Abstract
Central serous chorioretinopathy is an eye disease characterized by fluid buildup under the central retina whose etiology is not well understood. Abnormal choroidal veins in central serous chorioretinopathy patients have been shown to have similarities with varicose veins. To identify potential mechanisms, we analyzed genotype data from 1,477 patients and 455,449 controls in FinnGen. We identified an association for a low-frequency (allele frequency = 0.5%) missense variant (rs113791087) in PTPRB, the gene encoding vascular endothelial protein tyrosine phosphatase (odds ratio=2.85, P = 4.5 × 10-9). This was confirmed in a meta-analysis of 2,452 patients and 865,767 controls from 4 studies (odds ratio=3.06, P = 7.4 × 10-15). Rs113791087 was associated with a 56% higher prevalence of retinal abnormalities (35.3% vs 22.6%, P = 8.0 × 10-4) in 708 UK Biobank participants and, surprisingly, with increased risk of varicose veins (odds ratio=1.31, P = 2.3 × 10-11) and reduced risk of glaucoma (odds ratio=0.82, P = 6.9 × 10-9). Predicted loss-of-function variants in PTPRB, though rare in number, were associated with central serous chorioretinopathy in All of Us (odds ratio=17.09, P = 0.018). These findings highlight the significance of vascular endothelial protein tyrosine phosphatase in diverse ocular and systemic veno-vascular diseases.
Similar content being viewed by others
Introduction
Central serous chorioretinopathy (CSC) is a maculopathy associated with a thickened and dilated choroidal vasculature, retinal pigment epithelium (RPE) detachments and subretinal fluid (SRF). CSC often first manifests with decreased visual acuity in individuals between 30 and 50 years of age1. CSC is relatively common worldwide, with annual age-adjusted incidence estimates between 5.8:100,000 and 34:100,000 in different populations1,2; a prevalence as high as 1.7% has been reported in India3,4.
For some patients the SRF resolves spontaneously and quickly, but for others the SRF persists or recurs leading to photoreceptor damage and visual decline. Photodynamic therapy is a preferred treatment for chronic CSC but is ineffective in resolving SRF in a noteworthy percentage of patients2.
The etiology of CSC is not well understood, limiting therapeutic development and primary prevention. Imaging of the choroidal vessels via fluorescein and indocyanine green (ICG) angiography have revealed characteristic focal leakage and staining patterns representing leakage of dye from the choroid to the subretinal space5,6. Several risk factors have been noted, including endogenous and exogenous corticosteroids and pregnancy7; however, none of these have a clear known pathophysiologic mechanism causing CSC.
Original etiologic discussions focused on RPE hyperpermeability, but this theory does not explain the notably thickened choroid with dilated vasculature. Spaide and colleagues brought forth a convincing mechanistic explanation in 2022 which incorporated recent enhanced depth optical coherence tomography (OCT) and widefield ICG angiography findings and ascribed CSC at least in part to intrinsic venous outflow insufficiency8. The dilated vasculature, delayed choroidal filling and arterio-venous anastomoses noted on ICG angiography contribute toward a theory of CSC's being a venous overload choroidopathy. The vascular remodeling in CSC has been compared most closely to varicose veins, although the changes are on a much smaller scale. Varicose veins are a common disease affecting approximately 20% of the population (females more than males) and represent a weakening of the vessel wall with resultant vascular dilation in the lower extremity (typically in the greater and lesser saphenous veins)9.
Genome-wide analysis of variants associated with CSC has recently enjoyed success10,11,12. To date, these studies have identified 6 common variant loci linked to CSC. In a recent CSC meta-analysis12, we identified associated loci encoding genes that are preferentially expressed in the choroidal vasculature, which complements the theory of CSC's being a disorder of vascular competence. However, aside from the well-documented Complement Factor H locus—shared between CSC and age-related macular degeneration (AMD)10,11,12,13—no coding variants or clearly causal non-coding variants have robustly pointed to specific genes, and thus mechanistic understanding remains limited.
Here, we conduct a genome-wide association study of CSC in new data from the FinnGen study. We further evaluate the associations of a newly identified locus with CSC and co-associated traits across 5 studies, including over 1.3 million individuals. These findings provide an intriguing window into shared underlying pathophysiology between ocular diseases and systemic vascular dysfunction.
Results
A genome-wide association study of CSC in FinnGen identifies a missense variant in PTPRB
We began by conducting a case-control GWAS including 1477 patients with CSC and 455,449 controls in FinnGen (Supplementary Data 1; Supplementary Results). The prevalence of other ocular diseases among patients with CSC was low (Supplementary Data 2 and Supplementary Results). This analysis identified 3 loci at genome-wide significance (P < 5 × 10-8; Fig. 1 and Supplementary Fig. 1), 2 of which have been previously reported and are marked by common noncoding variants at CFH (Complement Factor H) and CD46 (Membrane cofactor protein). At the 12q15 locus, the lead variant was a low-frequency (AF = 0.5%) missense variant (rs113791087, 12:70559589:T:G) in the PTPRB (Protein Tyrosine Phosphatase Receptor Type B; NCBI Gene ID: 5787) gene that encodes the vascular endothelial protein tyrosine phosphatase (VE-PTP) protein (UniProt ID: P23467) (Supplementary Fig. 2–3). Rs113791087 was associated with an increased risk of CSC (OR = 2.85 [2.01–4.05] per G allele, P = 4.5 × 10−9). Fine-mapping with the sum of single effects (SuSiE) approach identified a single credible set in the locus that contained only the variant rs113791087 with a posterior inclusion probability of 0.995 (Supplementary Data 3).
A genome-wide association study of central serous chorioretinopathy was conducted, including 1477 patients with central serous chorioretinopathy and 455,449 controls from the FinnGen study. Each genomic variant is plotted as a data point, with P-values derived using logistic regression as implemented in Regenie (v2.2.4) shown on the y-axis on a logarithmic scale and chromosomal position shown on the x-axis. All P values are two-sided and were not adjusted for multiple comparisons. The genome-wide significance threshold (P = 5×10-8) is shown with a dashed line. In each of three loci reaching genome-wide significance, the nearest protein-coding gene to the lead variant is labeled (blue = previously reported loci, red = novel locus).
To ensure that this association was not confounded by AMD—which has certain overlapping phenotypic and genotypic features with CSC—we conducted a sensitivity analysis by excluding patients with AMD symmetrically from both CSC patients and controls and observed that the association of rs113791087 with CSC remained consistent (1092 patients and 485,392 controls, OR = 2.95 [1.97-4.43], P = 1.8 × 10-7) (Supplementary Datas 4–5). The effect size remained robust even after excluding 37 diagnosis codes that reflect potentially confounding causes of fluid maculopathy from patients and controls (769 patients and 452,038 controls, OR = 3.15 [1.97–5.05], P = 1.8 × 10-6) (Supplementary Datas 4–5)14.
Replication and meta-analysis of the association of PTPRB rs113791087 with CSC
We carried out replication of variants in the PTPRB locus in 3 independent studies, including newly analyzed data from the Million Veteran Program (MVP) participants of European ancestry (706 patients with CSC and 273,198 controls, Supplementary Data 1), newly analyzed data from All of Us participants of European ancestry (133 patients with CSC and 103,600 controls, Supplementary Data 1), and data from a previously reported European chronic CSC cohort (521 patients with chronic CSC and 3,577 population controls)10. Patients with AMD were excluded from all newly analyzed studies contributing to the meta-analysis and had been previously excluded from analyses of the chronic CSC cohort10. We observed consistently significant associations between rs113791087 and CSC in all studies (Fig. 2 and Supplementary Data 5) and in a cross-study meta-analysis of the locus (OR = 3.06 [95% CI 2.31-4.06], P = 7.3 × 10−15; 2452 patients with CSC and 865,767 controls) (Supplementary Fig. 4). Additionally, rs113791087 was the lead variant in its locus in the All of Us study even when considering all variants with a minor allele count > 40 uncovered by whole genome sequencing (Supplementary Fig. 5). The point estimate was higher in All Of Us (OR = 8.28 [95% CI 3.37–20.33]; 133 patients with CSC and 103,600 controls) compared with studies that included more patients with CSC, but confidence intervals were overlapping between all studies. All 133 patients with CSC in All of Us were unrelated.
The association of rs113791087 with central serous chorioretinopathy (CSC) was examined in 4 different studies. In all biobank-based studies (FinnGen, Million Veteran Program [MVP] and All of Us) included in the meta-analysis, patients with CSC were identified based on International Statistical Classification of Diseases codes, and all participants with age-related macular degeneration were excluded from patients and controls following a harmonized study protocol. In the chronic CSC (cCSC) cohort, patients were identified from ophthalmological clinics based on expert review. Odds ratios (OR) and P-values were derived using logistic regression as implemented in Regenie (v2.2.4) in FinnGen, SAIGE (v1.3.0) in MVP, and Regenie (v3.2.2) in All Of Us. Association estimates and two-sided p-values were previously derived in the European cCSC cohort using the firth bias-corrected likelihood ratio test. An inverse-weighted fixed-effects meta-analysis was also conducted to combine data from all studies. The point estimates of the odds ratios are shown with circles for the individual studies and with a square for the meta-analysis, and 95% confidence intervals (CI) are denoted with lines. For the All of Us cohort, the upper range of the confidence interval is truncated (arrow). We observed no statistically significant heterogeneity for rs113791087 in the meta-analysis (I2 = 0.48, Q-statistic = 5.8, Q-statistic P value = 0.12). All P-values are two-sided and were not adjusted for multiple comparisons.
In addition to the broad sensitivity analysis conducted in FinnGen, the granularity of ICD-10-CM and ICD-9-CM codes in MVP allowed us to examine the association of rs113791087 with dystrophies primarily involving the RPE (H35.54 and 362.76, respectively), an endpoint that includes pattern dystrophy which can pose a diagnostic challenge with respect to CSC. We observed no significant association of rs113791087 with this outcome (OR = 1.20 [0.78–1.86], P = 0.41; 2228 patients and 311,134 controls).
We also evaluated another missense variant in the PTPRB gene (rs61758735, 12:70555234:G > A, AF = 0.69%), which has previously been suggestively associated with CSC in 2 Dutch families in an exome sequencing study15. Rs113791087 is in linkage equilibrium with rs113791087 based on in-sample data from FinnGen (R2 = 0.0) and the 1000 Genomes European subpopulation reference panel (R2 = 0.0)16. In the meta-analysis of 4 studies, rs61758735 was associated with increased risk of CSC at nominal significance although with a smaller effect size than rs113791087 (OR = 1.63 [1.15–2.32], P = 0.006142).
No evidence of association between rs113791087 and circulating VE-PTP levels
We did not observe evidence of association between rs113791087 and circulating VE-PTP (the protein encoded by PTPRB) levels in FinnGen (Beta = -0.267 on the inverse rank normalized scale, SE = 0.205, P = 0.71), in a query of UK Biobank protein quantitative locus summary statistics published by Sun et al (Beta = -0.082 on the inverse rank normalized scale, SE = 0.053, P = 0.12)17, or in a Z score based meta-analysis of the two datasets (Z = -1.36, P = 0.17).
Rs113791087 is associated with retinal abnormalities on optical coherence tomography
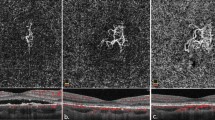
Genetic risk may manifest in subclinical differences even in the absence of diagnosed disease. To identify retinal characteristics potentially associated with the rs113791087 variant, we extracted optical coherence tomography (OCT) images from 266 participants who were heterozygous or homozygous for the G allele of rs113791087 and 442 age-matched participants with the TT allele in UK Biobank (UKB). These images were graded by 3 ophthalmologists blinded to genotype status. We observed a 56% relative increase in the prevalence of retinal abnormalities at the level of the RPE in participants with the GT or GG genotype compared to participants with the TT genotype (35.3% vs 22.6%, P = 0.00078 corrected for age, sex, examiner and first 10 PCs) (Fig. 3 and Supplementary Data 7).
Optical coherence tomography (OCT) images of 266 participants with the rs113791087 G allele (GG or GT genotype) and 442 age-matched participants lacking the G allele (TT genotype) were obtained from UKB. Images were independently evaluated by 3 retina specialists who were blinded to genotype. Each grader was tasked with identifying and categorizing retinal pigment epithelium (RPE) abnormalities according to the following categories: rare drusen (1−5), drusen (>5), pattern dystrophy, pigment epithelial detachment (PED) or nonspecific RPE irregularity, and other rare findings (subretinal fluid, pachychoroid pigment epitheliopathy, atrophy, intraretinal fluid, or evidence of central serous chorioretinopathy). Panel a depicts a B-scan superior to the fovea showing an area of RPE irregularity and pigment migration (arrowhead) from a randomly selected participant, who had the TT genotype. Panel b depicts a B-scan inferior to the fovea showing a second area of RPE irregularity and pigment migration (arrowhead) in the same participant. Panel c shows the prevalence of abnormalities (denoted with circles) for participants with the GG or GT genotype (red) and TT genotype (blue). Statistical significance was evaluated using logistic regression, including age, sex, examiner and the first 10 PCs as covariates. The P-value is two-sided and not adjusted for multiple comparisons. Binomial proportion 95% confidence intervals were calculated using the Agresti-Coull method. Optical coherence tomography images are reproduced by kind permission from UK Biobank ©.
Co-associations of PTPRB rs13791087 with varicose veins and glaucoma revealed by a phenome-wide association study in FinnGen
We conducted a phenome-wide association study of 2,469 phenotypes in FinnGen to identify potential pleiotropic effects of the PTPRB rs113791087 variant. We observed a genome-wide significant association with varicose veins of the lower extremity (OR = 1.39 [1.25–1.54], P = 3.1 × 10-10, 38,467 patients and 432,223 controls) (Fig. 4 and Supplementary Data 7). Among the most significant suggestive-level associations, we noted an increased risk of venous thromboembolism (OR = 1.29 [1.15–1.45], P = 1.8 × 10-5, 26,333 patients and 474,015 controls), an increased risk of pleural effusion (OR = 1.55 [1.24–1.94], P = 9.5 × 10-5, 6136 patients and 479,655 controls), and a reduced risk of glaucoma (OR = 0.79 [0.69–0.90], P = 5.1 × 10−4, 26,591 cases and 23,483 controls). Locus plots for these traits, including the PTPRB region are depicted in Supplementary Fig. 6–10. Results from fine mapping with SuSiE for the outcome of varicose veins are given in Supplementary Data 8; no credible sets were observed for the suggestive-level traits, possibly reflecting limited statistical power.
To identify potential pleiotropic associations of the rs113791087 variant, a phenome-wide association study was conducted including 2,469 phenotypes as outcomes using logistic regression as implemented in Regenie (v2.2.4). All P values are two-sided and were not adjusted for multiple comparisons. Data are shown for all phenotypes that were at least nominally associated (P < 0.05) with rs113791087. The negative common logarithm of each P value is shown on the y-axis. The genome-wide significance threshold (P = 5×10-8) is shown with a dashed line. The direction of association with the risk of each disease is denoted by symbols (arrow up = increased risk, arrow down = decreased risk).
In contrast with CSC and glaucoma, rs113791087 was not significantly associated with common eye diseases including wet AMD (OR = 1.05 [0.81–1.35], P = 0.72), dry AMD (OR = 1.03 [0.82–1.29], P = 0.79), or diabetic retinopathy (OR = 0.98 [0.82–1.18], P = 0.59). Because VE-PTP has been investigated as a specific target for diabetic macular edema18, we conducted further association analyses limited to participants with type 1 or type 2 diabetes, and observed no significant associations of rs113791087 with diabetic retinopathy among type 1 (OR = 2.14 [95% CI 0.88–5.20], P = 0.092; 2712 patients and 1116 controls) or type 2 diabetic patients (OR = 0.88 [95% CI 0.66–1.16], P = 0.36; 5443 patients and 77,435 controls), or with diabetic maculopathy among type 1 (OR = 1.40 [95% CI 0.53–3.71], P = 0.50; 715 patients and 3113 controls) or type 2 diabetic patients (OR = 0.79 [95% CI 0.49–1.29], P = 0.35; 1724 patients and 81,154 controls).
To better understand the co-association of rs113791087 with CSC and varicose veins of the lower extremity, we evaluated the association of these diseases in the general FinnGen population. We observed no difference in the risk of CSC among all participants with varicose veins (OR = 0.99 [95% CI 0.98–1.01], P = 0.43), suggesting that the shared association of rs113791087 with both outcomes may be reflective of true causal associations with both phenotypes rather than simple population-level correlation between the phenotypes.
The clinical manifestations of varicose veins of the lower extremity are broad-ranging and include, among others, asymptomatic venous dilation, edema, and ulceration. In analyses of ICD code-based varicose vein subtypes in FinnGen, we observed that rs113791087 was associated with uncomplicated varicose veins of the lower extremity (OR = 1.39 [1.25–1.53], P = 4.3×10-10; 38,025 patients and 455,679 controls), but not with ulcerated varicose veins of the lower extremity (OR = 0.93 [0.68–1.28], P = 0.67; 4075 patients and 455,679 controls).
Given that single-variant PheWAS cannot reliably differentiate between causal associations and confounding due to linkage disequilibrium with neighboring variants, we formally evaluated the probability that rs113791087 is the causal variant for each top trait from the FinnGen PheWAS using Coloc adapted Phenome-wide Scan (CoPheScan, Supplementary Methods) with in-sample linkage disequilibrium (LD) information in FinnGen19. CoPheScan assigned a high posterior probability (PP.Hc > 0.99) for rs113791087 being the causal variant for most top traits (glaucoma, venous thromboembolism, diseases of veins, lymphatic vessels and lymph nodes not elsewhere classified, and pleural effusion). CoPheScan assigned a lower probability for rs113791087 being causal for the outcome of any varicose veins (PP.Hc = 0.078) (Supplementary Data 9), but a high probability for rs113791087 being causal for the more specific outcome of uncomplicated varicose veins (PP.Hc > 0.99).
Cross-study meta-analyses of most significant disease associations for rs13791087
We evaluated the robustness of associations between rs113791087 and the top 5 distinct diseases from the FinnGen phenome-wide association study by conducting single-variant meta-analyses including data from FinnGen, MVP, UKB, and All of Us (Fig. 5 and Supplementary Datas 10-14). In this meta-analysis, rs113791087 was associated with an increased risk of uncomplicated varicose veins (OR = 1.31 [95% CI 1.21–1.42], P = 2.3×10−11; 62,621 patients and 1,341,326 controls) (Fig. 5 and Supplementary Data 14). We also observed an association between rs113791087 and a reduced risk of glaucoma at genome-wide significance (OR = 0.82 [95% CI 0.76–0.88], P = 6.9×10−9; 125,075 patients and 1,290,261 controls); the effect estimate was concordant for primary open-angle glaucoma (OR = 0.82 [95% CI 0.72–0.93], P = 0.0014; 35557 patients and 1366253 controls). Consistent with this observation, rs113791087 was also associated with 0.51 mmHg [95% CI 0.26–0.76 mmHg] lower mean intraocular pressure (P = 5.2×10-5) among 77,449 UKB participants.
The distinct diseases most significantly associated with rs113791087 in the phenome-wide association study of rs113791087 in FinnGen were carried forward for multi-study meta-analyses, including available data from FinnGen, UK Biobank (UKB), Million Veteran Program (MVP), and All of Us (AoU). Pulmonary embolism and deep vein thrombosis were examined separately as the major subtypes of venous thromboembolism, and primary open-angle glaucoma was examined separately as a major subtype of glaucoma. Odds ratios (OR) and P values were derived using logistic regression as implemented in Regenie (v2.2.4) in FinnGen, SAIGE (v1.3.0) in MVP, and Regenie (v3.2.2) in All Of Us. Association results from contributing studies were combined in an inverse variance-weighted meta-analysis. Point estimates of the odds ratios are shown with circles for each phenotype, and 95% confidence intervals (CI) are denoted with lines. All P-values are two-sided and were not adjusted for multiple comparisons.
In addition to the genome-wide significant associations, in the cross-study meta-analyses we observed a suggestive association between rs113791087 and an increased risk of venous thromboembolism (OR = 1.21 [95% 1.12-1.32], P = 4.6×10-6; 59887 patients and 1237762 controls). Association estimates for the constituent endpoints of pulmonary embolism (OR = 1.24 [95% CI 1.12–1.38], P = 2.9×10-5; 36456 patients and 1395022 controls) and deep vein thrombosis (OR = 1.15 [95% CI 1.00–1.31], P = 0.044; 23316 patients and 894706 controls) were directionally concordant.
Expression of PTPRB in ocular tissues and cultured choroidal endothelial cells
Given the broad spectrum of traits associated with rs113791087, we asked whether PTPRB is expressed in specific ocular tissues with plausible local disease relevance—as opposed to reflecting possible systemic effects on multiple outcomes. Using publicly available ocular single-cell RNA sequencing datasets, we observed pronounced expression of PTPRB in the endothelial cells of capillaries, arteries and veins, both in an integrated multi-tissue dataset of choroid, RPE and retina and in a choroid-specific dataset (Supplementary Fig. 7)20,21,22,23. PTPRB expression was also in the top 10% of genes (1,068/13,187) ranked by bulk expression in cultured choroidal endothelial cells from cadaveric human donors (Supplementary Fig. 11)24.
Predicted structural effect of rs113791087 on the interaction of VE-PTP with VE-Cadherin
Rs113791097 is an ILE → LEU change that lies at position 1272 in VE-PTP, the protein encoded by PTPRB. VE-PTP is a transmembrane protein that has an extracellular domain (where the variant lies) and an intracellular catalytic domain. The intracellular domain has been crystalized, while the extracellular domain has not been crystalized due to its inherently disordered nature. While the intracellular catalytic domain has been shown to be important for dephosphorylation of the Tie-2 receptor, the extracellular domain has been shown to be important in binding and stabilization of VE-Cadherin, specifically via reversal of VEGFR-2 mediated tyrosine phosphorylation of VE-Cadherin which leads to increased vascular permeability25. In fact, Nawroth et al. showed that the intracellular catalytic domain is not needed for VE-PTP to bind VE-Cadherin and reduce the permeability of endothelial cells. Though the extracellular domain has not been crystallized, we investigated whether an in silico structural prediction of the extracellular domain surrounding the VE-PTP Ile1272 amino acid is predicted to bind VE-Cadherin using AlphaFold 326. We found that the amino acid in question lies in a fibronectin type III-like domain and faces VE-Cadherin (Supplementary Fig. 12). We ran structure-based network analysis (SBNA, a published method shown to correctly assign pathogenicity to coding variants based on predicted structural effect27,28,29) to test whether Ile1272Leu in this configuration is predicted to have a pathogenic effect. The SBNA scores were in the range of pathogenicity (average SBNA score 2.83, range 1.7—4.8, based on five AlphaFold 3 models; SBNA scores for pathogenic variants in Hauser et al. had a median of 0.947, with higher being more pathogenic29).
Disease associations of predicted rare loss-of-function variants in PTPRB
To identify other genetic variation in PTPRB that might be associated with ocular or vascular systemic diseases, and to inform whether rs113791087 might act in a loss-of-function or gain-of-function manner, we identified participants who had rare pLOF PTPRB variants in UKB (n = 120 participants with a pLOF variant) and All of Us (n = 75 participants with a pLOF variant). While statistical power was limited, among traits highlighted by the rs113791087 phenome-wide association study, participants with a PTPRB pLOF variant had increased risk of CSC (OR = 17.09 [95% CI 1.93–63.49, P = 0.018), venous thromboembolism (OR = 2.22 [95% CI 1.29–3.83], P = 0.0039), pulmonary embolism (OR = 2.90 [95% 1.51–5.57], P = 0.0014), and deep vein thrombosis (OR = 2.15 [95% CI 1.08–4.27], P = 0.029) (Fig. 6 and Supplementary Data 15).
The risk of ocular and vascular diseases of interest was evaluated for participants with a predicted loss-of-function (pLOF) variant in PTPRB in All of Us and UK Biobank using logistic regression among unrelated individuals. Due to lack of ophthalmological outpatient clinic data in UK Biobank participants, patients with central serous chorioretinopathy were only evaluated in All of Us. Examined diseases were selected based on the most significant associations observed for rs113791087 in FinnGen. Pulmonary embolism and deep vein thrombosis were examined separately as the major subtypes of venous thromboembolism, and primary open-angle glaucoma was examined separately as a major subtype of glaucoma. Association results from UK Biobank and All of Us were combined in an inverse variance-weighted meta-analysis. Point estimates of the odds ratios (OR) are shown with circles for each phenotype, and 95% confidence intervals (CI) are denoted with lines. All P-values are two-sided and were not adjusted for multiple comparisons.
Discussion
Despite progress in characterizing common variant genetic risk loci for CSC, pointing to specific genes and mechanisms that explain pathophysiology remains challenging due to the inherent hurdles faced when interpreting noncoding lead variants in GWAS12. In contrast, here we identified a low-frequency missense variant (rs113791087) in PTPRB, the gene encoding VE-PTP, that is associated with a markedly increased risk of CSC (OR 3.06) and is prioritized as the likely causal variant by statistical fine-mapping. Unexpectedly, rs113791087 was also associated with an increased risk of varicose veins and with a reduced risk of glaucoma. These previously uncharacterized genetic associations point directly to VE-PTP and demonstrate its relevance in ocular and systemic vascular diseases, support a role for vascular dysfunction in CSC, and may eventually inform therapeutic development.
VE-PTP is predominantly expressed as a membrane-bound protein in venous and arterial vascular endothelial cells where it functions as an important regulator of angiogenesis, vascular integrity, and vascular permeability25,30,31. Among its binding partners, the actions of VE-PTP via Tie-2 and VE-cadherin are best characterized. Through its intracellular catalytic domain, VE-PTP acts as a dephosphorylating inhibitor of the tyrosine kinase receptor Tie-2, an essential regulator of angiogenesis and vascular maintenance. VE-PTP additionally acts as a supporting regulator of VE-cadherin—a key adhesion molecule of adherens junctions between vascular endothelial cells25,31,32—potentially through both dephosphorylation and direct non-enzymatic transmembrane binding. We hypothesize that the effect of rs113791087 is via interference with extracellular binding of VE-PTP to VE-Cadherin.
We are not aware of previous genome-wide significant associations between PTPRB variants and CSC, varicose veins, or glaucoma. Previously, a suggestive association with CSC—but not with varicose veins or glaucoma—has been reported for a different missense variant (rs61758735) in PTPRB based on an exome sequencing study of two families, and common intronic variants in the gene encoding the related VE-cadherin protein have been suggestively linked with CSC in patient cohorts15,33, but none of these reached genome-wide significance. An exome-wide significant association between rare genetic variants in PTPRB in aggregate and intraocular pressure has also been identified among UKB participants, although no direction of effect was reported34. In general, the role of PTPRB in human disease has not been well characterized, potentially reflecting intolerance to pLOF variation in humans (consistent with embryonic lethality observed with deletion of VE-PTP in mice) and a relative scarcity of common variant signals in the region30,31,35. Low-frequency variants such as rs113791087 may thus represent unique genetic instruments for assessing the associations of VE-PTP with clinical outcomes.
This genetic finding has illuminated the Tie-2 and VE-Cadherin pathways as important in CSC, an area of study that was not under investigation previously but that may be important for therapeutic advancement. At least partial relevance of Tie-2 signaling for glaucoma and venous dysfunction is suggested by the earlier identification of loss-of-function variants in the gene encoding Tie-2 in patients with primary congenital glaucoma36 and activating variants in patients with familial venous malformations with dilated vascular channels37,38. In mice, genetic insufficiency of Angiopoietin-1—a ligand of Tie-2—can inhibit normal formation of Shlemm’s canal and raise intraocular pressure39, genetic insufficiency of Tie-2 during embryogenesis may arrest venous development, and postnatal deletion of Tie-2 may lead to retinal venous degeneration with hemangioma-like vascular tufts40. In CSC, support for the potential benefit of increasing Angiopoietin-1 or Tie-2 activity comes from previous observations that patients with both acute and chronic forms of CSC have low circulating levels of Angiopoietin-1, the primary agonist of Tie-241. We note that following the discovery of this variant and the illumination of the importance of these pathways in CSC, we published a case series of patients with chronic CSC who were treated with intravitreal faricimab, an Ang-2 blocker (and thus a Tie-2 activator) that is proven to be safe and is FDA-approved for several other vascular eye diseases but has not been tested in CSC42. 14/16 treated eyes were found to have reduced disease activity in response to the medication, providing additional evidence that this genetic finding is highly relevant. Further randomized controlled trials are needed, however.
The underlying cause of CSC is unknown, but an analogy between CSC and varicose veins has been drawn based on findings of delayed choroidal filling, venous dilation, and vascular hyperpermeability in choroidal angiography of patients with CSC, as highlighted by Spaide et al.8. Statistical evidence exists for an overall enrichment of genes expressed in choroidal vascular endothelial cells among common variant CSC risk loci discovered to date12. Corticosteroid exposure—one of the classical triggers for CSC—may induce downregulation of VE-cadherin33, and VE-PTP may in general modulate vascular permeability in the context of increased inflammatory mediators43,44,45. Together with the identification of a CSC risk-increasing variant in VE-PTP in this study, these converging findings provide support for a primary role of choroidal vascular incompetence in CSC. In contrast, other features that are commonly observed in patients with CSC, such as disruption at the level of the RPE, may represent downstream sequelae of vascular dysfunction46.
In addition to the potential of this pathway in CSC, VE-PTP is also an existing investigational treatment target for more common ocular diseases. Angiopoietin/Tie-signalling is implicated in the pathogenesis of glaucoma36,39,47, and recent phase II clinical studies have evaluated razuprotafib (AKB-9778), an inhibitor of VE-PTP, as a novel treatment for glaucoma and diabetic macular edema, delivered through ophthalmic drops and subcutaneous injections, respectively18,48,49,50. The current findings and those of a recent exome sequencing study of UKB participants provide genetic support for VE-PTP as a therapeutic target for glaucoma34. The 18% reduction in the odds of glaucoma per minor allele of rs113791087 is greater than the effect estimate for any coding lead variant in a recent glaucoma GWAS meta-analysis and comparable with effect estimates reported for rare variants in ANGPTL751,52, another recent genetically motivated therapeutic target. However, we note that the discordant effect directions for CSC and glaucoma warrant further study and potential caution in therapeutic design to avoid unintended chorioretinal adverse effects with the use of VE-PTP inhibitors.
Lastly, somatic mutations in VE-PTP are implicated in angiosarcoma53, and inhibition of VE-PTP has been proposed as a potential treatment for selected high-grade cancers54,55,56. The rs113791087 variant has not to our knowledge been implicated as a somatic cancer driver mutation57. Germline genetic variants are generally not ideal instruments for complex oncological outcomes, and the lack of significant associations of rs113791087 with cancer endpoints in this study does not constitute strong evidence against the potential usefulness of VE-PTP inhibition in cancer treatment. However, the increased risk of varicose veins and venous thromboembolism in participants with rs113791087 raises concern for unintended venous dysfunction from systemic VE-PTP inhibition, particularly as many patients with cancer are already at high risk of venous thromboembolism58.
Our findings should be interpreted in the context of study design. First, the ascertainment of CSC patients in most of the study datasets was based on ICD-10 codes. Reassuringly, significant associations for rs113791087 across different healthcare systems and in an outpatient clinic cohort based on expert clinician review support a true association with CSC. Second, while directionally concordant associations of PTPRB pLOF variants with CSC suggest that rs113791087 may act via a loss-of-function mechanism, the functional impact of rs113791087 is not known, and mechanistic studies are needed to understand its potential effects on the expression and activity of VE-PTP in the choroid. Third, it is uncertain whether the mechanisms that underlie the observed disease associations are limited to early development or have continued relevance during later growth or adulthood. Fourth, analyses of rs113791087 were limited to patients of genetically inferred European ancestry due to the low frequency of the variant in other populations; however, we included all participants from UKB and All of Us when assessing rare pLOF variants.
In summary, we identified a low-frequency missense variant in the gene encoding VE-PTP that was associated with a significantly increased risk of CSC and varicose veins and with a reduced risk of glaucoma. Our findings provide support for a central role of venous dysfunction in CSC and implicate the interconnected vascular endothelial function regulators VE-PTP, Tie-2 and VE-cadherin as potential therapeutic targets in diverse ocular and systemic vascular diseases.
Methods
Study design
FinnGen is a public-private partnership research project that combines genotype data from newly collected and legacy samples administered by Finnish biobanks (https://www.finngen.fi/en) to provide novel insight into human diseases. This study includes genotype data from 500,348 individuals from FinnGen Data Freeze 12. The data were linked by unique national personal identification numbers to the national hospital discharge registry (available from 1968) and the specialist outpatient registry (1998–).
The Million Veteran Program is a population-scale biobank within the Department of Veterans Affairs. Voluntary enrollment of veterans receiving care in the VA began in 201159. All samples were scanned on the MVP 1.0 Axiom array (ThermoFisher); details on the design and QC are provided elsewhere60. Samples were phased using SHAPEIT4 and imputed to the TOPMed reference panel (version r2; N = 97,256 samples) using Minimac4. Phenotypes were based on electronic health record data with follow-up through the end of 2022.
The All of Us Research Program opened for enrollment in May 2018 and plans to enroll at least 1 million persons in the United States, collecting electronic health record data and biospecimens to advance the prevention and treatment of diseases61. In the current project, we used genetic data from 245,394 short-read whole genome-sequencing samples in the v7 release (Supplementary Methods), and excluded participants who did not have any electronic health record data.
Details for the European chronic CSC cohort have been reported previously10. For the genomic study, 521 European patients were recruited from the outpatient clinics of the Radboud University Medical Center (Netherlands), University Hospital of Cologne (Germany), and Leiden University Medical Center (Netherlands). Controls included 3,577 participants in the Nijmegen Biomedical Study. The patients with chronic CSC had subretinal fluid in at least one eye, retinal pigment epithelium irregularities with characteristic leakage on fluorescein angiography and corresponding hyperfluorescence on ICG.
UK Biobank (UKB) is a deeply phenotyped and genotyped prospective population-level cohort, which recruited approximately 500,000 participants aged 40–69 in the UK between 2006–201062. UKB participants were genotyped on the Affymetrix Applied Biosystems UK BiLEVE Axiom Array and the Affymetrix Applied Biosystems UKB Axiom Array. Sample and variant QC are described in detail by Bycroft et al. 62. rs113791087 was directly genotyped with high-quality statistics (missingness rate of 0.00198). PCA was performed using fastPCA as described and the first 10 PCs were downloaded and used as covariates in further analysis62. OCT imaging was performed for a subset of UKB participants63. For this study, UKB data was accessed under applications #50211 and #17488.
More details for genotyping, imputation and sequencing in the different studies are described in the Supplementary Methods.
Ethics statement
Study subjects in FinnGen provided informed consent for biobank research, based on the Finnish Biobank Act. Alternatively, separate research cohorts, collected between the Finnish Biobank Act coming into effect (in September 2013) and the start of FinnGen (August 2017), were collected as described in the Supplementary Methods. No compensation was provided for participating in the study.
The MVP024 study protocol was approved by the Veterans Affairs (VA) central Institutional Review Board (IRB). MVP participants provided written informed consent. No compensation was provided for participating in the study.
The general All of Us protocol has been approved by the All of Us Institutional Review Board. Use of All of Us data for this study was approved under a data use agreement between the Massachusetts General Hospital and the All of Us research program. Study participants provided written informed consent. No compensation was provided for participating in the collection of the data used in this study.
The study of European patients with chronic CSC was carried out in accordance with the tenets of the Declaration of Helsinki and was approved by the local ethics committees of the Radboudumc, Leiden University Medical Center, and University Hospital of Cologne. Written informed consent was obtained for all participants.
The UKB OCT substudy was approved by the North West Multi-centre Research Ethics Committee in accordance with the principles of the Declaration of Helsinki. Written, informed consent was obtained for all UKB participants.
Phenotype ascertainment
For the discovery GWAS, we identified patients with CSC based on the presence of at least one instance of the Finnish version of the International Classification of Diseases 10th revision (ICD-10) diagnosis code H35.7. Additionally, controls with retinal or choroidal disorders were excluded using the ICD-10 codes H30-H36, the ICD-9 codes 3610-3611, 3613, 3618-3619, 3621, 3622X, 3623A-3623D, 3624 A, 3625, 3626, 3627, 3628 A, 3628 A, 3628X, and 3630-3639, and the ICD−8 codes 3610-3611, 3626, 365, 3670, 376, 37701-37703, 37705, 37709-37712, 37798, and 3785.
We conducted additional sensitivity analyses to evaluate bias due to potential confounders. In these analyses, instead of control-specific exclusion criteria, we applied the following exclusion criteria to both patients and controls: 1) at least one instance of an ICD-10 code corresponding to age-related macular degeneration (H35.30 or H35.31), and 2) at least one instance of any ICD-10 code corresponding to a larger set of potentially confounding causes of fluid maculopathy (Supplementary Data 3).
In MVP and All of Us, patients with CSC were identified based on at least one instance of the ICD-10-CM code H35.71* or ICD−9-CM code 362.41, and all participants with AMD (ICD-10: H35.1*, H35.2*, H35.3*; ICD-9: 362.5, 362.51, 362.52) were excluded. Case definitions for the European chronic CSC cohort have been described previously10. Ages of individuals in All of Us were calculated at 2022/7/1 or date of death.
Genome-wide association study and regional meta-analysis of CSC
All FinnGen GWAS were conducted using Regenie v 2.2.464, with sex, age at death or end of follow-up, principal components (PCs) 1–10, genotyping array, and genotyping batch as fixed-effect covariates. An approximate Firth correction was used for variants reaching nominal significance (P < 0.01) in initial tests, and standard errors were computed based on the Firth beta estimate and Firth P-value. Fine mapping of 95% credible sets in FinnGen was performed using 3 megabase (Mb) windows around each lead variant (1.5 Mb in both directions) in genome-wide significant loci (lead variant P < 5×10-8) using in-sample dosage LD computed with LDstore 265, and the Sum of Single Effects (SuSiE) model with the maximum number of causal variants in a locus (L) set to 1066.
Genomic analysis of CSC in MVP was performed using SAIGE v1.3.067 on the set of samples classified as European ancestry using the HARE (Harmonized Ancestry and Race/Ethnicity) method68. We used leave-one-chromosome-out (LOCO) model fitting and enabled Firth effect size estimation for variants with P < 0.05. Sex, age at enrollment, mean-centered age-squared, and the first ten within-ancestry PCs were included as covariates.
Genomic analysis of CSC in All of Us was performed using Regenie v3.2.2 with age at death or end of follow-up, (age at death or end of follow-up)2, sex, and PCs 1−5 and 15 (based on association with CSC at p < 0.05 in a separate association test).
To evaluate whether rs113791087 was the lead variant in its locus on chromosome 12 even when combining data from 4 different studies, we performed an inverse variance weighted meta-analysis of CSC for all genomic variants in the region of rs113791087 with GWAMA (v2.2.2). LocusZoom was used to generate locus plots for the meta-analysis using the European ancestry reference panel69.
More detailed methods for participating cohorts are described in the Supplementary Information.
Analysis of Optical Coherence Tomography images in UKB
Deidentified OCT images of patients with the rs113791087 G allele (GG or GT) along with age-matched controls lacking the G allele (TT) were obtained from UKB (642 GG/GT OCTs, 1,058 TT OCTs; 10 age-matched participants with the TT genotype were pulled for every 1 participant with the GT or GG genotype, of which ~16% had an OCT)63. Repeat OCTs from the same patient were removed. Images were randomly chosen and independently evaluated by 3 retina specialists who were blinded to genotype. Because the scope of manual OCT review has practical limitations, only the left eye of each patient underwent assessment and 708 OCTs were reviewed. Each grader was tasked with identifying and categorizing various types of RPE abnormalities (drusen, pattern dystrophy, pigment epithelial detachment or nonspecific retinal pigment epithelium irregularity, subretinal fluid, pachychoroid pigment epitheliopathy, atrophy, intraretinal fluid and/or evidence of CSC). Statistical significance was evaluated using logistic regression in R, including age, sex, examiner and the first 10 PCs as covariates.
Phenome-wide association study, replication and meta-analyses
We evaluated the association of rs113791087 with 2,469 traits in FinnGen using Regenie v2.2.4 similarly to the association analyses for CSC64. Phenotype definitions and the characteristics of study participants for these traits are publicly available on https://risteys.finregistry.fi/.
For top traits identified in this phenome-wide association study, we pursued single-variant replication and meta-analyses including data from FinnGen, UKB, MVP and All of Us. Where possible, custom disease definitions to match these outcomes were created in UKB (Supplementary Data 11), All of Us (Supplementary Data 12), and Million Veteran Program (Supplementary Methods).
We evaluated the association of rs113791087 with each outcome in unrelated participants of genetically inferred European ancestry (excluding second-degree or closer relatives in UK Biobank and Million Veteran programs, and based on a kinship score cutoff of 0.1 in All of Us) with Firth logistic regression. Covariates in UK Biobank included genotyped sex, age, the first 10 PCs and genotyping array. Covariates in All of Us included age, age squared, sex, first 5 PCs, and additional outcome-related PCs (P < 0.05 in separated models). Covariates for analyses of uncomplicated varicose veins in MVP included age, age2, sex and the first 10 PCs. For other phenotypes in MVP, we used European-ancestry (HARE) summary statistics generated by the genome-wide PheWAS (gwPheWAS) project70. Briefly, outcomes were derived from phecodes following standard definitions71, surveys distributed to all MVP enrollees59, and clinical laboratory and vital signs measurements. A linear or logistic regression GWAS was performed on each phenotype in a modified version of SAIGE using sex, age, age2, and the first 10 PCs as covariates.
Results from different studies were combined in an inverse variance weighted meta-analysis as implemented in the metagen() function of the meta package (v6.5-0) in R (v4.2.0).
We additionally evaluated the association of rs113791087 with the mean of the left and right eye corneal-compensated intraocular pressure (IOP) measurements in UKB participants. Participants with a history of surgery for glaucoma were excluded and IOP readings less than 5 or greater than 60 were excluded. Linear regression analyses were adjusted by genotyped sex, age, the first 10 PCs and genotyping array.
Lastly, we identified rare predicted loss-of-function (pLOF) variants in PTPRB among UKB and All of Us participants by using the LOFTEE algorithm, excluding low-confidence or flagged pLOF variants and variants present at a frequency above 0.1% in either biobank or across gnomAD v2 super-populations35. Associations with disease outcomes were evaluated with Firth logistic regression using similar analysis designs as for rs113791087, except all individuals were included regardless of genetically inferred ancestry.
Additional epidemiological analyses
The association of CSC and varicose veins was evaluated in all available FinnGen participants using logistic regression as implemented in the glm function in R (v4.3.2) with CSC as the outcome and varicose veins, sex, and age at death or end of follow-up as independent predictors.
Analyses of RNA expression
We evaluated the expression of VE-PTP (PTPRB) using two single-cell sequencing datasets made publicly available by Voigt et al: an integrated dataset of studies between 2019–2020 (93,526 total cells from the retina, RPE and choroid) and a choroidal-specific dataset (37,070 cells from the choroid of 6 adults and 2 infants)20,21,22,23. Additionally, we evaluated the relative absolute bulk RNA expression level of VE-PTP (PTPRB) compared with all other detectable genes in a dataset of cultured choroidal endothelial cells from 10 human postmortem donors (five males and five females), protocols of which have been described previously24.
Structural protein analysis
The amino acid sequences for VE-cadherin and the Fibronectin type III-like domains 14, 15, and 16 of VE-PTP were used as inputs in AlphaFold 326. AlphaFold 3 generated 5 structural predictions of the binding between the two protein structures. Structure-based network analysis (SBNA) was performed on each of these structural predictions individually as previously described27,28,29. Briefly, atomic-level interactions were analyzed computationally to evaluate the extent to which altering a particular amino acid within the protein structure could affect the overall structure of the protein. The SBNA scores corresponding to these effect estimates were considered for Ile1272 in VE-PTP across the 5 structural prediction models. Visualizations of the structural prediction models were generated using PyMOL72.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The source data from FinnGen, UK Biobank and All of Us are available under restricted access due to the sensitive nature of the individual-level genotype and phenotype information. Individual-level genotypes and register data from FinnGen participants can be accessed by approved researchers via the Fingenious portal (https://site.fingenious.fi/en/) hosted by the Finnish Biobank Cooperative FinBB (https://finbb.fi/en/). Access to individual-level UKB data may be requested by researchers in academic, commercial, and charitable organizations (https://www.ukbiobank.ac.uk). This study used data from the All of Us Research Program’s Controlled Tier Dataset v7, available to authorized users on the Researcher Workbench (https://www.researchallofus.org). The raw Million Veteran Program data are protected and are not available due to data privacy laws. Summary-level association data from MVP used in this study are available through dbGaP, under accession code phs001672.v11.p1 [https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs001672.v11.p1].
Code availability
No custom software or analysis tools were created for this study; all analyses used existing software and tools as described in the Methods.
References
Kitzmann, A. S., Pulido, J. S., Diehl, N. N., Hodge, D. O. & Burke, J. P. The Incidence of Central Serous Chorioretinopathy in Olmsted County, Minnesota, 1980–2002. Ophthalmology 115, 169–173 (2008).
Feenstra H. M. A. et al. Central serous chorioretinopathy: An evidence-based treatment guideline. Prog Retin Eye Res. 101236 (2024).
van Rijssen, T. J. et al. Central serous chorioretinopathy: Towards an evidence-based treatment guideline. Prog. Retin Eye Res 73, 100770 (2019).
Sahoo, N. K. et al. Prevalence and Profile of Central Serous Chorioretinopathy in an Indian Cohort. Nepal J. Ophthalmol. 11, 5–10 (2019).
Montero, J. A. Ruiz-Moreno JM. Optical coherence tomography characterisation of idiopathic central serous chorioretinopathy. Br. J. Ophthalmol. 89, 562–564 (2005).
Gajdzik-Gajdecka, U. et al. Indocyanine green angiography in chronic central serous chorioretinopathy. Med Sci. Monit. 18, CR51–CR57 (2012).
Chatziralli, I. et al. Risk Factors for Central Serous Chorioretinopathy: Multivariate Approach in a Case-Control Study. Curr. Eye Res 42, 1069–1073 (2017).
Spaide, R. F. et al. Venous overload choroidopathy: A hypothetical framework for central serous chorioretinopathy and allied disorders. Prog. Retin Eye Res 86, 100973 (2022).
Piazza, G. Varicose veins. Circulation 130, 582–587 (2014).
Schellevis, R. L. et al. Role of the Complement System in Chronic Central Serous Chorioretinopathy: A Genome-Wide Association Study. JAMA Ophthalmol. 136, 1128–1136 (2018).
Hosoda, Y. et al. Genome-wide association analyses identify two susceptibility loci for pachychoroid disease central serous chorioretinopathy. Commun. Biol. 2, 468 (2019).
Rämö, J. T. et al. Overlap of genetic loci for central serous chorioretinopathy with age-related macular degeneration. JAMA Ophthalmol. 141, 449–457 (2023).
de Jong, E. K. et al. Chronic central serous chorioretinopathy is associated with genetic variants implicated in age-related macular degeneration. Ophthalmology 122, 562–570 (2015).
van Dijk, E. H. C. & Boon, C. J. F. Serous business: Delineating the broad spectrum of diseases with subretinal fluid in the macula. Prog. Retin Eye Res 84, 100955 (2021).
Schellevis, R. L. et al. Exome sequencing in families with chronic central serous chorioretinopathy. Mol. Genet Genom. Med 7, e00576 (2019).
1000 Genomes Project Consortium, Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Sun, B. B. et al. Plasma proteomic associations with genetics and health in the UK Biobank. Nature 622, 329–338 (2023).
Campochiaro, P. A. et al. Enhanced Benefit in Diabetic Macular Edema from AKB-9778 Tie2 Activation Combined with Vascular Endothelial Growth Factor Suppression. Ophthalmology 123, 1722–1730 (2016).
Manipur, I. et al. CoPheScan: phenome-wide association studies accounting for linkage disequilibrium. Nat. Commun. 15, 5862 (2024).
Voigt, A. P. et al. Single-cell transcriptomics of the human retinal pigment epithelium and choroid in health and macular degeneration. Proc. Natl Acad. Sci. USA 116, 24100–24107 (2019).
Voigt A. P. et al. Single-Cell RNA Sequencing in Human Retinal Degeneration Reveals Distinct Glial Cell Populations. Cells. https://doi.org/10.3390/cells9020438 (2020).
Voigt, A. P. et al. Bulk and single-cell gene expression analyses reveal aging human choriocapillaris has pro-inflammatory phenotype. Microvasc. Res 131, 104031 (2020).
Voigt, A. P. et al. Spectacle: An interactive resource for ocular single-cell RNA sequencing data analysis. Exp. Eye Res 200, 108204 (2020).
Brinks, J. et al. The Cortisol Response of Male and Female Choroidal Endothelial Cells: Implications for Central Serous Chorioretinopathy. J. Clin. Endocrinol. Metab. 107, 512–524 (2022).
Nawroth, R. et al. VE-PTP and VE-cadherin ectodomains interact to facilitate regulation of phosphorylation and cell contacts. EMBO J. 21, 4885–4895 (2002).
Abramson, J. et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500 (2024).
Gaiha, G. D. et al. Structural topology defines protective CD8+ T cell epitopes in the HIV proteome. Science 364, 480–484 (2019).
Nathan, A. et al. Structure-guided T cell vaccine design for SARS-CoV-2 variants and sarbecoviruses. Cell 184, 4401–4413.e10 (2021).
Hauser, B. M. et al. Structure-based network analysis predicts pathogenic variants in human proteins associated with inherited retinal disease. NPJ Genom. Med 9, 31 (2024).
Bäumer, S. et al. Vascular endothelial cell-specific phosphotyrosine phosphatase (VE-PTP) activity is required for blood vessel development. Blood 107, 4754–4762 (2006).
Dominguez, M. G. et al. Vascular endothelial tyrosine phosphatase (VE-PTP)-null mice undergo vasculogenesis but die embryonically because of defects in angiogenesis. Proc. Natl Acad. Sci. USA 104, 3243–3248 (2007).
Fachinger, G., Deutsch, U. & Risau, W. Functional interaction of vascular endothelial-protein-tyrosine phosphatase with the angiopoietin receptor Tie-2. Oncogene 18, 5948–5953 (1999).
Schubert, C. et al. Cadherin 5 is regulated by corticosteroids and associated with central serous chorioretinopathy. Hum. Mutat. 35, 859–867 (2014).
Gao, X. R., Chiariglione, M. & Arch, A. J. Whole-exome sequencing study identifies rare variants and genes associated with intraocular pressure and glaucoma. Nat. Commun. 13, 7376 (2022).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Souma, T. et al. Angiopoietin receptor TEK mutations underlie primary congenital glaucoma with variable expressivity. J. Clin. Invest 126, 2575–2587 (2016).
Vikkula, M. et al. Vascular dysmorphogenesis caused by an activating mutation in the receptor tyrosine kinase TIE2. Cell 87, 1181–1190 (1996).
Boscolo, E. et al. Rapamycin improves TIE2-mutated venous malformation in murine model and human subjects. J. Clin. Invest 125, 3491–3504 (2015).
Thomson, B. R. et al. Angiopoietin-1 is required for Schlemm’s canal development in mice and humans. J. Clin. Invest 127, 4421–4436 (2017).
Chu M., et al. Angiopoietin receptor Tie2 is required for vein specification and maintenance via regulating COUP-TFII. Elife. https://doi.org/10.7554/eLife.21032 (2016).
Karska-Basta I., et al. Imbalance in the Levels of Angiogenic Factors in Patients with Acute and Chronic Central Serous Chorioretinopathy. J. Clin. Med. Res. https://doi.org/10.3390/jcm10051087 (2021).
Rämö, J. T. et al. Targeting the Tie-2 receptor with faricimab in central serous chorioretinopathy: A case series motivated by a genetic finding. Am. J. Ophthalmol. 269, 246–254 (2024).
Shen, J. et al. Targeting VE-PTP activates TIE2 and stabilizes the ocular vasculature. J. Clin. Invest 124, 4564–4576 (2014).
Frye, M. et al. Interfering with VE-PTP stabilizes endothelial junctions in vivo via Tie-2 in the absence of VE-cadherin. J. Exp. Med 212, 2267–2287 (2015).
Braun, L. J. et al. VE-PTP inhibition stabilizes endothelial junctions by activating FGD5. EMBO Rep. 20, e47046 (2019).
Kanda, P. et al. Pathophysiology of central serous chorioretinopathy: a literature review with quality assessment. Eye 36, 941–962 (2022).
Thomson, B. R. et al. A lymphatic defect causes ocular hypertension and glaucoma in mice. J. Clin. Invest 124, 4320–4324 (2014).
Kim, J. et al. Impaired angiopoietin/Tie2 signaling compromises Schlemm’s canal integrity and induces glaucoma. J. Clin. Invest 127, 3877–3896 (2017).
Li, G. et al. A Small Molecule Inhibitor of VE-PTP Activates Tie2 in Schlemm’s Canal Increasing Outflow Facility and Reducing Intraocular Pressure. Invest Ophthalmol. Vis. Sci. 61, 12 (2020).
Brigell, M., Withers, B., Buch, A. & Peters, K. G. Tie2 Activation via VE-PTP Inhibition With Razuprotafib as an Adjunct to Latanoprost in Patients With Open Angle Glaucoma or Ocular Hypertension. Transl. Vis. Sci. Technol. 11, 7 (2022).
Tanigawa, Y. et al. Rare protein-altering variants in ANGPTL7 lower intraocular pressure and protect against glaucoma. PLoS Genet 16, e1008682 (2020).
Han, X. et al. Large-scale multitrait genome-wide association analyses identify hundreds of glaucoma risk loci. Nat. Genet 55, 1116–1125 (2023).
Behjati, S. et al. Recurrent PTPRB and PLCG1 mutations in angiosarcoma. Nat. Genet 46, 376–379 (2014).
Goel, S. et al. Effects of vascular-endothelial protein tyrosine phosphatase inhibition on breast cancer vasculature and metastatic progression. J. Natl Cancer Inst. 105, 1188–1201 (2013).
Li, G., Sachdev, U., Peters, K., Liang, X. & Lotze, M. T. The VE-PTP Inhibitor AKB−9778 Improves Antitumor Activity and Diminishes the Toxicity of Interleukin 2 (IL-2) Administration. J. Immunother. 42, 237–243 (2019).
Stanford, S. M. & Bottini, N. Targeting protein phosphatases in cancer immunotherapy and autoimmune disorders. Nat. Rev. Drug Discov. 22, 273–294 (2023).
Tate, J. G. et al. COSMIC: the Catalogue Of Somatic Mutations In Cancer. Nucleic Acids Res 47, D941–D947 (2019).
Lyman, G. H. et al. American Society of Hematology 2021 guidelines for management of venous thromboembolism: prevention and treatment in patients with cancer. Blood Adv. 5, 927–974 (2021).
Gaziano, J. M. et al. Million Veteran Program: A mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Hunter-Zinck, H. et al. Genotyping array design and data quality control in the Million Veteran Program. Am. J. Hum. Genet 106, 535–548 (2020).
All of Us Research Program Investigators, Denny, J. C. et al. The “All of Us” Research Program. N. Engl. J. Med 381, 668–676 (2019).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Keane, P. A. et al. Optical Coherence Tomography in the UK Biobank Study - Rapid Automated Analysis of Retinal Thickness for Large Population-Based Studies. PLoS One 11, e0164095 (2016).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet 53, 1097–1103 (2021).
Benner, C. et al. Prospects of Fine-Mapping Trait-Associated Genomic Regions by Using Summary Statistics from Genome-wide Association Studies. Am. J. Hum. Genet 101, 539–551 (2017).
Zou, Y., Carbonetto, P., Wang, G. & Stephens, M. Fine-mapping from summary data with the “Sum of Single Effects” model. PLoS Genet 18, e1010299 (2022).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet 50, 1335–1341 (2018).
Fang, H. et al. Harmonizing genetic ancestry and self-identified race/ethnicity in genome-wide association studies. Am. J. Hum. Genet 105, 763–772 (2019).
Boughton, A. P. et al. LocusZoom.js: interactive and embeddable visualization of genetic association study results. Bioinformatics 37, 3017–3018 (2021).
Verma, A. et al. Diversity and scale: Genetic architecture of 2068 traits in the VA Million Veteran Program. Science 385, eadj1182 (2024).
Denny, J. C. et al. PheWAS: demonstrating the feasibility of a phenome-wide scan to discover gene-disease associations. Bioinformatics 26, 1205–1210 (2010).
PyMOL [Internet]. 2020 [cited 2024 Sep 1]. Available from: http://www.pymol.org/pymol
Acknowledgements
We acknowledge funding from the National Institutes of Health and the Research to Prevent Blindness Career Development Award (K12EY016335 and 1K23EY035342) to E.J.R.. We acknowledge NIH Core Grant P30EY011373 to the Departments of Ophthalmology at Case Western Reserve University, NIH Core Grant P30EY025585 to Cleveland Clinic School of Medicine at Case Western Reserve University, and unrestricted support from Research to Prevent Blindness to the Departments of Ophthalmology at Case Western Reserve University, Cleveland Clinic School of Medicine at Case Western Reserve University. We acknowledge support from the International Retinal Research Foundation to S.K.I.. We want to acknowledge the participants and investigators of FinnGen study. The FinnGen project is funded by two grants from Business Finland (HUS 4685/31/2016 and UH 4386/31/2016) and the following industry partners: AbbVie Inc., AstraZeneca UK Ltd, Biogen MA Inc., Bristol Myers Squibb (and Celgene Corporation & Celgene International II Sàrl), Genentech Inc., Merck Sharp & Dohme LCC, Pfizer Inc., GlaxoSmithKline Intellectual Property Development Ltd., Sanofi US Services Inc., Maze Therapeutics Inc., Janssen Biotech Inc, Novartis Pharma AG, and Boehringer Ingelheim International GmbH. Following biobanks are acknowledged for delivering biobank samples to FinnGen: Auria Biobank (www.auria.fi/biopankki), THL Biobank (www.thl.fi/biobank), Helsinki Biobank (www.helsinginbiopankki.fi), Biobank Borealis of Northern Finland (https://www.ppshp.fi/Tutkimus-ja-opetus/Biopankki/Pages/Biobank-Borealis-briefly-in-English.aspx), Finnish Clinical Biobank Tampere (www.tays.fi/en-US/Research_and_development/Finnish_Clinical_Biobank_Tampere), Biobank of Eastern Finland (www.ita-suomenbiopankki.fi/en), Central Finland Biobank (www.ksshp.fi/fi-FI/Potilaalle/Biopankki), Finnish Red Cross Blood Service Biobank (www.veripalvelu.fi/verenluovutus/biopankkitoiminta), Terveystalo Biobank (www.terveystalo.com/fi/Yritystietoa/Terveystalo-Biopankki/Biopankki/) and Arctic Biobank (https://www.oulu.fi/en/university/faculties-and-units/faculty-medicine/northern-finland-birth-cohorts-and-arctic-biobank). All Finnish Biobanks are members of BBMRI.fi infrastructure (www.bbmri.fi). Finnish Biobank Cooperative -FINBB (https://finbb.fi/) is the coordinator of BBMRI-ERIC operations in Finland. We thank the U.S. veteran participants in MVP and MVP staff. This publication does not necessarily represent the views of the U.S. Department of Veterans Affairs or the United States Government. Support from the VA Office of Research & Development is acknowledged by N.S.P. (I01BX003364, I01 BX04557, IK6BX005233). We acknowledge the VA Million Veteran Program (MVP) and the VA-DOE genome-wide PheWAS core analytic team for generating the corresponding PheWAS summary statistics that were used in this manuscript. The All of Us Research Program is supported by the National Institutes of Health, Office of the Director: Regional Medical Centers: 1 OT2 OD026549; 1 OT2 OD026554; 1 OT2 OD026557; 1 OT2 OD026556; 1 OT2 OD026550; 1 OT2 OD 026552; 1 OT2 OD026553; 1 OT2 OD026548; 1 OT2 OD026551; 1 OT2 OD026555; IAA #: AOD 16037; Federally Qualified Health Centers: HHSN 263201600085U; Data and Research Center: 5 U2C OD023196; Biobank: 1 U24 OD023121; The Participant Center: U24 OD023176; Participant Technology Systems Center: 1 U24 OD023163; Communications and Engagement: 3 OT2 OD023205; 3 OT2 OD023206; and Community Partners: 1 OT2 OD025277; 3 OT2 OD025315; 1 OT2 OD025337; 1 OT2 OD025276. In addition, the All of Us Research Program would not be possible without the partnership of its participants. UK Biobank is generously supported by its founding funders the Wellcome Trust and UK Medical Research Council, as well as the British Heart Foundation, Cancer Research UK, Department of Health, Northwest Regional Development Agency and Scottish Government.
Author information
Authors and Affiliations
Consortia
Contributions
E.J.R., M.J.D., S.K.I., P.T.E, C.JF.B., J.A.T., N.S.P., J.T.R. and B.R.G. conceptualized or designed the study; J.T.R., B.R.G., L-C.W., S.J.J., X.W., SH.C., Y.L., S.H.K., and E.J.R. performed genomic or epidemiological analyses; E.J.R., P.S., and M.G.T. reviewed optical coherence tomography data; E.J.R., and B.M.H. analyzed structural protein predictions; E.H.C.vD., J.B., S.H.C., S.Y., C.J.F.B., A.P., M.J.D., C.W.H., S.K.I., S.A.A, D.P.R., J.A.T., K.K., W-C.W, M.B.G., C.L.N. and S.P. generated or collected data; E.J.R., M.J.D., S.K.I., P.T.E., C.J.F.B., N.S.P., A.P., and S.Y. acquired funding; E.J.R., M.J.D., S.K.I., P.T.E., C.JF.B., J.T.A., N.S.P. and L.S. supervised the analyses; J.T.R., E.J.R., B.R.G. and L-C.W. wrote the initial draft of the manuscript; all authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
Dr. Rossin and Dr. Rämö are named inventors on a provisional patent application that describes the secondary use of intravitreal Anti-Ang2 medications for use in the treatment of central serous chorioretinopathy. Dr. Ellinor receives sponsored research support from Bayer AG, IBM Research, Bristol Myers Squibb, Pfizer and Novo Nordisk; he has also served on advisory boards or consulted for MyoKardia and Bayer AG. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Dominic Furniss, Anand Swaroop and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Rämö, J.T., Gorman, B.R., Weng, LC. et al. Rare genetic variation in PTPRB is associated with central serous chorioretinopathy, varicose veins and glaucoma. Nat Commun 16, 4127 (2025). https://doi.org/10.1038/s41467-025-58686-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-58686-6
This article is cited by
-
Efficacy and safety of intravitreal faricimab for choroidal neovascularization secondary to central serous chorioretinopathy – a multicentre study
Graefe's Archive for Clinical and Experimental Ophthalmology (2026)