Abstract
Three-dimensional subcellular imaging is essential for biomedical research, but the diffraction limit of optical microscopy compromises axial resolution, hindering accurate three-dimensional structural analysis. This challenge is particularly pronounced in label-free imaging of thick, heterogeneous tissues, where assumptions about data distribution (e.g. sparsity, label-specific distribution, and lateral-axial similarity) and system priors (e.g. independent and identically distributed noise and linear shift-invariant point-spread functions are often invalid. Here, we introduce SSAI-3D, a weakly physics-informed, domain-shift-resistant framework for robust isotropic three-dimensional imaging. SSAI-3D enables robust axial deblurring by generating a diverse, noise-resilient, sample-informed training dataset and sparsely fine-tuning a large pre-trained blind deblurring network. SSAI-3D is applied to label-free nonlinear imaging of living organoids, freshly excised human endometrium tissue, and mouse whisker pads, and further validated in publicly available ground-truth-paired experimental datasets of three-dimensional heterogeneous biological tissues with unknown blurring and noise across different microscopy systems.
Similar content being viewed by others
Introduction
Three-dimensional (3D) sub-cellular imaging is highly desirable in biomedical research, as it reveals the intricate spatial organization of organelles, cells, and biological networks. Achieving high-resolution sub-cellular imaging across all three dimensions is crucial for accurately understanding complex biological processes, such as neural circuits, disease pathogenesis, and cellular responses to drugs1,2,3,4,5,6. However, the fundamental diffraction limit of optical microscopy results in axial resolution that is 2–5 times worse than lateral resolution. This resolution disparity severely hinders isotropic 3D imaging, limiting our ability to faithfully represent the true 3D structure of biological samples and fully realize the potential of 3D imaging in scientific research7,8,9.
While recent advances in microscopy theory and instrumentation enable isotropic 3D imaging with certain specialized systems, their reliance on specific hardware, contrast mechanisms, and/or sample preparation limits widespread adoption, particularly for thick, living, and intact biosystems10,11,12,13,14,15,16,17. To make isotropic imaging more accessible across different microscopy modalities, algorithmic approaches seek to address the inverse problem of axial deblurring. In recent years, deconvolution methods based on deep neural networks (DNN), given their effective and flexible learning of data priors, have shown unprecedented performance compared to classical deconvolution methods, such as Richardson-Lucy and fast iterative shrinkage thresholding algorithm (FISTA), on two-dimensional (2D) deconvolution18,19,20,21,22. However, axial resolution restoration is significantly more challenging since acquiring isotropic 3D imaging data as ground truth is impractical for most microscopes, especially in living tissues, making supervised learning methods unsuitable for general isotropic 3D imaging. Despite the existence of a few systems capable of collecting paired isotropic 3D imaging data and generously sharing it14,23, the potential for pre-training a network and generalizing to diverse microscopes and samples is hampered by the significant domain shifts in biomedical imaging24,25.
Domain shift, the difference in statistical distributions between training and inference datasets, can cause a model trained on one dataset to perform poorly on another26,27,28. This issue is especially pronounced in scientific imaging compared to general computer vision due to the high heterogeneity of microscopy modalities, system properties, sample variation across species and tissues, and the exploratory nature (unseen anomaly detection) of microscopy data25,29. To address these challenges and enable general isotropic 3D imaging, unsupervised generative adversarial network (GAN)-based methods have been proposed to enable deblurring without matched axial image pairs30,31,32. However, the inherent instability of saddle-point optimization in GAN-based methods leaves them vulnerable to network collapse, potentially generating hallucinations and inconsistencies33,34,35. Supervised methods leverage similarities between lateral and axial data distributions and employ explicit blurring forward models to achieve high-fidelity isotropic imaging without paired in-distribution data36,37. However, the practical applications of single-stack isotropic resolution recovery using supervised methods are limited for two primary reasons. First, these methods rely on an explicit point-spread function (PSF) model that is assumed to be precisely matched and consistent across the entire imaging volume. In reality, factors such as field curvature, misalignment, and tissue scattering contribute to optical aberrations, resulting in spatially varying PSFs that are sample- and system-dependent, making real-time calibration difficult. This modeling challenge is further exacerbated by noise, especially in low-light conditions38. Second, self-supervised methods require assumptions of lateral-axial similarity within single-stack data to learn axial deblurring from synthetic lateral deblurring. While this assumption holds for certain cell and tissue structures (e.g., neurons in the cortex and sinusoids in the liver, as shown in prior axial deblurring works31,32,36), it breaks down in tissues with high directionality, polarity, or anisotropy, which are common in biological systems such as developing embryos, epithelial glands, and collagen fibers in the extracellular matrix29.
These challenges are particularly pronounced in label-free imaging of thick, heterogeneous tissues39, where assumptions about data distribution (e.g. sparsity, label-specific distribution, and lateral-axial similarity) and system priors (e.g. independent and identically distributed (i.i.d.) noise, and linear shift-invariant (LSI) PSFs) are often invalid. To overcome these challenges, we propose SSAI-3D (System- and Sample-agnostic Isotropic 3D Microscopy), a weakly physics-informed, domain-shift-resistant axial deblurring framework for robust isotropic resolution recovery across diverse systems and samples. First, we formulate isotropic resolution recovery as a semi-blind deblurring problem and leverage a pre-trained blind deblurring network developed for natural scene images as the initial network. Then, we create a synthetic training dataset by blurring denoised lateral images with various PSFs of different sizes and orientations to adapt the deblurring network to the current 3D imaging stack. This allows us to train the network in a self-supervised manner, eliminating the need for pixel-wise co-registered data pairs. We demonstrate that the combination of a denoised and PSF-varying training dataset, coupled with leveraging existing knowledge from the experimentally pre-trained blind deblurring network, significantly alleviates susceptibility to spatially varying 3D blurring kernels and noise in incoherent real-valued PSF imaging.
Secondly, we sparsely fine-tune the large pre-trained deblurring network using this synthetic dataset. A surgeon network is employed to identify which layers are most critical for adapting to microscopic images, while other layers remain frozen to preserve the original deblurring capabilities learned from the much larger dataset of natural scene image pairs. This approach leverages prior knowledge of blind deblurring while efficiently adapting to microscopy data and minimizing assumptions about lateral-axial similarity, thus reducing overfitting and computational costs commonly associated with microscopy deconvolution problems. We demonstrate high-fidelity single-stack axial deblurring performance of SSAI-3D on open-source experimental datasets from various microscopy systems: confocal (with near-isotropic validation through hardware modification23), light-sheet (with near-isotropic validation through hardware modification14), wide-field, and nonlinear microscope39,40. The high-fidelity recovery across diverse microscopy systems (varying in aberrations, noise, and contrasts) and samples (with different degrees of lateral-axial dissimilarity) indicates that SSAI-3D can be a widely accessible and reliable tool for isotropic imaging, facilitating precise analysis of complex 3D biological structures and processes.
Results
Isotropic resolution recovery by self-supervised sparsely fine-tuned deblurring
The SSAI-3D framework for single-stack isotropic resolution recovery consists of two steps (Fig. 1c). First, a self-supervised training data is generated by applying a series of PSFs with varying sizes and orientations to denoised lateral images (see Methods and Supplementary Fig. S1 for details). Unlike methods that rely on an exact PSF model and thus have strong physics-based assumptions about the image generation forward model, these sets of weakly-physics-based synthetic pairs emphasize the general blurring action of the optical microscope rather than the precise forward model, thereby increasing the network’s robustness against unknown optical aberrations and noise.
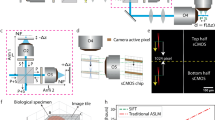
a SSAI-3D enables robust isotropic resolution recovery across diverse 3D imaging systems (confocal, light-sheet, wide-field, and nonlinear) and diverse 3D biological samples (organelles, cells, tissues, and organs). Created in BioRender. Liu, K. (2024) BioRender.com/f26y695. b The deblurring network is initialized with a large pre-trained network for blind deconvolution on a large dataset of natural image pairs. Example images were sourced from the REDS dataset52, released under a CC BY 4.0 license, which permits commercial use with proper attribution. c Starting with a single microscopy-specific image stack where the axial resolution is worse than the lateral resolution, lateral images are blurred with a series of PSFs of different sizes and orientations to generate a self-supervised dataset. Then, generating zero-shot metrics using ~1% of this dataset, a surgeon network is employed to select the critical layers to fine-tune in the large pre-trained deblurring network. Only ~10% layers are selected and sparsely fine-tuned according to the generated self-supervised dataset. Given unseen axial images, the fine-tuned deblurring network predicts deblurred images with isotropic resolution. Insets in the images represent the corresponding Fourier spectrums.
Secondly, we leverage knowledge learned from natural scene deblurring tasks via a large pre-trained deblurring network NAFNet41,42 (Fig. 1b, see Methods and Supplementary Fig. S2a for details of NAFNet). We then sparsely fine-tune the pre-trained NAFNet using the created self-supervised dataset, selecting and fine-tuning only ~10% of the layers while freezing the rest. Layer selection is guided by a surgeon network, which evaluates each layer’s contribution to the adaptation task using zero-shot metrics, including statistics related to the activation, gradient, loss function, and pre-trained weights (see Methods for details and Supplementary Fig. S2b for architecture). This surgeon network mitigates the stochastic instability of zero-shot metrics, while sparse fine-tuning avoids performance degradation in out-of-distribution data and the computational burden of full fine-tuning43. Our approach preserves the deblurring capability learned from the larger natural scene dataset, making only necessary adaptations to the microscopy image stack. This results in a computationally and data-efficient process that is less susceptible to overfitting to the lateral data distribution or potential mismatches in the forward model. Finally, with these two steps, in the inference stage, the axial images are sent to the adapted customized deblurring network that predicts deblurred images with isotropic resolution. Without strong assumptions about the imaging system or sample data distribution, SSAI-3D is anticipated to be a widely accessible and robust tool for isotropic 3D imaging across diverse microscopy modalities and a broad spectrum of biological applications (Fig. 1a).
We first compared SSAI-3D to the state-of-the-art unsupervised (SelfNet32) and supervised (CARE36) methods using two distinct synthetic objects, assuming a known and LSI PSF across the imaging volume. The first synthetic object comprised randomly seeded sub-diffraction-limited fluorescent beads within a imaging volume (Fig. 2a, see Methods for simulation data generation). All three methods demonstrated significant improvement in axial resolution (Fig. 2c), although Self-Net exhibited greater variability in bead shape and intensity compared to CARE and SSAI-3D, which achieved reliable axial resolution recovery (201.4 ± 2.7 nm and 200.7 ± 2.9 nm, respectively; n = 50 beads).
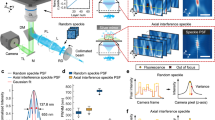
Simulation of bead (a) and strand structures (b) reveals the performance of SSAI-3D and existing methods (c, d). The simulation objects are visualized using depth-coded projections with the colorbar representing the actual depth. Reconstructed axial resolution (mean ± standard deviation, n = 50 beads) is labeled in c. e, f Characterization of reconstruction fidelity for beads (a) and strands (b). Box bounds represent the upper and lower quartiles, lines within boxes indicate medians, and whiskers extend to data points within 1.5 times the interquartile range (IQR), with outliers plotted individually beyond this range (n = 100 slices). g, h Ablation study on the performance of different levels of blurring for beads (a) and strands (b). i, j Robustness of reconstruction against PSF mismatch for beads (a) and strands (b). A blurring with a standard deviation of 5 is assumed to be known for CARE. k Comparison of the training time. Scale bars: 6 µm (a, c); 60 µm (b, d). Source data are provided as a Source Data file.
However, many biological structures do not exhibit perfect lateral-axial similarity like round beads. Therefore, we conducted a second simulation on a more heterogeneous 3D synthetic structure, comprising directional strands within an imaging volume (Fig. 2b, see Methods for simulation data generation). This denser object with directionality better simulates the complex 3D structures found in real biological samples, such as dendrites, glands, and fibers (neural, elastin, and collagen). In contrast to the bead simulation, existing methods exhibited significant performance degradation, while SSAI-3D maintained high-fidelity resolution recovery with accurate shape and intensity preservation (Fig. 2d and Supplementary Fig. S4). Quantitative analysis further confirmed that, even with a known and LSI PSF, existing algorithms significantly deteriorate in performance when applied to directional, 3D asymmetric structures, as a result of their reliance on the assumption of lateral-axial similarity. In contrast, SSAI-3D produced high-fidelity restorations with low mean square error (MSE) (Fig. 2e, f) and high structural similarity (SSIM) (Supplementary Fig. S5) across both simulations. Ablation studies confirmed the consistent axial deblurring fidelity of SSAI-3D across varying levels of blurring (Fig. 2g, h) and its robustness to PSF mismatch (Fig. 2i, j; see Methods for details). We also conducted experiments on simulating axial deblurring using non-Gaussian theoretical optical PSFs. These results demonstrate that SSAI-3D is capable of semi-blind deblurring and can handle non-Gaussian blur in real optical imaging. Additionally, we conducted experiments on simulated axial deblurring using non-Gaussian theoretical optical PSFs in a more realistic scenario, where the actual PSF is unknown during training and may have a very different shape. Results further demonstrated that SSAI-3D is capable of semi-blind deblurring and can effectively handle non-Gaussian blur in real optical imaging (Supplementary Fig. S6). Ablation studies further demonstrated that the dataset with multiple PSFs exhibiting variation is necessary to effectively address the challenging scenario of unknown and unmatched PSFs (Supplementary Fig. S7). Benefiting from sparse fine-tuning with only ~15 million parameters modified, SSAI-3D exhibited a 2.5–3.5-fold reduction in training time compared to existing methods, thereby enhancing its accessibility and reducing computational requirements (Fig. 2k).
Sparse fine-tuning on large pre-trained model relaxes assumptions on lateral-axial similarity of 3D data
To understand the performance differences between the two data types (beads vs. strands), we further investigated how lateral-axial data similarity influences algorithm performance. We quantified this relationship by plotting the accuracy on the reference lateral distribution against the accuracy on the potentially shifted axial distribution (n = 9 independent training and inference)44. In cases of high lateral-axial similarity, existing methods and SSAI-3D aligned with the identity line (y = x, Fig. 3b), indicating successful deblurring when the distribution shift in the test axial datasets is negligible. However, when lateral-axial similarity is low within the sample, CARE and SSAI-3D with full fine-tuning exhibited a significant gap between training MSE on lateral images and inference MSE on axial images. In contrast, SSAI-3D with sparse fine-tuning demonstrated robustness to distribution shifts, resulting in high-fidelity axial deblurring even in 3D heterogeneous samples45,46,47 (Fig. 3e; see Methods and Supplementary Figs. S8 and S9 for details and performance of pre-trained and fully fine-tuned network).
Simulation of sphere (high lateral-axial similarity, a) and cylinder (low lateral-axial similarity, d) reveals the performance of SSAI-3D and existing methods (c, f). b, e Relationship between training MSE on lateral images and inference MSE on axial images indicates robustness of different methods against distribution shifts. In label-free nonlinear imaging, biological tissues exhibit different levels of lateral-axial similarity (high in living and intact human blood-brain barrier microfluidic model (g) and low in freshly excised human endometrial tissue (i)). h, j Raw lateral and axial images of g and i, as well as restored axial images using SSAI-3D. Arrows: symmetric endothelial cells (h) and polarized epithelial glands (j). Scale bars: 50 µm. Source data are provided as a Source Data file.
In this experiment, we considered a simulation of imaging spheres and cylinders in 3D, representing cases of high (circles in both lateral and axial planes, Fig. 3a) and low (circles in lateral, rectangles in axial planes, Fig. 3d) lateral-axial similarity, respectively. Consistent with the results in Fig. 2, Self-Net, CARE, and SSAI-3D achieved comparable reconstructions under high lateral-axial similarity (Fig. 3c). However, in the low similarity case, performance diverged significantly, with SSAI-3D outperforming existing methods (Fig. 3f). This divergence arises because existing methods heavily rely on lateral-axial similarity, making them susceptible to significant performance decreases when distribution shifts occur in real axial test data. In contrast, the sparse finetuning in SSAI-3D preserves deblurring capabilities learned from the natural scene dataset while adapting minimally to the microscopy image stack, resulting in superior axial deblurring performance for 3D heterogeneous samples under lateral-axial distribution shifts.
In real-world biological imaging, lateral-axial similarity can vary significantly depending on the sample’s structure and imaging scale. For instance, neurons within a small volume may exhibit high lateral-axial similarity, whereas glands, dendrites, and fibers (neural, elastin, collagen) with high directionality, polarity, or anisotropy typically exhibit low lateral-axial similarity. We demonstrated this sample-dependent 3D heterogeneity using two thick and living samples imaged with label-free nonlinear imaging39. First, we imaged a living human blood-brain barrier microfluidic model where vascularized endothelial cells formed lumen structures with pericytes and astrocytes48. This 3D self-assembled multicellular model displayed high lateral-axial similarity (Fig. 3g). Secondly, we imaged a freshly excised human endometrial tissue, where directional collagen growth and the inherent polarity of the epithelial glands resulted in low lateral-axial similarity (Fig. 3i). SSAI-3D successfully restored fine structures such as endothelial cells (Fig. 3h; see Supplementary Movie 1 for the entire volume) and epithelial glands (Fig. 3j) in axial views, even in cases of low lateral-axial similarity. This robustness across samples with varying structural properties highlights the potential of SSAI-3D as a widely applicable tool for 3D biological imaging.
Generalizability to varying system imperfections and sample heterogeneity in simulation and experiments
Next, we investigate the robustness of SSAI-3D against various system imperfections compared to existing methods. In practical 3D biomedical imaging, imaging systems exhibit diverse PSFs in shape and size due to varying signal generation mechanisms and customized instrumentation and acquisition parameters. Furthermore, PSFs can be spatially varying across the imaging volume due to factors such as aberration, misalignment, and tissue scattering (Fig. 4a). Quantifying the precise PSFs for an arbitrary image stack is infeasible since they are system- and sample-dependent. Additionally, many imaging systems operate under photon-limited conditions due to biophysical and biochemical constraints, such as the need for high imaging speed or to minimize photobleaching, phototoxicity, and tissue heating49. The resulting low signal-to-noise ratio (SNR) images can significantly affect the accuracy of deconvolution methods38 (Fig. 4b). Given these challenges, we demonstrated the robustness of SSAI-3D against a spectrum of system imperfections, including spatially varying PSFs and noise, by leveraging a self-supervised dataset generated from PSFs with diverse sizes and orientations.
a Example of optical aberrations in real microscopy system (spatially-varying PSF in mesoSPIM23). b Example of noise in real microscopy system (low-light condition in dSLAM53). c Ground truth image for simulation on effect of optical aberrations and noise. Four different levels of optical aberrations and noise with the corresponding raw images (d Shift-invariant PSF (ideal case); e Shift-variant PSF; f Shift-variant PSF with rotations along z-direction; g Shift-variant PSF with rotations along z-direction and noise across the entire imaging volume). h–k Comparison of Self-Net, CARE, and SSAI-3D on deblurring performance using SSIM in scenarios d–g. Bottom: Blow-ups of the co-registered images within the white box. Scale bars: 2 mm (a); 25 µm (b–k).
We simulated randomly oriented tubular objects (high lateral-axial similarity) within an imaging volume (Fig. 4c, see Methods for simulation data generation). To evaluate axial deblurring performance, we initially applied a shift-invariant axial blur, assumed to be known for CARE (Fig. 4d). In this ideal scenario, Self-Net, CARE, and SSAI-3D all achieved high-quality resolution recovery with low MSE (Fig. 4h). However, in practice, obtaining an accurate PSF profile for the system is often challenging, and PSF mismatch can degrade the performance of CARE (Fig. 2i, j). Next, we progressively introduced system imperfections. Figure 4e, f incorporated variations in the extent and orientation of the axial blur. The non-uniform PSF profiles, combined with asymmetry between lateral and axial aberrations, led to decreased accuracy for Self-Net and CARE, particularly in regions with pronounced aberrations (Fig. 4i, j). To incorporate the effect of noise, we introduced both Gaussian and Poisson noise to the images after artificial axial blurring (Fig. 4g). To mitigate noise corruption and amplification during the forward blurring and inverse deblurring processes, SSAI-3D incorporates a denoising step prior to self-supervised dataset generation and the deblurring model, enhancing its robustness against noise in low-light conditions (Fig. 4k and Supplementary Fig. S10). Simulation across a gradient of noise levels consistently demonstrated superior performance for SSAI-3D, further confirming the applicability of SSAI-3D when handling noisy data (Supplementary Fig. S11).
Next, we investigate the performance of SSAI-3D on real-life biological data across various microscopy modalities, system properties, and sample types. As demonstrated in Fig. 5, SSAI-3D achieved consistent isotropic resolution recovery across a broad range of samples (living/fixed, human/animal, varying degrees of lateral-axial similarity) and imaging systems (diverse contrast mechanisms, commercial/custom-built, low/high resolution, low/high SNR). 3D images with varying degrees of anisotropy, acquired using light-sheet, confocal, wide-field, and nonlinear microscopy, all exhibited substantial axial resolution enhancement after restoration with SSAI-3D within 0.5 GPU hours of training (see Supplementary Table S1 and Methods for details). Fourier spectrum analysis quantified this improvement, revealing previously indistinguishable fine structures in low-resolution raw images, such as dendrites in mouse brain neurons (Fig. 5c) and nucleoli in human endometrial epithelial cells (Fig. 5f). Notably, the surgeon network selected different layers for fine-tuning in each imaging stack (Supplementary Fig. S12). These results highlight the potential of SSAI-3D as a generalizable and accessible tool for 3D imaging, enhancing axial resolution to facilitate the study of 3D tissue architecture and subcellular features.
Each example includes the raw and restored images (left), along with their respective blow-ups (highlighted in the white boxes, middle) and Fourier spectrums (right, labeled numbers represent the lateral and axial resolutions). a Cleared and stained mouse brain vasculature from light-sheet microscopy. b Freshly excised human endometrial tissue from multiphoton autofluorescence microscopy. c Cleared Thy1-GFP mouse brain neurons from confocal microscopy. d Fixed mouse whisker follicles from second harmonic generation (SHG) microscopy. e Cleared mouse liver from wide-field microscopy. f Freshly excised human endometrial tissue from third harmonic generation (THG) microscopy. Scale bars: 20 µm.
Validation of SSAI-3D with hardware-corrected isotropic imaging
To experimentally validate the high-fidelity axial deblurring of SSAI-3D and demonstrate its robustness to high sample heterogeneity and unknown system imperfections for downstream biological analysis, we presented two comparative studies against near-isotropic imaging references obtained via hardware advancements. First, we assessed SSAI-3D’s isotropic resolution recovery in light-sheet microscopy using co-registered raw images and a near-isotropic reference acquired by mesoscale selective plane-illumination microscopy (mesoSPIM)23 (Fig. 6a), which is a light-sheet microscope that achieves near-isotropic imaging by incorporating an axially scanned light-sheet (ASLM) module50. A whole mouse brain expressing tdTomato in vasoactive intestinal peptide (VIP) neurons was cleared and imaged with mesoSPIM51. At the mesoscale, the mouse brain exhibited high lateral-axial dissimilarity, and system aberrations varied significantly across the large field of view (FOV) (Fig. 4a). By comparing against the near-isotropic hardware-corrected images, SSAI-3D demonstrated high-fidelity 3D isotropic resolution recovery despite the high lateral-axial dissimilarity and the spatially varying PSFs, which degraded the performance of existing methods (please refer to Supplementary Fig. S13 for method comparison). To evaluate the effect of SSAI-3D on downstream biological analysis, we quantified the accuracy of neuron extraction and observed the segmentation accuracy of SSAI-3D closely matched that of the near-isotropic hardware-corrected mesoSPIM dataset (Fig. 6b; see Methods for detection algorithms).
a Comparison of software-corrected (SSAI-3D) and hardware-corrected (mesoSPIM) axial image of mouse brain from light-sheet microscopy. b Neuron detection accuracy using raw, software-corrected, and hardware-corrected images. Arrows in a and b are pointing to a same neuron. c Comparison of software-corrected (SSAI-3D) and hardware-corrected (multiview confocal) axial image of mitochondria from confocal microscopy. d Statistics of the volume of DNA puncta in mitochondria. Inset: overall statistics of raw data. Arrows in c represent the same DNA puncta with the volume marked in d. Scale bars: 1 mm (a, b); 1 µm (c, d). Source data are provided as a Source Data file.
To further evaluate generalizability across diverse system imperfections and sample geometries, we assessed SSAI-3D on a multi-modal confocal microscopy dataset, in addition to the light-sheet microscopy data. This dataset comprised co-registered raw images and a near-isotropic reference acquired by multiview confocal super-resolution microscopy14 (Fig. 6c). An expanded mitochondrion was imaged, with its outer membrane and DNA immunolabeled by two distinct dyes. Notably, axial and lateral data distributions, as well as those between channels, exhibited significant dissimilarity. Applying SSAI-3D to each channel individually with customized sparse fine-tuning, we found that only SSAI-3D achieved high-fidelity restoration compared to existing methods (please refer to Supplementary Fig. S14 for method comparison). Analysis of the detected DNA volume within the image stack revealed that volume statistics from the SSAI-3D reconstruction were highly consistent with those derived from the near-isotropic reference (Fig. 6d).
Discussion
SSAI-3D demonstrated robust isotropic resolution recovery across diverse real-world microscopy systems and samples (Fig. 5). In particular, this approach was applied to label-free nonlinear microscopy of intact, thick, and heterogeneous tissues, where assumptions about data distribution (e.g., sparsity, label-specific distribution, axial-lateral similarity) and system priors (e.g., i.i.d. noise, LSI PSFs) often fail. The axial deblurring pipeline was further validated using publicly available, ground-truth-paired experimental datasets featuring potentially high lateral-axial dissimilarity in biological tissues, as well as unknown blurring and noise characteristics across different microscopy systems (Fig. 6). SSAI-3D achieved this robust axial deblurring by generating a PSF-varying, noise-resilient, sample-informed training dataset and sparsely fine-tuning a large pre-trained blind deblurring network. Compared to existing computational methods, SSAI-3D does not require strong assumptions about the imaging system or sample data distribution, while maintaining low computational cost. Complementary to hardware-corrected isotropic imaging, SSAI-3D avoids the need for additional complex instrumentation and sample pre-processing, thus enabling broader applicability across biomedical applications, particularly for thick, intact, and living biosystems.
Compared to CARE36, the most widely used approach for isotropic resolution recovery, which relies on an ideal, LSI PSF across the imaging volume, SSAI-3D incorporates PSFs of varying sizes and orientations. This enhances reconstruction fidelity (Fig. 2e–h), robustness to PSF mismatch (Fig. 2i, j, Supplementary Fig. S6), and applicability to practical optical aberrations (Fig. 4h–j). Additionally, SSAI-3D pre-processes lateral data with a denoising step before self-supervised dataset generation, further improving robustness against noise (Fig. 4k), a step not included in CARE. Furthermore, while previous self-supervised axial deblurring methods like CARE assume lateral-axial similarity, SSAI-3D significantly reduces reliance on this assumption by employing a large pre-trained network (~150 million parameters41) leveraged from a natural scene dataset (~50,000 images52) with sparse finetuning. This approach demonstrates improved robustness against distribution shifts (low lateral-axial similarity, Fig. 3), effectively overcoming the limitations of training a small model (~1.5 million parameters) from scratch, as in CARE.
Imaging restoration has achieved remarkable progress in nature scene images by growing training datasets and larger neural networks. This work intends to build on these pre-trained models, such as NAFNet41,42, as powerful foundations for data-driven 3D microscopy restoration, which has been plagued by the challenge of curating large annotated datasets. However, the direct application of large pre-trained models to microscopy imaging using conventional fine-tuning is challenging due to the distribution shift from photography to microscopy, as well as the small or non-existing annotated microscopy data library. SSAI-3D, provides a framework for gentle customization of a blind photography deblurring network to a 3D microscopy restoration task with diverse data distributions and zero paired training data. Moreover, as a stand-alone module, sparse fine-tuning can be applied to other tasks where a large pre-trained model can be leveraged, including denoising, segmentation, and super-resolution, offering potential for broader applications within the microscopy community.
By utilizing a single image stack and without requiring precise knowledge of the system or sample, SSAI-3D provides computationally efficient access to high-fidelity isotropic resolution recovery for a wide range of microscopy applications. To facilitate broader adoption within the microscopy community, we offer a streamlined workflow (Supplementary Fig. S3) and have made our code and pre-trained surgeon network publicly available, along with detailed instructions and examples. We anticipate SSAI-3D will find widespread use in biomedical research, enabling new discoveries in fields such as neuroscience, developmental biology, and cancer research, where high-resolution 3D imaging is crucial.
Despite the variation in PSFs across different microscopy techniques, SSAI-3D has demonstrated robustness across a wide array of imaging modalities, effectively treating all microscopes as low-pass filters, regardless of the exact PSF shape (Supplementary Fig. S6, Figs. 5 and 6). This robustness is attributed to the deblurring features learned from the large pre-trained model and refined with a customized self-supervised dataset. Despite the robustness, the performance of SSAI-3D will likely degrade in these scenarios. First, if the original axial resolution is more than 7 times worse than the lateral resolution (roughly 2.5 µm lateral resolution assuming a Gaussian beam), the recovery fidelity deteriorates gradually, as the ablation experiment shown in Fig. 2i, j. Secondly, the denoising step has been critical for high-fidelity axial deblurring, likely similar to the noise amplification in the classical deconvolution problem. The use of SSAI-3D should be cautionary when it comes to highly noisy datasets, where either more acquisition time or a more effective denoising algorithm could help. Thirdly, with complex-valued coherent PSFs like OCT imaging, SSAI-3D may not achieve optimal axial deblurring without incorporating phase modeling into the self-supervised dataset generation. This could potentially be addressed by including more PSFs with varying phases to better model the forward image formation process.
Methods
Self-supervised dataset generation
SSAI-3D employed a self-supervision strategy for learning isotropic restoration using the raw 3D anisotropic data itself. High-resolution lateral images are first denoised using our previous denoising framework53. These denoised images are then artificially blurred by convolving with a set of 25 PSFs, combining five different sizes and five different orientations. To increase robustness to varying PSF sizes, Gaussian blur is applied to lateral images with kernel standard deviations ranging from 3 to 7 pixels. To increase robustness to varying orientations, the kernels are rotated within the range of -45° to 45°. This synthetic dataset, comprising the paired blurred images and their corresponding sharp originals, serves as the training data for the subsequent deep learning-based axial deblurring process. This self-supervised dataset generation process is visualized in Supplementary Fig. S1.
Contribution of different layers in fine-tuning
To determine which layers to fine-tune for adapting to the data distribution of a customized dataset, we assessed each layer’s contribution to the performance of the network using zero-shot metrics. In this paper, we employed the following 14 such metrics, obtainable via a single forward and backward pass of the pre-trained network, to predict the performance of fine-tuning each specific layer.
We denote the weights of the pre-trained network as \({{{\bf{\Theta }}}}\) and the loss function as \({{{\mathcal{L}}}}\). For layer \(l\), we denote the single pass forward activation and the single pass backward gradient at this layer as \({{{{\bf{A}}}}}^{(l)}\) and \({{{{\bf{G}}}}}^{(l)}\), respectively. Then, the 14 zero-shot metrics \({{{{\bf{U}}}}}^{(l)}=[{u}_{1}^{(l)},\ldots,{u}_{14}^{(l)}]\) of layer \(l\) can be represented as follows.
-
Metrics evaluating forward activation. These metrics aim to capture the compatibility of pre-trained parameters with the customized dataset. The metrics relate to activation values during the forward pass and include the average, standard deviation, average of absolute values, and standard deviation of absolute activation values:
$${u}_{1}^{(l)}{\mathbb{=}}{\mathbb{E}}\left\{{{{{\bf{A}}}}}^{(l)}\right\},{u}_{2}^{(l)}={{{\rm{std}}}}\left\{{{{{\bf{A}}}}}^{(l)}\right\},{u}_{3}^{(l)}{\mathbb{=}}{\mathbb{E}}\left\{\left|{{{{\bf{A}}}}}^{\left(l\right)}\right|\right\},{u}_{4}^{(l)}={{{\rm{std}}}}\left\{\left|{{{{\bf{A}}}}}^{\left(l\right)}\right|\right\}$$(1) -
Metrics evaluating backward gradient. These metrics aim to determine which pre-trained model parameters are most in need of adjustment. The metrics relate to the gradient of the model and include the average, standard deviation, average of absolute values, and standard deviation of absolute gradient values:
$${u}_{5}^{(l)}{\mathbb{=}}{\mathbb{E}}\left\{{{{{\bf{G}}}}}^{(l)}\right\},{u}_{6}^{(l)}={{{\rm{std}}}}\left\{{{{{\bf{G}}}}}^{(l)}\right\},{u}_{7}^{(l)}{\mathbb{=}}{\mathbb{E}}\left\{\left|{{{{\bf{G}}}}}^{\left(l\right)}\right|\right\},{u}_{8}^{(l)}={{{\rm{std}}}}\left\{\left|{{{{\bf{G}}}}}^{\left(l\right)}\right|\right\}$$(2)To evaluate the importance of each layer based on channel saliency, we also employed the fisher metric54. This metric assesses the impact of removing activation channels (and their corresponding parameters) that are estimated to have the least effect on the loss:
$${u}_{9}^{(l)}={\left({{{{\bf{A}}}}}^{\left(l\right)}\odot {{{{\bf{G}}}}}^{\left(l\right)}\right)}^{2}$$(3) -
Metrics related to the loss function. To evaluate the importance of each layer based on their sensitivity to the loss with respect to the network inputs, we employed the metric snip55 as
$${u}_{10}^{(l)}=\left|\frac{\partial {{{\mathcal{L}}}}}{\partial {{{{\bf{\Theta }}}}}^{(l)}}\odot {{{\bf{\Theta }}}}^{(l)}\right|$$(4)To evaluate the importance of each layer based on their sensitivity to the gradient (as opposed to loss in snip) with respect to the network inputs, we employed the metric grasp56 as
$${u}_{11}^{(l)}{\mathbb{=}}-{\mathbb{H}}\left\{\frac{\partial {{{\mathcal{L}}}}}{\partial {{{{\bf{\Theta }}}}}^{(l)}}\right\}\odot {{{\bf{\Theta }}}}^{(l)}$$(5)where \({\mathbb{H}}\left\{\cdot \right\}\) denotes the Hessian matrix. To evaluate the importance of each layer based on a loss function derived straightforwardly from the product of all the layer parameters (no need for input data to compute), we employed synflow57 as
$${u}_{12}^{(l)}=\frac{\partial {{{\mathcal{L}}}}}{\partial {{{{\bf{\Theta }}}}}^{(l)}}\odot {{{\bf{\Theta }}}}^{(l)}$$(6) -
Metrics related to the pre-trained network. We also included the metrics aim to capture the inherent knowledge from the parameters of the pre-trained network, including the average and the standard deviation:
Sparse fine-tuning with a surgeon network
Sparse fine-tuning was achieved by selecting specific layers in the large pretrained network for modification while freezing the remaining layers. To efficiently select these layers based on the aforementioned zero-shot metrics, we employed a small surgeon network. The surgeon network was a 6-layer multilayer perceptron (MLP) with 200k parameters (Supplementary Fig. S2). This network took the zero-shot metrics \({{{{\bf{U}}}}}^{(l)}\), along with the positional embedding \(l\), as inputs and outputs a score \({s}^{(l)}\), representing the contribution of layer \(l\) in the pre-trained deblurring network. The layers with the highest 10% scores were then selected for sparse fine-tuning. To pre-train the surgeon network using ground truth labels, we created a meta-dataset (~1GB) comprising 5 different stacks from a custom-built nonlinear microscope39. Using this meta-dataset, 300 input-output data pairs were generated, following the format \(\{[{{{{\bf{U}}}}}^{(l)},{l}],{s}^{(l)}\}\) as defined above. In training the surgeon network, we employed the Adam optimizer with a learning rate of 10−4, a weight decay of 10−6, and the pairwise ranking loss, as we were only interested in the relative performance of the layers. The pre-trained surgeon network is released along with the codes.
Architecture, training, and inference of deblurring network
The deblurring network employed a NAFNet (Nonlinear Activation Free Network) topology with an encoder-decoder architecture for end-to-end deconvolution, which is a state-of-the-art deep learning architecture for image restoration tasks41,42. Designed to be simple yet effective, it replaces traditional nonlinear activation functions with element-wise multiplication and channel attention mechanisms, facilitating efficient learning of image characteristics. In this work, we employ the pre-trained NAFNet (~150 million parameters), which has been trained on large natural scene datasets (~50,000 images from REDS52), to leverage its strong understanding of image deblurring features.
Due to the inherent nature of the method, the network was implemented in 2D, with input patches of 256 × 256 and a batch size of 8. For all presented experiments, network parameters were optimized using the Adam optimizer with a learning rate of 10−3 and weight decay of 10−4. Training SSAI-3D took approximately 30 min on a single Nvidia GeForce RTX 3080 card, while inference for a 300 × 300 × 300 tensor took roughly 50 s. To minimize memory consumption, the model was quantized to float16 precision. The employed deblurring network contained approximately 100 million parameters. Only the deblurring network was modified, with ~10% of the layers fine-tuned and the remaining ~90% frozen. During inference, to reconstruct the entire 3D volume, axial images in both XZ and YZ directions were first denoised using the same denoising network. They were then deblurred with the fine-tuned deblurring network, and finally averaged into a single stack with isotropic resolution. To adapt the grayscale input images (with size L × W) to the pretrained network requiring RGB input (with size L × W × 3), in the training stage, grayscale images were replicated across the third dimension to create tensors (with size L × W × 3), while in the inference stage, the output tensors were averaged along the third dimension to obtain the final grayscale predictions (with size L × W).
Simulation data generation
For the simulation of beads in Fig. 2a, the ground truth image volumes were generated with randomly located beads of random radius and intensity. The isotropic beads were generated according to a 3D Gaussian distribution with fixed standard deviations of 2.0 in all three dimensions. The volume for training the model contained 50 beads in a 30 µm × 30 µm × 30 µm volume, with 100 nm pixel size, assuming a lateral resolution of 200 nm and axial resolution of 700 nm. For the simulation of symmetric tubular strands in Fig. 4c, the ground truth image volumes contained strands facing random directions at random locations, with a mean thickness of 4 pixels and intensity values ranging from 15000 to 65535. The volume for training the model contained 5000 tubes in a 300 µm × 300 µm × 300 µm volume. A 3D elastic grid-based deformation field was then applied to deform the volume, creating the ground truth.
Different blur levels were generated using a 51 × 51 × 51 3D Gaussian kernel, elongated axially to serve as raw input, with a standard deviation of 0.5 in lateral directions and 4.0 in the axial direction. For the simulation of asymmetric tubular strands in Fig. 2b, the ground truth volume was generated via 3D Bezier curves, shaped as curves in the lateral direction and straight horizontal lines axially. This created semi-synthetic data with vastly different distributions across views. A total of 5000 strands were generated in a 300 µm × 300 µm × 300 µm volume, with a thickness of 1–4 pixels, a total length of around 1200 pixels, and binary brightness values. Similar to the previous simulation, blur was introduced using a 51 × 51 × 51 3D Gaussian kernel with anisotropic standard deviations in lateral and axial directions.
Sample preparation and imaging acquisition
All animal procedures were conducted in accordance with a protocol approved by the Institutional Animal Care and Use Committee at the Massachusetts Institute of Technology (2309000575).
In Figs. 3g, i and 5b, d, f, 3D imaging in living tissues was achieved using a custom-built nonlinear microscope39,58. A custom-built multimode fiber source at 1100 nm was employed for simultaneous label-free imaging of THG, NAD(P)H, SHG, and FAD39,40. To achieve deep imaging with uniform SNR across different depths, pulse energy was varied according to imaging depth, maintaining approximately the same pulse energy at the excitation focus39. In Fig. 3g, the blood-brain barrier microfluidic model was engineered using a tri-culture of primary human astrocytes (ScienCell, 1800), primary human brain pericytes (ScienCell, 1200), and induced pluripotent stem cell (iPSC)-derived vascular endothelial cells (Alstem, iPS11). Cells were expanded in the following media: Astrocyte Medium (ScienCell, 1801), Pericyte Medium (ScienCell, 1201), or VascuLife VEGF Endothelial Medium (Lifeline Cell Technology, LL-0003) supplemented with 10% fetal bovine serum (Thermo Fisher Scientific, 26140-079) and SB 431542 (Selleckchem, S1067)39. In Fig. 5d, mouse whisker pad tissues were obtained from 12-week-old adult C57BL/6 mice (n = 1 with 1 male) after heparinization and transcardial perfusion with sucrose solution. The tissue was dissected, shaved to remove the top skin layer, and fixed in 4% paraformaldehyde (PFA) solution at 4 °C. Animals were housed in the vivarium with a 12-h light/dark cycle and were given food and water ad libitum under room temperature and humidity. Findings were not sex-dependent in this study. In Figs. 3i and 5b, f, freshly excised human endometrial tissue was obtained from patient undergoing hysterectomy surgery (n = 1, with age 48). All participants provided informed consent, in accordance with protocols approved by the Partners Human Research Committee and the Massachusetts Institute of Technology Committee on the Use of Humans as Experimental Subjects53. Findings were not sex-dependent in this study.
In Figs. 5a, c, e, open-source datasets with raw images acquired from commercial 3D microscopes using mouse tissues were employed. In Fig. 5a, 3DISCO-cleared mouse brains were imaged with LaVision light-sheet microscopes at a voxel size of 1.63 µm × 1.63 µm × 3 µm. Brain structures were visualized using Alexa Fluor 59459. In Fig. 5c, a Thy1-GFP mouse was used, and the mouse brain was sectioned into 300-µm-thick tissue slices. The sections were optically cleared by CUBIC and then imaged using the Nikon Ni-E A1 microscope in confocal mode at a 0.21 µm × 0.21 µm × 1 µm voxel size32. In Fig. 5e, the mouse liver was sectioned into 200-µm-thick tissue slices. The sections were optically cleared by CUBIC-50 and imaged using a wide-field microscope. A water-immersion objective (Olympus, XLUMPLFLN) was used for acquisition, and the voxel size was 0.32 µm × 0.32 µm × 1 µm32.
In Fig. 6a, an open-source dataset of a whole mouse brain expressing tdTomato in VIP neurons was utilized, with raw images acquired and co-registered with near-isotropic resolution images from mesoSPIM for comparison. mesoSPIM is a light-sheet microscope that achieves near-isotropic imaging by incorporating the axially scanned light-sheet (ASLM) module, which scans the excitation beam and selectively illuminates the focal plane to improve axial resolution50. Detailed sample preparation and imaging acquisition procedures were presented in ref. 23. Briefly, a whole mouse brain expressing tdTomato in VIP neurons was transcardially perfused with PBS, followed by an ice-cold hydrogel solution for clearing. The brain was then equilibrated in refractive index matching solution for at least 4 days before imaging. During imaging, the brain was loaded into and immersed within a quartz cuvette filled with index-matching oil, allowing for sample movement and rotation along all directions23. The raw images and co-registered near-isotropic resolution images were acquired with the ASLM module turned on and off, respectively16.
In Fig. 6c, an open-source dataset of an expanded mitochondrion was utilized, with raw images acquired and co-registered with near-isotropic resolution images from multiview confocal microscopy for comparison. Detailed sample preparation and imaging acquisition procedures were presented in ref. 14. Briefly, U2OS cells were fixed with 4% PFA and 0.25% glutaraldehyde (Sigma) for 15 minutes and washed three times in PBS. Mitochondria were labeled using rabbit-ε-Tomm20 (Abcam), ε-rabbit-biotin (Jackson ImmunoResearch), and Alexa Fluor 488. DNA was labeled using 5 µg/mL of mouse-ε-DNA (Progen) and 1 µg/mL of ε-mouse-JF549 (Novus Biologicals), each for 1 h in PBS. The raw images and co-registered near-isotropic resolution images were acquired without and with the use of triple-view confocal imaging, respectively14.
Method comparison
We compared the performance of SSAI-3D with four baseline methods: Richardson-Lucy, OT-CycleGAN31, Self-Net32, and CARE36. These methods were all implemented by open-source codes released by the cited papers. For CARE, a 2D semi-synthetic dataset was created by applying an explicit PSF to blur the lateral images, and a 2D deblurring network was trained from scratch to deblur the axial images36. In Fig. 2i, j, a Gaussian blur with a standard deviation of 5 was assumed to be known for CARE. To assess robustness against PSF mismatch, the model trained with this known blur was applied to different levels of blurring for CARE, while a single model trained on varying blurs was used for SSAI-3D. For Self-Net, ablation of PSF mismatch in the training and inference stages is not applicable because, as an unsupervised method, it does not rely on PSF assumptions. In Fig. 6a, c, where the blurring was spatially varying, a single blur within the range of the axial degradation was selected. In Fig. 3b, e, and Supplementary Fig. S8, SSAI-3D with full fine-tuning was implemented by fine-tuning the entire pretrained deblurring network without freezing any layers. For all the experiments presented, the hyper-parameters and training settings were universal.
Evaluation metrics
The MSE used in the paper is defined as
where X denotes the restored image, X0 denotes the corresponding ground truth, and N denotes the number of pixels. The SSIM used in the paper is defined as
where \(\left\{\mu,{\mu }_{0}\right\}\) and \(\left\{\sigma,{\sigma }_{0}\right\}\) are the means and variances of X and \({{{{\bf{X}}}}}_{0}\), respectively. \(\sigma^{\prime}\) is the covariance of X and X0. The two constants \({C}_{1}={\left({k}_{1}L\right)}^{2}\) and \({C}_{2}={\left({k}_{2}L\right)}^{2}\) are chosen with \({k}_{1}=0.01\), \({k}_{2}=0.03\), and \(L=65535\) as the dynamic range of the pixel values in 16-bit images. In Fig. 2c, the axial resolution after isotropic resolution recovery was measured by the full width at half maximum (FWHM) of the sub-diffraction-limited beads in z-direction (mean ± standard deviation, n = 50 beads). In Fig. 5, the resolution of the image was characterized by the width in its Fourier spectrum. In Fig. 6b, d, the individual granules were detected by an universal threshold of pixel value in raw, software-corrected, and hardware-corrected images. The detection accuracy of image \({{{\bf{X}}}}\) was characterized by the normalized cross-correlation (NCC) against hardware-corrected image \(\overline{{{{\bf{X}}}}}\) as
Statistics and reproducibility
All box plots adhere to the standard format: box bounds represent the upper and lower quartiles, lines within boxes indicate medians, and whiskers extend to data points within 1.5 times the IQR, with outliers plotted individually beyond this range. Experiments in Figs. 4h–k, 5a–f, and 6a–d were independently repeated at least 3 times under different random seeds, all achieving similar results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The open-source datasets of mouse brain vasculature, brain neurons, and liver used in this study are available at https://doi.org/10.5281/zenodo.788251932 (Fig. 5a, c, e). The open-source dataset of light-sheet imaging of a whole mouse brain in this study is available at https://idr.openmicroscopy.org/23 (under accession number idr0066, Fig. 6a; download instructions can be found at https://idr.openmicroscopy.org/about/download.html). The open-source dataset of confocal imaging of a mitochondria in this study is available at https://zenodo.org/record/549595514 (Fig. 6c). The label-free nonlinear imaging datasets as well as the network checkpoints, including pre-trained NAFNet, sparsely fine-tuned NAFNet, and pre-trained surgeon network generated in this study are available at https://doi.org/10.5281/zenodo.1395254960. Source data are provided with this paper.
Code availability
The implementations of SSAI-3D as well as some examples are publicly available at https://github.com/You-Lab-MIT/SSAI-3D and https://doi.org/10.5281/zenodo.1395260360,61. Part of the code is built upon NAFNet, which is publicly available at https://github.com/megvii-research/NAFNet41,42.
References
Gobel, W., Kampa, B. M. & Helmchen, F. Imaging cellular network dynamics in three dimensions using fast 3D laser scanning. Nat. Methods 4, 73–79 (2007).
Wang, T. et al. Three-photon imaging of mouse brain structure and function through the intact skull. Nat. Methods 15, 789–792 (2018).
Hernandez, J. C. C. et al. Neutrophil adhesion in brain capillaries reduces cortical blood flow and impairs memory function in Alzheimer’s disease mouse models. Nat. Neurosci. 22, 413–420 (2019).
Olivier, N. et al. Cell lineage reconstruction of early zebrafish embryos using label-free nonlinear microscopy. Science 329, 967–971 (2010).
Pulous, F. E. et al. Cerebrospinal fluid can exit into the skull bone marrow and instruct cranial hematopoiesis in mice with bacterial meningitis. Nat. Neurosci. 25, 567–576 (2022).
Zhao, Z. et al. Two-photon synthetic aperture microscopy for minimally invasive fast 3D imaging of native subcellular behaviors in deep tissue. Cell 186, 2475–2491 (2023).
Boden, A. et al. Volumetric live cell imaging with three-dimensional parallelized RESOLFT microscopy. Nat. Biotechnol. 39, 609–618 (2021).
Verveer, P. J. et al. High-resolution three-dimensional imaging of large specimens with light sheet-based microscopy. Nat. Methods 4, 311–313 (2007).
Zipfel, W. R., Williams, R. M. & Webb, W. W. Nonlinear magic: Multiphoton microscopy in the biosciences. Nat. Biotechnol. 21, 1369–1377 (2003).
Planchon, T. A. et al. Rapid three-dimensional isotropic imaging of living cells using Bessel beam plane illumination. Nat. Methods 8, 417–423 (2011).
Aquino, D. et al. Two-color nanoscopy of three-dimensional volumes by 4Pi detection of stochastically switched fluorophores. Nat. Methods 8, 353–359 (2011).
Jia, S., Vaughan, J. C. & Zhuang, X. Isotropic three-dimensional superresolution imaging with a self-bending point spread function. Nat. Photon. 8, 302–306 (2014).
Chhetri, R. K. et al. Whole-animal functional and developmental imaging with isotropic spatial resolution. Nat. Methods 12, 1171–1178 (2015).
Wu, Y. et al. Multiview confocal super-resolution microscopy. Nature 600, 279–284 (2021).
Zhao, Y. et al. Isotropic super-resolution light-sheet microscopy of dynamic intracellular structures at subsecond timescales. Nat. Methods 19, 359–369 (2022).
Dean, K. M. et al. Isotropic imaging across spatial scales with axially swept light-sheet microscopy. Nat. Protoc. 17, 2025–2053 (2022).
Li, X. et al. Three-dimensional structured illumination microscopy with enhanced axial resolution. Nat. Biotechnol. 41, 1307–1319 (2023).
Ren, D., Zhang, K., Wang, Q., Hu, Q. & Zuo, W. Neural blind deconvolution using deep priors. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 3341–3350 (IEEE, 2020).
Khan, S. S., Sundar, V., Boominathan, V., Veeraraghavan, A. & Mitra, K. Flatnet: Towards photorealistic scene reconstruction from lensless measurements. IEEE Trans. Pattern Anal. Mach. Intell. 44, 1934–1948 (2020).
Whang, J. et al. Deblurring via stochastic refinement. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 16293–16303 (IEEE, 2022).
Yanny, K., Monakhova, K., Shuai, R. W. & Waller, L. Deep learning for fast spatially varying deconvolution. Optica 9, 96–99 (2022).
Kohli, A. et al. Ring deconvolution microscopy: An exact solution for spatially-varying aberration correction. Preprint at https://arxiv.org/abs/2206.08928 (2022).
Voigt, F. F. et al. The mesoSPIM initiative: Open-source light-sheet microscopes for imaging cleared tissue. Nat. Methods 16, 1105–1108 (2019).
Moor, M. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023).
Ma, C., Tan, W., He, R. & Yan, B. Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration. Nat. Methods 21, 1558–1567 (2024).
Luo, Y., Zheng, L., Guan, T., Yu, J. & Yang, Y. Taking a closer look at domain shift: Category-level adversaries for semantics consistent domain adaptation. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2507–2516 (IEEE, 2019).
Zhang, K., Scholkopf, B., Muandet, K. & Wang, Z. Domain adaptation under target and conditional shift. In International Conference on Machine Learning (ICML), 819–827 (ICML, 2013).
Saito, K., Ushiku, Y. & Harada, T. Asymmetric tri-training for unsupervised domain adaptation. In International Conference on Machine Learning (ICML), 2988–2997 (ICML, 2017).
Guo, M. et al. Deep learning-based aberration compensation improves contrast and resolution in fluorescence microscopy. bioRxiv (2023).
Li, X. et al. Unsupervised content-preserving transformation for optical microscopy. Light Sci. Appl. 10, 44 (2021).
Park, H. et al. Deep learning enables reference-free isotropic superresolution for volumetric fluorescence microscopy. Nat. Commun. 13, 3297 (2022).
Ning, K. et al. Deep self-learning enables fast, high-fidelity isotropic resolution restoration for volumetric fluorescence microscopy. Light Sci. Appl. 12, 204 (2023).
Yadav, A., Shah, S., Xu, Z., Jacobs, D. & Goldstein, T. Stabilizing adversarial nets with prediction methods. In International Conference on Learning Representations (ICLR), (ICLR, 2018).
Cohen, J. P., Luck, M. & Honari, S. Distribution matching losses can hallucinate features in medical image translation. In Medical Image Computing and Computer Assisted Intervention (MICCAI), 529–536 (Springer, 2018).
de Haan, K. et al. Deep learning-based transformation of H&E stained tissues into special stains. Nat. Commun. 12, 1–13 (2021).
Weigert, M. et al. Content-aware image restoration: Pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Chen, J. et al. Three-dimensional residual channel attention networks denoise and sharpen fluorescence microscopy image volumes. Nat. Methods 18, 678–687 (2021).
Guo, M. et al. Rapid image deconvolution and multiview fusion for optical microscopy. Nat. Biotechnol. 38, 1337–1346 (2020).
Liu, K. et al. “Deep and dynamic metabolic and structural imaging in living tissues.” Sci. Adv. 10, eadp2438 (2024).
You, S. et al. Intravital imaging by simultaneous label-free autofluorescence-multiharmonic microscopy. Nat. Commun. 9, 2125 (2018).
Chen, L., Chu, X., Zhang, X. & Sun, J. Simple baselines for image restoration. In European Conference on Computer Vision (ECCV), 17–33 (ECCV, 2022).
Wang, X. et al. BasicSR: Open source image and video restoration toolbox. https://github.com/XPixelGroup/BasicSR (2022).
Lee, Y. et al. Surgical fine-tuning improves adaptation to distribution shifts. In The Eleventh International Conference on Learning Representations (ICLR, 2022).
Wortsman, M. et al. Robust fine-tuning of zero-shot models. In Proceedings of the IEEE/CVF conference on Computer Vision and Pattern Recognition (CVPR), 7959–7971 (IEEE, 2022).
Hendrycks, D. & Gimpel, K. A baseline for detecting misclassified and out-of-distribution examples in neural networks. In International Conference on Learning Representations (ICLR) (ICLR, 2016).
Hendrycks, D. et al. The many faces of robustness: A critical analysis of out-of-distribution generalization. In Proceedings of the IEEE/CVF International Conference on Computer Vision (ICCV), 8340–8349 (ICCV, 2021).
Andreassen, A. J., Bahri, Y., Neyshabur, B. & Roelofs, R. The evolution of out-of-distribution robustness throughout fine-tuning. Trans. Mach. Learn. Res. (2022).
Hajal, C. et al. Engineered human blood-brain barrier microfluidic model for vascular permeability analyses. Nat. Protoc. 17, 95–128 (2022).
Li, X. et al. Real-time denoising enables high-sensitivity fluorescence time-lapse imaging beyond the shot-noise limit. Nat. Biotechnol. 41, 282–292 (2023).
Dean, K. M., Roudot, P., Welf, E. S., Danuser, G. & Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys. J. 108, 2807–2815 (2015).
Pi, H.-J. et al. Cortical interneurons that specialize in disinhibitory control. Nature 503, 521–524 (2013).
Liu, H. et al. Video super-resolution based on deep learning: A comprehensive survey. Artif. Intell. Rev. 55, 5981–6035 (2022).
Ye, C. T. et al. Learned, uncertainty-driven adaptive acquisition for photon-efficient multiphoton microscopy. Preprint at https://arxiv.org/abs/2310.16102 (2023).
Theis, L., Korshunova, I., Tejani, A. & Huszar, F. Faster gaze prediction with dense networks and fisher pruning. Preprint at https://arxiv.org/abs/1801.05787 (2018).
Lee, N., Ajanthan, T. & Torr, P. SNIP: Single-shot network pruning based on connection sensitivity. In International Conference on Learning Representations (ICLR) (ICLR, 2018).
Wang, C. et al. “Picking Winning Tickets Before Training by Preserving Gradient Flow.” In International Conference on Learning Representations (ICLR) (ICLR, 2020).
Tanaka, H., Kunin, D., Yamins, D. L. & Ganguli, S. Pruning neural networks without any data by iteratively conserving synaptic flow. Adv. Neural Inf. Process. Syst. 33, 6377–6389 (2020).
Qiu, T. et al. Spectral-temporal-spatial customization via modulating multimodal nonlinear pulse propagation. Nat. Commun. 15, 2031 (2024).
Todorov, M. I. et al. Machine learning analysis of whole mouse brain vasculature. Nat. Methods 17, 442–449 (2020).
Han, J., Liu, K. & You, S. System- and sample-agnostic isotropic three-dimensional microscopy by weakly physics-informed, domain-shift-resistant axial deblurring. URL https://doi.org/10.5281/zenodo.13952549 (2024).
Han, J., Liu, K. & You, S. System- and sample-agnostic isotropic three-dimensional microscopy by weakly physics-informed, domain-shift resistant axial deblurring. URL https://doi.org/10.5281/zenodo.13952603 (2024).
Acknowledgements
We thank Honghao Cao and Tong Qiu for assistance in nonlinear imaging, as well as helpful discussions from Matthew Yeung, Steven F. Nagle, and Haidong Feng. We thank Song Han for insights into sparse fine-tuning. We thank Roger D. Kamm, Sarah Spitz, and Francesca Michela Pramotton for providing the human blood-brain barrier microfluidic model for nonlinear imaging. We thank Ellen L. Kan for assistance in preparing the human endometrium tissue for nonlinear imaging. We thank Fan Wang, Manuel Levy, and Eva Lendaro for providing the mouse whisker pad tissue for nonlinear imaging. We acknowledge that Fig. 1a contains materials from BioRender. The work has been supported by MIT startup funds, NSF CAREER Award (2339338), and CZI Dynamic Imaging via Chan Zuckerberg Donor Advised Fund (DAF) through the Silicon Valley Community Foundation (SVCF). K.L. acknowledges support from the MIT Irwin Mark Jacobs (1957) and Joan Klein Jacobs Presidential Fellowship.
Author information
Authors and Affiliations
Contributions
J.H., K.L., and S.Y. conceived the idea of the project. J.H. designed detailed implementations in network training and inference. K.L. built the optical setup and performed the imaging experiments. J.H. and K.L. analyzed the data and prepared the figures with input from K.B.I., K.M., and L.G.G. K.B.I. and L.G.G. provided the samples for imaging and insights into the biomedical questions. J.H., K.L., and S.Y. wrote the manuscript with the input from all authors. S.Y. obtained the funding and supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jeroen Kalkman and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, J., Liu, K., Isaacson, K.B. et al. System- and sample-agnostic isotropic three-dimensional microscopy by weakly physics-informed, domain-shift-resistant axial deblurring. Nat Commun 16, 745 (2025). https://doi.org/10.1038/s41467-025-56078-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56078-4
This article is cited by
-
Advanced imaging strategies in cardiac organoids: bridging the gap between structural complexity and functional analysis
Cardiovascular Diabetology (2026)
-
Near-isotropic super-resolution microscopy with axial interference speckle illumination
Nature Communications (2025)