Abstract
The differentiation of post-migratory cranial neural crest cells (CNCCs) into distinct mesenchymal lineages is crucial for craniofacial development. Here we report a high-resolution spatiotemporal transcriptomic and cell-type atlas of CNCC-derived mesenchymal lineage diversification during mouse palatogenesis. We systematically defined each mesenchymal cell type by mapping their transcriptomic profiles to spatial identities. Integrative analysis of spatial transcriptomic data from E12.5 to E15.5 further revealed mesenchymal lineage establishment at or prior to initiation of palatogenesis. We also identified a heterogeneous Sox9+ mesenchymal progenitor population at the onset of palatal development, with subpopulations already activating early lineage-specific markers. In vivo lineage tracing using these early lineage-specific markers demonstrated that distinct mesenchymal populations are established as early as E10.5 to E11.5, preceding palatal development, and contribute to their respective lineages. Together, our findings reveal the comprehensive, dynamic molecular and cellular landscape of palate development and shed light on cell fate regulation during embryogenesis.
Similar content being viewed by others
Introduction
The development of the vertebrate head crucially depends on cranial neural crest cells (CNCCs), which initially diversify into neural and non-neural (ectomesenchymal) lineages in early embryonic development1. Subsequently, these ectomesenchymal cells migrate to the pharyngeal arches, contributing to the formation of various facial structures by differentiating into a variety of mesenchymal cell types, including those that form bone, cartilage, teeth, and connective tissue2,3,4,5. One major focus of research on CNCCs has been understanding how these cells are directed toward specific cell fates. While considerable advances have been made in elucidating the initial steps of these fate decisions, the mechanisms guiding post-migratory CNCC-derived ectomesenchyme in the pharyngeal arches to further differentiate into distinct mesenchymal populations remain poorly understood.
A key question in neural crest fate determination is whether individual neural crest cells (NCCs) are multipotent or predetermined to adopt specific fates before reaching their final destinations in vivo. NCCs exhibit multipotency under differentiation-inducing conditions in vitro; however, their multipotency in vivo appears to be context-dependent, with only a subset of CNCCs reported to demonstrate this capability in vivo6,7. Recent studies have also shown that although cranial and trunk NCCs can both give rise to neural lineages, the neuronal bias of trunk NCCs and the mesenchymal bias of CNCCs already emerge during delamination1. Furthermore, cell-lineage analysis using Confetti mice has revealed intrinsic heterogeneity within the early CNCC population, as indicated by CNCC derivatives forming spatially defined individual clones that contribute to shaping the early face5. However, the mechanisms by which these post-migratory CNCCs in specific regions further diversify into various mesenchymal cell types remain unclear.
In this study, we aimed to elucidate how post-migratory CNCCs fine-tune their fate and thus give rise to various cell types within specific anatomical regions. We utilized the development of the mouse palate as a model to trace the lineage specification and developmental trajectories of post-migratory CNCCs. The mouse palatal shelves are a well-defined craniofacial structure featuring an intricate architecture divided into the anterior hard palate and the posterior soft palate8. The anterior segment, which is bony, contains osteogenic mesenchymal cells, while the posterior segment, which is muscular, is enriched with perimysial cells, a CNCC-derived subpopulation crucial for supporting myogenic cells9. Understanding the development of the palatal region is also relevant to common birth defects such as cleft lip and/or palate, which adversely affect vital functions including speech, breathing, eating, and swallowing, as well as the quality of life8,10.
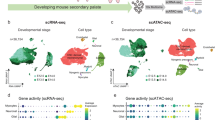
Several recent studies have demonstrated that single-cell and spatial omics are powerful tools for comprehensively analyzing cellular heterogeneity and fate regulation in the palatal mesenchyme9,11,12,13. For example, in the hard palate, Visium spatial RNA sequencing (spRNA-seq) analysis revealed dynamic activation of several late-stage osteogenic markers during osteoblast maturation12. In the soft palate region, single-cell RNA sequencing (scRNA-seq) analysis has enabled the identification of previously uncharacterized perimysial cells9. Moreover, in silico perturbation analysis identified SHOX2 and MEOX2 as key regulators of anterior and posterior palate development13, consistent with previously published in vivo findings using transgenic mouse models14,15. However, a comprehensive understanding of the mesenchymal progenitors, their lineage commitments, and subsequent cell fate diversification remains elusive. Furthermore, most current studies have limited capacity to integrate high-resolution, spatiotemporally dynamic data when analyzing the expression of multiple genes in relation to cell fate decisions.
In this study, we employed a high-resolution seqFISH-based spatial genomics platform16,17 to generate transcriptomic maps of distinct mesenchymal lineages during mouse palatogenesis. This approach facilitated the molecular identification of cell types based on their spatially resolved gene expression profiles in the maxillary region associated with palate development. We further performed integrative analysis of the spatial landscape across early and late palate development to visualize the in vivo dynamics of mesenchymal cell lineages as they approach their final anatomical locations, and uncovered early-stage heterogeneity within mesenchymal cells and progenitors for the palatal region, indicating the predetermination of cell fate before the establishment of anatomical structures. This early fate diversification of distinct mesenchymal lineages was validated in vivo using lineage tracing in transgenic mouse models, underscoring the robustness and significance of our approach. In summary, our research provides an example of how spatial transcriptomics can be utilized to gain a comprehensive spatiotemporal view of the fine-tuning of CNCC fate during later stages of craniofacial development, with important implications for more nuanced lineage-specific analyses of the etiology of congenital anomalies affecting the palate and other craniofacial structures.
Results
Identification of mesenchymal populations during palatogenesis
At the onset of palatogenesis at E12.5, post-migratory CNCCs arrive at the palatal primordium and undergo critical fate diversification to give rise to the various palatal structures before birth. To evaluate the molecular and cellular landscape of mesenchymal cell fate diversification of CNCCs in vivo, we employed scRNA-seq analysis alongside a seqFISH-based spatial genomics approach16,17 for high-resolution visualization of spatially resolved gene expression profiles (Fig. 1). To systematically identify specific marker genes for the various mesenchymal lineages, we first integrated scRNA-seq data across all stages of palatal development using Seurat integration analysis18. This dataset (79,151 cells in total) included E12.5 (12,027 cells), E13.5 (16,740 cells), E14.5 (15,550 cells), E15.5 (15,934 cells), and E18.5 (18,900 cells), with three biological replicates per stage (Fig. S1a–f).
a Schematic drawings of mouse palate development, showing scRNA-seq datasets collected from the mouse embryonic palate at stages E12.5, E13.5, E14.5, E15.5, and E18.5 (3 biological replicates per stage). b Schematic drawings of mouse embryonic head development, highlighting stages E12.5, E13.5, and E15.5, when spatial genomics datasets were collected using histological sections of anterior and posterior regions from these stages. Orange and green lines indicate the anterior and posterior head sections corresponding to the hard and soft palate regions, respectively. Abbreviation: palatal shelves (p).
We next performed differential gene expression analysis for each cluster to identify its associated cell type. Based on their marker gene expression, we distinguished all the cell types in the developing palate, including the non-mesenchymal cell clusters: epithelial (Krt14⁺/Krt5⁺/Krt15⁺, clusters 8, 9, 12, 24), endothelial (Cdh5⁺/Cldn5⁺/Sox7⁺, cluster 11), myogenic (Myod1⁺/Myog⁺/Pax7⁺, clusters 18, 25), neuronal (Tubb3⁺/Stmn2⁺/Myt1l⁺, cluster 17), glial (Sox10⁺/Plp1⁺/Foxd3⁺, cluster 16), erythroid (Hba-x⁺/Hba-a1⁺/Hba-a2⁺/Hbb-bt⁺, clusters 13, 20), myeloid (Lyz2⁺/C1qb⁺/C1qc⁺, clusters 14, 19), and mesenchymal marker-expressing cell clusters (0, 1, 2, 3, 4, 5, 6, 7, 10, 15, 21, 22, 23) identified by Prrx1⁺/Twist1⁺/Barx1⁺ expression9,13 (Figure S1g-i). To analyze the mesenchymal clusters in greater detail, we subsetted them from the remaining palatal cells. Following reclustering of the palatal mesenchyme, we further excluded three clusters beyond the scope of our study: non-CNCC-derived chondrocytes (Acan⁺/Matn3⁺/Snorc⁺, Cluster 14), vascular smooth muscle cells (Myh11⁺/Tinagl1⁺/Rgs5⁺, Cluster 16), and a very small population (34 cells) with low endothelial marker expression (Cluster 17) (Fig. 2a).
a UMAP plot showing mesenchymal cell clusters from integrated scRNA-seq data of E12.5 to E18.5 (E12.5, E13.5, E14.5, E15.5, and E18.5) palatal tissue. b UMAP plot of mesenchymal cell clusters subsetted from (a) after excluding clusters 14, 16, and 17 for a more detailed analysis. c Annotated mesenchymal cell types identified from clusters in (b). d Dot plot showing the expression of marker genes used to define each mesenchymal subpopulation in (c) across clusters. The size of each dot indicates the proportion of cells within the cluster expressing the respective gene, and the color intensity reflects the average expression level of the gene in the expressing cells. e UMAP plots illustrating combined and stage-specific distributions of mesenchymal cell clusters across E12.5 to E18.5. Note the reduction in progenitors (red) from E12.5 to E18.5. f Percentage distribution of mesenchymal cell types within each cluster, calculated relative to the total cell population at each developmental stage. More specifically, the proportion of the progenitor cluster (red) decreases from E12.5 to E18.5: 16.4 % at E12.5, 13.2% at E13.5, 7.20% at E14.5, 7.92% at E15.5, and 9.99% at E18.5. g Dot plot showing the expression of the gene Sox9 is enriched in putative progenitor populations across clusters. The size of each dot indicates the proportion of cells within the cluster expressing the respective gene, and the color intensity reflects the average expression level of the gene in the expressing cells.
We then performed differential gene expression analysis of all the remaining CNCC-derived mesenchymal clusters (Fig. 2b–d) and identified their corresponding cell types. Specifically, Clusters 5 and 11 expressed typical osteogenic markers (Runx2, Sp7, Ibsp, Alpl), along with Sox6, Smpd3, and Cfh. Clusters 7 and 8 expressed known markers (Lhx6, Msx1), along with Tfap2b, Plppr1, Grik1, and Nkain3, for the odontogenic cells adjacent to the palatal shelf. Perimysial clusters (3, 12) showed expression of Hic1, Tbx15, Aldh1a2, Fgf18, consistent with our previous studies9,19, along with additional markers such as Ebf2, Ebf3, and Spag17. Midline mesenchymal clusters (1, 4, 10), expressing marker genes associated with midline identify (Tbx22, Alx1, Meis2), also expressed Pax3. Consistent with our previous findings9, mitotic cells (Mki67, Top2a, Ccnb1, H1f5) in clusters 2, 6, 9, and 13 were distinguishable from the rest of the palatal mesenchyme due to their high expression of cell cycle genes, even after cell-cycle regression (Fig. 2b–d).
We also characterized a putative palatal mesenchymal progenitor population, which has not been previously studied. In the absence of known markers for these progenitors, we assessed cell distribution across developmental stages in the integrated E12.5-E18.5 scRNA-seq data. As embryonic development progresses, the progenitor population diminishes, leading us to hypothesize that a cell population enriched in early embryonic stages but less common in later stages might represent a progenitor population. Using this criterion, we identified a putative progenitor population (cluster 0, Fig. 2b) that is predominantly enriched in E12.5 cells and progressively declines during development (Fig. 2e). This trend was confirmed by decreased progenitor ratios across stages (Fig. 2f). This population is also enriched in expression of Sox9, which has been shown to be associated with early neural crest fate determination20, further supporting its putative progenitor fate (Fig. 2g). Taken together, our analyses comprehensively characterized all cell populations in the palatal region, with a focus on mesenchymal cells and their marker genes across all stages of palate development. This foundational work enabled the use of spatial genomics to map mesenchymal lineages in vivo from early to late developmental stages.
In vivo mapping of mesenchymal cells during palatogenesis
To accurately capture the cellular and molecular landscape of the palate and other craniofacial regions using seqFISH, we designed a customized gene panel with markers for all cell types relevant to palatal anatomy, with a particular emphasis on mesenchymal subpopulations. For each mesenchymal cell type, we included both established and newly discovered marker genes specific to osteogenic, odontogenic, perimysial, midline, and mitotic cells. Additionally, we incorporated pan-palatal mesenchymal markers (Barx1) and markers distinguishing anterior and posterior regions (Meox2 and Shox2). For other palatal cell types, marker genes for epithelial, endothelial, chondrogenic, myogenic, myeloid, and neuronal cells were also included. This generated a curated panel of 94 marker genes for seqFISH-based spatial transcriptomics (Supplementary Table 1).
We first performed seqFISH using our designed gene panel in the mouse head at E15.5. At this stage, these cell types have already reached their final anatomical locations but are not yet fully differentiated, likely retaining markers associated with both early and later differentiation stages. By extending our analysis to the entire anterior and posterior embryonic head, we were able to generate spatial transcriptomic data for the craniofacial region more broadly due to the similarity between the gene expression in the palate and other craniofacial cell populations. All seqFISH gene panel expression patterns are available through FaceBase to build a gene expression database for E15.5 craniofacial tissue (DOI: 10.25550/62-Y0VT [Full data]; Figs. S2 and S3 [Preview]).
To construct a comprehensive spatial atlas of cell types in transcriptionally mapped E15.5 mouse head sections, we further performed cell clustering analysis using spatial gene expression profiles obtained from seqFISH. Using Scanpy analysis21, we identified distinct cell clusters from the anterior and posterior head regions in UMAP plots (Fig. 3a, e). By further extracting spatial information for each cell cluster and visualizing these clusters with spatial plots, we identified distinct, spatially restricted clusters in the entire anterior and posterior head (Fig. 3b, f). In the hard palate region (Fig. 3c), we identified several distinct spatial clusters corresponding to populations observed in scRNA-seq analysis. Based on their respective anatomical locations, cluster 5 represents the osteogenic mesenchyme, clusters 18 and 36 are associated with the odontogenic mesenchyme, and cluster 13 corresponds to midline mesenchyme. The top-ranked marker genes for these spatial clusters (Fig. 3d) were expressed in the corresponding clusters identified in the scRNA-seq analysis (Fig. S4a), further validating their identity. Specifically, cluster 5 osteogenic mesenchyme is marked by Sp7, Ibsp, Ifitm5, and Sox6; clusters 18 and 36 odontogenic mesenchyme express Msx1, Tfap2b, Calca, and Lef1; and cluster 13 midline mesenchyme is defined by Vcam1, Osr1, Pax3, and Ntrk3. Additionally, we identified a spatially distinct group of cells (cluster 39) medial to the dental epithelium that was not identified in the E15.5 scRNA-seq analysis, tentatively defined as medial fibroblasts (Fig. 3c). These cells express Cxcl5, Ntrk2, Foxf1, and Dlx1, which are also markers of odontogenic mesenchyme based on E15.5 scRNA-seq data (Fig. S4a), suggesting that they may represent a subcluster of the odontogenic population identified in the scRNA-seq analysis.
a, b UMAP (a) and spatial (b) plots of the entire anterior head region at E15.5. c Spatial plot focused on the hard palate region, highlighting mesenchymal clusters and their spatial localization. Color keys indicate clusters in (a–c). d Hierarchical clustered dot plot of top-ranked gene expression in palatal mesenchymal clusters. Dot size represents the proportion of expressing cells, while color intensity indicates average expression levels. The dendrogram highlights hierarchical relationships among clusters. e, f UMAP (e) and spatial (f) plots of the entire posterior head region at E15.5. g UMAP plot focused on the soft palate region, highlighting mesenchymal clusters and their spatial localization. Color keys indicate clusters in (e–g). h Hierarchical clustered dot plot of top-ranked gene expression in palatal mesenchymal and myogenic clusters. Dot size represents the proportion of expressing cells, while color intensity indicates average expression levels. The dendrogram illustrates hierarchical relationships among clusters. Abbreviations: medial pterygoid muscle (MP), odontogenic epithelium (oe), and pterygoid plate (PP).
In the soft palate region, consistent with the scRNA-seq analysis, we identified cluster 13 as perimysial cells based on its location adjacent to the myogenic clusters (2 and 15, Myod1⁺), and we further identified midline clusters 27, 47, and 49 based on their central locations in the palatal shelf (Fig. 3g). By comparing the top differentially expressed marker genes from spatial data analysis (Fig. 3h) with scRNA-seq data (Fig. S4b), we confirmed the identities of these clusters. Perimysial cluster 13 expresses markers such as Spag17, which is specific to Hic1+/Tbx15+ perimysial cell populations observed in E15.5 scRNA-seq data (Fig. S4b). The midline clusters 27, 47, and 49 also express midline markers Tbx22 and Pax3, consistent with scRNA-seq data. Additionally, we identified two clusters (19 and 26) not detected in the scRNA-seq data, located between the midline and perimysial cells (Fig. 3g). These clusters are positioned closer to the nasal mesenchyme; thus, we tentatively identified them as nasal fibroblasts (Fig. 3h). Their markers (Tbx18, Cxcl5, Foxf1, Igfbp2) are associated with both perimysial and midline populations in E15.5 scRNA-seq data (Fig. S4b), suggesting they may have been grouped with either perimysial or midline cells in the scRNA-seq analysis.
Furthermore, distinct mesenchymal populations identified in other regions through spatial analysis also faithfully matched their anatomical identities (Fig. S5a). These included cell types similar to those in the palatal region in the same anterior section, such as osteogenic (os) mesenchyme associated with the orbito-sphenoid and mandibular bones and odontogenic (od) cells in the mandibular tooth region (Fig. S5b–d). This spatially resolved cell type map also revealed diversity in the putative fibroblasts under the oral mucosa compared to those under the dermis. For example, fibroblasts beneath the dermis (df) in the calvarial and mandibular regions (Fig. S5c, d) form a single cluster, while fibroblasts under the oral mucosa medial to the teeth (mf) in the palatal and mandibular regions (Fig. S5b, d) cluster together. Marker analysis revealed that the dermal and mucosal fibroblastic clusters have distinct gene expression profiles (Fig. S5e, f), suggesting differences in developmental origins or local environments.
Moreover, the mandibular bone’s osteogenic (os) mesenchyme, previously considered a single population, can now be divided into two spatially distinct clusters: one linked to the alveolar bone (c1) and the other to the mandibular body near Meckel’s cartilage (c2) (Fig. S5d). At the molecular level, the heterogeneity of these two clusters is further evidenced by differentially expressed markers (Fig. S5g, h). This distinction is supported by their functional differences: the porous alveolar bone supports teeth, while the dense mandibular body provides jaw stability and resists mechanical forces. Therefore, further exploration of these clusters could provide more insights into the structural and functional differences between the alveolar bone and the mandibular body.
In vivo mapping of mesenchymal markers during palatogenesis
To identify specific markers for in vivo mapping of each mesenchymal population, we screened the high-resolution gene expression patterns generated by seqFISH for the marker genes for each spatial cluster. Focusing on the genes with the highest spatial specificity for each mesenchymal cell type (Fig. 4a, j), we identified Sox6 and Sp7 as specific markers for osteogenic cells (Fig. 4b, c), Tfap2b and Msx1 for odontogenic cells (Fig. 4d, e), and Hic1 and Tbx15 for the perimysial population (Fig. 4k, l). Consistent with previous findings that anterior and posterior palatal mesenchyme exhibit distinct gene expression profiles during early development13,14,15, the midline mesenchymal population also showed anterior-posterior heterogeneity. While both midline regions expressed Pax3, Ntrk3 was detected in the hard palate, and Tbx22 was observed in the soft palate (Fig. 4f, g, m, n). Additionally, we identified marker genes for the newly resolved populations of fibroblasts: Cxcl5 and Ntrk2 mark the population medial to the teeth on the oral side of the hard palate (Fig. 4h, i), while Tbx18 and Foxf1 identify the population adjacent to the epithelium on the nasal side of the soft palate (Fig. 4o, p).
a Gene expression patterns produced using seqFISH show Tfap2b (orange), Sox6 (magenta), Ntrk3 (yellow), Ntrk2 (green), and Krt14 (light blue) expression in the E15.5 anterior head (hard palate) region. b–i Markers for osteogenic cells: Sox6 (magenta) (b) and Sp7 (magenta) (c); odontogenic cells: Tfap2b (orange) (d) and Msx1 (orange) (e); midline cells: Pax3 (yellow) (f) and Ntrk3 (yellow) (g); fibroblastic cells: Cxcl5 (green) (h) and Ntrk2 (green) (i); and epithelial cells: Krt14 (light blue) (b-i). j Gene expression patterns produced using seqFISH show Tbx15 (green), Pax3 (yellow), Tbx18 (magenta), Krt14 (light blue), Acan (orange), and Myod1 (purple) expression in the E15.5 posterior head (soft palate) region. k–p Markers for perimysial cells: Hic1 (green) (k) and Tbx15 (green) (l); midline cells: Pax3 (yellow) (m) and Tbx22 (yellow) (n); fibroblasts: Tbx18 (magenta) (o) and Foxf1 (magenta) (p); and epithelial cells: Krt14 (light blue) (k–p). Schematic drawings indicating the orientation and section levels are shown in light blue on top of the panels. Boxed areas in a and j are enlarged in b–i and k–p, respectively. White dashed lines in a–p outline palatal shelves; white triangles in b–i and k–p point to the palatal midline. White arrows indicate positive signals in b–i, and k–p. Abbreviations: fibroblasts (f), mandible (M), midline (m), medial pterygoid muscle (MP), nasal septum (NS), odontogenic region (od), odontogenic epithelium (oe), osteogenic region (os), perimysial region (p), pterygoid plate (PP/pp), salivary gland (SG), tensor veli palatini (TVP/tvp), tongue (T). Scale bars: 100 µm in a, j, b (for b–i), and k (for k–p).
As expected, we also uncovered the heterogeneity and diversity of marker gene expression in other craniofacial regions, such as the mandible (M), tongue (T), and nasal (NS) regions (Fig. 4a). For example, the osteogenic marker Sox6 is also expressed throughout the osteogenic mesenchyme clusters of the mandible (M), consistent with their osteogenic identity (Fig. 4a). Although the role of Sox6 in bone development has not been reported in the hard palate, it has been shown to promote osteogenic differentiation of osteogenic progenitors in the mandible22, highlighting the conserved roles of osteogenic-specific genes across craniofacial mesenchyme. These findings support the notion that mesenchymal tissues across the craniofacial complex share developmental similarities and suggest that specific subpopulations of CNCC-derived cells may contribute to multiple tissues and organs in the craniofacial region.
We also examined the in vivo expression patterns of markers for other non-CNCC-derived cell types, which were included in the panel as anatomical landmarks for identifying mesenchymal cell identities. The expression patterns of these typical chondrogenic (Acan), myogenic (Myod1), and epithelial (Krt14) markers all aligned with previous publications (Fig. 4a–p)8,23,24,25. Additionally, we validated the expression patterns detected with seqFISH-based spatial genomics using RNAScope in situ hybridization. We found consistent expression patterns between seqFISH and individual RNAscope in situ staining for several genes examined in this study, including Tfap2b, Sox6, and Pax3 in the hard palate region (Fig. S6a–d) and Hic1, Pax3, and Myod1 in the soft palate region (Fig. S6e–h). Thus, we have established a high-resolution spatially resolved transcriptomic atlas of all key cell populations involved in the development of the palate and potentially other craniofacial regions. Moreover, the lineage-specific markers identified here will facilitate precise labeling of individual cell types and enhance our understanding of mesenchymal fate diversification during early developmental stages.
Broadly, our findings showcase the complementarity of seqFISH-based high-resolution spatial and single-cell omics, through which we successfully established an in vivo atlas of distinct mesenchymal cell populations based on gene expression profiles. While our focus here was primarily on the palatal mesenchyme, these techniques are readily applicable to other craniofacial mesenchymal tissues, and our spatial gene expression profiles provide a foundation for further analysis. This method could enhance anatomical atlases at the molecular and cellular levels, particularly in early differentiation stages when anatomical structures are not yet well defined.
Dynamic mapping of mesenchymal lineages during palatogenesis
Developmental studies have traditionally relied on static histological sections, limiting the integration of anatomical data with gene expression profiles and hindering the spatial tracking of cell types during development. Thus, we employed spatial genomics analysis, which integrates both anatomical and gene expression data, to bioinformatically visualize how CNCC-derived mesenchymal progenitors undergo lineage diversification during palatogenesis. Using seqFISH, we performed spatial transcriptomic analysis of E12.5 and E13.5 mouse heads to include spatial gene expression profiles from earlier developmental stages. All gene panel expression patterns generated through this study are also available on FaceBase, serving as an E12.5 and E13.5 craniofacial tissue gene expression database (DOI: 10.25550/62-Y0VT [Full data]; Figs. S7–S10 [Preview]).
By clustering the same cell types across different stages using integrative spatial analysis, we can observe their dynamic changes in their in vivo distribution throughout development. We utilized this approach to analyze spatial transcriptomics data from E12.5, E13.5, and E15.5 sections of the embryonic head, to establish a dynamic cell type atlas from the onset of palatal shelf formation (E12.5) to the point where cell lineages are established in their final anatomical locations (E15.5). Through this integrative spatial analysis, we generated multi-stage, spatially resolved cell cluster maps illustrating the dynamics of specific cellular lineages, providing a comprehensive temporal and spatial cell type atlas of craniofacial development (Fig. S11). Given the structural differences between the anterior and posterior regions of the embryonic head, particularly in the palatal region, we integrated spatial transcriptomics data from these two regions independently (Fig. S11a–g and h–n, respectively). Furthermore, our panel of genes also labeled distinct cell lineages in the developing brain, potentially providing valuable lineage information for neural development as well (Fig. S11e-g and l-n). We have also shared the full version of this dynamic, integrated craniofacial cell type atlas on FaceBase (DOI: 10.25550/62-Y0VT), facilitating direct and interactive visualization and analysis of cell clusters using the open-source software TissUUmaps26.
We focused on mesenchymal cell populations associated with the hard and soft palate regions to study their spatial dynamics as the palate develops (Fig. 5). We first verified the in vivo identities of various integrated mesenchymal clusters at E15.5, a stage when anatomical structures are well established. The clustering was generally consistent with that observed using individual E15.5 hard and soft palate spatial data alone, although fewer distinct clusters per cell type were observed despite the higher resolution used for clustering. This may be due to the inclusion of clusters from earlier developmental stages in the integrated clustering and/or to differences between the Scanpy and scvi-tool algorithms. In the hard palate region, clusters 12 and 25 were associated with the osteogenic region, clusters 28 and 54 with the odontogenic region, and cluster 62 with the midline region (Fig. 5c, f, i). Similarly, in the soft palate region, clusters 51, 72, and 38 were associated with the perimysial, midline, and nasal fibroblast regions, respectively (Fig. 5l, o, r).
a–r TissUUmaps spatial plots of E12.5, E13.5, and E15.5 hard and soft palate regions from an integrative analysis of palatal spatial genomics data, with color keys displayed on the left. Different cell clusters are represented by distinct colors. For the hard palate region, individual clusters are shown as follows: osteogenic clusters 12 and 25 (a–c), odontogenic clusters 28 and 54 (d–f), and midline cluster 62 (g-i). For the soft palate region, individual clusters are represented as follows: perimysial cluster 51 (j–l), midline cluster 72 (m–o), and fibroblastic cluster 38 (p–r). Schematic drawings of section orientation and levels are depicted in light blue above the main panels. Black arrows indicate the locations of the respective cell clusters. Abbreviations: muscle mass (mm), odontogenic epithelium (oe), palatal epithelium (pe), and tensor veli palatini (tvp).
At E12.5, we found that all the main mesenchymal lineages had already been established in the palatal region at distinct anatomical locations (Fig. 5a, d, g, j, m, p), suggesting early fate diversification at or before CNCCs’ arrival in the palatal primordium. Furthermore, the distribution of these cell populations relative to the palatal shelves underwent notable changes from E12.5 to E15.5 (Fig. 5a–r). This observation aligns with the significant morphological transformations that occur during palatogenesis, such as the initial vertical growth of the palatal shelves, followed by their horizontal expansion. Specifically, we found that in the hard palate region, osteogenic cells, initially present in small numbers near the superior region of the palatal shelf, expanded to an inferior and medial location at a later stage (Fig. 5a–c). In contrast, odontogenic cells in the dental mesenchyme remained in more consistent locations on the oral side adjacent to the palatal shelf (Fig. 5d–f), but the two odontogenic populations, initially as a mixed group at E12.5, became anatomically distinct during development, forming two clusters: one linked to the dental papilla (cluster 28) and the other to the dental follicle (cluster 54), consistent with the odontogenic mesenchyme lineage bifurcation observed in our previous scRNA-seq analysis27. The midline population showed the most dynamic positioning, transitioning from the nasal region near the cranial base to the center of the palatal shelf (Fig. 5g–i).
In contrast, the spatial distribution of mesenchymal cell lineages in the soft palate region was less dynamic. For example, perimysial cells primarily expanded mediolaterally to support myogenic cells (Fig. 5j–l). The midline cells, initially located in the medial part of both palatal shelves, merged into a single population when the palatal shelves fused at the midline (Fig. 5m–o). Additionally, newly identified sub-nasal fibroblasts expanded inferiorly along the epithelium during palatal development (Fig. 5p–r). These findings demonstrate the feasibility of using integrative spatial analysis to globally visualize the dynamic locations and specifications of distinct mesenchymal cell populations during palate development. Our data also serves as a valuable resource for visualizing cell lineage dynamics during the development of other craniofacial organs.
Mesenchymal progenitor heterogeneity during palatogenesis
To investigate when palatal mesenchymal progenitors begin to activate lineage-specific markers and commit to distinct lineage trajectories, we further characterized the putative Sox9+ palatal mesenchymal progenitor population in vivo. We first examined the in vivo identity of these putative progenitors in the hard and soft palate regions at the onset of palatal development. At E12.5, the majority of SOX9 expression was observed around the superior region of the palatal shelves, close to the cranial base, in both the soft and hard palate regions, with a small minority associated with the odontogenic region (Fig. 6a–d). Notably, the medial and inferior regions of the palatal shelf primordium lacked SOX9 expression, suggesting that early fate diversification had already occurred between SOX9+ and SOX9- regions as early as E12.5 (Fig. 6b, d).
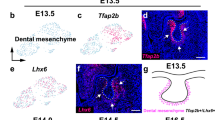
a–d Immunofluorescence of SOX9 (red) in coronal sections of the hard palate region (a, b) and soft palate region (c, d) at E12.5. Each experiment was independently repeated three times. e–i Immunofluorescence of tdTomato (red) (e–h, i, k), SP7 (green) (j), and MHC (green) (l) in coronal sections of the palate region of Sox9-CreER;tdTomato mice at E13.5 (e–h) and E16.5 (i–l). Each experiment was independently repeated three times. The injection protocol is illustrated below the figures. Schematic drawings indicating the orientation and section levels are shown in light blue on top of the panels. Boxed areas in a, c, e, g, i, and k are enlarged in b, d, f, h, j, and l, respectively. The boxed area in j is enlarged in the inset to highlight the colocalization of SP7 and tdTomato signals (white). White arrows indicate positive signals in the palate region in b, d, f, and h as well as in the osteogenic region in j and the perimysial region in l. Yellow arrows indicate signals in the odontogenic region in b, f, and i. Asterisks indicate the absence of expression in b, d, and j. White dashed lines in b, d, g, h, j, and l, outline the palatal shelves; yellow dashed lines in b, f, and i outline the odontogenic epithelium. Abbreviations: tamoxifen (TMX). Scale bars: 100 µm in a (for a and c), e (for e and g), i (for i and k), and b (for the remaining panels).
To validate that SOX9+ cells could serve as progenitors for various mesenchymal cell types in the palatal region, we next utilized Sox9-CreER;tdTomato reporter mice to trace Sox9+ cells and their progeny during palatogenesis. Given the approximately 16-24 hour window of effective tamoxifen induction during the embryonic stage28, we induced the labeling of Sox9+ cells at E11.5 and analyzed the contribution of Sox9+ cell derivatives (tdTomato+) from E12.5 onwards. By E13.5, Sox9+ cell derivatives had expanded into the inferior and medial parts of the palatal shelves (Fig. 6e–h), demonstrating growth potential indicative of progenitors. The Sox9+ cell progeny continued to increase in number and expand throughout most of the palatal shelves at E16.5, contributing to osteogenic and odontogenic regions in the hard palate region, as well as perimysial cells in the soft palate region (Fig. 6i–l). A small population on the oral side of the hard palate (Fig. 6j) was not labeled by the Sox9+ lineage, indicating that these cells may originate from a population that does not express Sox9 at E12.5. Overall, these findings suggest that Sox9+ cells indeed serve as progenitors for multiple mesenchymal cell lineages at E12.5.
Our integrative spatial analysis bioinformatically predicted that distinct cell lineages are established at the onset of palate development at E12.5. Thus, we hypothesized that Sox9+ mesenchymal progenitors also represent a heterogeneous population as early as E12.5, comprising several lineage-specific progenitor subpopulations, each characterized by the activation of early lineage-specific markers. To test this, we first identified early lineage-specific markers for cell types derived from SOX9+ cells. Using spatial gene expression patterns from seqFISH representing early (E12.5) and late (E15.5) populations, we investigated the spatiotemporal expression patterns of marker genes associated with osteogenic, odontogenic, and perimysial lineages (Fig. 7, Fig. S12). We identified distinct, dynamic expression patterns of these marker genes across developmental stages. We focused on genes actively expressed at early stages but with greater spatial specificity to the cell types of interest, making them ideal markers for lineage-specific progenitors and labeling distinct cell lineages. In this category, we identified Sox6 as a putative progenitor marker for the osteogenic lineage (Fig. 7a–d), Tfap2b for the odontogenic lineage (Fig. 7e–h), and Hic1 for the perimysial lineage (Fig. 7i–l).
a–d TissUUmaps spatial plots for osteogenic regions (a, b), showing the corresponding marker gene Sox6 (magenta) visualized by SeqFISH (c, d). e–h TissUUmaps spatial plots for odontogenic regions (e, f), showing the corresponding marker gene Tfap2b (orange) visualized by SeqFISH (g, h). i–l TissUUmaps spatial plots for perimysial regions (i, j), showing the corresponding marker gene Hic1 (green) visualized by SeqFISH (k, l). The epithelial marker Krt14 is shown in light blue in c, d, g, h, k, and l. White dashed lines outline the palatal shelves in c, d, g, h, k, and l, while yellow dashed lines in c, d, g, and h outline the odontogenic epithelium. Black arrows indicate the locations of the respective cell clusters in a, b, e, f, i, and j, while white arrows indicate positive signals in the remaining panels. Abbreviations: muscle mass (mm), odontogenic epithelium (oe), palatal epithelium (pe), and tensor veli palatini (tvp). Scale bars: 100 µm in c (for c and g), d (for d and h), k, and l.
We also found some genes that are activated early and broadly in the palatal region, with similarly extensive expression across anatomical structures and cell types at later stages, including known early markers like Runx2 (osteogenic) (Fig. S12c, d), Lhx6 (odontogenic) (Fig. S12k, l), and a newly identified perimysial-associated marker, Dlx2 (Fig. S12s, t). These genes likely represent an early activation profile associated with common progenitors rather than lineage-specific identity. In addition, another group of marker genes show low or undetectable expression at early stages but are significantly upregulated at later stages, suggesting their association with a more differentiated state. For late-stage osteoblast differentiation, examples include known markers such as Sp7 and Ibsp (Fig. S12e–h). For late odontogenic differentiation, we also identified Lef1 and Grik1 (Fig. S12m–p). Additionally, markers Ntrk2 and Ntrk3 are associated with late-stage perimysial fate determination (Fig. S12u–x). Collectively, these findings reveal a shift in molecular markers from broadly expressed, multi-lineage markers during early development to increasingly more specific markers along the differentiation trajectory. This shift underscores the complexity of developmental processes and emphasizes the importance of temporal and spatial specificity in identifying and utilizing molecular markers for cell lineage analysis.
Using the early lineage-specific markers we identified here: Sox6 for osteogenic, Tfap2b for odontogenic, and Hic1 for perimysial cells, we further evaluated the heterogeneity of Sox9+ cells at the onset of palate development. At E12.5, as observed earlier, these markers were already expressed in distinct domains in the hard and soft palate regions (Fig. 8a, c, and e), and they also overlapped with different subsets of Sox9+ progenitors in various regions (Fig. 8b, d, and f), supporting the molecular heterogeneity of the Sox9+ progenitor cells. Thus, the lineage diversification of CNCC-derived mesenchymal progenitors may have already begun before the CNCCs arrived at the palatal primordium at E12.5, with fate determination established prior to the onset of palatal development.
a–d Gene expression patterns produced using seqFISH showing co-expression of Sox6 (magenta) and Sox9 (white) (a, b) and Tfap2b (orange) and Sox9 (white) (c, d) in the hard palate region at E12.5. e, f Gene expression patterns produced using seqFISH showing co-expression of Hic1 (green), Myod1 (purple), and Sox9 (white) (e), and Hic1 (green) and Sox9 (white) (f) in the soft palate region at E12.5. Magenta, orange, and green boxes in a, c, and e are enlarged in b, d, and f, respectively. Magenta, orange, and green arrows in a, c, and e indicate regions expressing Sox6, Tfap2b, and Hic1, respectively. White arrows in b, d, and f indicate co-expression of both genes in the same cells. White dashed lines in a, c, and e outline the palatal shelves. g, j RNAScope in situ hybridization for Tfap2b (red) in the hard palate region (g), and Hic1 (red) and Myod1 (green) in the soft palate region (j) at E12.5. Each experiment was independently repeated three times. h, i Immunofluorescence of tdTomato (red) in coronal sections of the hard palate region of Tfap2b-CreER;tdTomato mice at E13.5 (h) and E16.5 (i). Each experiment was independently repeated three times. k, l Immunofluorescence of tdTomato (red) and MyoD (green) in coronal sections of the soft palate region of Hic1-CreER;tdTomato mice at E13.5 (k) and E16.5 (l). Each experiment was independently repeated three times. Injection protocol is illustrated below the figures. White dashed lines outline the palatal shelves in g–l, yellow dashed lines outline the odontogenic epithelium in g–i and the pterygoid plate in l. White arrows indicate positive cells in g–l. Schematic drawings of section orientation and levels are shown on the left for panels a-f and above for panels g–l. Abbreviations: muscle mass (mm), medial pterygoid muscle (mp), tensor veli palatini (tvp), and tamoxifen (TMX). Scale bars: 50 µm in a (for a and c), b (for b and d), e, f, and g (for g–l).
To determine whether the activation of these lineage-specific markers at early stages truly indicates the commitment to their respective cell fates, we further tested whether these early cells contribute to their corresponding late populations or if the late populations form de novo, independently of the early populations established at the onset of palatal development. We utilized transgenic mouse models to label early cell populations based on their marker gene expression. We traced two lineages derived from Sox9+ progenitors, namely odontogenic and perimysial cells, using Tfap2b-CreER;tdTomato and Hic1-CreER;tdTomato mice, respectively (Fig. 8g-l). Similarly to our analysis of Sox9-CreER;tdTomato mice, we performed tamoxifen induction at E11.5 to track these lineages from E12.5 onwards. Our in vivo lineage tracing results aligned with the integrative spatial analysis, showing that populations labeled by early lineage-specific markers indeed gave rise to the expected respective cell types at a later stage. Specifically, Tfap2b+ odontogenic cells labeled at E12.5 indeed gave rise to odontogenic cells from E13.5 onwards and occupied the entire odontogenic region by E16.5 (Fig. 8g–i). Hic1+ perimysial cells labeled at E12.5, initially restricted to the lateral side of the palatal shelf at E13.5, expanded medially into perimysial mesenchyme surrounding the myogenic cells by E16.5 (Fig. 8j–l). Thus, the heterogeneous expression of early lineage-specific markers indeed indicates that these mesenchymal progenitors have already diversified into various mesenchymal fates as early as E12.5.
Mesenchymal progenitor heterogeneity prior to palatogenesis
To further investigate the stage at which mesenchymal progenitors in the maxilla specify their fates, we examined the early development of the maxillary processes of the first branchial arch, which give rise to maxillary structures. We analyzed the anterior embryonic head region corresponding to the hard palate and the posterior region associated with the soft palate from E9.5 to E11.5 (Fig. 9), prior to the initiation of palatal shelf development. Specifically, we used Tfap2b (Fig. 9a–f) to label odontogenic progenitors in the anterior and Hic1 to identify perimysial cells in the posterior (Fig. 9g–l). We found that the establishment of region-specific expression for both Tfap2b and Hic1 in the maxillary process occurs between E10.5 and E11.5.
a–f RNAscope in situ hybridization for Tfap2b (red) in the anterior embryonic head from E9.5 to E11.5. Each experiment was independently repeated three times. Boxes in a–c are enlarged in d–f, respectively. Arrows indicate positive signals in the maxillary process in d and the oral region of the maxillary process in f. Arrowheads indicate positive signals in the aboral region of the maxillary process in e and f. White dashed lines in e and f outline the oral epithelium. g–l RNAscope in situ hybridization for Hic1 (red) in the posterior embryonic head from E9.5 to E11.5. Each experiment was independently repeated three times. Boxes in g–i are enlarged in j–l, respectively. Arrows mark signals in the maxillary process in j and close to the midline region of the maxillary process in l. Arrowheads indicate signals in the central region of the maxillary process in k and l. White dashed lines in k and l outline the oral epithelium. Scale bars: 100 µm in a (for a–c and g–i) and d (for the remaining panels).
Specifically, at E9.5, Tfap2b expression is widespread in the maxillary process (Fig. 9a, d), but by E10.5, it becomes restricted to the aboral region in the maxillary process, which is not typically associated with odontogenic fate (Fig. 9b, e). By E11.5, while this aboral expression reduces, Tfap2b expression begins to emerge in the oral region of the maxillary process, which is commonly associated with odontogenic fate (Fig. 9c, f). Similarly, at E9.5, in the posterior region, Hic1 expression is present extensively but at very low levels (Fig. 9g, j). By E10.5, Hic1 expression becomes enriched and localized to the central area of the maxillary process (Fig. 9h, k). At E11.5, this Hic1-expressing region expands to include the more medial area where the future soft palatal shelf will develop (Fig. 9i, l). This gradual restriction and activation of Tfap2b and Hic1 expression during E10.5 to E11.5 in regions associated with their future anatomical locations suggest potential fate determination of their respective lineages during this period.
To determine whether the Tfap2b+ or Hic1+ cells at E10.5-E11.5 can serve as progenitors for odontogenic and perimysial cells at later stages, we lineage-traced these two populations using Tfap2b-CreER;tdTomato and Hic1-CreER;tdTomato mice. As in other lineage tracing studies, tamoxifen injections were administered one day earlier, at E9.5 and E10.5, to label cells at E10.5 and E11.5, respectively. The labeled populations were then analyzed at E16.5 (Fig. 10). We found that while progenitors labeled from E10.5 onward (tamoxifen induced at E9.5) began contributing to their respective lineages, progenitors labeled from E11.5 (tamoxifen induced at E10.5) showed a more efficient and specific contribution.
a–d Immunofluorescence of tdTomato (red) in coronal sections of the hard palate region in E16.5 Tfap2b-CreER;tdTomato mice induced at E9.5 (a, b) and E10.5 (c, d). Each experiment was independently repeated three times. Boxes in a and c are enlarged in b and d, respectively. White dashed lines outline the palatal shelves, and yellow dashed lines outline the odontogenic epithelium. The arrow in a and the asterisk in c indicate the presence and absence, respectively, of tdTomato-positive cells in the non-odontogenic region. The arrows in b and d indicate the presence of tdTomato-positive cells in the odontogenic region. e–h Immunofluorescence of tdTomato (red) and MyoD (green) in coronal sections of the soft palate region at E16.5 from Hic1-CreER;tdTomato mice induced at E9.5 (e, f) and E10.5 (g, h). Each experiment was independently repeated three times. Boxes in e and g are enlarged in f and h, respectively. White dashed lines outline the palatal shelves in e and g, and yellow dashed lines in f and h outline the pterygoid plate. Arrows in f and h indicate the presence of tdTomato-positive cells in the perimysial region. Abbreviations: medial pterygoid muscle (mp), pterygoid plate (pp), tamoxifen (TMX), and tensor veli palatini (tvp). Scale bars: 100 µm in a (for a, c, e, and g) and in b (for the remaining panels).
In Tfap2b-CreER;tdTomato mice, when labeled at E10.5 (tamoxifen induced at E9.5), the Tfap2b+ lineage already started to contribute to the dental mesenchyme but also showed notable contributions to other mesenchymal populations in the palatal shelf (Fig. 10a, b), suggesting that specificity for the odontogenic lineages is still being established at this stage. In contrast, when labeled at E11.5 (Tamoxifen induced at E10.5), the labeled cells were more restricted to the dental mesenchyme, with very few detected in the palatal shelf (Fig. 10c, d). Similarly, in Hic1-CreER;tdTomato mice, when labeled at E10.5 (tamoxifen induced at E9.5), a few tdTomato+ cells started to be observed in the perimysial region surrounding the myogenic cells of the tensor veli palatini (TVP) (Fig. 10e, f). However, this perimysial contribution of Hic1+ cells significantly increased when labeled at E11.5 (tamoxifen induced at E10.5) (Fig. 10g, h), suggesting the activation of Hic1+ perimysial progenitor cells between E10.5-E11.5. These findings support the establishment of distinct mesenchymal cell types for the palatal region in the maxillary process of the first pharyngeal arch at E10.5-E11.5 and further emphasize that fate diversification of anterior and posterior palatal mesenchyme occurs before the initiation of palatal development. Taken together, our findings highlight the critical role of early developmental stages, prior to palatogenesis, in shaping cell fate decisions that influence distinct palatal mesenchymal cell types at later stages.
In summary, in this study, we demonstrated the application of spatially resolved bioinformatic predictions to identify lineage-specific progenitors and their early markers in the palatal mesenchyme and potentially other craniofacial mesenchymal lineages. These findings validate the accuracy of bioinformatically predicted dynamic lineage visualization through integrative spatial analysis, establishing a robust method and resource for tracing and understanding lineage dynamics in craniofacial mesenchymal development, thereby enhancing our ability to map the origins of crucial cell types during embryogenesis and providing comprehensive information on the dynamics of cell fate determination. Our findings also demonstrate the necessity of expanding the study window to include stages prior to the formation of anatomical structures, allowing for a more comprehensive investigation of developmental events, particularly those occurring at later stages.
Discussion
In this study, we demonstrate the utility of high-resolution seqFISH-based spatial genomics in mapping the molecular and cellular landscapes during craniofacial development. Focusing on development of the mouse palatal region as an example, we identified early lineage-specific markers that reveal heterogeneity is already present in mesenchymal cells and progenitors derived from post-migratory CNCCs at the onset of palatal development (E12.5). This early heterogeneity suggests that CNCCs undergo fate diversification prior to or immediately upon arriving at the regions associated with palatal development. We further identified mesenchymal lineage progenitor marker expression in the maxillary process of the first pharyngeal arch at developmental stages preceding palatal shelf formation. Moreover, cells expressing these mesenchymal progenitor markers in the maxillary process contribute to the corresponding cell types in the palatal region, as demonstrated by lineage tracing. These findings align with previous studies in early CNCC development, which indicate that early CNCCs also exhibit predetermined heterogeneity1,5. To elucidate the detailed mechanisms underlying palatal mesenchymal fate diversification in CNCCs, it will be essential to conduct bioinformatic analyses, including scRNA-seq, single cell ATAC-seq (scATAC-seq), and/or spatial omics, at developmental stages before palatal shelf formation, to investigate the regulatory mechanisms governing the establishment of lineage-specific progenitors. Additionally, for in vivo cell lineage analysis, recent barcoded lineage tracing transgenic mouse models29,30 would also facilitate a systematic mapping of the diversification and contributions of palatal and other craniofacial mesenchymal lineages derived from post-migratory CNCCs.
In addition to the high-resolution spatial transcriptome mapping using seqFISH-based techniques, the integrative analysis of spatial transcriptomics data across various developmental stages presented here has significantly advanced our understanding of craniofacial development. Historically, the relationships between cell types have been inferred from correspondences across histological sections, which are static representations of cell populations. This approach has limited power to reveal the dynamic interactions and migration patterns that are crucial during development. By adopting spatial transcriptomics, which integrates anatomical locations with gene expression profiles, we have been able to define cell types on a molecular basis even as they vary across different temporal and spatial contexts. This method allowed us to capture the continuous evolution of cell identities, providing insights into how cell types emerge and transition during development. It also provided important information on potential cell-cell interactions between neighboring mesenchymal populations and adjacent non-mesenchymal cells, which can serve as a foundation for future hypothesis-driven research that could reveal the molecular and cellular regulatory mechanisms of craniofacial development. Moreover, the ability to characterize cell types before the establishment of clear anatomical structures provides a clearer view into the sequential and spatial coordination of developmental processes than has previously been available. By analyzing the temporal and spatial patterns of lineage-specific markers within lineages of interest, we identified genes that are activated early and exhibit spatial specificity, facilitating the identification of early lineage fate regulation and the tracing of distinct progenitor populations. Importantly, our in vivo validation through lineage tracing not only strengthens the reliability of our findings but also highlights the power of integrating multiple technologies for understanding cell-cell interactions and their potential impact during embryonic development.
Through spatial integrative analysis during palate development, we further uncovered distinct and dynamic changes in the spatial distribution patterns of different mesenchymal lineages as the developing palatal region undergoes dramatic morphological changes. Cell lineage tracing using two early lineage-specific markers (Trap2b for odontogenic and Hic1 for perimysial cells) demonstrated that these early populations identified through integrative analysis contribute to their respective cell types at later stages. Therefore, it is likely that an early population may ultimately contribute to its later-stage cells in a different spatial location relative to the palatal shelf. Notably, in the integrative spatial analysis, the osteogenic and midline populations in the hard palate exhibited the most dramatic shifts in their anatomical location within these lineage trajectories. These populations shifted from the superior part of the palatal shelf primordium to the central and medial regions of the inferior part of the palatal shelf at later stages, likely involving intra-palatal migration to reach their final anatomical positions. This possibility aligns with our previous study on the dynamic movement of Gbx2+ progenitors during mandible development31, supporting the potential role of orderly migration in the differentiation of post-migratory CNCCs in vivo. During early CNCC migration, local microenvironmental cues are crucial for establishing and maintaining migratory streams32. We thus hypothesize that this mechanism may also be conserved in later stages, with distinct environmental signals regulating the dynamic migration patterns. The spatial and single-cell transcriptomic data from this study could be a valuable resource for investigating the regulatory cell-cell interactions driving CNCC lineage diversification and directed cell movements.
Although our initial goal was to construct a spatially resolved transcriptional map of the palatal mesenchyme, we expanded our scope to include the entire craniofacial complex using seqFISH-based spatial genomics. Consequently, the spatial transcriptomic maps we generated cover both the anterior and posterior craniofacial regions at developmental stages E12.5, E13.5, and E15.5. This comprehensive dataset provides the research community with a valuable resource for analyzing high-resolution gene expression patterns across the craniofacial landscape. The observed similarities among mesenchymal cells across different craniofacial regions suggest that these markers may be useful for studying and identifying similar cell types in other craniofacial mesenchymal tissues, such as those in the mandible and calvaria, in both developmental and disease contexts. For example, we identified previously unreported heterogeneity within the osteogenic mesenchyme in the mandibular region. This finding is supported by additional genes not included in our initial gene panel, such as Fgfr2, Gli1, Hand2, Satb2, Pax9, and Dlx3, which also exhibit variable expression patterns within the osteogenic mesenchyme at stages E12.5 and E14.5, as detailed in data available on FaceBase33 (DOI: 10.25550/1WWE). This data resource allows researchers to directly visualize the molecular and cellular landscapes identified in our analyses or to perform more specialized analyses tailored to their own studies. For example, by integrating this spatial data with scRNA-seq data from the literature, such as that from the developing mandible31 and calvaria34, future studies could advance our understanding of cellular heterogeneity and its maintenance within these regions.
In summary, we demonstrate the application of high-resolution spatial transcriptomic mapping to investigate the dynamic processes underlying craniofacial development. Leveraging these insights, our work establishes a dynamic molecular and cellular atlas, thereby enhancing our understanding of the molecular regulation of cell fate decisions and cell-cell interactions during embryogenesis. This atlas of the craniofacial region provides invaluable bioinformatic predictions for cell type establishment in the palatal region and beyond, serving as a crucial resource for future studies on craniofacial morphogenesis and related disorders. It will continue to expand our understanding of the dynamic cell fate determination process during embryonic development.
Methods
Animals
All animal handling procedures were conducted in compliance with all relevant ethical regulations approved by the Institutional Animal Care and Use Committee (IACUC) at the University of Southern California, protocol #9320. The following mouse strains were used in this study: C57BL/6 J (Jackson Laboratory, #000664), Sox9-CreER (Jackson laboratory, #035092)35, Tfap2b-CreER (generated by our group)36, Hic1-CreER (gift from Dr. T. Michael Underhill, University of British Columbia)37, and tdTomato reporter mice (Jackson laboratory, #007905)38. Adult C57BL/6 J male and female mice were crossed to generate C57BL/6 J embryos. For RNAScope in situ hybridization, scRNA-seq, and seqFISH experiments, C57BL/6 J embryos were collected at E9.5, E10.5, E11.5, E12.5, E13.5, E14.5, E15.5, and E18.5. Adult Sox9-CreER, Tfap2b-CreER, and Hic1-CreER mice were crossed with adult tdTomato reporter mice to generate Sox9-CreER;tdTomato, Tfap2b-CreER;tdTomato, and Hic1-CreER;tdTomato embryos. For cell lineage tracing at E11.5, all embryos received tamoxifen (Sigma, T5648) at a dosage of 1.5 mg/10 g body weight and were collected at E13.5 or E16.5. For cell lineage tracing at E9.5 and E10.5, all embryos received tamoxifen at a dosage of 1.0 mg/10 g body weight and were collected at E16.5. All mice were used for analysis without consideration of sex.
Tissue processing
Embryos at different developmental stages, regardless of sex, were fixed in 10% neutral buffered formalin solution (Sigma, HT501128) overnight at room temperature, followed by decalcification in 10% EDTA, depending on the stage. For cryosections, samples were dehydrated in a 15% sucrose/PBS solution, followed by 30% sucrose/50% Tissue-Tek OCT compound (Sakura, 4583). Samples were embedded in the OCT compound and frozen on a block of dry ice. Embedded samples were then sectioned at 8 μm using a cryostat (Leica, CM13050S).
Immunofluorescence
Cryosections were used for immunofluorescence staining. The slides were treated with citrate-based antigen unmasking solution (Vector, H-3300-250) in a steamer for 10 minutes, followed by hydrogen peroxide solution (Abcam, ab64218) for 10 minutes and 1% Triton X-100 in PBS for 10 minutes. The sections were treated with blocking reagent (Akoya Biosciences, FP1020) for one hour at room temperature before overnight incubation with the primary antibody diluted in the blocking reagent. The following day, the primary antibody was washed away with PBST (1×PBS with 0.1% Tween-20), and the slides were incubated with the secondary antibody diluted in the blocking reagent for two hours at room temperature. After washing away the secondary antibody with PBST, the slides were counterstained with DAPI (Thermo Fisher Scientific, D1306).
The primary antibodies used in this study were: rabbit recombinant monoclonal Sp7/Osterix (Abcam, ab209484, 1:100), mouse monoclonal myosin heavy chain (MHC) (DSHB, MF20, 1:20), rabbit monoclonal SOX9 (Abcam, ab185230, 1:100), rabbit polyclonal red fluorescent protein (RFP) (Rockland, 600-401-379, 1:500 with TSA), and rabbit monoclonal MyoD (Abcam, ab133627, 1:200 with TSA). The secondary antibodies were Alexa Fluor 488 Anti-Mouse (Thermo Fisher Scientific, A11001, 1:200), Alexa Fluor 568 Anti-Rabbit (Thermo Fisher Scientific, A11011, 1:200), and Anti-Rabbit HRP (Vector Laboratories, PI-1000, 1:200). For MyoD and RFP immunostaining, TSA Plus FITC or Cy3 (Akoya Bioscience, NEL7744001KT or NEL771B001KT) was applied for 3 minutes before DAPI counterstaining.
RNAScope in situ hybridization (ISH) assay
Cryosections were used for the RNAscope ISH assay. The slides were treated with a pre-heated target retrieval reagent (Advanced Cell Diagnostics, 322000) for 10 minutes in a steamer, and the signals were detected using the RNAscope Multiplex Fluorescent Detection Kit v2, following the manufacturer’s instructions (Advanced Cell Diagnostics, 323100). The following probes from Advanced Cell Diagnostics were used in this study: Mm-Tfap2b-O1 (535151), Mm-Sox6 (472061), Mm-Pax3 (455801), Mm-Hic1 (464131), and Mm-Myod1 (316081 or 316081-C2).
Imaging
The immunofluorescent and RNAScope in situ hybridization fluorescent signals on histological sections were captured using a Leica DMI3000 B research microscope.
scRNA-seq analysis
Tissues from the palatal region from E12.5, E13.5, E14.5, E15.5, and E18.5 embryos (3 biological replicates per stage) were dissected and dissociated using Multi Tissue Dissociation Kit 3 (130-110-204, Miltenyi Biotec) at 37 °C and 600 rpm for 15-30 minutes in a thermomixer (Thermo Fisher Scientific, 2231000269). The single-cell suspension was filtered using MACS® SmartStrainers (30 µm) (130-098-458 Miltenyi Biotec) and processed into a single-cell library using 10x Genomics Chromium single-cell 3’ v3 reagent kits (10x Genomics, PN-1000269) according to the manufacturer’s protocol. Sequencing was performed using a NovaSeq 6000 (Illumina) or NovaSeq XPlus (Illumina), and the quality control, sequence alignment, and read counts were processed and quantified using Cell Ranger (v9.0.0) through the 10x Genomics Cloud Analysis platform (10x Genomics). Integrative analysis of the scRNA-seq data from whole palatal tissue at E12.5, E13.5, E14.5, E15.5, and E18.5 was performed using the standard workflow of the Seurat v5 pipeline in RStudio18. Individual samples were processed as follows prior to integration. QC metrics were applied to retain cells with 200-7,500 unique genes (nFeature_RNA) and less than 5% mitochondrial reads (percent.mt), ensuring the removal of low-quality cells. Gene expression was normalized using LogNormalize, scaling to 10,000 counts per cell followed by log transformation. The top 2,000 most highly variable genes were identified for downstream analysis, and features were standardized through scaling. To correct for cell cycle effects, cells were scored for S and G2/M phases using the CellCycleScoring function, which assigned S.Score, G2M.Score, and phase identity (S, G2/M, or G1). Cell cycle effects were removed by regressing out S.Score and G2M.Score during data scaling with ScaleData, and the corrected data were used for downstream analyses. Cell clustering was performed using FindNeighbors (dims = 1:20) and FindClusters (resolution = 0.5), and UMAP was used to visualize clusters in low-dimensional space. The processed objects were saved as individual RDS files. For integration analysis, batch and grouped batch labels were assigned to individual RDS files based on sample identity and developmental stage, respectively. The datasets were merged into a single Seurat object using the merge function with unique cell IDs for each batch. Data preprocessing included normalization (NormalizeData), identification of highly variable features (FindVariableFeatures), and scaling (ScaleData). Dimensionality reduction was performed with PCA (RunPCA), and integration was achieved using IntegrateLayers with the CCA method to produce an integrated dataset (integrated.cca). Cell clustering was performed using FindNeighbors (dims = 1:20) and FindClusters (resolution = 0.5), with UMAP visualization (RunUMAP) to display clusters and batch information. FindMarkers and FindAllMarkers (min.pct = 0.25, logfc.threshold = 0.25, only.pos = TRUE) and were used to identify and differentially expressed positive marker genes for each cluster. The identified gene lists were further visualized with FeaturePlot and DotPlot, and the most specifically expressed genes were selected as cluster markers. Clusters were then renamed using RenameIdents to reflect their biological identities. Mesenchymal cell clusters were identified and extracted from the whole palate scRNA-seq data based on their specific cluster IDs. For more detailed analysis, datasets were processed using the same steps, including FindNeighbors (dim=20), FindClusters (resolution=0.5), and RunUMAP (dim=20). Similary, FindMarkers and FindAllMarkers (min.pct = 0.25, logfc.threshold = 0.25, only.pos = TRUE) were used to identified cluster-specific markers for each subpopulation, with their expression patterns visualized using FeaturePlot and DotPlot to further narrow down cluster-specific genes. Clusters were renamed using RenameIdents to reflect their biological identities. Metadata was grouped by developmental stages and cluster identities, and cell numbers were calculated using dplyr::summarize. Cell proportions were then determined by normalizing cluster counts to the total cell number per developmental stage using dplyr::mutate. To visualize cluster distributions across groups, a stacked bar chart was generated using ggplot2, with the geom_bar function displaying cluster-specific cell percentages across developmental stages.
SeqFISH analysis
SeqFISH-based spatial genomics imaging17 was performed by Spatial Genomics Inc. using a customized gene panel on cryosections of E12.5, E13.5, and E15.5 embryonic heads within a Gene Positioning System (Spatial Genomics). In vivo gene expression patterns of seqFISH were visualized using SGNlite software (v0.0.11) (Spatial Genomics). Clustering analysis was performed using the standard process in Scanpy (v1.10)21. The quality control criteria excluded cells with fewer than 7 expressed genes, fewer than 20 transcript counts, and areas smaller than 347 pixels (~3.7 µm²). A neighborhood graph was computed using principal components with 20 neighbors, and clustering was performed using the Leiden algorithm at a resolution of 1.0. The processed individual data were imported into SGNlite software (Spatial Genomics) to visualize spatially resolved clusters for each individual samples. The rank_genes_groups function was applied to identify differentially expressed genes between Leiden clusters, using the Wilcoxon rank-sum test. Dot plots were then generated to visualize the expression levels of the top differentially expressed genes across clusters, accompanied by a dendrogram depicting the hierarchical representation of the relationships between clusters. For integrative analysis of E12.5, E13.5 and E15.5 spatial genomic data, these data were integrated using the standard workflows of scvi-tools39. Cutoffs for cell filtering were determined using the formula: median ± ratio × MAD, with the ratio adjusted for each sample to account for variations in total transcript counts and cellular areas40. For anterior E12.5 samples, cells with fewer than 10 detected genes, total transcript counts <17 or >1,205, and cellular areas <930 or >19,187 μm² were excluded. For anterior E13.5, cells were filtered based on total transcript counts <13 or >1,115 and areas <235 or >20,318 μm². In anterior E15.5, cells with total transcript counts <10 or >926 and areas <318 or >20,539 μm² were removed. Similarly, for posterior E12.5, cells with fewer than 10 detected genes, total transcript counts <15 or >1,027, and areas <994 or >18,199 μm² were excluded. For posterior E13.5, thresholds were set at total transcript counts <15 or >920 and areas <753 or >18,498 μm², while posterior E15.5 cells with total transcript counts <11 or >1,051 and areas <917 or >20,638 μm² were removed. Integrated analysis of anterior and posterior sections from E12.5, E13.5, and E15.5 samples was performed using scvi-tools (v1.1) with default parameters (n_layers=2, n_hidden=30). The nearest neighbor graph was constructed using 20 neighbors, and Leiden clustering was applied with a resolution parameter of 4.0. To visualize spatial clusters at individual stages, cells from E12.5, E13.5, and E15.5 were filtered using the isin function, saved as separate H5AD files, and visualized with TissUUmaps 326. Spatial genomic samples processed in Scanpy were CPM normalized and log-transformed, whereas the integration of multiple spatial samples using scvi-tools utilized raw counts, with normalization managed by the variational autoencoder through iterative learning.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw and processed single-cell RNA sequencing (scRNA-seq) data from embryonic palatal tissue at different developmental stages (E12.5, E13.5, E14.5, E15.5, and E18.5) generated in this study has been deposited in the Gene Expression Omnibus (GEO) database under accession number GSE293181 and FaceBase under the DOI 10.25550/62-QZ1A41 [https://www.facebase.org/chaise/record/#1/isa:dataset/RID=62-QZ1A]. The raw and processed sequencing-FISH data for embryonic heads at E12.5, E13.5, and E15.5 are available in the FaceBase database under the DOI 10.25550/62-Y0VT42 [https://www.facebase.org/chaise/record/#1/isa:dataset/RID=62-Y0VT].
Code availability
All R and Python scripts supporting the findings of this paper are accessible in a GitHub repository (https://github.com/ChaiLabUSC/code.git) and on Zendo through the DOI 10.5281/zenodo.1508599643.
References
Soldatov, R. et al. Spatiotemporal structure of cell fate decisions in murine neural crest. Science 364, 10.1126/science.aas9536 (2019).
Achilleos, A. & Trainor, P. A. Neural crest stem cells: discovery, properties and potential for therapy. Cell Res 22, 288–304 (2012).
Chai, Y. et al. Fate of the mammalian cranial neural crest during tooth and mandibular morphogenesis. Development 127, 1671–1679 (2000).
Erickson, A. G., Kameneva, P. & Adameyko, I. The transcriptional portraits of the neural crest at the individual cell level. Semin Cell Dev Biol 138, 68–80 (2023).
Kaucka, M. et al. Analysis of neural crest-derived clones reveals novel aspects of facial development. Sci Adv 2, e1600060 (2016).
Baroffio, A., Dupin, E. & Le Douarin, N. M. Common precursors for neural and mesectodermal derivatives in the cephalic neural crest. Development 112, 301–305 (1991).
Bronner-Fraser, M. & Fraser, S. E. Cell lineage analysis reveals multipotency of some avian neural crest cells. Nature 335, 161–164 (1988).
Li, J. et al. Regulatory Mechanisms of Soft Palate Development and Malformations. J Dent Res 98, 959–967 (2019).
Han, X. et al. Runx2-Twist1 interaction coordinates cranial neural crest guidance of soft palate myogenesis. Elife 10, 10.7554/eLife.62387 (2021).
Dixon, M. J., Marazita, M. L., Beaty, T. H. & Murray, J. C. Cleft lip and palate: understanding genetic and environmental influences. Nat Rev Genet 12, 167–178 (2011).
Ozekin, Y. H., O’Rourke, R. & Bates, E. A. Single cell sequencing of the mouse anterior palate reveals mesenchymal heterogeneity. Dev Dyn 252, 713–727 (2023).
Piña, J. O. et al. Multimodal spatiotemporal transcriptomic resolution of embryonic palate osteogenesis. Nat Commun 14, 5687 (2023).
Yan, F. et al. Single-cell multiomics decodes regulatory programs for mouse secondary palate development. Nat Commun 15, 821 (2024).
Jin, J. Z. & Ding, J. Analysis of Meox-2 mutant mice reveals a novel postfusion-based cleft palate. Dev Dyn 235, 539–546 (2006).
Xu, J. et al. Shox2 regulates osteogenic differentiation and pattern formation during hard palate development in mice. J Biol Chem 294, 18294–18305 (2019).
Eng, C. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH. Nature 568, 235–239 (2019).
Lubeck, E., Coskun, A. F., Zhiyentayev, T., Ahmad, M. & Cai, L. Single-cell in situ RNA profiling by sequential hybridization. Nat Methods 11, 360–361 (2014).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol 42, 293–304 (2024).
Feng, J. et al. TGF-β signaling and Creb5 cooperatively regulate Fgf18 to control pharyngeal muscle development. Elife 11, 10.7554/eLife.80405 (2022).
Lee, Y. H. et al. Early requirement of the transcriptional activator Sox9 for neural crest specification in Xenopus. Dev Biol 275, 93–103 (2004).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol 19, 15 (2018).
Jin, A. et al. ScRNA-Seq Reveals a Distinct Osteogenic Progenitor of Alveolar Bone. J Dent Res 102, 645–655 (2023).
Li, I. M. H. et al. Differential tissue specific, temporal and spatial expression patterns of the Aggrecan gene is modulated by independent enhancer elements. Sci Rep 8, 950 (2018).
Liu, F. et al. Wnt/beta-catenin signaling directs multiple stages of tooth morphogenesis. Dev Biol 313, 210–224 (2008).
Ohki, S. et al. Transforming growth factor-beta and sonic hedgehog signaling in palatal epithelium regulate tenascin-c expression in palatal mesenchyme during soft palate development. Front Physiol 11, 532 (2020).
Pielawski, N. et al. TissUUmaps 3: Improvements in interactive visualization, exploration, and quality assessment of large-scale spatial omics data. Heliyon 9, e15306 (2023).
Jing, J. et al. Spatiotemporal single-cell regulatory atlas reveals neural crest lineage diversification and cellular function during tooth morphogenesis. Nat Commun 13, 4803 (2022).
Nakamura, E., Nguyen, M. T. & Mackem, S. Kinetics of tamoxifen-regulated Cre activity in mice using a cartilage-specific CreER(T) to assay temporal activity windows along the proximodistal limb skeleton. Dev Dyn 235, 2603–2612 (2006).
Biben, C. et al. In vivo clonal tracking reveals evidence of haemangioblast and haematomesoblast contribution to yolk sac haematopoiesis. Nat Commun 14, 41 (2023).
Li, L. et al. A mouse model with high clonal barcode diversity for joint lineage, transcriptomic, and epigenomic profiling in single cells. Cell 186, 5183–5199.e5122 (2023).
Yuan, Y. et al. Spatiotemporal cellular movement and fate decisions during first pharyngeal arch morphogenesis. Sci Adv 6, 10.1126/sciadv.abb0119 (2020).
Kulesa, P. M. & Gammill, L. S. Neural crest migration: patterns, phases and signals. Dev Biol 344, 566–568 (2010).
Chai, Y., Sanchez, P. & Thomas, P. Integrated research of functional genomics and craniofacial morphogenesis. FaceBase Consortium, https://doi.org/10.25550/1WWE (2017).
Farmer, D. T. et al. The developing mouse coronal suture at single-cell resolution. Nat Commun 12, 4797 (2021).
Xu, Z. et al. Embryonic attenuated Wnt/β-catenin signaling defines niche location and long-term stem cell fate in hair follicle. Elife 4, e10567 (2015).
Zhang, M., Feng, J., Li, Y., Qin, P. Z. & Chai, Y. Generation of tamoxifen-inducible Tfap2b-CreER. Genesis 62, e23582 (2024).
Scott, R. W., Arostegui, M., Schweitzer, R., Rossi, F. M. V. & Underhill, T. M. Hic1 Defines Quiescent Mesenchymal Progenitor Subpopulations with Distinct Functions and Fates in Skeletal Muscle Regeneration. Cell Stem Cell 25, 797–813.e799 (2019).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 13, 133–140 (2010).
Gayoso, A. et al. A Python library for probabilistic analysis of single-cell omics data. Nat Biotechnol 40, 163–166 (2022).
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat Rev Genet 24, 550–572 (2023).
Feng, J. & Chai, Y. scRNA-seq analysis of E12.5, E14.5, E15.5, and E18.5 mouse whole palate. FaceBase Consortium, https://doi.org/10.25550/62-QZ1A (2025).
Feng, J. & Chai, Y. High-resolution spatial profiling and cell lineage analysis of mouse palate at E12.5, E13.5, and E15.5. FaceBase Consortium, https://doi.org/10.25550/62-Y0VT (2025).
Feng, J. & Chai, Y. High-resolution spatial transcriptomics and cell lineage analysis reveal spatiotemporal cell fate determination during craniofacial development. Zenodo https://doi.org/10.5281/zenodo.15085996 (2025).
Acknowledgements
We are grateful to Bridget Samuels for critical reading of this manuscript and to Kimi Nakaki for the schematic illustrations. We acknowledge Spatial Genomics Inc. for their assistance with the bioinformatic analysis of the seqFISH data. This work was supported by the National Institute of Dental and Craniofacial Research, National Institutes of Health (R01 DE012711 and U01 DE028729 to Yang Chai).
Author information
Authors and Affiliations
Contributions
J.F., E.J., and Y.C. conceived the study. J.F., E.J., J.J.G., A.A.-V., and M.L. collected samples. J.F., E.J., T.G., H.Z., M.Z., S.C., and T.-V.H. analyzed the data. J.F., E.J., and Y.C. wrote the manuscript, with input from all authors. Y.C. acquired funding and supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Feng Bao, Robert Lipinski and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, J., Janečková, E., Guo, T. et al. High-resolution spatial transcriptomics and cell lineage analysis reveal spatiotemporal cell fate determination during craniofacial development. Nat Commun 16, 4396 (2025). https://doi.org/10.1038/s41467-025-59206-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59206-2