Abstract
Trained immunity refers to the long-term memory of the innate immune cells. However, little is known about how environmental nutrient availability influences trained immunity. This study finds that physiologic carbon sources impact glucose contribution to the tricarboxylic acid (TCA) cycle and enhance cytokine production of trained monocytes. Our experiments demonstrate that trained monocytes preferentially employe lactate over glucose as a TCA cycle substrate, and lactate metabolism is required for trained immune cell responses to bacterial and fungal infection. Except for the contribution to the TCA cycle, endogenous lactate or exogenous lactate also supports trained immunity by regulating histone lactylation. Further transcriptome analysis, ATAC-seq, and CUT&Tag-seq demonstrate that lactate enhance chromatin accessibility in a manner dependent histone lactylation. Inhibiting lactate-dependent metabolism by silencing lactate dehydrogenase A (LDHA) impairs both lactate fueled the TCA cycle and histone lactylation. These findings suggest that lactate is the hub of immunometabolic and epigenetic programs in trained immunity.
Similar content being viewed by others
Introduction
Reprogramming cellular metabolic processes has recently been proven essential for immune cells1,2,3. A widely accepted concept is that cell-intrinsic factors, such as critical molecules or metabolic enzymes, play a crucial role in the metabolic pathway in immune cells4. However, the availability of nutrients in the environment also significantly impacts metabolic utilization in immune cells5. For example, yeast substrate concentrations impacted metabolic output twice more than enzyme expression6. Cancer cells show changes in cell growth and stress resistance if cancer cells are cultured with a physiologic medium due to differences in substrate utilization7. Among nutrients in the environment, glucose is a significant nutrient that fuels cellular metabolic activities. After uptake through the glucose transporter, glucose flows through three distinctly metabolic pathways: glycolysis, the pentose phosphate pathway (PPP), and the hexosamine biosynthesis pathway8,9. Glucose is metabolized via glycolysis into pyruvate, which enters the TCA cycle when oxygen is available10. When oxygen is unavailable, pyruvate shifts to glycolysis, in which pyruvate converts to lactate10. Traditionally, lactate has been seen as a garbage of glucose metabolism11. However, a growing body of research shows that lactate is the primary circulating metabolite that fuels the TCA cycle and a significant substrate for gluconeogenesis12,13,14. A recent study identified a new epigenetic modification, histone lactylation, that regulates histones by adding lactyl groups to their lysine residues15. Histone lactylation has different temporal dynamics from acetylation and induces corresponding gene expression15. Subsequently, more than 30 histone lactylation sites have been identified in fungal, plant, mouse, and parasitic (protozoic) samples16.
Conventional wisdom says that the immune system is divided into the innate and the adaptive immune system. One of the main bases for this division is that the adaptive immune system is relatively slower and can build specific immunological memory17. By contrast, the innate immune system is rapid and non-specific and cannot build immunological memory17. However, recent studies have shown that innate immune cells, such as natural killer cells and monocytes, can make a non-specific memory that challenges this paradigm18,19. Specifically, the functional adaptation of innate immunity after a primary insult, including Bacille Calmette-Guerin vaccination or microorganism infection, represents a de facto innate immune memory20. Once these innate immune cells encounter a secondary non-specific (heterologous) stimulus, their response is altered, which results in a more robust response21. The induction of a non-specific memory results in a stepped-up function of the innate immune cells termed trained immunity22. The biological relevance of innate immune memory, such as cephalopods, plants, insects, and mammals, is widespread23. Analysis of differentially regulated pathways in trained immunity showed that mTOR-HIF1α regulated oxidative phosphorylation to glycolysis, the central regulatory mechanism for β-glucan, a fungal cell wall component, induced trained immunity24. Nevertheless, how environmental nutrient availability impacts trained immunity metabolism and function remains poorly understood.
In this study, we found that physiologic carbon sources (PCSs) influence the glucose utilization of trained monocytes. Our experiments have demonstrated that lactate is a physiological fuel for trained immunity, is preferred over glucose, and regulates the cytokine production of trained immune cells. Mechanistically, some intrinsic lactate function is a significant substrate for acetyl-CoA synthesis and fuels the TCA cycle. The other inherent lactate moves into the cell nucleus and regulates cytokine production through the effects of histone lactylation. Our data identifies the dominant metabolite (lactate) in trained immunity, contributes to our understanding of innate immune memory, and opens new therapeutic avenues.
Results
PCSs impact glucose utilization in monocytes during the induction of trained immunity
Traditional cell culture mediums, such as the Roswell Park Memorial Institute (RPMI) 1640 medium, are designed to promote cell growth and survival ex vivo25. Thus, the concentration of some nutrients in traditional cell culture medium is supraphysiological (Supplementary Fig. 1a). In addition, the conventional medium also lacks most standard culture media, such as lactate, citrate, acetate, β-hydroxybutyrate (βOHB), and pyruvate25. In this study, we referred to those metabolites as PCSs (Supplementary Fig. 1a). Previous studies show that these carbon sources are oxidizable substrates in vivo26,27. Notably, in some situations, these carbon sources exhibit circulatory turnover fluxes exceeding traditional sources, such as glucose and glutamine26,27. Thus, we designed the modified RPMI medium (MM) with approximate polar metabolite concentrations in mouse serum (Supplementary Fig. 1a).
To obtain trained immune cells, we treated purified monocytes with β-glucan for 24 hrs, followed by washing and resting for five days in a culture medium. Then, cells were restimulated with β-glucan or LPS for 24 hrs as previously described24 (Supplementary Fig. 1b). 13C-glucose labeling into lactate and intracellular TCA cycle intermediates, including citrate and fumarate, within 0-8 hrs in β-glucan treated monocytes (Supplementary Figs. 1c). Analysis of metabolite levels in culture medium over time revealed several trends (Supplementary Figs. 1d-1f). As expected, glucose and glutamine were consumed within 12-24 hrs during the trained immunity process, and PCS treatment mildly affected glucose and glutamine consumption in β-glucan treated monocytes (Supplementary Figs. 1d and 1e). A slower decrease of other carbon sources, including OHB, citrate, and pyruvate, was observed in the PCS-supplemented medium compared to the glucose-supplemented medium (Supplementary Fig. 1f). Interestingly, we observed accumulation of several TCA cycle intermediates (αKG and fumarate) in extracellular medium over time, indicated the export of these metabolites (Supplementary Fig. 1f).
Using 13C-based metabolic tracers, we investigate the role of PCSs on glucose metabolism in trained immunity (Fig. 1a). Mass spectrometry (MS) assay shows that 13C-glucose extensively labels intermediates of glycolysis and the TCA cycle in β-glucan treated monocytes (Figs. 1b-1d). By contrast, in the presence of PCSs at a physiologic concentration in β-glucan treated monocytes, 13C-glucose contribution into TCA cycle intermediates was significantly inhibited (Figs. 1b-1d). Intriguingly, the presence of PCSs at physiologic concentration directs glucose flux to the PPP and the serine synthesis pathway (SSP) (Figs. 1e and 1d). Similar results were obtained in metabolites arising from TCA cataplerosis, such as glutamate and aspartate (Fig. 1g). We observe similar 13C labeling patterns in β-glucan treated monocytes cultured PCSs and 13C-glucose for long (6 days) (Supplementary Fig. 1g). To investigate the contribution of PCSs to TCA cycle metabolism, β-glucan treated monocytes were cultured in MM supplemented with separated 13C-labeled PCSs at physiologic concentrations. We observe that 13C-glucose and 13C-glutamine contributed carbon to the TCA cycle intermediates (Supplementary Fig. 1h). Intriguingly, TCA cycle intermediates were also labeled from some PCSs, including βOHB and lactate, in β-glucan treated monocytes (Supplementary Fig. 1h). Results showed that βOHB and lactate contribute carbon to the TCA cycle by 2-carbon entry via citrate synthase (M + 2 citrate) and carry through the entire cycle in β-glucan treated monocytes (Fig. 1h). Of note, when considering the lower concentration of lactate (1 mM) compared with the concentration of glucose (5 mM), trained monocytes prefer to use PCSs over glucose for TCA cycle metabolism (Fig. 1h). Because it has been reported that alanine did not contribute to the TCA cycle28, the presence of 13C-alanine was included as a negative control for comparison (Supplementary Fig. 1h). Taken together, these findings suggest that PCSs reduced the contribution of 13C-glucose to the TCA cycle, and drives 13C-glucose switch from glycolysis to PPP and SSP, in trained immunity.
a Schematic of 13C-glucose carbon labeling through the TCA cycle, the pentose phosphate pathway (PPP), and the serine synthesis pathway (SSP). b–g Monocytes were cultured in MM containing 13C-glucose (±PCSs) and treated with β-glucan (5 μg/ml) for 4 hrs, followed by mass spectrometry analyzes. h Monocytes were cultured in MM containing the indicated 13C-labeled carbon source and treated with β-glucan (5 μg/ml) for 4 hrs. The relative contributions of 13C from indicated 13C-labeled substrates into metabolites of central carbon metabolism are shown. Data in (b-h) are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. Medium: Phosphate buffered saline. See also Supplementary Fig. 1.
Lactate activates trained immunity
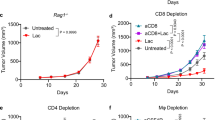
Because among these above PCSs, lactate has the most significant regulation on the TCA cycle (Fig. 1h), we next investigated the role of lactate on trained immunity in response to C. albicans infection (Fig. 2a). In line with a previous study29, C. albicans infection enhances the production of lactate in a time- and dose-dependent manner (Supplementary Fig. 2a). Since lactate dehydrogenase (Ldh) interconverts lactate and pyruvate by using NAD+/NADH as a co-factor30, we asked whether LDHA plays a role in trained immunity. Firstly, we employed the tamoxifen-inducible LDHA knockout mice (LDHA-/-), and LDHA protein expression in diverse organs of LDHA-/- mice was abrogated upon tamoxifen treatment (Supplementary Figs. 2b and 2c). To investigate the role of lactate on trained monocyte function, we trained mice with β-glucan followed by a challenge with C. albicans (Fig. 2a). As expected, IL-6 and TNF-α production was induced in β-glucan-trained Wt mice, but not in LDHA-/- mice (Fig. 2b). Histological analysis of the lung tissue showed that prior β-glucan treatment limits lung damage during subsequent infection (Fig. 2c). Compared with β-glucan-trained Wt mice, more severe lung injury was demonstrated in LDHA-/- mice (Fig. 2c). In this vein, LDHA-/- mice significantly susceptible to C. albicans-induced lethality, accompanied by higher bacterial burden in the lung and kidney tissue (Figs. 2d and 2e). A previous study reported that trained immunity is a protective host against secondary C. albicans infection that relies on monocytes induced by the same pathogen31. Similar results were obtained in C. albicans-trained Wt and LDHA-/- mice (Supplementary Figs. 2d-2g). To determine whether LDHA regulates trained immunity in vitro, we employed two specific small interfering RNAs (siRNAs) for LDHA. We confirmed siRNA efficiency (Supplementary Fig. 2h). SiRNA-LDHA #2 was selected for the following experiments. As shown in Supplementary Fig. 2i, LDHA knockdown inhibits β-glucan-induced expression of IL-6 and TNF-α in human peripheral blood mononuclear cells (PBMCs).
a Schematic representation of in vivo trained immunity experimental setup. b–d Wt and Ldha-/- mice were trained with or without β-glucan. Six days later, mice were infected with or without a secondary C. albicans lethal dose (3 × 106) for 24 hrs, followed by measuring IL-6 and TNF-α levels (b), lung injury(c), and kidney and lung fungal burden (d). e Wt and Ldha-/- mice were trained with or without β-glucan. Six days later, mice were infected with or without a secondary C. albicans lethal dose (3 × 106) for indicated times. Survival curves show data collected until day ten after infection. Statistical analysis was performed using the log-rank test (n = 8 for each group). f–h C57BL/6 mice were trained with or without β-glucan. Six days later, mice were secondarily infected without or with C. albicans (3 × 106) and infused with or without lactate (1 mM) for 24 hrs, followed by measuring IL-6 and TNF-α levels (f), kidney fungal burden (g), and lactate levels in serum (h). i C57BL/6 mice were trained with or without β-glucan. Six days later, mice were infected with or without a secondary C. albicans lethal dose (3 × 106) and infused with or without lactate (1 mM) for indicated times. Survival curves show data collected until day 12 after infection. Statistical analysis was performed using the log-rank test (n = 8 for each group). Data in (b–d, and f–h) are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Figs. 2.
Next, we investigated the role of lactate on trained immunity in response to C. albicans infection. As shown in Supplementary Fig. 2j, lactate enhances β-glucan-induced expression of IL-6 and TNF-α in a dose-dependent manner in human PBMCs. Of note, to rule out the possibility that lactate affects trained immunity by changing the acidity13, the pH of the medium was adjusted to an equal level in all in vitro experiments of this manuscript. The presence of lactate increased C. albicans-induced levels of IL-6 and TNF-α, accompanied by lower fungal burden in the kidney and lung tissue (Figs. 2f-2h). Strikingly, lactate increased the protective response of trained monocytes against lethal systemic C. albicans infection (Fig. 2i). These findings suggest that lactate is indispensable for the protective reaction of trained immunity against C. albicans infection.
Lactate is a physiologic fuel during the induction of trained immunity
Next, we investigate the mechanisms by which lactate regulates trained immunity. Firstly, we measured the timeline of 13C-glucose incorporation during β-glucan treatment. β-glucan treated monocytes displayed higher 13C-glucose incorporation into glucose-6-phosphate (G6P), fructose-6-phosphate (F6P), 3-phosphoglycerate (3PG), pyruvate, lactate, citrate, succinate, malate, glutamate and aspartate within 0-4 hrs (Supplementary Figs. 3a-3f), indicates increased glycolytic flow to the TCA cycle. 13C-glucose incorporation flow from oxidative phosphorylation (OxPhos) to SSP and PPP, as well as ATP synthesis and S-adenosylmethionine (SAM) generation, after 4 hrs of β-glucan treatment (Supplementary Figs. 3a-3g). However, 13C-glucose incorporation into glycolysis continued to increase throughout the experiment (Supplementary Fig. 3b). In addition, β-glucan also induced hexokinase 2 (HK2) enzymes and LDHA enzyme activity in a time- and dose-dependent manner (Supplementary Figs. 3h and 3i). Those data suggest β-glucan treated monocytes prefer to use glucose as physiologic fuel early. Still, β-glucan treated monocytes prefer to use glucose for biosynthesis later.
To evaluate the competition between glucose and lactate for TCA cycle metabolism, treated monocytes were cultured with 13C-glucose and unlabeled lactate with gradually increased concentration. Even in the presence of physiologic glucose concentrations (5 mM), β-glucan treated monocytes use low concentrations of lactate (1-2 mM) as a carbon source for intermediates in the TCA cycle (Figs. 3a-3d). By contrast, a low concentration of lactate (1 mM) directs glucose (5 mM) flux from OxPhos to SSP and PPP (Figs. 3a-3f). A substantial fraction of glucose-derived labeled SAM, including m + 5 (via the PPP) and m + 6-9 (via both the PPP and SSP), significantly increased in lactate-treated monocytes (Figs. 3g and 3h), indicates glucose is used for biosynthesis-in the presence lactate. Because lactate contributes to TCA cycle metabolism, we next investigated the role of lactate on trained monocytes bioenergetics. By using Forster resonance energy transfer (FRET)-based ATP sensors targeted to mitochondria (Mito, mtAT1.03), we found that the presence of lactate or β-glucan treatment enhances ATP production (Fig. 3i). The combination of β-glucan and lactate treatment resulted in a significant synergistic induction of ATP production (Fig. 3i). Similarly, the presence of lactate enhances their basal oxygen consumption rate (OCR) and ATP production from OxPhos (Supplementary Fig. 3j). Consistent with 13C metabolite data, lactate increases β-glucan-induced levels of acetyl-CoA, citrate, and malate (Fig. 3j). Finally, we investigate the role of glucose on the lactate contributed TCA cycle metabolism during β-glucan treatment. As shown in Fig. 3k, a low lactate concentration (1 mM) still contributes to TCA cycle intermediates in glucose (1-10 mM) contained culture medium during β-glucan treatment. When the glucose concentration is 25-fold higher than lactate levels, 13C-lactate could not contribute carbon to the TCA cycle (Fig. 3k). These findings suggest lactate is a TCA cycle substrate for trained monocytes under physiologic conditions, even when glucose is available.
a–f Monocytes were cultured in MM containing 13C-glucose (5 mM), indicating unlabeled lactate concentration, and treated with β-glucan (5 μg/ml) for 4 hrs, followed by mass spectrometry analyzes. g Schematic of derivation and contribution of carbon atoms in SAM synthesis. h Experiments were performed as described in (a). 13C-glucose labeling into ATP or SAM is shown. i Monocytes were treated with β-glucan and lactate (1 mM) for 30 min; mitochondrial ATP was measured by the mtAT 1.03 probe. j Monocytes were treated with or without β-glucan and lactate (1 mM) for 12 hrs, then acetyl-CoA, citrate, and malate levels were measured. k Monocytes were cultured in MM containing 13C-lactate (1 mM), indicating unlabeled glucose concentration, and treated with β-glucan (5 μg/ml) for 4 hrs, followed by mass spectrometry analyzes. Data in (a–f and h–k) are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Figs. 3.
Lactate fuels the trained immunity via LDHA
Next, we investigate the role of LDHA on the bioenergetic function of lactate in trained immunity. As shown in Figs. 4a and 4b, decreased abundance of 13C-lactate-derived citrate, succinate, malate, glutamate, and aspartate was observed in LDHA-/- monocytes in response to β-glucan treatment. 13C-lactate also contributed to acetyl-CoA-related metabolites, including Ac-carnitine and Ac-alanine, in β-glucan treated Wt monocytes, but not in β-glucan treated LDHA-/- monocytes (Fig. 4c). By using the mitochondrial ATP probe mtAT, β-glucan treatment enhances and ATP production in Wt monocytes, but not in β-glucan treated LDHA-/- monocytes (Fig. 4d). Consistently, LDHA knockout inhibited β-glucan-induced levels of acetyl-CoA, citrate, malate (Fig. 4e). Sodium oxamate (Oxa) is widely used to inhibit lactate production by regulating the activities of LDHA32. Consistently, β-glucan-induced levels of acetyl-CoA, citrate, and malate were inhibited in Oxa-treated trained monocytes (Supplementary Fig. 4a). We next explored whether the host used lactate as a fuel in trained immunity in vivo by infusing 13C metabolite. In our in vivo system, ~80% enrichment of fully labeled (M + 3) 13C-lactate in serum and lung (Figs. 4f and 4g). Consistent with ex vivo data, infused 13C-lactate readily labeled TCA cycle-derived metabolites in vivo in response to β-glucan treated Wt mice, but not in LDHA-/- mice (Figs. 4h and 4i). We further investigated β-glucan-regulated glucose metabolism in lactate metabolism inhibited system. Intriguingly, β-glucan treatment directs glucose flux from PPP and SSP back to OxPhos when lactate metabolism is inhibited (Supplementary Figs. 4b-4f), suggesting glucose is an alternative carbon source for the TCA cycle in trained immunity. These findings indicate that lactate processing via LDHA is a critical metabolic node in trained immunity.
a–c Monocytes from Wt and Ldha-/- mice were cultured in MM containing 13C-lactate (1 mM) and treated with β-glucan (5 μg/ml) for 12 hrs, followed by mass spectrometry analyzes. d Monocytes from Wt and Ldha-/- mice were cultured in lactate media (0.5 mM glucose, 0.5 mM glutamine, and 2 mM lactate), and treated with β-glucan for 30 min, mitochondrial ATP was measured by the mtAT 1.03 probe. e Monocytes from Wt and Ldha-/- mice were treated with or without β-glucan and lactate (1 mM) for 12 h, followed by measuring acetyl-CoA, citrate, and malate levels. f, g C57BL/6 mice were trained with or without β-glucan. Six days later, mice were infused with 13C-lactate for 12 hrs, followed by mass spectrometry analyzes. h, i Wt and Ldha-/- mice were trained with or without β-glucan. Six days later, mice were infused with 13C-lactate for 12 hrs; monocytes were isolated and followed by mass spectrometry analyzes. All data are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Fig. 4.
Lactate regulates histone lactylation via LDHA in monocytes during the induction of trained immunity
A previous study showed that lysine lactylation (Kla) is a new type of histone mark that can be stimulated by lactate15 (Supplementary Fig. 5a); we suspected that β-glucan regulated high level of lactate impact on histone lactylation in trained immunity. Immunoblots analysis revealed that histone Kla levels were increased; however, histone acetylation (Kac) levels were mildly increased, in a time-dependent fashion in β-glucan treated monocytes (Fig. 5a and Supplementary Fig. 5b). Histone Kla levels elevation was detected as early as 6 hrs after β-glucan treatment (Fig. 5a and Supplementary Fig. 5b). Intriguingly, high histone Kla levels were maintained from days 1 to 9 (Fig. 5b and Supplementary Fig. 5c). As expected, β-glucan challenge induce histone Kla levels, mildly induce Kac levels, in dose-dependent manner (Fig. 5c and Supplementary Fig. 5d). Given that replenishing cells with fresh media did not affect Kla levels, the increase of histone Kla during trained immunity is intrinsic and not due to paracrine effects (Supplementary Fig. 5e). Consistent with previous results15, MS/MS analysis showed that a mass shift of 72.021 Da on lysine residues in the histone was observed in β-glucan treated monocytes (Supplementary Figs. 5f and 5g). In this vein, metabolic labeling experiments using 13C-glucose or 13C-lactate followed by MS/MS analysis demonstrated that β-glucan promotes histone lactylation directly from glucose or exogenous lactate (Figs. 5d-5g). In addition, β-glucan treatment increased histone Kla levels in Wt monocytes but not in LDHA-/- monocytes (Fig. 5h and Supplementary Fig. 5h). These findings suggest high intracellular lactate levels in trained immunity-regulated histone lactylation.
a, b Monocytes were treated with or without β-glucan (5 μg/ml) for the indicated time before western blotting. Quantification of the immunoblots is shown on the right panel. c Monocytes were treated with or without β-glucan at indicated concentrations for 24 hrs before western blotting. Quantification of the immunoblots is shown on the right panel. d, e MS/MS spectra of a 13C-glucose labeled histone Kla peptide and its unlabeled counterpart from β-glucan treated monocytes. f, g Experiments were performed as described in (d,e), except 13C-lactate was used. h Monocytes from Wt and Ldha-/- mice were treated with or without β-glucan (5 μg/ml) for 24 hrs before western blotting. Quantification of the immunoblots is shown on the right panel. All experiments were repeated at least three times. Data in (a-c and h) are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Fig. 5.
Lactate maintains chromatin accessibility via histone lactylation in monocytes during the induction of trained immunity
To investigate the mechanical effect of lactate production on trained immunity, we performed the Assay for Transposase-Accessible Chromatin with high throughput sequencing (ATAC-seq) analysis to compare chromatin region accessibility changes between Wt monocytes, Wt, and LDHA-/- trained monocytes. We identified 2,848 chromatin regions exhibiting accessibility changes (Fig. 6a). To evaluate the functional impact of lactate on transcription, we performed RNA-seq in Wt monocytes, Wt, and LDHA-/- trained monocytes in biological duplicates (Fig. 6a). We first explored the relationship between changes in chromatin accessibility and gene expression. We assigned each differentially accessible regions (DARs) to its nearest transcription start site and examined the gene expression differences. We observed significant correlations between changes in chromatin accessibility and expression of the neighboring gene (Fig. 6b). Next, we identified 836 genes that were upregulated in Wt monocytes, Wt, and LDHA-/- trained monocytes, which were also marked with chromatin accessibility alterations (Fig. 6c). Gene ontology (GO) biological process analysis of these genes revealed significant enrichment of biological pathways, including immune response, lactate metabolism, and HIF signaling (Fig. 6d). We choose H3 lysine 27 (H3K27la), this lysine residue is also acetylated (H3K27ac), to elucidate the potential functional significance of histone lactylation in trained immunity by using CUT&Tag, a novel genome-wide immunotethering assay. Genome-wide distribution analysis reveals that H3K27la and H3K27ac histone modifications were located mainly within the promoter regions (Fig. 6e). As anticipated, H3K27la peaks were increased near transcription start sites (TSSs) in Wt-trained monocytes, but not LDHA-/- trained monocytes (Fig. 6f). However, H3K27ac peaks were slightly increased near TSSs in Wt-trained monocytes compare with Wt non-trained monocytes (Supplementary Fig. 6a). We compared the expression of H3K27la-marked genes in Wt non-trained monocytes, Wt and LDHA-/- trained monocytes. Among the total 7,547 shared peaks between the three groups, we obtained 248 peaks that were significantly changed (Fig. 6g). GO pathway analysis revealed that biological processes such as inflammatory response, lactate metabolism, and cell adhesion, which is consistent with ATAC-seq data (Fig. 6h). To test the role of H3K27la and H3K27ac in the function of trained monocytes, we integrated our ATAC-seq datasets and CUT&Tag datasets into the Integrative Genomics Viewer to examine their binding profiles on the genes. As shown in Supplementary Fig. 6b, H3K27ac peaks were slightly enhanced at the promoters of IL-6 and TNF-α in Wt-trained monocytes compared with Wt non-trained monocytes. However, H3K27la peaks were highly enriched at the promoters of critical genes of inflammatory factors and cell adhesion molecules, including IL-6, TNF-α, PSGL, and ICAM-1, in Wt-trained monocytes, but not in LDHA-/- trained monocytes (Figs. 6i, 6j and Supplementary Figs. 6c and 6d). We then used a chromatin immunoprecipitation (ChIP)-qPCR assay to assess the effects of H3K27la and H3K27ac on the stabilized effector gene expression. Indeed, H3K27la recruitment to the promoter regions of IL-6, TNF-α, PSGL, and ICAM-1 was increased in Wt-trained monocytes but not in LDHA-/- trained monocytes (Fig. 6k). Only minor differences existed between the from the H3K27ac recruitment to the promoter regions of IL-6, TNF-α, PSGL, and ICAM-1 in Wt-trained monocytes compare with the Wt non-trained monocytes (Fig. 6l). The role of histone modifications on the activation of the promoter is not restricted to one lysine, as similar results were obtained by H3K18la and H3K18ac (Supplementary Figs. 6e and 6f). In line with these observations, blocking CBP/p300 histone acetyltransferase (HAT) activity with HAT inhibitor (HATi) also decreased H3K27la and H3K18la interaction with the promoter of IL-6, TNF-α, PSGL, and ICAM-1 (Supplementary Figs. 6g and 6h). These findings suggest that intrinsic lactate production alters acetylation-dependent epigenetic programming at the promoter of inflammatory factors and cell adhesion molecules in trained monocytes.
a Heatmap representing the differentially accessible regions (DARs) of ATAC-seq (left panel) and differential gene expression of RNA-seq (right panel) in monocytes from Wt monocytes, Wt trained monocytes, and Ldha-/- trained monocytes trained as described in Fig S1b. b DARs of ATAC-seq and differential gene expression of RNA-seq subjected to Pearson’s correlation analysis. Correlation coefficient (r) and P values were calculated by Pearson’s correlation analysis. c Venn diagram depicting the number of candidate genes in ATAC-seq and upregulated genes in RNA-seq of Wt monocytes, Wt trained monocytes, and Ldha-/- trained monocytes. d GO analysis of significantly different genes in (c). The most significant and nonredundant biological processes are shown, with respective gene numbers and P values. P values by two-tailed Student’s t-test. e Pie chart showing the distribution of H3K27la and H3K27ac at annotated genomic regions in trained monocytes. f Heatmap showing the genomic occupancy of H3K27la ± 3 kb flanking TSSs in trained monocytes from Wt and Ldha-/- mice. The genes shown in rows are sorted in descending order by signal strength. g Heatmap of chromatin peak accessibility for significantly different peaks in trained monocytes from Wt and Ldha-/- mice. Each row represents a Z score of log2-transferred normalized read counts within each sample using CUT&Tag-seq. h GO analysis of significantly different genes in (g). Most significant and nonredundant biological processes with respective gene numbers and P values are shown. P values two-tailed Student’s t-test. i, j Representative ATAC-seq and CUT&Tag-seq genome browser tracks for H3K27la peak enrichment at IL-6 and TNF-α gene loci for trained monocytes from Wt and Ldha-/- mice. k, l Monocytes from Wt and Ldha-/- mice were trained as described in Fig S1b, followed by ChIP analysis. Data in (k and l) are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Fig. 6.
Exogenous lactate supports trained immunity through contribution to the TCA cycle and histone lactylation
Intracellular lactate can be produced by LDHA or imported by monocarboxylate transporter 1 (MCT1; also known as SLC16A1)12. Thus, we sought to explore the effect of MCT1 on the trained immunity. As shown in Supplementary Fig. 7a, IL-6, and TNF-α production was induced in β-glucan-trained Wt mice, but not in MCT1 inhibitor AZD3965 (AZD) treated mice. In this vein, AZD-treated mice were significantly susceptible to C. albicans-induced lethality, with higher bacterial burden in the lung and kidney tissue (Supplementary Figs. 7b and 7c). Next, we evaluate the role of MCT1 on the lactate-fueled TCA cycle by treating trained monocytes with 13C-lactate. As shown in Supplementary Figs. 7d-7f, AZD treatment abolished lactate entry into the TCA cycle. As expected, β-glucan treatment increased histone Kla levels in Wt monocytes but not in AZD-treated monocytes (Supplementary Fig. 7g). ChIP-qPCR assay indicated that AZD treatment inhibit H3K27la and H3K18la recruitment to the promoter regions of IL-6, TNF-α, PSGL, and ICAM-1 (Supplementary Figs. 7h and 7i). Hence, MCT1-mediated lactate uptake service as a TCA cycle substrate regulated cytokine production via histone lactylation.
Our data revealed that β-glucan mildly induced histone acetylation. We hypothesized that lactate is taken up and readily converted to pyruvic acid through LDHA and subsequently converted to cytosolic acetyl-CoA in two enzymatic catalyzed reactions by pyruvate dehydrogenase (PDHA) and ATP-citrate lyase (ACLY), respectively33 (Fig. 7a). To this purpose; we compare the role of glucose and lactate on acetyl-CoA generation. As shown in Fig. 7b, neither high glucose (HG) nor low glucose (LG) affect β-glucan-regulated acetyl-CoA generation-however, lactate treatment induced β-glucan regulated acetyl-CoA generation, indicating that β-glucan promotes acetyl-CoA generation via lactate. Still, not glucose (Fig. 7b). To dissect the key step of lactate metabolism involved in acetyl-CoA generation, we employed a series of pharmacologically targeting LDHA, PDHA and ACLY by the LDHA inhibitor (sodium oxamate, Oxa), pyruvate transporter inhibitor (UK5099), and ACLY inhibitor (BMS303141, BMS), respectively (Fig. 7a). Interesting, treatment of Oxa, UK5099, and BMS inhibit β-glucan induced acetyl-CoA generation (Fig. 7c). LC/MS analysis focused on isotopologues of acetyl-CoA demonstrate that lactate carbons labeled the majority of acetyl-CoA. The m + 2 acetyl-CoA isotopologue revealed the most substantial labeling pattern derived from lactate carbons (Figs. 7d and 7e). To pinpoint the pathway in how lactate facilitates histone acetylation, we hypothesized that this involves oxidative metabolism, particularly ACLY. Strikingly, BMS counteracted β-glucan-mediated histone acetylation but not histone lactylation (Fig. 7f). Surprising, the ACLY inhibitor did not affect β-glucan-mediated IL-6 and TNF-α production, and bacterial burden in the lung and kidney tissue in vitro and in vivo (Figs. 7g-7i). In addition, BMS treatment inhibits H3K27ac and H3K18ac, but not H3K27la and H3K18la, recruitment to the promoter regions of IL-6, TNF-α, and PSGL (Figs. 7j-7m). These results suggest that lactate is a significant substrate for acetyl-CoA production. Compared with lactate-mediated histone acetylation, lactate-mediated histone lactylation plays a dominant role in trained immunity.
a Schematic of 13C-lactate carbon labeling through the TCA cycle. b Monocytes were cultured with high glucose (HG) (15 mM), low glucose (LG) (3 mM), and/or lactate (1 mM), and treated with or without β-glucan (5 μg/ml) for 24 hrs, followed by measuring acetyl-CoA levels. c Monocytes were cultured in lactate media (0.5 mM glucose, 0.5 mM glutamine, and 2 mM lactate) and treated with or without β-glucan (5 μg/ml), sodium oxamate (20 mM), UK5099 (10 μM), BMS303141 (BMS) (10 μM) for 24 hrs, followed by measuring acetyl-CoA levels. d, e Monocytes were cultured in MM containing 13C-lactate (1 mM) and treated with β-glucan (5 μg/ml) for 4 hrs, followed by LC/MS analysis. Shown are labeled and unlabeled fractions of acetyl-CoA. f Monocytes were cultured in lactate media (0.5 mM glucose, 0.5 mM glutamine, and 2 mM lactate) and treated with or without β-glucan (5 μg/ml), sodium oxamate (20 mM), UK5099 (10 μM), BMS (10 μM) for 24 hrs before western blotting. Quantification of the immunoblots is shown on the right panel. Experiments were repeated at least three times. g Monocytes were cultured in lactate media (0.5 mM glucose, 0.5 mM glutamine, and 2 mM lactate), treated with or without β-glucan (5 μg/ml), BMS (10 μM) for 24 hrs, and then IL-6 and TNF-α levels were measured. h, i Wt mice were trained with or without β-glucan. Six days later, mice were infected with or without a secondary C. albicans lethal dose (3 × 106) and/or BMS (50 mg/kg) for 24 hrs, followed by measuring IL-6 and TNF-α levels (h) and fungal burden (i). j–m Monocytes from Wt mice were treated as described in (g), followed by ChIP analysis. Data are expressed as means ± SEMs, n = 3 mice per condition, two-way ANOVA. See also Supplementary Fig. 7.
Discussion
Trained immunity refers to the long-lasting memory traits of innate immunity34. How environmental nutrient availability impacts trained immunity still needs to be better understood. Here, for the first time, we show that trained monocytes exhibit a high degree of plasticity in fuel choice, not only conventional fuels such as glucose but also PCSs. Exploring this, we demonstrated that cell-intrinsic lactate is a prominent fuel in trained monocytes, contributing to the TCA cycle, even when glucose is available. Mechanistically, lactate was a significant substrate for acetyl-CoA production in trained monocytes and regulated trained monocytes’ function through effects on histone lactylation. Our study highlights a previously underappreciated role of lactate in trained immunity.
Immunological memory has long been ascribed only to adaptive T and B cells35,36. The recently emerged concept of trained immunity is defined as causing long-term alterations in innate immune cells, enabling a robust innate host response to a secondary stimulus35,36. Analysis of differentially regulated pathways in trained monocytes demonstrated an essential role in changes in intracellular metabolism, especially glucose metabolism24. The mTOR-HIF1α pathway regulated a shift from oxidative phosphorylation to glycolysis, the central regulatory mechanism for trained immunity24. This study observed the high lactate levels, an end product of glycolysis, in trained immunity, but the physiological significance of overexpressed lactate was not explored. We show lactate is a physiologic fuel for trained immunity, serving as a preferred TCA cycle and biosynthetic substrate. Additionally, lactate regulates cytokine production through its effects on histone lactylation. Our results strongly complement the previous work, filling the downstream signaling pathways of trained immunity. We propose a working model of the most prominent regulated pathways in trained immunity based on prior studies and our present data. According to this model, trained innate immune cells activate the Akt-mTOR-HIF-1α pathway, leading to a shift in metabolism from oxidative phosphorylation to glycolysis and high lactate accumulation. A portion of elevated lactate levels serves as a physiologic fuel for the TCA cycle. The other part of elevated levels of lactate enters into the nucleus and binds the loci of cytokines promoter, leading to maintaining the accessibility of chromatin. If trained innate immune cells encounter a secondary stimulus, accessible chromatin permits transcription factors to quickly bind to the corresponding promoter, leading to a faster and more robust response.
Recently, some research has revealed the link between the stimulation of innate immune pathways and the induction of epigenetic and metabolic changes in trained immune cells37,38. Rob et al. show that glutaminolysis is another metabolic pathway involved in β-glucan induced trained immunity, linked to enrichment in H3K4me3, which is essential for trained immunity37. However, there was no detectable difference in the expression of methyltransferases between non-trained and trained immune cells. Siroon et al. show that the cholesterol/mevalonate synthesis pathway is elevated in β-glucan-trained mice, which is linked to enrichment in H3K27ac38. However, the entire genome H3K27ac study does not reveal a complete overlap between β-glucan and mevalonate-induced trained immunity. We suspect part of this phenomenon is due to the cell culture medium. The current cell culture medium is designed to promote cell growth and survival. Thus, the current cell culture medium contains supraphysiological concentrations of some nutrients while absenting other nutrients relevant to cellular metabolism. An important implication of this study is that glucose utilization by trained monocytes is highly influenced by other non-glucose substrates, precisely a few of the nutrients absent from most medium formulations today. Our study established that lactate is a primary source of acetyl-CoA production in trained monocytes. This directly contributes to histone lactylation and promotes permissive lysine lactylation in histones at cytokines gene loci. Mechanistically, we show that LDHA, which controls lactate metabolism via regulating lactate/pyruvate conversion, is required for a lactate-fueled TCA cycle and regulated histone lactylation. Our study demonstrates that lactate is the hub between metabolism, immunity, and epigenetics in trained monocytes.
It has yet to slip our attention that there are several important caveats. (i) The distinctions and linkages of histone lactylation, histone methylation, and histone lactylation in trained immunity still need to be explored. (ii) We only focus on the role of higher concentration (exceeding 100 μM) PCS on the glucose utilization of trained immunity. Lower concentration PCS may also serve as bioenergetic fuels in trained immune cells when the nutrients are unavailable. (iii) We did not explore whether lactate metabolism is an effective target in clinical treatment. (iv) The direct causal effect of lactylation on trained immunity is challenging to demonstrate, as p300 inhibitors also block histone acetylation. (v) LDHA knockout has a significant impact on survival in this model, irrespective of β-glucan treatment (Fig. 2e). This likely arises from the critical role of LDHA in supporting glycolytic responses to infection of all innate immune cells (in the absence of immune training). (vi) Comparison of histone accessibility between WT and LDHA cells to infer lactate-mediated changes (without lactate-treated controls) is challenging, because loss of LDHA will significantly alter the metabolic and functional status of these cells (for example by limiting glycolytic flux and NAD/H redox) in addition to its role permitting lactate generation and subsequent usage. When considering the next step, studies exploring these questions would greatly help clarify the role of lactate in trained immunity.
Methods
Ethics statement
Consistent with the Helsinki Declaration, the Institutional Review Board of Wuhan University approved the collection of clinical samples following guidelines for protecting human subjects. All study participants provided written informed consent for collecting samples and subsequent analyzes.
All animal experiments followed the National Institutes of Health Guide for the Care and Use of Laboratory Animals. The protocols and procedures were approved by the Institutional Animal Care and Use Committee of Wuhan University (project license WDSKY0201302).
Monocyte and peripheral blood mononuclear cell isolation
Peripheral blood mononuclear cells (PBMCs) were isolated from the blood of healthy donors by standard Ficoll gradient centrifugation and cultured in RPMI 1640 without FBS and antibiotics. Percoll (Sigma-Aldrich) isolation of monocytes was performed as previously described39. Briefly, 100 − 300 × 106 PBMCs were layered on top of a hyper-osmotic personal solution (48.5% Percoll, 41.5% H2O, and 0.16 M NaCl) and centrifuged for 15 min at 580 × g. The interphase layer was isolated, and cells were washed with cold Phosphate-Buffered Saline (PBS).
Western blot analysis
Western blot analyzes were performed as previously described40. Briefly, cells were harvested by low-speed centrifugation and washed with PBS. Cells were lysed in RIPA Buffer (Cell Signaling Technology, Boston, MA, US), and protein concentrations were determined using BCA assays. Forty micrograms of each protein sample were separated using 12% SDS-PAGE and transferred to nitrocellulose membranes (Bio-Rad). Membranes were blocked with 1×Tris-buffered saline with Tween 20 (TBST) and 5% (w/v) non-fat milk for 1 hour at room temperature. Then, membranes were incubated with primary antibodies overnight at 4 °C. Subsequently, blots were incubated with horseradish peroxidase-linked secondary antibodies (Jackson ImmunoResearch) for an additional 1 hour. Immunoreactive bands were visualized using an enhanced chemiluminescence system (GE Healthcare).
Cytokine measurements
Cytokines were determined in PBMC culture supernatants by a multiplex Procartaplex assay (Thermo Fisher Scientific, USA) and acquired on a Luminex 200 System (Thermo Fisher Scientific, USA) according to the manufacturer’s protocol.
Chromatin immunoprecipitation sequencing (ChIP-seq) analysis
Samples were cross-linked in suspension for 5 min in 10 mL of 0.75% formaldehyde (Pierce, #28906) in PBS at room temperature. Nuclei were separated using the cell nucleus isolation kit (Thermo Fisher Scientific, #78835). Lysates were diluted to 500 mL by the addition of binding buffer (25 mM HEPES pH 7.5, 100 mM NaCl, 0.1% NP-40). 5 mL of RNAse A/T1 (Thermo Fisher Scientific, #EN0551) was added, and the sample was incubated at 37 °C for 25 min. Then, CaCl2 was added to a final concentration of 40 mM, followed by 75 U of micrococcal nuclease (Worthington Biochemical), and incubated at 37 °C for 5 min. MNase was quenched by adding 40 mM EDTA, and the total volume was brought to 1.2 mL with binding buffer. Next, insolubilities were removed by centrifugation at max speed (21,000 RCF) at 4 °C for 5 min, and the supernatant containing soluble chromatin was collected. At this stage, 5 mL of chromatin was measured using the Qubit dsDNA HS Assay Kit (Invitrogen, #Q32851).
Samples were diluted with binding buffer to ensure similar chromatin concentrations and to match IP conditions. 50 mL of chromatin was set aside as input for each sample. For each IP, 25 mL of Protein A-coated magnetic beads (Invitrogen, #10008D) were washed once with binding buffer and incubated with either 0, 2.5, or 10 mL of H3K27Ac antibody. The total volume of bead plus antibody was brought to 200 mL using the binding buffer and was rotated at room temperature for 30 min. To compare post-translational modifications between samples, we performed one biological replicate of all samples on the same day. We made a master mix of bead plus antibodies, scaling up all components by the number of samples. Buffer-containing antibody was removed, and bead plus antibody was resuspended in 200 mL of soluble chromatin followed by 30 min rotation at room temperature. Unbound chromatin was removed, and beads were vortexed with 500 mL of binding buffer. Buffer was removed, and bound material was eluted from beads by vortexing in 133 mL of elution buffer. At this time, the input was brought to 133 mL by adding 83 mL elution buffer. Proteinase K (Invitrogen, #25530015) was added to a final concentration of 15 mM, and the sample was incubated overnight at 37 °C.
The following morning, each DNA sample was purified using a MinElute PCR Kit (Qiagen, #28004) and eluted in 30 mL of Buffer EB. 5 mL of DNA was quantified by Qubit dsDNA HS Assay Kit. The remaining 25 mL of DNA was frozen at -20 °C until library preparation. Libraries were sequenced using a NextSeq 500 (Illumina) using 75 bp paired-end sequencing (5 × 107 reads/sample for IPs, 1 × 108 reads/sample for input). Next-generation sequencing (NGS) data were aligned to the mm 10 genome. Bed files generated by the NGS alignment were processed using the latest siQ-ChIP release (https://github.com/BradleyDickson/siQ-ChIP), and siQ-ChIP quantification was performed. Responses were computed automatically by the siQ-ChIP software as the area ratio under overlapping peaks for any pair of tracks being compared. Individual ChIP tracks were visualized using IGV.81.
CUT&Tag sequencing analysis
The library preparation for CUT&Tag was performed as previously reported41. In brief, cells were washed twice with PBS, concanavalin A-coated magnetic beads (Bangs Laboratories, BP531) in 500 μL Wash Buffer [20 mM Hepes, pH 7.5, 150 mM NaCl, 0.5 mM Spermidine, and protease inhibitor Cells were incubated with 10 μL of the activated mixture (Roche)] for 10 min at room temperature. Cell-bound beads were collected and resuspended with 50 μL Dig-Wash Buffer (20 mM Hepes pH 7.5, 150 mM NaCl, 0.5 mM Spermidine, protease inhibitor mixture, and 0.05% digitonin) containing 2 mM EDTA, 0.1%BSA, and a 1:50 dilution of the primary antibody, and incubated at 4 °C overnight. A secondary antibody diluted at 1:100 in 100 μL of Dig-Wash Buffer was then administered into the beads and incubated for 60 min at room temperature following primary antibody removal with a magnet stand (Vazyme, CM101). The pG-Tn5 adapter complex was prepared according to the manufacturer’s instruction with Hyperactive pG-Tn5 Transposase for CUT&Tag (Vazyme, S602). Standard segmentation and amplification were performed as reported previously3. Amplified DNA libraries were purified with VAHTS DNA Clean Beads (Vazyme, N411) and shipped for next-generation sequencing (NGS) sequencing by Annoroad Gene Technology. Reads were filtered and mapped to the mm10 genome, and peaks were then called to generate a peak matrix with a pipeline similar to the ATAC-seq analysis mentioned above. Differential peaks were identified by log2FC > 0.5, P < 0.05, and normalized read counts > 0.
ATAC sequencing analysis
The ATAC sequencing was performed as previously reported42. In brief, cells were lysed with lysis buffer (10 mM Tris-HCl, pH 7.4, 10 mM NaCl, 3 mM MgCl2, and 0.1% (vol/vol) Igepal CA-630) at 4 °C for 10 min. Segmentation and amplification were done following the manufacturer’s instructions with TruePrep DNA library Prep Kit V2 (Vazymes, TD501). Annoroad Gene Technology performed single-end sequencing, and clean reads with trimmed adapters were aligned to the mm10 reference genome with the Bowtie2 (2.3.5.1) package43. Broad peaks were called by the Macs2 (2.1.2) package44 using the parameter (-nomodel-shift-100-extsize 200 -B-broad), and differential accessed peaks were called using the Deseq2 (v.1.20.0) package45. Differential accessed peaks were called by log2FC > 0.5 and P < 0.05 and normalized read counts > 0. Motif enrichment analysis and peak-associated gene annotation were performed using HOMER (v4.11.1) using peaks filtered by log2FC > 0.5, P < 0.05 and normalized read counts > 0. Browser tracks were visualized by IGV Browser (v2.8.2) after normalizing the read from each sample to its library size. Ataqv (v1.0.0, https://github.com/ParkerLab/ataqv) package developed by the Parker Laboratory from the University of Michigan was used to perform the ATAC-seq data quality control analysis.
Mice
Wild-type (Wt) C57BL/6 mice were purchased from Sanxia University (Yichang, Hubei). Ldha-/- mice on a C57BL/6 J background were previously described46. Littermates of the same sex were randomly assigned to experimental groups. Mice were randomly allocated to experimental groups, and the investigator was blinded to the group allocation during the experiment. 6-8 weeks-old mice were treated with tamoxifen (20 mg/ml in corn oil) or coin oil by intraperitoneal injection. According to protocols provided by the Shanghai Model Organisms Center (Shanghai, China) for inducible Cre driver lines, LDHA exon2 deletion was confirmed by genotyping 10 days after the final injection. All the analyzes were performed blindly. Ldha-/- mice were randomly allocated into experimental groups for further treatment, and cell samples were assigned based on the genotype of interest. All mice were bred under specific pathogen-free conditions in the accredited animal facility at Wuhan University.
Bacterial culture
Mycobacterium tuberculosis (Mtb) H37Rv were grown in 7H9 broth (BD) supplemented with 0.2% glycerol (Wisent), 0.05% Tween80 (Fisher), and 10% albumin-dextrose-catalase under constant shaking at 37 °C. For in vitro and in vivo experiments, Mtb (H37Rv) bacteria in log growing phase (OD 0.4-0.9) were centrifuged (4000 RPM, 15 min) and resuspended in RPMI without penicillin/streptomycin or sterile PBS.
Chromatin immunoprecipitation (ChIP)-quantitative PCR (qPCR) assay
Formaldehyde was added to the culture medium to a final concentration of 1%. The cells were then washed twice with PBS, scraped, and lysed in lysis buffer (1% SDS, 10 mM Tris-HCl [pH 8.0], 10% protease inhibitor cocktail, 50 mg/ml each of aprotinin and leupeptin) for 10 min at 4 °C. The lysates were sonicated on ice, and the debris was removed by centrifugation at 12,000 rpm for 15 min at 4 °C. One-fourth of the supernatant was used as the DNA input control. The remaining sample was diluted 10-fold with dilution buffer (0.01% SDS, 1% Triton X-100, 1 mM EDTA, 10 mM Tris-HCl [pH 8.0], and 150 mM NaCl) followed by incubation with antibodies overnight at 4 °C. Immunoprecipitated complexes were collected using protein A/G-Sepharose. The pellets were washed four times with dialysis buffer containing 2 mM EDTA and 50 mM Tris-HCl, pH 8.0. After washing, the precipitates were incubated with an elution buffer (1% SDS and 0.1 M NaHCO3) at room temperature. Supernatants were transferred to clean tubes, and RNase A was added to destroy the bound RNA in the sample. Samples were incubated at 65 °C for 5 hrs to reverse the formaldehyde cross-links, and DNA was precipitated with ethanol and extracted two times with phenol-chloroform. Finally, pellets were resuspended in Tris-EDTA (TE) buffer and subjected to PCR amplification. Quantitative real-time RT-PCR was performed using a LightCycler 480 (Roche) and the SYBR green system (Applied Biosystems). GAPDH was amplified as an internal control.
HPLC-MS/MS analysis
Mass spectrometry and metabolite identification were performed on 80% methanol and 20% LC/MS-grade water-extracted metabolites. Analyzes were performed using High-Performance Liquid Chromatography, High-Resolution Mass Spectrometry, and Tandem Mass Spectrometry (HPLC-MS/MS). The system consisted of a Thermo Q-Exactive in line with an electrospray source and an Ultimate 3000 (Thermo) series HPLC consisting of a degasser, a binary pump, and an autosampler outfitted with an Xbridge Amide column (dimensions of 4.6 mm × 100 mm and a 3.5 μm particle size). The mobile phase A: 20 mM ammonium acetate (pH 9.0), 20 mM ammonium hydroxide, 95% (v/v) water, and 5% (v/v) acetonitrile. The mobile phase B: 100% Acetonitrile. The gradient was as follows: 15% A (0 min); 30% A (2.5 min); 43% A (7 min); 62% A (16 min); 75% A (15–20 min); 15% A (15–20 min); with a flow rate of 400 μL/min. The capillary of the ESI source was set to 275 °C, with sheath gas at 45 arbitrary units, auxiliary gas at five arbitrary units, and the spray voltage at 4.0 kV. The top 5 precursor ions were fragmented using the higher energy collisional dissociation cell set to 30 % normalized collision energy in MS2. Data acquisition and analysis were carried out by Tracefinder 2.1 software (Thermo Fisher Scientific) and Xcalibur 4.0 software (Thermo Fisher Scientific).
In vivo 13C-tracer infusions
Mice were infused intravenously with 13C-labeled tracers. 13C-lactate infusion rate is 1 mL/min for 4 h for a 20 g mouse. Then, serum and lung were isolated and processed for metabolomic analysis.
Quantification of mitochondrial ATP level by flow cytometry
For FRET-based ATP determination, a Novocyte 3000 flow cytometer (ACEA BioSciences) was used to record ERAT fluorescence at channels with Ex/Em filters set as follows: (1) 405 nm/445 nm (bandwidth: 45). (2) 405 nm/530 nm (bandwidth: 30). Mitochondrial ATP levels for individual cells were defined as the ratio of fluorescence intensity of channel 2 (Ex/Em: 405/530) divided by that of channel 1 (Ex/Em: 405/445), similar to parameters used previously47,48.
Reagents and critical commercial assay kits
The details of critical reagents, antibodies, and commercial assay kits are listed in Supplementary Table 1 and Supplementary Table 2. Unless specified otherwise, all biochemical reagents were purchased from Sigma-Aldrich.
Statistical analysis
Data were obtained from three independent reproducible experiments. Data were expressed as mean ± standard deviations or mean ± the standard error of the mean. Statistical significance was determined using Student’s unpaired two-tailed t-test, one-way or two-way ANOVA multiple comparison tests as indicated in the legend. A p-value < 0.05 was considered significant and was marked with an asterisk (*).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data presented in the study are included in this article and supplementary material. ChiP-seq/CUT&Tag sequencing and ATAC-sequencing datas are deposited in NCBI under accession numbers: SAMN47310247 [https://www.ncbi.nlm.nih.gov/biosample/47310247], SAMN47310248 [https://www.ncbi.nlm.nih.gov/biosample/47310248], SAMN47310249 [https://www.ncbi.nlm.nih.gov/biosample/47310249], SAMN47310250 [https://www.ncbi.nlm.nih.gov/biosample/47310250], SAMN47310251 [https://www.ncbi.nlm.nih.gov/biosample/47310251], SAMN47310252 [https://www.ncbi.nlm.nih.gov/biosample/47310252]). The metabolomics data was deposited in the Metabolomics Workbench (Study ID: ST003833, https://dev.metabolomicsworkbench.org:22222/data/DRCCMetadata.php?Mode=Study&StudyID=ST003833). Further inquiries can be directed to the corresponding author. Source data are provided with this paper.
References
Stienstra, R., Netea-Maier, R. T., Riksen, N. P., Joosten, L. A. B. & Netea, M. G. Specific and complex reprogramming of cellular metabolism in myeloid cells during innate immune responses. Cell Metab. 26, 142–156 (2017).
Zhou, L. et al. Hepatitis B virus rigs the cellular metabolome to avoid innate immune recognition. Nat. Commun. 12, 98 (2021).
He, Q. et al. MAVS integrates glucose metabolism and RIG-I-like receptor signaling. Nat. Commun. 14, 5343 (2023).
Pan, C., Li, B. & Simon, M. C. Moonlighting functions of metabolic enzymes and metabolites in cancer. Mol. Cell 81, 3760–3774 (2021).
Muir, A., Danai, L. V. & Vander Heiden, M. G. Microenvironmental regulation of cancer cell metabolism: implications for experimental design and translational studies. Dis. Model Mech. 11, dmm035758 (2018).
Hackett, S. R. et al. Systems-level analysis of mechanisms regulating yeast metabolic flux. Science 354, aaf2786 (2016).
Rossiter, N. J. et al. CRISPR screens in physiologic medium reveal conditionally essential genes in human cells. Cell Metab. 33, 1248–1263.e9 (2021).
Puleston, D. J., Villa, M. & Pearce, E. L. Ancillary activity: beyond core metabolism in immune cells. Cell Metab. 26, 131–141 (2017).
Xu, S. et al. Interleukin-6 classic and trans-signaling utilize glucose metabolism reprogramming to achieve anti- or pro-inflammatory effects. Metabolism 155, 155832 (2024).
Peng, M. et al. Aerobic glycolysis promotes T helper 1 cell differentiation through an epigenetic mechanism. Science 354, 481–484 (2016).
Lin, J., Liu, G., Chen, L., Kwok, H. F. & Lin, Y. Targeting lactate-related cell cycle activities for cancer therapy. Semin Cancer Biol. 86, 1231–1243 (2022).
Faubert, B. et al. Lactate metabolism in human lung tumors. Cell 171, 358–371.e9 (2017).
Zhang, W. et al. Lactate is a natural suppressor of RLR signaling by targeting MAVS. Cell 178, 176–189.e15 (2019).
Hui et al. Glucose feeds the TCA cycle via circulating lactate. Nature 551, 115–118 (2017).
Zhang, D. et al. Metabolic regulation of gene expression by histone lactylation. Nature 574, 575–580 (2019).
Chen, A.-N. et al. Lactylation, a novel metabolic reprogramming code: current status and prospects. Front Immunol. 12, 688910 (2021).
Khan, N. et al. M. tuberculosis reprograms hematopoietic stem cells to limit myelopoiesis and impair trained immunity. Cell 183, 752–770.e22 (2020).
Mitroulis, I. et al. Modulation of myelopoiesis progenitors is an integral component of trained immunity. Cell 172, 147–161.e12 (2018).
Kalafati, L. et al. Innate immune training of granulopoiesis promotes anti-tumor activity. Cell 183, 771–785.e12 (2020).
Cirovic, B. et al. BCG vaccination in humans elicits trained immunity via the hematopoietic progenitor compartment. Cell Host Microbe 28, 322–334.e5 (2020).
Netea, M. G. et al. Trained immunity: a program of innate immune memory in health and disease. Science 352, aaf1098 (2016).
Netea, M. G. & van der Meer, J. W. M. Trained immunity: an ancient way of remembering. Cell Host Microbe 21, 297–300 (2017).
Netea, M. G. et al. Defining trained immunity and its role in health and disease. Nat. Rev. Immunol. 20, 375–388 (2020).
Cheng, S.-C. et al. mTOR- and HIF-1α–mediated aerobic glycolysis as metabolic basis for trained immunity. Science 345, 1250684 (2014).
Cantor, J. R. et al. Physiologic medium rewires cellular metabolism and reveals uric acid as an endogenous inhibitor of UMP synthase. Cell 169, 258–272.e17 (2017).
Kaymak, I. et al. Carbon source availability drives nutrient utilization in CD8+ T cells. Cell Metab. 34, 1298–1311.e6 (2022).
Luda, K. M. et al. Ketolysis drives CD8+ T cell effector function through effects on histone acetylation. Immunity 56, 2021–2035.e8 (2023).
Ron-Harel, N. et al. T cell activation depends on extracellular alanine. Cell Rep. 28, 3011–3021.e4 (2019).
Su, H. et al. Glutathione synthesis primes monocytes metabolic and epigenetic pathway for β-glucan-trained immunity. Redox Biol. 48, 102206 (2021).
Luengo, A. et al. Increased demand for NAD+ relative to ATP drives aerobic glycolysis. Mol. Cell 81, 691–707.e6 (2021).
Peignier, A. & Parker, D. Trained immunity and host-pathogen interactions. Cell Microbiol 22, e13261 (2020).
Zhao, Y. et al. In vivo monitoring of cellular energy metabolism using SoNar, a highly responsive sensor for NAD(+)/NADH redox state. Nat. Protoc. 11, 1345–1359 (2016).
Cheng, S.-C. et al. Broad defects in the energy metabolism of leukocytes underlie immunoparalysis in sepsis. Nat. Immunol. 17, 406–413 (2016).
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Farber, D. L., Netea, M. G., Radbruch, A., Rajewsky, K. & Zinkernagel, R. M. Immunological memory: lessons from the past and a look to the future. Nat. Rev. Immunol. 16, 124–128 (2016).
Netea, M. G., Quintin, J. & van der Meer, J. W. M. Trained immunity: a memory for innate host defense. Cell Host Microbe 9, 355–361 (2011).
Arts, R. J. W. et al. Glutaminolysis and fumarate accumulation integrate immunometabolic and epigenetic programs in trained immunity. Cell Metab. 24, 807–819 (2016).
Bekkering, S. et al. Metabolic induction of trained immunity through the mevalonate pathway. Cell 172, 135–146.e9 (2018).
Repnik, U., Knezevic, M. & Jeras, M. Simple and cost-effective isolation of monocytes from buffy coats. J. Immunol. Methods 278, 283–292 (2003).
Yu, H. et al. Major vault protein promotes hepatocellular carcinoma through targeting interferon regulatory factor 2 and decreasing p53 activity. Hepatology 72, 518–534 (2020).
Kaya, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 29, 1930 (2019).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Xie, H. et al. Targeting lactate dehydrogenase-a inhibits tumorigenesis and tumor progression in mouse models of lung cancer and impacts tumor-initiating cells. Cell Metab. 19, 795–809 (2014).
Imamura, H. et al. Visualization of ATP levels inside single living cells with fluorescence resonance energy transfer-based genetically encodedindicators. PNAS 106, 15651–15656 (2009).
Vishnu, N. et al. ATP increases within the lumen of the endoplasmic reticulum upon intracellular Ca2+ release. Mol. Biol. Cell 25, 368–379 (2014).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (grant numbers: 2021YFC2701800, 2021YFC2701804, 2023YFC2307800), the National Natural Science Foundation of China (grant number U22A20335), the Fundamental Research Funds for the Central Universities (grant number: 2042024kf0011, 2042022dx0003), the Natural Science Foundation of Wuhan (grant number: 2024040701010031), the Huxiang high-level talents gather engineering innovation and entrepreneurship talents in Hunan province (grant number: 2021RC5006), the funds from Science and Technology Department of Hubei Province (grant number: 2022CFB624), the Knowledge Innovation Program of Wuhan (grant number: 2022020801010116), the Fundamental Research Funds for the Central Universities (grant number: 2042021kf023).
Author information
Authors and Affiliations
Contributions
H.C., X.C., Y.L.: Data curation, Project administration, Writing–original draft. Y.C., G.Z.: Data curation, Project administration, Resources. X.C.: Resources. S.R., H.Z.: Data curation. L.Z.: Project administration. Z.Li.: Supervision. A.L.: Conceptualization. X.Z.: Project administration. W.X., Li.Z.: Data curation. C.G.: Resources. Y.Z.: Resources. K.D.: Resources. Z.Lu., H.R.: Resources. H.Y., Z.C., K.Xu.: Conceptualization, Funding acquisition, Resources, Supervision. F.W., S.L.: Conceptualization, Funding acquisition, Resources, Supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks, the anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cai, H., Chen, X., Liu, Y. et al. Lactate activates trained immunity by fueling the tricarboxylic acid cycle and regulating histone lactylation. Nat Commun 16, 3230 (2025). https://doi.org/10.1038/s41467-025-58563-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58563-2
This article is cited by
-
Lactate: elucidating its indispensable role in human health
Molecular Cancer (2026)
-
Injectable hydrogels for osteomyelitis treatment induce metabolic reprogramming for protection against reinfection
Nature Communications (2026)
-
PANoptosis in diabetes: immunometabolic insights and treatments
Apoptosis (2026)
-
Lactylation at the crossroads of immune metabolism and epigenetic regulation: revealing its role in rheumatic immune diseases
Journal of Translational Medicine (2025)
-
Hexokinase 2-mediated histone H3K18la promotes PAI-1-dependent thrombosis in acute myeloid leukemia via tumor-endothelial crosstalk
Nature Communications (2025)