Abstract
Faithful meiotic segregation requires pairwise alignment of the homologous chromosomes and their synaptonemal complex (SC) mediated stabilization. Here, we investigate factors that promote and coordinate these events during C. elegans meiosis. We identify BRA-2 (BMP Receptor Associated family member 2) as an interactor of HIM-17, previously shown to promote double-strand break formation. We found that loss of bra-2 impairs synapsis elongation without affecting homolog recognition, chromosome movement or SC maintenance. Epistasis analyses reveal previously unrecognized activities for HIM-17 in regulating homolog pairing and SC assembly in a partially overlapping manner with BRA-2. We show that removing bra-2 or him-17 restores nuclear clustering, recruitment of PLK-2 at the nuclear periphery, and abrogation of ectopic synapsis in htp-1 mutants, suggesting intact CHK-2-mediated signaling and presence of a barrier that prevents SC polymerization in the absence of homology. Our findings shed light on the regulatory mechanisms ensuring faithful pairing and synapsis.
Similar content being viewed by others
Introduction
The formation of euploid gametes depends on the faithful execution of several disparate tasks during meiosis, all meticulously coordinated to ensure correct chromosome segregation into the daughter cells. Diploid genomes are composed of two homologous chromosomes, the paternal and the maternal copy, which must recognize one another (pairing) and recombine to achieve the formation of physical tethers called crossovers (COs). COs are essential to achieve successful segregation of the homologous chromosomes into the gametes1,2.
Pairwise chromosome alignment depends on microtubule-driven forces transmitted from the cytosol to the nucleus through the SUN-KASH protein bridge3,4,5,6. Once the chromosomes are connected to this protein module spanning the nuclear membranes, active movement along the nuclear envelope is triggered, enabling the chromosomes to travel within the nucleus until the synaptonemal complex (SC) has been established between homologous chromosome pairs. Therefore, chromosome movement is essential for pairing and effective homologous synapsis, as well as to resolve chromosome interlocks3,7,8,9.
In Caenorhabditis elegans, active chromosome movement begins at meiotic entry, defining a specific region of the germ line called “transition zone” (TZ) that loosely corresponds to leptotene/ zygotene stages7,10,11,12. Upon entry into the TZ, cytoplasmic motor proteins promote robust movement of the chromosome ends tethered to the nuclear envelope by a family of zinc finger proteins that associate with chromosome-specific sub-telomeric regions called pairing centers (PCs)13,14,15,16. ZIM-1, -2 or -3 individually bind to the PCs of the autosomes, while HIM-8 is recruited to the PC of the X chromosome. Both the PCs and their cognate PC-binding protein associate with the inner nuclear envelope proteins SUN-1 and MJL-1, a recently identified C. elegans homolog of mammalian MAJIN, essential for the fusion of the chromosome ends to the nuclear envelope17. Homolog pairing is established within these clustered PCs, and once homology is satisfied, the SC forms (synapsis) between homologs, first nucleated at PCs, and then elongated in between each chromosome pair9. However, synapsis can take place irrespective of homology, and therefore, mechanisms must be in place to ensure that SC assembly is strictly achieved between homologous partners.
Previous work has shown that cytoplasmic dynein plays essential roles in promoting the establishment of synapsis strictly between homologous partners, by exerting opposing forces to promiscuous associations between non-homologous chromosomes7. Under impaired dynein function, homologs can still achieve pairing, although with delayed kinetics, but fail in establishing synapsis, since SC subunits accumulate into nuclear polycomplexes and do not elongate. These observations have led to a model whereby dynein activity imposes a barrier to synapsis that is contingent to homology7.
The polarized-clustered, crescent shape configuration of the chromatin as typically observed in TZ, is acquired in part by the recruitment of chromosomes ends to the nuclear periphery18 and in part by the destabilization of the nuclear lamina19.
PCs recruit the Polo-like kinase PLK-2, which leads to SUN-1 phosphorylation and likely other targets, and is crucial for faithful pairing and synapsis15,20. Importantly, PLK-2 and phosphorylated SUN-1 concentrated at the PCs are part of a surveillance system that can extend the TZ stage in response to defects in synapsis and recombination15,20,21 and, therefore these markers can be used as a proxy to monitor meiotic entry and homology search competence.
The SC is a meiosis-specific proteinaceous structure composed of lateral and central elements that is conserved in most species. The SC acts as a zipper-like scaffold that allows physical exchange of DNA molecules between the homologs during recombination, and thus it is essential for CO formation22. In worms, a family of four HORMA-domain containing proteins is recruited to the chromosome axes, where these factors play different roles during SC assembly. HTP-3 serves as a scaffold and promotes axes morphogenesis and also physiological DNA double-strand break (DSB) induction23,24, HIM-3 is required for synapsis25, and HTP-1/2 play regulatory roles by coordinating homologous pairing with SC assembly26. Lack of HTP-3 or HIM-3 abrogates chromosome pairing, chromatin clustering and synapsis altogether, while removal of HTP-1 triggers precocious exit from the homology search state resulting in severely reduced chromosome clustering, defective pairing, and extensive non-homologous synapsis. Removal of htp-1 or sun-1 bypasses the dynein-dependent release of SC assembly resulting in ectopic synapsis7, indicating that HTP-1, as well as SUN-1, are essential to maintain this barrier in an active state. Abrogation of htp-2 functions on its own does not result in aberrations of pairing or synapsis, however its removal in htp-1 mutants exacerbates synapsis defects27, suggesting a partial redundancy.
While being an essential requirement for CO formation in C. elegans, the SC alone is not sufficient to promote COs, whose formation depends on homologous recombination-mediated repair of deliberately induced DSBs by the Topoisomerase-like protein SPO-1128,29. SPO-11 acts in conjunction with numerous accessory factors shown to be essential for achieving optimal DSB levels, and while SPO-11 itself is highly conserved, auxiliary pro-DSB proteins have widely diverged across species30. Importantly, unlike in most species, DSB induction and SC formation occur independently of one another in C. elegans29, allowing the study of either process without confounding effects.
We previously generated functional tagged lines by CRISPR of several accessory factors required for DSB formation31,32, which we exploited via a biochemical approach aimed at identifying proteins interacting with the DSB promoting machinery. In this work we focus on putative interactors identified in pull-downs experiments performed by using HIM-17::3XHA31 as bait, followed by mass spectrometry analysis on HA-immunoprecipitated complexes. We identify the uncharacterized MYND-type zinc finger protein BRA-2 as a putative HIM-17 interactor, and we report its roles during early meiotic progression in this study. We observe that BRA-2 and HIM-17 form complexes in vivo and occasionally colocalize in the germ line, yet their chromatin loading is not interdependent. Loss of BRA-2 severely reduces chiasmata formation and impairs synapsis, by preventing efficient SC elongation upon homology assessment. Strikingly, removal of HIM-17 from BRA-2-depleted animals completely abolishes pairing and synapsis along the autosomes but not on the X chromosome, although X-chromosome synapsis is only achieved in late pachytene nuclei. BRA-2 removal restores a transition zone nuclear morphology and recruitment of PLK-2 to the chromosome PCs in htp-1 mutants, without improving pairing levels, supporting further evidence that BRA-2 has an early regulatory activity at the level of SC elongation after homolog pairing has been successfully achieved. Furthermore, we discovered that removal of him-17 in htp-1 mutants delays SC formation and greatly improves pairing, which is lost in htp-1; him-17 worms depleted for BRA-2, indicating partially overlapping functions between BRA-2 and HIM-17 in regulating chromosome pairing and SC formation. Our data suggest that BRA-2 cooperates with HIM-17 at meiotic entry to promote pairing and allow SC polymerization between homologs and unveil an as yet undescribed role for the chromatin associated protein HIM-17 beyond DSB formation.
Results
BRA-2, but not its paralog BRA-1, is required for robust crossover formation
In a mass spectrometry analysis aimed at identifying interactors of the THAP (THanatos Associated Protein)-domain containing protein HIM-17, we identified BRA−2 (BMP Receptor Associated family member 2) as a putative binding partner (Suppementary Table 1), for which no roles in meiosis were known.
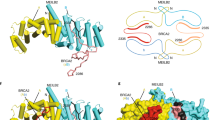
BRA-2 is a small protein (~25kD) harboring a MYND-type zinc finger domain (ZnF) at its C-terminal region, conserved in several proteins across species. BLAST search identified the mammalian putative histone H3.3K36me3 reader ZMYND1133 as the closest homolog to C. elegans BRA-2 (Fig. 1a). In mammals, it has been shown that recognition and binding to trimethylated histone H3.3K36 by ZMNYD11 is important to regulate pre-mRNA processing and elongation, and further, mutations in human ZMYND11 have been linked to a wide spectrum of intellectual disability syndromes34. However, functional analyses in the germ line have never been carried out. It is important to note that while the MYND-ZnF domain is highly conserved between worms and mammals, the protein domains directly involved in the recognition and binding of the chromatin modification (PHD-RING, PWWP- and BROMO- domain) are not present in C. elegans BRA-2, possibly indicating a different function or an alternative mode of action in nematodes.
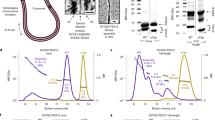
a Schematic representation of C. elegans BRA-1/2 and mammalian ZMYND11 with putative protein domains highlighted. b Exon/Intron organization of bra-1 and bra-2 loci. Red lines indicate deleted regions in the respective mutant alleles. c Quantification of viability levels in the indicated genetic backgrounds. The number of worms scored for each genotype was WT (7), bra-2(ok1171) (12), bra-1(nk1) (11), HA::AID::bra-2; TIR1::mRuby (5), 3XFLAG::bra-1 (5). Bars indicate mean with ±SD. d Left: quantification of DAPI-bodies number in diakinesis nuclei of the indicated genotypes. Nuclei analyzed: WT = 47, bra-2 = 85, bra-1 = 47. Representative examples are shown to the right. Scale bar 2 μm. Bars indicate average with ±SD. e Quantification of OLLAS::COSA-1 foci in the indicated genotypes (left) and representative images of late pachytene nuclei stained for SYP-1 (blue)/OLLAS (COSA-1, red), and counterstained by DAPI (gray). At least three germ lines for each genetic background were used for the quantifications. The number of nuclei analyzed was: OLLAS::cosa-1 (138), bra-2 OLLAS::cosa-1 (153). Bars indicate average with ±SD. Scale bar 5 μm.
To assess the phenotypes triggered by loss of BRA-2, we exploited the bra-2(ok1171) mutant strain, which carries a large deletion that removes the entire bra-2 locus, thus resulting in a null allele (Fig. 1b). Assessment of viability levels revealed high embryonic lethality in bra-2 mutants (Fig. 1c), consistent with aneuploidy arising from chromosome missegregation.
In nuclei at diakinesis, the last stage of meiotic prophase I, chromosomes become highly condensed and appear as six DAPI bodies in WT animals, each representing a pair of homologous chromosomes held together by a chiasma. Defects in recombination and DNA repair can cause an increase/reduction in the DAPI bodies number and/or variation in their morphology, thus diakinesis nuclei can be used as a powerful read-out for the proper completion of upstream meiotic tasks.
Consistent with elevated embryonic lethality, diakinesis nuclei in bra-2(ok1171) mutants displayed a large proportion of achiasmatic univalents (89.4% of nuclei contained >6 DAPI bodies), suggesting defective CO establishment (Fig. 1d). Quantification of CO-designation sites by assessing COSA-1 foci35 revealed a dramatic reduction compared to controls, with an average of 2.4 COSA-1 foci/nucleus in the bra-2 mutants versus 5.98 in the controls (Fig. 1e), and with 40% of nuclei that did not show any COSA-1 foci in the bra-2 mutants. This indicates that BRA-2 is essential for efficient CO formation.
The C. elegans genome encodes a closely related paralog of bra-2 called bra-136. These proteins share 62% identity and like BRA-2, BRA-1 also carries only a MYND-type ZnF domain (Fig. 1a). Given their high similarity, we sought to investigate whether BRA-1 might also be involved in promoting CO formation. We employed the bra-1(nk1) mutant background, which carries a large deletion that removes most of the bra-1 locus36 (Fig. 1b), likely resulting in a null allele. Surprisingly, both viability as well as number of DAPI bodies in the diakinesis nuclei were indistinguishable from WT animals (Fig. 1c, d). This indicates that BRA-1 is not essential for chiasmata formation and that only BRA-2 exerts essential functions in promoting efficient establishment of COs.
BRA-2 is enriched on the autosomes and its loading is independent of synapsis and recombination
To gain further insight into BRA-2 localization and its functions, we engineered the bra-2 endogenous locus by introducing an auxin-induced degradation tag (AID) in tandem with an HA tag after the start codon. This allows detection of BRA-2 as well as its conditional depletion upon exposure to auxin, mediated by TIR1 binding and subsequent targeting of HA::AID::BRA-2 for proteosome-dependent degradation37. HA::AID::bra-2; TIR1::mRuby animals displayed normal levels of embryonic viability (Fig. 1c), indicating that the HA::AID fusion tag did not compromise protein functionality. We assessed BRA-2 depletion efficiency by Western Blot on total protein extracts, which showed that exposure to auxin can rapidly and efficiently deplete >95% of HA::AID::BRA-2 within 3 hours (Supplementary Fig. 1a).
Immunostaining using anti-HA antibodies showed that BRA-2 localizes as a nuclear factor throughout meiotic prophase I (Fig. 2a). The nuclear localization was recapitulated by Western blot on fractionated protein extracts, which indeed showed that HA::AID::BRA-2 is enriched in both soluble and chromatin-bound nuclear fractions (Fig. 2b). Cytological analysis indicated that HA::AID::BRA-2 was not evenly distributed on chromatin, as one chromosome was devoid of signal. Co-staining using anti-HIM-8 antibodies revealed that HA::AID::BRA-2 is prominently excluded from the X chromosome (Fig. 2c), as also corroborated by co-localization with H3K4me2, XND-1 and phosphorylated-RNA Pol IIS2 (Supplementary Fig. 1b), all known to be enriched on the autosomes38,39,40. Furthermore, assessment of HA::AID::BRA-2 localization in the spo-11(ok79) and syp-2(ok307) mutants showed that loading of BRA-2 is independent of DSBs and synapsis respectively (Supplementary Fig. 1c).
a Top: whole mount gonad stained with anti-HA (red) and anti-HIM-8 (green), showing expression of BRA-2 throughout the germ line and its enrichment on the autosomes. Scale bar 20 μm. b Western blot on fractionated protein extracts probed with anti-HA antibodies showing enrichment of BRA-2 in the nuclear-soluble and chromatin-bound fractions. Detection of GAPDH and Histone H3 was used as loading controls of cytosolic and chromatin-bound fraction respectively. kD indicates molecular weight of the marker bands (kiloDalton). c Magnified mid-pachytene nuclei showing more detailed staining of HA::AID::BRA-2 (red)/HIM-8 (green). Arrowheads indicate the X chromosome. Scale bar 5 μm. d Co-immunoprecipitation assay showing physical interaction between HIM-17::GFP and HA::AID::BRA-2. kD indicates molecular weight of the marker bands (kiloDalton). e Early pachytene nuclei stained for GFP (HIM-17, cyan), HA (BRA-2, magenta) and DAPI (green), showing similar diffuse chromatin localization. Scale bar 5 μm. f Early-mid pachytene nuclei immunoassayed for HA (yellow), SYP-1 (magenta) and DAPI (cyan) displaying him-17-independent recruitment of BRA-2 onto the DNA. Scale bar 5 μm. g Early-mid pachytene nuclei immunoassayed for HA (magenta), XND-1 (yellow) and DAPI (green) displaying bra-2-independent recruitment of HIM-17 onto the DNA. Scale bar 5 μm. All shown analyses were performed in biological duplicates.
Although we observed neither chromosome segregation defects in the bra-1(nk1) mutants nor redundancy between bra-1 and bra-2 (Fig. 1c, d), we wished to investigate BRA-1 localization and monitor potential changes in the absence of BRA-2. To this end, we generated a 3XFLAG::bra-1 tagged line (Fig. 1c) and combined it with HA::AID::bra-2 to analyze their localization.
HA::AID::BRA-2 and 3XFLAG::BRA-1 display distinct temporal loading patterns in the germ line, as the former is expressed throughout the gonad and the latter appears only at pachytene exit (Supplementary Fig. 1d). BRA-1 and BRA-2 did not display interdependent loading, as 3XFLAG::BRA-1 was normally loaded in BRA-2-depleted animals and conversely, HA::AID::BRA-2 did not display any abnormalities in the bra-1(nk1) knockout mutants (Supplementary Fig. 1e). Lastly, Western blot analysis on whole cell extracts showed that BRA-1/2 proteins are not destabilized in the reciprocal mutant background (Supplementary Fig. 1f, g), further corroborating that these two factors act in distinct pathways and that BRA-1 has a negligible or no role in the germ line.
Given the putative interaction identified by mass spectrometry, we next sought to investigate whether BRA-2 and HIM-17 interact in vivo and if their loading is affected in the reciprocal mutant background. Immunoprecipitation of HIM-17::GFP41 identified HA::AID::BRA-2 (Fig. 2d), confirming our mass spectrometry data and indicating that they are found in a complex. Cytological analysis showed that BRA-2 and HIM-17 display a very similar localization in the gonad (Fig. 2e), accumulating diffusely on the chromatin without showing obvious enrichment along the chromosome axes/SCs. We noticed that the two proteins also displayed foci of varying intensity which did not co-localize, while the remaining pools occasionally displayed some overlapping staining. Furthermore, we observed that BRA-2 was still loaded in him-17(e2707) mutants (Fig. 2f) and conversely, HIM-17::3XHA appropriately localized in the bra-2(ok1171) mutants (Fig. 2g). Thus, while displaying a similar localization pattern and being found together in protein complexes, BRA-2 and HIM-17 do not undergo interdependent loading.
BRA-2 is largely dispensable for homolog pairing but is essential for normal SC assembly
DAPI-staining analysis of diakinesis nuclei and quantification of CO-designation sites number revealed defective CO formation in bra-2(ok1171) (Fig. 1d, e). This can originate from defects in pairing, synapsis, or recombination11,26,29,42,43; thus to distinguish between these possibilities, we first assayed pairing levels by performing FISH analysis using specific probes recognizing different autosomes, as well as immunostaining with antibodies directed against the PC-binding protein HIM-8 to identify the chromosome X. We divided the gonads into five equal regions from Transition Zone (TZ) to Late Pachytene (LP) and assessed pairing of the indicated chromosomes in each nucleus across these zones (Fig. 3a). Pairing was largely achieved in bra-2 mutants, although it peaked at slightly reduced levels compared to control animals, most prominently for the autosomes scored (Fig. 3a, b). This indicates that although not being essential, BRA-2 is required to attain normal levels of homolog pairing.
a Schematic representation of the C. elegans germline with indicated Prophase I stages and zones division employed for FISH quantification. Charts show paring quantification of the indicated chromosomes in WT and bra-2(ok1171) mutants. Scale bars report S.E.M., and asterisk denotes statistical significance as calculated by Χ2 test (two-sided, CI. = 0.05, p values zone 2-5: Ch. III – 0.000059, 0.009, 0.000374, 0.000026; Ch. V - <0.00001, 0.000018, 0.000108, 0.007386; Ch. X – 0.004118, 0.14855, 0.176575, 0.018383; Ch. IV (zone 1-5) – 0.00001, 0.00003, 0.000012, 0.003951, 0.00072). At least three germ lines for each genetic background were used for the quantifications. The following number of nuclei was analyzed in WT and bra-2(ok1171), respectively: Ch. III - zone 1 (142, 188), zone 2 (169, 167), zone 3 (150, 166), zone 4 (148, 130), zone 5 (123, 102); Ch. IV - zone 1 (145, 164), zone 2 (122, 178), zone 3 (95, 153), zone 4 (84, 117), zone 5 (57, 114); Ch. V - zone 1 (143, 188), zone 2 (169, 167), zone 3 (152, 168), zone 4 (147, 131), zone 5 (123, 102); Ch. X - zone 1 (150, 261), zone 2 (153, 231), zone 3 (129, 221), zone 4 (106, 197), zone 5 (94, 163). b Mid-pachytene nuclei displaying indicated probes signal (Ch. III, red - Ch. V, green – Ch. IV, red - DAPI in blue) and antibodies (SYP-1, magenta - HIM-8, yellow – DAPI, cyan). Scale bar 5 μm. Analysis was performed in biological triplicates. c Quantification of chromosome movement indexes in live animals assessed by tracking SUN-1::GFP in untreated HA::AID::bra-2; TIR1::mRuby worms and after auxin exposure. Bars indicate ±SD with mean. Statistical analysis was performed by two-tailed Mann-Whitney test (ns= non-significant, *p = 0.02, **p = 0.004). The number of nuclei analyzed was 13 and 20 (-auxin and +auxin, respectively). The experiment was per formed in biological triplicates. d Early-pachytene nuclei showing anti-SYP-1 (yellow) and FISH probe labeling Ch. III (red) and (e) anti-SYP-1 (magenta) and FISH probe labeling Ch. V (yellow). Analysis was performed in biological triplicates. Scale bar 5 μm.
To further understand the role of BRA-2 during establishment of pairing, we investigated the movement of chromosome ends by monitoring SUN-1 dynamics in live animals, since this poses an essential requirement for successfully achieving the installation of the SC between homologous chromosomes7,18. We then used auxin-depleted HA::AID::BRA-2; TIR1::mRuby animals carrying a functional sun-1::GFP transgene and tracked the dynamic behavior of GFP aggregates, which mark chromosome ends at the nuclear periphery, in live worms12. This analysis showed that the speed, split/fusion events and coalescence times of SUN-1::GFP aggregates followed normal kinetics in absence of BRA-2 (Fig. 3c). However, we found a mild, although statistically significant increase in the area covered and the total distance traveled by the SUN-1::GFP patches, suggesting a slight increase in chromosome movements. These results indicate that the reduced pairing levels observed in bra-2 mutants are not a consequence of gross defects in chromosome motion during the pairing stages.
Once homology is satisfied, the physical interaction between the coaligned chromosomes is stabilized by the SC in C. elegans42,44,45. At meiotic entry, axis proteins are loaded first (HTP-1, -2 and −3, HIM-3, cohesins), and then central components (SYP-1, -2, -3, -4, -5 and -6, SKR-1, and SKR-2) polymerize between the chromosome axes23,25,44. By early pachytene (EP), the SC is fully built along the entire length of the chromosomes, as defined by complete overlap between axial and central elements, coupled with loss of clustered chromatin configuration. DNA regions decorated by axial components but devoid of SYPs, indicate absence of synapsis.
The bra-2(ok1171) mutants displayed a dramatic impairment of synapsis, as SYP-1 loading between TZ and EP was barely detectable (Fig. 4a and Supplementary Fig. 2, SYP-1 is seen mostly in puncta and small patches). In MP-LP stages (corresponding to zones 5-6 in the gonad), SYP-1 staining appeared more robust and elongated, however full synapsis was only rarely observed, indicating that BRA-2 is essential for normal SC assembly. We note that the chromatin in TZ nuclei displayed the characteristic clustered morphology, and that appropriate HTP-3 localization was seen in bra-2 mutants, suggesting successful meiotic entry and that loss of BRA-2 confers a genuine synapsis defect. We also analyzed synapsis in bra-1 and bra-2; bra-1 double mutants by HTP-3/SYP-1 immunostaining, which showed no dramatic defects in the former and no differences in comparison to bra-2 single mutants in the latter (Fig. 4a and Supplementary Fig. 2). This further corroborates that only BRA-2 exerts meiotic functions. By combining FISH analysis (or HIM-8 detection) with SYP-1 immunostaining, we were able to observe that paired chromosomes X, III and V and were associated with some SYP-1 protein in pachytene nuclei (Fig. 3b, d, e), indicating that the residual SC installation does not occur between non-homologous chromosomes in absence of BRA-2.
a Left: nuclei from the indicated mutants and stage stained for axial (HTP-3, red) and central (SYP-1, green) elements of the SC, counterstained by DAPI (blue). Note extensive regions of HTP-3 devoid of SYP-1 signal, indicating the absence of synapsis. Scale bar 5 μm. Right: schematic representation depicting zoning of the germ line used to quantify synapsis. The chart shows quantification of nuclei with full SC in the indicated zones and genetic backgrounds. Bars show S.E.M., and asterisks denote statistical significance assessed by two-sided Χ2 test (ns= non-significant, **p = 0.004, ****p < 0.0001). At least three germ lines were analyzed and the number of nuclei scored in WT, bra-1 and bra-2 mutants respectively was: zone 1 (229, 224, 306), zone 2 (235, 301, 249), zone 3 (204, 243, 235), zone 4 (199, 227, 230), zone 5 (167, 186, 180), zone 6 (143, 146, 134). b In vivo imaging of GFP::SYP-3 in untreated worms and after exposure to auxin indicates slightly reduced formation of SYP-3 foci coupled with (c) impaired elongation proficiency and starkly reduced SYP-3 polymerization speed. 100 nuclei were analysed for each condition. Statistical significance was calculated by the Mann-Whitney test (two-sided). d Left: representative images of nuclei from HA::AID::bra-2; TIR1::mRuby animals stained for HTP-3 (red)/SYP-1 (green) at the indicated stages (TZ=Transition Zone, EP=Early Pachytene, LP=Late Pachytene) and conditions of exposure to auxin. Scale bar 5 μm. Right: Quantification of nuclei displaying full synapsis across the germ line in the indicated backgrounds and exposure conditions to auxin. 2-3 germ lines were analyzed and the number of nuclei scored in -auxin, 19 h auxin and 16 h recovery conditions respectively, was: zone 1 (254, 113, 269), zone 2 (295, 111, 332), zone 3 (248, 146, 289), zone 4 (270, 172, 231), zone 5 (229, 147, 214), zone 6 (194, 118, 177). Bars show S.E.M. and asterisks denote statistical significance assessed by two-sided Χ2 test (ns= non-significant, ****p < 0.0001). e Schematization of the assay performed to track stage-specific cells before and after BRA-2 depletion. (PMT= Premeiotic Tip, TZ=Tranistion Zone). f Co-staining of SYP-1 (green)/pHTP-3S285 (red) with EdU (gray) in the indicated conditions of exposure to auxin. Arrowheads point to nuclei that are positive for EdU incorporation and show incomplete SC. Scale bar 10 μm. Analysis was performed in biological duplicates.
The SC is assembled gradually at meiotic entry, with the central elements nucleated at the PC sites first, and then rapidly elongated along the whole length of the chromosomes9. To discern whether the largely impaired SC assembly observed in bra-2 mutants was originating from defective recruitment of SC subunits or their polymerization along the axes, we carried out live imaging of GFP::SYP-3 at meiosis onset in live worms9. The initial recruitment of SYP-3 as bright foci, indicating SC nucleation sites, was only slightly reduced (Fig. 4b). Strikingly, while GFP::SYP-3 foci underwent elongation in the presence of BRA-2 (-auxin), we found that in about half of the cells, GFP::SYP-3 foci did not show elongation upon BRA-2 depletion, and the remaining portion displayed a severe reduction in the elongation rate (∼65%; 474 nm/m in the -auxin vs 169 nm/m in the +auxin) (Fig. 4c). While we cannot distinguish whether the GFP::SYP-3 foci that did not show elongation were also localizing at PCs, these data show that BRA-2 is important for stimulating their polymerization along the axes.
Loss of BRA-2 destabilizes SC subunits and hinders their polymerization
We next considered the possibility that impaired SC assembly observed in bra-2 mutants could be resulting from defective cell-cycle progression. To address this, we co-stained WAPL-1 and the Cyclin E CYE-1, two markers expressed in the progenitor zone (which harbors nuclei in premeiotic divisions and pre-meiotic replication)46,47 (Supplementary Fig. 3a). Moreover, we performed an assay to monitor incorporation of 5-ethynyl-2′-deoxyuridine (EdU) to assess ongoing replication (Supplementary Fig. 3b). For all these markers we did not find any difference between WT and bra-2 mutants, indicating that germ cells enter meiosis with unperturbed kinetics and undergo normal mitotic and pre-meiotic replication respectively.
Once we assessed that the mitosis-to-meiosis developmental switch was not impacted by lack of BRA-2, we combined our HA::AID::bra-2; TIR1::mRuby strain with available functional tagged lines of other SC central elements to monitor loading of SYP-2::V5, V5::SYP-448, and GFP::SYP-39.
HA::AID::bra-2; TIR1::mRuby animals exposed to auxin for 24 hours recapitulated the synapsis defects observed in the bra-2(ok1171) null mutants, whereby almost no loading of endogenous SYP-1 was observed until EP (Supplementary Fig. 3c). We found that similar to defective SYP-1 loading, BRA-2-depleted animals also displayed impaired localization of SYP-2::V5 (Supplementary Fig. 3d), GFP::SYP-3 (Supplementary Fig. 3e) and V5::SYP-4 (Supplementary Fig. 3f), as expected by their interdependent localization. Interestingly, Western blot analysis showed that SYP-2::V5 and V5::SYP-4 were reduced in absence of BRA-2 (Supplementary Fig. 3g) and assessment of transcript levels of endogenous syp-1 and syp-4 revealed no detectable abnormalities in expression (Supplementary Fig. 4a). We expanded this analysis by performing RNAseq on gonads dissected from HA::AID::bra-2; TIR1::mRuby animals exposed to auxin for 48 h. These studies did not reveal bra-2 as a major contributor to global transcriptional regulation of known meiotic factors, except for reductions in syp-2 and dsb-2 transcript levels (Supplementary Fig. 4b–f). The reduced expression of syp-2 is unlikely to be the origin of the severe loss of synapsis associated with BRA-2 removal, since synapsis is unaffected in mutants with similarly decreased levels of syp-2 expression, and highly reduced levels of SYP-2 can still support SC formation49,50,51,52 (see Discussion). These data strongly suggest that BRA-2 is not a critical regulator of global meiotic genes transcription and that the synapsis defects observed in its absence are not a secondary consequence of aberrant gene expression.
BRA-2 promotes elongation of the SC but not its maintenance
We observed that in HA::AID::bra-2; TIR1::mCherry animals exposed to auxin for 24 hours, nuclei at meiotic entry displayed similar SC defects to those observed in bra-2 null animals, however, those at later stages (LP) remained fully synapsed (Supplementary Fig. 3h).
Germ cells advance synchronously from the distal tip of the gonad onwards, with a pace of approximately one nuclear row/hour53,54. Therefore, we reasoned that impaired synapsis in nuclei specifically at TZ-EP would be consistent with BRA-2 exerting more prominent roles at earlier rather than later stages. In support of this, we observed that if exposure to auxin was prolonged for 48 hours, thus allowing meiocytes to travel further within the gonad, then synapsis defects were in fact detected throughout pachytene, fully phenocopying the bra-2 null mutants (Supplementary Fig. 5a).
To further corroborate this, we undertook two different approaches: first, we exposed HA::AID::bra-2; TIR1::mRuby animals to auxin for 3 hours to achieve nearly complete BRA-2 depletion in the whole germ line (Supplementary Fig. 1a) and then we moved the animals onto plates without auxin to allow for BRA-2 resynthesis. We reasoned that analyzing the depleted animals at different times during recovery/resynthesis would allow us to assess how cells that were at different substages of meiotic progression during auxin exposure responded to BRA-2 depletion.
Given that it takes 12–14 hours for nuclei to travel from the mitotic tip to TZ53,54,55,56, we let the depleted animals recover for slightly longer (16 hrs) to ensure that cells had enough time to enter meiosis. As shown in Fig. 4d, nuclei in zones 3 and 4 (corresponding to TZ/EP), that were in the premeiotic tip/meiotic entry at the time of the depletion, displayed comparable SC impairment to worms constantly exposed to auxin without recovery. Nuclei in zones 5 and 6 (corresponding to MP/LP), depleted for BRA-2 when they were already beyond TZ, instead displayed levels of synapsis comparable to control animals.
In a second approach, we treated the HA::AID::bra-2; TIR1::mRuby animals as above, but we included incubation with EdU during the last 20 minutes of the 3h-exposure to auxin, followed by the 16 h of recovery. Since the EdU is incorporated into the DNA only during premeiotic DNA replication, this allowed us to clearly distinguish the germ cells that engaged meiosis after depletion of BRA-2 (EdU-positive) from those that had entered meiosis before/during the depletion (EdU-negative) (Fig. 4e). As shown in Fig. 4f, the cells displaying unsynapsed chromosomes were also positive for EdU (arrowheads) and conversely, EdU-negative cells showed full synapsis. Importantly, Western blot analysis showed that significant resynthesis of BRA-2 protein levels occurred within 24-48 h (Supplementary Fig. 5b), ruling out any major contribution as a result of its reappearance and resumption of its function within the 16 hours of the recovery time window. Altogether, these data indicate that BRA-2-mediated regulation of chromosome synapsis occurs at meiotic entry and suggests that BRA-2 is essential for SC polymerization rather than its maintenance.
BRA-2 and HIM-17 promote autosome pairing and synapsis in a partially redundant manner
Our results so far indicate that BRA-2 exerts crucial roles in promoting SC polymerization upon successful homology assessment, and given its physical interaction with HIM-17, we sought to investigate whether bra-2 and him-17 may also genetically interact. We employed different alleles of him-17, and we also generated a functional him-17::AID tagged line by CRISPR to facilitate genetic manipulation of strains.
We built the HA::AID::bra-2; TIR1::mRuby; him-17::AID strain and assessed pairing and synapsis before and after auxin exposure. FISH analysis revealed that, strikingly, chromosome V pairing levels were dramatically impaired when BRA-2 and HIM-17 (Fig. 5a) were simultaneously absent, reaching only 30% in comparison to the normal pairing levels observed in HA::AID::bra-2; TIR1::mRuby; him-17::AID controls (-auxin). Previous studies have shown that while being essential for meiotic DSB induction, him-17 is dispensable for both pairing and synapsis41, and therefore this result was unexpected.
a Left: quantification of chromosome V pairing in HA::AID::bra-2; TIR1::mRuby; him-17::AID worms before and after auxin exposure. At least three germ lines were analyzed and the number of nuclei scored (-auxin vs +auxin respectively) was: zone 1 (155, 255), zone 2 (172, 260), zone 3 (211, 253), zone 4 (173, 239), zone 5 (106, 143). Bars indicated mean with S.E.M. and asterisks denote statistical significance assessed by the Χ2 test (two-sided, CI. = 0.05, *p < 0.0001). Right: representative examples of early pachytene nuclei analyzed by FISH (Ch. V FISH probe in yellow, DNA in blue). Scale bar 5 μm. b Analysis of phosphorylated-SUN-1S8 (green) in whole-mount gonads in the indicated genetic backgrounds, before and after exposure to auxin. DAPI is shown in purple. Scale bar 10 μm. The analysis was performed in biological duplicates. c SYP-1 (green)/HTP-3 (red) immunofluorescence in early-mid pachytene nuclei of HA::AID::bra-2; TIR1::mRuby control animals and in him-17(e2707) mutants, before and after exposure to auxin. DAPI is shown in blue. Scale bar 5 μm. Analysis was performed in biological triplicates. d Co-staining of SYP-1 (magenta)-HIM-8 (yellow) reveals that only the X chromosome is proficient in SC assembly upon contemporary absence of BRA-2 and HIM-17. DAPI shown in blue. Scale bar 5 μm. Analysis was performed in biological triplicates.
We also combined the HA::AID::bra-2; TIR1::mRuby with him-17(ok424) deletion allele, likely to be a him-17 null, and assessed localization of phosphorylated-SUN-1S8 to assess whether the pairing defects that we observed resulted from impaired NE recruitment of the protein machinery required for chromosome movement. We found that phosphoSUN-1S8 was abundantly present and extended until late pachytene stage (Fig. 5b) as similarly observed in synapsis-defective mutants11,21, indicating efficient SUN-1-mediated chromosome motion but delayed exit from the homology search state.
Analysis of SC assembly revealed that loss of both BRA-2 and HIM-17 abrogated synapsis from the TZ and mid-pachytene nuclei, and SYP-1 was detected only in late pachytene nuclei mostly as a single stretch, possibly suggesting synapsis of a single chromosome pair (Fig. 5c and Supplementary Fig. 6a). In fact, co-staining using anti-HIM-8 antibodies revealed that the X chromosome was the only one retaining synapsis at late pachytene stage in him-17 mutants depleted for BRA-2 (Fig. 5d, Supplementary Fig. 6b). These results indicate that both proteins are essential for pairing and synapsis of the autosomes but not for the sex chromosomes. This was further corroborated by the finding that X chromosome synapsis was only mildly affected upon loss of BRA-2 alone (Supplementary Fig. 6c) (consistent with the global synapsis impairment triggered by loss of BRA-2) and indicating that the X chromosomes do not have a specific tendency to being unsynapsed.
We ruled out that lack of pairing and synapsis originated from impaired meiotic entry by co-staining WAPL-1 and phosphorylated SUN-1S8, since the former is lost from the nucleus upon entry into meiosis while the latter appears at the nuclear envelope. WAPL-1 and pSUN-1S8 displayed mutually exclusive localization before and after exposure to auxin (Supplementary Fig. 6d), further indicating that pairing and SC defects are not caused by a defective mitosis-to-meiosis switch.
It has been previously shown that him-17 establishes epistatic relationships with two other pro-DSB factors in C. elegans, namely xnd-1 and him-557. Thus, we wondered whether the synthetic synapsis defects observed under simultaneous removal of BRA-2-HIM-17 could be similarly triggered by lack of BRA-2 and these proteins as well. Removal of HIM-5 or XND-1 in BRA-2-depleted animals did not further impair synapsis (Supplementary Fig. 6e-f), indicating a specific role for him-17 in promoting pairing and SC assembly in BRA-2-depleted animals.
HIM-17 functions in promoting synapsis are independent of its roles in DSB induction
We next sought to investigate whether the impaired synapsis observed under simultaneous removal of HIM-17 and BRA-2 was linked to reduced DSB formation, since him-17 activity has been shown to be essential for break induction41. We used RAD-51 (the C. elegans ortholog of E. coli RecA58,59) foci as an indirect read out of DSBs, since a direct marker for meiotic breaks is not available in worms. Meiotic DSB repair in C. elegans is entirely dependent on RAD-51, since DMC1 is not present. Once DSBs are formed and resected, RAD-51 is recruited to the recombination intermediates where it promotes displacement and invasion of the homologous template to achieve recombination. Cytological detection of RAD-51 foci follows a reproducible pattern in WT animals, and in the absence of SPO-11-mediated DSB induction, no RAD-51 foci are detectable in the germ line (except for occasional replication-dependent DNA damage). Under conditions in which synapsis or CO formation are compromised, RAD-51 foci accumulate to high numbers and their clearance from the chromatin is strongly delayed42.
First, we assessed whether BRA-2 has a role in DSB formation, and we found that HA::AID::bra-2; TIR1::mRuby animals exposed to auxin for 24 h displayed a dramatic accumulation of RAD-51 foci as pachytene progressed, consistent with impaired SC formation42 (Fig. 6a, b). This shows that DSBs are generated upon BRA-2 depletion, indicating that unlike HIM-17, BRA-2 is not required for DSB induction. As expected, depletion of HIM-17 severely abrogated RAD-51 foci formation, which was similarly impaired under HIM-17-BRA-2 co-depletion (Fig. 6a, b).
a Analysis of RAD-51 foci formation in the indicated backgrounds, in absence or presence of auxin. Exposure to auxin triggers extensive RAD-51 accumulation in BRA-2-depleted animals, which is suppressed by HIM-17 co-depletion. Bars indicate S.E.M. At least three germ lines for each genetic background were employed, and the number of nuclei analysed in (-auxin) samples from zone 1 to 7 respectively, was: HA::AID::bra-2 (191, 182, 235, 203, 194, 163, 104), him-17::AID (234, 266, 182, 177, 146, 137, 125), HA::AID::bra-2; him-17::AID (180, 241, 209, 171, 151, 134, 113). The number of nuclei analysed in (+auxin) samples from zone 1 to 7 respectively, was: HA::AID::bra-2 (243, 215, 154, 167, 152, 114, 94), him-17::AID (164, 259, 218, 181, 154, 106, 101), HA::AID::bra-2; him-17::AID (158, 219, 226, 221, 206, 141, 112). b Representative images of mid-pachytene nuclei in the indicated backgrounds and conditions, stained for RAD-51 (red)/SYP-1 (green) and counterstained by DAPI. Scale bar 5 μm. The analysis was performed in biological duplicates. c Mid-pachytene nuclei of worms of the indicated genotype exposed to auxin before and after IR showing that lack of synapsis does not depend on DSBs. HTP-3 is shown in red and SYP-1 in green. DNA is shown in blue. Scale bar 5 μm. The analysis was performed in biological duplicates. d Mid-pachytene nuclei of worms of the indicated genotype exposed to auxin showing that lack of synapsis does not depend on SPO-11 activity. HTP-3 is shown in red and SYP-1 in green. DNA shown in blue. Scale bar 5 μm. The analysis was performed in biological duplicates.
Next, we exposed worms lacking both HIM-17 and BRA-2 to ionizing radiation to assess whether exogenous DSBs could suppress the SC defects by performing analysis of HTP-3 and SYP-1 staining 4 hours after irradiation. This revealed no differences between test and non-irradiated BRA-2-depleted animals and indicates that the SC defects are independent of DSB formation (Fig. 6c).
As a complementary approach, we simultaneously depleted SPO-11 and BRA-2, as we reasoned that if lack of meiotic DSBs caused synapsis defects in the bra-2; him-17 then this phenotype should be recapitulated in the bra-2; spo-11. Also in this case, synapsis defects were indistinguishable from animals depleted only for BRA-2 (Fig. 6d), once more demonstrating that the synapsis defects occurred independently of DSBs per se. These results suggest that HIM-17 exerts roles in promoting SC assembly which are not linked to its functions in mediating DSB induction, unveiling a previously unknown role for this protein in promoting pairing and synapsis.
Loss of BRA-2 restores CHK-2 activity in htp-1 mutants
Previous work has shown that successful synapsis between homologous chromosomes requires functional dynein, since under conditions of impaired dynein function, homologs are still capable of attaining pairing (although with delayed kinetics) but chromosomes fail to synapse, resulting in the accumulation of SC subunits into polycomplexes and a persistent clustering of the chromatin7. As previously mentioned, sun-1 and htp-1 have been shown to be essential for this mechanism, since in their absence, the constraints on SC polymerization imposed by lack of dynein are bypassed and synapsis ensues between non-homologous partners7.
In bra-2 mutants, SC elongation is defective but pairing is largely successful, indicating that an active barrier to prevent ectopic synapsis is intact, and therefore we wondered whether removal of htp-1 would circumvent the defects in SC assembly.
To this end, we combined the HA::AID::bra-2; TIR1::mRuby with htp-1(gk174) mutants and assessed PLK-2/SYP-1 staining. PLK-2 undergoes a dynamic localization in the germ line: it concentrates into aggregates at the nuclear periphery during TZ stage; and, it redistributes along the SC in pachytene in response to CO site designation and to promote timely inactivation of CHK-215,20,60. Importantly, PLK-2 can delay exit from TZ (visible as prolonged nuclear clustering) in the presence of defects in synapsis or recombination, allowing the CHK-2-dependent signaling to be extended15,20,21,61.
Abrogation of htp-1 function triggers precocious exit from TZ and extensive non-homologous synapsis, therefore htp-1 null mutants lack a defined region of the gonad containing nuclei with clustered chromatin26,27 and display a dramatic reduction of PLK-2/pSUN-1S8 at the nuclear envelope61.
PLK-2 aggregates at the nuclear periphery were barely detectable at meiotic entry in htp-1 mutants (Fig. 7a, TZ, -auxin, and Supplementary Fig. 7), and no clearly defined nuclear polarization was observed, recapitulating previous findings61. Exposure of HA::AID::bra-2; TIR1::mRuby animals to auxin triggered retention of PLK-2 at the PCs (Fig. 7a TZ and MP, +auxin,), in line with lack of synapsis and indicating a prolonged state of chromosome movement. Strikingly, removal of BRA-2 in htp-1 mutants restored chromatin clustering, which consistently overlapped with persistent PLK-2 localization at the nuclear envelope until MP stage (Fig. 7a, TZ and MP, +auxin, and Supplementary Fig. 7). SYP-1 loading appeared greatly diminished compared to both BRA-2-depleted worms and htp-1 single mutants, as only one or two SYP-1 tracks were detectable in MP nuclei of worms lacking both BRA-2 and HTP-1 (Fig. 7b), consistent with previous data that HTP-1 is important not only for homologous pairing but also for SC processivity7. This indicates that lack of BRA-2 largely abrogates the indiscriminate SC assembly triggered by absence of HTP-1 and restores establishment and maintenance of chromosome clustering, indicating active CHK-2-mediated signaling. Previous work has shown that removal of htp-2 in htp-1 null mutants reduces SC formation (and consequently non-homologous synapsis)27, and slightly shortens the CHK-2 active zone when synapsis is also abrogated62. Thus, we generated the HA::AID::bra-2; TIR1::mRuby htp-1 htp-2 strain and assessed PLK-2 localization to address whether HTP-2 was playing any roles in sustaining the checkpoint activation triggered by BRA-2 depletion in htp-1 mutants. As shown in Supplementary Fig. 7, PLK-2 localization in htp-1 htp-2 double mutants does not dramatically differ from htp-1 nulls, except for a prominent accumulation as a single bright aggregate which is likely to correspond to the X chromosome PC21,61,63. A similar PLK-2 localization pattern has been also recently shown by Barroso and colleagues upon over-expression of HTP-2 in worms lacking htp-1, indicating that HTP-2 is unable to functionally replace HTP-1 regardless of its abundance along the chromosome axes64. Depletion of BRA-2 in htp-1 htp-2 double mutants triggered a comparable response as observed in the htp-1 single mutants, indicating that HTP-2 does not function in promoting checkpoint activation in htp-1 mutants depleted for BRA-2.
a Nuclei in transition zone (TZ) or mid-pachytene (MP) stained for PLK-2 (yellow)/SYP-1 (magenta) counterstained by DAPI (cyan) in HA::AID::bra-2; TIR1::mRuby control animals and htp-1(gk174) mutants, before and after auxin exposure (24 h). Scale bar 5 μm. Analysis was performed in biological duplicates. b Mid-pachytene (MP) nuclei stained for HTP-3 (red)/SYP-1 (green) in the indicated genetic backgrounds before and after exposure to auxin. Scale bar 5 μm. Analysis was performed in biological duplicates. c Quantification of nuclei with paired signals for Ch. V in the indicated genetic backgrounds before and after exposure to auxin. Bars depict S.E.M. and asterisks indicate statistical significance as assessed by Χ2 test (two-sided, CI. = 0.05, *=p < 0.0001, ns= non- significant). At least three germ lines for each genetic background were employed for quantification, and the number of nuclei scored was: HA::AID::bra-2 (-auxin, zone 1-5) 150, 179, 175, 146, 110; HA::AID::bra-2; htp-1 (-auxin, zone 1-5) 128, 139, 148, 174, 122; HA::AID::bra-2 (+auxin, zone 1-5) 198, 208, 200, 200, 126; HA::AID::bra-2; htp-1 (+auxin, zone 1-5) 155, 174, 175, 163, 116. d Quantification of non-homologous synapsis in the indicated genetic backgrounds and exposure conditions to auxin only in the pachytene region. Quantifications were performed on the same samples quantified in (c), but only in nuclei spanning the pachytene region and corresponding to zones 4 (EP/MP) and 5 (MP/LP). Bars indicate S.E.M. and asterisk denotes statistical significance assessed by Χ2 test (two-sided, CI. = 0.05, ns= non-significant, *p < 0.0001). e Representative images of mid-pachytene nuclei showing FISH for Ch. V (orange) and anti-SYP-1 (light green) immunostaining, counterstained by DAPI (purple). Scale bar 5 μm. Analysis was performed in biological triplicates.
Given that chromatin clustering results from vigorous chromosome movement, which in turn is essential for robust synapsis between homologous chromosomes, we wondered whether the rescue of TZ coupled with efficient PLK-2 recruitment at the nuclear envelope were also indicative of improved pairing levels in the htp-1 mutants depleted for BRA-2. To this end, we performed FISH analysis to monitor pairing of chromosome V. To make sure that LP nuclei were affected by BRA-2 depletion, we exposed the HA::AID::bra-2; htp-1 TIR1::mRuby animals to auxin for 48 h instead of 24 h, as we have previously shown that at this time point also cells at later stages display SC defects (Supplementary Fig. 5a).
While initial pairing of chromosome V was improved (zone 1 - 43.8% in BRA-2-depleted htp-1 mutants versus 10.1% in non-depleted), removal of BRA-2 did not bypass the constraint imposed by htp-1 function in achieving robust chromosome coalignment in the rest of the germ line (Fig. 7c), since the percentage of nuclei with unpaired chromosomes from zone 2-6 was comparable to htp-1 mutants. We also noticed that in the HA::AID::bra-2; TIR1::mRuby animals, there was a higher frequency of nuclei with unpaired signals compared to bra-2(ok1171) nulls (Fig. 3). We hypothesize that this might be a consequence of the age difference of the worms analyzed between the two experiments, as worms were analyzed 24 h post-L4 when the bra-2(ok1171) deletion mutant was used, versus 48 h post-L4 in the bra-2 degron strain. When we combined FISH with SYP-1 immunostaining we found that chromosome V was engaged in non-homologous synapsis (assessed by monitoring whether one or both probe signals were associated with SYP-1) in all pachytene nuclei scored in the htp-1 null mutants, recapitulating previous data26,27. In contrast, non-homologous synapsis was reduced to levels observed in control animals upon BRA-2 depletion in pachytene cells (Fig. 7d, e), consistent with the fact that BRA-2 removal abrogates indiscriminate synapsis triggered by htp-1 loss of function.
These results indicate that when homology is not detected, the removal of BRA-2 restores a barrier to indiscriminate SC assembly by bypassing the constraints imposed by HTP-1, although it is not sufficient to suppress the loss of pairing caused by the abrogation of htp-1 function.
Loss of HIM-17 partially alleviates pairing and synapsis defects in htp-1 mutants
Unlike bra-2, him-17 is dispensable for synapsis41,65 (this study). However, the synthetic effects that we observed when BRA-2 and HIM-17 were simultaneously removed suggest that HIM-17 could be part of an accessory/parallel pathway that is activated upon dysfunctional pairing/synapsis, or that it could play partially overlapping roles with BRA-2.
To investigate these possibilities, we assessed whether removal of him-17 in htp-1 mutants altered SC dynamics and/or pairing as similarly observed upon BRA-2 depletion (Fig. 7). Indeed, htp-1; him-17 double mutants showed a well-defined TZ with nuclei displaying a clustered chromatin morphology (Fig. 8a). This was coupled with reduced SYP-1 loading, which localized predominantly as chromosome-associated puncta at meiotic entry/EP, and wild-type-like clusters of PLK-2-labeled chromosome ends at the nuclear periphery (Fig. 8b). Compared to htp-1 animals depleted for BRA-2 however, the window of PLK-2 localization at the nuclear envelope was narrower in the htp-1; him-17 double mutants, extending to a size comparable to WT animals (Fig. 8b). Strikingly, FISH analysis revealed that in the htp-1; him-17 double mutants the pairing of chromosome V in TZ (corresponding to Zone 1) was not different from controls, and it was steadily maintained throughout the germ line (Fig. 8c, d). Combined analysis of FISH and immunostaining for SYP-1 also showed that paired chromosome V was largely synapsed (Supplementary Fig. 8e). Thus, our data suggest that removal of HIM-17 in htp-1 mutants not only restores chromosome clustering early on, but also allows the homologs to align and remain synapsed thereby preventing non-homologous synapsis.
a Analysis of SYP-1 localization in the indicated mutant backgrounds. Red asterisks indicate nuclei with clustered chromatin. Scale bar 5 μm. Analysis was performed in biological duplicates. b Top: SYP-1 (magenta)/PLK-2 (yellow) immunofluorescence in the indicated mutants and stages. (TZ= Transition Zone, EP= Early Pachytene, MP= Mid-Pachytene and LP= Late-Pachytene). Scale bar 10 μm. Bottom: quantification of PLK-2 in the indicated genotypes. The number of germ lines analyzed was WT (3), him-17(ok424) (6), htp-1(gk174) (4), htp-1(gk174); him-17(ok424) (5). Bars indicate S.E.M., and asterisks show statistical significance assessed by T test (two-tailed, 95%). WT=htp-1 (p = 0.0057), WT=him-17 (p = 0.023), him-17=htp-1 and him-17= htp-1; him-17 (p = 0.0095), htp-1= htp-1; him-17 (p = 0.028). The quantifications were performed in biological duplicates, and at least three germ lines were employed for quantifications. c Quantification of Ch. V pairing in the indicated mutants. At least four germ lines for each genetic background were employed for quantification, and the number of nuclei scored was (zone 1-5): WT -126, 152, 128, 107, 80; htp-1(gk174) - 357, 361, 361, 331, 214; him-17(ok424) – 197, 209, 211, 145, 114; htp-1(gk174); him-17(ok424) – 377, 380, 361, 303, 202. Bars indicate S.E.M., and asterisks denote statistical significance as assessed by Χ2 test (two-sided, CI = 95%, *p < 0.0001). d Representative images of Ch. V FISH (yellow) and DAPI (cyan) in mid-pachytene nuclei in the indicated mutants. Scale bar 5 μm. Analysis was performed in biological triplicates. e Ch. V FISH (green) and SYP-1 (magenta) immunostaining in the indicated mutant backgrounds. Scale bar 5 μm. The analysis was performed in biological duplicates. f Mid-pachytene nuclei stained for HTP-3 (red)/SYP-1 (green) in the indicated genetic backgrounds and conditions of exposure to auxin. All strains analyzed carried the TIR1::mRuby expressing transgene in the background. Scale bar 5 μm. The analysis was performed in biological duplicates.
Next, we asked whether the phenotypes observed in the htp-1; him-17 double mutants might originate from impaired DSB formation, given that HIM-17 is required for break induction. To this end, we generated the spo-11(ok79) htp-1(gk174) double mutants and analyzed PLK-2 localization. This showed that removal of SPO-11 did not affect the extent of PLK-2 recruitment in the absence of htp-1 nor restored presence of nuclei with clustered chromatin in TZ (Supplementary Fig. 8), indicating that the accumulation of PLK-2 and the delay in SYP-1 loading observed in htp-1; him-17 double mutants are not caused by absence of meiotic DSBs.
The pairing and synapsis phenotypes observed in the htp-1; him-17 mutants differed from those observed in the bra-2; htp-1 animals, since in the former we found only delayed SYP-1 loading associated with strongly improved pairing levels, while in the latter, synapsis was almost entirely abrogated but pairing levels were not different from htp-1 single mutants. Therefore, we decided to deplete both BRA-2 and HIM-17 to assess their epistatic relationships in influencing pairing and synapsis establishment in germ lines lacking htp-1. The SC impairment observed in worms co-depleted for BRA-2 and HIM-17 was unchanged after removal of htp-1 (Fig. 8f), as well as the pairing defects (Supplementary Fig. 8b), suggesting that BRA-2-HIM-17-mediated roles in regulating pairing and synapsis occur upstream/irrespectively of HTP-1 presence.
In conclusion, our work identifies regulatory circuits that promote homologs pairing and release constraints to SC elongation upon homology assessment. These activities are differently stimulated by absence of BRA-2 and HIM-17 and importantly, their loss reinstates a barrier to uncontrolled synapsis contingent to homology, that bypasses the requirements imposed by HTP-1 function.
Discussion
Our work identifies the MYND-ZnF domain containing protein BRA-2 as a HIM-17 interactor and shows that it exerts crucial functions in regulating SC polymerization. Lack of BRA-2 impairs SC assembly with only minor effects on homolog pairing, resulting in defective CO establishment and reduced chiasmata formation. Loss of BRA-2 reduces the polymerization rate of SC central components concomitant with protein destabilization. Live imaging analysis of synapsis establishment and conditional degron-mediated depletion of BRA-2 reveal that the key function of the protein during synapsis is in regulating SC elongation upon homology assessment rather than SC maintenance, and strikingly, co-depletion with HIM-17 reveals partially overlapping activities that coordinate homolog pairing with establishment of synapsis.
BRA-2 as a relay for SC assembly upon homology assessment
At meiotic entry, homologous chromosomes must align pairwise to successfully undergo synapsis and recombine. However, SC assembly can take place irrespective of homology and consequently controlling systems must be in place to properly coordinate processive SC polymerization only once homology assessment has been satisfied. How co-aligning companions recognize one another, remains one of the most important unanswered questions in the field, and although our data do not directly address homology assessment, they reveal important aspects of the regulatory mechanisms that ensure the release of the constraints to SC assembly at the right time and between the right partners.
We observed that the absence of BRA-2 does not grossly compromise homolog coalignment, as evidenced by largely normal pairing levels (Fig. 3), that were steadily maintained throughout the pachytene stage, indicating a stable association between the homologs. Synapsis on the other hand, was dramatically impaired, although chromosome ends containing the PC were associated with some residual SYP-1, as observed by FISH combined with immunofluorescence (Fig. 3). This would suggest that while BRA-2 is dispensable to achieve pairing and initial synapsis at chromosome ends, it is essential for transmitting a downstream “pairing OK” signal which allows polymerization of the SC along the entire chromosome length.
Previous work has shown that indeed the SC is nucleated first at PC sites and then is polymerized along the entire length of the chromosomes with high processivity9. In line with this, our analysis of GFP::SYP-3 dynamics in live animals has revealed that the formation of the putative SYP-3 nucleation sites shows only a slight reduction under BRA-2 depletion, however 50% of these foci never undergo elongation and those that manage to linearize, do so with a substantially reduced elongation rate (Fig. 4).
Our RNA-seq data indicate that loss of BRA-2 causes reduced expression of dsb-2, involved in DSB formation66, and syp-2, one of the central elements of the SC that is essential for establishment of synapsis42. In C. elegans, all central elements (SYP-1, -2, -3, -4, -5, -6 and SKR-1, -2) are loaded inter-dependently42,44,67,68,69,70,71, and therefore, reduced expression/loading of one of the subunits could drive the destabilization of all others. Consequently, the defective SC polymerization we observe in BRA-2-depleted animals could be explained by a limiting pool of SYP proteins. However, while we cannot definitively exclude a contribution of the reduced syp-2 expression to the synapsis defects that we observe in bra-2 mutants, several lines of evidence support the interpretation that these phenotypes are unlikely to be a function of reduced synapsis per se.
Recent work has shown that absence of him-17 causes a reduction of syp-2 expression comparable to that we observe in the absence of BRA-252, however, him-17(ok424) null mutants display normal synapsis41,65 (this study), indicating that such a reduction in syp-2 expression does not trigger, per se, SC assembly defects. Furthermore, any possible consequences of SYP-2 reduction that occur in bra-2 mutants are also unlike those previously reported for syp-2/+ heterozygotes, and for syp-1 partial RNAi conditions. Libuda and colleagues50 have shown that reduced syp-1 levels (comparable to SYP-2 reduction in our analysis) cause an overall decrease in intensity of SYP-1 signal relative to the axis (~65%), coupled with some missing or undetectable short stretches of SC. In the case of syp-2/+ germlines, the reduction of syp-2 resulted in an increase in the number of small gaps in long SYP-1-stained segments that were of wild-type intensity, and were limited to early pachytene, as at later stages, full length SC was maintained51. In bra-2 mutants, some TZ nuclei showed one or two short bright SYP-1 segments while only 10% of pachytene nuclei achieved full synapsis (Fig. 4). Consequently, our phenotype significantly deviates from previously published observations in that the reduction of SYP-2 is not evidenced as gaps or an overall decrease in SYP-1 levels at the SC, but instead SYP-1 appears robust and contiguously localized to a reduced number of axes only at advanced pachytene stages. The more robust SC staining that we observe in late pachytene nuclei in bra-2 mutants is likely to also be a consequence of the relaxation of the constraints imposed on SC assembly that occur at this stage26.
A further crucial piece of evidence in support of our model, is provided by the rescued nuclear clustering and restored PLK-2 loading that we observe in the bra-2; htp-1 double mutants, which was not observed in syp-2; htp-1 double mutants26,27,64. This indicates that if the phenotypes observed in bra-2 mutants were mainly caused by reduced/impaired synapsis per se, we then would have expected that the htp-1 worms depleted for BRA-2 would show a similar phenotype to htp-1; syp-2 double mutants, which is in stark contrast with our observations (Fig. 7 and Supplementary Fig. 7).
Lastly, it has been shown that early pairing takes place independently of the SC, which becomes essential for stabilizing the association between the homologs later on42,44,69,70. Unlike what has been observed in synapsis-defective mutants42,44,67,69,70, we find steady-state pairing levels in the bra-2 mutants despite the dramatic impairment in synapsis. This additionally reinforces the interpretation that BRA-2 is not required for the initial stabilization and maintenance of the PCs associations, and that its activity is instead necessary to promote elongation of the SC along the entire chromosome length following homolog recognition.
HTP-1 activity in preserving a homology-contingent barrier to synapsis becomes dispensable in absence of BRA-2 or HIM-17
Several studies have shown that in C. elegans, pairing-defective mutants fail to acquire a clustered chromatin configuration at meiotic entry. This holds true in the case of mutations in factors directly involved in the assembly of the SUN/KASH module SUN-1/ZYG-127,11, but also in mutants of genes involved in regulatory activities of meiotic progression such as prom-1 and him-1972,73, the meiotic kinases mutants CHK-2 and PLK-215,20,45, and in the HORMA-domain containing protein family23,25,26. Together with impaired nuclear polarization, several pairing-defective mutants display deregulated establishment of synapsis, which occurs between non-homologous chromosomes. Lack of synapsis alone on the other hand, triggers prolonged nuclear clustering coupled with extended recruitment of PLK-2 at the PCs and phosphorylation of SUN-1 (a marker for the time window when chromosome movements take place). Impaired SC formation, however, is not always sufficient to trigger asymmetric chromatin reorganization, as indicated by lack of a defined TZ in the axial element mutants htp-3 or him-3, both structurally essential for the loading of SC central elements23,24,25. This suggests that multiple levels of control are integrated both at the nuclear envelope, as well as at the interface with the chromosome axes to allow chromatin redistribution and prolonged permanence into a homology search state when synapsis and/or recombination are defective.
Previous work has established the model wherein dynein-driven forces exerted on the PCs promote timely establishment of pairing and allow SC polymerization once homology has been satisfied7, resulting in the exit from homology search state and redistribution of chromatin in the nuclear space. Under impaired dynein function, homologs still manage to attain (delayed) pairing coupled with chromosome clustering, however synapsis is prevented and the SC subunits form nuclear polycomplexes. SUN-1 and the axial protein HTP-1 have been shown to be essential for establishing and/or maintaining this barrier to SC formation7,11,18,26,27.
Our data indicate that loss of BRA-2 or HIM-17 is sufficient to restore the barrier that prevents illegitimate SC assembly, since both bra-2; htp-1 and htp-1; him-17 double mutants, as well as bra-2; htp-1; him-17 triple mutants, show extensive nuclear clustering and non-homologous synapsis is abrogated despite complete loss of autosome pairing. Interestingly, in the bra-2; htp-1 mutant background pairing showed an initial improvement in TZ, which was not maintained throughout pachytene, where it reached comparable levels to those observed in the htp-1 single mutants (Fig. 7c). This could indicate that abrogation of illegitimate synapsis caused by removal of BRA-2 in htp-1 mutants might facilitate the initial coalignment of the homologs which, however, lacks sufficient stabilization due to complete absence of SC, since overall synapsis in bra-2; htp-1 doubles was further reduced in comparison to bra-2 or htp-1 single mutants. This highlights the essential roles that HTP-1 plays in promoting homolog coalignment and also in stimulating SC processivity, as previously reported7,26,27. In alternative, simultaneous abrogation of bra-2 and htp-1 function could result in changes into chromatin configuration that could favor early homologs association but not maintenance of their interaction. Importantly, failure in attaining pairing was not caused by impaired movement of chromosome ends, as indicated by extended nuclear polarization and recruitment of PLK-2/pSUN-1S8. Homolog pairing and subsequent homologous SC assembly were strongly improved in htp-1; him-17 double mutants, suggesting that BRA-2 and HIM-17 play separable roles during pairing and have partially overlapping functions during SC establishment.
The loss of autosomal pairing and synapsis along the autosomes observed in the bra-2; him-17 double mutant could be due to a failure at the level of homolog recognition, which in turn prevents SC polymerization via an intact dynein-mediated functional barrier. Alternatively, loss of BRA-2-HIM-17 might weaken/prevent a stable association between the PCs, resulting in a deficiency in overcoming the forces opposed by dynein function, thereby prolonging the window of the homology search state and blocking SC assembly. This would be in line with our finding that both PLK-2 and pSUN-1S8 are abundantly recruited at the nuclear envelope in absence of bra-2 and him-17, indicative of active CHK-2 signaling and no obstacles to chromosome movement. Therefore, BRA-2 and HIM-17 might cooperate to promote the stabilization of the initial interaction between the PCs and/or support the homology assessment step, or perhaps they could play roles in limiting CHK-2 activity. Work in mammals has shown that ZMYND11 is important to assemble a co-repressor protein surface that in turn regulates RNA pol II activity during elongation33,34 rather than directly tweaking transcription levels. It is tantalizing to envision a scenario where perhaps loss of BRA-2-HIM-17 might release a constraint on factor(s) that promote/signal exit from the homology search state, which would result in prolonged permanence in TZ in the absence of BRA-2 and HIM-17.
Alternatively, previous work in budding yeast has suggested a mechanical model which could also provide insights into the possible interplay between BRA-2/HIM-17 and HTP-1 function. In this scenario, chromatin loop expansion extends the axis and blocks SC formation until the axis is relaxed, an event that can be triggered by CO designation and results in local SC nucleation and spreading at a distance1,74,75,76. To ensure that full SC polymerization is accomplished, global axis relaxation occurs at late zygotene stage. In this context, we could envision a scenario whereby chromatin-localized BRA-2 (and to some extent HIM-17), negatively regulates chromatin expansion, while axis-localized HTP-1 blocks axis relaxation and concomitant SC formation. Consequently, bra-2 mutants allow more chromatin expansion and axis extension that impedes SC formation, while htp-1 mutants have constitutively relaxed axes that permit nonhomologous SC formation. This is consistent with the phenotypes that we observe both in the bra-2; htp-1 and htp-1; him-17. In conclusion, our work provides evidence of a complex, multilayered regulatory network tasked with unleashing synapsis to stabilize homolog interactions with precise timing (Fig. 9), and further studies will be necessary to elucidate the molecular mechanisms underlying these processes and to frame BRA-2-HIM-17 activities during the establishment of pairing and synapsis.
Graphical schematization and table summarizing pairing, synapsis, and extent of chromosome clustering observed in the indicated genetic backgrounds.
Methods
Strain maintenance and genetics
All strains were cultured according to standard procedures77. Worms were grown on nematode growth medium (NGM) plates seeded with OP50 E. coli as a food source and maintained at 20 °C for all the experiments. To assess embryonic lethality and male progeny, L4 worms of the relevant genotype were individually picked, and mothers were moved onto fresh plates every 24 hours. Dead-eggs/hatched-eggs were scored the day after the mothers were moved and male progeny three days later. The percentage of embryonic lethality was calculated as a total number of dead eggs/total number of laid eggs. CRISPR-tagged strains were generated by SunyBiotech (https://www.sunybiotech.com/), the him-17::3XHA tagged line had been previously generated by CRISPR31, and the him-17::GFP line was generated by biolistic bombardment41. All strains generated by CRISPR, were outcrossed twice to WT worms prior usage. A complete list of strains used in this study is reported in Supplementary Data 2.
Immunofluorescence studies
Worms of the indicated age and genotype were dissected in 15μl of 1XPBS containing 0.1% Tween (PBST) on a 22×22 coverslip. Gonads were fixed by adding 15μl of 2% PFA in 1XPBST and fixation was carried out for 5 minutes at room temperature. Coverslips were flicked off after briefly dipping the slides into liquid nitrogen and samples were transferred to methanol at −20 °C for at least 5 minutes. Slides were washed three times in 1X PBST and blocked for 1 h at room temperature in 1% BSA in 1XPBST. Primary antibodies were allowed to incubate overnight at room temperature in a humid chamber and the following day, slides were washed three times in 1XPBST for 10 minutes each. Incubation of secondary antibodies was carried out for 2 h at room temperature in a humid chamber in the dark. Slides were washed three times in 1XPBST for 10 minutes each in the dark and a 60μl of a DAPI solution in water (2μg/ml) was added on top of the dissected worms. After 1 minute, slides were washed in 1XPBST for at least 20 minutes in the dark, mounted with Vectashield and coverslip were sealed with nail polish. A full list of the antibodies used in this study is reported in Supplementary Data 3.
Samples were acquired with a fully motorized widefield upright microscope Zeiss AxioImager.Z2, equipped with a monochromatic camera Hamamatsu ORCA Fusion, sCMOS sensor, 2304 × 2304 pixels, 6.5 × 6.5 μm size. Z-stacks were set at 0.25 μm thickness and images were deconvolved with ZEN Blue Software using the “constrained iterative” algorithm set at maximum strength. Whole projections of deconvolved images were generated with Fiji and processed in Photoshop, where some false coloring was applied.
Auxin Treatment
Worms of the indicated genotype were selected as L4s or young adults (24 h post L4 stage) and picked onto NGM plates with and without 1 mM indole-3-acetic acid (auxin, Sigma). Exposure to auxin was carried out for the indicated times. Given that auxin inhibits bacterial growth, OP50 liquid cultures were concentrated 2X and spotted onto auxin plates a few days in advance and air-dried before placing the worms. Seeded auxin plates were kept refrigerated at 4 °C protected from light and used within three weeks. All degron strains were crossed into the CA1199 background37, which carries the TIR1::mRuby transgene expressed under sun-1 regulatory elements and integrated on chromosome IV.
Biochemistry
For whole cell extracts, 100 synchronized young adult worms (24 h post-L4 stage) were picked into 30μl of 1XTE (Tris-EDTA Buffer, pH=8) containing 1X cOmplete Protease Inhibitor (Roche) and flash-frozen in liquid nitrogen. After thawing, 10μl of 4X Laemmli Buffer was added, and worms were boiled for 10 minutes with mild shaking. Samples were spun down at maximum speed for 1 minute and run on precast 4-20% gradient acrylamide gels (BioRad) in 1X Tris-Glycine buffer containing 0.1% SDS. Proteins were transferred onto a nitrocellulose membrane (Amersham) in 1X Tris-Glycine buffer containing 20% methanol for 1 h at 4 °C. The Membrane was rinsed in 1XTBS containing 0.1% Tween (TBST) and blocked in 1XTBST containing 5% milk. Incubation of primary and secondary antibodies was carried out in the same buffer overnight at 4 °C and 2 h at room temperature respectively. For nuclear protein fractionation, extracts were produced as in32 without modifications and co-immunoprecipitation was performed by incubating 1-2 mg of nuclear extract with pre-equilibrated anti-HA Affinity Matrix (50μl, Sigma) or agarose GFP traps (30 μl, Chromotek) in Buffer D (20 mM HEPES pH 7.9, 150 mM KCl, 20% glycerol, 0.2 mM EDTA, 0.2% Triton X-100 and complete Roche inhibitor) overnight at 4 °C. The following day, beads were extensively washed in Buffer D and resuspended in 40μl of 2X Laemmli Buffer and boiled for 10 minutes. Samples were spun down for 1 minute at maximum speed to pellet the beads, and proteins were loaded onto precast 4–20% gradient acrylamide gels.
In-gel digestion and LC-MS/MS analysis
Selected 1D gel bands were excised manually and after de-staining and washing procedures each band was subjected to protein reduction (10 mM DTT in 25 mM NH4HCO3, 45 min, 56 °C, 750 rpm) and alkylation (55 mM IAA in 25 mM NH4HCO3; 30 min, laboratory temperature, 750 rpm) step. After further washing by 50% ACN/NH4HCO3 and pure ACN, the gel pieces were incubated with 125 ng trypsin (sequencing grade; Promega) in 50 mM NH4HCO3. The digestion was performed overnight at 37 °C on a Thermomixer (750 rpm; Eppendorf). Tryptic peptides were extracted into LC-MS vials by 2.5% formic acid (FA) in 50% ACN with the addition of polyethylene glycol (final concentration 0.001%)78 and concentrated in a SpeedVac concentrator (Thermo Fisher Scientific). LC-MS/MS analyses on HIM-17::3XHA pull downs were performed using RSLCnano system (Thermo Fisher Scientific) connected to Orbitrap Fusion Lumos Tribrid (Thermo Fisher Scientific). Prior to LC separation, tryptic digests were concentrated and desalted using trapping column (300 μm × 5 mm, μPrecolumn, 5μm particles, Acclaim PepMap 100 C18, Thermo Fisher Scientific). After washing of trapping column with 0.1% FA, the peptides were eluted in backflush mode (flow 500 nl/min) from the trapping column onto Acclaim PepMap100 C18 column (3 µm particles, 75 μm × 500 mm; Thermo Fisher Scientific) by 120 min gradient program (mobile phase A: 0.1% FA in water; mobile phase B: 0.1% FA in 80% ACN). Both columns were heated to 40 °C. The analysis of the mass spectrometric RAW data files was carried out using the MaxQuant software (version 1.6.10.43) using default settings unless otherwise noted. MS/MS ion searches were done against modified cRAP database (based on http://www.thegpm.org/crap, 112 protein sequences), and UniProtKB protein database for Caenorhabditis_elegans (27,028 protein sequences). Trypsin/P enzyme specificity with 2 allowed miss cleavages and minimal peptide length 6 amino acids were set. Only peptides and proteins with FDR threshold <0.01 and proteins having at least one unique or razor peptide were considered.
EdU staining
Synchronized young adult worms were hand-picked in 50μl of M9 containing 0.1% tween, to which 50μl of 1 mM EdU dissolved in DMSO were added before placing the samples at 20 °C protected from light. For the experiment shown in Supplementary Fig. 2b, worms were left soaking in M9 containing EdU for 40 minutes, after which they were allowed to recover for 1.5 hrs at 20 °C on seeded plates before dissections and staining. To detect EdU, worms were dissected in 1XPBST and fixed in PFA at 4% final concentration for 10 minutes at room temperature. Coverslips were freeze-cracked in liquid nitrogen and slides were placed in methanol at −20 °C for 10 minutes. After three washes in 1XPBST, EdU was detected with the Click-IT reaction kit (ThermoFisher) according to manufacturer protocol.
For the experiment in Fig. 3, worms were incubated in M9 containing EdU for 20 minutes and then placed either on non-auxin plates (untreated and recovery) or auxin plates for the indicated times. For EdU detection combined to immunofluorescence, samples were first processed as for normal immunostaining including incubation with secondary antibodies, after which, the Click-IT protocol was applied according to manufacturer instructions. We and others79 have noticed that the Click-IT protocol significantly lowers antibody detection efficiency, therefore after testing several markers for the chromosome axes, we chose to employ the anti-phosphoHTP-3S285 antibody that we previously generated80, which localizes identically as pan anti-HTP-3 but displayed better staining quality.
Quantification of synapsis, PLK-2, COSA-1 and DAPI bodies
To quantify synapsis, germlines of the indicated genotype were divided into six equal regions spanning the mitotic tip to diplotene entry. Nuclei were considered synapsed only if SYP-1 fully overlapped with HTP-3. At least three gonads for each genotype were used for quantifications. For PLK-2 quantification, nuclear cell rows from transition zone to late pachytene stage were counted and considered as “positive” only if at least 50% of the cells in a given row displayed PLK-2 staining in >1 aggregate, as previously shown61. At least three germlines were used for the quantification. For DAPI bodies counts, only -1 and -2 diakinesis nuclei were scored. For OLLAS::COSA-1 staining, only the nuclei in the last seven rows before the diplotene stage were included in the quantification31.
SUN-1::GFP tracking
For the analysis of SUN-1 aggregates in live animals, worms were anaesthetized with M9 buffer containing 10 mM Tetramisole, mounted on 2% agarose pads and covered with coverslips, which were sealed with melted petroleum jelly. SUN-1 aggregates marking chromosome end attachments at the nuclear periphery were filmed as previously described12. Before filming adult hermaphrodites, preselected HA::AID::bra-2; TIR1::mRuby L4s were incubated at 20°C for 24 hours on plates containing 1 mM auxin or ethanol plates, which served as a control. Images were acquired as 0.8 µm thick optical sections every 5 seconds for 5 minutes with a DeltaVision Ultra Epifluorescence Microscope, using a 60x silicone objective. The softWoRx suite was used for deconvolution. The images were analyzed using ImageJ (NIH) with StackReg and Manual Tracking plugins.
GFP::SYP-3 live imaging
Preselected L4s syp-3(ok758); GFP::syp-3; HA::AID::bra-2; TIR1::mRuby worms were placed either on an OP50 spread NGM plate or on an NGM plate containing auxin with 3 drops of OP50. The worms were allowed to grow for 24 hours to reach adulthood and begin oogenesis. Live worm mount slides were prepared by placing a 1 mm thick 10% agar pad on frosted slides and allowed to cure. At 24 hours post L4 a single one-day old adult was placed in a 0.75 µl drop of beads on the agar pad. A cover slip was placed over the pad, and the position of the worm marked on the back of the slide. Imaging was done with the DeltaVision deconvolution microscope. A 1024 × 1024 time-lapse 3D stacked image was taken of the full thickness of meiotic nuclei using FITC for 5 minutes. The region imaged was centered on the transition from partial accumulation (foci) of SYP-3::GFP and elongated SYP-3::GFP. After imaging, movies were deconvolved and evaluated for SYP-3::GFP foci initiating an elongation events.
Fluorescence in situ hybridization
FISH experiments to detect chromosomes III and V was performed as in32 without modifications. To label the pairing center end of chromosome III, the T17A3 cosmid was used to generate a digoxigenin-labeled DNA probe with the Digoxigenin-nick translation mix (Roche) according with manufacturer instructions. The chromosome V was visualized with a biotin-labeled probe generated by amplifying the endogenous 5 s rDNA locus by PCR, and labeling was performed with the Biotin-nick translation mix (Roche) according with manufacturer instructions. Labeled probes were added to FISH-mix (10% dextran sulfate, 50% formamide, 2XSSCT), and slides were incubated for 3´at 92 °C followed by overnight incubation at 37 °C. To combine FISH with SYP-1 staining, we first performed FISH protocol, and after probe incubation and post-hybridization washes, slides were washed thrice with 2XSSCT for 10 minutes each at room temperature and blocked in 1% BSA in 2XSSCT for 30 minutes. Incubation with primary chicken anti-SYP-1 antibody61 was carried out overnight at room temperature in a humid chamber, and the following day the slides were washed three times with 2XSCCT for 10 minutes each. Goat anti-chicken secondary antibody was combined with FITC-conjugated anti Biotin or Rhodamine-conjugated anti Digoxigenin and left to incubate for 3 h at room temperature in the dark. Slides were washed, and DAPI staining was performed as for regular immunostaining.
For the detection of non-PC of chromosome IV, we employed the same two Cy3-conjugated oligoprobes corresponding to “IV-3” in81 and performed FISH “in tube” with minor modifications. Briefly, worms were dissected on a 22×22 coverslip in 30μl of 1XEGG buffer containing 0.1% Tween and fixed by adding an equal volume of 8% PFA in 1XEGG buffer containing 0.1% Tween. Samples were fixed for 4 minutes at room temperature and then transferred to 1.5-mL Eppendorff tubes, in which 1 mL of cold methanol was added. Tubes were placed at −20 °C until the completion of dissections. Methanol was replaced with 1 mL of 2XSSC containing 0.1% Tween to perform three washes at room temperature. After the last wash, samples were transferred to 0.1 mL PCR tubes, and 100 μl of 50% formamide in 2XSSC containing 0.1% Tween was added to carry out a pre-hybridization step of the samples in a PCR thermocycler at 37 °C overnight. The following day, formamide was removed and 40μl of FISH mix containing 2μl of each oligoprobe from a 10 mM stock were added. Probe hybridization was carried out in a thermocycler at 92 °C for 2´30´´, 72 °C for 2´ and 37 °C overnight. The following day, FISH mix was removed, samples were washed three times in 2XSSCT for 5 minutes each at room temperature in the dark. DAPI was allowed to stain in the tubes for 2 minutes, and after two 10-minutes wash in 2XSCCT, the dissected germ lines were transferred to a Superfrost positively charged slide in minimal volume and mounted with 12μl of Vecatshield.
RNA-Seq
125mM K-NAA was dissolved in water and filtered and added to cooled MyoB media to a final concentraton of 4 mM. Plates were dried overnight and seeded with OP50 prior to use. 4×100 L4 worms were picked and aged for 2 days on either MyoB or MyoB + auxin. Worms were dissected in M9 buffer (for 1 L: 3 g KH2PO4, 6 g Na2HPO4, 5 g NaCl, 1 ml 1 M MgSO4) containing 200 nM levamisole and gonads were collected using a pulled micropipette to obtain the mitotic region through diplotene onset. Gonads were directly pipetted into 500μl of Trizol on ice. 50 gonads/ genotypes/treatment were collected and stored at -80 °C. RNA isolation and library was performed as in ref. 82. Samples were thawed on ice, and 100μl of 10 mM Tris was added to each tube. Samples were vortexed for 15 seconds and subjected to 3 freeze-thaw cycles with dry ice/37 °C water bath. 50μl of 1-Bromo-3-chloropropane (BCP) was added than transferred to Phase Lock Gel-Heavy 2 ml tubes (Brinkmann Instruments) and spun 15 min at 12,000 g at 4 °C. The aqueous phase was transferred to a new tube, 0.5μl of linear polyacrylamide (LPA) was added, and the sample was vortexed 10 sec and incubated at −80 °C for 2 hrs before spinning at 14,000 g for 30 min at 4 °C. The supernatant was aspirated and discarded, and pellet was washed in 1 ml ice-cold ethanol before centrifuging again for 10 min at 4 °C. The EtOH wash was repeated 2 more times. After the last wash and spin, the supernatant was removed with a P10 and the pellet air-dried with open cap for 3 minutes until clear. Pellet was dissolved in 15 μl RNase-free water, gently mixed, and stored at -80 °C. Concentrations were determined by Qubit and RNA quality was assessed by TapeStation. All samples obtained RIN scores of >8, confirming RNA quality. The NEB-Next Poly(A) mRNA magnetic isolation module and (NEB #E7490) Ultra II RNA library pre kit (NEB #E7770S) was used to prepare polyA+ RNA and cDNA libraries according to the manufacturer’s protocols. NEB-Next adapters were used for barcoding. Library quality was assessed by TapeStation and libraries were stored at -20 °C prior to sequencing. Libraries were pooled prior to sequencing on a NextSeq 2000 Sequences with P2 100 Flow Cell.
These high-quality reads, generated by NEBNext® Ultra™ II RNA Library Prep Kit, were trimmed of the universal TruSeq adapters using Cutadapt (v 2.10). The trimmed reads were mapped against the Ensembl C. elegans reference genome (WBCel235) using the STAR (v 2.7.9a) mapping tool. The output file from STAR was converted from SAM format to BAM format using SAMtools (v 1.14). Counts for expressed genes were generated using HT-Seq (v 0.13.5), and output was generated in text format. These count text files were then imported into the Bioconductor R package, edgeR (v 4.0.16). The edgeR package was then utilized to identify differentially expressed genes between comparison groups. Gene ontology analysis of differentially expressed genes was performed using Panther GO83.
Generation of RAD-51 antibody
The synthetic peptide SAQASRQKKSDQEQRA corresponding to aminoacids 40-55 of C. elegans RAD-51 isoform A was used to immunize four Wistar Rats according to standard procedures (Genscript). After four rounds of immunization, the raw serum from each animal was pooled and purified by protein G-sepharose. The specificity of the antibody was assessed by immunofluorescence in WT animals, as well as in rad-51(Ig8701) and spo-11(ok79) mutant worms, in which no RAD-51 foci were observed.
Statistical analyses
All statistical analyses were performed using the GraphPad Prism 7 software. Two-tailed Mann-Whitney or two-sided Chi-square tests were performed were appropriate. Sample sizes were chosen according to common standards applied in the C. elegans meiosis community, and all the experiments were repeated twice or more to ensure the reproducibility of the results.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed in this study are included in this published article (and its Supplementary information files). Source data are provided with this paper. The data set generated in the mass spectrometry analysis of HIM-17::3XHA pull downs is provided in the Source Data and has been deposited in the PRIDE public repository under the accession code PXD053401. The RNA sequencing data are provided in the Source Data and have been deposited in the GEO public repository with the accession code GSE275668. Source data are provided with this paper.
References
Zickler, D. & Kleckner, N. Meiotic chromosomes: integrating structure and function. Annu. Rev. Genet. 33, 603–754 (1999).
Zickler, D. & Kleckner, N. Meiosis: Dances between homologs. Annu. Rev. Genet. 57, 1–63 (2023).
Link, J. & Jantsch, V. Meiotic chromosomes in motion: a perspective from Mus musculus and Caenorhabditis elegans. Chromosoma 128, 317–330, https://doi.org/10.1007/s00412-019-00698-5 (2019).
Burke, B. LINC complexes as regulators of meiosis. Curr. Opin. Cell Biol. 52, 22–29 (2018).
Alleva, B. & Smolikove, S. Moving and stopping: Regulation of chromosome movement to promote meiotic chromosome pairing and synapsis. Nucleus 8, 613–624 (2017).
Kim, H. J., Liu, C. & Dernburg, A. F. How and why chromosomes interact with the cytoskeleton during meiosis. Genes 13, 901 (2022).
Sato, A. et al. Cytoskeletal forces span the nuclear envelope to coordinate meiotic chromosome pairing and synapsis. Cell 139, 907–919 (2009).
Koszul, R. & Kleckner, N. Dynamic chromosome movements during meiosis: a way to eliminate unwanted connections? Trends Cell Biol. 19, 716–724 (2009).
Rog, O. & Dernburg, A. F. Direct visualization reveals kinetics of meiotic chromosome synapsis. Cell Rep. 10, 1639–1645 (2015).
Wynne, D. J., Rog, O., Carlton, P. M. & Dernburg, A. F. Dynein-dependent processive chromosome motions promote homologous pairing in C. elegans meiosis. J. Cell Biol. 196, 47–64 (2012).
Penkner, A. M. et al. Meiotic chromosome homology search involves modifications of the nuclear envelope protein Matefin/SUN-1. Cell 139, 920–933 (2009).
Baudrimont, A. et al. Leptotene/Zygotene chromosome movement via the SUN/KASH protein bridge in caenorhabditis elegans. PLoS Genet 6, e1001219 (2010).
Phillips, C. M. et al. Identification of chromosome sequence motifs that mediate meiotic pairing and synapsis in C. elegans. Nat. Cell Biol. 11, 934–942 (2009).
Phillips, C. M. et al. HIM-8 binds to the X Chromosome pairing center and mediates chromosome-specific meiotic synapsis. Cell 123, 1051–1063 (2005).
Harper, N. C. et al. Pairing centers recruit a polo-like kinase to orchestrate meiotic chromosome dynamics in C. elegans. Dev. Cell 21, 934–947 (2011).
Phillips, C. M. & Dernburg, A. F. A family of zinc-finger proteins is required for chromosome-specific pairing and synapsis during meiosis in C. elegans. Dev. Cell 11, 817–829 (2006).
Kim, H. J., Liu, C., Zhang, L. & Dernburg, A. F. MJL-1 is a nuclear envelope protein required for homologous chromosome pairing and regulation of synapsis during meiosis in C. elegans. Sci. Adv. 9, eadd1453 (2023).
Penkner, A. et al. The nuclear envelope protein Matefin/SUN-1 is required for homologous pairing in C. elegans Meiosis. Dev. Cell 12, 873–885 (2007).
Liu, C. & Dernburg, A. F. Chemically induced proximity reveals a Piezo-dependent meiotic checkpoint at the oocyte nuclear envelope. Science. 386, eadm7969 (2024).
Labella, S., Woglar, A., Jantsch, V. & Zetka, M. Polo kinases establish links between meiotic chromosomes and cytoskeletal forces essential for homolog pairing. Dev. Cell 21, 948–958 (2011).
Woglar, A. et al. Matefin/SUN-1 phosphorylation is part of a surveillance mechanism to coordinate chromosome synapsis and recombination with meiotic progression and chromosome movement. PLoS Genet 9, e1003335 (2013).
Page, S. L. & Hawley, R. S. The genetics and molecular biology of the synaptonemal complex. Annu. Rev. Cell Dev. Biol. 20, 525–558 (2004).
Goodyer, W. et al. HTP-3 Links DSB formation with homolog pairing and crossing over during C. elegans Meiosis. Dev. Cell 14, 263–274 (2008).
Severson, A. F., Ling, L., van Zuylen, V. & Meyer, B. J. The axial element protein HTP-3 promotes cohesin loading and meiotic axis assembly in C. elegans to implement the meiotic program of chromosome segregation. Genes Dev. 23, 1763–1778 (2009).
Couteau, F., Nabeshima, K., Villeneuve, A. & Zetka, M. A component of C. elegans meiotic chromosome axes at the interface of homolog alignment, synapsis, nuclear reorganization, and recombination. Curr. Biol. 14, 585–592 (2004).
Martinez-Perez, E. HTP-1-dependent constraints coordinate homolog pairing and synapsis and promote chiasma formation during C. elegans meiosis. Genes Dev. 19, 2727–2743 (2005).
Couteau, F. HTP-1 coordinates synaptonemal complex assembly with homolog alignment during meiosis in C. elegans. Genes Dev. 19, 2744–2756 (2005).
Keeney, S., Giroux, C. N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997).
Dernburg, A. F. et al. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell 94, 387–398 (1998).
Lam, I. & Keeney, S. Mechanism and regulation of meiotic recombination initiation. Cold Spring Harb. Perspect. Biol. 7, a016634 (2014).
Janisiw, E. et al. Poly(ADP-ribose) glycohydrolase coordinates meiotic DNA double-strand break induction and repair independent of its catalytic activity. Nat. Commun. 11, 4869 (2020).
Trivedi, S., Blazícková, J. & Silva, N. PARG and BRCA1–BARD1 cooperative function regulates DNA repair pathway choice during gametogenesis. Nucleic Acids Res. 50, 12291–12308 (2022).
Guo, R. et al. BS69/ZMYND11 reads and connects Histone H3.3 Lysine 36 Trimethylation-decorated chromatin to regulated Pre-mRNA processing. Mol. Cell 56, 298–310 (2014).
Yates, T. M. et al. ZMYND11 ‐related syndromic intellectual disability: 16 patients delineating and expanding the phenotypic spectrum. Hum. Mutat. 41, 1042–1050 (2020).
Yokoo, R. et al. COSA-1 reveals robust homeostasis and separable licensing and reinforcement steps governing meiotic crossovers. Cell 149, 75–87 (2012).
Morita, K., Shimizu, M., Shibuya, H. & Ueno, N. A DAF-1-binding protein BRA-1 is a negative regulator of DAF-7 TGF- signaling. Proc. Natl Acad. Sci. 98, 6284–6288 (2001).
Zhang, L., Ward, J.D., Cheng, Z. & Dernburg, A.F. (2015). The auxin-inducible degradation (AID) system enables versatile conditional protein depletion in C. elegans. Development, dev.129635. https://doi.org/10.1242/dev.129635.
Wagner, C. R., Kuervers, L., Baillie, D. L. & Yanowitz, J. L. xnd-1 regulates the global recombination landscape in Caenorhabditis elegans. Nature 467, 839–843 (2010).
Kelly, W. G. et al. X-chromosome silencing in the germline of C. elegans. Development 129, 479–492 (2002).
Larson, B. J., Van, M. V., Nakayama, T. & Engebrecht, J. Plasticity in the meiotic epigenetic landscape of sex chromosomes in caenorhabditis species. Genetics 203, 1641–1658 (2016).
Reddy, K. C. & Villeneuve, A. M. C. elegans HIM-17 links chromatin modification and competence for initiation of meiotic recombination. Cell 118, 439–452 (2004).
Colaiácovo, M. P. et al. Synaptonemal complex assembly in C. elegans is dispensable for loading strand-exchange proteins but critical for proper completion of recombination. Dev. Cell 5, 463–474 (2003).
Kelly, K. O., Dernburg, A. F., Stanfield, G. M. & Villeneuve, A. M. Caenorhabditis elegans msh-5 is required for both normal and radiation-induced meiotic crossing over but not for completion of meiosis. Genetics 156, 617–630 (2000).
MacQueen, A. J. Synapsis-dependent and -independent mechanisms stabilize homolog pairing during meiotic prophase in C. elegans. Genes Dev. 16, 2428–2442 (2002).
MacQueen, A. J. Nuclear reorganization and homologous chromosome pairing during meiotic prophase require C. elegans chk-2. Genes Dev. 15, 1674–1687 (2001).
Crawley, O. et al. Cohesin-interacting protein WAPL-1 regulates meiotic chromosome structure and cohesion by antagonizing specific cohesin complexes. eLife 5, e10851 (2016).
Brodigan, T. M., Liu, J. I., Park, M., Kipreos, E. T. & Krause, M. Cyclin E expression during development in caenorhabditis elegans. Dev. Biol. 254, 102–115 (2003).
Alleva, B., Clausen, S., Koury, E., Hefel, A. & Smolikove, S. CRL4 regulates recombination and synaptonemal complex aggregation in the Caenorhabditis elegans germline. PLoS Genet 15, e1008486 (2019).
Hayashi, M., Mlynarczyk-Evans, S. & Villeneuve, A. M. The synaptonemal complex shapes the crossover landscape through cooperative assembly, crossover promotion and crossover inhibition during Caenorhabditis elegans Meiosis. Genetics 186, 45–58 (2010).
Libuda, D. E., Uzawa, S., Meyer, B. J. & Villeneuve, A. M. Meiotic chromosome structures constrain and respond to designation of crossover sites. Nature 502, 703–706 (2013).
Cahoon, C. K., Richter, C. M., Dayton, A. E. & Libuda, D. E. Sexual dimorphic regulation of recombination by the synaptonemal complex in C. elegans. eLife 12, e84538 (2023).
Carelli, F. N. et al. Widespread transposon co-option in the Caenorhabditis germline regulatory network. Sci. Adv. 8, eabo4082 (2022).
Jaramillo-Lambert, A., Ellefson, M., Villeneuve, A. M. & Engebrecht, J. Differential timing of S phases, X chromosome replication, and meiotic prophase in the C. elegans germ line. Dev. Biol. 308, 206–221 (2007).
Tolkin, T. & Hubbard, E. J. A. Germline stem and progenitor cell aging in C. elegans. Front. Cell Dev. Biol. 9, 699671 (2021).
Hillers, K.J. (2017). Meiosis. WormBook, 1–43. https://doi.org/10.1895/wormbook.1.178.1.
Hicks, T. et al. Continuous double-strand break induction and their differential processing sustain chiasma formation during Caenorhabditis elegans meiosis. Cell Rep. 40, 111403 (2022).
Meneely, P. M., McGovern, O. L., Heinis, F. I. & Yanowitz, J. L. Crossover distribution and frequency are regulated by him-5 in Caenorhabditis elegans. Genetics 190, 1251–1266 (2012).
Rinaldo, C., Bazzicalupo, P., Ederle, S., Hilliard, M. & La Volpe, A. Roles for Caenorhabditis elegans rad-51 in meiosis and in resistance to ionizing radiation during development. Genetics 160, 471–479 (2002).
Alpi, A., Pasierbek, P., Gartner, A. & Loidl, J. Genetic and cytological characterization of the recombination protein RAD-51 in Caenorhabditis elegans. Chromosoma 112, 6–16 (2003).
Zhang, L. et al. Recruitment of Polo-like kinase couples synapsis to meiotic progression via inactivation of CHK-2. Elife 12, e84492 (2023).
Silva, N. et al. The fidelity of synaptonemal complex assembly is regulated by a signaling mechanism that controls early meiotic progression. Dev. Cell 31, 503–511 (2014).
Kim, Y., Kostow, N. & Dernburg, A. F. The chromosome axis mediates feedback control of CHK-2 to ensure crossover formation in C. elegans. Dev. Cell 35, 247–261 (2015).
Castellano-Pozo, M. et al. Surveillance of cohesin-supported chromosome structure controls meiotic progression. Nat. Commun. 11, 4345 (2020).
Barroso, C. et al. (2024). Two structurally mobile regions control the conformation and function of metamorphic meiotic HORMAD proteins. Preprint, https://doi.org/10.1101/2024.08.05.606648.
Nadarajan, S., Altendorfer, E., Saito, T. T., Martinez-Garcia, M. & Colaiácovo, M. P. HIM-17 regulates the position of recombination events and GSP-1/2 localization to establish short arm identity on bivalents in meiosis. Proc. Natl Acad. Sci. USA 118, e2016363118 (2021).
Rosu, S. et al. The C. elegans DSB-2 protein reveals a regulatory network that controls competence for meiotic DSB formation and promotes crossover assurance. PLoS Genet 9, e1003674 (2013).
Hurlock, M. E. et al. Identification of novel synaptonemal complex components in C. elegans. J. Cell Biol. 219, e201910043 (2020).
Zhang, Z. et al. Multivalent weak interactions between assembly units drive synaptonemal complex formation. J. Cell Biol. 219, e201910086 (2020).
Smolikov, S., Schild-Prüfert, K. & Colaiácovo, M. P. A yeast two-hybrid screen for SYP-3 interactors identifies SYP-4, a component required for synaptonemal complex assembly and Chiasma Formation in Caenorhabditis elegans Meiosis. PLoS Genet 5, e1000669 (2009).
Smolikov, S. et al. SYP-3 restricts synaptonemal complex assembly to bridge paired chromosome axes during meiosis in Caenorhabditis elegans. Genetics 176, 2015–2025 (2007).
Blundon, J. M. et al. Skp1 proteins are structural components of the synaptonemal complex in C. elegans. Sci. Adv. 10, eadl4876 (2024).
Jantsch, V. et al. Caenorhabditis elegans prom-1 is required for meiotic prophase progression and homologous chromosome pairing. MBoC 18, 4911–4920 (2007).
Tang, L. et al. Mutations in Caenorhabditis elegans him-19 show meiotic defects that worsen with age. MBoC 21, 885–896 (2010).
Kleckner, N. et al. A mechanical basis for chromosome function. Proc. Natl Acad. Sci. USA. 101, 12592–12597 (2004).
Börner, G. V., Kleckner, N. & Hunter, N. Crossover/Noncrossover differentiation, synaptonemal complex formation, and regulatory surveillance at the Leptotene/Zygotene transition of meiosis. Cell 117, 29–45 (2004).
Zickler, D. & Kleckner, N. The Leptotene-Zygotene transition of meiosis. Annu. Rev. Genet. 32, 619–697 (1998).
Brenner, S. The genetics of Caenorhabditis elegans. Genetics 77, 71–94 (1974).
Stejskal, K., Potěšil, D. & Zdráhal, Z. Suppression of Peptide sample losses in autosampler vials. J. Proteome Res. 12, 3057–3062 (2013).
Almanzar, D. E., Hamrick, A. & Rog, O. Single-sister labeling in the C. elegans germline using the nucleotide analog EdU. STAR Protoc. 3, 101344 (2022).
Das, D., Trivedi, S., Blazícková, J., Arur, S. & Silva, N. (2022). Phosphorylation of HORMA-domain protein HTP-3 at Serine 285 is dispensable for crossover formation. G3 Genes|Genomes|Genetics, jkac079. https://doi.org/10.1093/g3journal/jkac079.
Adilardi, R. S. & Dernburg, A. F. Robust, versatile DNA FISH probes for chromosome-specific repeats in Caenorhabditis elegans and Pristionchus pacificus. G3 Genes|Genomes|Genet. 12, jkac121 (2022).
Cockrum, C. S. & Strome, S. Maternal H3K36 and H3K27 HMTs protect germline development via regulation of the transcription factor LIN-15B. eLife 11, e77951 (2022).
Thomas, P. D. et al. PANTHER: Making genome‐scale phylogenetics accessible to all. Protein Sci. 31, 8–22 (2022).
Acknowledgements
We are grateful to E. Martinez-Perez, S. Arur, A. Villeneuve, W. Kelly, Y. Kim, E. Kipreos, M. Colaiacovo and A. Dernburg for valuable strains and reagents. We thank Chantal Wicky for sharing unpublished data, Anne Villeneuve for critical reading of the manuscript, and Marilina Raices for assistance in gonad dissections for RNA-Seq studies. We acknowledge the core facility CELLIM supported by the Czech-BioImaging large RI project (LM2023050 funded by MEYS CR) for their support with obtaining scientific data presented in this paper. CIISB, Instruct-CZ Centre of Instruct-ERIC EU consortium, funded by MEYS CR infrastructure project LM2023042, is gratefully acknowledged for the financial support of the measurements at the CEITEC Proteomics Core Facility. Computational resources were provided by the e-INFRA CZ project (ID:90254), supported by MEYS CR. In vivo filming of chromosome-nuclear envelope end attachments was performed by the BioOptics Facility at Max Perutz Labs using the VBCF instrument pool. RNAseq analysis was supported by the Genomics Analysis Core, a shared resource funded by the University of Pittsburgh School of Medicine. Some strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). Research in the Silva lab is funded by the Czech Science Foundation (GA23-04918S), the Smolikove lab is funded by NSF (2027955), the Jantsch lab is funded by the FWF (SFB F 8805-B), the Zetka lab by the Canadian Institutes of Health Research (PJT-173381) and the Yanowitz lab by NIH grant R01GM104007.
Author information
Authors and Affiliations
Contributions
N.S. acquired funding, designed the research and performed most of the experiments with the technical support of J.B. and S.T.; R.B. performed in vivo imaging of GFP::SYP-3; S.S.G. performed in vivo analysis of SUN-1::GFP; S.S. generated multiple strains; M.S., A.C., U.R.C., J.L.Y. performed the RNAseq and analyzed the data; N.S., J.L.Y., S.Sm., V.J. and M.Z. provided supervision; N.S. wrote the manuscript, with editing inputs from V.J., M.Z., S.Sm. and J.L.Y.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Hiroki Shibuya, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Blazickova, J., Trivedi, S., Bowman, R. et al. Overlapping and separable activities of BRA-2 and HIM-17 promote occurrence and regulation of pairing and synapsis during Caenorhabditis elegans meiosis. Nat Commun 16, 2516 (2025). https://doi.org/10.1038/s41467-025-57862-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-57862-y
This article is cited by
-
A nuclear TRiC/CCT chaperonin assembles meiotic HORMAD proteins into chromosome axes competent for crossing over
Nature Communications (2025)