Abstract
Pathological retinal neovascularization (RNV) is one of the leading causes of blindness worldwide; however, its underlying mechanism remains unclear. Here, we found that the expression of endothelial protein C receptor (Epcr) was increased during RNV, and its ligand was elevated in the serum or vitreous body of patients with proliferative diabetic retinopathy. Deleting endothelial Epcr or using an EPCR-neutralizing antibody ameliorated pathological retinal angiogenesis. EPCR promoted endothelial heme catabolism and carbon monoxide release through heme oxygenase 1 (HO-1). Inhibition of heme catabolism by deleting endothelial Ho-1 or using an HO-1 inhibitor suppressed pathological angiogenesis in retinopathy. Conversely, supplementation with carbon monoxide rescued the angiogenic defects after endothelial Epcr or Ho-1 deletion. Our results identified EPCR-dependent endothelial heme catabolism as an important contributor to pathological angiogenesis, which may serve as a potential target for treating vasoproliferative retinopathy.
Similar content being viewed by others
Introduction
Pathological retinal neovascularization (RNV) is a major cause of vision loss worldwide1,2. The most common causes include diabetic retinopathy, retinopathy of prematurity (ROP), and retinal vein occlusion, which affect millions of people3,4,5. The aberrant vessel growth usually leads to macular edema, fibrotic scarring, and retinal detachment2,6. These conditions are the primary causes of blindness in working-age and pediatric populations in developed countries6,7. Endothelial cells (ECs) rarely proliferate and remain in a quiescent state in the adult organism, whereas uncontrolled proliferation of ECs is initiated and drives aberrant vessel growth under pathological stimuli8. Vascular endothelial growth factor (VEGF) is extensively studied in the regulation of EC proliferation and angiogenesis9,10. Anti-VEGF regents are first-line therapy for RNV, but resistance occurs in certain patients11,12. Hence, it is necessary to identify novel therapeutic targets of RNV.
Endothelial protein C receptor (EPCR, also known as PROCR) is a single-pass transmembrane glycoprotein expressed in ECs and stem cells13,14. In addition to its anti-coagulation role15, EPCR mediates intracellular signaling of activated protein C (APC)16. Activation of downstream protein kinase B (AKT) promotes cell growth, improves cardiac function, and favors tumor progression16,17,18,19,20. Lineage tracing of Epcr-positive cells indicate that Epcr is specifically expressed in the ECs of the retinal vasculature21. Epcr transcription is activated in the retinal vessels during the angiogenic expansion stage but not during the quiescent stage21. Ablation of Epcr-positive cells leads to delayed retinal vascular expansion in postnatal mice, suggesting a role for EPCR in retinal angiogenesis22. However, the role of EPCR in RNV pathology remains unclear.
This study found that Epcr transcription was activated in ECs during RNV. Conditional knockout of Epcr in ECs or using an EPCR-neutralizing antibody ameliorated pathological RNV. Further data showed that EPCR affected heme catabolism by regulating heme oxygenase 1 (HMOX1, also known as HO-1). Conditional knockout of Ho-1 in ECs or treatment with an HO-1 enzymatic inhibitor suppressed pathological angiogenesis in retinopathy. Carbon monoxide-releasing molecule-3 (CORM3) partially rescued the reduced neovascularization caused by the deletion of endothelial Epcr and Ho-1, indicating a role for carbon monoxide (CO). Clinically, levels of APC, which is an EPCR ligand, were elevated in proliferative diabetic retinopathy (PDR) patients’ serum and vitreous body. These results indicate that EPCR-dependent endothelial heme catabolism is an important contributor to RNV.
Results
EPCR is highly expressed in retinal ECs during RNV
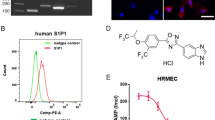
The mouse model of oxygen-induced retinopathy (OIR) serves as a proxy for human pathological RNV, including PDR and ROP23. The neovascular response peaked on postnatal day 17 (P17) when the mice were returned to room air for five days (Fig. 1a). To identify the potential genes involved in pathological neovascularization, we performed transcriptomic analyses of the retinas of P17 mice of OIR (GSE241239). To minimize the RNA-seq bias, we further analyzed transcriptomic data from two other independent studies (GSE194176 and GSE158799)24,25. A combined analysis revealed 303 upregulated genes (Fig. 1b, c). Among these candidate genes, we decided to focus on Epcr because EPCR was reported to be specifically detected on the surface of vascular ECs and stem cells13,14. Further data confirmed that Epcr expression was upregulated in the retinas of P17 mice in the OIR (Fig. 1d, e). We sorted ECs from the mouse retina to evaluate the expression of Epcr in the retinal vasculature. Endothelial Epcr expression was much higher in the OIR group than in the control group (Fig. 1f, g).
a Illustration of mouse model of OIR and different phases of retinopathy. Created in BioRender. Song, H. (2025) https://BioRender.com/g81i639. b Combined analysis of upregulated genes of three independent RNA-seq data in the retina of OIR mice. GSE241239 was performed in the present study. c Volcano plot of Log2 fold-changed genes of three RNA-seq data. d The protein levels of EPCR in the retina throughout the progression of OIR. n = 3 biological replicates. Relative to β-Actin, Mean ± SD, p = 0.1864, 0.0102, 0.0064 (two-tailed Student’s t-test). Relative to CD31, Mean ± SD, p = 0.8608, 0.0362, 0.0010 (two-tailed Student’s t-test). e The mRNA levels of Epcr of the retina determined by qRT-PCR throughout the progression of OIR. n = 3 biological replicates. Mean ± SD, p = 0.7809, 0.0002, 0.0025 (two-tailed Student’s t-test). f FACS-isolated retinal endothelial cells from OIR mice at P17 showed increased expression of mRNA levels of Epcr. n = 3 biological replicates. Mean ± SD, p = 0.0133 (two-tailed Student’s t-test). g The representative protein levels of EPCR in FACS-isolated retinal endothelial cells from OIR mice. n = 3 biological replicates. Mean ± SD, p = 0.0136 (two-tailed Student’s t-test). h Representative images of lineage tracing of Epcr-positive cells in P17 mice using Epcr-CreERT2; Rosa26tdT mice. Scale bar 500 μm. i Quantification of the fluorescent intensity of Epcr-positive cells. n = 3 biological replicates. Mean ± SD, p = 0.0018 (two-tailed Student’s t-test). j WT (n = 30) and OIR (n = 30) mice serum APC levels were determined using ELISA. Mean ± SD, p = 0.0070 (two-tailed Student’s t-test). k Ctrl (n = 20) and PDR patients (n = 20) serum APC levels were determined with ELISA. Mean ± SD, p = 0.0080 (two-tailed Student’s t-test). l APC levels in Ctrl (n = 10) and PDR (n = 10) vitreous body were determined with ELISA. Mean ± SD, p = 0.0014 (two-tailed Student’s t-test). ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
Next, we examined the expression pattern of EPCR in the retina. Immunofluorescence staining with an anti-EPCR antibody showed that EPCR was selectively detected in the retinal ECs of P17 mice under physiological conditions (Supplementary Fig. 1a). In the OIR model, EPCR was detected only in retinal ECs, and the fluorescence intensity of EPCR signals was much higher in the neovascular tufts (Supplementary Fig. 1b, c). Epcr-CreERT2 (also known as Procr-CreERT2) mice were previously used to mark proliferative ECs in the retinal vasculature21. Here, we used Epcr-CreERT2 and Rosa26tdT mice to perform genetic lineage tracing of Epcr-positive cells in the retina of P17 mice. Tamoxifen (TAM) was administered for three consecutive days at P10, P11, and P12, and the retina was analyzed at P17 (Supplementary Fig. 1d). Under physiological conditions, a very small proportion of Epcr+ cells were observed (Fig. 1h). There was a large proportion of Epcr+ cells in the retina of OIR mice, and Epcr+ cells mainly colocalized with neovascular tufts (Fig. 1h, i, Supplementary Fig. 1e). These data indicated that hypoxia significantly increased the transcription of Epcr. We also assessed the protein levels of EPCR in the retina of db/db mice with early-stage diabetes. The data showed that there was no difference in the protein levels of EPCR between db/db mice and db/m mice (Supplementary Fig. 1f–h). Hypoxia is also known to play a vital role in the retinal vascular development process26. Therefore, the expression pattern of EPCR in the retinal vasculature of postnatal mice was then assessed. Immunofluorescence staining with anti-EPCR showed that the levels of EPCR were much higher in the margin of the retinal vasculature (Supplementary Fig. 1i). Consistent data were acquired using Epcr-CreERT2 and Rosa26tdT mice (Supplementary Fig. 1j, k). Collectively, these data suggest that EPCR is specifically detected in ECs of the retina and is upregulated in neovascularization in the OIR model and during retinal vascular expansion.
EPCR activates downstream signaling pathways by binding to its ligands. Thus, we measured the serum concentrations of APC, a major ligand for EPCR, in OIR mice. The data showed that the levels of APC in the serum of OIR mice were higher than those in control mice (Fig. 1j). To evaluate whether the findings in mice hold promise for human patients with RNV, we investigated the levels of APC in the vitreous humor and serum in patients with PDR. The concentrations of APC in the vitreous humor and serum were significantly higher in patients with PDR than in the control with a macular hole or idiopathic epiretinal membrane (Fig. 1k, l).
EPCR contributes to RNV in vivo and in vitro
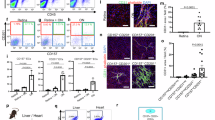
To determine the role of EPCR in RNV, we intercrossed the Cdh5-CreERT2 mouse with Epcrflox/flox mouse to generate EC-specific Epcr KO mice (EpcriΔEC). The protein levels of EPCR were almost completely ablated in ECs sorted from EpcriΔEC mice (Supplementary Fig. 2a, b). Cre toxicity has been reported in another study27, whereas our data showed that this mice strain did not cause vascular changes in the OIR model (Supplementary Fig. 2c–e). To assess the role of EPCR in pathological RNV, we examined the changes in the retinal vasculature of EpcriΔEC mice of OIR at P17. Our data showed that the area of neovascular tufts in the retina of EpcriΔEC mice was smaller than that in EpcrWT mice (Fig. 2a, b). In contrast, the avascular area in EpcriΔEC mice’s retina was larger than that in EpcrWT mice (Fig. 2a, c). Furthermore, we used 5-ethynyl-2’-deoxyuridine (EdU) to label proliferative cells in the retina and used E26 transformation-specific (ETS)-related gene (ERG) as an EC marker. The result showed that EdU+ ECs in the retina decreased significantly in EpcriΔEC mice compared to EpcrWT mice (Fig. 2d, e). Retinal vascular leakage was also attenuated in EpcriΔEC mice’s retina compared to the retina of EpcrWT mice (Supplementary Fig. 2f, g). Meanwhile, the data indicated that the deletion of endothelial Epcr did not affect the mature physiological angiogenesis of mice at P17 (Supplementary Fig. 2h, i). These in vivo results suggest that upregulated EPCR in pathological vasculature drives EC growth and angiogenesis in OIR.
a Representative images of retinal vasculature stained with IB4 in EpcrWT and EpcriΔEC mice at P17 using the OIR mice model. The orange space indicates the avascular area for two left images and neovascular tufts for two right images. Scale bar 1000 μm and 500 μm. b Quantification of the area of neovascular tufts in EpcrWT (n = 8) and EpcriΔEC (n = 7) mice. Mean ± SD, p = 0.0004 (two-tailed Student’s t-test). c Quantification of the avascular area in EpcrWT (n = 8) and EpcriΔEC (n = 7) mice. Mean ± SD, p = 0.0218 (two-tailed Student’s t-test). d Immunofluorescence staining for IB4 (blue), E26 transformation-specific (ETS)-related gene (red), and EdU (green) in the retina of EpcrWT and EpcriΔEC mice. Proliferating (E26 transformation-specific (ETS)-related gene and EdU double positive) ECs are shown in yellow. Scale bar 50 μm. e Quantification of the proliferating ECs in the retina of EpcrWT (n = 6) and EpcriΔEC (n = 6) mice. Mean ± SD, p < 0.0001 (two-tailed Student’s t-test). f EdU-incorporation in EPCR-overexpressed human umbilical vein endothelial cells (HUVECs). DAPI (blue) was used to identify ECs nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. g Quantification of EdU-incorporation in EPCR-overexpressed HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0046 (two-tailed Student’s t-test). h Representative images of HUVECs sprouting in EPCR-overexpressed HUVECs. Scale bar 50 μm. i Quantification of sprouts length in EPCR-overexpressed HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0375 (two-tailed Student’s t-test). j Wound healing of EPCR-overexpressed HUVECs. Scale bar 100 μm. k Quantification of wound healing ability in EPCR-overexpressed HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p < 0.0001 (two-tailed Student’s t-test). l EdU-incorporation in EPCR-depleted HUVECs. DAPI (blue) is used to identify EC nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. m Quantification of EdU-incorporation in EPCR-depleted HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0061 (two-tailed Student’s t-test). n Representative images of sprouting in EPCR-depleted HUVECs. Scale bar 50 μm. o Quantification of sprouts length in EPCR-depleted HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0056 (two-tailed Student’s t-test). p Wound healing of EPCR-depleted HUVECs. Scale bar 100 μm. q Quantification of wound healing ability in EPCR-depleted HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0094 (two-tailed Student’s t-test). ∗∗∗∗p < 0.0001, ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
Next, we determined the effect of Epcr deletion on retinal vascular expansion in postnatal retinas. EC-specific deletion of Epcr led to a sparse vascular network and reduced radial expansion of the superficial retinal vascular plexus (Supplementary Fig 3a, b). Tip cells’ filopodia and branch points were decreased in the retina of endothelial Epcr KO mice (Supplementary Fig. 3c–e). Meanwhile, EC proliferation was also significantly inhibited in EpcriΔEC mice (Supplementary Fig. 3f, g).
To further evaluate the impact of elevated EPCR on angiogenesis, we used an adenovirus to overexpress EPCR and assessed the angiogenic behavior of ECs in vitro. EPCR adenovirus treatment significantly increased the protein levels of EPCR in ECs, which promoted EC proliferation in an EdU staining assay (Fig. 2f, g, Supplementary Fig. 3h). A three-dimensional endothelial spheroid assay showed that the length of endothelial sprouts were increased in the EPCR adenovirus treatment group (Fig. 2h, i). Moreover, a wound scratch assay revealed that EPCR adenovirus-treated ECs were more motile than control adenovirus-treated ECs (Fig. 2j, k). Furthermore, the knockdown of EPCR using siRNA suppressed EC proliferation, migration, and sprouting (Fig. 2l–q, Supplementary Fig. 3i). Together with in vivo results, these data indicated that EPCR promotes vascular growth.
Pharmacological inhibition of EPCR ameliorates pathological retinal neovascularization
To evaluate the translational potential of targeting EPCR in anti-angiogenic therapy, we assessed the effects of a neutralizing antibody against EPCR on pathological angiogenesis in vitro and in vivo. We found that a single dose of EPCR antibody significantly reduced the area of neovascular tufts compared to the vehicle and anti-IgG groups (Fig. 3a, b). Whereas the avascular area remained unchanged after EPCR antibody treatment (Fig. 3a, c). Suppressed proliferation of ECs in retinal neovascular tufts was also observed after treatment with the EPCR antibody (Fig. 3d, e). Similar results were obtained in vitro as EPCR antibody treatment significantly inhibited EC proliferation, migration, and sprouting (Fig. 3f–k). These results were consistent with our findings using EpcriΔEC mice, suggesting the translational potential of anti-EPCR in the treatment of retinal neovascularization.
a Representative images of retinal vasculature stained with IB4 in PBS, anti-IgG and anti-EPCR-treated mice at P17 using the OIR mice model. The orange space indicates the avascular area for three lower images and neovascular tufts for three upper images. Scale bar 1 mm and 500 μm. b Quantification of the area of neovascular tufts in PBS (n = 7), anti-IgG (n = 8) and anti-EPCR-treated (n = 9) mice. Mean ± SD, p = 0.2614, 0.0164, 0.0002 (one-way ANOVA with Tukey’s multiple comparisons test). c Quantification of the avascular area in PBS (n = 7), anti-IgG (n = 8) and anti-EPCR-treated (n = 9) mice. Mean ± SD, p = 0.9994, 0.1934, 0.1622 (one-way ANOVA with Tukey’s multiple comparisons test). d Immunofluorescence staining for IB4 (blue), E26 transformation-specific (ETS)-related gene (red), and EdU (green) in anti-EPCR-treated mice. Proliferating (E26 transformation-specific (ETS)-related gene and EdU double positive) ECs are shown in yellow. Scale bar 100 μm. e Quantification of the proliferating ECs in PBS (n = 6), anti-IgG (n = 6) and anti-EPCR-treated (n = 6) mice. Mean ± SD, p = 0.4990, 0.0008 (one-way ANOVA with Tukey’s multiple comparisons test). f EdU-incorporation in anti-EPCR-treated ECs (C166). DAPI (blue) is used to identify EC nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. g Quantification of EdU-incorporation in anti-EPCR-treated ECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0005, p = 0.0297 (one-way ANOVA with Tukey’s multiple comparisons test). h Representative images of anti-EPCR-treated ECs sprouting. Scale bar 50 μm. i Quantification of sprouts length in anti-EPCR-treated ECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0026, p = 0.0183 (one-way ANOVA with Tukey’s multiple comparisons test). j Wound healing of anti-EPCR-treated ECs. Scale bar 100 μm. k Quantification of wound healing ability in anti-EPCR-treated ECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0007, p = 0.0089 (one-way ANOVA with Tukey’s multiple comparisons test). ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
EPCR regulates the expression of HO-1 through NRF2/KEAP1
To reveal the possible molecular mechanisms underlying EPCR-regulated angiogenesis, we performed transcriptomic analyses of EPCR siRNA- or control siRNA-treated ECs (GSE249130, Supplementary Fig. 4a). These data were then combined and analyzed with the mice retina results with OIR (Supplementary Fig. 4b). Thirty genes were upregulated in the retina of OIR mice and downregulated in Epcr-depleted ECs (Fig. 4a). Among these genes, Ho-1 was the most significantly affected (Fig. 4a). Increased mRNA and protein levels of Ho-1 were confirmed in the retinas of OIR mice (Supplementary Fig. 4c, d). Further data showed that the expression of Ho-1 was suppressed at the mRNA and protein levels in ECs treated with EPCR siRNA (Fig. 4b, c). Additionally, the expression of Ho-1 was upregulated at the mRNA and protein levels in ECs treated with EPCR adenovirus (Fig. 4d, e). The expression of Ho-1 in retinal ECs of OIR mice was then evaluated. We sorted ECs from the retina of OIR mice using flow cytometry. The results indicated that the expression of Ho-1 in ECs was upregulated in the OIR group (Supplementary Fig. 4e, f). This was further confirmed by immunofluorescence staining. The expression of HO-1 in the physiological retina did not show an apparent pattern, while it was highly expressed in neovascular tufts of OIR retinas (Supplementary Fig. 4g, h). Further data showed that the expression of Ho-1 in ECs sorted from the retina of EpcriΔEC mice decreased compared with that in Epcrwt mice (Fig. 4f, g). Collectively, these results suggested that HO-1 was a downstream molecule of EPCR in regulating angiogenesis.
a Heatmap of genes (p-value) downregulated in RNA-seq data of EPCR-depleted HUVECs and upregulated in the RNA-seq data of the retinas from OIR mice. Heatmap was generated and analyzed using RNA-seq data from EPCR-depleted HUVECs. b Representative immunoblots and quantification for HO-1 levels in EPCR-depleted HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.0077, 0.0008 (two-tailed Student’s t-test). c The mRNA levels of HO-1 in EPCR-depleted HUVECs determined by qRT-PCR. n = 3 biological replicates. Mean ± SD, p = 0.0008, 0.0016 (two-tailed Student’s t-test). d Representative immunoblots and quantification for HO-1 levels of EPCR adenovirus-treated HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.0025, 0.0016 (two-tailed Student’s t-test). e The mRNA levels of HO-1 in EPCR adenovirus-treated HUVECs determined by qRT-PCR. n = 3 biological replicates. Mean ± SD, p = 0.0058, <0.0001 (two-tailed Student’s t-test). f Representative immunoblots and quantification for HO-1 levels in ECs sorted from the retina of EpcriΔEC mice. n = 3 biological replicates. Mean ± SD, p < 0.0001, p = 0.0004 (two-tailed Student’s t-test). g The mRNA levels of Ho-1 in ECs sorted from the retina of EpcriΔEC mice. n = 3 biological replicates. Mean ± SD, p < 0.0001, p = 0.0001 (two-tailed Student’s t-test). h Representative immunoblots and quantification for HO-1, NRF2, KEAP1, p-AKT(S473), and EPCR levels in EPCR-depleted HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.017, 0.0021, 0.0031, 0.0007, 0.0014 (two-tailed Student’s t-test). i Representative immunoblots and quantification for HO-1, NRF2, KEAP1, p-AKT(S473), and EPCR levels in EPCR adenovirus-treated HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.0075, 0.0063, 0.0001, 0.0008, 0.0079 (two-tailed Student’s t-test). j Representative immunoblots and quantification for HO-1, NRF2, KEAP1, and p-AKT(S473) levels in EPCR adenovirus and LY294002-treated HUVECs. n = 3 biological replicates. Mean ± SD, p < 0.0001, p = 0.0002, 0.0002, 0.0128, <0.0001, 0.0003, 0.0003, 0.0029 (one-way ANOVA with Tukey’s multiple comparisons test). k Representative immunoblots and quantification for HO-1, NRF2 and KEAP1 levels in EPCR-depleted and KEAP1-depleted HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.0013, p = 0.0221, <0.0001, 0.0015, 0.0002, 0.0013 (one-way ANOVA with Tukey’s multiple comparisons test). l Representative immunoblots and quantification for HO-1 and NRF2 levels in EPCR-depleted and KEAP1 inhibitor-treated HUVECs. n = 3 biological replicates. Mean ± SD, p = 0.0247, 0.0395, 0.0068, 0.0426 (one-way ANOVA with Tukey’s multiple comparisons test). ∗∗∗∗p < 0.0001, ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
AKT is recognized as a target of EPCR signaling28. High EPCR expression or stimulation with APC usually leads to AKT activation16,19. We found that overexpression of EPCR using an adenovirus promoted the activation of AKT, whereas silencing EPCR with siRNA inhibited AKT phosphorylation (Fig. 4h, i). Nuclear factor erythroid 2-related factor 2 (NRF2) is a ubiquitous transcription factor directly regulating HO-129. NRF2 is tightly regulated by Kelch-like ECH-associated protein 1 (KEAP1) through ubiquitination and proteasome-dependent degradation30. Decreased expression of NRF2 and HO-1, and increased expression of KEAP1 were observed after EPCR knockdown in ECs (Fig. 4). Therefore, we assessed the effects of EPCR on the expression of KEAP1 and NRF2. Overexpression of EPCR in ECs promoted the levels of NRF2 and HO-1 and inhibited the expression of KEAP1 (Fig. 4i). Meanwhile, APC treatment stimulated the activation of AKT and increased the expression of NRF2 and HO-1 while suppressing the expression of KEAP1 (Supplementary Fig. 4i).
AKT activates NRF2 in multiple cells31,32. Our data showed that inactivating AKT with LY294002 (a PI3K inhibitor) blocked the effect of EPCR overexpression on NRF2, KEAP1, and HO-1 (Fig. 4j). Consistent results were observed when ECs were treated with APC (Supplementary Fig. 4j). These results suggested that AKT plays an important role in mediating EPCR signaling. Furthermore, we observed that depleting the expression of KEAP1 using siRNA rescued the downregulation of NRF2 and HO-1 caused by EPCR siRNA (Fig. 4k, Supplementary Fig. 4k). Consistent results were obtained after treatment with Ki696 (a KEAP1 inhibitor) (Fig. 4l). These data suggested that EPCR controlled HO-1 expression via the AKT-KEAP1-NRF2 pathway.
Considering the critical role of VEGF in promoting RNV, we assessed the effect of EPCR signaling on VEGF. The results showed that neither knockdown nor overexpressing EPCR affected the protein levels of VEGFA (Supplementary Fig. 5a, b). The data suggested that endothelial EPCR signaling might be VEGF-independent. BTB and CNC Homology 1 (BACH1) is a transcription factor that regulates oxidative stress response and heme metabolism29. Stromal Cell-Derived Factor 1 (SDF1), also known as CXCL12, has been shown to promote angiogenesis through HO-133. Thus, we further evaluated the role of BACH1 and SDF1 in EPCR signaling. The results showed that depletion of BACH1 in ECs could increase the protein levels of HO-1 (Supplementary Fig. 5c). However, the protein levels of BACH1 did not change in the retina of OIR mice compared to control mice (Supplementary Fig. 5d), and neither knockdown nor overexpressing EPCR affected the protein levels of BACH1 (Supplementary Fig. 5e, f). The data indicated that BACH1 did not participate in the endothelial EPCR signaling pathway. Consistent with a previous report34,35, the results showed that the protein levels of SDF1 increased significantly in the retinal neo-vasculature in OIR mice (Supplementary Fig. 6a, b), and knockdown of SDF-1 decreased the expression of HO-1 (Supplementary Fig. 6c). SDF-1 usually interacts with C-X-C motif chemokine receptor 4 and activates downstream AKT signaling pathways to promote angiogenesis33. Knockdown of EPCR in ECs could suppress the expression of SDF1, while overexpression of EPCR could increase the expression of SDF1 (Supplementary Fig. 6d, e). These data suggested that EPCR might regulate the expression of SDF-1 through AKT signaling. This was further confirmed as ECs treated with LY294002 exhibited reduced SDF-1 expression (Supplementary Fig. 6f). However, the knockdown of SDF1 did not reverse the increased expression of HO-1 induced by EPCR overexpression (Supplementary Fig. 6g). All these data suggested that EPCR regulated the expression of SDF-1 through of AKT signaling, and SDF-1 did not contribute to the expression of HO-1 regulated by EPCR.
Genetic ablation of endothelial Ho-1 attenuates angiogenesis
To evaluate the role of HO-1 in retinal angiogenesis, we bred Cdh5-CreERT2 mice with Ho-1flox/flox mice to generate EC-specific Ho-1 KO mice (Ho-1iΔEC). The deletion efficiency was determined using flow cytometry-sorted ECs, and the protein levels of HO-1 in ECs sorted from Ho-1iΔEC mice were significantly decreased (Supplementary Fig. 7a). The role of HO-1 in pathological retinal angiogenesis was then determined in the OIR mice. Our data showed that the area of neovascular tufts in the retina of Ho-1iΔEC mice was smaller, while the avascular area in the retina of Ho-1iΔEC mice was larger than that in Ho-1WT mice (Fig. 5a–c). Furthermore, our results showed that EdU+ ECs decreased significantly in the retina of Ho-1iΔEC mice than in Ho-1WT mice (Fig. 5d, e).
a Representative images of retinal vasculature stained with IB4 in the retina of Ho-1WT and Ho-1iΔEC mice at P17 using the OIR mice model. The orange space indicates the avascular area for two left images and neovascular tufts for two right images. Scale bar 1000 μm and 500 μm. b Quantification of the area of neovascular tufts in Ho-1WT (n = 8) and Ho-1iΔEC (n = 8) mice. Mean ± SD, p = 0.0049 (two-tailed Student’s t-test). c Quantification of the avascular area in Ho-1WT (n = 8) and Ho-1iΔEC (n = 8) mice. Mean ± SD, p = 0.0067 (two-tailed Student’s t-test). d Immunofluorescence staining for IB4 (blue), E26 transformation-specific (ETS)-related gene (red), and EdU (green) in the retina of Ho-1WT and Ho-1iΔEC mice. Proliferating (E26 transformation-specific (ETS)-related gene and EdU double positive) ECs are shown in yellow. Scale bar 50 μm. e Quantification of the proliferating ECs in the retina Ho-1WT (n = 6) and Ho-1iΔEC (n = 6) mice. Mean ± SD, p = 0.0018 (two-tailed Student’s t-test). f EdU-incorporation in HUVECs treated with EPCR adenovirus and HO-1 siRNA. DAPI (blue) identifies ECs nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. g Quantification of EdU-incorporation in HUVECs treated with EPCR adenovirus and HO-1 siRNA. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0009, p = 0.0300 (one-way ANOVA with Tukey’s multiple comparisons test). h Representative sprouting images in HUVECs treated with EPCR adenovirus and HO-1 siRNA. Scale bar 50 μm. i Quantification of sprouts length in HUVECs treated with EPCR adenovirus and HO-1 siRNA. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0013, p = 0.0320 (one-way ANOVA with Tukey’s multiple comparisons test). j Representative wound healing images in HUVECs treated with EPCR adenovirus and HO-1 siRNA. Scale bar 100 μm. k Quantification of wound healing ability in HUVECs treated with EPCR adenovirus and HO-1 siRNA. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0012, p = 0.0471 (one-way ANOVA with Tukey’s multiple comparisons test). ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
Physiological retinal vascular development was also affected in Ho-1iΔEC mice. Deleting endothelial Ho-1 reduced radial expansion of the superficial retinal vascular plexus (Supplementary Fig. 7b, c). Meanwhile, the tip cells’ filopodia and branch points were decreased in the retina of Ho-1iΔEC mice (Supplementary Fig. 7d–f). Additionally, EdU+ ECs decreased significantly in the retina of Ho-1iΔEC mice than in Ho-1WT mice (Supplementary Fig. 7g, h).
The role of HO-1 in angiogenesis was further assessed in vitro. First, the knockdown efficiency of HO-1 siRNA was evaluated (Supplementary Fig. 7i). The EdU assay showed that silencing HO-1 inhibited EC proliferation induced by the EPCR adenovirus (Fig. 5f, g). A three-dimensional endothelial spheroid assay was used to evaluate the angiogenic potential of ECs treated with EPCR adenovirus and HO-1 siRNAs. Fewer and shorter sprouts were observed in HO-1 siRNA-treated ECs than in EPCR adenovirus-treated ECs (Fig. 5h, i). Besides, ECs treated with HO-1 siRNAs exhibited impaired migration ability compared to the EPCR adenovirus treatment group (Fig. 5j, k). We also assessed the effects of HO-1 knockdown on ECs. The results showed that silencing HO-1 suppressed EC proliferation, migration, and sprouting (Supplementary Fig. 7j–o). These results collectively revealed the vital role of HO-1 in developmental angiogenesis and RNV.
Endothelial heme catabolism impacts angiogenesis
HO-1 catalyzes heme degradation into CO, biliverdin, and iron36. Catabolism of cytotoxic labile heme and the generation of CO exert cytoprotective effects37. Thus, we further assessed whether the angiogenic role of HO-1 was mediated by its enzymatic function. Here, a heme oxygenase (both HO-1 and HO-2) enzymatic inhibitor, zinc protoporphyrin IX (ZnPPIX), was used to determine its effect on pathological RNV. The data showed that ZnPPIX treatment decreased the area of neovascular tufts in the retina and increased the avascular area (Fig. 6a–c). Besides, ZnPPIX suppressed EC proliferation as EdU+ retinal ECs decreased significantly in mice treated with ZnPPIX (Fig. 6d, e).
a Representative images of retinal vasculature stained with IB4 in DMSO or ZnPPIX-treated mice at P17 using the OIR mice model. The orange space indicates the avascular area for two left images and neovascular tufts for two right images. Scale bar 1000 μm and 500 μm. b Quantification of the area of neovascular tufts in DMSO (n = 8) and ZnPPIX-treated (n = 9) mice. Mean ± SD, p = 0.0002 (two-tailed Student’s t-test). c Quantification of the avascular area in the retina of DMSO (n = 8) and ZnPPIX-treated (n = 9) mice. Mean ± SD, p = 0.0291 (two-tailed Student’s t-test). d Immunofluorescence staining for IB4 (blue), E26 transformation-specific (ETS)-related gene (red), and EdU (green) in the retina of DMSO and ZnPPIX-treated mice. Proliferating (E26 transformation-specific (ETS)-related gene and EdU double positive) ECs are shown in yellow. Scale bar 50 μm. e Quantification of the proliferating ECs in the retina of DMSO (n = 6) and ZnPPIX-treated (n = 6) mice. Mean ± SD, p = 0.0013 (two-tailed Student’s t-test). f EdU-incorporation in EPCR adenovirus and ZnPPIX-treated HUVECs. DAPI (blue) identifies EC nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. g Quantification of EdU-incorporation in EPCR adenovirus and ZnPPIX-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0020, p = 0.0422 (one-way ANOVA with Tukey’s multiple comparisons test). h Representative sprouting images of EPCR adenovirus and ZnPPIX-treated HUVECs. Scale bar 50 μm. i Quantification of sprouts length in EPCR adenovirus and ZnPPIX-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0008, p = 0.0498 (one-way ANOVA with Tukey’s multiple comparisons test). j Wound healing of EPCR adenovirus and ZnPPIX-treated HUVECs. Scale bar 100 μm. k Quantification of wound healing ability in EPCR adenovirus and ZnPPIX-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0021, p = 0.0184 (one-way ANOVA with Tukey’s multiple comparisons test). ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
Next, we determined the effect of ZnPPIX on retinal vascular expansion. ZnPPIX treatment led to a sparse vascular network and reduced radial expansion of the superficial retinal vascular plexus (Supplementary Fig. 8a, b). The tip cells’ filopodia and branch points were decreased in the retina of mice treated with ZnPPIX (Supplementary Fig. 8c–e). Meanwhile, ZnPPIX significantly suppressed ECs proliferation in the retina (Supplementary Fig. 8f, g).
Further data showed that EPCR-induced cell proliferation, migration, and sprouting of ECs in vitro were inhibited by ZnPPIX (Fig. 6f–k). Additionally, we evaluated the effect of ZnPPIX on ECs treated with HO-1 siRNA. The result showed that ZnPPIX did not affect EC proliferation, migration, and sprouting after HO-1 knockdown (Supplementary Fig. 8h–m). These data indicated that endothelial heme catabolism contributes to EPCR-mediated RNV.
Heme catabolism-derived CO contributes to pathological angiogenesis
Endogenous CO is a signaling molecule that promotes tumor growth and angiogenesis38,39. Heme catabolism in macrophages increases tumor angiogenesis through CO production, suggesting a role for CO in regulating angiogenesis37. Here, we showed that pathological retinal neovascular tufts in carbon monoxide-releasing molecule-3 (CORM3)-treated Ho-1iΔEC mice significantly increased compared with that in Ho-1iΔEC mice, while the avascular area was decreased (Fig. 7a–c). Furthermore, the data showed that EdU+ ECs also increased significantly in the CORM3-treated retina of Ho-1iΔEC mice than that in Ho-1iΔEC mice (Fig. 7d, e). Pathological retinal neovascular tufts in CORM3-treated EpcriΔEC mice also significantly increased, and avascular area decreased compared with that in EpcriΔEC mice (Supplementary Fig. 9a–c). Consistent data were acquired for the number of EdU+ ECs (Supplementary Fig. 9d, e). Meanwhile, the role of other byproducts of heme catabolism was evaluated. The data showed that bilirubin and iron supplements did not reverse the vascular changes caused by endothelial Epcr deletion (Supplementary Fig. 10).
a Representative images of retinal vasculature stained with IB4 in Ho-1WT, Ho-1iΔEC and CORM3-treated mice at P17 using the OIR mice model. The orange space indicates the avascular area for three lower images and neovascular tufts for three upper images. Scale bar 1 mm and 500 μm. b Quantification of the area of neovascular tufts in the retina of Ho-1WT (n = 8), Ho-1iΔEC (n = 8) and CORM3-treated (n = 7) mice. Mean ± SD, p < 0.0001, p = 0.0017 (one-way ANOVA with Tukey’s multiple comparisons test). c Quantification of the avascular area in the retina of Ho-1WT (n = 8), Ho-1iΔEC (n = 8) and CORM3-treated (n = 7) mice. Mean ± SD, p = 0.0076, p = 0.0423 (one-way ANOVA with Tukey’s multiple comparisons test). d Immunofluorescence staining for IB4 (blue), E26 transformation-specific (ETS)-related gene (red), and EdU (green) in the retina of Ho-1iΔEC and CORM3-treated mice. Proliferating (E26 transformation-specific (ETS)-related gene and EdU double positive) ECs are shown in yellow. Scale bar 50 μm. e Quantification of the proliferating ECs in the retina of Ho-1WT (n = 6), Ho-1iΔEC (n = 6) and CORM3-treated (n = 6) mice. Mean ± SD, p = 0.0005, p = 0.0056 (one-way ANOVA with Tukey’s multiple comparisons test). f EdU-incorporation in HO-1-depleted and CORM3-treated HUVECs. DAPI (blue) identifies EC nuclei. EdU (red) staining indicates proliferating ECs. Scale bar 200 μm. g Quantification of EdU-incorporation in HO-1-depleted and CORM3-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0024, p = 0.0341 (one-way ANOVA with Tukey’s multiple comparisons test). h Representative images of HO-1-depleted and CORM3-treated HUVECs sprouting. Scale bar 50 μm. i Quantification of sprouts length in HO-1-depleted and CORM3-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p = 0.0001, p = 0.0089 (one-way ANOVA with Tukey’s multiple comparisons test). j Wound healing of HO-1-depleted and CORM3-treated HUVECs. Scale bar 100 μm. k Quantification of wound healing ability in HO-1-depleted and CORM3-treated HUVECs. n = 3 replicates, and the experiments were repeated 3 times. Mean ± SD, p < 0.0001, p = 0.0075 (one-way ANOVA with Tukey’s multiple comparisons test). ∗∗∗∗p < 0.0001, ∗∗∗p < 0.001, ∗∗p < 0.01, ∗p < 0.05. Source data are provided as a Source Data file.
We then assessed the effect of CORM3 on postnatal retinal vasculature in Ho-1iΔEC and EpcriΔEC mice. Treatment with CORM3 could rescue the sparse vascular network and increase the radial expansion of superficial retinal vascular plexus in Ho-1iΔEC mice (Supplementary Fig. 11a, b). Compared with Ho-1iΔEC mice, tip cells’ filopodia and branch points increased in the retina of Ho-1iΔEC mice treated with CORM3 (Supplementary Fig. 11c–e). Consistent results were acquired in CORM3-treated EpcriΔEC mice (Supplementary Fig. 11f–j).
Furthermore, CORM3 rescued HO-1 silence-caused inhibition of EC proliferation, migration, and sprouting (Fig. 7f–k). Additionally, the effect of CORM3 on EPCR-deleted ECs was evaluated. The results showed that EPCR silencing inhibited EC proliferation, migration, and sprouting, which was partially reversed after treatment with CORM3 (Supplementary Fig. 12a–f). These results suggested that CO was an important contributor to EPCR-dependent endothelial heme catabolism-regulated angiogenesis.
Discussion
Therapeutic strategies for RNV are effective, but off-target effect and resistance limit their efficacy12,40. The reason is that these therapies cannot distinguish between healthy and pathological vessels. Our current study showed that EPCR was selectively detected in ECs of the retina, and EPCR transcription was activated under angiogenic stimulus. Conditional deletion of Epcr in ECs or anti-EPCR neutralizing antibody treatment attenuated pathological retinopathy. Mechanistically, EPCR-controlled endothelial heme catabolism through HO-1 and heme catabolism-derived CO plays a vital role. Notably, the EPCR ligand levels were elevated in the vitreous body or serum of patients with PDR. These observations revealed the potential therapeutic value of targeting EPCR for suppressing pathological RNV (Fig. 8).
a EPCR is specifically upregulated in retinal endothelial cells during RNV, and endothelial deletion of Epcr ameliorates RNV. b EPCR regulates the expression of HO-1 and heme catabolism-derived CO through AKT/NRF2/KEAP1 signaling. EPCR blockade decreases CO levels and inhibits retinal angiogenesis. Created in BioRender. Song, H. (2025) https://BioRender.com/r06f399.
Here, we observed that the expression of EPCR was detected specifically in the retinal ECs and was upregulated in neovascular tufts using anti-EPCR staining. EPCR was not detected in retinal pigment epithelia, photoreceptor cells, or ganglion cells. Consistent with our previous work21, we found that Epcr transcription is activated during the vessel expansion stage in the retina. Meanwhile, Epcr transcription was activated in ECs during RNV using Epcr-CreERT2; Rosa26tdT mice. These findings revealed that EPCR expression was selectively detected on retinal ECs and was upregulated under angiogenic stimulus. Vascular endothelial growth factor receptor 2 (VEGFR2) is the major receptor for VEGF guiding angiogenesis41. However, VEGFR2 is widely detected in the retina, including ECs and neurons42. Sustained suppression of VEGF would damage the photoreceptors and lead to loss of vision43,44. It is observed in the clinic that retinal atrophy occurs in patients who received anti-VEGF treatment45. The specific expression pattern of EPCR in the retina indicates that targeting EPCR is unlikely to induce off-target effects.
It is reported that Epcr + ECs are highly proliferative and major contributors to vessel development21. Conditional expression of diphtheria toxin (DTA) in Epcr+ cells delays retinal vessel extension in postnatal mice, indicating a role for EPCR in angiogenesis22. Consistent data was acquired in our study that conditional deletion of Epcr in ECs suppressed the expansion of the retinal vascular plexus. Using the OIR mice model, we showed that endothelial deletion of Epcr or EPCR neutralization antibody reduced ECs proliferation and attenuated pathological RNV. The angiogenic role of EPCR in neovascularization is also observed in hindlimb ischemia mice46. Deletion of Epcr in ECs using Tie2-Cre; Epcrflox/flox mice suppresses new vessel formation46. Given that Epcr transcription is specifically activated in proliferating ECs21, EPCR blockade could be a novel option for treating RNV with minimal effects on quiescent ECs. However, it is also noteworthy that endothelial deletion of Epcr or Ho-1 could impair the physiological vascular regeneration during OIR, as evidenced by increased vaso-obliteration observed at P17. Consistent data were acquired that inhibition of EPCR or HO-1 suppresses retinal physiological vascular development at P7. Therefore, the EPCR blocking strategy could be specific to proliferating ECs but not to pathological angiogenesis.
The mechanism by which EPCR contributes to RNV remains to be revealed. We found that the deletion of EPCR in ECs reduced the expression of HO-1 through the AKT/KEAP1/NRF2 pathway. In comparison, EPCR ligand (APC) treatment upregulated protein levels of HO-1 via AKT/KEAP1/NRF2. The data were consistent with studies showing that EPCR activates AKT signaling in the heart and multiple cells16,17,18,19. The levels of the EPCR ligand were elevated in the serum or vitreous body of patients with PDR, suggesting that it activates EPCR signaling to favor retinal neovascularization. KEAP1 is a negative regulator of NRF2, affecting the expression of NRF2 in ECs47. NRF2 is known to directly regulate the expression of HO-1 during tumor growth and angiogenesis29,48. However, its role in retinal angiogenesis is complex. NRF2 is detected in retinal ganglion cells, astrocytes, müller glial cells, and ECs, with prominent expression in retinal ganglion cells and müller glial cells during retinopathy49,50. Enhancing NRF2 activity in retinal neurons can attenuate diabetic retinopathy and inhibit RNV49. Meanwhile, endothelial deletion of Nrf2 delays vascular expansion and decreases vascular density51. Our data showed that the deletion of EPCR in ECs decreased the protein levels of NRF2 and HO-1 and inhibited the angiogenic function of ECs. These results indicate a notable difference in the cellular context of NRF2 action, and NRF2 plays a vital role in regulating HO-1 after endothelial deletion of EPCR.
Here, we showed that EC-specific deletion of Ho-1 suppressed retinal neovascularization in OIR and postnatal mice’s retina. HO-1 catalyzes heme degradation to CO, ferrous iron, and biliverdin52. While initially considered a waste product, CO is increasingly recognized as a cytoprotective and homeostatic molecule53. Heme catabolism-derived CO is a major source of endogenous CO54, which contributes to tumor metastasis and angiogenesis37. In ECs, CO is reported to increase cell growth in vitro, whereas its role in vivo is unknown38. Here, our data indicate that heme catabolism-derived CO promoted retinal angiogenesis, revealing a vital role of CO in angiogenesis. Clinically, the levels of exhaled CO significantly increase in patients with diabetes and correlate with blood glucose levels and disease duration55,56. PDR, characterized by uncontrolled RNV, is a late-stage microvascular complication of diabetes3. Elevated levels of APC in the serum or vitreous body of patients with PDR could activate EPCR signaling. This data suggests that this upregulated EPCR signaling may enhance heme catabolism and increase CO production, potentially contributing to RNV. However, the precise roles of EPCR and heme catabolism in the retinas of patients with PDR require further investigation.
In summary, we determined that EPCR is highly expressed in retinal ECs and that blocking EPCR attenuates pathological retinal angiogenesis. Mechanistically, EPCR controls heme catabolism via the KEAP1/NRF2/HO-1 pathway, and CO derived from heme catabolism plays a vital role in RNV. Hence, our findings elucidate a novel mechanism of RNV and suggest that EPCR may be a promising therapeutic target for translational application. However, these mouse models do not accurately represent the pathological progression of retinopathy in primates. Therefore, determining the therapeutic efficiency of anti-EPCR for pathological retinal angiogenesis in non-human primates is indispensable in the future. Moreover, the expression pattern of EPCR in the human retina is unknown. For translational significance, it is important to determine the expression pattern of EPCR in post-mortem retinas under diverse pathophysiological conditions.
Methods
Patient samples
The patients diagnosed with proliferative diabetic retinopathy (PDR), idiopathic epiretinal membrane (ERM), and idiopathic macular hole (MH) were included. Vitreous samples were obtained from patients who underwent vitrectomy surgery by the same surgeon. Details of the patient information are shown in Supplementary Table 1. The collected vitreous samples were placed on ice immediately and centrifuged to exclude the debris. The supernatants were aliquoted into sterile tubes and stored in liquid nitrogen. Serum samples were obtained from patients who were diagnosed with PDR. The control serum samples were obtained from patients who were diagnosed with ERM and MH. Details of the patient information are shown in Supplementary Table 2. The obtained serum samples were centrifuged at 4 °C, and the supernatants were stored at −80 °C. All surgeries were performed, and human samples were harvested in accordance with the principles in the Declaration of Helsinki. This study was approved by the institutional research ethics committee of Shanghai General Hospital ([2022]-109). Individuals provided written informed consent prior to their participation in the study.
Mice
All the experimental animals were accommodated in animal facilities, where they were subjected to 12-hour cycles of light and darkness. These animals were provided with unrestricted access to standard chow and water. The C57BL/6 J mice were purchased from Jihui Laboratory Animal Care Co., Ltd. Epcr flox/flox mice (the Cyagen Biotechnology Co., Ltd.) and Ho-1flox/flox mice (the Cyagen Biotechnology Co., Ltd.) were crossed with Cdh5 CreERT2 mice57 respectively to generate conditional knockout mice. For the lineage tracing of Epcr+ cells, the Epcr CreERT2 mice21 were crossed to Rosa26tdTomato reporter mice (the Cyagen Biotechnology Co., Ltd.) The body weight measurement and body weight analysis of mice were shown in Supplementary Table 3, 4. All animal studies were approved by the Institutional Animal Care and Use Committee at Shanghai Changhai Hospital (CHEC (A.E)2023-024).
Oxygen-induced retinopathy (OIR)
Oxygen-induced retinopathy (OIR) was performed as previously reported25. In brief, the pups were exposed to 75% oxygen with their nursing mothers for five days from P7 to P12. Then, the pups were returned to room air at P12. When the mice were returned to room air, the hypoxia-induced neovascularization was initiated and peaked on P17. Animals were euthanized, and the retinas were harvested at P17. To activate CreERT2, 50 μl of tamoxifen (2 mg/ml) was injected intraperitoneally from P10 to P12. To evaluate the anti-angiogenic role of EPCR neutralizing antibody, mice were randomly divided into three groups and injected intravitreally with 1 μl anti-EPCR antibody, PBS, and anti-IgG, respectively, at P12. Ocular surface anesthesia was performed using one drop of 0.5% proparacaine hydrochloride (ALCON-COUVREUR N.V.) before injection. ZnPPIX (25 mg/kg) and CORM3 (40 mg/kg) were injected intraperitoneally twice at P12 and P13. To detect proliferating cells, 50 μl EdU (50 mg/kg) was administered intraperitoneally 6 h before euthanasia. The neovascular tuft and avascular areas were measured using the Freehand selection and Measurement tools in Image J software and then divided by the total retinal area. Mice (P17) with body weight lower than 6 g were excluded from the analysis.
RNA-Seq sample preparation and sequencing
Total RNA from retinas and cells was extracted with RNA Easy Fast Tissue/Cell Kit (TIANGEN). Agilent 2100 Bioanalyzer and RNA nano 6000 assay kit (Agilent Technologies) were used to evaluate the quality of RNA samples. The transcriptome sequencing library was constructed through 1 ng/µl RNA randomly fragmentation, cDNA strand 1 / strand 2 synthesis, end repair, A-tailing, ligation of sequencing adapters, size selection, and library PCR enrichment. The integrity of cDNA was evaluated by Agilent 2100 Bioanalyzer. The library preparations were sequenced on an Illumina HiSeq 2500 platform (Illumina, USA). The raw data of sequences were uploaded to the NCBI Sequence Read Archive (SRA) database (GSE241239 for the retinas of OIR mice; GSE249130 for HUVECs with EPCR deletion).
Isolation of retinal ECs
Isolation of retinal ECs was performed according to protocols we previously described with some modifications21. The retinas were separated from eyeballs then minced into fine fragments and digested in Iscove’s Modified Dulbecco’s MediumIs (IMDM, LI1090-500, BioAgrio) with 1.5 mg/ml Collagenase H (11087789001, Roche), 1.25 mM CaCl2, 0.4 mM MgCl2, 1% penicillin/streptomycin and 3.5 μg/ml DNaseI (D4263, Sigma) at 37 °C for 45 min. The samples were pipetted every 10 min using a 1 ml pipette tip to ensure even digestion. The digestion mix was passed through a 40 μm nylon mesh to prepare single-cell suspensions. Cells were then incubated for 20 min with FITC-conjugated anti-CD31 (11-0317-82, Invitrogen, diluted 1:100) and PE-Cy7 conjugated anti-CD45 (25-0451-82, Invitrogen, diluted 1:200) in PBS with 5% FBS. Then, the single-cell suspensions were subjected to FACS using SONY ID7000 or LSR Fortessa (BD Biosciences). FACS data were analyzed by FlowJo software. The purity of sorted ECs was routinely checked and ensured to be more than 95%.
Real-time PCR analysis
Retinas were homogenized completely through a tissue grinding apparatus (Bioprep-24R, ALLSHENG). Total RNA was extracted from homogenized retina and HUVECs using MolPure® TRIeasy Plus Total RNA Kit (19211ES60, YEASEN). cDNA was prepared using Hifair® III 1st Strand cDNA Synthesis SuperMix for qPCR (11141ES, YEASEN) following manufacturer instructions. Quantitative PCR performed with Hieff UNICON® Universal Blue qPCR AYBR Green Master Mix (11184ES, YEASEN) in Bio-Rad CFX Manager 2.1 Detection system (Bio-Rad, Hercules, CA, USA). Relative gene expression was calculated with the ΔΔCT method and normalized to the expression of β-actin. Primer sequences are provided in Supplementary Table 5.
Retinal dissection and whole-mount staining
Eyes were fixed in 4% paraformaldehyde (PFA) for 0.5 h at RT. After dissection, retinas were blocked in a blocking buffer (1% BSA, 5% FBS, and 0.5% Triton-X-100 in PBS) for 1 h at RT. Primary antibodies were diluted in PBS containing 1% BSA and incubated with the retinas at 4 °C overnight. After being washed twice with PBS, retinas were incubated with secondary antibodies at RT for 2 h. Then, the retinas were washed and flat-mounted with Fluoromount-G (SouthernBiotech). For the labeling of EdU, an additional step was performed to detect EdU-labeled proliferative cells using the Click-It EdU kit (C10338, Invitrogen) prior to mounting. Images were acquired with a Mica confocal microscope (Leica Microsystems). For the quantification of proliferating ECs in the OIR model, high-resolution confocal images (20x magnification) of IB4, ERG, and EdU labeled retinal vasculature were acquired. Four fields were taken for quantification per retina, and the total number of EdU and ERG double-positive cells were counted by image-J software per field. Then, the mean of each retina was calculated subsequently. The mean from each group was statistically compared using GraphPad Prism, and a student’s t-test was performed to calculate the p-value.
Quantification and statistical analysis
All calculations were performed using GraphPad Prism (GraphPad software 9.0, GraphPad, Bethesda, MD, USA). Statistical analysis was performed using the two-tailed Student’s t-test and one-way analysis of variance (ANOVA), where appropriate, to compare different groups. For all bar graphs, data were presented as mean ± SD. All experiments were repeated at least three times, and P values of <0.05 were considered as statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The datasets generated during the current study are available in the GEO repository, GSE 241239 and GSE 249130. Source data are provided with this paper. The RNA-seq data generated in this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE 241239 and GSE 249130. Source data are provided with this paper.
References
Binet, F. et al. Neutrophil extracellular traps target senescent vasculature for tissue remodeling in retinopathy. Science 369, (2020).
Campochiaro, P. A. Molecular pathogenesis of retinal and choroidal vascular diseases. Prog. Retin Eye Res 49, 67–81 (2015).
Antoszyk, A. N. et al. Effect of intravitreous aflibercept vs vitrectomy with panretinal photocoagulation on visual acuity in patients with vitreous hemorrhage from proliferative diabetic retinopathy: a randomized clinical trial. Jama 324, 2383–2395 (2020).
Stahl, A. et al. Effect of intravitreal aflibercept vs laser photocoagulation on treatment success of retinopathy of prematurity: the FIREFLEYE randomized clinical trial. Jama 328, 348–359 (2022).
Yeh, S. et al. Therapies for macular edema associated with central retinal vein occlusion: a report by the American Academy of Ophthalmology. Ophthalmology 122, 769–778 (2015).
Gariano, R. F. & Gardner, T. W. Retinal angiogenesis in development and disease. Nature 438, 960–966 (2005).
Selvam, S., Kumar, T. & Fruttiger, M. Retinal vasculature development in health and disease. Prog. Retin Eye Res 63, 1–19 (2018).
Eelen, G., Treps, L., Li, X. & Carmeliet, P. Basic and therapeutic aspects of angiogenesis updated. Circ. Res 127, 310–329 (2020).
Apte, R. S., Chen, D. S. & Ferrara, N. VEGF in signaling and disease: beyond discovery and development. Cell 176, 1248–1264 (2019).
Simons, M., Gordon, E. & Claesson-Welsh, L. Mechanisms and regulation of endothelial VEGF receptor signalling. Nat. Rev. Mol. Cell Biol. 17, 611–625 (2016).
Arima, M. et al. Claudin-5 redistribution induced by inflammation leads to anti-VEGF-resistant Diabetic Macular Edema. Diabetes 69, 981–999 (2020).
Wallsh, J. O. & Gallemore, R. P. Anti-VEGF-resistant retinal diseases: a review of the latest treatment options. Cells 10, 1049 (2021).
Wang, D. et al. Long-term expansion of pancreatic islet organoids from resident procr(+) progenitors. Cell 180, 1198–1211.e1119 (2020).
Mohan Rao, L. V., Esmon, C. T. & Pendurthi, U. R. Endothelial cell protein C receptor: a multiliganded and multifunctional receptor. Blood 124, 1553–1562 (2014).
Magisetty, J. et al. Selective inhibition of activated protein C anticoagulant activity protects against hemophilic arthropathy in mice. Blood 139, 2830–2841 (2022).
Ren, D. et al. Activated protein C strengthens cardiac tolerance to ischemic insults in aging. Circ. Res 130, 252–272 (2022).
Yang, X. V. et al. Activated protein C ligation of ApoER2 (LRP8) causes Dab1-dependent signaling in U937 cells. Proc. Natl Acad. Sci. USA 106, 274–279 (2009).
Sinha, R. K. et al. Apolipoprotein E receptor 2 mediates activated protein C-induced endothelial AKT activation and endothelial barrier stabilization. Arterioscler Thromb. Vasc. Biol. 36, 518–524 (2016).
Wang, D. et al. Protein C receptor stimulates multiple signaling pathways in breast cancer cells. J. Biol. Chem. 293, 1413–1424 (2018).
Wang, D. et al. Protein C receptor is a therapeutic stem cell target in a distinct group of breast cancers. Cell Res 29, 832–845 (2019).
Yu, Q. C. et al. Activation of Wnt/β-catenin signaling by Zeb1 in endothelial progenitors induces vascular quiescence entry. Cell Rep. 41, 111694 (2022).
Yu, Q. C., Song, W., Wang, D. & Zeng, Y. A. Identification of blood vascular endothelial stem cells by the expression of protein C receptor. Cell Res 26, 1079–1098 (2016).
Connor, K. M. et al. Quantification of oxygen-induced retinopathy in the mouse: a model of vessel loss, vessel regrowth and pathological angiogenesis. Nat. Protoc. 4, 1565–1573 (2009).
Zou, J. et al. Interleukin-19 promotes retinal neovascularization in a mouse model of oxygen-induced retinopathy. Invest Ophthalmol. Vis. Sci. 63, 9 (2022).
Crespo-Garcia, S. et al. Pathological angiogenesis in retinopathy engages cellular senescence and is amenable to therapeutic elimination via BCL-xL inhibition. Cell Metab. 33, 818–832.e817 (2021).
Caprara, C. & Grimm, C. From oxygen to erythropoietin: relevance of hypoxia for retinal development, health and disease. Prog. Retin Eye Res 31, 89–119 (2012).
Brash, J. T. et al. Tamoxifen-activated CreERT impairs retinal angiogenesis independently of gene deletion. Circ. Res 127, 849–850 (2020).
Liu, C. et al. Procr functions as a signaling receptor and is essential for the maintenance and self-renewal of mammary stem cells. Cell Rep. 38, 110548 (2022).
Lignitto, L. et al. Nrf2 activation promotes lung cancer metastasis by inhibiting the degradation of Bach1. Cell 178, 316–329.e318 (2019).
Mills, E. L. et al. Itaconate is an anti-inflammatory metabolite that activates Nrf2 via alkylation of KEAP1. Nature 556, 113–117 (2018).
Dai, X. et al. Elevating CXCR7 improves angiogenic function of EPCs via Akt/GSK-3β/Fyn-mediated Nrf2 activation in diabetic limb ischemia. Circ. Res 120, e7–e23 (2017).
Lien, E. C. et al. Glutathione biosynthesis is a metabolic vulnerability in PI(3)K/Akt-driven breast cancer. Nat. Cell Biol. 18, 572–578 (2016).
Deshane, J. et al. Stromal cell-derived factor 1 promotes angiogenesis via a heme oxygenase 1-dependent mechanism. J. Exp. Med 204, 605–618 (2007).
Butler, J. M. et al. SDF-1 is both necessary and sufficient to promote proliferative retinopathy. J. Clin. Invest 115, 86–93 (2005).
Lima e Silva, R. et al. The SDF-1/CXCR4 ligand/receptor pair is an important contributor to several types of ocular neovascularization. Faseb j. 21, 3219–3230 (2007).
Otterbein, L. E., Soares, M. P., Yamashita, K. & Bach, F. H. Heme oxygenase-1: unleashing the protective properties of heme. Trends Immunol. 24, 449–455 (2003).
Consonni, F. M. et al. Heme catabolism by tumor-associated macrophages controls metastasis formation. Nat. Immunol. 22, 595–606 (2021).
Dulak, J., Deshane, J., Jozkowicz, A. & Agarwal, A. Heme oxygenase-1 and carbon monoxide in vascular pathobiology: focus on angiogenesis. Circulation 117, 231–241 (2008).
Loboda, A., Jozkowicz, A. & Dulak, J. HO-1/CO system in tumor growth, angiogenesis and metabolism - Targeting HO-1 as an anti-tumor therapy. Vasc. Pharm. 74, 11–22 (2015).
Fogli, S. et al. Clinical pharmacology of intravitreal anti-VEGF drugs. Eye (Lond.) 32, 1010–1020 (2018).
Pérez-Gutiérrez, L., Ferrara, N. Biology and therapeutic targeting of vascular endothelial growth factor A. Nat Rev Mol Cell Biol (2023).
Okabe, K. et al. Neurons limit angiogenesis by titrating VEGF in retina. Cell 159, 584–596 (2014).
Usui, Y. et al. Neurovascular crosstalk between interneurons and capillaries is required for vision. J. Clin. Invest 125, 2335–2346 (2015).
Bucher, F. et al. Antibody-mediated inhibition of tspan12 ameliorates vasoproliferative retinopathy through suppression of β-catenin signaling. Circulation 136, 180–195 (2017).
Sadda, S. R., Guymer, R., Monés, J. M., Tufail, A. & Jaffe, G. J. Anti-vascular endothelial growth factor use and atrophy in neovascular age-related macular degeneration: systematic literature review and expert opinion. Ophthalmology 127, 648–659 (2020).
Bochenek, M. L. et al. EPCR-PAR1 biased signaling regulates perfusion recovery and neovascularization in peripheral ischemia. JCI Insight 7 (2022).
Kopacz, A. et al. Keap1 governs ageing-induced protein aggregation in endothelial cells. Redox Biol. 34, 101572 (2020).
Huang, Z. et al. Nattokinase attenuates retinal neovascularization via modulation of Nrf2/HO-1 and glial activation. Invest Ophthalmol. Vis. Sci. 62, 25 (2021).
Wei, Y. et al. Nrf2 in ischemic neurons promotes retinal vascular regeneration through regulation of semaphorin 6A. Proc. Natl Acad. Sci. USA 112, E6927–E6936 (2015).
Xu, Z. et al. NRF2 plays a protective role in diabetic retinopathy in mice. Diabetologia 57, 204–213 (2014).
Wei, Y. et al. Nrf2 acts cell-autonomously in endothelium to regulate tip cell formation and vascular branching. Proc. Natl Acad. Sci. USA 110, E3910–E3918 (2013).
Ayer, A., Zarjou, A., Agarwal, A. & Stocker, R. Heme oxygenases in cardiovascular health and disease. Physiological Rev. 96, 1449–1508 (2016).
Yuan, Z., De La Cruz, L. K., Yang, X. & Wang, B. Carbon monoxide signaling: examining its engagement with various molecular targets in the context of binding affinity, concentration, and biologic response. Pharm. Rev. 74, 823–873 (2022).
Otterbein, L. E., Foresti, R. & Motterlini, R. Heme oxygenase-1 and carbon monoxide in the heart: the balancing act between danger signaling and pro-survival. Cir. c. Res 118, 1940–1959 (2016).
Paredi, P., Biernacki, W., Invernizzi, G., Kharitonov, S. A. & Barnes, P. J. Exhaled carbon monoxide levels elevated in diabetes and correlated with glucose concentration in blood: a new test for monitoring the disease? Chest 116, 1007–1011 (1999).
Cheng, S. et al. Exhaled carbon monoxide and risk of metabolic syndrome and cardiovascular disease in the community. Circulation 122, 1470–1477 (2010).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486 (2010).
Acknowledgements
Thanks to Yi Arial Zeng (State Key Laboratory of Cell Biology, Chinese Academy of Sciences) for kindly providing Epcr-CreERT2 mice and technical assistance; to Youheng Wei (State Key Laboratory of Genetic Engineering, Fudan University, Shanghai, China) for providing technical assistance. This work was supported by grants from National Natural Science Foundation of China (82171081, 82271106, 82388101, U22A20311, 82171076), National Key R&D Program (2022YFC2502800), Shanghai Municipal Education Commission (2023ZKZD18), Shanghai Science and Technology Committee (22ZR1478200), Shanghai Changhai Hospital (2023YQ01). Figure 1a, Fig. 8, and Supplementary Fig. 1d, j were created using BioRender.com.
Author information
Authors and Affiliations
Contributions
Conceptualization, H.S., X.D.S., W.S., H.J.Z., and F.L.; Methodology, Q.L., X.S., and A.G.; investigation, H.S., Q.L., X.G., Z.F., W.Z., M.W., Y.L., H.R.Z., Z.N., L.Z., H.M.Z., and Y.J.; Funding acquisition, X.D.S., H.S. and W.S.; Writing-original draft, H.S and Q.L.; Writing-review & editing, X.D.S., F.Z., F.L., and X.L.; Supervision, H.S., X.D.S., X.L. and F.Z. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Jian-xing Ma, Abolfazl Zarjou and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, H., Li, Q., Gui, X. et al. Endothelial protein C receptor promotes retinal neovascularization through heme catabolism. Nat Commun 16, 1603 (2025). https://doi.org/10.1038/s41467-025-56810-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-56810-0
This article is cited by
-
Multi-omics integrated analysis identifies causal risk factors and therapeutic targets for diabetic retinopathy
Journal of Translational Medicine (2025)