Abstract
Chromosome synapsis is an evolutionarily conserved process essential for meiotic recombination. HORMAD1 and HORMAD2, which monitor chromosome asynapsis by localizing to unsynapsed chromosome axes, are removed from synapsed chromosome axes by TRIP13, though the biological significance of this process remains unclear. We show that when HORMAD1 and HORMAD2 are retained on synapsed chromosome axes, they recruit BRCA1, activate chromosome asynapsis checkpoint, and trigger oocyte elimination. Unexpectedly, N-terminal tagging retains HORMAD1 and HORMAD2 on synapsed chromosome axes without triggering oocyte elimination due to defective BRCA1 recruitment. Mechanistically, HORMAD1 co-immunoprecipitates with BRCA1 readily, not through the canonical closure motif-binding mode but via an interface on its HORMA domain near the N-terminus. HORMAD2 co-immunoprecipitates with BRCA1 weakly but also regulates its recruitment. Collectively, the TRIP13-dependent removal of HORMAD1 and HORMAD2 from synapsed chromosome axes is essential for female fertility, preventing aberrant chromosome asynapsis checkpoint activation and unintended oocyte elimination.
Similar content being viewed by others
Introduction
Together with the generation of programmed DNA double-strand breaks (DSBs) in the meiotic prophase, synapsis occurs between homologous chromosomes, which are used as templates for DSB repair through homologous recombination1. This process, known as meiotic recombination, is essential for generating the obligate crossovers between homologous chromosomes, which are crucial for proper chromosome segregation during the first meiotic division2. Unrepaired DSBs and chromosome asynapsis accumulate in recombination-defective meiocytes, significantly threatening genomic integrity if meiosis continues3. Therefore, meiotic recombination is tightly monitored by checkpoints that trigger the elimination of recombination-defective meiocytes.
In females, recombination-defective oocytes are eliminated before establishing the primordial follicle pool soon after birth4,5,6,7,8, but the mechanisms of oocyte meiotic recombination checkpoints are not fully understood. Unrepaired DSBs are believed to be the major trigger for eliminating recombination-defective oocytes, which activate CHK2-dependent DNA damage checkpoint to promote oocyte apoptosis5,6,8. Deficiency for CHK2 or downstream apoptotic proteins rescues the survival of recombination-defective oocytes, such as Dmc1−/− oocytes5,8. Besides unrepaired DSBs, chromosome asynapsis also exists in recombination-defective oocytes. We have shown recently that BRCA1-dependent chromosome asynapsis checkpoint plays a predominant role in eliminating recombination-defective oocytes9. Brca1−/− rescues the survival of Dmc1−/− oocytes much more efficiently than Chk2−/−. Although BRCA1 localizes to DSBs in somatic cells, it localizes to unsynapsed chromosome axes in a DSB-independent manner9. BRCA1 promotes ATR activity on unsynapsed chromosome axes, possibly critical for chromosome asynapsis checkpoint activation9. It remains elusive how this checkpoint is activated by chromosome asynapsis.
HORMAD1 and HORMAD2, two meiotic proteins with HORMA domains that localize specifically at unsynapsed chromosome axes, are likely the most upstream regulators of the chromosome asynapsis checkpoint by monitoring chromosome asynapsis10,11,12,13. Consistent with this idea, HORMAD1 regulates BRCA1’s localization at unsynapsed chromosome axes in oocytes9. In addition, Hormad1−/− fully rescues the survival of Dmc1−/− oocytes and Hormad2−/− partially rescues the survival of Dmc1−/− oocytes6,14. However, it remains elusive how HORMAD1, and whether HORMAD2, regulates BRCA1-dependent chromosome asynapsis checkpoint in oocytes. In particular, since HORMAD1 also promotes DSB formation and HORMAD1 and HORMAD2 suppress inter-sister chromatid recombination6,7,15, it is still unclear if HORMAD1 and HORMAD2 directly activate the chromosome asynapsis checkpoint.
Upon completion of chromosome synapsis, HORMAD1 and HORMAD2 are removed from synapsed chromosome axes by TRIP13, which engages the N-terminus of HORMAD1 and HORMAD2 and remodels their HORMA domains from “closed” to “open” conformation to release them from their partner proteins on chromosome axes16. However, the biological significance of TRIP13-dependent removal of HORMAD1 and HORMAD2 from synapsed chromosome axes is unclear. As expected, HORMAD1 and HORMAD2 are retained on synapsed chromosome axes in Trip13 mutant mice17,18,19. Female Trip13 mutant mice are infertile due to rapid oocyte elimination soon after birth17,19, but the contribution of HORMAD1 and HORMAD2 on synapsed chromosome axes to the elimination of Trip13 mutant oocytes has not been examined. Intriguingly, Trip13 mutant mice also have DSB repair defects for unknown reasons17,19. In Trip13Gt/Gt mice (also known as Trip13Mod/Mod mice) that are homozygous for a Trip13 hypomorphic allele, DSB repair defects but not chromosome synapsis defects exist, and γH2AX signals are observed before Trip13Gt/Gt oocytes are eliminated17,19. Together with the observation that Chk2−/− rescues the survival of Trip13Gt/Gt oocytes, it is believed that Trip13Gt/Gt oocytes are eliminated by the CHK2-dependent DNA damage checkpoint5. Similar to Chk2−/−, Hormad2−/− also rescues the survival of Trip13Gt/Gt oocytes6. Using RAD51 as a marker for DSBs, it has been shown that Hormad2−/− lifts the suppression on inter-sister chromatid recombination so that DSBs are repaired, and DNA damage checkpoint is not activated in Trip13Gt/Gt Hormad2−/− oocytes6. However, a recent study challenges this idea by showing that RAD51 is not a suitable maker for DSB20, suggesting that Hormad2−/− may rescue the survival of Trip13Gt/Gt oocytes through other mechanisms.
In this study, we examine the potential direct contribution of HORMAD1 and HORMAD2 on synapsed chromosome axes to eliminating Trip13−/− oocytes independent of the DNA damage checkpoint. We generate Trip13−/− mice and show that DSB repair is delayed but completed before Trip13−/− oocytes are eliminated, suggesting that Trip13−/− oocytes are not eliminated by the DNA damage checkpoint. Instead, HORMAD1 and HORMAD2 recruit BRCA1 to synapsed chromosome axes, promoting the elimination of Trip13−/− oocytes by activating the chromosome asynapsis checkpoint. Therefore, TRIP13-dependent removal of HORMAD1 and HORMAD2 from synapsed chromosome axes is required to prevent aberrant activation of chromosome asynapsis checkpoint that triggers oocyte elimination. In addition, we reveal that HORMAD1 co-immunoprecipitates with BRCA1 through a unique interface on its HORMA domain that is likely blocked by N-terminal tagging, providing mechanistic insight into how BRCA1-dependent chromosome asynapsis checkpoint is activated in Trip13−/− oocytes.
Results
Trip13 −/− oocytes are not eliminated by the DNA damage checkpoint
To examine the mechanisms underlying oocyte elimination caused by TRIP13 deficiency, we generated Trip13−/− mice (Fig. S1a, b). Like Trip13Gt/Gt mice, female Trip13−/− mice were infertile, and atrophied ovaries were observed at postnatal day (PD) 21 (Fig. 1a). We continued to examine the DSB repair status in the meiotic prophase of Trip13−/− oocytes. In contrast to wild-type (WT) oocytes, several DSB markers, including RAD51, DMC1, and RPA2, were retained in the pachytene stages of Trip13−/− oocytes (Fig. 1b–g). Therefore, consistent with the observations in Trip13Gt/Gt oocytes, Trip13−/− oocytes have DSB repair defects in the meiotic prophase.
a Representative images of H&E staining of ovary sections from control and Trip13−/− mice at PD21. Scale bar, 100 μm. Representative images of the RPA2 and SYCP3 staining (b), RAD51 and SYCP3 staining (d), and DMC1 and SYCP3 staining (f) in chromosome spreads of control and Trip13−/− oocytes at pachytene and diplotene stages. Scale bar, 10 μm. Quantification of RPA2 foci (c), RAD51 foci (e), and DMC1 foci (g) in chromosome spreads of control and Trip13-/- oocytes at pachytene and diplotene stages. 25 pachytene nuclei and 20 diplotene nuclei of each genotype were analyzed. Representative images of γH2AX and SYCP3 staining in chromosome spreads of control and Trip13-/- oocytes at pachytene (h) and diplotene (i) stages. Scale bar, 10 μm. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
Despite the presence of unrepaired DSBs in pachytene stages, Trip13−/− oocytes progressed to diplotene stages. Unexpectedly, all DSB markers disappeared in the diplotene stages of Trip13−/− oocytes (Fig. 1b–g), suggesting that the DSB repair is delayed but is eventually completed before Trip13−/− oocytes are eliminated. Since no DSBs exist before oocyte elimination, Trip13−/− oocytes are unlikely to be eliminated by the DNA damage checkpoint.
Accompanied by the completion of DSB repair in pachytene stages of WT oocytes, γH2AX signals were diminished. γH2AX signals were observed in the pachytene stages of Trip13−/− oocytes (Fig. 1h), which were believed to be caused by the unrepaired DSBs at this stage. Surprisingly, γH2AX signals persisted when DSB repair was completed in the diplotene stages of Trip13−/− oocytes (Fig. 1i). Therefore, the γH2AX signals in Trip13−/− oocytes are caused not by unrepaired DSBs but by other signaling pathways that potentially contribute to the elimination of Trip13−/− oocytes.
N-terminal tagging retains HORMAD1 and HORMAD2 on synapsed chromosome axes but does not trigger oocyte elimination
Besides DSB repair defects, Trip13−/− oocytes also have ectopic retention of HORMAD1 and HORMAD2 on synapsed chromosome axes18,19. Since Trip13−/− oocytes are unlikely to be eliminated by the DNA damage checkpoint, HORMAD1 and HORMAD2 on synapsed chromosome axes may trigger the elimination of Trip13−/− oocytes. To test this possibility, we attempted to retain HORMAD1 and HORMAD2 on synapsed chromosome axes without generating DSB repair defects. Previous studies have revealed that bulky N-terminal tagging disrupts the release of HORMAD1 and HORMAD2 from synapsed chromosome axes, presumably by blocking the engagement of TRIP1316. We generated Hormad1SFB/SFB and Hormad2SFB/SFB mice by adding S-FLAG-streptavidin binding protein (SFB) tags to the N-termini of HORMAD1 and HORMAD2, respectively (Figure S2a, b). As predicted, HORMAD1 was retained on synapsed chromosome axes of Hormad1SFB/SFB oocytes (Figure S2c, d), and HORMAD2 was retained on synapsed chromosome axes of Hormad2SFB/SFB oocytes (Figure S2e,f). Hormad1SFB/SFB Hormad2SFB/SFB mice (from now on referred to as Hormad1&2SFB/SFB mice) were then generated by crossing these two mice.
Similar to Trip13−/− oocytes, both HORMAD1 and HORMAD2 were retained on synapsed chromosome axes of Hormad1&2SFB/SFB oocytes (Fig. 2a, b). Unlike Trip13−/− oocytes with DSB repair defects, the DSB repair kinetics were indistinguishable between WT and Hormad1&2SFB/SFB oocytes (Fig. 2c–h). Therefore, Hormad1&2SFB/SFB oocytes are suitable for examining if ectopic retention of HORMAD1 and HORMAD2 on synapsed chromosome axes leads to oocyte elimination. Unexpectedly, in contrast to the atrophied ovaries in Trip13−/− female mice, oocytes at all developmental stages could be observed in the ovaries of Hormad1&2SFB/SFB female mice at PD21 (Fig. 2i). The numbers of follicles were similar between control and Hormad1&2SFB/SFB female mice at PD21 (Fig. 2j). The numbers of oocytes were also similar between control and Hormad1&2SFB/SFB female mice at PD4, one day before the establishment of the primordial follicle pool (Figure S3a, b). Therefore, ectopic retention of HORMAD1 and HORMAD2 on synapsed chromosome axes by N-terminal tagging does not trigger oocyte elimination.
Representative images of HORMAD1 and SYCP3 staining (a) and HORMAD2 and SYCP3 staining (b) in chromosome spreads of control, Trip13−/−, and Hormad1&2SFB/SFB oocytes at pachytene stages. Scale bar, 10 μm. Representative images of RPA2 and SYCP3 staining (c), RAD51 and SYCP3 staining (e), and DMC1 and SYCP3 staining (g) in chromosome spreads of control and Hormad1&2SFB/SFB oocytes at pachytene and diplotene stages. Scale bar, 10 μm. Quantification of RPA2 foci (d), RAD51 foci (f), and DMC1 foci (h) in chromosome spreads of control and Hormad1&2SFB/SFB oocytes at pachytene and diplotene stages. 25 pachytene nuclei and 20 diplotene nuclei of each genotype were analyzed. i. Representative images of H&E staining of ovary sections from control, Trip13−/−, and Hormad1&2SFB/SFB mice at PD21. Scale bar, 100 μm. j. Quantification of primordial and total follicle numbers in control, Trip13−/−, and Hormad1SFB/SFB mice at P21. 3 mice of each genotype were analyzed. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
N-terminal tagging of HORMAD1 and HORMAD2 rescues the survival of Trip13 −/− oocytes
The viability of Hormad1&2SFB/SFB oocytes challenges the idea that HORMAD1 and HORMAD2 on synapsed chromosome axes trigger the elimination of Trip13−/− oocytes. To test if Trip13−/− oocytes are eliminated by signaling pathways unrelated to HORMAD1 and HORMAD2 on synapsed chromosome axes, we generated Trip13−/− Hormad1SFB/SFB Hormad2SFB/SFB mice (from now on referred to as Trip13−/− Hormad1&2SFB/SFB mice) and examined if Trip13−/− oocytes are still eliminated when HORMAD1 and HORMAD2 are tagged at their N-termini. Interestingly, oocytes at all developmental stages could be observed in the ovaries of Trip13−/− Hormad1&2SFB/SFB female mice at PD21 (Fig. 3a). Unlike Trip13−/− female mice, many follicles were present in Trip13−/− Hormad1&2SFB/SFB female mice at PD21 (Fig. 3b), and many oocytes were also present in Trip13−/− Hormad1&2SFB/SFB female mice at PD4 as well (Figure S4a-b). Like Trip13−/− oocytes, DSB repair was delayed but was eventually completed in Trip13−/− Hormad1&2SFB/SFB oocytes (Fig. 3c–h). Therefore, N-terminal tagging of HORMAD1 and HORMAD2 rescues the survival of Trip13−/− oocytes without promoting DNA repair, which further supports the idea that Trip13−/− oocytes are not eliminated by the DNA damage checkpoint.
a Representative images of the H&E staining of ovary sections from control, Trip13−/−, and Trip13−/− Hormad1&2SFB/SFB mice at PD21. Scale bar, 100 μm. b Quantification of primordial and total follicle numbers in control, Trip13−/−, and Trip13−/− Hormad1SFB/SFB mice at P21. The control and Trip13−/− data were the same as in Fig. 2j. 3 mice of each genotype were analyzed. Representative images of RPA2 and SYCP3 staining (c), RAD51 and SYCP3 staining (e), and DMC1 and SYCP3 staining (g) in chromosome spreads of Trip13−/− and Trip13−/− Hormad1&2SFB/SFB oocytes at pachytene and diplotene stages. Scale bar, 10 μm. Quantification of RPA2 foci (d), RAD51 foci (f), and DMC1 foci (h) in chromosome spreads of Trip13−/− and Trip13−/− Hormad1&2SFB/SFB oocytes at pachytene and diplotene stages. 25 pachytene nuclei and 20 diplotene nuclei of each genotype were analyzed. Representative images of γH2AX and SYCP3 staining in chromosome spreads of control, Trip13−/−, Hormad1&2SFB/SFB, and Trip13−/− Hormad1&2SFB/SFB oocytes at pachytene (i) and diplotene (j) stages. Scale bar, 10 μm. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
Besides blocking TRIP13’s remodeling, N-terminal tagging of HORMAD1 and HORMAD2 may also disrupt other functions of HORMAD1 and HORMAD2 and allow the survival of Hormad1&2SFB/SFB oocytes. Before Trip13−/− oocytes were eliminated, γH2AX signals persisted while DSB repair was completed. Interestingly, γH2AX signals were diminished in pachytene and diplotene stages of both Hormad1&2SFB/SFB and Trip13−/− Hormad1&2SFB/SFB oocytes (Fig. 3i, j). Therefore, although not promoting DSB repair, N-terminal tagging of HORMAD1 and HORMAD2 disrupts the signaling that is responsible for the γH2AX signals in Trip13−/− oocytes, which is potentially critical for the elimination of Trip13−/− oocytes. Taken together, HORMAD1 and HORMAD2 on synapsed chromosome axes likely trigger the elimination of Trip13−/− oocytes, but N-terminal tagging of HORMAD1 and HORMAD2 disrupts the key signaling for this process.
Trip13 −/− oocytes are eliminated by BRCA1-dependent chromosome asynapsis checkpoint
In WT oocytes, HORMAD1 and HORMAD2 monitor chromosome asynapsis by localizing specifically at unsynapsed chromosome axes10,11,12,13. By recruiting BRCA1 to unsynapsed chromosome axes in oocytes, HORMAD1 is a potential regulator of BRCA1-dependent chromosome asynapsis checkpoint, which triggers the elimination of recombination-defective oocytes by activating ATR9. Interestingly, besides HORMAD1 and HORMAD2, BRCA1 and ATR were also identified on synapsed chromosome axes in Trip13−/− oocytes (Fig. 4a, b). These observations suggest that BRCA1-dependent chromosome asynapsis checkpoint is activated on synapsed chromosome axes, which may trigger the elimination of Trip13−/− oocytes. In contrast to Trip13−/− oocytes, BRCA1 and ATR were not retained on synapsed chromosome axes in Hormad1&2SFB/SFB oocytes or Trip13−/− Hormad1&2SFB/SFB oocytes (Fig. 4a, b). It is possible that N-terminal tagging of HORMAD1 and HORMAD2 disrupts the recruitment of BRCA1, prevents the activation of BRCA1-dependent chromosome asynapsis checkpoint, and rescues the survival of Trip13−/− oocytes.
Representative images of BRCA1 and SYCP3 staining (a) and ATR and SYCP3 staining (b) in chromosome spreads of control, Trip13−/−, Hormad1&2SFB/SFB, and Trip13−/− Hormad1&2SFB/SFB oocytes at pachytene stages. Scale bar, 10 μm. c. Representative images of H&E staining of ovary sections from control, Trip13−/−, and Trip13−/− Brca1f/- Stra8-GFPCre mice at PD21. Scale bar, 100 μm. d Quantification of primordial and total follicle numbers in control, Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre mice at P21. The control and Trip13−/− data were the same as in Fig. 2j. 3 mice of each genotype were analyzed. Representative images of RPA2 and SYCP3 staining (e), RAD51 and SYCP3 staining (g), and DMC1 and SYCP3 staining (i) in chromosome spreads of Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre oocytes at pachytene and diplotene stages. Scale bar, 10 μm. Quantification of RPA2 foci (f), RAD51 foci (h), and DMC1 foci (j) in chromosome spreads of Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre oocytes at pachytene and diplotene stages. 25 pachytene nuclei and 20 diplotene nuclei of each genotype were analyzed. Representative images of BRCA1 and SYCP3 staining (k) and ATR and SYCP3 staining (l) in chromosome spreads of Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre oocytes at pachytene stages. Scale bar, 10 μm. Representative images of γH2AX and SYCP3 staining in chromosome spreads of Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre oocytes at pachytene (m) and diplotene (n) stages. Scale bar, 10 μm. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
To further interrogate if the BRCA1-dependent chromosome asynapsis checkpoint is indeed responsible for the elimination of Trip13−/− oocytes, we examined if Brca1−/− rescues the survival of Trip13−/− oocytes by conditional BRCA1 depletion in oocytes using Stra8-GFPCre, which has been shown to disrupt BRCA1-dependent chromosome asynapsis checkpoint without affecting meiotic recombination9. Similar to Trip13−/− Hormad1&2SFB/SFB female mice, oocytes at all developmental stages could be observed in the ovaries of Trip13−/− Brca1f/- Stra8-GFPCre female mice at PD21 (Fig. 4c). Many follicles were present in Trip13−/− Brca1f/- Stra8-GFPCre female mice at PD21 (Fig. 4d), and many oocytes were also present in Trip13−/− Brca1f/- Stra8-GFPCre female mice at PD4 as well (Figure S5a, b). Compared with Trip13−/− oocytes, the DSB repair kinetics were not altered (Fig. 4e–j), but BRCA1 or ATR were not retained on synapsed chromosome axes in Trip13−/− Brca1f/- Stra8-GFPCre oocytes (Fig. 4k, l). Therefore, Trip13−/− oocytes are indeed eliminated by the BRCA1-dependent chromosome asynapsis checkpoint.
Similar to Trip13−/− Hormad1&2SFB/SFB female mice, γH2AX signals were diminished in pachytene and diplotene stages of Trip13−/− Brca1f/- Stra8-GFPCre oocytes (Fig. 4m, n). Therefore, the BRCA1-dependent chromosome asynapsis checkpoint is responsible for the γH2AX signals in Trip13−/− oocytes. Since H2AX is a known substrate of ATR in DNA damage response21, BRCA1-dependent ATR recruitment on synapsed chromosome axes is likely responsible for the γH2AX signals in Trip13−/− oocytes, which is disrupted by N-terminal tagging of HORMAD1 and HORMAD2.
Hormad2 −/− rescues the survival of Trip13 −/− oocytes by disrupting BRCA1-dependent chromosome asynapsis checkpoint
Previous studies have shown that Hormad2−/− rescues the survival of Trip13Gt/Gt oocytes by promoting DSB repair through inter-sister chromatid recombination so that the DNA damage checkpoint is not activated6. Since we have shown that the BRCA1-dependent chromosome asynapsis checkpoint, but not the DNA damage checkpoint, is responsible for the elimination of Trip13−/− oocytes, we re-examined if and how Hormad2−/− rescues the survival of Trip13−/− oocytes.
Similar to previous studies, many follicles were still present in Trip13−/− Hormad2−/− female mice at PD21 (Fig. 5a, b). However, there was no difference in DSB repair kinetics between Trip13−/− and Trip13−/− Hormad2−/− oocytes (Fig. 5c–h), suggesting that Hormad2−/− rescues the survival of Trip13−/− oocytes without promoting DSB repair. On the contrary, the signals of BRCA1 and ATR were significantly decreased on synapsed chromosome axes in Trip13−/− Hormad2−/− oocytes (Fig. 5i, j), and γH2AX signals were largely diminished in pachytene and diplotene stages of Trip13−/− Hormad2−/− oocytes (Fig. 5k, l). Therefore, Hormad2−/− rescues the survival of Trip13−/− oocytes by disrupting the BRCA1-dependent chromosome asynapsis checkpoint.
a Representative images of H&E staining of ovary sections from control, Trip13−/−, and Trip13−/− Hormad2−/− mice at PD21. Scale bar, 100 μm. b Quantification of primordial and total follicle numbers in control, Trip13−/−, and Trip13−/− Hormad2−/− mice at P21. The control and Trip13−/− data were the same as in Fig. 2j. 3 mice of each genotype were analyzed. Representative images of RPA2 and SYCP3 staining (c), RAD51 and SYCP3 staining (e), and DMC1 and SYCP3 staining (g) in chromosome spreads of Trip13−/− and Trip13−/− Hormad2−/− oocytes at pachytene and diplotene stages. Scale bar, 10 μm. Quantification of RPA2 foci (d), RAD51 foci (f), and DMC1 foci (h) in chromosome spreads of Trip13−/− and Trip13−/− Hormad2−/− oocytes at pachytene and diplotene stages. 25 pachytene nuclei and 20 diplotene nuclei of each genotype were analyzed. Representative images of BRCA1 and SYCP3 staining (i) and ATR and SYCP3 staining (j) in chromosome spreads of Trip13−/− and Trip13−/− Hormad2−/− oocytes at pachytene stages. Scale bar, 10 μm. Representative images of γH2AX and SYCP3 staining in chromosome spreads of Trip13−/− and Trip13−/− Hormad2−/− oocytes at pachytene (k) and diplotene (l) stages. Scale bar, 10 μm. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
N-terminal tagging of HORMAD1 and HORMAD2 disrupts BRCA1’s localization on unsynapsed chromosome axes in both sexes but specifically compromises MSCI in spermatocytes
In WT oocytes, HORMAD1 and HORMAD2 localize to unsynapsed chromosome axes and recruit BRCA1, which is critical for meiotic recombination9,13. In Hormad1&2SFB/SFB oocytes, N-terminal tagged HORMAD1 and HORMAD2 failed to recruit BRCA1 when retained on synapsed chromosome axes. Interestingly, BRCA1’s signal at unsynapsed chromosome axes was also significantly decreased in Hormad1&2SFB/SFB oocytes (Fig. S6a, b), suggesting that N-terminal tagging of HORMAD1 and HORMAD2 disrupts BRCA1’s localization on both synapsed and unsynapsed chromosome axes in oocytes. Although BRCA1’s localization at unsynapsed chromosome axes was compromised, meiotic recombinase loading and DSB repair were normal in Hormad1&2SFB/SFB oocytes (Fig. 2c–h). It is likely that the residual BRCA1 at unsynapsed chromosome axes is sufficient for supporting meiotic recombination in Hormad1&2SFB/SFB oocytes.
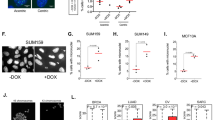
Unlike Hormad1&2SFB/SFB female mice, Hormad1&2SFB/SFB male mice were infertile. No sperm was present in the epididymides, and the testes were smaller than those in WT mice (Fig. 6a–c). The meiotic progression in Hormad1&2SFB/SFB male mice was arrested at mid-pachytene stages, as spermatocytes at late pachytene stages could not be observed (Fig. 6d, e). Similar to Hormad1&2SFB/SFB oocytes, BRCA1’s signal at unsynapsed chromosome axes was significantly decreased in Hormad1&2SFB/SFB spermatocytes (Fig. S6c, d), but meiotic recombinase loading and DSB repair on autosomes were normal (Fig. 6f–i). No γH2AX signals were present on autosomes in Hormad1&2SFB/SFB spermatocytes at mid-pachytene stages (Fig. 6e). The residual BRCA1 at unsynapsed chromosome axes is likely sufficient for supporting meiotic recombination on autosomes of Hormad1&2SFB/SFB spermatocytes as well.
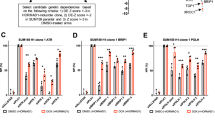
a Representative images of H&E staining of cauda epididymis sections from control and Hormad1&2SFB/SFB mice at PD60. Scale bars, 100 μm. b Representative images of control and Hormad1&2SFB/SFB testes at PD28. c The ratios of testis weight to body weight in control and Hormad1&2SFB/SFB mice at PD28. 3 mice of each genotype were analyzed. d Representative images of H&E staining of testis sections from control and Hormad1&2SFB/SFB mice at PD60. Scale bars, 100 μm. e Representative images of γH2AX and SYCP3 staining in chromosome spreads of control and Hormad1&2SFB/SFB spermatocytes at leptotene, zygotene, pachytene, and diplotene stages. Scale bar, 10 μm. Representative images of DMC1 and SYCP3 staining (f) and RAD51 and SYCP3 staining (h) in chromosome spreads of control and Hormad1&2SFB/SFB spermatocytes at zygotene stages. Scale bar, 10 μm. Quantification of DMC1 foci (g) and RAD51 foci (i) in chromosome spreads of control and Hormad1&2SFB/SFB spermatocytes at zygotene stages. 20 zygotene nuclei of each genotype were analyzed. j The mean of Log2 fold change of gene expression level on each chromosome in control and Hormad1&2SFB/SFB spermatocytes. Representative images of BRCA1 and SYCP3 staining (k) and ATR and SYCP3 staining (l) in chromosome spreads of control and Hormad1&2SFB/SFB spermatocytes at pachytene stages. Scale bar, 10 μm. All statistical data are presented as mean ± SD. P value, two-tailed Student’s t test. Source data are provided as a Source Data file.
In the male meiotic prophase, the X and Y chromosomes form an XY body and become transcriptionally silenced, a phenomenon known as meiotic sex chromosome inactivation (MSCI)22. In Hormad1&2SFB/SFB spermatocytes, no intact XY body could be observed (Fig. 6e). RNA-sequencing revealed that the transcription of X and Y chromosomes, but not autosomes, were specifically elevated (Fig. 6j). Therefore, MSCI was defective in Hormad1&2SFB/SFB spermatocytes, which is likely the cause for the arrest of meiotic progression. BRCA1 is known to be important for MSCI. BRCA1 localizes to unsynapsed axes of XY chromosomes and initiates ATR-dependent signaling, which spreads from chromosome axes to chromosome loops to silence the entire XY chromosomes. Like autosomes, BRCA1’s signal at unsynapsed axes of XY chromosomes was significantly decreased in Hormad1&2SFB/SFB spermatocytes (Fig. 6k). ATR’s signal, found at both unsynapsed axes and loops of XY chromosomes in WT spermatocytes, was diminished in Hormad1&2SFB/SFB spermatocytes (Fig. 6l). Therefore, the defective BRCA1-dependent ATR signaling is responsible for the MSCI defects in Hormad1&2SFB/SFB spermatocytes. Taken together, N-terminal tagging of HORMAD1 and HORMAD2 disrupts the recruitment of BRCA1 to unsynapsed chromosome axes. Despite being able to support meiotic recombinase loading and DSB repair on autosomes, the residual BRCA1 is not sufficient for initiating MSCI on XY chromosomes.
HORMAD1 co-immunoprecipitates with BRCA1 through a unique interface that is likely blocked by N-terminal tagging
To examine how HORMAD1 and HORMAD2 recruit BRCA1 to chromosome axes and why N-terminal tagging of HORMAD1 and HORMAD2 disrupts the recruitment, we examined the interacting proteins of HORMAD1 and HORMAD2 through tandem affinity purifications followed by mass spectrometry analyses using Hormad1SFB/+ testes and Hormad2SFB/SFB testes, respectively. Consistent with the known interaction between HORMAD1 and HORMAD2, they were identified in reciprocal purifications (Fig. 7a, b). Some other known interacting proteins of HORMAD1 and HORMAD2 were also identified, including SYCP2 and SYCP3, axial/lateral elements of the synaptonemal complex, as well as STAG3 and SMC3, components of meiotic cohesins (Fig. 7a, b)23. Interestingly, BRCA1 was identified in the purifications of both HORMAD1 and HORMAD2 (Fig. 7a, b), suggesting that HORMAD1 and HORMAD2 may recruit BRCA1 to chromosome axes through physical interactions. Given that N-terminal tagging of HORMAD1 and HORMAD2 disrupts the recruitment of BRCA1 to chromosome axes, we examined if HORMAD1 and HORMAD2 indeed co-immunoprecipitate BRCA1 using both N-terminal and C-terminal SFB-tagged proteins. When expressed in 293 T cells, C-terminal SFB-tagged HORMAD1 co-immunoprecipitates with BRCA1 readily, but N-terminal SFB-tagged HORMAD1 co-immunoprecipitates with BRCA1 weakly (Fig. 7c). Compared with HORMAD1, both N-terminal and C-terminal SFB-tagged HORMAD2 co-immunoprecipitates with BRCA1 weakly (Fig. 7c). These observations suggest that HORMAD1, but not HORMAD2, predominantly co-immunoprecipitates with BRCA1, and N-terminal tagging of HORMAD1 disrupts the co-immunoprecipitation. However, it remains to be studied if HORMAD1 directly binds BRCA1.
Lists of proteins identified by mass spectrometry after tandem affinity purifications using Hormad1SFB/+ testes (a) and Hormad2SFB/SFB testes (b). The complete lists of proteins identified by mass spectrometry are provided in Supplementary Data 1, 2. c Co-IP analyses of N-terminal SFB-tagged (SFBN) and C-terminal SFB-tagged (SFBC) HORMAD1 and HORMAD2 with BRCA1 in 293 T cells. d A schematic representation of WT and deletion mutants of HORMAD1. e. Co-IP analyses of C-terminal SFB-tagged (SFBC) HORMAD1 deletion mutants with BRCA1 in 293 T cells. f Co-IP analyses of C-terminal SFB-tagged (SFBC) HORMAD1 carrying mutations at key residues critical for closure motif binding with BRCA1 in 293 T cells. g Analysis of a previously reported crystal structure of human HORMAD1 (8J69). The N-terminus (brown) of HORMAD1 is far away from the closure motif (yellow)-binding region of the HORMA domain (cyan). h Co-IP analyses of C-terminal SFB-tagged (SFBC) HORMAD1 carrying mutations at key residues critical for closure motif binding with HA-tagged SYCP2 fragment (amino acids 390-490) in 293 T cells. i Analysis of two potential BRCA1-binding interfaces adjacent to the N-terminus of HORMAD1 using a previously reported crystal structure of human HORMAD1 (8J69). Solvent-exposed residues on interface I (magenta) and II (green) mutated in this study are highlighted. j Co-IP analyses of C-terminal SFB-tagged (SFBC) HORMAD1 carrying mutations at solvent-exposed residues on interface I and II with BRCA1 in 293 T cells. k Co-IP analyses of C-terminal SFB-tagged (SFBC) HORMAD1 carrying mutations at solvent-exposed residues on interface I with HA-tagged SYCP2 fragment (amino acids 390-490) in 293 T cells. Source data are provided as a Source Data file.
To explore how HORMAD1 co-immunoprecipitates with BRCA1, we first mapped the region of HORMAD1 that co-immunoprecipitates with BRCA1. Removing the HORMA domain, but not the N-terminal or C-terminal regions, of HORMAD1 completely abolished the co-immunoprecipitation (Fig. 7d, e), suggesting that HORMAD1 uses its HORMA domain to co-immunoprecipitates with BRCA1. When interacting with their binding partners, HORMA domains typically adopt a “closed” conformation by wrapping a short closure motif on the partner proteins24. A previously reported crystal structure of human HORMAD1 has revealed that its HORMA domain adopts a similar “closed” conformation but wraps a short closure motif on its own C-terminus25. Interestingly, mutations of key residues on HORMAD1 critical for closure motif binding, Y188A and F193A, had no impact on the co-immunoprecipitation with BRCA1 (Fig. 7f), suggesting that HORMAD1’s HORMA domain does not co-immunoprecipitates with BRCA1 using the closure motif-binding mode. We performed further analysis of the previously reported crystal structure of human HORMAD1 and revealed that the N-terminus of HORMAD1 was far away from the closure motif-binding region on the HORMA domain (Fig. 7g)25. Since N-terminal tagging of HORMAD1 disrupts the co-immunoprecipitation with BRCA1, HORMAD1’s HORMA domain likely co-immunoprecipitates with BRCA1 using an interface adjacent to the N-terminus of HORMAD1.
To further interrogate this idea, we examined whether N-terminal tagging of HORMAD1 disrupts the interaction between HORMAD1 and other partner proteins bound at the closure motif-binding region. SYCP2 is a known binding partner of HORMAD123. Although the molecular mechanism underlying the interaction between HORMAD1 and SYCP2 is unclear, AlphaFold predicted that HORMAD1’s HORMA domain adopts a closure motif-binding mode to interact with amino acids 400-406 of SYCP2, which is within the 100 amino acids region of SYCP2 (390-490) that has been shown to interact with HORMAD2 in a yeast two-hybrid assay (Fig. S7)26. Therefore, we tested the interaction between HORMAD1 and a fragment of SYCP2 (amino acids 390-490) containing the putative closure motif. Consistent with the prediction, mutations of key residues on HORMAD1 critical for closure motif binding, Y188A and F193A, disrupted the co-immunoprecipitation with SYCP2390-490 (Fig. 7h). On the contrary, both N-terminal and C-terminal SFB-tagged HORMAD1 strongly co-immunoprecipitated SYCP2390-490 (Fig. 7h). Although it is unclear if HORMAD1 binds the exact sequence of SYCP2 as predicted by AlphaFold, these observations suggest that N-terminal tagging of HORMAD1 does not disrupt SYCP2 binding. Taken together, N-terminal tagging and closure motif-binding region mutations had different impacts on the co-immunoprecipitation of HORMAD1 with BRCA1 and SYCP2, respectively, further confirming that HORMAD1’s HORMA domain does not use the closure motif-binding mode to co-immunoprecipitate BRCA1.
To investigate how HORMA1’s HORMA domain co-immunoprecipitates with BRCA1, we analyzed the previously reported crystal structure of human HORMAD1 and identified two interfaces adjacent to the N-terminus of HORMAD1 that potentially contribute to the co-immunoprecipitation with BRCA1 (Fig. 7i)25. We generated mutations on solvent-exposed residues on both interfaces of HORMAD1 and examined their impacts on the co-immunoprecipitation with BRCA1. Interestingly, mutation of two residues on interface I, R56E and L83A, significantly disrupted the co-immunoprecipitation with BRCA1 (Fig. 7j). On the contrary, mutations of three residues on interface II, K148E, R155E, and Q162K, did not affect the co-immunoprecipitation with BRCA1 (Fig. 7j). Importantly, R56E and L83A did not impact the co-immunoprecipitation with SYCP2 (Fig. 7k), suggesting that these two mutations specifically disrupted the co-immunoprecipitation with BRCA1 but not those bound by the closure motif-binding region. HORMAD1 likely uses interface I to co-immunoprecipitate BRCA1, and N-terminal tagging of HORMAD1 blocks the accessibility of BRCA1 to this interface.
We continued to explore the regions on BRCA1 that are critical for the co-immunoprecipitation with HORMAD1. More than half of BRCA1’s total amino acids are encoded by exon 11 but lack defined functional domains (Fig. S8a). For this reason, the BRCA1ΔExon11 protein is frequently used in cellular studies as a substitute for full-length BRCA1, retaining all functional domains in a shorter form. Unexpectedly, both the exon 11 region of BRCA1 and the BRCA1ΔExon11 protein coimmunoprecipitated with HORMAD1 (Fig. S8b). This observation suggests that the HORMAD1-BRCA1 coimmunoprecipitation could involve multiple regions of BRCA1, which remain unidentified. Further investigation is required to reveal the molecular details underlying the co-immunoprecipitation between HORMAD1 and BRCA1, particularly if they bind directly, and how N-terminal tagging of HORMAD1 disrupts the co-immunoprecipitation.
Discussion
Failure to remove HORMAD1 and HORMAD2 from synapsed chromosome axes triggers oocyte elimination by activating chromosome asynapsis checkpoint
In the meiotic prophase, HORMAD1 and HORMAD2 localize specifically at unsynapsed chromosome axes. Upon completion of chromosome synapsis, they are removed from synapsed chromosome axes by TRIP13 (Fig. 8)19, but the biological significance of this process is unclear. In this study, we show that HORMAD1 and HORMAD2, when retained on synapsed chromosome axes in Trip13−/− oocytes, recruit BRCA1, activate chromosome asynapsis checkpoint, and trigger oocyte elimination (Fig. 8). Disrupting BRCA1-dependent chromosome asynapsis checkpoint activation efficiently rescues the survival of Trip13−/− oocytes. Therefore, TRIP13-dependent removal of HORMAD1 and HORMAD2 from synapsed chromosome axes is essential for primordial follicle pool establishment and female fertility by preventing aberrant activation of chromosome asynapsis checkpoint and unintended elimination of normal oocytes. Disruption of this critical step in the meiotic prophase could diminish the ovarian reserve and cause premature ovarian insufficiency in humans.
Top, in the meiotic prophase of WT oocytes, HORMAD1 and HORMAD2 localize specifically at unsynapsed chromosome axes and recruit BRCA1, which promotes meiotic recombination. Upon completion of chromosome synapsis, TRIP13 engages the N-terminus of HORMAD1 and HORMAD2 and removes them from synapsed chromosome axes. BRCA1 is removed from synapsed chromosome axes, too. Middle, in the meiotic prophase of Trip13−/− oocytes, HORMAD1 and HORMAD2 are retained on synapsed chromosome axes, which recruit BRCA1, activate chromosome asynapsis checkpoint aberrantly, and trigger oocyte elimination. Bottom, in the meiotic prophase of Hormad1&2SFB/SFB oocytes, N-terminal tagging of HORMAD1 and HORMAD2 disrupts the recruitment of BRCA1 to unsynapsed chromosome axes, but the residual BRCA1 is sufficient for meiotic recombination. N-terminal tagging of HORMAD1 and HORMAD2 also blocks TRIP13’s engagement and causes their retention at synapsed chromosome axes. However, Hormad1&2SFB/SFB oocytes are not eliminated because BRCA1 is not retained on synapsed chromosome axes, and the chromosome asynapsis checkpoint is not activated.
Our study suggests that the elimination of Trip13−/− oocytes is a direct consequence of aberrant retention of HORMAD1 and HORMAD2 on synapsed chromosome axes, which is different from the conclusions drawn by previous studies. Based on the DSB repair defects and the γH2AX signals before oocyte elimination, previous studies suggest that Trip13Gt/Gt oocytes are eliminated by the DNA damage checkpoint6. This idea is further supported by the observation that Chk2−/−, in which a canonical member of the DNA damage checkpoint is removed, rescues the survival of Trip13Gt/Gt oocytes5. However, using several markers for DSBs, we show that DSB repair is delayed but completed before Trip13−/− oocytes are eliminated. On the contrary, the γH2AX signals persist after the completion of DSB repair in Trip13−/− oocytes and become diminished only when the recruitment of BRCA1 to synapsed chromosome axes is disrupted. We speculate that ATR, which is also recruited to synapsed chromosome axes in a BRCA1-dependent manner, is responsible for the γH2AX signals in Trip13−/− oocytes in the absence of DSBs. ATM is primarily responsible for CHK2 phosphorylation and activation in somatic cells27, but ATR can also phosphorylate CHK228. Therefore, besides its canonical role in DNA damage checkpoint in oocytes, CHK2 may also function downstream ATR in chromosome asynapsis checkpoint to promote the elimination of Trip13−/− oocytes in the absence of DSBs. Taken together, Trip13−/− oocytes are not eliminated by DNA damage checkpoint but by chromosome asynapsis checkpoint aberrantly activated by HORMAD1 and HORMAD2 on synapsed chromosome axes.
It should be noted that disrupting the BRCA1-dependent chromosome asynapsis checkpoint by N-terminal tagging of HORMAD1&2, removing BRCA1 in oocytes, or removing HORMAD2 significantly, but not fully, rescues the ovarian reserve of Trip13−/− mice. This finding is consistent with our previous study, which has demonstrated that disrupting the BRCA1-dependent chromosome asynapsis checkpoint by removing BRCA1 in meiotic cells significantly, but not fully, rescues the ovarian reserve in Dmc1−/− and Spo11−/− mice9. These observations reinforce the conclusion that BRCA1 is a key component of the chromosome asynapsis checkpoint. However, they also indicate the presence of other, as-yet-unidentified regulators of this checkpoint.
HORMAD1 and HORMAD2 do not disrupt the completion of DSB repair on synapsed chromosome axes
It has been proposed that HORMAD1 and HORMAD2 actively suppress inter-sister chromatid recombination on unsynapsed chromosome axes6,7. Previous studies have also shown that Hormad2−/− rescues the survival of Trip13Gt/Gt oocytes by promoting DSB repair on synapsed chromosome axes6, raising a possibility that HORMAD1 and HORMAD2 disrupt the completion of DSB repair on synapsed chromosome axes. However, DSB repair is completed in Trip13−/− oocytes while HORMAD1 and HORMAD2 persist on synapsed chromosome axes. Instead of using RAD51 alone as a marker for DSBs, whose suitability has been challenged20, we analyze several markers for DSBs and show that the DSB repair kinetics are similar between Trip13−/− and Trip13−/− Hormad2−/− oocytes, which demonstrates that Hormad2−/− rescues the survival of Trip13−/− oocytes without promoting DSB repair. On the contrary, we show that Hormad2−/− disrupts chromosome asynapsis checkpoint activation by compromising the recruitment of BRCA1 to synapsed chromosome axes.
Like HORMAD1 and HORMAD2, BRCA1 persists on synapsed chromosome axes, but DSB repair is completed in Trip13−/− oocytes, suggesting that BRCA1 does not disrupt the completion of DSB repair either. No difference in DSB repair kinetics is observed between Trip13−/− and Trip13−/− Brca1f/- Stra8-GFPCre oocytes either, confirming that Brca1−/− rescues the survival of Trip13−/− oocytes without promoting DSB repair too, which is consistent with our previous observation that Brca1−/− does not promote DSB repair in Dmc1−/− oocytes9. Therefore, disrupting BRCA1-dependent chromosome asynapsis checkpoint activation alone rescues the survival of Trip13−/− oocytes without promoting DSB repair. Taken together, HORMAD1, HORMAD2, and BRCA1 do not disrupt the completion of DSB repair when retained on synapsed chromosome axes.
HORMAD1 and HORMAD2 mediate DSB-independent recruitment of BRCA1 to chromosome axes
In Trip13−/− oocytes, BRCA1’s localization on synapsed chromosome axes is not restricted to sites of unrepaired DSBs, and BRCA1 persists on synapsed chromosome axes when DSB repair is completed. Therefore, despite localizing to DSBs in somatic cells, BRCA1’s localization on synapsed chromosome axes in Trip13−/− oocytes is DSB-independent, which is consistent with the observations that BRCA1 localizes to unsynapsed chromosome axes in a DSB-independent manner in normal or recombination-defective oocytes9. In addition, HORMAD1 and HORMAD2 not only regulate BRCA1’s localization at unsynapsed chromosome axes in normal or recombination-defective oocytes but also tether BRCA1 to synapsed chromosome axes where they are retained in Trip13−/− oocytes. BRCA1 is likely recruited to both unsynapsed and synapsed chromosome axes similarly by the coordinated actions of HORMAD1 and HORMAD2.
Although both HORMAD1 and HORMAD2 regulate BRCA1’s recruitment to chromosome axes in oocytes, their loss has different impacts on BRCA1’s localization. We have shown that BRCA1 is almost completely absent in Hormad1−/− oocytes9, and we show in this study that BRCA1 can be weakly observed on unsynapsed chromosome axes in Hormad2−/− oocytes. Since Hormad1−/− but not Hormad2−/− oocytes have DSB repair and chromosome synapsis defects, it is unfair to compare if they rescue the survival of Trip13−/− oocytes with similar efficiency. However, consistent with their differential impacts on BRCA1’s localization on unsynapsed chromosome axes, Hormad1−/− rescues the survival of recombination-defective Dmc1−/− oocytes much more efficiently than Hormad2−/− 6,14.
It might not be surprising that loss of HORMAD1 generates more significant impacts on DSB-independent recruitment of BRCA1 to unsynapsed chromosome axes than loss of HORMAD2 because HORMAD2’s localization at unsynapsed chromosome axes requires HORMAD1 but not vice versa12. However, we reveal an intrinsic difference between HORMAD1 and HORMAD2 in their abilities to co-immunoprecipitate BRCA1, which might be more critical in determining the different impacts on BRCA1’s localization when they are lost. HORMAD1 co-immunoprecipitates with BRCA1 readily, but HORMAD2 co-immunoprecipitates with BRCA1 weakly. Structure analyses are needed to clarify the mechanism underlying this difference. Besides physical interaction, it is also unclear if HORMAD2 regulates BRCA1’s localization by other unknown mechanisms. Despite these differences, HORMAD1 and HORMAD2 likely cooperate to recruit BRCA1 to chromosome axes.
HORMAD1 co-immunoprecipitates with BRCA1 through a unique interface on its HORMA domain
Consistent with previous studies, N-terminal tagging of HORMAD1 and HORMAD2 blocks TRIP13’s remodeling and causes their retention at synapsed chromosome axes16. Unlike Trip13−/− oocytes, Hormad1&2SFB/SFB oocytes are not eliminated because N-terminal tagging of HORMAD1 and HORMAD2 also disrupts the recruitment of BRCA1 to both synapsed and unsynapsed chromosome axes (Fig. 8). Mechanistically, N-terminal tagging of HORMAD1 disrupts the co-immunoprecipitation with BRCA1. Since HORMAD2 co-immunoprecipitates with BRCA1 weakly, N-terminal tagging of HORMAD2 has a much less pronounced effect on the co-immunoprecipitation with BRCA1. Therefore, the disrupted co-immunoprecipitation between HORMAD1 and BRCA1 is the primary reason for the defective recruitment of BRCA1 to chromosome axes in Hormad1&2SFB/SFB oocytes and spermatocytes. Consistent with the normal localization of HORMAD1 on chromosome axes in Hormad1&2SFB/SFB oocytes and spermatocytes, N-terminal tagging of HORMAD1 does not interfere with the co-immunoprecipitation with SYCP2, a member of axial/lateral elements of the synaptonemal complex. Therefore, N-terminal tagging of HORMAD1 specifically disrupts the co-immunoprecipitation with BRCA1 and compromises the recruitment of BRCA1 to chromosome axes.
HORMAD1 co-immunoprecipitates with BRCA1 through its HORMA domain, but the canonical closure motif-binding region that binds SYCP2 is not responsible for the co-immunoprecipitation. Instead, we identify a unique interface on HORMAD1’s HORMA domain that mediates the co-immunoprecipitation with BRCA1. Since the N-terminal deletion of HORMAD1 does not affect the co-immunoprecipitation with BRCA1, N-terminal tagging of HORMAD1 likely blocks the accessibility of BRCA1 to this unique surface due to steric hindrance. The unique interface for BRCA1 co-immunoprecipitation and the closure motif-binding region for SYCP2 binding are on different sides of the HORMA domain, with the former being close to the N-terminus of HORMAD1, which is the reason why N-terminal tagging of HORMAD1 disrupts the co-immunoprecipitation with BRCA1 but not with SYCP2. The presence of two independent interfaces on the HORMA domain perfectly explains how HORMAD1 recruits BRCA1 while remaining tethered to chromosome axes by binding chromosome axes proteins such as SYCP2, providing mechanistic insights into how BRCA1 functions on chromosome axes in the meiotic prophase and how BRCA1-dependent chromosome asynapsis checkpoint is aberrantly activated in Trip13−/− oocytes.
Although we have identified two residues on the unique interface on HORMAD1’s HORMA domain that are critical for the co-immunoprecipitation with BRCA1, the molecular details of the interaction still require further investigation. In particular, it is important to determine if HORMAD1 binds BRCA1 directly. Besides BRCA1, HORMAD1 may exploit the newly identified interface on its HORMA domain to interact with other proteins in the meiotic prophase. Revealing how HORMAD1 co-immunoprecipitates with BRCA1 will shed light on the shared mechanism through which HORMAD1 binds different partner proteins using the newly identified interface. Besides being expressed in the meiotic prophase, HORMAD1 is aberrantly expressed in many cancers. We have shown before that HORMAD1 binds and compromises the function of the MCM8-9 complex in cancer cells29. The identification of a new binding mode of HORMAD1’s HORMA domain also provides mechanistic insight into the functions of HORMAD1 in cancer cells, which is critical for targeted therapy.
Methods
Mice
Hormad1SFB/+ and Hormad2SFB/+ mice were generated at GemPharmatech by inserting the DNA coding sequence for the S-FLAG-streptavidin binding protein (SFB) tag after the first ATG of the coding sequence of Hormad1 and Hormad2 gene, respectively. The guide RNAs (gRNAs) map to the first coding exon of the Hormad1 gene (TGTCCTCTGCAACTGCATAG) and Hormad2 gene (ATCCTGATTCATTCTCACGA), respectively. The sgRNAs were generated by in vitro transcription using T7 RNA polymerase. The donor vector containing homology arms flanking the DNA coding sequence for the SFB tag was also generated. A mixture of Cas9 protein, sgRNA, and donor vector was injected into zygotes of C57BL/6 J background, and the survived zygotes were transferred into the oviducts of 0.5 days post coitum (dpc) pseudopregnant surrogate mothers the same day. PCR and Sanger sequencing were used to identify the targeted mice of the F0 generation, which were crossed to C57BL/6 J mice to obtain targeted mice of the F1 generation.
Trip13f/f mice (Strain No. T015909) were obtained from GemPharmatech. Trip13+/- mice were generated by crossing Trip13f/+ mice with Ddx4-Cre mice (Strain No. 006954) obtained from The Jackson Laboratory. Brca1f/f mice (Strain No. 01XB8) were obtained from NCI Mouse Repository. Hormad2−/− mice were obtained from RIKEN BRC (Strain No. CDB0576K). Stra8-GFPCre mice were kind gifts of Ming-Han Tong (Chinese Academy of Sciences)30.
All experimental procedures were approved by the Zhejiang University Animal Care and Use Committee (File nos. ZJU20190144 and ZJU20220228). All mice were housed under an appropriately controlled environment (temperature 20-22°C, humidity 50-70%, and 12 h light/12 h dark cycle) with access to water and food ad libitum. Mouse genotyping was performed by PCR using mouse toes or tails. Primer sequences for mouse genotyping are shown in Table S1.
Preparation of meiotic spreads
For spreads of oocytes at the meiotic prophase, ovaries were isolated from fetal or newborn female mice. After washing with ice-cold PBS three times, ovaries were transferred into hypotonic extraction buffer (30 mM Tris-HCl pH 8.0, 50 mM sucrose, and 17 mM sodium citrate) and incubated at room temperature (RT) for 25 min. The ovaries were then transferred into 100 mM sucrose buffer droplets, and oocytes were released by mincing the ovaries. After removing tissue pieces, the cell suspension was mixed with an equal volume of fixation buffer (1% PFA, 0.15% Triton X-100, and 10 mM borate buffer pH 9.2). The mixture was smeared onto coverslips and air-dried.
For spreads of spermatocytes at the meiotic prophase, testes were dissected, and tunica albuginea were removed. Seminiferous tubules were digested in collagenase IV solution (1 mg/ml) at 37°C for 25 min. Cell pellets were obtained by centrifugation and then resuspended in hypotonic extraction buffer at RT for 10 min. After incubation, cell pellets were obtained by centrifugation and resuspended in 100 mM sucrose buffer at RT for 5 min. Finally, the cell suspension was mixed with an equal volume of fixation buffer. The mixture was smeared onto coverslips and air-dried.
Immunofluorescent (IF) staining and imaging
For IF staining of meiotic spreads, coverslips were treated with 0.5% TritonX-100 at RT for 10 min. After washing with PBS three times, coverslips were incubated with primary antibodies at RT for 90 min. Coverslips were washed with PBS and incubated with secondary antibodies at RT for 1 h. After washing with PBS three times, coverslips were incubated with Hoechst 33342 solution for 5 min and then mounted with an anti-fade reagent. Images were captured using a fluorescence microscope with a CCD camera (ECLIPSE Ti2-E, Nikon). The fluorescence intensity was quantified by ImageJ.
For IF staining in frozen sections of PD4 ovaries, samples were fixed overnight in 4% PFA solution at 4°C, then dehydrated in 30% sucrose solution, embedded in OCT, and stored at −80°C before sectioning at 5 μm thickness. IF staining of sections was performed as described above. For oocyte number quantification of PD4 females, MVH-positive oocytes were counted in every fifth section. Counts were multiplied by five to estimate the total number of oocytes per ovary. Only one ovary per animal was used for oocyte quantification. Antibodies used in this study are listed in Table S2.
Histological analysis
Ovaries, testes, and epididymides were dissected and fixed in 4% PFA at 4°C overnight. Fixed tissues were dehydrated using increasing concentrations of ethanol and then embedded in paraffin for serial sectioning at 5 μm thickness. Tissue sections were stained with hematoxylin and eosin (H&E). For follicle number quantification of PD21 females, counts were made for every fifth section. Only one ovary per animal was used for follicle quantification. Histological images were taken by the Grundium Ocus microscope slide scanners (OCUS01, Grundium).
RNA sequencing and data analysis
Testes were collected and kept in liquid nitrogen before processing. Three pairs of testes from each mouse strain were used. Testes were homogenized in TRIzol reagent (Invitrogen), and total RNA was extracted. Sequencing libraries were generated using NEBNext Ultra RNA Library Prep Kit for Illumina (New England Biolabs) following the manufacturer’s protocols. RNA sequencing was performed using NovaSeq 6000 (Illumina). Raw data were generated from the Illumina platform. Sequence artifacts were excluded before downstream analysis, including low-quality reads, unrecognizable nucleotides, and adapter contamination. The filtered clean reads were mapped to Mus musculus genome build mm10 using STAR (version 2.6.0) with default parameters31. Read counts were extracted and each chromosome’s read counts were calculated. The RNA sequencing data reported in this study has been deposited in the Gene Expression Omnibus website with accession code GSE274491.
Tandem affinity purification and mass spectrometry analysis
For each tandem affinity purification, testes were dissected from 5 Hormad1SFB/+ mice or Hormad2SFB/+ mice, respectively, at PD28. After removing tunica albuginea, seminiferous tubules were lysed in NETN300 buffer (50 mM Tris-HCl pH8.0, 300 mM NaCl, 0.5 mM EDTA, 0.5% Nonidet P-40) on ice for 10 min. After centrifugation, the supernatant was collected and diluted with an equal volume of ice-cold H2O. The mixture was incubated with streptavidin-conjugated beads (NUPTEC) at 4°C for 2 h. The beads were collected by centrifugation and were washed with NETN100 buffer (50 mM Tris-HCl pH 8.0, 100 mM NaCl, 0.5 mM EDTA, 0.5% Nonidet P-40) three times. The bound proteins were eluted with saturated biotin (Sigma-Aldrich) in NETN100 buffer at 4°C for 30 min. After centrifugation, the supernatant was collected and incubated with S beads (Millipore) at 4°C for 2 h. The beads were collected by centrifugation and were washed with NETN100 buffer three times. The bound proteins were eluted by SDS loading buffer at 95°C for 10 min and were briefly separated by SDS-PAGE. The entire protein bands (less than 1 cm) were excised and subjected to in-gel trypsin digestion overnight. The peptides were extracted with acetonitrile and vacuum-dried.
Mass spectrometry (MS) analysis of samples was performed at the core facility, Fudan University. MS analysis was performed once (n = 1) with no technical or biological replicates for peptides obtained from Hormad1SFB/+ mice and Hormad2SFB/+ mice, respectively. No control was included in MS analysis. Briefly, samples were loaded onto Proxeon EASY-nLC 1200 system (Thermo Fisher Scientific) packed with Acclaim 120 C18 column (Thermo Fisher Scientific). The mobile phases consisted of solvent A (0.1% formic acid) and solvent B (0.1% formic acid in 80% ACN). The peptides were eluted using the following gradients: 5–8% B in 3 min, 8–44% B in 45 min, 44–100% B in 2 min, and 100% B for 10 min at a flow rate of 200 nL/min. The eluates were then subjected to Orbitrap Exploris 480 mass spectrometer (Thermo Fisher Scientific) for data acquisition. Data acquisition mode was set to obtain one MS scan followed by HCD-MS/MS acquisitions with a cycle time of 2 seconds. For the MS scans, the mass range was set as 300–1800 m/z at a resolution of 70,000, the automatic gain control (AGC) target was set as standard, maximum ion injection time was 50 ms, and dynamic exclusion time was 30 s. For the MS/MS scans, the resolution was set as 17,500, the precursor isolation window was 1.8 m/z, maximum ion injection time was 100 ms, and the dynamic exclusion was 30 s.
The MS/MS spectra raw data were analyzed by matching a UniProtKB Mus musculus protein databases (released on Jan. 28, 2021) with the acquired fragmentation patterns using Proteome Discoverer (v1.4 & v2.0, Thermo Fisher Scientific). 10 ppm mass tolerance for precursor ions and 0.05 Da for fragments. Trypsin was defined as cleavage enzyme; Carboxyamidomethyl for cysteine and oxidation for methionine residues were set as static modifications and variable modifications, respectively. A decoy database containing the reversed sequences of all proteins was appended to the target database to validate the probabilities of identified peptides with a filtration of false discovery rate (FDR) < 1% according to the target-bait method. The complete lists of proteins identified by mass spectrometry are shown in Supplementary Data 1, 2.
Cell culture and transfection
293 T cells were obtained from the American Type Culture Collection (CRL-3216) and cultured in DMEM medium (Thermo Fisher) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin and streptomycin at 37°C with 5%CO2. Cell transfection was performed using Lipofectamine 3000 (Invitrogen) according to the manufacturer’s instructions.
Plasmid construction
The cDNA for HORMAD1, HORMAD2, SYCP2 (390-490), and BRCA1 was synthesized and cloned into either pCAGGS-SFB-IRSE2-puro vector or pCMA-HA vector. Overlapping PCR was performed to generate mutants in indicated genes according to standard procedures. All plasmid constructions were confirmed by Sanger sequencing.
Co-immunoprecipitation and Western blotting
Cells were harvested and lysed in NETN300 buffer on ice for 10 min. After centrifugation, the supernatant was collected and diluted with an equal volume of ice-cold H2O. The mixture was incubated with streptavidin-conjugated beads (NUPTEC) at 4°C for 2 h. The beads were collected by centrifugation and were washed with NETN300 buffer three times. The bound proteins were eluted by SDS loading buffer at 95°C for 10 min, separated by SDS-PAGE, and transferred to PVDF membranes. The membranes were blocked with 5% non-fat milk at RT for 1 h. After washing with TBST buffer three times, the membranes were incubated with primary antibodies at 4°C overnight. The membranes were washed with TBST buffer three times and then incubated with HRP-linked secondary antibodies at RT for 1 h. After washing with TBST buffer three times, the membranes were analyzed using an enhanced chemiluminescence system. The antibodies used in this study are listed in Table S2. Unprocessed Western blotting images are provided in the Source Data file.
Structural analysis
Analysis of a previously reported crystal structure of human HORMAD1 (PDB accession code 8J69) was performed using PyMOL. Structural prediction of the HORMAD1-SYCP2 complex was performed using AlphaFold332. Predicted aligned error (PAE) plots were generated using PAE Viewer33.
Statistics and reproducibility
Each experiment was repeated independently at least 3 times with similar results. At least 3 independent samples were included in each experiment. Statistical analysis was performed by GraphPad Prism 6, and the results were presented as means ± SD. To calculate statistical significance between different groups, two-tailed unpaired Student’s t-tests were performed.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNA sequencing data reported in this study has been deposited in the Gene Expression Omnibus website with accession code GSE274491. The raw mass spectrometry data are not available. We did not request them at the time of analysis, and they were removed from the Fudan University core facility after 2 years as part of routine data cleanup. The processed data are available as Supplementary Data. Source data are provided with this paper.
References
Kowalczykowski, S. C. An Overview of the Molecular Mechanisms of Recombinational DNA Repair. Cold Spring Harb. Perspect. Biol. 7, a016410 (2015).
Zickler, D. & Kleckner, N. Recombination, Pairing, and Synapsis of Homologs during Meiosis. Cold Spring Harb. Perspect. Biol. 7, a016626 (2015).
Hunter, N. Meiotic Recombination: The Essence of Heredity. Cold Spring Harb. Perspect. Biol. 7, a016618 (2015).
Di Giacomo, M. et al. Distinct DNA-damage-dependent and -independent responses drive the loss of oocytes in recombination-defective mouse mutants. Proc. Natl Acad. Sci. USA 102, 737–742 (2005).
Bolcun-Filas, E., Rinaldi, V. D., White, M. E. & Schimenti, J. C. Reversal of female infertility by Chk2 ablation reveals the oocyte DNA damage checkpoint pathway. Science 343, 533–536 (2014).
Rinaldi, V. D., Bolcun-Filas, E., Kogo, H., Kurahashi, H. & Schimenti, J. C. The DNA Damage Checkpoint Eliminates Mouse Oocytes with Chromosome Synapsis Failure. Mol. Cell 67, 1026–1036.e1022 (2017).
Qiao, H. et al. Impeding DNA Break Repair Enables Oocyte Quality Control. Mol. Cell 72, 211–221.e213 (2018).
ElInati, E. et al. The BCL-2 pathway preserves mammalian genome integrity by eliminating recombination-defective oocytes. Nat. Commun. 11, 2598 (2020).
Bai, L. et al. BRCA1 safeguards genome integrity by activating chromosome asynapsis checkpoint to eliminate recombination-defective oocytes. Proc. Natl Acad. Sci. USA 121, e2401386121 (2024).
Daniel, K. et al. Meiotic homologue alignment and its quality surveillance are controlled by mouse HORMAD1. Nat. Cell Biol. 13, 599–610 (2011).
Kogo, H. et al. HORMAD1-dependent checkpoint/surveillance mechanism eliminates asynaptic oocytes. Genes Cells 17, 439–454 (2012).
Wojtasz, L. et al. Meiotic DNA double-strand breaks and chromosome asynapsis in mice are monitored by distinct HORMAD2-independent and -dependent mechanisms. Genes Dev. 26, 958–973 (2012).
Kogo, H. et al. HORMAD2 is essential for synapsis surveillance during meiotic prophase via the recruitment of ATR activity. Genes Cells 17, 897–912 (2012).
Shin, Y. H., McGuire, M. M. & Rajkovic, A. Mouse HORMAD1 is a meiosis i checkpoint protein that modulates DNA double- strand break repair during female meiosis. Biol. Reprod. 89, 29 (2013).
Shin, Y. H. et al. Hormad1 mutation disrupts synaptonemal complex formation, recombination, and chromosome segregation in mammalian meiosis. PLoS Genet 6, e1001190 (2010).
Ye, Q. et al. The AAA+ ATPase TRIP13 remodels HORMA domains through N-terminal engagement and unfolding. EMBO J. 36, 2419–2434 (2017).
Li, X. C. & Schimenti, J. C. Mouse pachytene checkpoint 2 (trip13) is required for completing meiotic recombination but not synapsis. PLoS Genet. 3, e130 (2007).
Wojtasz, L. et al. Mouse HORMAD1 and HORMAD2, two conserved meiotic chromosomal proteins, are depleted from synapsed chromosome axes with the help of TRIP13 AAA-ATPase. PLoS Genet 5, e1000702 (2009).
Roig, I. et al. Mouse TRIP13/PCH2 is required for recombination and normal higher-order chromosome structure during meiosis. PLoS Genet. 6, e1001062 (2010).
Ravindranathan, R., Raveendran, K., Papanikos, F., San-Segundo, P. A. & Toth, A. Chromosomal synapsis defects can trigger oocyte apoptosis without elevating numbers of persistent DNA breaks above wild-type levels. Nucleic Acids Res. 50, 5617–5634 (2022).
Motoyama, N. & Naka, K. DNA damage tumor suppressor genes and genomic instability. Curr. Opin. Genet. Dev. 14, 11–16 (2004).
Turner, J. M. Meiotic silencing in mammals. Annu. Rev. Genet. 49, 395–412 (2015).
Fujiwara, Y. et al. Meiotic cohesins mediate initial loading of HORMAD1 to the chromosomes and coordinate SC formation during meiotic prophase. PLoS Genet. 16, e1009048 (2020).
Rosenberg, S. C. & Corbett, K. D. The multifaceted roles of the HORMA domain in cellular signaling. J. Cell Biol. 211, 745–755 (2015).
Wang, H. et al. Structural and biochemical insights into the interaction mechanism underlying HORMAD1 and its partner proteins. Structure 31, 1578–1588.e1573 (2023).
West, A. M. et al. A conserved filamentous assembly underlies the structure of the meiotic chromosome axis. Elife. 8, e40372 (2019).
Matsuoka, S., Huang, M. & Elledge, S. J. Linkage of ATM to cell cycle regulation by the Chk2 protein kinase. Science 282, 1893–1897 (1998).
Pabla, N., Huang, S., Mi, Q. S., Daniel, R. & Dong, Z. ATR-Chk2 signaling in p53 activation and DNA damage response during cisplatin-induced apoptosis. J. Biol. Chem. 283, 6572–6583 (2008).
Liu, K. et al. Aberrantly expressed HORMAD1 disrupts nuclear localization of MCM8-MCM9 complex and compromises DNA mismatch repair in cancer cells. Cell Death Dis. 11, 519 (2020).
Lin, Z. et al. Mettl3-/Mettl14-mediated mRNA N(6)-methyladenosine modulates murine spermatogenesis. Cell Res. 27, 1216–1230 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Elfmann, C. & Stulke, J. PAE viewer: a webserver for the interactive visualization of the predicted aligned error for multimer structure predictions and crosslinks. Nucleic Acids Res. 51, W404–W410 (2023).
Acknowledgements
We thank Ming-Han Tong for sharing mouse strains. We thank Quanfeng Zhu for helping with mouse genotyping. We thank Jiaxue Wu for helping with the mass spectrometry analysis. We thank Chao Bi and Xiaoli Hong from the Core Facilities, Zhejiang University School of Medicine for their technical support. We thank Xiaofen Jin, Yumiao Niu, and Feiting Xie from the Central Laboratory, Women’s Hospital, Zhejiang University School of Medicine for their technical support. This work is funded by National Natural Science Foundation of China (32070829 and 32370900 to L.L. and 32200692 to X.J.).
Author information
Authors and Affiliations
Contributions
Y.L. and L.L. designed the research. X.J., Z.L., J.L., and L.B. performed the research. X.J., Z.L., J.L., J.X., Y.L., and L. L. analyzed the data. X.J., Y.L., and L. L. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Owen Davie, Jayakrishnan Nandakumar, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jiao, X., Liang, Z., Li, J. et al. Aberrant activation of chromosome asynapsis checkpoint triggers oocyte elimination. Nat Commun 16, 2260 (2025). https://doi.org/10.1038/s41467-025-57702-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57702-z