Abstract
Functional circadian clock is critical to the adaptation and survival of organisms. In land plants, the comprehensive profiling of circadian gene expression at the single-cell level is largely unknown partly due to the challenges in obtaining precisely-timed single cells embedded within cell walls. To bridge this gap, we employ time-lapse single-nucleus RNA sequencing (snRNA-seq) on Arabidopsis seedlings collected over a 48-hour window at 4-hour intervals, as well as over a 24-hour day at 2-hour intervals, yielding a total of over 77,142 and 130,000 nuclei. Here, we find that four cell clusters in the shoot share a coherent rhythm, while around 3000 genes display cell-type specific rhythmic expression. Our analysis indicates that genes encoding circadian regulators oscillate in multiple cell types, and the majority of them are well-documented core clock genes, suggesting the snRNA-seq circadian data could be used to identify more clock components oscillating in a cell-autonomous way. We identify ABF1 as a circadian regulator, whose overexpression and shortens the circadian period. Our data provides a comprehensive resource for plant circadian rhythmicity at the single-cell level (hosted at https://zhailab.bio.sustech.edu.cn/sc_circadian).
Similar content being viewed by others
Introduction
The circadian clock is a time-keeping mechanism that coordinates 24 h rhythms synchronized with environmental cycles, which is essential for the temporal organization of physiology in organisms1,2,3. The core clock genes form the transcriptional-translational feedback loops (TTFLs), which regulates gene expression from dawn till late night, driving metabolism and growth in synchrony with the day-night cyclic cues4,5,6,7. Within plant systems, the circadian clocks regulate distinct rhythmicity and functions among different tissues and cell types8,9,10,11,12,13.
In plants, studies on circadian rhythms have extensively utilized techniques such as bioluminescence tracking of promoter:LUC and real-time quantitative PCR (qRT-PCR) to monitor the promoter dynamic activity of the reporter genes or the transcripts rhythmic enrichment in whole seedlings and intact tissues14,15,16,17,18,19. Despite these findings, these studies obscured the differences in circadian rhythms at the individual cell level. Previous studies using long-term imaging approaches have provided glimpses into this complexity18,20,21,22,23,24,25,26. However, these methods, while informative, have lacked the throughput to comprehensively dissect the operation of the clock system at the single-cell level.
The advent of single-cell RNA sequencing (scRNA-seq) has revolutionized our understanding of cellular heterogeneity, providing a lens to view gene expression dynamics at an unprecedented resolution27,28,29. In recent years, scRNA-seq has been used to delineate circadian gene expression in animals, providing insight into the temporal regulation within individual cells with samples collected in 4 h intervals30,31 or at two time points32,33. A recent study in plants also used scRNA-seq to examine protoplasts of Arabidopsis shoots and roots collected at the end of the day or end of night34. However, plant cells, encased by rigid cell walls, pose unique challenges for short-interval time-series analysis, because the protoplasting process required for plant single cell isolation can take several hours and therefore unideal for capturing the time-series transient transcript changes of circadian regulation. To overcome issues induced by protoplasting, a recent study by Torii et al. manually extracted cellular content from 216 single cells collected at different time points using microcapillary and constructed scRNA-seq libraries for each cell, and their results shed light on the pivotal role of the circadian clock in leaf cell differentiation processes35. However, a comprehensive atlas of single-cell circadian gene expression for a whole Arabidopsis seedling remains to be developed.
In this study, we have overcome these limitations by employing single-nucleus RNA sequencing (snRNA-seq), a technique less impeded by the physical constraints of plant cell walls. By applying snRNA-seq to Arabidopsis seedlings, we have mapped circadian gene expression in different cell types with high temporal resolution at two-hour and four-hour intervals. Our 24 h snRNA-seq results reveal significant oscillator similarities among four shoot cell clusters and identify approximately 3000 genes that oscillate within a specific cell cluster. Furthermore, we detected genes oscillating in multiple cell types, the majority of them being well-known core clock genes, highlighting the potential of using snRNA-seq to study the hierarchical clock network in plants. Among these, we identified ABF1 as a novel circadian regulator. Overexpression of ABF1 shortens the circadian period, indicating its role in modulating the plant circadian rhythm.
Through this comprehensive dataset, we provide an invaluable resource for further investigation into plant circadian rhythmicity at the single-cell level, offering a new dimension to understanding how temporal regulation can influence plant biology at the most fundamental level. Our findings represent a considerable step forward in plant chronobiology, promising to enrich our understanding of plant physiology and potentially guiding agricultural practices to harness the innate timing mechanisms of crops for optimized growth and yield.
Results and discussion single-nucleus RNA-seq reveals circadian rhythms in Arabidopsis
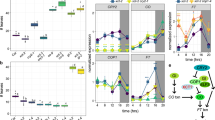
To craft a detailed single-cell transcriptomic blueprint of temporal mRNA rhythmic accumulation via the circadian clock, we leveraged the 10X Genomics platform to execute snRNA-seq on whole seedlings, which were harvested every 4 h across 13 circadian time points, as well as every 2 h across 12 circadian time points under constant light (LL; free-running conditions), respectively (Fig. 1a). After performing quality control and computing the expression matrix, we acquired a 24 h dataset of 131,152 high-quality single-nucleus transcriptomes spread across twelve libraries, encompassing 24,503 genes, with an average of 847 genes and 1058 UMI detected per nucleus (Supplementary Data 1). We also obtained a 48-hour dataset of 77,142 nuclei, encompassing 28,813 genes, with an average of 2359 genes and 7131 UMI detected per nucleus (Supplementary Data 1). Both of our data revealed pronounced circadian rhythm patterns, with increased transcriptomic similarity between adjacent circadian time points, regardless of the sample they originated from (Fig. 1b, c). This confirmed that our approach effectively captured the dynamic changes in clock gene expression. To further validate our methodology, we employed a pseudo-bulking approach to aggregate gene counts in each sample. As expected, the expression patterns of 17 core clock genes7 (CCA1, LHY, RVE4, RVE8, RVE6, LNK1, LNK2, PRR9, PRR7, PRR5, PRR3, TOC1, GI, LUX, ELF4, ELF3 and ZTL) showed robust oscillations peaking at each anticipated circadian phase (Fig. 1d, e), such as CCA1 and TOC1, peaking at subjective dawn and dusk, respectively (Fig. 1f, g). Collectively, these findings underscore the feasibility of our approach in analyzing circadian rhythms using single-nucleus transcriptional profiling.
a Experimental design and methodology. Schematic representation of the experimental setup for circadian entrainment, sampling, and snRNA-seq processing. Seedlings were entrained under a 12 h light/12 h dark cycle before being transferred to continuous light conditions for circadian rhythmicity analysis. The snRNA-seq libraries were conducted using the 10X Genomics platform, followed by high-throughput sequencing and detailed data analysis. b, c Cross-sample correlation analysis in the 24 h dataset (b) and the 48 h dataset (c). The heatmap presents alignment scores between each sample pair, where these scores represent the proportion of cells identified as mutual nearest neighbors across datasets. Higher scores signify greater transcriptomic similarity between datasets. Source data are provided as a Source Data file. d, e Heatmap showing the min-max normalized average expression profiles of core clock genes across 12 circadian time points (d) and 13 circadian time points (e). Source data are provided as a Source Data file. f, g Line plots illustrating the transcript accumulation of CCA1 and TOC1 in the 24-hour dataset (f) and the 48 h dataset (g), with each point representing the mean expression across all nuclei in a sample. Source data are provided as a Source Data file.
A recent study in mammals suggests that the circadian clock can affect cell clustering, thus interfering with subsequent analysis30. The similarity between samples at adjacent time points suggested the presence of such a phenomenon in our data (Fig. 1b, c) and the UMAP visualization also indicates that the baseline is different among samples (Supplementary Fig. 1). To eliminate this effect and improve clustering results, we evaluated five data integration strategies, which can be categorized into four groups (Supplementary Fig. 2a): (1) Direct application of the Harmony method to integrate data; (2) Following the approach proposed by Wen et al. 30, exclusion of clock-regulated genes during the clustering step; (3) Individual application of SCTransform36 on each sample to eliminate baseline variations across the data; (4) Employing a procedure analogous to the removal of cell cycle effects37, performing SCTransform separately on each sample followed by linear regression to regress out the expression of core clock genes7. Next, we evaluated these strategies from the perspective of conserving biological variation and eliminating batch effects. We first performed clustering on each dataset individually (Supplementary Fig. 3), and then used the silhouette score to evaluate the performance of these clustering results on the cell distance matrices derived from various strategies (Supplementary Fig. 4). This score is a metric to gauge the quality of clustering results by considering distances between clusters and clusters. Clustering performed on each library individually is expected to be more sensitive to biological variations. Therefore, a lower silhouette score suggests that the original cell clusters become indistinguishable within the integrated dataset, indicating a loss of discernible biological differences. Consequently, in this context, the silhouette score acted as an indicator for preserving biological variability. In addition, we employed the kBET38 method to measure the concordance of different samples in the final distance matrix, serving as a metric for evaluating the batch effect removal (Supplementary Fig. 4). We observed that the exclusion of clock-regulated genes enhanced the performance of batch effect removal, and removing an equal number of randomly selected genes does not have the impact on batch effect removal (Supplementary Fig. 2b). This indicates that circadian rhythms are the main cause of sample heterogeneity. In addition, the removal of clock-regulated genes resulted in a huge decrease in silhouette score, indicating the impaired biological variation (Supplementary Fig. 2b). However, the result with the removal of randomly selected genes indicated that this decrease is not solely due to the reduction in the number of genes. Strategies 3 and 4 both exhibited enhancements in both biological conservation and batch effect removal when compared to directly integrating datasets, thus achieving a balance between preserving biological characteristics and removing batch effects. Given that Strategy 3 yielded better biological conservation scores and batch correction scores than Strategy 4 (Supplementary Fig. 4), we selected Strategy 3 as our final strategy.
Following data integration and clustering, we identified 40 distinct clusters from the 48-hour dataset and 27 clusters from the 24 h dataset (Fig. 2a and Supplementary Fig. 8b). Each cluster contained nuclei from all time points with minimal variance (Supplementary Fig. 5). To annotate the identities of each cluster, we first identified cluster-specific expression genes (Supplementary Fig. 6 and Supplementary Data 2), then constructed a pseudo-seedling single-cell dataset by merging two public single-cell root and single-cell shoot datasets39,40. Within this synthesized dataset, we identified genes specifically expressed in each cluster and employed the Jaccard Index as a metric to assess the overlap between these specifically expressed genes in our dataset and the pseudo-seedling single-cell dataset. This method guided our annotation, assigning cell types to clusters based on the highest overlap in gene expression (Supplementary Fig. 7). Through this process, we were able to identify various cell types from mesophyll, stomata, epidermis, cortex, endodermis, vascular bundles, and the meristem (Supplementary Fig. 7). Furthermore, we were able to distinguish the root or shoot origin for all clusters, except for those pertaining to vascular tissue (Supplementary Fig. 7), aligning with a previous report that vascular cells from shoots and roots tend to cluster together39. Clusters are labeled with “R”, “S”, and “V” to represent cell types derived from roots, shoots, and vascular tissues, respectively (Fig. 2a and Supplementary Fig. 8a). Clusters labeled “U” correspond to cells with origins from the shoot and root that remain to be determined (Fig. 2a). The term “Unknown” is reserved for cells with unidentified cell types (Fig. 2a and Supplementary Fig. 8a). For example, cluster U2 represents unknown cell types, while cluster U1 consists of epidermal cells whose tissue origins are unknown (Fig. 2a). We have also organized clusters by cell type to ensure that clusters representing the same cell type are grouped together (Fig. 2a and Supplementary Fig. 8a). As mentioned earlier, the 48 h dataset is of higher quality, with most clusters successfully annotated (Fig. 2a). To identify previously unannotated cell types, we conducted parallel annotation of the higher-quality 48 h dataset. We applied the scANVI method to project the high-sampling-density 24 h dataset onto the 48 h dataset, obtaining label-transferred annotations. Our analysis revealed that the label transfer results for the majority of clusters were consistent with our previous annotations (Supplementary Fig. 9). In addition, this approach enabled us to annotate some previously unannotated clusters, such as clusters S8 and S10, identified as mesophyll cells, and cluster R5, annotated as endodermal cells (Supplementary Fig. 9). Furthermore, RCA (RUBISCO ACTIVASE), a mesophyll cell marker identified in the stereo-seq dataset41, showed relatively highly expressed in mesophyll-related clusters in the 24 h dataset, including cluster S8, S0, S10, S2 and S4 (Supplementary Fig. 10b), further supporting our annotation of cluster S8 and cluster S10. In addition, we found a subset of cells within cluster S8 (24 h dataset) expressing YAB1 when we generated subclusters (cluster S6, S8 and V3) according to the expression of YAB1 and SWEET12 (phloem cell marker42) (Supplementary Fig. 10a). An in situ hybridization study have shown that YAB1 is expressed in the peripheral region of young leaf primordia and later in the abaxial side of developing leaves and floral organs43. We also analyzed stereo-seq data from leaves collected from 6-week-old Arabidopsis plants41, which revealed that YAB1 is specifically expressed in mesophyll and epidermal cells (Supplementary Fig. 10c). In addition, we selected two cluster S8-specific genes, AT1G56650 and AT4G14090, which were detectable in the stereo-seq dataset and had the highest specific scores based on cellex analysis (Supplementary Data 2). Both genes showed specific expression in mesophyll and epidermal cells of leaves. Based on this comprehensive analysis, we have reannotated cluster S8 (24-hour dataset) as mesophyll cell and epidermal cells of primordia of leaves (Supplementary Fig. 8). Furthermore, we incorporated 28 well-characterized marker genes (Supplementary Data 3) to verify cluster classifications for both datasets (Supplementary Fig. 11). Through this analysis, we confirmed the cell identities of the major cell types, and identified cluster U1 (48 h data) as endodermal cells and cluster U0 (48 h data) as epidermal cells (Fig. 2a and Supplementary Fig. 11b). We also performed the β-glucuronidase (GUS) staining of three cluster-specific genes, AT1G66100, CEPD1 (AT1G06830), and KCS6 (AT1G68530) to further validate our annotations. The results from these experiments showed expression patterns consistent with our annotations. Specifically, GUS staining of CEPD1 (Supplementary Fig. 12e) demonstrated its enrichment in cluster V2 (24 h data) (Supplementary Fig. 12d), corresponding to vascular cells, while AT1G66100 (Fig. 3H) and KCS6 (Supplementary Fig. 12a) were enriched in cluster S1 (24 h data), corresponding to epidermal cells (Fig. 3e and Supplementary Fig. 12b and c). Taken together, we effectively revealed cell heterogeneity and classified the major cell types in Arabidopsis seedlings.
a UMAP plot of 40 clusters in the 48 h dataset. Clusters are labeled with “R”, “S”, and “V” to represent cell types derived from roots, shoots, and vascular tissues, respectively. Clusters labeled “U” correspond to cells with origins from the shoot and root that remain to be determined. The term “Unknown” is reserved for cells with unidentified cell types. b Expression of CCA1 and TOC1 across different cell types. Heatmaps representing the mean expression level of the same gene across different clusters at 13 circadian time points. An asterisk (*) indicates clusters where the gene is oscillating. Source data are provided as a Source Data file.
a–c Line plots showing the average rhythms of RBCS2B:LUC-GUS luminescence (a) AT1G66100:LUC-GUS luminescence (b) and CAB2:LUC-GUS luminescence (c) in transgenic plants (blue lines) with two biological replicates. These rhythms are compared with the pseudo-bulk expression from snRNA-seq data (orange lines), highlighting similarities and discrepancies between in vivo luminescence and sequencing data. Source data are provided as a Source Data file. d–f UMAP plots representing the specific expression patterns of RBCS2B (AT5G38420) (d), AT1G66100 (e) and CAB2 (AT1G29920) (f) within the 24 h snRNA-seq dataset. g–i Histochemical GUS assays of transgenic seedlings harboring RBCS2B:LUC-GUS (g), AT1G66100:LUC-GUS (h) and CAB2:LUC-GUS (i) reporter constructs, demonstrating cell-type-specific expression patterns. Each experiment was independently repeated three times (n = 9 samples), yielding similar results.
Analyzing clock gene expression in different cell types
To explore circadian rhythm variations across different cell types, we collected 13,256 known clock-regulated genes identified from three global studies44,45,46. Using the JTK-cycle algorithm47, we assessed the rhythmicity of these genes across both datasets. Our analysis revealed that approximately 40% (5406/13,256) of circadian clock-regulated genes displayed transcript rhythmic accumulation in at least one cluster in the 24 h dataset (Supplementary Data 4), and 9832 genes oscillated in the 48 h dataset (Supplementary Data 4). We found that shoot clusters demonstrated a higher prevalence of oscillating genes compared to root clusters. Specifically, mesophyll cells in cluster S2 from the 48 h dataset contained 3956 oscillating genes, while mesophyll cells in cluster S0 from the 24 h dataset also showed a high number, with roughly 2400 oscillating genes (Supplementary Fig. 13). In addition, our data could also be used to identify transcripts that are stable and highly expressed across all cell types. We analyzed both datasets by calculating the average expression (CPM) for each gene across all samples and selected the top 5% as highly expressed. Among these high-expression genes, we computed the standard deviation of log₂-transformed CPM values across all cell clusters and samples, identifying those with a standard deviation below 0.5 in the 48 h dataset and below 0.2 in the 24 h dataset as stably and highly expressed. This resulted in 50 and 47 stably expressed genes in the 48 h and 24 h dataset, respectively (Supplementary Data 5), which could serve as valuable references for normalizing qPCR results and RNA-seq experiments. Next, we compared the oscillation patterns of core clock genes across different cell types. Our findings revealed that these genes exhibited different oscillation patterns across a variable number of clusters (Supplementary Fig. 14 and Supplementary Fig. 15). We selected CCA1 and TOC1 as representative clock genes, which displayed expression patterns with maximum expression phases at subjective dawn and dusk, respectively (Fig. 2b and Supplementary Fig. 8b). Although these genes were detected in nearly all clusters, the extent of their oscillation across clusters showed considerable variation. Specifically, CCA1 oscillated in 20 clusters in the 24 h dataset and 28 clusters in the 48 h dataset, while TOC1 was confined to oscillations in only 13 and 23 clusters, respectively (Fig. 2b and Supplementary Fig. 8b). This diversity partially underscores the heterogeneity of the functional circadian clock across various cell types. The phase and amplitude analysis of the 24 h rhythms for each cluster in the 24 h dataset revealed that several genes including CCA1, LHY, and PRR3 showed more robust oscillation in a mesophyll subcluster (cluster S0 in both datasets) compared with other cell types (Supplementary Fig. 16a and Supplementary Fig. 17a). In addition, we also observed phase shifts among clock gene oscillations across different cell types, such as the transcript accumulation of TOC1, which displays a phase delay in shoot epidermal cells (cluster S3, S5 and S14 in the 48 h dataset, cluster S1 in the 24 h dataset) relative to other shoot cell types (Supplementary Fig. 16b and Supplementary Fig. 17b). Furthermore, we observed considerable variations in phase and amplitude among different vascular cell types for rhythmic expression of clock genes in multiple vascular clusters (Supplementary Fig. 16 and Supplementary Fig. 17). Collectively, these findings provide insights that advance our understanding of cell-type specific oscillators and the multi-oscillator nature and organization of the plant circadian network.
To corroborate the oscillatory patterns found in our dataset, we generated transgenic lines expressing luciferase (LUC) and GUS encoding genes driven by the promoters of RBCS2B (AT5G38420), CEPD1, CAB2 (AT1G29920), KSC6, and AT1G66100. This enabled us to spatially map expression locations and concurrently monitor the rhythmicity of these genes (Fig. 3). We measured the expression levels of these genes under LL conditions and observed a high degree of reproducibility across two biological replicates for each gene (Fig. 3a–c). For RBCS2B, CAB2, and AT1G66100, which oscillated in our snRNA-seq data, we assessed their promoter activity and found that these genes exhibited a largely similar oscillating pattern with the 24 h snRNA-seq data (Fig. 3a–c and Supplementary Fig. 18). It is worth noting that the promoter activity measured at the protein level by LUC assay may not always be the same as RNA-seq measured at the transcript level, and a certain degree of discrepancies are to be expected. In addition, histochemical GUS assays for tissue specificity also aligned with our cell type annotations, supporting the accuracy of our cluster classification (Fig. 3g–i, Supplementary Fig. 12 and Supplementary Fig. 18). The expression of RBCS2B and CAB2, as indicated by GUS staining, was predominant in shoot-specific cell types (Fig. 3g, i), corroborating our snRNA-seq findings (Fig. 3d, f and Supplementary Fig. 18a, c). Similarly, KCS6 and AT1G66100 confirmed their epidermal localization (Fig. 3h and Supplementary Fig. 12b, c), which was in agreement with our initial cluster classification (Fig. 3e and Supplementary Figs. 12a, 18b). Collectively, these validations underscore the reliability of snRNA-seq for capturing the intricate spatiotemporal expression of clock-regulated genes within Arabidopsis.
In addition, we observed the overall circadian rhythm peaks in our snRNA-seq data are broader than those observed in the LUC experiments. To investigate whether this broadening is due to clusters containing multiple cell types with different phases, we performed an analysis focusing on cells expressing well-known cell-type-specific markers. We selected seven marker genes representing seven distinct cell types (Supplementary Fig. 19 and Supplementary Fig. 20). We compared the expression patterns of two genes from our LUC results (as shown in Fig. 3), AT1G6610 and RBCS2B, in cells expressing these markers versus the corresponding annotated clusters. For the epidermal cell-specific gene AT1G6610 (cluster S1 in the 24 h dataset, and cluster S4 in the 48 h dataset), we found that its expression in cells expressing the epidermal marker gene AMTL148 closely matched its expression trend within the annotated cluster S1 (Supplementary Fig. 19a and Supplementary Fig. 20a). For RBCS2B, which is expressed across all cell types in the shoot (Fig. 3g), we observed that its expression patterns in most cell types showed similar trends between cluster-based and marker-based analyses (Supplementary Fig. 19b–k and Supplementary Fig. 20b–r). These results suggest that the broader circadian rhythm peaks observed in our cluster-based analysis are not primarily due to clusters containing multiple cell types with distinct phases. Instead, the accuracy of our cluster separation appears robust, as evidenced by the consistent expression patterns of cell-type-specific marker genes. In addition, we analyzed marker gene expression across different clusters. We found that most clusters predominantly express markers for a single cell type (Supplementary Fig. 11). This further supports that most clusters consist of homogeneous cell populations.
Four mesophyll cell types exhibit synchronized oscillation
In a comparative analysis of gene oscillation across different cell types, we employed the S-MOD method49, to statistically assess the overlap of oscillating genes (Fig. 4a and Supplementary Fig. 21a). This analysis delineated a clear demarcation between shoot and root cells in both datasets, with a pronounced intra-tissue oscillating gene congruence within shoots (Fig. 4b and Supplementary Fig. 21a). We observed higher statistical relatedness among four mesophyll cell types (clusters S0, S1, S2, and S7) in the 48 h dataset. Furthermore, pairwise phase analysis of common oscillating genes in these clusters revealed a phase lag in clusters S0 and S1 compared to S2 and S7 (Fig. 4c). Specifically, there were 2092, 1394, 1922, and 1305 common oscillating genes were identified when comparing clusters S0 and S2, clusters S0 and S7, clusters S1 and S2, clusters S1 and S7, respectively. Of these, 581, 324, 344, and 208 genes exhibited significant phase shifts, with most showing delayed phases in clusters S0 and S1 (Fig. 4c). However, the phase of 95% of genes between clusters S0 and S1, and 92% between clusters S2 and S7, showed no significant difference (Fig. 4c). Furthermore, a subset of shoot cells in the 24 h dataset – mesophyll, proliferating, meristematic, endodermal cells (clusters S2, S4, S6 and S8) – exhibited substantial relatedness of oscillating gene content (Supplementary Fig. 21a). Subsequently, pairwise phase analysis of the common oscillating genes identified from these clusters was performed (Supplementary Fig. 21b). Our findings showed that between proliferating and mesophyll cells (cluster S2) and endodermis and meristem cells (cluster S6), proliferating and mesophyll cells (cluster S4) and endodermis and meristem cells (cluster S6), proliferating and mesophyll cells (cluster S2) and mesophyll cells (cluster S8), proliferating and mesophyll cells (cluster S4) and mesophyll cells (cluster S8), endodermis and meristem cells (cluster S6) and mesophyll cells (cluster S8), and proliferating and mesophyll cells (cluster S2) and proliferating and mesophyll cells (cluster S4), there were 125, 110, 208, 179, 124, and 270 common oscillating genes, respectively, with 27, 12, 53, 26, 14 and 19 of these genes exhibiting substantial phase shifts. Predominantly, most of the genes with phase differences showed a significant phase lag mainly in endodermis and meristem cells (cluster S6) when compared to proliferating and mesophyll cells (cluster S2 and S4), and mesophyll cells (cluster S8) (Supplementary Fig. 21b). We also observed a phase delay of most genes in a subtype of mesophyll cells (cluster S8) compared to proliferating and mesophyll cells (cluster S2 and S4) (Supplementary Fig. 21b). These results suggest a delayed oscillatory phase in endodermis and meristem cells compared to a subtype of mesophyll cells, which itself has a delayed phase when compared to proliferating and mesophyll cells (Supplementary Fig. 21b). The 48 h dataset contains more clusters and finer resolution (Fig. 2a), which may contribute to the differences observed in the S-MOD results compared to the 24 h dataset. Conclusively, our study delineates a clear distinction in oscillating gene content between shoot and root cells, with subsets of shoot cells exhibiting markedly enhanced oscillator coherence, underscoring a complex, tissue-specific regulation of gene oscillation.
a Summary of the S-MOD calculation for pairwise statistical overlap between oscillating genes. The significance of the overlap (p-value) between two clusters was calculated using various parameters: n1 and n2 represent the number of oscillating genes in the two respective gene sets (A1 and A2), m is the count of common oscillating genes shared by the clusters, and N denotes the total number of oscillating genes identified in the entire dataset. The S-MOD score is defined as the negative logarithm of the p-value, offering a clear measure of the statistical significance of the overlap. b Heatmap displaying the S-MOD scores for pairwise comparisons of oscillating genes across 40 clusters in the 48 h dataset. To facilitate easy differentiation, clusters originating from the shoot, root, stele, and unknown cell types are color-coded in blue, red, pink, and gray, respectively. Source data are provided as a Source Data file. c Pairwise phase analysis between four mesophyll cell subclusters (cluster S0, S1, S2, and S7) of the 48-hour dataset. The number of genes (n) showing significant phase shifts is indicated above each plot, with the blue dots representing genes with statistically significant phase differences (Sig), and gray dots indicating non-significant phase relationships (NS). For this analysis, the significance of phase differences was assessed using a two-tailed z-test. Source data are provided as a Source Data file.
snRNA-seq identifies the circadian clock-controlled genes in various cell types
Observing the rhythmic expression of multiple core clock genes across various cell types, we next quantified the number of clusters where each gene oscillated. Our findings indicated that approximately 30% (2940/9832) of genes in the 48 h dataset, and 61% (3286/5406) of genes in the 24 h dataset only oscillated in a specific cluster (Fig. 5a and Supplementary Fig. 23), ~18% (532/2940) and 11% (374/3286) of which are cluster-specific genes in the 48 h and 24 h dataset, respectively (Supplementary Fig. 22). This finding of cell-type specific oscillation is consistent with a previous bulk microarray study in Arabidopsis, which indicated limited clock-regulated gene overlap between mesophyll cells and the vasculature ones19. A similar phenotype has also been reported in the Drosophila clock system, where the oscillating genes exhibit minimal consistency across cell types31.
a Left: Bar plot illustrating the quantity of oscillating genes identified across varying numbers of clusters in the 48 h dataset. Right: Scatter plot representing the correlation between the pseudo-bulk expression level of each oscillating gene and the corresponding count of clusters where the gene is identified as oscillating. Core clock genes are highlighted in dark blue, other regulators within the top 50 genes are highlighted in orange, ABF1 is highlighted in red, other genes are shown in light blue. b Ranking of transcription factors from PlantTFDB in the 48 h dataset (upper panel) and 24 h dataset (lower panel). Core clock genes are highlighted in dark blue; genes regulating circadian rhythms are colored in orange; other transcription factors are shown in light blue, and ABF1 is highlighted in red. c Bioluminescence analysis of pCCA1::LUC in the Col-0 and ABF1 overexpression (ABF1ox) backgrounds under driven (2 days, 16 h light:8 h darkness photoperiods) and free-running (5 days, constant light) conditions. White and gray bars represent day and night, respectively (16–64 h). From 64 h on, white and dotted bars represent subjective day and subjective night, respectively. Mean ± SEM, n = 12 and 9. d Period length estimations of pCCA1::LUC reporter expression in the experiment shown in (c), as analyzed by FFT-NLLS, two-sided Student’s t test, n = 12 samples (Col-0) and 9 samples (ABF1ox), *p-value is 0.0275. The box plots display the median (horizontal line), second to third quartiles (box), with the maxima and minima as whiskers.
Furthermore, we have conducted a comprehensive analysis of circadian genes specific to each cell cluster. In each dataset, we identified around 3000 genes that exhibit oscillations exclusively within individual cell clusters (Fig. 5a and Supplementary Fig. 23a) and performed Gene Ontology (GO) analysis on these genes (Supplementary Data 6). Due to the higher time point resolution of the 24 h dataset, we used this dataset for detailed analysis. For mesophyll cells (cluster S0), which are essential for energy conversion, we observed an enrichment in genes involved in the response to light stimulus, with peak oscillation phases around CT18 to CT0 (Supplementary Fig. 24). For example, Chloroplast ATP/ADP transporter 1 (NTT1), which encodes an ADP/ATP transporter responsible for the exchange of ADP and ATP within chloroplasts, peaked at CT0 (Supplementary Data 6), aligning with its role in nocturnal energy supply during the night50,51,52. In guard cells (cluster S7), which are crucial for stress response, we identified genes enriched in salicylic acid (SA) responses, peaking at CT16 (Supplementary Fig. 24). This peak phase aligns with the oscillation of SA, which peaks in the middle of the night53. Among these genes, AtMYBS1/AtMYBL, acting as a negative regulator of heat tolerance, highlights the functional relevance of these oscillations54. In addition, in phloem companion cells (cluster V2), responsible for the transport of organic compounds and crucial for responding to environmental stresses, we observed genes involved in cellular responses to organic substances, with peak oscillation at CT4 (Supplementary Fig. 24). For instance, leucine-rich repeat extensin 3 (LRX3) encodes a cell wall-localized protein essential for salt tolerance55. We also identified auxin-responsive genes that oscillate exclusively in the shoot epidermis (cluster S1), peaking at CT4 (Supplementary Fig. 24).
Further GO analysis of oscillating genes peaking at different phases within each cluster uncovered both common and distinct terms among different cell types (Supplementary Data 7). Specifically, genes involved in light response and photosynthesis were enriched among the shoot cell types (Supplementary Fig. 25). Moreover, we also observed a GO term related to auxin response was predominantly enriched in shoot epidermal cells around CT4 (Supplementary Fig. 26a), involving a set of genes associated with the auxin signaling pathway, including six Small Auxin-Upregulated RNA (SAUR) family genes (SAUR10, 15, 19, 21, 22, 25). These genes were found to oscillate only in shoot epidermal cells, peaking in the early morning hours (Supplementary Fig. 26b), and were specifically expressed in these cells (Supplementary Fig. 26c). Moreover, a recent study56 reported upregulation of these SAUR genes in shoot epidermal cells exposed to light after constant dark treatment, consistent with our findings that these genes are upregulated at subjective morning. Furthermore, among these SAUR genes, SAUR19 and SAUR24 are known to promote cell expansion mediated by auxin and play significant roles in plant growth57. While our GO analysis provides an overview of potential functional pathways, further functional validation of these cell-type-specific oscillating genes is necessary to elucidate their precise roles in circadian rhythm regulation.
A data-driven approach for identifying key circadian regulators
Our findings further indicate that core clock genes, including CCA1 (28), LNK2 (33), PRR7 (29), LHY (28), and RVE8 (25) are oscillating in most of the 40 clusters in the 48-hour dataset (Fig. 5a) and in most of the 27 clusters in the 24-hour dataset (CCA1 (20), LNK2 (19), PRR7 (18), LHY (16), and RVE8 (18) (Supplementary Fig. 23a). In both datasets, genes that oscillate in multiple clusters were predominantly circadian regulators (Fig. 5a and Supplementary Fig. 23a). In addition to these core clock genes, we also found several genes involved in regulating circadian rhythm that also oscillate in many clusters. For example, jumonji domain containing 5 (JMJD5)58, glycine-rich RNA-binding proteins 7 (GRP7)59, LNK360, LNK460, PHYTOCHROME INTERACTING FACTOR4 (PIF4)61, and PIF562 oscillated in more than 30 clusters in the 48 h dataset (Fig. 5a). Then, shifting focus to the 14 genes that oscillated in over 12 clusters in the 24 h dataset, we found that half of them is reported as circadian regulators, including 6 core clock genes, and other 3 genes including GRP759, RVE263, and HY5-HOMOLOG (HYH)64 (Supplementary Fig. 23b). Hence, genes found to oscillate in multiple cell types are indeed genes that play a critical role in the regulation of circadian rhythms. Given that the average number of detectable genes per cell is just over 847 in the 24-hour dataset, we sought to determine whether the genes exhibiting rhythmicity across all clusters were simply those with the highest expression levels. We analyzed the relationship between gene expression levels (measured as CPM) and the number of clusters in which genes exhibit rhythmicity in both datasets. Our analysis revealed a slight positive correlation, with a Kendall tau coefficient of 0.21 (48-hour dataset) and 0.25 (24 h dataset) (Supplementary Fig. 27a, b). However, genes displaying rhythms across all clusters are not exclusively the highest-expressed genes. For instance, LNK2 is the highest-expressed core clock gene and ranks third in the number of rhythmic clusters in the 24-hour dataset (Supplementary Fig. 23a). Yet, it ranks 80th in CPM among all genes (Supplementary Fig. 23a). When we applied a stringent filter by selecting genes with CPM greater than 100, the correlation coefficient dropped to 0.15 in the 24 h dataset and to − 0.03 in the 48 h dataset (Supplementary Fig. 27a, b). These findings indicate that while there is a modest association between gene expression levels and rhythmicity detection, our approach is not solely biased toward highly expressed genes. Since many core clock genes in Arabidopsis are transcription factors65, we ranked all transcription factors from PlantTFDB66,67,68,69 based on the number of clusters in which they oscillate (Supplementary Data 8). ABRE-binding factor 1 (ABF1), a transcription factor involved in the ABA signaling pathway70, ranks relatively high among transcription factors in both the 24 h and 48 h datasets, ranking 6th and 30th, respectively (Fig. 5b). Moreover, we also ranked transcription factors known to bind to the CCA1 promoter region71 in both datasets. The results revealed that ABF1 ranks exceptionally high, at 1st in the 24 h dataset and 7th in the 48 h dataset (Supplementary Fig. 23c, d). To investigate ABF1’s impact on the circadian clock, we overexpressed ABF1 in the pCCA1::LUC transgenic line, and we found that ABF1 overexpression shortens the circadian period under free-running conditions (Fig. 5d). In addition, a recent study by Liang et al.72 observed that overexpression of ABF3, a member of the same gene family, also shortens the circadian period. In summary, our data provide a comprehensive atlas for studying cell-type-specific circadian rhythms in Arabidopsis and introduce novel circadian regulators. This demonstrates that single-nucleus sequencing technology can reveal insights into circadian clock mechanisms.
Considering that clock components exhibit significant evolutionary conservation across angiosperms73,74, our snRNA-seq data on circadian expression can be leveraged beyond Arabidopsis research and serve as a robust tool to pinpoint circadian regulators in a diverse range of plant species. Moreover, for the many plant species in which the circadian components remain partially unclear75,76,77, our time-series snRNA-seq approach could be applied to quickly generate a potential gene list of key circadian regulators ranked by the number of cell types that they are oscillating in.
Augmenting the utility of our study, we have curated an online website that provides easy access to gene expression data from our research (Fig. 6). The portal facilitates searches using gene IDs, with immediate visualization and access to the expression profile of each gene, including UMAP plots, bar plots, and line plots detailing expression patterns across circadian time points at both the whole plant and cell-type levels in the 24-hour and the 48-hour datasets (Fig. 6d–g), thus providing a valuable resource for the community and fostering a collaborative environment for furthering research on plant chronobiology. In addition, our website (https://zhailab.bio.sustech.edu.cn/sc_circadian) allows users to download the CPM values of each gene of interest in each cluster.
a UMAP plot representing a visualization of the identified cell types. b The interface provides a user-input field for querying specific gene IDs of interest. c Options for selecting snRNA-seq datasets: “24 h data with 2 h intervals” and “48 h data with 4 h intervals,” allowing users to choose the datasets they wish to explore. d Users can select the preferred gene expression unit for visualization through the ‘Layer’ option, which includes Counts Per Million (CPM) or logarithm base 2 of CPM (log2CPM). In addition, the ‘Share y Limit’ feature allows for consistent y-axis scaling across multiple plots, enhancing comparability, as exemplified in the bar plots of Fig. 6h–j. Genes detected in the snRNA-seq data are shown below. e A download function allows users to download the CPM values of each gene of interest in each cell, enhancing the usability and accessibility of our data. f UMAP plot representing the gene expression profiles across cell clusters. g UMAP plots representing the expression profiles across different samples. h Bar plots representing gene expression profiles within each cluster. i Line plot showing the overall expression profiles at the whole seedling level. j Line plots showing expression profiles across the clusters. Source data are provided as a Source Data file.
In conclusion, our time-series high-throughput snRNA-seq data provide a perspective on circadian gene expression profiles within individual cell types of plants. This method represents a significant step forward in understanding the plant circadian clock at the cellular level, offering insights into the complexities of the clock network. The potential future application of this technique to other plant species could contribute to a broader understanding of circadian rhythms in plant biology and potentially inform agricultural practices.
Methods
Plant materials
Plant growth and material harvesting were performed as previously described with minor modifications78. The Arabidopsis thaliana used in this study was the Columbia-0 ecotype. Seeds were sown on MS agar plates containing 3% sucrose and stratified at 4 °C for 2 days, then placed in growth chambers at 22 °C. Plants were entrained under a 12 h light and 12 h dark (12 L/12D, 100 μmol m−2 s−1) photoperiod for 7 days, and then released into constant light (100 μmol m−2 s−1) conditions. They were harvested every 2 h over a 24 h period, and every 4 h over a 48 h period starting from subjective dawn on day 9.
Tissue-specific LUC-GUS transgenic plant construction, GUS staining, monitoring circadian rhythms
The promoter regions of RBCS2B (AT5G38420), AT1G66100, CAB2 (AT1G29920), CEPD1 (AT1G06830), and KCS6 (AT1G68530) which encode a PR (pathogenesis-related) protein were amplified using the Phusion High-Fidelity DNA Polymerase (New England Biolabs, M0530L) from Col-0 genomic DNA with two primer pairs RBCS2B-BglII-PF (5’-CCAGATCTGGAATATTCAATGTTGACTATC-3’) and RBCS2B-Eco91I-PR (5’-CGTCGGTGACCATGGCTTCTTCTTGTTGTTTCTCTTC-3’), AT1G66100-BamHI-PF (5′-GTGGATCCAGTCCATTAATGTCATAAATCTG-3’) and AT1G66100-Eco91I-PR (5′-CGTCGGTGACCATCTTTTGATTGATTAGTTG-3’), CEPD1-SalI-F1 (5’-CTCAATGGACGAAGTTGTAAATATGTG-3’) and CEPD1-KpnI-R1 (5’- GGGGTACCTATCTTGCATAAGGTGAAGAAG-3’), KCS6-BamHI-F1 (5’- GTGGATCCAAAAGCAGAGTGGGTGATTC-3’) and KCS6-NcoI/BstEII-R1 (5’-CGTCGGTGACCATGGTCGGAGAGTTTTAATG-3’), CAB2-SalI-F1 (5’-CGTATACGAATAGATCTCTCTATCCTTATG-3’) and CAB2-Eco91I-R1 (5’-CGTCGGTGACCATGGTTGAGGCGGCCATTGAAACTTTTTTGTG-3’). PCR products of RBCS2B promoter region (2,364 bps) and PCR products of AT1G66100 promoter region (2600 bps) were double digested with BamHI (Thermo Fisher, FD0054) and Eco91I (Thermo Fisher, FD0394) or BglII (Thermo Fisher, FD0084) and Eco91I (Thermo Fisher, FD0394), PCR products of CEPD1 promoter region (3128 bps) was digested with SalI (Thermo Fisher, FD0644) and KpnI (Thermo Fisher, FD0524), and PCR products of KCS6 promoter region (5263 bps) was digested with BamHI (Thermo Fisher, FD0054) and BstEII (Thermo Fisher, FD0394), PCR products of CAB2 promoter region (1904 bps) was digested with SalI (Thermo Fisher, FD0644) and Eco91I (Thermo Fisher, FD0394), then the promoter-driven LUC-GusP fusion reporter plasmids were made through insertion of the digested PCR-amplified promoter regions into a modified pENTR 2B vector with LUC-GusP gene at the multiple cloning site (MCS). Finally, The promoter:LUC-GusP constructs were recombined into a modified pH2GW7∆79 from which the 35S promoter had been deleted via Gateway cloning technology. RBCS2B:LUC-GusP, AT1G66100:LUC-GusP, CEPD:LUC-GusP and KCS6:LUC-GusP transgenic (T3) lines were used to determine the tissue-specific expression pattern via histochemical GUS reporter activity80. For firefly LUC measurement, two reporter transgenic lines were entrained for 10 days in LD (12 h light/12 h dark, 22 °C) before release to the free-running conditions for luciferase activity analysis with an EM-CCD cooled camera system (Andor Technology). The raw data are shown in the Source Data file.
Nuclei isolation and 10X single-nucleus RNA-seq library preparation
Frozen samples from 9-day-old Arabidopsis seedlings were used for single-nucleus preparation. The nuclei were isolated as following these steps. Firstly, the seedlings were chopped in ice-cold 1 x Nuclei Isolation Buffer (NIB, MilliporeSigma, CELLYTPN1) supplemented with 1 mM dithiothreitol (DTT, Thermo Fisher, R0861), 1 × protease inhibitor (Sigma, 4693132001), and 0.4 μ/μl Murine RNase inhibitor (Vazyme, R301-03). Thereafter, the lysate was filtered through a 40 μm strainer and centrifuged at 4 °C for 5 minutes at 1000 × g. The supernatant was then removed, and the nuclei pellet was resuspended with 500 μl NIB. To further remove the debris, the nuclei suspension was filtered with a 20 μm strainer and stained with 4,6-Diamidino-2-phenylindole (DAPI, 1 mg/ml) before fluorescent activated cell sorting (FACS). The isolated nuclei were run on an MA900 flow cytometer (SONY) with a 100 μm nozzle. Sorting was conducted into a tube containing 1 ml of collection buffer (1 × PBS [Corning, 21-040-CVC] supplemented with 1% BSA [VWR, VWRV0332] and 0.4 μ/μl Murine RNase inhibitor). The entire nucleus isolation and FACS sorting process took less than 2 h. At least 100,000 nuclei were sorted per sample based on the DAPI signal intensity and the nuclear size. The single-nucleus suspensions were pelleted at 4 °C for 5 min at 1000 × g and then resuspended in 45 μl collection buffer. The quality of the nuclei was checked under the microscope. Approximately 20,000 intact nuclei were loaded onto the Chromium Next GEM Chip G, followed by the library construction for Illumina sequencing according to the manufacturer’s protocol for the 10X ChromiumTM Single Cell 3’ Library Kit.
ABF1 effect on clock activity
To analyze the effect ABF1 overexpression on clock activity, an Arabidopsis thaliana line carrying the pCCA1::LUC reporter was transformed with a construct harboring the ABF1 coding sequence under control of the 35S promoter. To this end, the MultiSite Gateway Three-Fragment Vector Technology (Invitrogen, 12537-023) was used. A gBlockTM gene fragment consisting of the full-length coding sequence of ABF1 (without the stop codon) flanked by the attB1 and attB2 sites was ordered from IDT and subsequently cloned into the pDONR207 vector by Gateway BP recombination reaction (Invitrogen). Next, a MultiSite Gateway reaction (Invitrogen) was set with this construct, a pDONR P4-P1R vector containing the 35S promoter sequence, a pDONR P2R-P3 vector containing a 6xHA tag, and the pK7m34GW binary destination vector (https://gateway.psb.ugent.be) to generate the final 35S::ABF1-6xHA construct, which was used for floral dip transformation of Arabidopsis. For circadian rhythm measurements, control (pCCA1::LUC) and T1 ABF1 overexpressing (pCCA1::LUC;ABF1ox) seedlings were grown on 0.5x MS plates, which in the case of the ABF1 overexpression lines contained kanamycin (50 µg/ml) for transformant selection. Seedlings were grown for 7 days under long day conditions (LD: 16 h light, 8 h dark), after which control and positive T1 seedlings were transferred 96-well plates containing 0.5x MS and acclimated for 2 days. After the growth and entrainment period, 500 µM luciferin (GoldBio, LUCK-100) was added to the wells, and the plates were transferred to a luminometer (Tecan Infinite 200 PRO plate reader). Bioluminescence was recorded every 1 h for 2 days under LD and 5 days under constant light (80 mmol m-2 s-1) conditions, and data were analyzed by fast Fourier transformed non-linear least squares (FFT-NLLS) using BioDare281,82 (the whole 5 days in LL were used for the analysis (0–120 h). The experiment was repeated three times with similar results. The raw data are shown in the Source Data file.
Single-nucleus RNA-seq data analysis
We first utilized Cell Ranger for preprocessing the raw Fastq files. All parameters are set to their default values, except for the “--include-introns” flag to accommodate the high intron retention ratio feature of the nucleus transcriptome. The gene annotation file employed in our study is based on the Araport11 version, which itself relies on the TAIR10 version of the genome sequence. Next, we use scdblfinder83 to eliminate potential doublets and SoupX84 to remove ambient RNA contamination, thereby obtaining an initially filtered matrix. The obtained matrix was then subjected to additional filtering using the median absolute deviation (MAD) method recommended in the Single-Cell Best Practices85. Furthermore, cells with an organelle transcript proportion exceeding 10% are further removed from the matrix.
After completing quality control steps for each library, we merge the filtered results to obtain the final gene expression matrix. Subsequently, we tested multiple integration strategies to identify a suitable method for our data: (1) Performing PCA directly on the concatenated dataset, either with or without the removal of known circadian genes. This is followed by the construction of a nearest neighbor graph and clustering. (2) Applying Harmony86 to the concatenated dataset to alleviate batch effects. This is followed by the construction of a nearest neighbor graph, either with or without the removal of known clock-regulated genes. (3) Performing SCTransform separately on each sample to remove the baseline difference, followed by the Harmony method.
Among all the methods, the number of highly variable genes was set to 3000. PCA and the construction of a nearest neighbor graph were performed using the functions provided in Scanpy87 with default parameters. In addition, we also performed clustering on each sample separately. Apart from setting the number of highly variable genes to 3000, all other parameters were consistent with the tutorial of Scanpy. The clustering results will be evaluated using the silhouette coefficient, implemented with the functions wrapped in sklearn88, on the cell distance matrix obtained from the aforementioned methods. We also evaluated the effectiveness of batch effect removal using the kBET38 method provided in scIB89.
After the clustering method is determined, we employ the UMAP method90 (scanpy.tl.umap, min_dist = 0.3) to obtain a two-dimensional embedding of the cells, and the Leiden method91 (scanpy.tl.leiden, resolution = 0.4) to obtain clustering results. Upon careful examination of the expression patterns of known cell type marker genes and Jaccard index results (Supplementary Figs. 7, 11), we further subdivided clusters that may contain multiple cell types, resulting in subclusters V1/S3/R4, V3/S6/S8, and S9/S11, each originating from a single initial cluster.
The gene expression matrix of the 48-hour dataset was conducted like that of the 24 h dataset. During quality control, we retained only nuclei with more than 700 detected genes, over 1000 UMI counts, and less than 10% of transcripts derived from organelles. We then applied the optimal integration strategy 3 (Supplementary Fig. 4) from the 24 h dataset to integrate gene expression matrices and used the same approach to generate UMAP embeddings and clustering results. In addition, we subdivided clusters that may contain multiple cell types according to the marker genes’ expression and Jaccard index results, identifying subclusters R3/R4/R5/R6, R8/R9/R10, V0/V1/V2, and V3/S10, with each subcluster derived from a single initial cluster (Supplementary Figs. 7, 11).
We utilized the cellex method to identify cluster-specific genes92. This method integrates various indicators such as gene expression ratios, T-test results, and gene enrichment scores to calculate a specificity score for each gene in different clusters, ranging from 0 to 1, and we use a threshold of 0.9 as the final cutoff. In order to annotate the identity of each cluster, we combined the Root data from Shahan et al. with the Shoot data from Zhang et al.39,40. Subsequently, we also utilized the cellex method to identify cluster-specific genes in this pseudo-seedling dataset. To assess the similarity between our clusters and the clusters in the pseudo-seedling dataset, we employed the Jaccard Index as a measure.
To determine which genes driven by the circadian clock and exhibit oscillation in our dataset, we collected clock-regulated genes identified in three different sources of transcriptomic data. We then followed the approach proposed by Wen et al. to identify the subset of genes oscillating in our data. Initially, we merged all cells within each cluster to obtain pseudo-bulk data. Next, after performing CPM normalization and log2-transformation, we used the JTKcycle method wrapped in the metacycle package to calculate the corresponding p-values47. In this step, the circadian periodicity was set to 24 h. Genes with adjusted p-values below 0.05 were considered as oscillating. We then employed a nonlinear regression to fit the cosine curve and determine the phase, amplitude, and corresponding standard deviation errors of these oscillating genes. We also utilized the z-score to compute the statistical significance of phase and amplitude as described by Wen et al.30. In general, the z-score is obtained using the formula: \(z=\frac{|{\varphi }_{1}-{\varphi }_{2}|}{\sqrt{{S}_{1}^{2}+{S}_{2}^{2}}}\), where φ represents phase or amplitude, and S represents standard deviation. Subsequently, the two-tailed p-value is calculated using the scipy.stats.norm.cdf function.
Statistics and reproducibility
No statistical method was used to predetermine the sample size. Cells that did not meet quality criteria were removed during the quality control process (see above for details); no additional data were excluded from the analyses. The experiments were not randomized, and the investigators were not blinded to the allocation during the experiments and outcome assessment.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequencing data generated in this study have been deposited in the China National Center for Bioinformation database under accession code PRJCA021408. The source data for Figs. 1–6 are provided as a Source Data file. Source data are provided in this paper.
Code availability
The source code to reproduce this project can be accessed at https://github.com/ZhaiLab-SUSTech/circadian_notebooks_updated93.
References
Young, M. W. & Kay, S. A. Time zones: a comparative genetics of circadian clocks. Nat. Rev. Genet. 2, 702–715 (2001).
Harmer, S. L. The circadian system in higher plants. Annu. Rev. Plant Biol. 60, 357–377 (2009).
Xu, X., Yuan, L. & Xie, Q. The circadian clock ticks in plant stress responses. Stress Biol. 2, 15 (2022).
Panda, S. Circadian physiology of metabolism. Science 354, 1008–1015 (2016).
Nohales, M. A. & Kay, S. A. Molecular mechanisms at the core of the plant circadian oscillator. Nat. Struct. Mol. Biol. 23, 1061–1069 (2016).
Patke, A., Young, M. W. & Axelrod, S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell Biol. 21, 67–84 (2020).
Xu, X. et al. Circadian clock in plants: Linking timing to fitness. J. Inter. Plant Biol. 64, 792–811 (2022).
Endo, M. Tissue-specific circadian clocks in plants. Curr. Opin. Plant Biol. 29, 44–49 (2016).
Inoue, K., Araki, T. & Endo, M. Oscillator networks with tissue-specific circadian clocks in plants. Semin. Cell Dev. Biol. 83, 78–85 (2018).
Sorkin, M. L. & Nusinow, D. A. Time will tell: Intercellular communication in the plant clock. Trends Plant Sci. 26, 706–719 (2021).
Swift, J., Greenham, K., Ecker, J. R., Coruzzi, G. M. & Robertson McClung, C. The biology of time: dynamic responses of cell types to developmental, circadian and environmental cues. Plant J. 109, 764–778 (2022).
Perez-Garcia, P., Serrano-Ron, L. & Moreno-Risueno, M. A. The nature of the root clock at single cell resolution: Principles of communication and similarities with plant and animal pulsatile and circadian mechanisms. Curr. Opin. Cell Biol. 77, 102102 (2022).
Davis, W., Endo, M. & Locke, J. C. W. Spatially specific mechanisms and functions of the plant circadian clock. Plant Physiol. 190, 938–951 (2022).
Shimizu, H., Araki, T. & Endo, M. Photoperiod sensitivity of the Arabidopsis circadian clock is tissue-specific. Plant Signal. Behav. 10, e1010933 (2015).
Shimizu, H. et al. Decentralized circadian clocks process thermal and photoperiodic cues in specific tissues. Nat. Plants 1, 15163 (2015).
James, A. B. et al. The circadian clock in Arabidopsis roots is a simplified slave version of the clock in shoots. Science 322, 1832–1835 (2008).
Lee, H. G. & Seo, P. J. Dependence and independence of the root clock on the shoot clock in Arabidopsis. Genes Genom. 40, 1063–1068 (2018).
Takahashi, N., Hirata, Y., Aihara, K. & Mas, P. A hierarchical multi-oscillator network orchestrates the arabidopsis circadian system. Cell 163, 148–159 (2015).
Endo, M., Shimizu, H., Nohales, M. A., Araki, T. & Kay, S. A. Tissue-specific clocks in Arabidopsis show asymmetric coupling. Nature 515, 419–422 (2014).
Yakir, E. et al. Cell autonomous and cell-type specific circadian rhythms in Arabidopsis: Circadian rhythms in individual plant cells. Plant J. 68, 520–531 (2011).
Gould, P. D. et al. Coordination of robust single cell rhythms in the Arabidopsis circadian clock via spatial waves of gene expression. Elife 7, e31700 (2018).
Nakamura, S. & Oyama, T. Adaptive diversification in the cellular circadian behavior of arabidopsis leaf- and root-derived cells. Plant Cell Physiol. 63, 421–432 (2022).
Nakamura, S. & Oyama, T. Long-term monitoring of bioluminescence circadian rhythms of cells in a transgenic Arabidopsis mesophyll protoplast culture. Plant Biotechnol. 35, 291–295 (2018).
Muranaka, T. & Oyama, T. Heterogeneity of cellular circadian clocks in intact plants and its correction under light-dark cycles. Sci. Adv. 2, e1600500 (2016).
Muranaka, T. & Oyama, T. in Bioluminescent Imaging. (Springer US, New York, NY, 2020).
Isoda, M. & Oyama, T. Use of a duckweed species, Wolffiella hyalina, for whole-plant observation of physiological behavior at the single-cell level. Plant Biotechnol. 35, 387–391 (2018).
Saliba, A.-E., Westermann, A. J., Gorski, S. A. & Vogel, J. Single-cell RNA-seq: advances and future challenges. Nucleic Acids Res. 42, 8845–8860 (2014).
Stark, R., Grzelak, M. & Hadfield, J. RNA sequencing: the teenage years. Nat. Rev. Genet. 20, 631–656 (2019).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
Wen, S. et al. Spatiotemporal single-cell analysis of gene expression in the mouse suprachiasmatic nucleus. Nat. Neurosci. 23, 456–467 (2020).
Ma, D. et al. A transcriptomic taxonomy of Drosophila circadian neurons around the clock. ELife 10, e63056 (2021).
Morris, E. L. et al. Single‐cell transcriptomics of suprachiasmatic nuclei reveal a Prokineticin‐driven circadian network. EMBO J. 40, e108614 (2021).
Xiao, T. et al. T regs in visceral adipose tissue up-regulate circadian-clock expression to promote fitness and enforce a diurnal rhythm of lipolysis. Sci. Immunol. 7, eabl7641 (2022).
Apelt, F. et al. Shoot and root single cell sequencing reveals tissue- and daytime-specific transcriptome profiles. Plant Physiol. 188, 861–878 (2022).
Torii, K. et al. A guiding role of the Arabidopsis circadian clock in cell differentiation revealed by time-series single-cell RNA sequencing. Cell Rep. 40, 111059 (2022).
Choudhary, S. & Satija, R. Comparison and evaluation of statistical error models for scRNA-seq. Genome Biol. 23, 27 (2022).
Luecken, M. D. & Theis, F. J. Current best practices in single‐cell RNA‐seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Büttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Zhang, T.-Q., Chen, Y. & Wang, J.-W. A single-cell analysis of the Arabidopsis vegetative shoot apex. Dev. Cell 56, 1056–1074 (2021).
Shahan, R. et al. A single-cell Arabidopsis root atlas reveals developmental trajectories in wild-type and cell identity mutants. Dev. Cell 57, 543–560 (2022).
Xia, K. et al. The single-cell stereo-seq reveals region-specific cell subtypes and transcriptome profiling in Arabidopsis leaves. Dev. Cell 57, 1299–1310 (2022).
Chen, L.-Q. et al. Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335, 207–211 (2012).
Sawa, S. et al. FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Genes Dev. 13, 1079–1088 (1999).
Covington, M. F., Maloof, J. N., Straume, M., Kay, S. A. & Harmer, S. L. Global transcriptome analysis reveals circadian regulation of key pathways in plant growth and development. Genome Biol. 9, R130 (2008).
Hsu, P. Y. & Harmer, S. L. Circadian phase has profound effects on differential expression analysis. PLoS ONE 7, e49853 (2012).
Romanowski, A., Schlaen, R. G., Perez‐Santangelo, S., Mancini, E. & Yanovsky, M. J. Global transcriptome analysis reveals circadian control of splicing events in Arabidopsis thaliana. Plant J. 103, 889–902 (2020).
Hughes, M. E., Hogenesch, J. B. & Kornacker, K. JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. Rhythms 25, 372–380 (2010).
Lu, P., Porat, R., Nadeau, J. A. & O’Neill, S. D. Identification of a meristem L1 layer-specific gene in Arabidopsis that is expressed during embryonic pattern formation and defines a new class of homeobox genes. Plant Cell 8, 2155–2168 (1996).
Zhang, Y. et al. Large-scale comparative epigenomics reveals hierarchical regulation of non-CG methylation in Arabidopsis. Proc. Natl. Acad. Sci. USA. 115, https://doi.org/10.1073/pnas.1716300115 (2018).
Linka, N. & Weber, A. P. M. Intracellular metabolite transporters in plants. Mol. Plant 3, 21–53 (2010).
Trentmann, O., Jung, B., Neuhaus, H. E. & Haferkamp, I. Nonmitochondrial ATP/ADP transporters accept phosphate as third substrate. J. Biol. Chem. 283, 36486–36493 (2008).
Neuhaus, H. E., Thom, E., Möhlmann, T., Steup, M. & Kampfenkel, K. Characterization of a novel eukaryotic ATP/ADP translocator located in the plastid envelope of Arabidopsis thaliana L. Plant J. 11, 73–82 (1997).
Zheng, X. et al. Spatial and temporal regulation of biosynthesis of the plant immune signal salicylic acid. Proc. Natl Acad. Sci. USA 112, 9166–9173 (2015).
Li, X. et al. AtMYBS1 negatively regulates heat tolerance by directly repressing the expression of MAX1 required for strigolactone biosynthesis in Arabidopsis. Plant Commun. 4, 100675 (2023).
Zhao, C. et al. Leucine-rich repeat extensin proteins regulate plant salt tolerance in Arabidopsis. Proc. Natl. Acad. Sci. USA 115, 13123–13128 (2018).
Han, X. et al. Time series single-cell transcriptional atlases reveal cell fate differentiation driven by light in Arabidopsis seedlings. Nat. Plants 9, 2095–2109 (2023).
Spartz, A. K. et al. The SAUR19 subfamily of SMALL AUXIN UP RNA genes promote cell expansion. Plant J. 70, 978–990 (2012).
Jones, M. A. et al. Jumonji domain protein JMJD5 functions in both the plant and human circadian systems. Proc. Natl. Acad. Sci. USA 107, 21623–21628 (2010).
Heintzen, C., Nater, M., Apel, K. & Staiger, D. At GRP7, a nuclear RNA-binding protein as a component of a circadian-regulated negative feedback loop in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 94, 8515–8520 (1997).
de Leone, M. J. et al. The LNK gene family: At the crossroad between light signaling and the circadian clock. Genes 10, 2 (2018).
Dornbusch, T., Michaud, O., Xenarios, I. & Fankhauser, C. Differentially phased leaf growth and movements in Arabidopsis depend on coordinated circadian and light regulation. Plant Cell 26, 3911–3921 (2014).
Shor, E., Paik, I., Kangisser, S., Green, R. & Huq, E. PHYTOCHROME INTERACTING FACTORS mediate metabolic control of the circadian system in Arabidopsis. N. Phytol. 215, 217–228 (2017).
Zhang, X. et al. Constitutive expression of CIR1 (RVE2) affects several circadian‐regulated processes and seed germination in Arabidopsis. Plant J. (2007).
Hajdu, A. et al. ELONGATED HYPOCOTYL 5 mediates blue light signalling to the Arabidopsis circadian clock. Plant J. 96, 1242–1254 (2018).
McClung, C. The plant circadian oscillator. Biology 8, 14 (2019).
Jin, J., Zhang, H., Kong, L., Gao, G. & Luo, J. PlantTFDB 3.0: a portal for the functional and evolutionary study of plant transcription factors. Nucl. Acids Res. 42, D1182–D1187 (2014).
Jin, J. et al. PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 45, D1040–D1045 (2017).
Tian, F., Yang, D.-C., Meng, Y.-Q., Jin, J. & Gao, G. PlantRegMap: charting functional regulatory maps in plants. Nucleic Acids Res. 48, D1104–D1113 (2019).
Jin, J. et al. An Arabidopsis transcriptional regulatory map reveals distinct functional and evolutionary features of novel transcription factors. Mol. Biol. Evol. 32, 1767–1773 (2015).
Choi, H., Hong, J., Ha, J., Kang, J. & Kim, S. Y. ABFs, a Family of ABA-responsive element binding factors. J. Biol. Chem. 275, 1723–1730 (2000).
Pruneda-Paz, J. L. et al. A genome-scale resource for the functional characterization of arabidopsis transcription factors. Cell Rep. 8, 622–632 (2014).
Liang, T., Yu, S., Pan, Y., Wang, J. & Kay, S. A. The interplay between the circadian clock and abiotic stress responses mediated by ABF3 and CCA1/LHY. Proc. Natl. Acad. Sci. USA 121, e2316825121 (2024).
Song, Y. H., Ito, S. & Imaizumi, T. Similarities in the circadian clock and photoperiodism in plants. Curr. Opin. Plant Biol. 13, 594–603 (2010).
McClung, C. R. Beyond Arabidopsis: The circadian clock in non-model plant species. Semin. Cell Dev. Biol. 24, 430–436 (2013).
Laosuntisuk, K., Elorriaga, E. & Doherty, C. J. The game of timing: Circadian rhythms intersect with changing environments. Annu. Rev. Plant Biol. 74, 511–538 (2023).
Francisco, M. & Rodríguez, V. M. Importance of daily rhythms on brassicaceae phytochemicals. Agronomy 11, 639 (2021).
Rees, H. et al. Circadian regulation of the transcriptome in a complex polyploid crop. PLoS Biol. 20, e3001802 (2022).
Harmer, S. L. et al. Orchestrated transcription of key pathways in Arabidopsis by the circadian clock. Science 290, 2110–2113 (2000).
Karimi, M., Inzé, D. & Depicker, A. GATEWAYTM vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 7, 193–195 (2002).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Zieliński, T., Hay, J. & Millar, A. J. in Plant Circadian Networks. (Springer US, New York, NY, 2022).
Zielinski, T., Moore, A. M., Troup, E., Halliday, K. J. & Millar, A. J. Strengths and limitations of period estimation methods for circadian data. PLoS ONE 9, e96462 (2014).
Germain, P.-L., Lun, A., Garcia Meixide, C., Macnair, W. & Robinson, M. D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 10, 979 (2022).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. GigaScience 9, giaa151 (2020).
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat. Rev. Genet. 24, 550–572 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform Manifold Approximation and Projection. JOSS 3, 861 (2018).
Traag, V. A., Waltman, L. & Van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Timshel, P. N., Thompson, J. J. & Pers, T. H. Genetic mapping of etiologic brain cell types for obesity. ELife 9, e55851 (2020).
Liu, Z. & Qin, Y. 48-Hour and 24-Hour Time-lapse Single-nucleus Transcriptomics Reveal Cell-type specific Circadian Rhythms in Arabidopsis. Zenodo https://doi.org/10.5281/ZENODO.15045784 (2025).
Acknowledgements
The group of J.Z. is supported by the Biological Breeding-National Science and Technology Major Project (2023ZD04073), National Natural Science Foundation of China (32325031, 32470601), the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (2016ZT06S172); Y.L. is supported by the National Natural Science Foundation of China (32300479), Natural Science Foundation of the Higher Education Institutions of Guangdong Province, China (2022KTSCX115) and Natural Science Foundation of Guangdong Province of China (2023A1515011997); and the National Natural Science Foundation of China to Q.X. (32170259, 31670285) and X.X. (U1904202, 32170275); Central Plains Science and Technology Innovation Leadership Talent Program to X.X. (234200510023); Z.L. is supported by the Fundamental Research Fund for Central Universities (2412023YQ005); M.A.N. is supported by MCIN/AEI/10.13039/501100011033 PID2020-119491GA-I00 (grant PID2020-119491GA-I00) and Generalitat Valenciana (grant CIDEGENT/2020/037). C.M.-V. is the recipient of a predoctoral fellowship from Generalitat Valenciana (CIACIF/2021/329). This work was supported by the Shenzhen Science and Technology Program (Grant No. ZDSYS20230626091659010).
Author information
Authors and Affiliations
Contributions
Y.Q., J.Z., Q.X., X.X., Y.L., and M.A.N. designed the experiments. Y.Q., Y.L., X.Z., S.G., and C.M-V. performed the experiments. Y.Q., Z.L., X.Z., Y.L., and M.A.N. analyzed the data. Q.X., Y.G., and C.M.-V. Generated the constructs. Y.Q., Z.L., J.Z., M.A.N., and X.X. wrote the manuscript. J.Z., M.A.N., Q.X., and X.X. oversaw the study. B.L. and Q.X. provided conceptual insight.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Motomu Endo, Dmitri Nusinow, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Qin, Y., Liu, Z., Gao, S. et al. 48-Hour and 24-Hour Time-lapse Single-nucleus Transcriptomics Reveal Cell-type specific Circadian Rhythms in Arabidopsis. Nat Commun 16, 4171 (2025). https://doi.org/10.1038/s41467-025-59424-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59424-8
This article is cited by
-
Unveiling the early defense response dynamics in grapevines against Plasmopara viticola by single-cell transcriptomics
Genome Biology (2026)
-
High resolution diel transcriptomes of autotetraploid potato reveal expression and sequence conservation among rhythmic genes
BMC Genomics (2025)
-
FX-Cell: a method for single-cell RNA sequencing on difficult-to-digest and cryopreserved plant samples
Nature Methods (2025)