Abstract
Recurrent respiratory papillomatosis (RRP) is a chronic airway disease caused by Human Papillomavirus (HPV). INO-3107, DNA immunotherapy designed to elicit T-cells against HPV-6 and HPV-11, was evaluated in a 52-week Phase 1/2 study for efficacy, safety, and immunogenicity (NCT04398433). Thirty-two eligible adults with HPV-6 and/or HPV-11 RRP, requiring ≥2 surgical interventions in the year preceding dosing were enrolled between October 2020 and November 2021 and administered 4 INO-3107 doses by intramuscular injection followed by electroporation. The primary endpoint was safety and tolerability, as assessed by treatment-emergent adverse events (TEAEs). Secondary endpoints included surgical intervention frequency and change in RRP Severity Score (modified) post-INO-3107 and assessment of immune responses. 81% (26/32) of patients experienced surgery reduction following INO-3107 compared with the year prior to treatment. Blood assessments revealed HPV-6 and HPV-11 antigen-specific T-cell induction. RNA sequencing identified an inflammatory response in papillomas, inclusive of cytolytic CD8 + T-cell signatures. T-cell receptor sequencing revealed emergent T-cell clones in blood and confirmed trafficking to papillomas. Treatment-related adverse events (AEs) were reported in 13/32 (41%) patients, all low-grade. INO-3107 provides clinical benefit to HPV-6 and/or HPV-11-associated RRP adults and is well-tolerated. Importantly, treatment-induced peripheral T-cell responses traffic to airway tissue and are associated with clinical response.
Similar content being viewed by others
Introduction
Human Papillomavirus (HPV)-associated diseases are an emerging global epidemic1,2. The role of HPV-6 and HPV-11 in the etiology of recurrent respiratory papillomatosis (RRP), the most common benign tumor of the laryngeal epithelium, is well-established3,4. RRP lesions can occur in the oropharynx, larynx, and elsewhere in the respiratory tract5, significantly impacting voice production and even breathing in severe cases. Rarely, papillomas can undergo malignant transformation in the pulmonary tract, most notably in the pulmonary parenchyma6. RRP is a chronic, debilitating disease that is associated with substantial patient burden. Repeated surgical removal of papillomas for symptomatic management remains the mainstay of treatment and can result in significant morbidity considering the number of surgical procedures required, impact on voice, effects of repetitive anesthesia, as well as psychological impact, and financial burden7. Additionally, the risk of laryngeal injury increases with each surgery8.
Current treatment of HPV-6- and HPV-11-related RRP could be improved with the addition of HPV-specific immunotherapy, which has the therapeutic potential to control or eliminate papillomas by generating immunity against HPV itself. Patients suffering from RRP have been characterized as exhibiting suppressive or regulatory immune-related functions within papillomas, as opposed to mounting a cytotoxic response against HPV infected cells9,10,11,12,13. Our previous studies of DNA immunotherapies directed against HPV have demonstrated the induction of potent immune responses for HPV in a pilot trial of adults with HPV-6-related RRP and in HPV-related head and neck squamous cell carcinoma as well as cervical disease, inclusive of the generation of HPV-specific cytotoxic T-cells as well as CD8+ immune cell infiltration into tumors14,15,16,17,18. Additionally, other groups have conducted clinical studies of RRP treatment from an immunological approach via antibody and T-cell-related platforms, with results that support an immune-based impact on disease state19,20. INO-3107 is a DNA immunotherapy encoding for antigens from both HPV-6 and HPV-11 as well as encoding for an interleukin-12 (IL-12) immune adjuvant. Here, we report full study results from a DNA-based therapy in development for RRP from a completed Phase 1/2 trial demonstrating the safety, efficacy, and immune responses of INO-3107.
Results
Patient demographics
A total of 32 patients were enrolled between October 2020 and November 2021, with follow-up completed through December 2022; seven (7) patients had juvenile-onset (JO) RRP and 25 had adult-onset (AO) RRP (Table 1). All patients completed all clinical trial visits up to Week 52. Demographics of the trial population are presented in Table 1. Sixteen (16, 50.0%) patients were infected with HPV-6 only, ten (10, 31.3%) were infected with HPV-11 only, and two (2, 6.3%) were co-infected with both HPV-6 and HPV-11. Four (4, 12.5%) patients’ genotypes could not be distinguished between HPV-6 and HPV-11 per assay standards at Screening but were known from prior diagnostic history to be HPV-6 and/or HPV-11 positive, which was subsequently confirmed through on-study assessment.
All patients in the trial had complete follow-ups for efficacy and safety data.
INO-3107 reduces or eliminates surgical intervention in the treatment of RRP
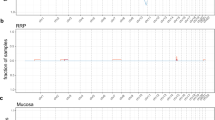
Of the 32 patients, 26 had a clinical response (81.3%), defined as a reduction of at least one surgical procedure in the year following Day 0 (overall clinical response; OCR). Nine patients (28.1%) had a complete response (CR), defined as no surgical interventions in the year following Day 0, and 14 additional patients (43.8%) had a partial response (PR), defined as a reduction in the number of surgical interventions by 50–99% in the year following Day 0 (Fig. 1). Together this represents an overall response rate (ORR) of 71.9%; 23 of 32 patients. Reduction in surgical frequency was independent of HPV genotype as responses were noted for patients exhibiting HPV-6 and/or HPV-11 per genotypic assessment (ORR of 66.7% for HPV-6, ORR of 83.4% for HPV-11).
A Waterfall plot indicating a change in the frequency of clinically indicated surgical interventions during the year of study as compared with the year before. Change in the number of interventions is indicated for each individual patient, and responding patients are color-coded. Complete responders (CR) are labeled dark blue, partial responders (PR) are labeled light blue, and patients who exhibited a decrease in surgical intervention but contributed only to the calculation of overall clinical response (OCR) are green. Non-responders are labeled gray (increase in surgery) or with 0 (no change). B Top left panel: Airway pictures taken before treatment with INO-3107 or after the completion of treatment. The top row is a complete responder before treatment and at Week 52, the middle and bottom rows are partial responders taken before treatment and Week 26. Top right panel: Bar graphs indicating efficacy breakouts for patients infected with HPV-6, patients infected with HPV-11, and all patients in the study. Complete (dark blue) and partial response (light blue) are separated within the overall response rate (ORR) stack for clarity. OCR (all patients exhibiting a decrease in surgical frequency of any magnitude in green) is graphed separately to allow for comparison. Bottom panel: Responses are also broken out in tabular fashion. HPV human papillomavirus; % percent.
When comparing the median number of surgeries in the year prior to treatment with the year following Day 0, in the year prior, patients had a median of four surgeries (range 2–8), while in the year following treatment, there was a median decrease of three surgeries with a 95% confidence interval of 2 to 3, which indicates a statistically significant decrease.
INO-3107 improves RRP severity scores in adults with RRP
Treatment with INO-3107 resulted in reductions in RRP Severity Score (modified) from pretreatment through the treatment assessment period, as assessed by laryngoscopy throughout the trial. Anatomical disease severity was measured by a modified Derkay Score. Scores were obtained prior to surgery at or near Day 0, and again at each visit when laryngoscopy was performed. Between Day 365 and Day 0, there was a mean change in total site score of −10.7 (95% CI −16.3, −5.0) and a mean change in total symptom score of −0.4 (95% CI −0.7, 0.0). The mean change in total clinical score (total site and total symptom score) was −11.0 (95% CI −16.7, −5.3).
INO-3107 is well tolerated in adults with RRP
Treatment with INO-3107 was well tolerated in this trial. Twenty patients (62.5%) reported treatment-emergent adverse events (TEAEs), most of which were treatment-related (35/58 events), and almost all of which were Common Terminology Criteria for Adverse Events (CTCAE) Grade 1 or 2 in severity (54/58 events (Table 2)). All related TEAEs were Grade 1 or 2 in severity, and the most common related TEAE was injection site pain (10 patients); all other related TEAEs were seen in three patients or fewer (Table 2).
Grade 3 TEAEs were observed in four (4) patients (12.5%), all of which were unrelated. Serious adverse events (SAE) were observed in three (3) patients (9.4%), none of which were considered treatment-related. No patient experienced a Grade 4 or 5 TEAE or TEAE leading to treatment discontinuation or death.
INO-3107 induces HPV-6 and HPV-11-specific T-cell responses and drives peripheral T-cell clonal expansion
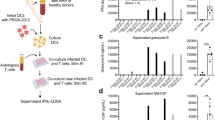
T-cell responses induced by INO-3107 were first gauged via ex vivo interferon gamma (IFNγ) ELISpot. T-cell reactivity against INO-3107 antigens was noted in 94% of patients (30/32) over the course of the trial and reactivity was confirmed individually for HPV-6 as well as HPV-11 (Fig. 2A and Supplementary Fig. 2).
A Interferon-gamma ELISpot T-cell response displayed as peak log2 fold change above pretreatment responses against the HPV-6 (open black circles) and HPV-11 (open black squares) components for N = 32 patients, each represented by a symbol. Box and whiskers extend from 25th–75th percentile and minima to maxima, respectively; line at median, + at mean. B Top left panel: Representative staining of markers assessed for lytic granule loading (LGL) in HPV-specific multiparametric flow cytometry of CD8 + T-cells. Percentile of indicated cell populations are noted in each quadrant. Top right panel: peak frequency (%) of HPV-specific cytotoxic CD8 + T-cells (CTLs) above pretreatment responses as determined by flow cytometry and observed for N = 32 study patients, each represented by an open black diamond. Values at zero represent true zero values as well as negative values normalized to zero based on Y axis description. Box and whiskers extend from 25th–75th percentile and minima to maxima, respectively; line at median, + at mean. Bottom left panel: Peripheral T-cell profiling employing uniform manifold approximation and projection (UMAP) on all CD8 + T-cells and with activation, antigen experience, and effector marker identification, as determined by flow cytometry at Week 26 for n = 29 patients; no Week 26 data available for three patients. Bottom right panel: Heatmap of HPV-specific peak changes (Δ) in frequencies (%) of indicated parameters of CD8 + T-cells grouped by clinical response, as determined by flow cytometry. Infecting strain(s) of HPV noted in papilloma tissue are also indicated for n = 31 patients; one outlier is absent for improved visualization. C Left panel: Longitudinal frequency (%) of significantly expanded T-cell clonal populations as defined by CloneTrack for n = 27 patients, each represented by an open black diamond at any timepoint. Data were normalized to Screening (pretreatment) to specifically visualize on-study clonal expansion. Line denotes point-to-point geometric mean. Right panel: Absolute number of unique significantly expanded clonal populations observed during the study on a per-patient basis for n = 27 patients, each represented by an open black diamond. D Tracking of CDR3 clonotypes longitudinally across the study via Immunarch. Representative patient shown. E Break out of peripheral T-cell response frequencies by assay and in aggregate. log logarithm, HPV human papillomavirus, IFN-γ interferon-gamma, SFU spot forming units, Grz granzyme, Prf perforin, Gnly granulysin, TCRβ T-cell receptor beta chain, PBMC peripheral blood mononuclear cells, N number of patients, % percentile.
Upon confirmation of antigen-specific T-cell responses in blood, multiparametric flow cytometry was employed to further elucidate T-cell reactivity to INO-3107 antigens. Following stimulation with HPV-6 or HPV-11 peptides, assessment of markers indicating antigen-specific activation (CD137, CD69, CD38, and Ki67), antigen experience and immune regulation (PD-1, TIM-3, and LAG-3) and lytic capability (GrzA – Granzyme A, GrzB – Granzyme B, Gnly - Granulysin, Prf - Perforin) was performed on CD8 + T-cells as previously reported14. Heatmap visualization did not suggest broad trends between marker expression and clinical response (Fig. 2B). However, a supervised assessment of these data based on biological relevance and a previous study of immunotherapy for HPV revealed that a CD8 + T-cell subset consisting of cells expressing CD38, PD-1, and Prf was elevated in 83% of patients (26/32)14,15,21,22 (Fig. 2B and Supplementary Figs. 3, 4). The relevance of this phenotype was further supported through a uniform manifold approximation and projection (UMAP) assessment, which indicated the presence of a cluster that exhibited strong co-localization of all three of these markers (Fig. 2B). Responders exhibited shorter kinetics to the induction of response, higher overall response magnitude and superior persistence of response when compared to non-responders although the low proportion of non-responders in the trial (six patients) prevented a strong statistical comparison of these differences (Supplementary Fig. 5).
To further assess changes in the peripheral T-cell compartment, sequencing of complementarity-determining region 3 (CDR3) of the beta chain of the T-cell receptor (TCRβ) was performed. The CloneTrack system is a unique mathematical and immunological analytical platform that has been employed previously to identify the expansion of high-magnitude T-cell clones in blood after the administration of immunotherapy through assessment of TCRβ sequencing data23. In the current study, CloneTrack analysis identified T-cell clones that were expanded significantly following INO-3107 treatment and could be detected through the end of the study (Week 52) (Fig. 2C). Interestingly, the persistence of expanded T-cell clones differed between non-responders and responders, where the former exhibited an expansion at study Week 6 followed by a steady decline to Week 52, while the latter showed an expansion at study Week 6 that was sustained thereafter (Supplementary Fig. 6). Additional assessment of clonotype tracking of T-cells using Immunarch24 confirmed persistently expanded CDR3 sequences within responders, as identified via CloneTrack (Fig. 2D and Supplementary Fig. 7). Notably, 100% (27/27) of patients with sufficient testable samples exhibited T-cell expansion relative to their matched pretreatment assessment (range 2–119 significantly expanded unique clonal populations) (Fig. 2D), indicating engagement of peripheral T-cell clonal populations after treatment with INO-3107.
Aggregate assessment of ELISpot, flow cytometry and significant T-cell clonal expansion data determined that 100% of patients (32/32) exhibited peripheral immune response during the conduct of the study (Fig. 2E). Taken together these results suggest that INO-3107 robustly engaged peripheral T-cells, inclusive of the induction of HPV-6 and HPV-11-specific activity.
Treatment with INO-3107 results in a significantly enriched antiviral immune status in responder airways
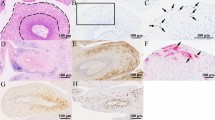
Once the induction of INO-3107-driven HPV-6 and HPV-11-specific immune responses in peripheral blood was confirmed, the determination of immune status in airway tissues of patients obtained prior to INO-3107 dosing as well as at the end of the study was analyzed via RNA sequencing (RNAseq) of airway tissue samples. Gene set enrichment analysis (GSEA) was first employed in the assessment of hallmark signatures for interferon-gamma response (IGR), interferon alpha response (IAR), and inflammatory response (IR) as defined by others25,26. The output of these analyses did not suggest significant pretreatment differences in papillomas of responders when compared to non-responders in any signature (Supplementary Fig. 8). However, after treatment with INO-3107, significant enrichment was observed for all three signatures exclusively in patients exhibiting clinical responses (IGR p = 0.000, q = 0.000; IAR p = 0.000, q = 0.001; IR p = 0.000, q = 0.014), (Fig. 3A). Core enrichment of these response signatures was driven by the induction of pro-inflammatory cytokine and chemokine genes (IL7, IL15, XCL1, CXCL10, CXCL11, others), and upregulation of genes associated with the presence of innate and adaptive immune cells (CSF1, CD14, CCL5, TBX21, others) (Fig. 3A and Supplementary Fig. 9).
A Gene set enrichment analysis (GSEA) results for hallmark interferon gamma, interferon alpha, and inflammatory signatures comparing pretreatment papilloma tissue with end of study tissue, inclusive of p value and q value, as determined by one-sided false discovery rate (FDR) correction for multiple comparisons, shown in the inset table. Core enriched genes are displayed with an intensity of color-coding tracking to rank metric scores of those genes. Non-core enriched genes are found in the supplement. B Heatmap of 280 differentially expressed genes (DEGs) in responders using one minus Spearman rank correlation with average linkage for hierarchical clustering and samples grouped by study visit; this gene set was analyzed using ingenuity pathway analysis (IPA), which confirmed the activation of interferon-gamma and interferon alpha signaling pathways in responder tissue at end of study versus screening via p value and z-score, as determined using one-sided Fisher’s exact test and reported in the inset table. HPV human papillomavirus, TMM trimmed mean of M-values, CPM counts per million.
While GSEA allows for testing of sets of genes across different variables, Ingenuity Pathway Analysis (IPA) allows for the synthesis of RNAseq data such that more complex questions can be answered regarding larger functional networks27. To that end, IPA was employed for deeper analysis of differentially expressed genes (DEGs) identified when comparing Screening (prior to treatment) to End of Study (Fig. 3B), inclusive of assessment of interferon responses identified by GSEA as well as querying specific immune cell functional networks. IPA confirmed interferon-gamma (IFNγ) and interferon-alpha (IFNα) activity as being significantly enriched in the airways of responding patients at the end of the study as compared to matched pretreatment papilloma samples (IFNγː z = 4.521, p = 4.95e-20; IFNαː z = 2.932, p = 1.12e−05, Fig. 3B and Supplementary Fig. 10). Further analyses of this dataset identified a variety of significantly enriched immune cell-associated functional networks, all of which were represented specifically in patients exhibiting clinical responses. These networks included multiple functions of antigen presenting cells (APCs) inclusive of chemotaxis (z = 2.715, p = 3.03e−06), binding (z = 2.479, p = 1.33e−11), and response (z = 3.021 p = 5.85e−07) among others (Table 3, Fig. 4A, and Supplementary Figs. 11–13). Encouragingly, a large number of T-cell associated networks were also noted as significantly enriched including T-cell homing (z = 2.358, p = 7.77e−06), T-cell activation (z = 2.238, p = 1.29e−26) and cytotoxicity of T-cells (z = 2.596, p = 1.18e−06), which collectively included enriched genes such as CD3E, CD3G, CXCR3, CCR5, LCK, SH2D2A, GZMA, and PRF1 amongst others (Table 3 and Fig. 4).
A Bubble plot of various IPA outputs relevant to inflammation as well as adaptive and innate immune responses that are enriched in airway tissue after INO-3107 treatment. p value and z-score (orange = activated; blue = inhibited) determined by one-sided Fisher’s exact test. B Network diagram of IPA data from tissue-localized T-cell activity, inclusive of p value and z-score as determined using one-sided Fisher’s exact test and reported in the inset table. Networks are defined by central nodes with bold names, differentially expressed genes (DEGs) that are upregulated (red) or downregulated (green), labeled with gene names, and connected to nodes they contribute to with dashed lines. Log logarithm, MHC major histocompatibility complex, Th helper T-cell.
INO-3107 drives significant T-cell infiltration into papillomas predominantly from emergent T-cell clones that traffic from blood
Due to IPA suggesting a variety of active T-cell-related pathways in patient airways, further exploration was undertaken to elucidate the type and extent of T-cell infiltration. Single sample GSEA (ssGSEA) was employed as a means of more specifically describing tissue-localized CD8 + T-cell populations on a per-patient basis. Results of these assessments identified the presence of various CD8 + T-cell infiltration signatures, inclusive of those bearing transcriptional hallmarks of both central memory (Tcm) and effector memory (Tem) maturation status28,29 (Fig. 5). When comparing pretreatment tissue to end of study tissue, increases in total CD8 + T-cell, CD8+ Tcm, and CD8+ Tem enrichment scores were observed, inclusive of a statistically significant increase specifically for the total CD8 + T population (Fig. 5, p < 0.001). To further profile tissue-localized CD8 + T-cells, a transcriptional signature for cytotoxic CD8 + T-cells was employed30,31. Results revealed a significant increase in the enrichment score for this cytotoxic population after treatment with INO-3107 (Fig. 5, p < 0.0005). When assessed by clinical response, responders had significantly higher total CD8 + T-cell, CD8+ Tem, and CD8 + T cytotoxic scores in end of study tissue as compared to pretreatment, while non-responders did not (Supplementary Fig. 14). Interestingly, while non-responders as a group did not reach statistical significance in the enrichment of CD8 + T-cell signatures, we did note increases in enrichment scores for four out of five of these patients (Supplementary Fig. 15). This result would suggest that increased CD8 + T-cell activity was occurring in the papilloma of some non-responding patients albeit to a lesser extent than responders. Taken together the results described here suggest that treatment with INO-3107 induced CD8+ enrichment in patient airways inclusive of signatures for cytotoxic status.
Single sample GSEA comparison of Screening (pretreatment) to end of study for total CD8+ (top left), CD8+ effector memory (top right), CD8+ central memory (bottom left) and CD8+ cytotoxic (bottom right) T-cell signatures. Selected core enriched genes for these assessments are identified in the inset table: CD3D/E/G and CD8A are T-cell identification markers. CD27 encodes a receptor required for activation of CD8 + T-cells. EOMES and TBX21 are transcription factors expressed in activated CD8 + T-cells. CTSW, GZMA, GZMK, FASLG, and PRF1 encode lytic proteins that induce T-cell killing of target cells. Both CX3CR1 and CXCR6 chemokines promote the survival, maintenance, and migration of CD8 + T-cells. Box and whiskers extend from 25th–75th percentile and minima to maxima, respectively; line at median, + at mean. p values are from a two-sided Wilcoxon signed-rank test of n = 17 patients, each represented by an open black diamond at both timepoints.
As an orthogonal method of confirming increased T-cell infiltration as suggested by IPA and ssGSEA, direct TCRβ sequencing of patient tissues was employed as a means of assessing tissue-localized T-cells through enumeration of TCRβ clone counts. To determine if T-cell infiltration prior to receiving INO-3107 was associated with clinical efficacy, TCRβ clone counts from papillomas resected prior to treatment were assessed against the percent change in surgical frequency during the study. No association was noted (Fig. 6A), and assessment of median TCRβ clone counts between non-responders (1130 TCRβ clones) and responders (1076 TCRβ clones) at this timepoint were similar (Supplementary Fig. 16). When the same assessment was performed on tissue obtained at the end of the study, a trend was noted between higher clone counts and the likelihood of a reduction in the surgical intervention (Fig. 6A), which was also evidenced by higher median clone counts in responders (4195 TCRβ clones) than non-responders (1218 TCRβ clones) at this timepoint (Supplementary Fig. 16). Further analysis focusing on the change in the number of TCRβ clone counts in the end of study tissue relative to pretreatment tissue revealed a clear and significant increase after INO-3107 treatment (Fig. 6B, p < 0.02). This pattern was found to be persistent when assessed in the context of clinical response status (Fig. 6B, p = 0.001). Further analysis revealed that the degree of change in T-cell infiltration in airway tissues after treatment with INO-3107 associates strongly with the change in surgery frequency on trial (Fig. 6C, p = 0.019 and Supplementary Fig. 17, p = 0.015).
A Regression analysis for TCRβ sequencing of tissue-localized T-cells in papilloma tissue from screening (pretreatment, left panel) and end of study (right panel) assessed against percent (%) change in surgical frequency during the study year (post-) relative to the year prior (pre-) to study start. Data were displayed as total TCRβ clone counts as determined by summed total reads from n = 17 patients, each represented by an open black diamond. p values and r values are determined by two-sided Spearman correlation analysis. B Left panel: Direct comparison of total TCRβ clone reads in papilloma tissue from Screening (pretreatment) and end of study from n = 17 patients, each represented by an open black diamond. Right panel: TCRβ clone reads in papilloma tissue from Screening (pretreatment) and end of study additionally broken out by clinical outcome where n = 5 non-responders (open red circles) and n = 12 Responders (open green squares). Box and whiskers extend from 25th–75th percentile and minima to maxima, respectively; line at median, + at mean. p values are determined by a two-sided Wilcoxon signed-rank test. Only significant p values are displayed. C Regression analysis of change (Δ) in TCRβ clone reads above Screening (pretreatment) to end of study against change in surgical frequency (%) on the study (post-) as compared to the year prior (pre-) to study start. p value and r value are determined by a two-sided Spearman correlation analysis of n = 17 patients, each represented by an open black diamond. TCRβ T-cell receptor beta chain, Tx treatment.
Analysis of CDR3 regions of TCRs present in airway tissue taken at the end of the study in clinical responders suggested a low degree of overlap with those sequences noted in papillomas resected prior to INO-3107 administration (Fig. 7A). To determine where TCRβ sequences discrepant between these two timepoints originated, peripheral blood of patients was assessed for the presence of these newly detected sequences. Results of this assessment revealed that such sequences were present in the blood prior to the end of the study, suggesting the trafficking of T-cell clones from circulation into papilloma tissue. Most encouragingly, the majority of these T-cell clones were found to be emergent clones that were not detectable in blood or papilloma tissue prior to administration of INO-3107 but became detectable only after initiation of therapy (Fig. 7B). Assessment of the degree of increase in emergent clones suggested a statistically significant expansion relative to prior to the initiation of INO-3107 treatment (Fig. 7B, p = 0.005). Comparing emergent clones in peripheral blood to infiltrating T-cell clonal populations in airway tissue taken at the end of the study revealed that the majority of infiltrating tissue-localized TCRβ sequences were specifically emergent T-cell clonal populations (Fig. 7C). These TCRβ results confirm the increased T-cell presence indicated by IPA and ssGSEA in the airways of treated patients. Furthermore, these results elucidate that clinical response is associated with the degree of increase in TCRβ clone counts in patient airways and that the majority of these T-cell clones are emergent T-cells that are detectable in blood only after administration of INO-3107, followed by trafficking to papillomas.
A Venn diagram displaying limited overlap (25 shared clones) between TCRβ clones identified in end of study papilloma tissue as compared to Screening (pretreatment) tissue for n = 13 patients. B Left panel: Emergent T-cell clones are identified and tracked longitudinally for n = 14 on study patients, each represented by lines connecting open diamonds color-coded by the left panel. No clone was detectable at Screening (pretreatment). Right panel: Assessment of emergent T-cell clone peak frequencies show statistical significance via two-sided Wilcoxon signed-rank test for n = 14 patients, each represented by a green line connecting an open black diamond at each timepoint. C Left panel: peak emergent T-cell clones in blood identified on a per-patient basis for n = 14 patients each represented by a color-coded bar and black diamond. Right panel: Venn diagram of considerable overlap between unique TCRβ clones in end of study airway tissue with emergent clones identified in the blood of n = 13 patients. TCRβ T-cell receptor beta chain, PBMC peripheral blood mononuclear cells.
Discussion
Here, we report results for a DNA immunotherapy for RRP from a completed Phase 1/2 trial demonstrating the safety, efficacy, and immune responses of INO-3107. When compared to the year prior to treatment with INO-3107, 26 RRP patients (81.3%) treated with INO-3107 exhibited reductions in clinically indicated surgical interventions in the year following treatment. Clinical response was seen across both HPV-6 and/or HPV-11 infections, suggesting that INO-3107 efficiently addresses both viral genotypes. Despite the small trial population of 32 patients, enrolled patients were inclusive of a broad age group ranging from 25 to 82, with HPV-6 and HPV-11, and both juvenile or adult symptom onset, representing the general adult RRP population and supporting the generalizability of the results.
The importance of reducing the surgical burden of this disease cannot be overstated. In this study, 81% of patients saw a reduction in number of surgeries, when comparing surgical treatments in the year prior to treatment with the year following Day 0, and there was a median decrease in the number of surgeries by three. A recent report from a long-term study of RRP patients undergoing recurrent surgeries reported an increased incidence of permanent damage to vocal cords and airway as a direct function of surgical intervention and specifically highlighted the need for non-surgical interventions in RRP patients8. Additionally, while non-surgical interventions such as monoclonal antibodies to various targets have been attempted in the treatment of RRP, they have shown variable impact with respect to control of disease once therapy is removed, and continued therapy has known associated toxicities32,33,34,35. INO-3107 treatment was well-tolerated and allowed for partial or complete control of disease in the context of reducing surgical interventions during the monitoring period of the trial, with a four-dose regimen spanning only 9 weeks.
Limitations of this study should be acknowledged. This study was single-arm and not placebo-controlled. The decision to proceed with the surgical intervention was the same during the 52-week pre- and post-treatment periods and was left to the individual investigators and patients, introducing some variability into the study from site to site. Additionally, as the study was conducted in 32 patients, study size should be considered. The current study focused on RRP disease in adults only; future investigations could include studies of INO-3107 in the pediatric population. For correlative analyses, end of study tissue was not available from all complete responders and the contribution of the induction of CD4 + T-cell responses, while previously described after treatment with INO-310736, are not expanded on here as the current report is focused on CD8 + T-cell-based contribution to the proposed mechanism of action.
The induction of anti-HPV-6 and HPV-11 immune responses has been identified in patients who have shown varying degrees of response to adjuvant antibody therapy inclusive of PD-1/PDL-1, TGFβ, and VEGFA10,37,38,39, identifying a likely connection between the immune system and control of the disease. This concept is further supported by work from other groups using T-cell inducing immunotherapy for the treatment of RRP, which have shown promising clinical results, although the application of viral vectors for the induction of anti-HPV immune responses may have been limited by anti-vector immune responses and a difficult to treat papilloma microenvironment19. Here, we report not only the induction of HPV-6 and HPV-11 targeted immune responses, but also track newly identified T-cell clonal populations from the blood into patient airways after INO-3107 treatment. Post-treatment biopsies showed an increased inflammatory state, consisting of interferon signaling pathways and innate immune cells as well as CD8 + T-cell signatures, inclusive of cytotoxic immune responses. As the lack of such response is believed to be involved in maintaining papilloma disease state9,10,11,12,13, it was expected and confirmed that the induction of these responses by INO-3107 were identified as being associated with clinical benefit in our study. The maintenance of this immune response should, therefore, be expected to be associated with a prolonged impact on the disease state, although this remains to be confirmed. Of interest, patients who did not show a decrease in surgical intervention frequency while on this trial still showed immunological changes in their papilloma tissue with respect to these parameters, albeit less than patients who responded clinically. As the DNA platform allows multiple administrations without the risk of generating anti-vector immunity, and as it has been previously shown that adding “booster” doses outside the primary series of DNA therapies can augment immune responses40, the possibility remains open for re-administration of INO-3107 to augment the immune response to further improve clinical response. Such an undertaking would also allow for continued research into immune responsiveness in RRP patients over time, which may have broader implications for understanding the mechanisms underpinning changes in disease state.
The results presented here make a strong rational case for use of INO-3107 as a non-surgical approach to RRP treatment from both a clinical and mechanistic standpoint. To that end, future clinical trials are being planned to further address prolonged efficacy, immune response, and redosing to expand the potential impact of this approach for the benefit of RRP patients.
Methods
Study design and patients
The study was conducted in accordance with the Declaration of Helsinki, Good Clinical Practice guidelines, and local and national regulatory requirements. The protocol was reviewed and approved by the relevant Institutional Review Board/Ethics Committee at each participating site (NYU Langone; University of Texas Southwestern; Washington University School of Medicine; University of California, Davis; Johns Hopkins University School of Medicine; Mayo Clinic Arizona; Emory University; University of Cincinnati Medical Center). All patients provided written informed consent.
RRP-001 (ClinicalTrials.gov identifier: NCT04398433) was a Phase 1/2, single-arm, open-label study conducted across eight centers in the United States, for which an interim efficacy assessment for standard needle injection was reported36. As the addition of an experimental side port needle was not noted to achieve differences in efficacy or immune response, the trial is reported here in its entirety as a single treatment paradigm. This was a single dose-level trial using a six-patient safety run-in with a 1-week waiting period between each patient (see Supplementary Fig. 1).
Eligible patients were men and women aged ≥18 years with histologically documented HPV-6 and/or HPV-11 positive disease, historically and/or at baseline as assessed by SPF10 LiPA25 HPV DNA genotyping followed by HPV-6 and HPV-11 specific qPCR conducted by DDL Diagnostic Laboratory (Rijswijk, Netherlands) and were enrolled across eight sites to reduce potential selection bias. Patients could have presented as either Juvenile-onset (JO; diagnosed before age 12 years) or Adult-onset (AO; diagnosed at or after 12 years of age) RRP, who exhibited moderate to severe disease based on a history of required frequent RRP interventions to remove papilloma, defined as ≥2 RRP surgical interventions (in the operating room or office-based) in the year prior to treatment. Patients must have been judged by the investigator to be appropriate candidates for an upcoming surgical intervention. Key exclusion criteria included the use of off-label adjuvant therapy for RRP disease (e.g., antiviral medications, chemotherapy, anti-angiogenic therapy [including bevacizumab], and prophylactic HPV vaccination) within 3 months of study start; immunosuppression; high risk of bleeding; and any illness or condition that may affect patient safety or evaluation of study endpoints (full inclusion and exclusion criteria are listed in Supplementary Information and full protocol is available on clinicaltrials.gov and appended at the end of the Supplementary Information). All patients were required to undergo a clinically necessary surgical intervention, which included predominantly laser-based techniques or microdebridement, within 14 days of the first dose (Day 0) to maximize standardization of baseline disease burden across patients. All patients underwent an assessment of the RRP Severity Score (modified) immediately prior to baseline surgery (see below). No compensation was provided for patient participation.
Treatment
Patients received one 6.25 mg dose of INO-3107 (a DNA plasmid-based immunotherapy composed of a plasmid encoding for HPV-6 and HPV-11 E6 and E7 antigens as well as a plasmid encoding for interleukin-12) on Day 0 and Weeks 3, 6, and 9. INO-3107 was administered intramuscularly (IM) in the deltoid (or anterolateral quadriceps muscle, if the deltoid muscle was not possible or appropriate) in a 1 mL volume, followed immediately by electroporation (EP) using the CELLECTRA proprietary device designed to enable local transfection of DNA plasmids into the cells (Inovio Pharmaceuticals; Plymouth Meeting, PA).
Endpoints and assessments
The primary endpoint was safety and tolerability, assessed by reported TEAEs and serious adverse events (SAE). TEAEs were defined as those occurring within 30 days of receipt of the last dose of INO-3107. Safety was assessed by monitoring adverse events (AEs) from the time of signing informed consent through Week 52 or last visit, including SAEs, unanticipated (serious) adverse device effects (UADE), and dose-limiting toxicities (DLTs; see Supplementary Information). Adverse events were graded according to the National Cancer Institute CTCAE Version 5.041. Clinically significant changes in laboratory parameters and vital signs from baseline assessments were also assessed. There were no missing or incomplete safety data.
Secondary endpoints included: (1) efficacy as assessed by number of RRP surgical interventions in the year following Day 0 compared with the prior year (surgeries on Day 0 were considered to have been performed during the prior year); (2) efficacy as assessed by changes in staging assessments measured by the RRP Severity Score (modified) over time; and (3) antigen-specific cellular immune responses assessed by T-cell phenotype and lytic potential in peripheral blood mononuclear cells (PBMCs) by flow cytometry. There were no missing or incomplete efficacy data. Data was collected according to GCP/ICH standards at each of the enrolling investigational sites by qualified personnel representative of the Sponsor.
Clinical assessment
Clinical response to treatment with INO-3107 was defined as a reduction in clinically indicated surgical interventions during the study period (52 weeks) as compared to the number of interventions recorded in the year prior to the first dose of INO-3107. Clinical response designation was sub-categorized as complete response19 (no surgeries in the year following the first dose of INO-3107), partial response19 (reduction of 50–99% in the frequency of surgeries in the year following the first dose, when compared with the year prior), overall response rate19 (complete response plus partial response), and overall clinical response (reduction of at least one surgery in the year following first dose when compared with the year prior). The decision to perform surgery among individual patients was made by their treating laryngologist, in order to maintain consistency in the time period prior to and following administration of INO-3107. The clinical trial protocol mandated that prior to surgery, both anatomical and symptom severity (modified Derkay) scores be obtained, and that the rationale for surgery be clearly documented in the case report forms.
RRP severity score (modified)
To standardize and increase precision in the grading of papillomas identified during each flexible laryngoscopy, the RRP staging system initially published by ref. 42, was revised by Pransky (S.M.P.) to provide a more detailed anatomical map of sites to be scored. Guidelines were issued to all investigators to assess lesions as surface (<3 mm), raised (4–8 mm), or bulky (>8 mm). Photo documentation of each laryngoscopy was required. The assessment form subdivided the true vocal cords (using both the free edge and superior surface as separate locations), ventricles, and false vocal cords into three subsections from anterior to posterior. The epiglottis and posterior glottis were divided into left and right. The aryepiglottic folds were clearly delineated, along with the left and right arytenoids. In addition, a definition and grading score were created for an extensive carpet of papilloma. If operative endoscopy was performed, the same laryngeal assessment form was utilized, along with assessment of the subglottis (divided into left and right, and anterior, lateral, and posterior segments) and the trachea (divided into superior, middle, and distal segments). The staging assessment was further expanded to include a symptom score that identified the presence of voice changes, stridor, respiratory difficulty, and symptoms such as cough, throat clearing, globus, or pain. Laryngoscopies were photographed, and the severity scores were retrospectively verified by independent otolaryngologists with extensive experience in RRP not participating in the study. Patients underwent office laryngoscopy and staging at Screening and Weeks 6, 11, 26, and 52.
Statistical analysis
No formal power analysis was applicable to the study; descriptive statistics were used to summarize data. The trial planned to treat approximately 30 patients, which would enable the trial to provide 95% confidence that the true incidence of SAEs was <12% if no SAEs were observed. For the primary endpoint, the main summary of safety data was based on TEAEs, with a frequency of preferred term events computed. The safety population comprised all patients who received at least 1 dose of INO-3107.
For secondary efficacy endpoints, the number of RRP surgical interventions in the year following the first dose of INO-3107 compared with the number in the year prior on a per-patient basis was summarized descriptively using median change and associated 95% confidence interval (CI). Mean changes and associated 95% CIs were computed for RRP severity scores from baseline pre-dose to each post-dose evaluation. The modified intention-to-treat population, comprising all patients who received at least one dose of INO-3107, was used for the secondary efficacy analyses. Due to the sample size (N = 32), separate analyses based on sex, race, or ethnicity were not planned.
Correlative analysis
Immunological and correlative statistical analysis was post hoc in nature. As sample was limited, data represent single assay runs from distinct samples obtained from each individual patient at specific timepoints during the conduct of the study. For peripheral blood analysis, data represent N = 32 patients for immunology assays and n = 27 patients with both pre- and post-treatment samples remaining for TCR sequencing, unless otherwise stated in the figure legend due to sample availability. For paired tissue analysis, 17 patients had sufficient samples for use: n = 5 non-responders and n = 12 responders per the overall clinical response definition. Graphing and statistical analyses were performed using GraphPad Prism (v9) software, unless stated otherwise.
Peripheral blood mononuclear cells (PBMCs) and serum were cryopreserved for immune analysis in batches. T-cell responses to HPV-6 and HPV-11 were determined by interferon-γ (IFN-γ) ELISpot and flow cytometry14. Briefly, 1 × 106 PBMCs were stimulated in vitro with pools of 15-mer peptides overlapping by 11 amino acids spanning HPV-6 or HPV-11 E6/E7 antigens. For ELISpot, PBMCs were stimulated with peptides for 18–24 h followed by quantification of IFN-γ spot forming units (SFU) per 106 PBMCs using ImmunoSpot® S6 counter and Professional (v7.037.0) software. For multiparametric flow cytometry, PBMCs were stimulated with peptides for five days without exogenous co-stimulation or cytokines, stained for CD3, CD4, CD8, CD137, CD69, CD38, Ki67, TIM-3, LAG-3, PD-1, Granzyme A, Granzyme B, Granulysin and Perforin, then acquired using BD LSRFortessa™ and FACSDiva™ (v9.0) software, and analyzed with FlowJo™ (v10.5.3). T-cell clonotype tracking was assessed using the bioinformatic tool Immunarch (v0.9.0) in R and CloneTrack assessment for statistically significant elevations in peripheral T-cell clones was performed23,24,43. Briefly, T cell clones were identified and tracked via their CDR3 nucleotide sequence. The total number of effective cells sequenced, N, was estimated as the summation of all productive TCRβ reads for a given sample, productive reads being defined as sequences in frame and without stop codons. An individual T cell clone count, nx, was estimated as the number of reads of the corresponding CDR3 nucleotide sequence with the frequency of that clone defined as \({f}_{x}=\frac{{n}_{x}}{N}\). We required that a specific clone demonstrate at least a twofold increase compared to pretreatment visits (M) to warrant further investigation. Clones demonstrating a fold change \( < 2(\frac{{m}_{x}}{M} < 2\times \frac{{n}_{x}}{N})\) were assigned default p values = 1, otherwise p values were calculated using an adapted Fisher’s exact test with the repertoire size of the initial sample rescaled by half. Furthermore, p values were adjusted using Bonferroni correction:\({P}_{{adjusted}}=P\times \left|N\cup M\right|\), where \(\left|N\cup M\right|\) is the number of unique clones in the union of the two samples being tested. A TCRβ clone was considered expanded were Padjusted < 0.05.
Logistical considerations prevented the possibility of isolating fresh tissue for viable tumor-infiltrating lymphocyte (TIL) isolation. In place of fresh TIL isolation, tissue obtained as papilloma resection at Screening (prior to INO-3107 treatment) as well as tissue obtained from resection or biopsy at the end of study was formalin-fixed and paraffin-embedded (FFPE) for use in RNA sequencing (RNAseq) and TCR sequencing (TCRseq) at Personalis, Inc. (Menlo Park, CA, USA). Dual isolation of nucleic acids was performed utilizing the AllPrep DNA/RNA FFPE Tissue kit (Qiagen). RNA input for sequencing was standardized across samples. Exome capture was performed using SureSelect Clinical Research Exome v2 (SSCRv2) (Agilent), according to the manufacturer’s recommendations. Additional supplementation with Personalis ACE proprietary target probes was performed to enhance coverage. Briefly, manufacturer protocols were modified to adjust the average library insert length to ~250 bp and the use of the Stranded RNA Sequencing kit (Kapa Biosystems) for RNA sequencing (RNAseq). Sequencing was performed on NovaSeq™ 6000 (Illumina, San Diego, CA, USA). Reads that mapped to the transcript of each gene were counted to measure expression levels for genes from each tumor sample. Raw strand-specific counts per gene were generated by the STAR aligner. Counts per million mapped reads (CPM) were calculated and globally normalized across samples using the trimmed mean of M values (TMM) using the Bioconductor package edgeR44,45,46,47. RNA-based T-cell receptor (TCR) clonotype nucleotide sequences were derived from both FFPE tissue and a minimum of 1 × 106 viable PBMCs using a universal software platform to analyze and process raw TCR repertoire sequencing data. Default settings and downstream filters were applied at reporting levels to remove false positives. Only productive TCR sequences are analyzed and reported here.
Immunological enrichment in tissue was assessed through gene set enrichment analysis (GSEA) and ingenuity pathway analysis (IPA) (QIAGEN Inc., https://digitalinsights.qiagen.com/IPA)25,26. Gene set enrichment analysis (GSEA)25,26,27 was used to describe enriched pathways and gene sets based on clinical outcomes following treatment. Differentially expressed genes (DEGs) were determined based on set quantitative thresholds between paired timepoints within the responder (those exhibiting an Overall Clinical Response) group: p < 0.05; Median TMM-adjusted CPM >1.5-fold change; median TMM-adjusted CPM >0.5 at either timepoint. Enrichment of DEGs was evaluated in IPA with respect to the user dataset of 22,955 genes detectable in our analysis27. For GSEA, the significance of enrichment was determined according to a one-sided, false discovery rate (FDR) and/or p value, while for IPA, significance was determined using a one-sided Fisher’s exact test per the software’s internal assessment. Single sample GSEA (ssGSEA) was performed in R using GSVA (v1.50.5)43,48. The sequencing data for this study has been deposited into the gene expression omnibus database under accession code GSE275788. Statistical analysis on comparative TCRβ frequencies and ssGSEA data was performed using GraphPad Prism (v9) software and employed two-sided Wilcoxon rank-sum or Wilcoxon signed-rank tests as indicated. Heatmap visualizations and hierarchical clustering by one minus Spearman rank correlation with average linkage were performed using Morpheus (https://software.broadinstitute.org/morpheus).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The sequencing data generated in this study have been deposited in the Gene Expression Omnibus database under accession code GSE275788. The de-identified individual patient clinical data are available under restricted access for ongoing research, regulatory, and privacy reasons due to ethical concerns related to the sensitive nature of the participant information. Interested investigators can obtain and certify a data transfer agreement and submit requests by emailing Jeffrey Skolnik (Jeffrey.Skolnik@inovio.com). The raw individual patient data were protected and are not available due to data privacy laws. The processed de-identified immunology data are available under restricted access for privacy, ethical, and ongoing research concerns. Interested investigators can obtain and certify a data transfer agreement and submit requests by emailing Matthew Morrow (Matthew.Morrow@inovio.com). Responses to data requests will be provided within 30 days of receipt.
Code availability
Custom code used in this analysis can be found on GitHub (https://github.com/Inovio-Pharmaceutical/CloneTrack.Adopted.git).
References
Plotzker, R. E. et al. Sexually transmitted human papillomavirus: update in epidemiology, prevention, and management. Infect. Dis. Clin. North Am. 37, 289–310 (2023).
Payaradka, R. et al. Oncogenic viruses as etiological risk factors for head and neck cancers: an overview on prevalence, mechanism of infection and clinical relevance. Arch. Oral. Biol. 143, 105526 (2022).
Derkay, C. S. & Bluher, A. E. Update on recurrent respiratory papillomatosis. Otolaryngol. Clin. North Am. 52, 669–679 (2019).
Major, T. et al. The characteristics of human papillomavirus DNA in head and neck cancers and papillomas. J. Clin. Pathol. 58, 51–55 (2005).
Ouda, A. M. et al. HPV and recurrent respiratory papillomatosis: a brief review. Life 11, 1279 (2021).
Sechi, I. et al. Pulmonary involvement in recurrent respiratory papillomatosis: a systematic review. Infect. Dis. Rep. 16, 200–215 (2024).
Rosenberg, T. et al. Therapeutic use of the human papillomavirus vaccine on recurrent respiratory papillomatosis: a systematic review and meta-analysis. J. Infect. Dis. 219, 1016–1025 (2019).
So, R. J. et al. Factors associated with Iatrogenic laryngeal injury in recurrent respiratory papillomatosis. Otolaryngol. Head. Neck Surg. 170, 1091–1098 (2024).
Skolnik, J. M. & Morrow, M. P. Vaccines for HPV-associated diseases. Mol. Asp. Med. 94, 101224 (2023).
Hatam, L. J. et al. Immune suppression in premalignant respiratory papillomas: enriched functional CD4+Foxp3+ regulatory T cells and PD-1/PD-L1/L2 expression. Clin. Cancer Res. 18, 1925–1935 (2012).
Israr, M. et al. Altered monocyte and Langerhans cell innate immunity in patients with recurrent respiratory papillomatosis (RRP). Front. Immunol. 11, 336 (2020).
Lucs, A. V. et al. Immune dysregulation in patients persistently infected with human papillomaviruses 6 and 11. J. Clin. Med. 4, 375–388 (2015).
DeVoti, J. et al. Decreased Langerhans cell responses to IL-36γ: altered innate immunity in patients with recurrent respiratory papillomatosis. Mol. Med. 20, 372–380 (2014).
Aggarwal, C. et al. Immunotherapy targeting HPV16/18 generates potent immune responses in HPV-associated head and neck cancer. Clin. Cancer Res. 25, 110–124 (2019).
Aggarwal, C. et al. Immune therapy targeting E6/E7 oncogenes of human papillomavirus type 6 (HPV-6) reduces or eliminates the need for surgical intervention in the treatment of HPV-6 associated recurrent respiratory papillomatosis. Vaccines 8, 56 (2020).
Aggarwal, C. et al. Safety and efficacy of MEDI0457 plus durvalumab in patients with human papillomavirus-associated recurrent/metastatic head and neck squamous cell carcinoma. Clin. Cancer Res. 29, 560–570 (2023).
Trimble, C. L. et al. Safety, efficacy, and immunogenicity of VGX-3100, a therapeutic synthetic DNA vaccine targeting human papillomavirus 16 and 18 E6 and E7 proteins for cervical intraepithelial neoplasia 2/3: a randomised, double-blind, placebo-controlled phase 2b trial. Lancet 386, 2078–2088 (2015).
Bagarazzi, M. L. et al. Immunotherapy against HPV16/18 generates potent TH1 and cytotoxic cellular immune responses. Sci. Transl. Med. 4, 155ra138 (2012).
Norberg, S. M. et al. The tumor microenvironment state associates with response to HPV therapeutic vaccination in patients with respiratory papillomatosis. Sci. Transl. Med. 15, eadj0740 (2023).
Smahelova, J. et al. Outcomes after human papillomavirus vaccination in patients with recurrent respiratory papillomatosis: a nonrandomized clinical trial. JAMA Otolaryngol. Head Neck Surg. 148, 654–661 (2022).
Eberhardt, C. S. et al. Functional HPV-specific PD-1+ stem-like CD8 T-cells in head and neck cancer. Nature 597, 279–284 (2021).
Pokrývková, B. et al. PD1+CD8+ cells are an independent prognostic marker in patients with head and neck cancer. Biomedicines 10, 2794 (2022).
Rojas, L. A. et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature 618, 144–150 (2023).
Nazarov, V. I. et al. immunarch: bioinformatics analysis of T-cell and B-cell immune repertoires. R package version 0.9.0. https://CRAN.R-project.org/package=immunarch (2022).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Krämer, A., Green, J., Pollard, J. Jr. & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Regev, A. et al. The human cell atlas. Elife 6, e27041 (2017).
Aran, D., Hu, Z. & Butte, A. J. xCell: digitally portraying the tissue cellular heterogeneity landscape. Genome Biol. 18, 220 (2017).
Yan, K. et al. 9-Gene signature correlated with CD8+ T cell infiltration activated by IFN-γ: a biomarker of immune checkpoint therapy response in melanoma. Front. Immunol. 12, 622563 (2021).
Szabo, P. A. et al. Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease. Nat. Commun. 10, 4706 (2019).
Robbins, Y. et al. Dual PD-L1 and TGF-b blockade in patients with recurrent respiratory papillomatosis. J. Immunother. Cancer 9, e003113 (2021).
Warner, B. M. et al. Sicca syndrome associated with immune checkpoint inhibitor therapy. Oncologist 24, 1259–1269 (2019).
Kalra, N. et al. Angiogenesis inhibitor drug-induced benign migratory glossitis in a patient of juvenile-onset recurrent respiratory papillomatosis under maintenance therapy. Int. J. Clin. Pediatr. Dent. 17, 92–96 (2024).
Robinson, C. H. et al. Renal implications of long-term systemic bevacizumab for recurrent respiratory papillomatosis. Ann. Otol. Rhinol. Laryngol. 133, 119–123 (2024).
Mau, T. et al. Interim results of a phase 1/2 open-label study of INO-3107 for HPV-6 and/or HPV-11-associated recurrent respiratory papillomatosis. Laryngoscope 133, 3087–3093 (2023).
Lam, B. et al. Profiling of VEGF receptors and immune checkpoints in recurrent respiratory papillomatosis. Laryngoscope 134, 2819–2825 (2024).
Bai, K. et al. Durable response in a patient with recurrent respiratory papillomatosis treated with immune checkpoint blockade. Head Neck 44, E31–E37 (2022).
Liu, T. et al. PD-L1 expression and CD8+ infiltration shows heterogeneity in juvenile recurrent respiratory papillomatosis. Int J. Pediatr. Otorhinolaryngol. 95, 133–138 (2017).
Morrow, M. P. et al. Augmentation of cellular and humoral immune responses to HPV16 and HPV18 E6 and E7 antigens by VGX-3100. Mol. Ther. Oncolytics 3, 16025 (2016).
US Department of Health and Human Services, National Institutes of Health, National Cancer Institute. Common Terminology Criteria for Adverse Events (CTCAE), Version 5. (2017).
Derkay, C. S. et al. A staging system for assessing severity of disease and response to therapy in recurrent respiratory papillomatosis. Laryngoscope 108, 935–937 (1998).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing. https://www.R-project.org/ (2022).
Chen, Y. et al. edgeR v4: powerful differential analysis of sequencing data with expanded functionality and improved support for small counts and larger datasets. Nucleic Acids Res. 53, gkaf018 (2025).
Chen, Y., Lun, A. T. L. & Smyth, G. K. From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Res. 5, 1438 (2016).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Hänzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Acknowledgements
This study was funded by INOVIO Pharmaceuticals. The sponsor designed the study, collected and analyzed the data, and drafted the manuscript. The authors would like to thank the RRP-001 trial patients and site staff involved in this study, as well as Jan Pawlicki, Kimberly Kraynyak, and Alex Dolgoter, for their contributions.
Author information
Authors and Affiliations
Contributions
M.P.M. and J.M.S. contributed to the study design and development concept and drafted the manuscript. E.G. and A.S. directed analyses of all immune and sequencing assays. M.R.A., P.C.B., S.R.B., A.D.F., A.M.K., D.G.L., T.M., R.C.P., and N.F.S. were clinical investigators of the study. G.S.T. oversaw HPV typing for study inclusion. S.W., S.A.M., E.L.R., and K.S.R. were responsible for analyses of immune and sequencing data via CloneTrack, Immunarch, ssGSEA, GSEA, and IPA. S.M.P. and D.B.W. contributed to the conception of the program, interpretation of the data, as well as editing the manuscript. M.D. performed statistical analysis. All authors contributed to the acquisition, analysis or interpretation of the data, and performed critical revision of the manuscript for intellectual content.
Corresponding author
Ethics declarations
Competing interests
M.P.M., E.G., A.S., G.S.T., S.W., S.A.M., E.L.R., K.S.R., M.D., and J.M.S. are employees of and may hold stock and/or stock options in Inovio Pharmaceuticals. T.M., P.C.B., S.R.B., and D.G.L. received honoraria for consultancy for Inovio Pharmaceuticals. M.R.A. and R.C.P. have no funding, financial relationships, or conflicts of interest to disclose. A.D.F. received honoraria for consultancy for Inovio Pharmaceuticals and Precigen, Inc. A.M.K. receives support from the RRP Foundation for attending an RRPF Research meeting. S.M.P. receives honoraria for consultancy and advice for Inovio Pharmaceuticals; and is a member of the Recurrent Respiratory Task Force of the American Society of Pediatric Otolaryngology. N.F.S. provides advice for Inovio Pharmaceuticals; and provided compensated or non-compensated advisory role for Astra Zeneca, Merck, CUE Biopharma, and PDS. D.B.W. received remuneration as a member of the Board of Directors and consultant for Inovio Pharmaceuticals, and received honoraria for consultancy for Geneos, Flagship, and Advaccine, and as an advisor for Geneos, Astra Zeneca, BBI/Sumitomo, and Pfizer, and as a speaker for Astra Zeneca. All authors declare to have competing interests. None of the other authors have any competing interests.
Peer review
Peer review information
Nature Communications thanks Zhenkun Yu, who co-reviewed with Liu Kai; Everardo Saad, Ruth Tachezy and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Morrow, M.P., Gillespie, E., Sylvester, A. et al. DNA immunotherapy for recurrent respiratory papillomatosis (RRP): phase 1/2 study assessing efficacy, safety, and immunogenicity of INO-3107. Nat Commun 16, 1518 (2025). https://doi.org/10.1038/s41467-025-56729-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56729-6