Abstract
Spatial transcriptomics is an essential application for investigating cellular structures and interactions and requires multimodal information to precisely study spatial domains. Here, we propose STAIG, a deep-learning model that integrates gene expression, spatial coordinates, and histological images using graph-contrastive learning coupled with high-performance feature extraction. STAIG can integrate tissue slices without prealignment and remove batch effects. Moreover, it is designed to accept data acquired from various platforms, with or without histological images. By performing extensive benchmarks, we demonstrate the capability of STAIG to recognize spatial regions with high precision and uncover new insights into tumor microenvironments, highlighting its promising potential in deciphering spatial biological intricates.
Similar content being viewed by others
Introduction
Biological tissues are intricate networks of various cell types that perform essential functions through their unique spatial configurations. Recent spatial transcriptomics (ST) techniques, such as 10x Visium1, Slide-seq2, Stereo-seq3, and STARmap4, have enhanced our ability to map genetic data within these configurations, providing deeper insights into the genetic organization of tissues in health and disease, and advancing our understanding of molecular and physiological intricacies.
ST heavily relies on the identification of spatial domains with uniform gene expression and histology. Currently, two main identification approaches exist, nonspatial and spatial clustering approaches5. Non-spatial clustering methods such as K-means6, Louvain7, and Seurat8 derive clustering results based solely on gene expression, often resulting in disjointed clusters that poorly reflect the actual spatial patterns of tissues. In contrast, emerging spatial clustering methods integrate genetic and spatial information using graph convolutional models. For instance, SEDR9 employs a deep autoencoder alongside a variational graph autoencoder to map gene data. However, this class of methods may have potential pitfalls. Specifically, the transformation of ST data into graph structures primarily relies on artificially defined distance criteria, which can introduce biases. These biases arise from the uniform construction of edges between regions that belong to different biological domains but meet arbitrary criteria.
Although some methods like STAGATE10 address this issue by assigning weights to edges based on cell gene similarity, they do not integrate the image information. Moreover, other methods, such as GraphST11 and ConST12 utilize contrastive learning frameworks that preserve the original structure and involve a random shuffling of nodes during training, leaving the structure intact throughout the process. Concurrently, these methods fail to consider the exclusion of potentially similar nodes from the negative samples during comparisons, resulting in biases from false negatives impacting the outcomes.
In addition, utilizing histological images, such as those stained with Hematoxylin and Eosin (H&E), presents significant challenges. Methods like stLearn13 and SpaGCN14 directly use information derived from image RGB values, making them susceptible to variations in staining quality. Techniques like DeepST15 and ConST12 attempt to enhance gene matrices by extracting image features using neural networks and the pre-trained MAE16 visual model, respectively. However, the limited availability of histological images restricts the pre-training of these models, thereby affecting the quality of feature extraction and diminishing their broad applicability.
Lastly, batch integration in most current methods still necessitates manual intervention, such as manually aligning coordinates or relying on additional tools17. Exploring how models can identify commonalities in gene expression through local contrast rather than global node shuffling, offers a promising avenue for eliminating batch effects in the feature space.
To overcome these limitations, we propose STAIG (Spatial Transcriptomics Analysis via Image-Aided Graph Contrastive Learning), a deep learning framework that integrates gene expression, spatial data, and histological images without relying on alignment. STAIG extracts features from H&E-stained images using a self-supervised model that does not require pre-training on extensive histology datasets. In addition, STAIG dynamically adjusts the graph structure during training and selectively excludes homologous negative samples, guided by information from histological images, thus minimizing biases from initial construction. Finally, STAIG performs end-to-end batch integration by recognizing gene expression commonalities through local contrast, eliminating the need for manual coordinate alignment and reducing batch effects. We have evaluated STAIG using diverse datasets and observed its promising capabilities in spatial domain identification. Our results demonstrate that STAIG effectively reveals detailed spatial and genetic information within tumor microenvironments, advancing our understanding of complex biological systems.
Results
Overview of STAIG
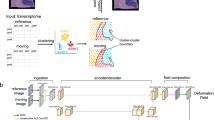
To reduce noise and uneven staining, STAIG first segments histological images into patches that align with the spatial locations of data spots, and subsequently refines these images with a band-pass filter. Image embeddings were extracted using the Bootstrap Your Own Latent (BYOL)18 self-supervised model, and an adjacency matrix was simultaneously constructed based on the spatial distances between the spots (Fig. 1a). To integrate the data from different tissue slices, the image embeddings for multiple tissue slices were stacked vertically (Fig. 1b), and the adjacency matrices were merged using a diagonal placement method (Fig. 1c).
a STAIG begins with spatial transcriptomic (ST) data. Each slice includes spots with spatial coordinates, gene data, and optional Hematoxylin and Eosin (HE) stained images. Image patches at these spots undergo noise reduction, including bandpass filtering, before being processed for image embeddings via the BYOL framework. Parallelly, an adjacency matrix is created using the spatial data of the spots. b For multiple slices, the image embeddings from each slice are vertically merged, forming an image embedding space where spots are distributed, with dotted lines indicating their Euclidean distances. c Adjacency matrices from each section are combined diagonally to form an integrated adjacency matrix. This matrix is then used to construct a graph, with gene expression data represented as node information. d For spots connected by edges, distances are calculated in the image embedding space. These distances are then transformed into probabilities of random edge removal using a SoftMax function. The original graph undergoes two rounds of edge random removal based on these probabilities, creating two augmented views. Subsequently, features of nodes in these graph views are randomly masked. e A graph neural network (GNN) processes these augmented views, concentrating on node-level differences. This step is encapsulated within a dotted box, emphasizing the neighbor contrastive strategy. f The derived embeddings from the GNN are then utilized for spatial domain identification and integration.
Next, an original graph was built from the integrated adjacency matrix, where nodes represent gene expression and edges reflect adjacency (Fig. 1c). To generate two independent augmented views from this graph, STAIG employs a parallel and independent augmentation process (Fig. 1d). For each augmented view, the process involved randomly removes edges from the original graph, guided by an image-driven probability (or a fixed probability in the absence of images). This is followed by the random masking of gene features, entailing the zeroing of a subset of gene values. More importantly, the probability of edge removal is estimated based on the Euclidean distance between nodes in the image feature space, thereby introducing image-informed augmentation into the graph.
The augmented views are processed using a shared GNN guided by a neighboring contrastive objective. This approach aims to closely align nodes and their adjacent neighbors in two graph views while distancing non-neighboring nodes (Fig. 1e). In addition, when images are available, the Debiased Strategy (DS)19 leverages image embeddings as prior knowledge to ensure nodes that are closer based on image features remain proximate (details in the Methods section). Finally, the trained GNN produced embeddings to identify the spatial regions and minimize batch effects across consecutive tissue slices (Fig. 1f).
Performance in the identification of brain regions
To assess the performance of STAIG in the identification of tissue regions, we prepared 10x Visium human and mouse brain ST datasets: 12 human dorsolateral prefrontal cortex (DLPFC)20 slices that were manually annotated into cortical layers L1‒L6 and white matter (WM), and mouse anterior and posterior brain sections. We benchmarked STAIG with existing methods such as Seurat, GraphST, DeepST, STAGATE, SpaGCN, SEDR, ConST, MuCoST21, and stLearn. To quantify the performance, the Adjusted Rand Index (ARI)22 and Normalized Mutual Information (NMI)23 were used for manually annotated datasets, and the Silhouette Coefficient (SC)24 and Davies-Bouldin Index (DB)25 were employed for others (refer to Baseline methods and evaluation metrics in the Supplementary Notes).
Overall, for the human brain datasets, STAIG achieved the highest median ARI (0.69 across all slices) and NMI (0.71) (Fig. 2a and Supplementary Figs. S1 and S2); in particular, the ARI reached 0.84 with a corresponding NMI of 0.78 in slice #151672 (Fig. 2b). Regarding slice #151673 used in previous studies10,11,15, STAIG achieved the highest ARI of 0.68 and NMI of 0.74, not only distinctly separating tissue layers L1‒L6 and WM in uniform manifold approximation and projection (UMAP) visualizations but also closely matched manual labels and accurately recognized layer proportions (Fig. 2c). Conversely, the results demonstrate that the existing methods provide relatively lower ARIs and NMIs with misclassified spots and inaccuracies in layer proportions (Fig. 2d, e): stLearn yielded a missing layer and misclassifying spots; GraphST achieved an ARI of 0.64 and an NMI of 0.73 but had discrepancies in the positioning of layers 4 and 5; other methods recorded ARIs between 0.25 and 0.57, with corresponding NMIs ranging from 0.42 to 0.69, owing to inaccuracies in layer proportions.
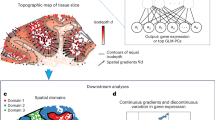
a Boxplots of adjusted rand index (ARI) and normalized mutual information (NMI) of the nine methods applied to all 12 DLPFC slices (n = 12 biological replicates). In the boxplot, the center line denotes the median, box limits denote the upper and lower quartiles, black dots denote individual slices, and whiskers denote the 1.5 × interquartile range. b Manual annotations and clustering results with ARI and NMI by STAIG on DLPFC slice #151672. c Manual annotations, H&E stained image, clustering result with ARI, NMI, and UMAP visualization by STAIG on DLPFC slice #151673. d UMAP visualizations by baseline methods (stLearn, Seurat, STAGATE, SpaGCN, DeepST, conST, SEDR, GraphST, MuCoST) on DLPFC slice #151673. e Clustering results with ARI and NMI by baseline methods on DLPFC slice #151673. Manual annotations and clustering results of the other DLPFC slices are shown in Supplementary Figs. S1 and S2. f H&E stained image, anatomical annotations from the Allen Reference Atlas, and clustering results with Silhouette Coefficient (SC) and Davies-Bouldin Index (DB) by STAIG on mouse brain posterior tissue. The solid line box denotes the cerebellar cortex, and the dashed line box denotes the hippocampus area with Cornu Ammonis (CA) and Dentate Gyrus (DG). Clustering results from baseline methods are shown In Supplementary Fig. S3. g Manual annotations from Long et al. and bar charts of ARI by STAIG and baseline methods on mouse anterior tissue. The x-axis represents methods, and the y-axis represents ARI. Clustering results from baseline methods and bar charts of NMI are shown in Supplementary Fig. S4.
In the mouse posterior tissue dataset (Fig. 2f and Supplementary Fig. S3), STAIG successfully identified the cerebellar cortex and hippocampus region, further distinguishing the Cornu Ammonis (CA) and dentate gyrus sections. This was consistent with the Allen Mouse Brain Atlas annotations26. Although the overall accuracy decreased in the absence of manual labels, STAIG achieved the highest SC of 0.31 and the lowest DB of 1.11. In the mouse anterior tissue dataset (Fig. 2g and Supplementary Fig. S4), STAIG accurately demarcated areas, including the olfactory bulb and dorsal pallium, yielding the highest ARI of 0.44 and NMI of 0.72 when we used manual labels from Long et al.11 as a reference.
Efficacy of image feature extraction
To investigate the impact of the usage of image features, we applied the k-nearest neighbors algorithm (KNN)27 to the image features from STAIG and those from the comparison algorithms, that is, stLearn, DeepST, and ConST. For instance, in the analysis of slice #151507 (Fig. 3a), we found that the image features from stLearn were heavily influenced by stain intensity and led to mismatches with the actual layer annotations. Despite utilizing deep learning models, DeepST and ConST failed to capture the intricate texture characteristics of brain tissues.
a H&E stained image, manual annotations, and KNN clustering results based purely on image features by STAIG and three other image-based methods (stLearn, DeepST, ConST) on DLPFC slice #151507. Semi-transparent masks have been extracted from class labels to facilitate the comparison of layer stratification. b H&E stained image, manual annotations, and KNN clustering results based purely on image features by STAIG and three other image-based methods on DLPFC slice #151673. Semi-transparent masks extracted from manual annotations are utilized to compare layer stratification. c H&E stained image, visual interpretation-based manual annotations, and KNN clustering results based purely on image features by STAIG and three other image-based methods on the human breast cancer dataset. The gray contour lines, extracted from manual annotations, are utilized to compare clustering outcomes.
STAIG was the only method that aligned closely with the manually annotated layers in terms of specific distribution areas, although some boundaries were still somewhat indistinct despite minimal influence from staining variations (Fig. 3a). It displays the colored blocks composing Layer 6, clearly separating Layer 5 and the WM region, with only a small part on the left affected by staining. Moreover, the WM region was further divided into a smaller subregion, which is consistent with the appearance of the Histology image, where the bottommost area of the WM exhibits the darkest staining and cells with morphologies distinct from other regions. In addition, STAIG accurately distributed Layer 3 on either side of Layer 1. Furthermore, the performance of STAIG image features on slice #151673 was equally effective (Fig. 3b). Interestingly, the comparative methods for this slice generally succeeded in differentiating only between the WM and non-WM layers, whereas clustering of STAIG within the non-WM layers successfully identified Layer 3 and Layer 4.
We further investigated the effectiveness of the image features using a human breast cancer H&E-stained image (Fig. 3c); a pathologist manually annotated regions solely based on visual interpretation9. The results indicated that the image features from stLearn mixed tumor and normal regions, while those from ConST, although appearing to partition the image into various regions, exhibited inconsistencies with the manually annotated contours upon closer examination. DeepST failed to capture image features. Conversely, STAIG clustering precisely identified the tumor regions and maintained spatial region coherence in the clustering results, with the segmented regions nearly perfectly aligning with the manually annotated contours, suggesting effective image feature extraction capabilities in STAIG.
Identifying spatial domains without histological image data
Because ST experiments lacking image information are not unusual, we assessed the predictive performance with datasets where only gene expression and spatial information are available. Initially, we prepared a mouse coronal olfactory bulb dataset of Stereo-seq3 annotated using DAPI-stained images9 (Fig. 4a): the olfactory nerve layer (ONL), glomerular layer (GL), external plexiform layer (EPL), mitral cell layer (MCL), internal plexiform layer (IPL), granule cell layer (GCL), and rostral migratory stream (RMS).
a DAPI-stained image with annotated laminar organization and clustering results with SC and DB by STAIG, STAGATE, GraphST, and Seurat on Stereo-seq mouse olfactory bulb tissue. Clustering results from other baseline methods are shown in Supplementary Fig. S5. b Visualization of spatial domains identified by STAIG and the corresponding marker gene expressions on the Stereo-seq mouse olfactory bulb tissue. c Bar chart of SC by STAIG and baseline methods on Stereo-seq mouse olfactory bulb tissue. The x-axis are methods, and the y-axis represents SC. d Annotations from the Allen Reference Atlas and clustering results with SC and DB by STAIG on Slide-seqV2 mouse olfactory bulb tissue. Clustering results from baseline methods are shown in Supplementary Fig. S6. e Bar chart of SC by STAIG and baseline methods on Slide-seqV2 mouse olfactory bulb tissue. The x-axis represents SC, and the y-axis represents methods. f Annotations from the Allen Reference Atlas, clustering results with SC, and visualization of CA1, CA3, and DG domains identified by STAIG and the corresponding marker gene expressions on Slide-seqV2 mouse hippocampus tissue. Clustering results from baseline methods are shown in Supplementary Fig. S7. g Manual annotations and clustering results with ARI by STAIG, STAGATE, GraphST, and Seurat on STARmap mouse visual cortex dataset. Clustering results from other baseline methods are shown in Supplementary Fig. S8. h Manual annotations and clustering results with SC and DB by STAIG, GraphST, and STAGATE on MERFISH human middle temporal gyrus dataset (upper) and mouse visual cortex (VIS) dataset (lower). Clustering results from other baseline methods are shown in Supplementary Figs. S9 and S10.
All algorithms provided the overall trends of spatial stratification for the dataset, whereas disparities in the annotations were observed (Fig. 4a and Supplementary Fig. S5a). For example, Seurat and stLearn failed to distinguish between layers, and STAGATE and GraphST mix the IPL and GCL. DeepST yielded distinct stratifications but failed to discern the GL and IPL layers. STAIG successfully delineated all layers, including thinner layers such as the MCL and IPL. As shown in Fig. 4b, the expression of layer-specific marker genes28,29,30,31,32,33 supported the high-resolution predictions of STAIG, outperforming the existing algorithms (Fig. 4c and Supplementary Fig. S5b).
Next, we tested the algorithms using the mouse olfactory bulb tissue dataset Slide-seqV234 (Fig. 4d and Supplementary Fig. S6a). All algorithms, except STAIG, GraphST, and STAGATE, struggled to discern the stratification of the RMS, accessory olfactory bulb (AOB), and granular layer of the accessory olfactory bulb (AOBgr). In contrast, STAIG displayed a higher performance in interlayer clustering (Fig. 4e and Supplementary Fig. S6b), and the results of STAIG matched well with the expression of layer-specific marker genes35,36 (Supplementary Fig. S6c).
Using the mouse hippocampus dataset34 from Slide-seqV2 (Fig. 4f and Supplementary Fig. S7) were further assessed. Notably, several methods, such as Seurat, stLearn, SpaGCN, and SEDR, display unclear clustering results that lack spatial cohesion. However, STAIG, GraphST, STAGATE, and DeepST delineated spatially consistent clusters for the major anatomical regions, particularly the CA, which was accurately categorized into CA1 and CA3 sections. Again, STAIG had the highest SC of 0.40 and the lowest DB of 0.87, which was validated by the expression of marker genes37,38,39.
Furthermore, using the mouse visual cortex dataset of STARmap dataset4 with inherent annotations, we found that STAIG outperformed all other methods, achieving the highest ARI of 0.67 and NMI of 0.72, which markedly surpassed the next-best method, STAGATE (Fig. 4g and Supplementary Fig. S8). Moreover, the UMAP plots highlight the ability of STAIG to completely separate all neocortical layer clusters.
Lastly, we applied STAIG to two datasets40 from the MERFISH platform: one featuring 4000 genes in the human middle temporal gyrus (MTG) and another with 243 genes in the mouse visual cortex (VIS). The analysis incorporated both the manually annotated cellular classes and layer-specific labels from the original research (Fig. 4h). In the human MTG dataset, STAIG not only achieved the highest SC of 0.28 and the lowest DB of 1.26 compared to other algorithms but was also the only method that distinctly identified each layer (Fig. 4h and Supplementary Fig. S9). Other algorithms failed to clearly delineate the layers, with many clustering various layers together. Notably, although STAGATE performed relatively well, it still confused layers L4, L5, and L6. In the mouse VIS dataset, STAIG continued to outperform all other methods in terms of metric scores (SC:0.32, DB:1.09) (Fig. 4h and Supplementary Fig. S10). The clustering results from STAIG were nearly identical to the real manual annotations. Another algorithm that showed significant layer differentiation was GraphST; however, it incorrectly classified some samples from layer L1 as belonging to layer L4.
Integrating slices without prior spatial alignment
STAIG can achieve alignment-free integration of spatial transcriptomic slices. To better evaluate the batch effects after integration, we introduced the BatchKL41 metric and the ILISI42 to assess the numerical distribution of the integrated data, in addition to the existing evaluation metrics. Furthermore, we included STAligner43, a machine-learning method specifically designed for integrating spatial transcriptomic slices, in our comparisons.
We first tested vertical and horizontal integration on samples from the same origin. For vertical integration, we selected adjacent continuous slices #151675 and #151676 from the same donor in the DLPFC dataset. The UMAP results showed that the two slices were well-integrated (BatchKL: 0.14, ILISI: 1.88), with STAIG achieving the highest ARI of 0.64 and the highest NMI of 0.71 (Fig. 5a and Supplementary Fig. S11).
a Vertical integration of STAIG (DLPFC slice #151675 and #151676). Includes manual annotations, integrated UMAP, ARI, NMI, BatchKL, and ILISI. Baseline methods results are detailed in Supplementary Fig. S11. b Horizontal integration of STAIG. Includes SC, DB, BatchKL, ILISI, H&E stained images, annotations from the Allen Reference Atlas, integration results by STAIG on mouse anterior and posterior brain tissue slices, and UMAP. The solid line box highlights the hippocampus zones in both slices. Baseline methods results are shown in Supplementary Fig. S12. c Integration of STAIG for slices from different sources (DLPFC #151507, #151508 and #151675, #151676), including manual annotations, ARI, NMI, BatchKL, ILSI, and UMAP visualizations. Baseline methods results are shown in Supplementary Fig. S13. d Integration of STAIG for slices from different sources(MERFISH, VIS regions of two mice. The results include manual annotations, SC, DB, BatchKL, ILISI, clustering results, and UMAP visualizations. Baseline methods results are shown in Supplementary Fig. S14. e Cross-platform integration of STAIG (Stereo-seq and Slide-seqV2, mouse olfactory bulb dataset). The results include SC, DB, BatchKL, ILISI, clustering results, and UMAP visualizations. Results from baseline methods are presented in Supplementary Fig. S15.
Next, we tested horizontal integration using anterior and posterior mouse brain slices. Unlike the methods SpaGCN and GraphST, which require pre-alignment of slice edges, STAIG precisely identified key brain structures, such as the cerebral cortex layers, cerebellum, and hippocampus, without prior spatial alignment. The results aligned well with the annotation of the Allen Brain Atlas (Fig. 5b) and showed accuracy comparable to the existing methods (Supplementary Fig. S12). In addition, the BatchKL and the ILISI were consistent with those of the comparative algorithms.
Further observation of the UMAP distribution after horizontal integration revealed that regions common to both the anterior and posterior mouse brain, such as the isocortex and thalamus, completely overlapped in the UMAP. In contrast, regions unique to each slice were separated in the UMAP. This is a normal phenomenon, as the two tissue slices have distinct functional regions, which demonstrates the effectiveness of the integration—neither completely separated nor completely overlapping. The UMAP results of the comparative methods GraphST and SpaGCN, which directly link the two slice graphs, also exhibited this pattern.
To evaluate the effectiveness of integration performance, we integrated slices from different individuals using the DLPFC dataset, specifically slices #151507, #151508, #151675, and #151676 from two donors. We compared this integration with methods that can directly integrate different sources, as well as with the STAGATE combined with Harmony42 (Fig. 5c and Supplementary Fig. S13). The UMAP from direct outputs of STAGATE revealed significant batch effects among samples from different individuals, with the BatchKL of 1.30 and the ILISI of 1.43. After applying Harmony, both the ARI and NMI of STAGATE increased, and the BatchKL decreased to 1.10. Despite these improvements, notable separation was still visible on the UMAP. The UMAP generated by the DeepST method, which employs Domain Adversarial Networks (DAN) for alignment, also exhibited clear separation. In contrast, STAIG, along with STAligner, achieved complete overlap on UMAP. STAIG consistently demonstrated the best integration, with the highest ARI of 0.52, NMI of 0.62, ILISI of 2.95, and the lowest BatchKL of 0.33.
In addition to the Visium platform, we extended our method comparisons to the MERFISH platform. We integrated datasets from the VIS region of two different mice (Fig. 5d and Supplementary Fig. S14). The two slices originated from different individuals, featuring non-identical tissue distributions. The slice from mouse 1 included a larger brain sample encompassing the VIS region, whereas the slice from mouse 2 comprised solely the VIS region. STAIG successfully achieved integration across these datasets. Among the comparative methods, only STAIG and STALigner managed to obtain clear layer separation that aligned with the original cell annotations. However, STAIG stood out with the highest SC of 0.40, the lowest DB of 0.88, and a reduced BatchKL of 0.26. Notably, while the STAGATE, post-integration with Harmony, showed improvements in metrics, the actual clustering quality deteriorated, with blurred demarcations between the WM and L6 regions.
Finally, we performed cross-platform integration of the SlideV2 and Stereo-seq platforms (Fig. 5e and Supplementary Fig. S15). STAIG successfully integrated the mouse olfactory bulb datasets from the two platforms. Compared to STAligner, which is also designed for cross-platform integration, our BatchKL was lower. Moreover, the shape of the UMAP showed that different layers were separately clustered, achieving higher SC and ILISI, with lower DB.
Specifying the tumor microenvironment from human breast cancer ST
In the analysis of the human breast cancer dataset, we found that the STAIG results closely matched the manual annotations and achieved the highest ARI of 0.64 and NMI of 0.70 (Supplementary Fig. S16). Notably, STAIG proposed a slightly different, yet more refined spatial stratification. In particular, for the region ‘Healthy_1’ in the manual annotation (Fig. 6a), STAIG dissected it into subclusters Clusters 3 and 4 (Fig. 6b). We noticed that the differentially expressed genes (DEGs) in Cluster 3 compared with other clusters (> 0.25 log2FoldChange, denotes as log2FC) (Fig. 6c) were involved in biological processes such as extracellular matrix organization, wound healing, and collagen fibril organization (Fig. 6d), with statistical significance (Gene Ontology enrichment analysis, adjusted p-value < 0.05).
a Manual annotation of human breast cancer dataset based on the HE-stain image. b Clustering results with ARI and NMI by STAIG on human breast cancer dataset. Clustering results from other baseline methods are shown in Supplementary Fig. S16. c Differential Gene Expression (DGE) analysis of Cluster 3 versus other clusters. Each point represents a gene, the vertical axis represents the -log10 of the p-value and the horizontal axis represents the log2FoldChange (log2 FC). P-values were derived from a two-sided Wilcoxon rank-sum test. The significance thresholds were set at |log2FC| > 0.25 and p-value < 0.05. d Gene Ontology (GO) analysis for Cluster 3 versus other clusters. The vertical axis represents the GO terms, and the horizontal axis represents the -log10 of the p-value. GO enrichment analysis was performed using the one-sided hypergeometric test, and p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. e Violin plots and the visualization of expression of CAF marker genes (COL6A1, COL1A2, VIM, PDGFRB, S100A4) in Cluster 3 (n = 166 spots) versus other clusters (n = 3632 spots). The vertical axis represents gene expression levels. Each violin represents the distribution of expression for a particular gene, with the width indicating frequency. The central white dot denotes the median expression level, the thick black bar within each violin represents the interquartile range (IQR; 25th to 75th percentile), and the thin black line (whiskers) extends to 1.5 × IQR from the 25th and 75th percentiles, capturing the data range excluding outliers. Data represent biological replicates (individual spots).
Interestingly, DEGs included DCN, CCL19, and CCL21, which are known to function in cancer-associated fibroblasts (CAFs)44,45. Moreover, the CAF marker genes (e.g., COL6A1, COL1A2, VIM, PDFGRB, S100A4)46,47,48 were upregulated in Cluster 3 (log2FC > 0.25, Wilcoxon rank-sum test, p-value < 0.05) (Fig. 6e) but not in Cluster 4 (Supplementary Fig. S17a and b).
Taken together, through the integration of multimodality by STAIG, we found that Cluster 3 shapes a tumor microenvironment densely populated by CAFs and revealed the molecular properties of CAF-rich areas.
Identifying tumor-adjacent tissue junctions in zebrafish melanoma ST
To delineate the tumor microenvironment in zebrafish melanoma, we analyzed tissue slices A and B (Fig. 7a). While the original study segmented interface clusters into muscle-like and tumor-like subclusters using scRNA-seq data, standalone ST data analysis proved insufficient for precise cluster identification, especially in smaller regions without integrating scRNA data. In simple slice configurations, such as slice A, all comparative algorithms identified the interface (Supplementary Fig. S18). However, when organ complexity increased, algorithms struggled to accurately demarcate these interfaces (Supplementary Fig. S19). Notably, GraphST, conST, DeepST, and Seurat inaccurately included normal muscle tissue within interface cluster boundaries, and SpaGCN and STAGATE identified larger interface areas. They missed smaller scattered interfaces encircled by the tumor that STAIG accurately aligned with the original study results (Fig. 7b).
a H&E stained image and annotation of zebrafish melanoma on slices A and B from Hunter et al. (2021). b Interface domains identified by STAIG on slices A and B with SC and DB. Baseline methods results are shown in Supplementary Figs. S18 and S19. c DGE analysis of the interface domain versus other domains on slice A. Each point represents a gene, the vertical axis represents the -log10 of the p-value and the horizontal axis represents the log2FoldChange (log2 FC). P-values were derived from a two-sided Wilcoxon rank-sum test. The significance thresholds were set at |log2FC| > 0.25 and p-value < 0.05. d GO analysis for the identified interface domain versus other domains on slice A. The vertical axis represents the GO terms, and the horizontal axis represents the -log10 of the p-value. GO enrichment analysis was performed using the one-sided hypergeometric test, and p-values were adjusted for multiple comparisons using the Benjamini-Hochberg method. e UMAP visualization of clustering results on slices A and B by STAIG. The solid outline indicates clusters of normal tissue, while the dashed outline denotes clusters of tumor tissue. f Fine-grained subregions (muscle-proximal and tumor-proximal) of the interface identified by STAIG and the gene expression patterns of specific genes (ckmb, ckma, BRAFV600E, neb).
Furthermore, we incorporated TESLA49, a tool designed to enhance gene expression matrices to the super-pixel level based on imaging, into our comparative framework for analyzing the tumor microenvironment. When cross-referencing cancer-associated gene lists from original research, TESLA was unable to identify interfaces with the same precision as STAIG. For instance, in slice A, TESLA missed the interface area highlighted by the yellow frame, whereas STAIG successfully recognized it (Supplementary Fig. S20a). In slice B, STAIG also accurately identified the interface areas (Supplementary Fig. S20b). Conversely, TESLA misclassified the interface locations as the core areas of the tumor (the white frame), incorrectly placing them away from the edges, which contradicts the actual observations.
In addition, the DEGs found at the interfaces by STAIG included known genes, such as si:dkey-153m14.1, zgc:158463, RPL41, and hspb9 (Fig. 7c and Supplementary Fig. S21a) and were associated with mRNA metabolic processes (Fig. 7d and Supplementary Fig. S21b). These results highlight the active transcriptional and translational processes in these regions, thus supporting previous findings.
Subsequently, STAIG identified Cluster 12 in slices A and 7 in slice B as being closer to the tumor area, whereas Cluster 5 in slices A and 9 in slice B were positioned nearer to the muscle tissue. As shown in the UMAP plots (Fig. 7e), although Clusters 12 and 5 in slice A were interconnected, Cluster 12 was aligned with tumor-associated clusters, and Cluster 5 had normal tissue clusters. A similar pattern was observed in Clusters 7 and 9 in slice B. Notably, the muscle-proximal subcluster showed high expression of muscle marker genes50,51, such as ckmb and ckma (Fig. 7f and Supplementary Fig. S21c, d), and the tumor-proximal subcluster exhibited high expression of tumor-associated genes52,53, for example, BRAFV600E and neb. These marker genes exhibited differential functional involvement (Supplementary Fig. S21e): the subcluster farther from the tumor showed an enrichment of muscle structure and metabolism, whereas those closer to the tumor were involved in protein folding and responding to temperature changes, which are indicative of increased cellular activity and stress responses typically found in tumor cells54.
In summary, we characterized the tumor borders of zebrafish melanoma and found that the cluster near tumors resembled a tumor-like interface, whereas that closer to normal tissue was akin to a muscle-like interface. These results highlight the necessity for ST annotation and dissection at a higher resolution to enhance our understanding of intricate cancer invasion.
Discussion
In this study, we proposed STAIG, a deep learning model that efficiently integrates gene expression, spatial coordinates, and histological images using graph contrastive learning coupled with high-performance feature extraction. Unlike other methods, our approach first processes images using Gaussian blurring and bandpass filtering, which are essential for eliminating noise and enabling the model to focus on texture rather than color. Therefore, our model successfully extracted spatial features from an entire H&E-stained image without relying on additional image data. These features were processed by graph augmentation along with a unique contrastive loss function to focus on neighbor node consistency, which contributed to improving the performance of STAIG, even in settings without image data.
The batch integration capability of STAIG stems from the generalization power of the contrastive learning framework, enabling the model to extract meaningful information across multiple similar datasets. While a comprehensive theoretical explanation for why sample-level contrastive learning leads to effective self-supervised clustering and generalization is still lacking55, we investigated this phenomenon from the perspective of changes in the positions of samples in the latent space. Specifically, we observed that, in the early stages after dimensionality reduction through GNN and graph augmentation, the latent space exhibited a directional shift between samples from different slices due to batch effects. However, the angular differences between samples of different categories remained consistent across slices. That is, while all samples are initially very close to each other in the latent space, during each iteration, the contrastive learning at the node level ensures that the model focuses on the angular differences between categories rather than the differences introduced by batch effects when processing consecutive slices. This alignment enables samples from the same category, regardless of the slice, to move in similar directions and distances. In subsequent iterations, the model increasingly learns and extracts features relevant to category distinctions, further mitigating the impact of batch effects. Ultimately, samples of the same category, irrespective of their slice origin, cluster together, leading to embeddings that are largely free from batch effects. Detailed data can be found in the Supplementary Materials section titled The principle of STAIG integration in latent space.
We further evaluated the integration capabilities of STAIG by utilizing scDesign356 to generate simulated data with controllable batch effect magnitudes. The detailed results can be found in the supplementary materials under Exploring the integration capacity of STAIG with artificially modified batch effects. The results demonstrate that STAIG can effectively perform integration under varying degrees of batch effects. By leveraging scDesign3, users can assess whether their data falls within STAIG’s integration capacity limits. In addition, we compared the performance of STAIG between joint training of multiple datasets and independent training approaches. The experimental results confirm that the shared weight updating mechanism in GNN is crucial for integration capability (refer to supplementary materials Comparison between whole batch and separate training in STAIG).
By extensively benchmarking various ST datasets, we demonstrated the effectiveness of STAIG by utilizing advanced image feature extraction if histological images were available. In the human breast cancer and zebrafish melanoma datasets, STAIG not only identified spatial regions with high resolution compared to existing studies but also uncovered previously challenging-to-identify regions, offering deeper depth into tumor microenvironments. In addition, STAIG demonstrated strong capability in delineating tumor boundaries, outperforming others in identifying transitional zones with precision. This effectiveness is also linked to the contrast strategy of the STAIG, which captures vital local information.
We acknowledge a potential stability issue in the sampling process of \({G}^{1}\) and \({G}^{2}\). To address this, we implemented a resampling method that generates n augmented graphs and averages their structures to create the final enhanced graph (see Supplementary Materials: Stability and efficiency of the averaged structure). While resampling increases algorithmic complexity, our ablation studies show stable results when n was kept below 10 (see Supplementary Fig. S31d–f). Therefore, we believe that the current methodology is robust across various datasets and scenarios. Nonetheless, this approach may not completely eliminate stability issues in every test or dataset. To mitigate potential biases, we provide users with the option to apply the resampling process to their own datasets, which we anticipate will help reduce any sampling-related bias.
In conclusion, we confirmed the impact of combining advanced machine learning with multimodal spatial information and the promising potential of STAIG in deciphering cellular architectures, which facilitates our understanding of spatial biological intricacies. In future work, because constructing a whole-graph representation is resource-intensive, especially in systems with limited GPU memory, optimizing matrix compression during graph construction and effectively applying mini-batch approaches to optimize GNNs should be further considered.
Methods
Data description
Publicly available ST datasets and histological images acquired from various platforms were downloaded (Supplementary Table S1). From the 10x Visium platform, the DLPFC dataset included 12 sections from three individuals, with each individual contributing four sections sampled at 10 μm and 300 μm intervals. The spot counts for these datasets varied between 3498 and 4789 per section, with the DLPFC dataset featuring annotations for its six layers and white matter. The human breast cancer dataset had 3798 spots, and the mouse brain dataset comprised the anterior and posterior sections with 2695 and 3355 spots, respectively. For zebrafish melanoma, two tissue slices, A and B, with 2179 and 2677 spots, respectively, were analyzed. For the integration experiments, the DLPFC and mouse brain were used.
The Stereo-seq dataset of the mouse olfactory bulb included 19,109 spots with a 14 μm resolution. Slide-seqV2 datasets featured a 10 μm resolution with the mouse hippocampus (18,765 spots from the central quarter radius) and the mouse olfactory bulb (19,285 spots). The STARmap dataset comprised 1207 spots. For the MERFISH datasets, the human MTG contained 3970 spots, while the VIS regions of mouse1 and mouse2 had 5995 and 2479 spots respectively. For the integration experiments, the two mouse VIS regions from the MERFISH datasets, along with the mouse olfactory bulb from both Stereo-seq and Slide-seqV2 were used. These datasets lacked images, and image annotations were downloaded from the Allen Brain Atlas website (Supplementary Table S2).
Data preprocessing
Raw gene expression counts were log-transformed and normalized using the SCANPY57 package. After normalization, the data were scaled to achieve a mean of zero and unit variance. For our experiments, we selected a subset of \(F\) highly variable genes (HVGs), where \(F\) was designated as 3000.
In addition to the SCANPY default VST (variance of standardized counts) method, we provided four categories of gene selection approaches for users to choose from. The first category comprises empirical distribution-based methods58, from which we selected the MVP (mean-variance plot) method that employs Z statistics of mean and variance. The second category encompasses methods for minimizing redundancy in gene selection59,60, where we implemented scGeneClust. The third category is based on co-expression gene networks61,62,63, for which we selected PyWGCNA as our implementation. The fourth category focuses on spatially variable gene selection methods14,64, where we employed SpatialDE. For detailed performance evaluation, please refer to the supplementary materials section STAIG’s performance with different feature gene selection methods.
Basic graph construction
For each slice, we constructed a basic undirected neighborhood graph, \(G=\left(V,E\right)\), where \(V\) represents the set of \(N\) spots \(\{{v}_{1},{v}_{2},\ldots {v}_{N}\}\), and \(E\subseteq V\times V\) denotes the edges between these spots. The corresponding adjacency matrix, \(A \in \{{0,1}\}^{N\times N}\), is defined such that \({A}_{{ij}}=1\) indicates the presence of an edge \(\left({v}_{i},{v}_{j}\right)\in E\). We determined the edges by computing a distance matrix \(D\in {{\mathbb{R}}}^{N\times N}\), with \({D}_{{ij}}\) representing the Euclidean distance between any two spots \({v}_{i}\) and \({v}_{j}\). KNN was employed to connect each spot with its top \(k\) closest spots, forming neighboring nodes. The gene expression matrix \(X\in {{\mathbb{R}}}^{N\times F}\) was also established, where each row corresponds to the expression profile of HVGs for each spot.
Histological image feature extraction
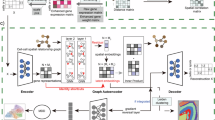
For the ST data with H&E-stained slices, extracting image features from each spot was critical. We began by applying Gaussian blurring to the entire H&E-stained image using a 7 × 7 Gaussian kernel (see Supplementary Material, Gaussian blur parameter selection) to reduce texture noise and staining biases. Then, similar to HistToGene65, we cropped the images into patches centered on the spot coordinates. The size of the patches was based on a dataset-specific parameter, fiducial_diameter_fullres (denoted as d), which is the pixel diameter of a fiducial spot in the full-resolution image. According to our tests, the default patch size was set to 3.5 d (see Supplementary Material, Patch size selection). Subsequently, we resized the patches to 512 × 512. These N patches then underwent a band-pass filter to highlight the cellular nuclear morphology. The default parameters for the band-pass were 245–275 (see Supplementary Material, Bandpass filter parameters selection).
We utilized BYOL18 with a ResNet5066 backbone for advanced self-supervised image feature extraction due to its ability to learn robust features without the need for negative samples, ideal for spatial transcriptomics where class labels are absent. Unlike BYOL, U-Net, traditionally employed for tasks like cell nucleus counting and segmentation, requires supervised learning with annotated masks and is not designed to extract embeddings. This limitation makes U-Net67 unsuitable for integrating complex biological information from HE images in spatial transcriptomics, which goes beyond simple cellular structures.
During the initialization phase of BYOL, we employed the official PyTorch-pretrained ResNet50 weights as the initial model without any further fine-tuning on additional datasets. In BYOL, the images were resized to 224 × 224 pixels and subjected to two random augmentations68: horizontal flipping, color distortion, Gaussian blurring, and solarization. These images were fed into two networks: the online network, which comprises an encoder, projector, and predictor, and the target network, which is similar but lacks the predictor. Both networks utilized the ResNet architecture for encoding and multilayer perceptrons for projecting and predicting. Embeddings from these networks were synchronized using an L2 loss to ensure consistency.
After training, the encoder of the online network produced a 2048-dimensional feature vector for each image, which was reduced to 64 components using Principal Component Analysis69. The resulting matrix \(C\in {{\mathbb{R}}}^{N\times 64}\) represented the image features, with each row \({c}_{i}\) corresponding to the features of spot \({v}_{i}\).
Integration of slices
Each slice was processed individually while dealing with multiple tissue slices. At the initial stage of integration, we adopted the same approach as STAligner to individually process each slice. The raw gene expression counts of each slice were log-transformed and normalized, and then the data were scaled to achieve a mean of zero and unit variance. We also selected a default of 5000 highly variable genes (HVGs) for each slice. Subsequently, we took the intersection of all the HVGs as the final set of common \(F\) HVGs. After filtering for the common HVGs in each slice, we proceeded to the next step of integration.
Given m slices with the respective numbers of spots \({N}_{1},{N}_{2},\ldots,{N}_{m}\), the total spot count \(T\) is the sum of all \({N}_{i}\). We then obtained distinct adjacency matrices \({A}^{1}\) to \({A}^{m}\) and image feature matrices \({C}^{1}\) to \({C}^{m}\) for each slice. Batch integration commenced with the vertical concatenation of common gene expression into a single matrix \({{{\mathcal{X}}}}\in {{\mathbb{R}}}^{T\times F}\).
In the next step, we integrate these matrices. The adjacency matrices were combined into a block-diagonal matrix \({{{\mathcal{A}}}}\), as shown in Eq. (1), and the image feature matrices were vertically concatenated to form matrix \({{{\mathcal{C}}}}\), as shown in Eq. (2): These integrated matrices formed the basis for further analysis.
During this process, each spot \({v}_{i}\) was assigned a slice identifier \(s({v}_{i})\), indicating the specific slice to which the spot belongs.
Graph augmentation
The contrastive learning workflow in the STAIG is based on the GCA framework70. Each iteration augmented the input graph, resulting in two modified graphs, \({G}^{1}\) and \({G}^{2}\), and their associated gene expression matrices, \({{{{\mathcal{X}}}}}^{1}\) and \({{{{\mathcal{X}}}}}^{2}\). These were derived from the original graph \(G\), which possessed an edge set \({{{\mathcal{E}}}}\), through a specific augmentation strategy applied to the integrated adjacency matrix \({{{\mathcal{A}}}}\) and gene expression matrix \({{{\mathcal{X}}}}\).
Utilizing the integrated image feature matrix \({{{\mathcal{C}}}}\), we computed the image spatial distance matrix \({D}^{{img}}\in {R}^{T\times T}\), with \({D}_{{ij}}^{{img}}\) indicating the Euclidean distance between spots \({v}_{i}\) and \({v}_{j}\) in the image feature space (Supplementary material, Methods for generating the image distance matrix). The edge weight matrix \(W\) is defined \({D}^{{img}}\) as expressed in Eq. (3).
\(W\) was then scaled to a probability matrix \(P\in {{\mathbb{R}}}^{T\times T}\), with each element \({P}_{i,j}\) being calculated as per Eq. (4):
During each iteration, the augmented graphs \({G}^{1}\) and \({G}^{2}\) maintained the node set \(V\) of original graph \(G\). However, their sets of edges, \({{{{\mathcal{E}}}}}^{1}\) and \({{{{\mathcal{E}}}}}^{2}\) were independently formed by probabilistically removing each edge from \({{{\mathcal{E}}}}\) (for example, an edge \(\left({v}_{i},{v}_{j}\right)\in {{{\mathcal{E}}}}\)), based on the probability \({p}_{{ij}}^{e}\) derived from \(P\).
In scenarios lacking image data, STAIG offers two methods of edge removal: fixed edge removal with a set probability, and adaptive edge removal based on gene similarity. For fixed edge removal, STAIG’s default parameter \(p\) is set to 0.2. When performing adaptive edge removal based on gene similarity, we proceed as follows: PCA is used to reduce the preprocessed gene expression data of each spot to 64 dimensions, and this reduced data is then used to compute the cosine similarity between spots.
The matrix of cosine similarities between spots is directly used as the edge weight matrix \(W\) in the computation of Eq. (4). For a comparison of the effects of random edge deletion and adaptive edge deletion based on gene similarity, as well as the choice of distance formula for gene similarity, see the supplementary material Comparing Adaptive Edge Perturbation Based on Gene Similarity with Random Perturbation. In the paper, we conducted tests using random edge deletion.
The gene expression matrix \({{{\mathcal{X}}}}\) was augmented by introducing variabilities such as salt-and-pepper noise into the images. This involves two independent random vectors, \({r}^{1}\) and \({r}^{2}\), each within \({\{{{\mathrm{0,1}}}\}}^{F}\). The elements in the vectors are sampled using a Bernoulli distribution. STAIG’s default parameter for this is set at 10% (see Supplementary material, Masked ratio of features in graph augmentation). The augmented matrices \({{{{\mathcal{X}}}}}^{1}\) and \({{{{\mathcal{X}}}}}^{2}\) are then formulated by multiplying each row \({x}_{i}\) in \({{{\mathcal{X}}}}\) by \({r}^{1}\) and \({r}^{2}\) respectively, as described in Eq. (5).
Here, \({x}_{i}^{1}\) and \({x}_{i}^{2}\) corresponded to the respective rows in \({{{{\mathcal{X}}}}}^{1}\) and \({{{{\mathcal{X}}}}}^{2}\). The symbol \(\odot\) represented the element-wise multiplication.
Contrastive learning in the STAIG framework based on the GCA architecture
In every iteration, \({G}^{1}\) and \({G}^{2}\) are processed through a shared GNN structured with \(l\) layers of GCNs, followed by two layers of fully connected networks, yielding embeddings \({H}^{1}\) and \({H}^{2}\) for each graph as follows:
where \({F}^{{\prime} }\ll F\), and the rows \({h}_{i}^{1}\) and \({h}_{i}^{2}\) in \({H}^{1}\) and \({H}^{2}\) represented the embeddings of spot \({v}_{i}\) in the augmented graphs \({G}^{1}\) and \({G}^{2}\), respectively.
In the STAIG model, neighbor contrastive learning loss71 was employed to ensure the consistency and distinctiveness of the embeddings in \({H}^{1}\) and \({H}^{2}\). This involved constructing and comparing positive and negative sample pairs.
For positive pairs, we considered the embedding \({h}_{i}^{1}\) of \({v}_{i}\) in \({G}^{1}\) as an example. Positive pairs were sourced from: (1) The embedding \({h}_{i}^{2}\) of the same spot \({v}_{i}\) in \({G}^{2}\); (2) embeddings of neighboring nodes of \({v}_{i}\) in \({G}^{1}\), denoted as \({{h}_{j}^{1}{|v}}_{j}\in {{{{\mathcal{N}}}}}_{i}^{1}\), where \({{{{\mathcal{N}}}}}_{i}^{1}\) represented the set of neighboring nodes of \({v}_{i}\) in \({G}^{1}\); and (3) embeddings of neighboring nodes of \({v}_{i}\) in \({G}^{2}\), expressed as \({{h}_{j}^{2}{|v}}_{j}\in {{{{\mathcal{N}}}}}_{i}^{2}\), with \({{{{\mathcal{N}}}}}_{i}^{2}\) being the set of neighbors of \({v}_{i}\) in \({G}^{2}\). The total number of positive pairs for anchor \({h}_{i}^{1}\) is \({\hat{{{{\mathcal{N}}}}}}_{i}=|{{{{\mathcal{N}}}}}_{i}^{1}|+|{{{{\mathcal{N}}}}}_{i}^{2}|+1\), where \(|{{{{\mathcal{N}}}}}_{i}^{1}|\) and \(|{{{{\mathcal{N}}}}}_{i}^{2}|\) denote the counts of neighboring nodes of \({v}_{i}\) in \({G}_{1}\) and \({G}_{2}\), respectively.
By contrast, negative sample pairs are generally defined as all pairs other than the positive ones. However, selecting negative samples without considering class information led to a sampling bias19, where many negative samples were incorrectly classified as belonging to the same class. This negatively affected the clustering results. To address this issue, when the images were available, we implemented the DS19. DS filters out false negatives based on the similarity of the image features \({c}_{i}\) of each spot \({v}_{i}\). Initially, the spots were clustered into \(Q\) classes using the k-means method based on their image features, creating pseudo-labels \({Y}_{p}=\{{y}_{i}\}_{i=1}^{T}\). During the construction of negative samples, any \({v}_{i}\) sharing the same pseudo-label were excluded. The default parameter for \(Q\) is 40 (see Supplementary Materials, ‘Cluster Number in Debiased Strategy’).”
The neighbor contrastive loss associated with anchor \({h}_{i}^{1}\) between \({G}^{1}\) and \({G}^{2}\) is given by
here, \({f}^{{{\mathrm{1,1}}}}\left(i,j\right)={e}^{\theta \left({h}_{i}^{1},{h}_{j}^{1}\right)/\tau }\), \({f}^{{{\mathrm{1,2}}}}\left(i,j\right)={e}^{\theta \left({h}_{i}^{1},{h}_{j}^{2}\right)/\tau }\), \(\tau\) was a temperature parameter, and \({{{\rm{\theta }}}}\left(\cdot \right)\) represented a similarity measure (in our work, the inner product is employed). The first term of the denominator is an inter-graph positive pair for the same node. The last two terms in the denominator of Eq. (7) can be decomposed as:
where nodes that are not directly connected to \({v}_{i}\) in both \({G}^{1}\) and \({G}^{2}\) are considered as negative pairs within the same augmented graphs (intra-graph) and across different augmented graphs (inter-graph), respectively. By minimizing Eq. (7), the objective is to enhance the alignment among positive pairs while reducing the similarity among negative pairs.
For the task of integration, we have made some modifications to Eq. (7), specifically limiting the selection of negative samples to within the same slice while avoiding the DS due to the batch effects present in image features from different slices. Details regarding how each component of the loss function contributes to integration are provided in the supplementary materials ‘Ablation studies for integration’. The neighbor contrastive loss is described as follows:
Given the structural similarity between \({G}^{1}\) and \({G}^{2}\), computation of \(l\left({h}_{i}^{2}\right)\) mirrored that of \(l\left({h}_{i}^{1}\right)\). The final neighbor contrastive loss averaged over both augmented graphs is expressed as
Spatial domain identification through clustering and refinement
Upon completion of training, the spot embeddings in both augmented graphs were averaged to derive the final low-dimensional representation \(H\) given as follows:
The \(H\) was clustered using the mclust72 algorithm based on a Gaussian Mixture Model. For datasets with manual annotations, the number of clusters was set to match the number of labeled classes. In the absence of such annotations, SC determines the number of clusters, with the count corresponding to the highest SC designated as the final number for the spatial domain.
Consistent with previous baseline methods, we employed a refinement technique for the DLPFC dataset20 to further mitigate noise and ensure smoother delineation of clustering boundaries. As part of this refinement, for a given \({v}_{i}\), all spots within a radius \(d\) in its spatial vicinity were assigned to the same class as \({v}_{i}\).
Regarding multi-slice integration, because all data were amalgamated into a single matrix prior to model input, the embedding of spots across different slices outputted by the GNN eliminated batch differences. The procedure for spatial identification mirrored that of single-slice processing.
The overall architecture of STAIG
The architecture of the STAIG features a GNN with one GCN layer and two layers of a fully connected network, producing a 64-dimensional \(F^{\prime}\). It employed a Parametric Rectified Linear Unit activation function and an Adam optimizer with an initial learning rate of \(5\times {10}^{-4}\) and a weight decay of \(1\times {10}^{-5}\). The model ran for 400 epochs, with a temperature parameter τ set to 10 and a neighbor count \(k\) of 5 by default. Further details on hyperparameter optimization can be found in the Hyperparameter optimization on 10x Visium and Hyperparameter optimization on other platforms of the Supplementary Information. STAIG was tested on an NVIDIA A100 GPU installed in a Linux system with an AMD EPYC 7742 CPU and 64GB of memory. Detailed runtime costs are provided in Supplementary Table S3.
The ablation studies of STAIG
To assess the effectiveness of the STAIG based solely on its novel contrastive learning framework and the efficacy of the graph augmentation strategy utilizing image information, we conducted ablation experiments on the DLPFC dataset. The experimental setup included a basic framework without image data, using a fixed graph edge removal probability and DS with image data, employing only adaptive random edge removal based on image features, and implementing a complete framework.
The results demonstrated the consistency of STAIG’s superiority in the median ARI compared with other methods, even in the absence of image data. Both the adaptive random edge removal strategy and DS were grounded in significantly improved average and median ARI and NMI values. Comprehensive details of these ablation studies are provided in the ‘Ablation studies on the overall framework’ of the Supplementary Information.
In addition, we conducted ablation studies on STAIG’s image feature extraction phase to validate the effectiveness of the filtering process and the BYOL framework. Detailed information can be found in the Supplementary Material, ‘BYOL ablation study for histology feature extraction’.
Visualization and functional analysis of spatial domain
The detected spatial domains were visualized using UMAP. Differential gene expression was analyzed using the Wilcoxon test for SCANPY. Gene ontology analysis was conducted using the clusterProfiler73 package, and the statistical tests were corrected using the Benjamini-Hochberg (BH) method. For differentially expressed gene analysis, |log2FC | > 0.25 and p-value < 0.05 were used as the marker gene selection thresholds. P-values were calculated using the Wilcoxon rank-sum test. A comprehensive discussion regarding the selection of the log2FC threshold is provided in the Supplementary Materials.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets examined in our study are publicly available and can be accessed in their unprocessed form directly from the original sources, as detailed in Supplementary Tables S1 and S2. The data used in this study are available in the Zenodo under accession code [https://doi.org/10.5281/zenodo.10277127].
Code availability
The source code for STAIG is openly available for academic and noncommercial use. This information can be accessed from the following GitHub repository74: [https://github.com/y-itao/STAIG]. This repository includes all necessary instructions for installation, usage, and a detailed README for guidance. It is also deposited at Zenodo [https://doi.org/10.5281/ZENODO.14238885].
References
Rao, N., Clark, S. & Habern, O. Bridging genomics and tissue pathology: 10x Genomics explores new frontiers with the Visium Spatial Gene Expression Solution. Gen. Eng. Biotechnol. News 40, 50–51 (2020).
Rodriques, S. G. et al. Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Chen, A. et al. Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185, 1777–1792 (2022).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Dries, R. et al. Advances in spatial transcriptomic data analysis. Genome Res. 31, 1706–1718 (2021).
Hartigan, J. A. & Wong, M. A. Algorithm AS 136: A K-means clustering algorithm. Appl. Stat. 28, 100 (1979).
Traag, V. A., Waltman, L. & Van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Xu, H. et al. Unsupervised spatially embedded deep representation of spatial transcriptomics. Genome. Med. 16, 12 (2024).
Dong, K. & Zhang, S. Deciphering spatial domains from spatially resolved transcriptomics with an adaptive graph attention auto-encoder. Nat. Commun. 13, 1739 (2022).
Long, Y. et al. Spatially informed clustering, integration, and deconvolution of spatial transcriptomics with GraphST. Nat. Commun. 14, 1155 (2023).
Zong, Y. et al. conST: An interpretable multi-modal contrastive learning framework for spatial transcriptomics. Preprint at https://doi.org/10.1101/2022.01.14.476408 (2022).
Pham, D. et al. Robust mapping of spatiotemporal trajectories and cell–cell interactions in healthy and diseased tissues. Nat. Commun. 14, 7739 (2023).
Hu, J. et al. SpaGCN: Integrating gene expression, spatial location and histology to identify spatial domains and spatially variable genes by graph convolutional network. Nat. Methods 18, 1342–1351 (2021).
Xu, C. et al. DeepST: identifying spatial domains in spatial transcriptomics by deep learning. Nucleic Acids Res. 50, e131–e131 (2022).
He, K. et al. Masked Autoencoders Are Scalable Vision Learners. in 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 15979–15988 https://doi.org/10.1109/CVPR52688.2022.01553 (IEEE, New Orleans, LA, USA, 2022).
Zeira, R., Land, M., Strzalkowski, A. & Raphael, B. J. Alignment and integration of spatial transcriptomics data. Nat Methods 19, 567–575 (2022).
Grill, J.-B. et al. Bootstrap your own latent a new approach to self-supervised learning. In Proceedings of the 34th International Conference on Neural Information Processing Systems (Curran Associates Inc., Red Hook, NY, USA, 2020).
Zhao, H., Yang, X., Wang, Z., Yang, E. & Deng, C. Graph debiased contrastive learning with joint representation clustering. In Proceedings of the Thirtieth International Joint Conference on Artificial Intelligence 3434–3440 (International Joint Conferences on Artificial Intelligence Organization, Montreal, Canada, 2021).
Maynard, K. R. et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 24, 425–436 (2021).
Zhang, L., Liang, S. & Wan, L. A multi-view graph contrastive learning framework for deciphering spatially resolved transcriptomics data. Brief. Bioinform. 25, bbae255 (2024).
Steinley, D. Properties of the hubert-arable adjusted rand index. Psychol. Methods 9, 386 (2004).
Estevez, P. A., Tesmer, M., Perez, C. A. & Zurada, J. M. Normalized mutual information feature selection. IEEE Trans. Neural Netw. 20, 189–201 (2009).
Shahapure, K. R. & Nicholas, C. Cluster quality analysis using silhouette score. In 2020 IEEE 7th international conference on data science and advanced analytics (DSAA) 747–748 (IEEE, 2020).
Vergani, A. A. & Binaghi, E. A soft davies-bouldin separation measure. In 2018 IEEE International Conference on Fuzzy Systems (FUZZ-IEEE) 1–8 (IEEE, Rio de Janeiro, 2018).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Keller, J. M., Gray, M. R. & Givens, J. A. A fuzzy K-nearest neighbor algorithm. IEEE Trans. Syst. Man Cybern. SMC-15, 580–585 (1985).
Seroogy, K. B., Brecha, N. & Gall, C. Distribution of cholecystokinin‐like immunoreactivity in the rat main olfactory bulb. J. Comp. Neurol. 239, 373–383 (1985).
Farmer, W. T. & Murai, K. Resolving astrocyte heterogeneity in the CNS. Front. Cell. Neurosci. 11, 300 (2017).
Deans, M. R. et al. Control of neuronal morphology by the atypical cadherin Fat3. Neuron 71, 820–832 (2011).
Paul, A., Chaker, Z. & Doetsch, F. Hypothalamic regulation of regionally distinct adult neural stem cells and neurogenesis. Science 356, 1383–1386 (2017).
Tabar, V. et al. Migration and differentiation of neural precursors derived from human embryonic stem cells in the rat brain. Nat. Biotechnol. 23, 601–606 (2005).
Mamoor, S. The α1 subunit of the γ-aminobutyric acid receptor, Gabra1, is differentially expressed in the brains of patients with schizophrenia. Preprint at https://doi.org/10.31219/osf.io/m93ya (2020).
Stickels, R. R. et al. Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol. 39, 313–319 (2021).
Biesemann, C. et al. Proteomic screening of glutamatergic mouse brain synaptosomes isolated by fluorescence activated sorting. EMBO J. 33, 157–170 (2014).
Calì, T., Brini, M. & Carafoli, E. The PMCA pumps in genetically determined neuronal pathologies. Neurosci. Lett. 663, 2–11 (2018).
Iijima, T., Miura, E., Watanabe, M. & Yuzaki, M. Distinct expression of C1q‐like family mRNAs in mouse brain and biochemical characterization of their encoded proteins. Eur. J. Neurosci. 31, 1606–1615 (2010).
Takeda, K. WFS1 (Wolfram syndrome 1) gene product: predominant subcellular localization to endoplasmic reticulum in cultured cells and neuronal expression in rat brain. Hum. Mol. Genet. 10, 477–484 (2001).
Newrzella, D. et al. The functional genome of CA1 and CA3 neurons under native conditions and in response to ischemia. BMC Genomics 8, 370 (2007).
Fang, R. et al. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56–62 (2022).
Li, X. et al. Deep learning enables accurate clustering with batch effect removal in single-cell RNA-seq analysis. Nat. Commun. 11, 2338 (2020).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Zhou, X., Dong, K. & Zhang, S. Integrating spatial transcriptomics data across different conditions, technologies and developmental stages. Nat. Comput. Sci. 3, 894–906 (2023).
Bockstal, M. V. et al. Differential regulation of extracellular matrix protein expression in carcinoma-associated fibroblasts by TGF-β1 regulates cancer cell spreading but not adhesion. Oncoscience 1, 634–648 (2014).
Ozga, A. J., Chow, M. T. & Luster, A. D. Chemokines and the immune response to cancer. Immunity 54, 859–874 (2021).
Han, C., Liu, T. & Yin, R. Biomarkers for cancer-associated fibroblasts. Biomark. Res. 8, 64 (2020).
Zhang, H. et al. Define cancer-associated fibroblasts (CAFs) in the tumor microenvironment: new opportunities in cancer immunotherapy and advances in clinical trials. Mol. Cancer 22, 159 (2023).
Kay, E. J. et al. Cancer-associated fibroblasts require proline synthesis by PYCR1 for the deposition of pro-tumorigenic extracellular matrix. Nat. Metab. 4, 693–710 (2022).
Hu, J. et al. Deciphering tumor ecosystems at super resolution from spatial transcriptomics with TESLA. Cell Syst. 14, 404–417.e4 (2023).
Jin, S. et al. Ebf factors and MyoD cooperate to regulate muscle relaxation via Atp2a1. Nat. Commun. 5, 3793 (2014).
Bayir, M., Arslan, G. & Oğuzhan Yildiz, P. Characterization, identification and phylogeny of the creatine kinase (ckma) gene in medaka (Oryzias latipes). Mar. Sci. Technol. Bull. 9, 15–22 (2020).
Ascierto, P. A. et al. The role of BRAF V600 mutation in melanoma. J. Transl. Med. 10, 85 (2012).
Ohnishi, Y., Tajima, S. & Ishibashi, A. Coordinate expression of membrane type-matrix metalloproteinases-2 and 3 (MT2-MMP and MT3-MMP) and matrix metalloproteinase-2 (MMP-2) in primary and metastatic melanoma cells. Eur. J. Dermatol. 11, 420–423 (2001).
Wang, M., Law, M. E., Castellano, R. K. & Law, B. K. The unfolded protein response as a target for anticancer therapeutics. Crit. Rev. Oncol. Hematol. 127, 66–79 (2018).
Huang, W., Yi, M., Zhao, X. & Jiang, Z. Towards the generalization of contrastive self-supervised learning. In The Eleventh International Conference on Learning Representations, ICLR 2023, Kigali, Rwanda, May 1-5, 2023 (OpenReview.net, 2023).
Song, D. et al. scDesign3 generates realistic in silico data for multimodal single-cell and spatial omics. Nat. Biotechnol. 42, 247–252 (2023).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Chen, H.-I. H., Jin, Y., Huang, Y. & Chen, Y. Detection of high variability in gene expression from single-cell RNA-seq profiling. BMC Genomics 17, 508 (2016).
Deng, T. et al. A cofunctional grouping-based approach for non-redundant feature gene selection in unannotated single-cell RNA-seq analysis. Brief. Bioinform. 24, bbad042 (2023).
Ding, C. & Peng, H. Minimum redundancy feature selection from microarray gene expression data. J. Bioinform. Comput. Biol. 03, 185–205 (2005).
Zhang, T. & Wong, G. Gene expression data analysis using Hellinger correlation in weighted gene co-expression networks (WGCNA). Comput. Struct. Biotechnol. J. 20, 3851–3863 (2022).
Rezaie, N., Reese, F. & Mortazavi, A. PyWGCNA: a Python package for weighted gene co-expression network analysis. Bioinformatics 39, btad415 (2023).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Svensson, V., Teichmann, S. A. & Stegle, O. SpatialDE: identification of spatially variable genes. Nat. Methods 15, 343–346 (2018).
Pang, M., Su, K. & Li, M. Leveraging information in spatial transcriptomics to predict super-resolution gene expression from histology images in tumors. Preprint at https://doi.org/10.1101/2021.11.28.470212 (2021).
Wu, Z., Shen, C. & Van Den Hengel, A. Wider or deeper: Revisiting the ResNet model for visual recognition. Pattern Recognit. 90, 119–133 (2019).
Falk, T. et al. U-Net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2019).
Punn, N. S. & Agarwal, S. BT-Unet: A self-supervised learning framework for biomedical image segmentation using barlow twins with U-net models. Mach. Learn. 111, 4585–4600 (2022).
Wold, S., Esbensen, K. & Geladi, P. Principal component analysis. Chemom. Intell. Lab. Syst. 2, 37–52 (1987).
Zhu, Y. et al. Graph contrastive learning with adaptive augmentation. In Proceedings of the Web Conference 2021 2069–2080 (ACM, Ljubljana Slovenia, 2021).
Shen, X., Sun, D., Pan, S., Zhou, X. & Yang, L. T. Neighbor contrastive learning on learnable graph augmentation. AAAI 37, 9782–9791 (2023).
Scrucca, L., Fop, M., Murphy, T. B. & Raftery, A. E. mclust 5: clustering, classification and density estimation using Gaussian finite mixture models. R J. 8, 289 (2016).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Yang, Y. et al. STAIG: Spatial transcriptomics analysis via image-aided graph contrastive learning for domain exploration and alignment-free integration. STAIG https://doi.org/10.5281/ZENODO.14238885 (2024).
Acknowledgements
Computational resources were provided by the supercomputer system SHIROKANE at the Human Genome Center, the Institute of Medical Science, the University of Tokyo. This study was supported by the JSPS KAKENHI [22K06189 to K.N., JP22K21301 to M.L., and 20H05940 to S.P.] and the JST SPRING [JPMJSP2108 to X.Z.].
Author information
Authors and Affiliations
Contributions
Y. Y. and K. N. conceived and designed the study. Y.Y. and Y.C. performed the data analysis. Y.Y., Y.C., and K.N. drafted the manuscript. S.P., M.L., X.Z., and Y.Z. provided guidance for data analysis. All the authors have read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Zhi-Jie Cao, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Y., Cui, Y., Zeng, X. et al. STAIG: Spatial transcriptomics analysis via image-aided graph contrastive learning for domain exploration and alignment-free integration. Nat Commun 16, 1067 (2025). https://doi.org/10.1038/s41467-025-56276-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56276-0