated at 37 °C for 5 mins, and subsequently fixed in pre-cooled methanol (Cat # 34860, Sigma) at −20 °C for 30 mins. Once the fixation was completed, the chip was placed under a ventilation hood to allow residual methanol to air dry. The tissue section on the chip was then stained with nucleic acid reagent (Cat # Q10212, 0.5% v/v, Invitrogen) for 5 mins and subsequently washed with 0.1X SSC buffer (Cat # AM9770, 0.05 U/mL RNase inhibitor, Ambion). The nuclei images were captured using a Zeiss Axio Scan Z1 microscope (at EGFP wavelength). Subsequently, the tissue section was incubated in the permeabilization buffer (Cat # 111KP118, STOmics America Ltd) for 12 mins at 37 °C. Stereo-seq transcriptomics chip-captured RNAs from the permeated tissue were then reverse transcribed for 3 hours at 42°C. Next, the tissue was removed and the cDNAs were released from the chip using the transcriptomics reagent kit (Cat # 111KT114, STOmics America Ltd). After the cDNA obtained was size-selected, amplified, and purified, the concentration was quantified by Qubit dsDNA HS assay kit (Cat # Q32854, Invitrogen). Next, 20 ng of cDNA from each sample were used for library construction using the library preparation kit (Cat # 111KL114, STOmics America Ltd) and subsequently for DNB generation. Finally, the DNBs were sequenced on the DNBSEQTM T7 sequencing platform (Complete Genomics, San Jose, USA) with 50 bp read1 and 100 bp read2 (Cat # 100008555, Complete Genomics).
Tissue preparation, cryosection, and immunohistochemistry
Cryosections were cut at a thickness of 10 µm and mounted onto slides. The sections were fixed and permeabilized with pre-cooled acetone at −20 °C for 10 mins, and then rinsed three times with 1X PBS. Sections were placed under a vented hood for air drying prior to AT8 and Aβ42 staining. Phospho-Tau (Ser202, Thr205) Monoclonal Antibody (AT8) (Cat # MN1020, Invitrogen), beta Amyloid1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42 Polyclonal Antibody (Cat # 44-344, Invitrogen), and Tyramide SuperBoostTM Kit (Cat # B40915, Cat # B40962, and Cat # B40953, Invitrogen) were used for immunohistochemical staining according to the vendor’s instructions (Invitrogen).
Stereo-seq data processing
The fastq files from Stereo-seq were processed following the standard pipeline (https://github.com/STOmics/SAW). Initially, the first reads containing coordinate identity (CID) sequences underwent alignment to the designed coordinates of the Stereo-seq chip, based on the results from the first round of sequencing. This step allowed for a maximum of one base mismatch during the alignment process to account for potential sequencing errors. After alignment, reads exhibiting molecular identifiers (MIDs) possessing more than two bases with a quality score below 10 were excluded to ensure data integrity. The associated CID and MID for each qualified read were then incorporated into the read header. Retained reads were aligned to the reference genome using STAR119 and only reads achieving a mapping quality score above 10 were considered for gene annotation, ensuring the accuracy of gene expression profiling. The unique molecular identifiers (UMIs) sharing identical CIDs were combined into a single UMI. This step allowed for a single mismatch, facilitating the correction of sequencing and PCR errors. Finally, the CID-containing expression profile matrix was constructed.
Bioinformatics analysis of stereo-seq dataset
Stereo-seq data integration and dimension reduction
At the bin110, bin50, and single-nuclei resolution, all the pseudo-spots/nuclei across 12 samples were merged and the 2000 genes with the highest dispersion (variance/mean) were selected for principal component analysis (PCA). To remove potential batch effects, we have applied the Harmony algorithm23 to transform the top 50 PCs of the pseudo-spot/nuclei and project all pseudo-spot/nuclei into a shared embedding panel. UMAP27 was used to project the transformed PCs into a two-dimensional panel.
Clustering analysis
To identify the laminar architecture of human PFC at the bin110 resolution, the spatial constrained clustering algorithm was employed using the ‘tl.spatial_neighbors’ and the ‘tl.leiden’ functions in Stereopy package (v1.0.0)24 with the default parameter. At the bin50 and single-nucleus resolution, ‘sc.tl.leiden’ function in Scanpy120 was used with the default parameter for pseudo-spot and nuclei clustering. All the clusters of pseudo-spots/nuclei were annotated by their specific gene markers. To further confirm our annotation results for each cluster, we plotted the annotated pseudo-spots/nuclei on the two-dimensional panel based on their coordinate information for each sample, and mapped the annotated pseudo-spots/nuclei on the corresponding area of the H&E sections. The orientation of each sample was confirmed by identifying the border between grey matter and adjacent white matter (WM) using the H&E stained sections coupled with the gene distribution profiles within the section (Fig. S4A).
DGE analysis
The “sc.tl.rank_genes_groups” function from the Scanpy package (v1.9.3) was utilized for DGE analysis. A two-sided t-test with FDR adjustment was applied to calculate the adjusted P-values., and the genes with adjusted P-value ≤ 0.05, fold-change ≥ 0.25, and mean expression across all pseudo-spots/nuclei ≥ 0.25 were considered as the DEGs.
Layer-layer and cell-cell communication analysis
Layer-layer and cell-cell interactions based on the expression of known LR pairs in different layer/cell types were inferred using CellChat (v2.1.0)48. In this algorithm, the communication strength is computed by modelling ligand-receptor interactions for the Bin110 pseudo-spot in the NA, moderate, and severe AD groups, separately, using the Law of Mass Action, incorporating gene expression profiles projected onto a protein-protein interactions network. Further, this model has incorporated the cell proportion information to avoid the bias due to the unbalanced comparison of the cell-cell interactions. We followed the official workflow and applied the data processing functions ‘identifyOverExpressedGenes’, ‘identifyOverExpressedInteractions’. The layer and cellular communication networks were inferred by the function ‘computeCommuProb’. Function ‘netVisual_heatmap’ was used for the pairwise comparison of interactions, and the function ‘netVisual_bubble’ was used to compare the communication probabilities mediated by L-R from certain layer/cell group to other groups. All the analyses were performed with the default parameter setting.
Nucleus segmentation with nucleic acid staining
We followed the analysis pipeline from Stereopy24 to capture the spatial transcriptional profiles at single-nucleus resolution. Briefly, the nucleic acid staining image of the same section for the Stereo-seq library preparation was used to project the nuclei images on the transcriptional atlas. Deep cell model24 was used to segment the nuclei on the transcriptional atlas, and the UMI from all DNB within the corresponding segmented nuclei were aggregated per-gene and then summed to generate a nucleus by gene matrix for downstream analysis.
Concentric circles analysis
To understand how gene expression varies with proximity to the high stress area in PFC, we mapped the high stress focal point (identified at bin50 resolution) on a two-dimensional panel based on their coordinator for each sample, respectively, and drew three concentric circles around each high stress focal point to differentiate nuclei distances. We first selected nuclei located within 2500 pixels (1250 µm) from the high-stress focal point in each sample. Nuclei within a radius of 500 units of the pixel (approximately 250 µm) are categorized as level I, those between 500 and 1500 units (approximately 250-750 µm) as level II, and those in 1500 to 2500 units (approximately 750-1250 µm) as level III. Nuclei intersecting circles from multiple high stress points were assigned to the closest level (Fig. 4D).
Gene co-expression network analysis
We performed nuclei-type specific weighted gene co-expression network analysis (WGCNA) based on the nuclei from level I to III using the hdWGCNA package (v0.3.00)94. We first applied the MetacellsByGroups function with default parameter (“k = 25” and “max_shared = 10”) to convert our gene expression matrix into metacell expression matrix for reducing the data sparsity. After that, “SetDatExpr” function was applied to extract the cell type-specific gene expression matrix for each nuclei type. We then used the “TestSftPowers” function with default parameters to determine the appropriate soft-power threshold for reducing noise during the co-expression module construction processes. In our study, the soft-power threshold was determined to be seven for Ex and five for Inh. We then constructed the co-expression networks for Ex and Inh neurons using the “ConstructNetwork” function with “soft_power = 7” and “soft_power = 5”, respectively. To calculate module eigengenes and module connectivity, we employed the “ModuleEigengenes” and “ModuleConnectivity” functions, both with default parameters. We also used the “GetModuleTraitCorrelation” function with default parameters to assess the correlation between co-expression modules and AD levels (NA, moderate, and severe), as well as between the co-expression modules and the distance to stressed neurons (from level III to I). Finally, we utilized the “MofiScan” function with “EnsDb.Hsapiens.V86” as the reference and the “OverlapModulesMotifs” function to identify potential transcription factors regulating the modules.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The Stereo-seq data from six AD cases and six controls generated in this study have been deposited in GEO database (GSE269906). The raw FASTQ data have been deposited in the SRA database and are accessible for download through the accession number PRJNA1120963.
Code availability
The codes are available and can be downloaded from GitHub (https://github.com/Wonderangela123/AD-Stereo-Seq-Project.git) and Zenodo (https://zenodo.org/records/14048103).
Change history
06 May 2025
In the original version of this article, the given and family names of S. Michal Jazwinski were incorrectly structured. The name was displayed correctly in all versions at the time of publication. The original article has been corrected.
References
Knopman, D. S. et al. Alzheimer disease. Nat. Rev. Dis. Prim. 7, 33 (2021).
Zhang, W., Xiao, D., Mao, Q. & Xia, H. Role of neuroinflammation in neurodegeneration development. Signal Transduct. Target Ther. 8, 267 (2023).
Goel, P. et al. Neuronal cell death mechanisms in Alzheimer’s disease: An insight. Front Mol. Neurosci. 15, 937133 (2022).
Gustavsson, A. et al. Global estimates on the number of persons across the Alzheimer’s disease continuum. Alzheimers Dement 19, 658–670 (2023).
Hampel, H. et al. The Amyloid-beta Pathway in Alzheimer’s Disease. Mol. Psychiatry 26, 5481–5503 (2021).
Grundke-Iqbal, I. et al. Microtubule-associated protein tau. A component of Alzheimer paired helical filaments. J. Biol. Chem. 261, 6084–6089 (1986).
Busche, M. A. & Hyman, B. T. Synergy between amyloid-beta and tau in Alzheimer’s disease. Nat. Neurosci. 23, 1183–1193 (2020).
Crary, J. F. et al. Primary age-related tauopathy (PART): a common pathology associated with human aging. Acta Neuropathol. 128, 755–766 (2014).
Rodrigue, K. M. et al. beta-Amyloid burden in healthy aging: regional distribution and cognitive consequences. Neurology 78, 387–395 (2012).
Sziraki, A. et al. A global view of aging and Alzheimer’s pathogenesis-associated cell population dynamics and molecular signatures in human and mouse brains. Nat. Genet 55, 2104–2116 (2023).
Xia, X., Jiang, Q., McDermott, J. & Han, J. J. Aging and Alzheimer’s disease: Comparison and associations from molecular to system level. Aging Cell 17, e12802 (2018).
Liu, R. M. Aging, Cellular Senescence, and Alzheimer’s Disease. Int J. Mol. Sci. 23, 1989 (2022).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Chen, S. et al. Spatially resolved transcriptomics reveals genes associated with the vulnerability of middle temporal gyrus in Alzheimer’s disease. Acta Neuropathol. Commun. 10, 188 (2022).
Grubman, A. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat. Neurosci. 22, 2087–2097 (2019).
Chen, W. T. et al. Spatial Transcriptomics and In Situ Sequencing to Study Alzheimer’s Disease. Cell 182, 976–91.e19 (2020).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–92.e21 (2022).
Salat, D. H., Kaye, J. A. & Janowsky, J. S. Selective preservation and degeneration within the prefrontal cortex in aging and Alzheimer disease. Arch. Neurol. 58, 1403–1408 (2001).
Gouras, G. K. et al. Intraneuronal Abeta42 accumulation in human brain. Am. J. Pathol. 156, 15–20 (2000).
Guy, J. & Staiger, J. F. The Functioning of a Cortex without Layers. Front Neuroanat. 11, 54 (2017).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Huuki-Myers, L. A. et al. A data-driven single-cell and spatial transcriptomic map of the human prefrontal cortex. Science 384, eadh1938 (2024).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Fang S., et al. Stereopy: modeling comparative and spatiotemporal cellular heterogeneity via multi-sample spatial transcriptomics. bioRxiv. 2023:2023.12.04.569485.
Yang, L., Liu, J., Lu, Q., Riggs, A. D. & Wu, X. SAIC: an iterative clustering approach for analysis of single cell RNA-seq data. BMC Genomics 18, 689 (2017).
Lin, P., Troup, M. & Ho, J. W. CIDR: Ultrafast and accurate clustering through imputation for single-cell RNA-seq data. Genome Biol. 18, 59 (2017).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Phipson, B. et al. propeller: testing for differences in cell type proportions in single cell data. Bioinformatics 38, 4720–4726 (2022).
Scahill, R. I., Schott, J. M., Stevens, J. M., Rossor, M. N. & Fox, N. C. Mapping the evolution of regional atrophy in Alzheimer’s disease: unbiased analysis of fluid-registered serial MRI. Proc. Natl Acad. Sci. USA 99, 4703–4707 (2002).
Vogt, B. A., Van Hoesen, G. W. & Vogt, L. J. Laminar distribution of neuron degeneration in posterior cingulate cortex in Alzheimer’s disease. Acta Neuropathol. 80, 581–589 (1990).
Lawrence, J. M., Schardien, K., Wigdahl, B. & Nonnemacher, M. R. Roles of neuropathology-associated reactive astrocytes: a systematic review. Acta Neuropathol. Commun. 11, 42 (2023).
Nasrabady, S. E., Rizvi, B., Goldman, J. E. & Brickman, A. M. White matter changes in Alzheimer’s disease: a focus on myelin and oligodendrocytes. Acta Neuropathol. Commun. 6, 22 (2018).
Wang, W., Zhao, F., Ma, X., Perry, G. & Zhu, X. Mitochondria dysfunction in the pathogenesis of Alzheimer’s disease: recent advances. Mol. Neurodegener. 15, 30 (2020).
Suganuma, Y. et al. Quinonoid dihydropteridine reductase, a tetrahydrobiopterin-recycling enzyme, contributes to 5-hydroxytryptamine-associated platelet aggregation in mice. J. Pharm. Sci. 150, 173–179 (2022).
Fanet, H. et al. Tetrahydrobiopterin Improves Recognition Memory in the Triple-Transgenic Mouse Model of Alzheimer’s Disease, Without Altering Amyloid-beta and Tau Pathologies. J. Alzheimers Dis. 79, 709–727 (2021).
Al-Dalahmah, O. et al. Osteopontin drives neuroinflammation and cell loss in MAPT-N279K frontotemporal dementia patient neurons. Cell Stem Cell 31, 676–693.e10 (2024).
Krokowski, D. et al. Stress-induced perturbations in intracellular amino acids reprogram mRNA translation in osmoadaptation independently of the ISR. Cell Rep. 40, 111092 (2022).
Wang, N., Yang, W., Li, L. & Tian, M. MEF2D upregulation protects neurons from oxygen-glucose deprivation/re-oxygenation-induced injury by enhancing Nrf2 activation. Brain Res 1741, 146878 (2020).
Masood, M. I., Schafer, K. H., Naseem, M., Weyland, M. & Meiser, P. Troxerutin flavonoid has neuroprotective properties and increases neurite outgrowth and migration of neural stem cells from the subventricular zone. PLoS One 15, e0237025 (2020).
Leng, K. et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat. Neurosci. 24, 276–287 (2021).
Baird, L. & Yamamoto, M. The Molecular Mechanisms Regulating the KEAP1-NRF2 Pathway. Mol. Cell Biol. 40, 13 (2020).
Li, L., Venkataraman, L., Chen, S. & Fu, H. Function of WFS1 and WFS2 in the Central Nervous System: Implications for Wolfram Syndrome and Alzheimer’s disease. Neurosci. Biobehav Rev. 118, 775–783 (2020).
Xu, L. Z., Li, B. Q. & Jia, J. P. DAPK1: a Novel Pathology and Treatment Target for Alzheimer’s Disease. Mol. Neurobiol. 56, 2838–2844 (2019).
Wang, X., Lee, G., Liebhaber, S. A. & Cooke, N. E. Human cysteine-rich protein. A member of the LIM/double-finger family displaying coordinate serum induction with c-myc. J. Biol. Chem. 267, 9176–9184 (1992).
Parihar, M. S. & Brewer, G. J. Amyloid-beta as a modulator of synaptic plasticity. J. Alzheimers Dis. 22, 741–763 (2010).
Muralidhar, S., Wang, Y. & Markram, H. Synaptic and cellular organization of layer 1 of the developing rat somatosensory cortex. Front Neuroanat. 7, 52 (2013).
Larkum, M. E., Petro, L. S., Sachdev, R. N. S. & Muckli, L. A Perspective on Cortical Layering and Layer-Spanning Neuronal Elements. Front Neuroanat. 12, 56 (2018).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Scala, F. et al. Layer 4 of mouse neocortex differs in cell types and circuit organization between sensory areas. Nat. Commun. 10, 4174 (2019).
Kim, E. J., Juavinett, A. L., Kyubwa, E. M., Jacobs, M. W. & Callaway, E. M. Three Types of Cortical Layer 5 Neurons That Differ in Brain-wide Connectivity and Function. Neuron 88, 1253–1267 (2015).
Yu, M., Sporns, O. & Saykin, A. J. The human connectome in Alzheimer disease - relationship to biomarkers and genetics. Nat. Rev. Neurol. 17, 545–563 (2021).
Martinez-Lozada, Z. & Ortega, A. Glutamatergic Transmission: A Matter of Three. Neural Plast. 2015, 787396 (2015).
Sudhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008).
Moberg, S. & Takahashi, N. Neocortical layer 5 subclasses: From cellular properties to roles in behavior. Front Synaptic Neurosci. 14, 1006773 (2022).
Fiorentino, A., Sharp, S. I. & McQuillin, A. Association of rare variation in the glutamate receptor gene SLC1A2 with susceptibility to bipolar disorder and schizophrenia. Eur. J. Hum. Genet 23, 1200–1206 (2015).
Sonnewald, U. & Schousboe, A. Introduction to the Glutamate-Glutamine Cycle. Adv. Neurobiol. 13, 1–7 (2016).
Salpietro, V. et al. AMPA receptor GluA2 subunit defects are a cause of neurodevelopmental disorders. Nat. Commun. 10, 3094 (2019).
Swanson, C. J. et al. Metabotropic glutamate receptors as novel targets for anxiety and stress disorders. Nat. Rev. Drug Discov. 4, 131–144 (2005).
Kim, H. et al. Calsyntenin-3 interacts with both alpha- and beta-neurexins in the regulation of excitatory synaptic innervation in specific Schaffer collateral pathways. J. Biol. Chem. 295, 9244–9262 (2020).
Sheng, M., Sabatini, B. L. & Südhof, T. C. Synapses and Alzheimer's disease. Cold Spring Harb. Perspect. Biol. 4, a005777 (2012).
Castelli, V. et al. Neuronal Cells Rearrangement During Aging and Neurodegenerative Disease: Metabolism, Oxidative Stress and Organelles Dynamic. Front Mol. Neurosci. 12, 132 (2019).
Dehkordi, S. K. et al. Profiling senescent cells in human brains reveals neurons with CDKN2D/p19 and tau neuropathology. Nat. Aging 1, 1107–1116 (2021).
Alavi Naini, S. M. & Soussi-Yanicostas, N. Tau Hyperphosphorylation and Oxidative Stress, a Critical Vicious Circle in Neurodegenerative Tauopathies? Oxid. Med Cell Longev. 2015, 151979 (2015).
Marquez-Jurado, S. et al. Mitochondrial levels determine variability in cell death by modulating apoptotic gene expression. Nat. Commun. 9, 389 (2018).
Ries, M. & Sastre, M. Mechanisms of Abeta Clearance and Degradation by Glial Cells. Front Aging Neurosci. 8, 160 (2016).
Mrdjen, D. et al. The basis of cellular and regional vulnerability in Alzheimer’s disease. Acta Neuropathol. 138, 729–749 (2019).
Rackham, O. & Filipovska, A. Organization and expression of the mammalian mitochondrial genome. Nat. Rev. Genet 23, 606–623 (2022).
Tonnies, E. & Trushina, E. Oxidative Stress, Synaptic Dysfunction, and Alzheimer’s Disease. J. Alzheimers Dis. 57, 1105–1121 (2017).
Hol, E. M. et al. Neuronal expression of GFAP in patients with Alzheimer pathology and identification of novel GFAP splice forms. Mol. Psychiatry 8, 786–796 (2003).
Silva, I., Silva, J., Ferreira, R. & Trigo, D. Glymphatic system, AQP4, and their implications in Alzheimer’s disease. Neurol. Res Pr. 3, 5 (2021).
Chen, T., Gai, W. P. & Abbott, C. A. Dipeptidyl peptidase 10 (DPP10(789)): a voltage gated potassium channel associated protein is abnormally expressed in Alzheimer’s and other neurodegenerative diseases. Biomed. Res Int 2014, 209398 (2014).
Jauregui-Huerta, F. et al. Responses of glial cells to stress and glucocorticoids. Curr. Immunol. Rev. 6, 195–204 (2010).
Gns, H. S., Rajalekshmi, S. G. & Burri, R. R. Revelation of Pivotal Genes Pertinent to Alzheimer’s Pathogenesis: A Methodical Evaluation of 32 GEO Datasets. J. Mol. Neurosci. 72, 303–322 (2022).
Liu, L. et al. Microarray Analysis of Differential Gene Expression in Alzheimer’s Disease Identifies Potential Biomarkers with Diagnostic Value. Med Sci. Monit. 26, e919249 (2020).
Plagman, A. et al. Cholecystokinin and Alzheimer’s disease: a biomarker of metabolic function, neural integrity, and cognitive performance. Neurobiol. Aging 76, 201–207 (2019).
Emilsson, L., Saetre, P. & Jazin, E. Low mRNA levels of RGS4 splice variants in Alzheimer’s disease: association between a rare haplotype and decreased mRNA expression. Synapse 59, 173–176 (2006).
Ham, S. & Lee, S. V. Advances in transcriptome analysis of human brain aging. Exp. Mol. Med 52, 1787–1797 (2020).
Iatrou, A., Clark, E. M. & Wang, Y. Nuclear dynamics and stress responses in Alzheimer’s disease. Mol. Neurodegener. 16, 65 (2021).
Jin, M. & Cai, S. Q. Mechanisms Underlying Brain Aging Under Normal and Pathological Conditions. Neurosci. Bull. 39, 303–314 (2023).
Lago-Baldaia, I., Fernandes, V. M. & Ackerman, S. D. More Than Mortar: Glia as Architects of Nervous System Development and Disease. Front Cell Dev. Biol. 8, 611269 (2020).
Soliman, N., Okuse, K. & Rice, A. S. C. VGF: a biomarker and potential target for the treatment of neuropathic pain? Pain. Rep. 4, e786 (2019).
Schock, S. C. et al. Rescue of neurons from ischemic injury by peroxisome proliferator-activated receptor-gamma requires a novel essential cofactor LMO4. J. Neurosci. 28, 12433–12444 (2008).
Hurst, C. et al. Integrated Proteomics to Understand the Role of Neuritin (NRN1) as a Mediator of Cognitive Resilience to Alzheimer’s Disease. Mol. Cell Proteom. 22, 100542 (2023).
Campanella C. et al. Heat Shock Proteins in Alzheimer’s Disease: Role and Targeting. Int J Mol Sci. 19, (2018).
Carlo, A. S. et al. The pro-neurotrophin receptor sortilin is a major neuronal apolipoprotein E receptor for catabolism of amyloid-beta peptide in the brain. J. Neurosci. 33, 358–370 (2013).
Dong, L. X. et al. LncRNA NEAT1 promotes Alzheimer’s disease by down regulating micro-27a-3p. Am. J. Transl. Res 13, 8885–8896 (2021).
Kanai, Y. et al. The SLC1 high-affinity glutamate and neutral amino acid transporter family. Mol. Asp. Med 34, 108–120 (2013).
Sikora, J., Harzer, K. & Elleder, M. Neurolysosomal pathology in human prosaposin deficiency suggests essential neurotrophic function of prosaposin. Acta Neuropathol. 113, 163–175 (2007).
Chen, Y. & Yu, Y. Tau and neuroinflammation in Alzheimer’s disease: interplay mechanisms and clinical translation. J. Neuroinflammation 20, 165 (2023).
Zhang, C. et al. Comprehensive analyses of brain cell communications based on multiple scRNA-seq and snRNA-seq datasets for revealing novel mechanism in neurodegenerative diseases. CNS Neurosci. Ther. 29, 2775–2786 (2023).
Liu, A., Fernandes, B. S., Citu, C. & Zhao, Z. Unraveling the intercellular communication disruption and key pathways in Alzheimer’s disease: an integrative study of single-nucleus transcriptomes and genetic association. Alzheimers Res Ther. 16, 3 (2024).
Liu, B. et al. Glio- and neuro-protection by prosaposin is mediated by orphan G-protein coupled receptors GPR37L1 and GPR37. Glia 66, 2414–2426 (2018).
Matsuda, S., Matsuda, Y. & D’Adamio, L. CD74 interacts with APP and suppresses the production of Abeta. Mol. Neurodegener. 4, 41 (2009).
Morabito, S., Reese, F., Rahimzadeh, N., Miyoshi, E. & Swarup, V. hdWGCNA identifies co-expression networks in high-dimensional transcriptomics data. Cell Rep. Methods 3, 100498 (2023).
Guglielmotto, M. et al. The Decrease of Uch-L1 Activity Is a Common Mechanism Responsible for Abeta 42 Accumulation in Alzheimer’s and Vascular Disease. Front Aging Neurosci. 9, 320 (2017).
Liao, W. et al. Identification of candidate genes associated with clinical onset of Alzheimer’s disease. Front Neurosci. 16, 1060111 (2022).
Sze, C. I., Bi, H., Kleinschmidt-DeMasters, B. K., Filley, C. M. & Martin, L. J. Selective regional loss of exocytotic presynaptic vesicle proteins in Alzheimer’s disease brains. J. Neurol. Sci. 175, 81–90 (2000).
Carulla, P. et al. Involvement of PrP(C) in kainate-induced excitotoxicity in several mouse strains. Sci. Rep. 5, 11971 (2015).
Kellett, K. A. & Hooper, N. M. Prion protein and Alzheimer disease. Prion 3, 190–194 (2009).
Wu, E. et al. HSPA8 acts as an amyloidase to suppress necroptosis by inhibiting and reversing functional amyloid formation. Cell Res 33, 851–866 (2023).
Gao, R. & Penzes, P. Common mechanisms of excitatory and inhibitory imbalance in schizophrenia and autism spectrum disorders. Curr. Mol. Med 15, 146–167 (2015).
Kane, M. J. & Citron, B. A. Transcription factors as therapeutic targets in CNS disorders. Recent Pat. CNS Drug Discov. 4, 190–199 (2009).
Sun, H. et al. Quantitative integration of epigenomic variation and transcription factor binding using MAmotif toolkit identifies an important role of IRF2 as transcription activator at gene promoters. Cell Discov. 4, 38 (2018).
Fang, X., Zhong, X., Yu, G., Shao, S. & Yang, Q. Vascular protective effects of KLF2 on Abeta-induced toxicity: Implications for Alzheimer’s disease. Brain Res 1663, 174–183 (2017).
Li, L., Zi, X., Hou, D. & Tu, Q. Kruppel-like factor 4 regulates amyloid-beta (Abeta)-induced neuroinflammation in Alzheimer’s disease. Neurosci. Lett. 643, 131–137 (2017).
Wang, Y. et al. Kruppel-like factor 5 accelerates the pathogenesis of Alzheimer’s disease via BACE1-mediated APP processing. Alzheimers Res Ther. 14, 103 (2022).
Miyoshi, E. et al. Spatial and single-nucleus transcriptomic analysis of genetic and sporadic forms of Alzheimer’s Disease. Nat. Genet. 56, 2704–2717 (2024).
Pini, L. et al. Brain atrophy in Alzheimer’s Disease and aging. Ageing Res Rev. 30, 25–48 (2016).
Smit, T. et al. Reactive astrocytes as treatment targets in Alzheimer’s disease-Systematic review of studies using the APPswePS1dE9 mouse model. Glia 69, 1852–1881 (2021).
Lutz, M. W. & Chiba-Falek, O. Bioinformatics pipeline to guide late-onset Alzheimer’s disease (LOAD) post-GWAS studies: Prioritizing transcription regulatory variants within LOAD-associated regions. Alzheimers Dement (N. Y) 8, e12244 (2022).
Shibata, M. et al. Clearance of Alzheimer’s amyloid-ss(1-40) peptide from brain by LDL receptor-related protein-1 at the blood-brain barrier. J. Clin. Investig. 106, 1489–1499 (2000).
Li, Y. et al. Decreased CSF clearance and increased brain amyloid in Alzheimer’s disease. Fluids Barriers CNS 19, 21 (2022).
Alonso, A. D., Grundke-Iqbal, I., Barra, H. S. & Iqbal, K. Abnormal phosphorylation of tau and the mechanism of Alzheimer neurofibrillary degeneration: sequestration of microtubule-associated proteins 1 and 2 and the disassembly of microtubules by the abnormal tau. Proc. Natl Acad. Sci. USA 94, 298–303 (1997).
An, L. & Liu, Y. ZNF460 mediates epithelial-mesenchymal transition to promote gastric cancer progression by transactivating APOC1 expression. Exp. Cell Res 422, 113452 (2023).
Rouland, A. et al. Role of apolipoprotein C1 in lipoprotein metabolism, atherosclerosis and diabetes: a systematic review. Cardiovasc Diabetol. 21, 272 (2022).
Poduslo, S. E., Neal, M., Herring, K. & Shelly, J. The apolipoprotein CI A allele as a risk factor for Alzheimer’s disease. Neurochem Res 23, 361–367 (1998).
Braak, H., Alafuzoff, I., Arzberger, T., Kretzschmar, H. & Del Tredici, K. Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol. 112, 389–404 (2006).
Chen, A. et al. Single-cell spatial transcriptome reveals cell-type organization in the macaque cortex. Cell 186, 3726–43.e24 (2023).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Acknowledgements
The authors are grateful to STOmics Americas Ltd for technical assistance with the data generation. This work was supported by STOmics Grant Program and partially benefited from National Institutes of Health under grant U19AG055373 (to H.W.D. and H.S.), R01AG061917 (to H.W.D.), R01AG068232 (to H.W.D), P30GM145498 (to H.W.D.), P20GM109036 (to H.W.D.), P30AG072973 (to R.H.S.).
Author information
Authors and Affiliations
Contributions
Y.G. conducted the major data analysis and wrote the main manuscript text; M.H. and R.H.S. collected the human brain samples and associated clinical information; Xiao Z., Y.L., and Y.P.C. performed the biological experiments; M.H., R.H.S., H.S., H.D.W, S.M.J., K.Z., Xiang Z., X.W., A.L., L.J., D.W., Q.Z., and X.Y. provided valuable suggestions throughout the study implementation; H.S. and H.W.D. were responsible for conceiving, designing, initiating, directing, supervising, language proofreading, and securing fundings for this study. All authors participated in the discussions of the project and reviewed and/or revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gong, Y., Haeri, M., Zhang, X. et al. Stereo-seq of the prefrontal cortex in aging and Alzheimer’s disease. Nat Commun 16, 482 (2025). https://doi.org/10.1038/s41467-024-54715-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-024-54715-y
This article is cited by
-
Identification of Chlamydia pneumoniae and NLRP3 inflammasome activation in Alzheimer’s disease retina
Nature Communications (2026)
-
Microglia networks within the tapestry of alzheimer’s disease through spatial transcriptomics
Molecular Neurodegeneration (2025)
-
Advancements in the investigation of the mechanisms underlying cognitive aging
Biogerontology (2025)