Abstract
Chimeric Antigen Receptor (CAR) T cell therapy is a pivotal treatment for hematological malignancies. However, CAR T cell products exhibit batch-to-batch variability in cell number, quality, and in vivo efficacy due to donor-to-donor heterogeneity, and pre/post-manufacturing processes, and the manufacturing of such products necessitates careful testing, both post-manufacturing and pre-infusion. Here, we introduce the Cell Trajectory Modulation (CTM) assay, a microfluidic, label-free approach for the rapid evaluation of the functional attributes of CAR T cells based on biophysical features (i.e., size, deformability). CTM assay correlates with phenotypic metrics, including CD4:CD8 ratio, memory subtypes, and cytotoxic activity. Validated across multiple donors and culture platforms, the CTM assay requires fewer than 10,000 cells and delivers results within 10 minutes. Compared to labeled flow cytometry processing, the CTM assay offers real-time data to guide adaptive manufacturing workflows. Thus, the CTM assay offers an improvement over existing phenotypic assessments, marking a step forward in advancing CAR T cell therapy manufacturing.
Similar content being viewed by others
Introduction
Chimeric antigen receptor (CAR) T cell therapy has emerged as a breakthrough medicine for treating various hematological malignancies. It shows unprecedented success in clinical trials, leading to its rapid adoption and approval for clinical use. However, as CAR T cell therapy gains broader acceptance, challenges and gaps associated with consistent manufacturing have become apparent. Current autologous CAR T cells are extremely complex to produce, with each personalized batch of Food and Drug Administration (FDA) approved Axicaptagene Ciloleucel and Tisagenlecleucel products costing around $500,0001. Unfortunately, 2–9% of these products are rejected due to manufacturing issues such as insufficient CAR T cells, low effector cell numbers, or low cell viability2,3,4. In some cases, a new batch of CAR T product is manufactured, resulting in an additional 3–4 weeks' delay of the critical therapy.
The personalized nature of autologous CAR T cell manufacturing presents significant hurdles in managing scalability, cost-effectiveness, treatment efficacy, and toxicity5,6,7. Current CAR T manufacturing methods in bioreactors are akin to a black box, where product functional phenotypes are not monitored until final product release tests. These single-end-point measurements of cell quality attributes do not correlate well with clinical response8,9. This inevitably results in CAR T cells that are manufactured with large variations in cell number, phenotypes, in vivo efficacy, and potential side effects8,10,11.
Critically, manufactured CAR T potency and functional tests are only performed before the addition of cryopreservation media, and are not performed during the post-thaw, pre-product infusion4,12. Thus, the CAR T cell attributes measured at the release test can be significantly altered at pre-infusion, potentially impacting in vivo efficacy. Both Tisagenlecleucel and Axicaptagene Ciloleucel products utilize IFN-γ secretion as an indirect assay as part of potency assessment for product release4,12. However, Novartis reported that IFN-γ secretion varied greatly from batch to batch, suggesting that there may be significant uncontrolled variations in the safety, potency, and clinical effect of these products4. These variations in measured quality are further confounded by downstream cellular changes during DMSO cryopreservation13,14, shipping, and thawing15, which potentially affect clinical efficacy. Coupled with inherent patient-to-patient heterogeneity16, there is a strong need for new technologies with more specific and rapid cell functional phenotype measurements during, at the end of the manufacturing, and arguably at pre-infusion of CAR T cells. Insights into the T cell properties during production, which could enable adaptive manufacturing processes for early detection of adverse production events17,18, could significantly improve the consistency and efficacy of cell therapy products.
Here, we developed a microfluidic T cell biophysical size and deformability profiling assay, called the cell trajectory modulation (CTM) assay, to assess the quality of manufactured T cells, which shows strong correlations with immune phenotypes, activation, and CAR T potency. The CMT assay adopts specific micropillar structures engineered for precision cell interaction and measurement. The readout is based on the output of T cell trajectories corresponding to unique biophysical signatures. The CTM assay handles 20 µL of unprocessed and unlabeled culture samples, using <10,000 cells, and completes the profiling within <10 min, making it suitable for integration into the current CAR T manufacturing workflow, and enabling rapid label-free measurements of the CAR T cells.
When benchmarked against flow cytometry cell profiling, the CTM assay reveals bioreactor-specific, donor-specific, and cell processing (cryopreservation) variations of the CAR T product. The nature of CAR T cell manufacturing remains dynamic and heterogeneous, and insights into the temporal trends during manufacturing are not currently adopted as critical quality attributes (CQA). The CTM assay offers a streamlined, label‐free method for monitoring cell quality during CAR T cell production. By integrating this label-free assay into existing manufacturing workflows, the CTM assay provides timely insights into cell characteristics that may inform future process refinement and analytics. These findings highlight the potential of the CTM assay to deepen our understanding of product variability and support efforts toward achieving greater consistency in cell therapy manufacturing.
Results
CTM assay profiles T cell biophysical features
The CTM assay evolved from observations in cell sorting by deterministic lateral displacement (DLD) techniques, where, for a given microfluidic pillar arrangement, sorting trajectories of biological particles can be resolved at 20 nm resolution19,20. Researchers noticed that cell sorting was significantly impacted at varying flow rates and with different DLD pillar structures21. While these observations of variations are often deemed undesirable for sorting applications, this assay adapts these unique cell trajectories in differing conditions to characterize the T cell’s inherent biophysical properties, as shown in Fig. 1a. These CTM conditions result in unique trajectories of cells flowing within the device, and the interacting T cells would finally flow into the designated location bins within the microfluidic chip (Fig. 1a). The cumulative frequency histogram plots showing the modulated biophysical signatures result in the different histogram maps with H1 and H2 linked to size profiling at a slow flow rate, while H3 and H4 map the deformation profiles at higher flow rates. H1-4 denotes the 4 different conditions of the biophysical assay. The device used in this work is a new design for T cells with a designed cell trajectory specification for a size range from 6 to 12 µm. We have achieved a higher level of system integration and automation compared with our earlier studies on whole blood22, as shown in Supplementary Figs. 1 and 2, a significant step in deploying and enabling direct sample profiling of cells from bioreactors.
a T cells are injected into the microfluidic device, and a CTM assay is performed by subjecting cells to varying flow parameters and microstructures (L and L-flipped) to modulate cell interactions and flow trajectories within the device. The modulated trajectory results in unique cell output positions and signatures for each condition 1, 2, 3, and 4, corresponding to 4 different histograms H1, H2, H3, and H4, respectively. b describes the downstream processing of the signatures using a transform function, resulting in various features based on the modulated biophysical signatures heatmap H1, H2, H3, and H4. These CTM features are grouped into size (S), deformation (D), and combined (C) categories. The CTM features are used in machine classifier models for correlations to T cell functions such as phenotyping and potency.
The modulated biophysical signatures are processed in custom transformation functions (see Fig. 1b and Supplementary Tables 1–3), with each function mapping and integrating the histogram array data into a single value corresponding to the T cell feature of size (S), deformability (D) or a combined (C) feature involving both size and deformability histograms. These biophysical assay features are finally fed into data classification models and statistical analysis to study the correlation to T cell activation, functional phenotype, and potency.
Distinct T cell biophysical states in the CTM assay
We first experimentally checked if the CTM assay can profile distinct physical morphology of T cells (n = 5) with known exposure to anti-CD3/CD28 24-h activation or chemical triggers such as 10% (v/v) DMSO for an additional 15 min post 24-h culture (Fig. 2a–c). Exposure to DMSO is common in CAR T production due to the cryopreservation process, and it is known to cause mechanical changes to cells23,24.
a–c show the H1-4 CTM assay biophysical signatures for three T cell groups at 24 h for unstimulated (control), CD3/28 activation state, and exposure to DMSO for an additional 15 min (mimicking cryopreservation), respectively. d shows unsupervised hierarchical clustering of the used to group the T cell states with respect to scaled CTM heatmap features grouped using a second unsupervised hierarchical clustering. e depicts the grouped CTM features in a Pearson’s correlation heatmap. f Selected features from the different clusters are shown here, with distinct differences in CTM features between all three T cell conditions. The box plots show the bounded 1st and 3rd quartiles with the median as center line; whiskers show the minimum and maximum values. g a principal component analysis (PCA) plot showed clear clusters of the cell conditions. All statistical mean analyses were performed using a two-tailed paired Student’s t-test with *, **, and **** denoting p-values < 0.05, 0.01, and 0.0001, respectively. The p-values are denoted below the significance symbols. Figure data included in Source Data.
The unstimulated samples show sharper singular peaks for all H1-4 conditions, while T cell size profiles (H1 and H2) increase distinctively during CD3/CD28 bead activation25,26 with multiple peaks and an increase in histogram spread. Interestingly, exposure to DMSO for just 15 min changes the biophysical properties of the cells, with significantly reduced size and deformability differences across H1 to H4. This suggests a decrease in cell deformability in high-flow conditions, which is expected upon exposure to DMSO24. Changes in cell deformability are expected since these cryoprotectants interact with the cell membrane and intracellular structures23. These DMSO concentrations are comparable to the concentrations spiked into CAR T products for storage and transportation. For example, Kymriah products add 7.5% (v/v) DMSO in the cryopreservation protocol and may be infused up to 30 min post product thaw. Prolonged DMSO in in vitro cell cultivation was shown to impact the properties of the T cell function; however, in vivo studies have also been shown to be superior to freshly produced CAR T cells.
The CTM features show distinct profiles of these three groups in separate clusters via unsupervised hierarchical clustering, and the features correlation heatmap suggests cluster groups of similar features (Fig. 2d, e). Selected CTM features also show significant differences among the three test groups (Fig. 2f). For example, CTM feature D6 shows low expression levels for unstimulated samples, while C18 shows an inverse relationship. The principal component analysis (PCA) in Fig. 2g shows segregated clusters for all three groups (see Supplementary Fig. 3), which demonstrates a proof of concept that biophysical properties can be used for sensitively probing changes in the T cell state.
Culturing T cells in bioreactors is a highly dynamic process with constantly changing media conditions, and autocrine and paracrine signaling, affecting the cell phenotype27. A single-end-point release test might fail to detect variance during CAR T manufacturing, which precludes any possibility of optimizing cell potency based on attributes measured during production. While the activated cells show clear differences at 24 h using CD3/CD28 beads, the detection of cellular changes early in the activation cycle is of critical importance. These biophysical changes can be detected in the CTM assay as early as 6 h post-CD3/CD28 activation (Supplementary Fig. 4).
Biophysical signatures in cell cultures comparable to flow cytometry forward scatter plots
T cell biophysical morphologies change dynamically over time. These changes can be tracked via flow cytometry or commercial microscopes. Here, we study the utility of the CTM assay for rapid profiling of T cell temporal biophysical signatures under culture conditions. CD3+ T cells were isolated from the peripheral blood mononuclear cells (PBMC), activated, and cultured in a 12-well plate in AIM V media containing interleukin-2 (IL-2). The temporal cell dynamics were profiled every 2 days using flow cytometry, CTM biophysical assay, and quantitative phase imaging (QPI) for cell morphological characterization (Fig. 3a, b, and Supplementary Fig. 5). The cell biophysical features dynamically changed with multiple peaks across the CTM histogram in Fig. 3c. These dynamic cell changes are in line with known activation/expansion dynamics of CAR T production28,29 and were reflected in the CTM histograms.
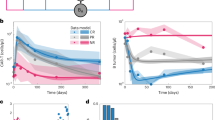
a describes the experiment set up with alternate-day profiling of T cells in a 14-day culture of activated T cell cultures using flow cytometry and CTM assay. b At each time point, cells were imaged using quantitative phase imaging (QPI) in the well plates with a bar scale representing 20 µm. c the frequency distribution and H1-4 histograms of the CTM assay were plotted. d Flow cytometry forward scatter plots of the corresponding cells in culture. e Interpolated forward scatter where the flow cytometry forward scatter data is interpolated into 35 bins for relative comparison with the 35 bins of CTM histogram (H4). Pearson’s correlation was performed between interpolated forward scatter data and CTM histogram, with Pearson’s coefficient shown in ρ and significance of ***, ****, and ns representing two-tailed p-value < 0.001, 0.0001, and not significant, respectively. Figure data included in Source Data.
Corresponding flow cytometry profiles were measured for T cells via forward scatter (FSC) plots, commonly used as a measure of cell size distribution. The FSC plots in Fig. 3d show similar histogram peaks and distributions compared to CTM features. The Pearson’s correlation comparing the CTM H4 profile and interpolated FSC shows that H4 describes a deformed cell profile closely correlated to FSC cell profiles (Fig. 3e). The strong correlation to deformable H4 CTM histogram shows that CTM H4 reflect the shear stress immune cells undergo under high-flow conditions akin to those in flow cytometry. These results demonstrate the use of biophysical size and deformability markers as a viable means the rapidly profiling of T cells in a label-free manner.
CAR T biophysical signatures distinguish CAR T from different bioreactors
We studied whether the biophysical CTM profiling can be used to detect any differences between CAR T cells manufactured using different bioreactors, particularly G-Rex gas-permeable well plates and the Breez perfusion-based microbioreactor. G-Rex is one of the current FDA-approved culture platforms for CAR T cell manufacturing. The Breez perfusion-based microbioreactor was recently used for the manufacturing of CAR T cells, demonstrating comparable expression of activation markers and CAR to other CAR T cell manufacturing methods30.
The CAR T cell manufacturing protocol is performed via bead-based negative isolation of CD3+ cells from commercial frozen PBMC samples from 3 donors (Fig. 4a and “Methods” section). Technical triplicates were performed for each donor and profiled using the biophysical assay. The T cells were transduced with lentivirus to produce anti-CD19 CAR T cells. In both bioreactors, the CAR T cells were expanded for 12 days, and the final products were profiled using CTM, flow cytometry, and a potency cytotoxicity assay to measure the lysis of NALM6 CD19+ cells, which were later correlated with CTM-measured biophysical attributes.
a shows the schematics of CAR T production in G-Rex and Breez from PBMC-isolated CD3+ T cells and the end point 12-day manufactured CD19 CAR T. The end point CAR T product passes through a functional potency cytotoxic assay, flow cytometry profiling, and CTM assay. b Multiple runs (n = 3) of the 3 donors for CAR T cells were performed using the CTM assay, resulting in the scaled heatmap values of CTM features between the two bioreactor groups. c shows the scaled heatmap values of expression levels of selected flow cytometry phenotypes with a comparative significance study shown between the two bioreactor groups. d A scatter volcano plot with significance in Pearson’s correlation and statistical fold difference between CTM and flow cytometry features for Breez and G-Rex CAR T cells. The PCA plot for all the CTM and flow cytometry features was visualized for Breez and G-Rex CAR T cells (e). The fraction of respective features that are significant (p > 0.05) is denoted in the legend. All statistical mean analyses were performed using a two-tailed paired Student’s t-test with *, **, and *** denoting p-value < 0.05, 0.01, and 0.001, respectively. Figure data included in Source Data.
CAR T cells manufactured from the same donor samples using G-Rex or Breez microbioreactor presented distinctly different biophysical signatures, with 21 out of 88 (23.9%) CTM features showing significant differences (Fig. 4b). Unsupervised hierarchical clustering found distinct clusters in the CTM feature heatmap, showing clear regions of high feature expression for Breez but low expression in G-Rex. This suggests that the manufacturing process may impact product attributes given the same or similar starting material, which is in line with other studies on the variation of CAR T in different bioreactors29,31.
Biophysical and phenotype distinction between bioreactors
A panel of T cell phenotypes was profiled (Supplementary Fig. 6) with selected markers of flow cytometry shown in Fig. 4c. Donor-to-donor-specific profiles were dominant in the flow cytometry markers, and distinct phenotypes were seen in the heatmap. 22.4% (34/152) of all flow cytometry markers showed significance between the CAR T cells of the two bioreactors in Fig. 4d. Particularly, the expression of CAR between the two bioreactor modalities did not show a significant difference, contrary to the expression levels of CD127+ and PD1+.
A clear distinction can be appreciated in the unsupervised PCA plots (Fig. 4e) showing donor-specific clusters with distinct CTM profiles, separated by bioreactor types, similarly to flow cytometry PCA plots (see Supplementary Fig. 7). Thus, the biophysical profiling distinguishes the products manufactured from two different bioreactors as well as donor-specific CAR T profiles.
CAR T biophysical markers correlate with CAR T cell phenotypes
During CAR T cell manufacturing, T cells are cultured for up to 12 days, and often, the temporal dynamics of cell functional phenotypes (by flow cytometry) are not explicitly monitored. Here, we explored if CTM markers were functionally correlated with manufactured CAR T cell phenotypes. Interestingly, there were strong correlations between CTM biophysical features and T cell phenotypic characteristics (Fig. 5a). Selected correlation plots critical in CAR T manufacturing, such as CD127+ %, CD4:CD8 ratios, and CAR %, were shown in Fig. 5b–e with strong Pearson’s correlation of −0.93, 0.85, −0.79 and 0.89, respectively.
a shows a Pearson’s correlation heatmap comparing flow cytometry phenotype and CTM biophysical features with significant correlations, two-tailed Pearson’s correlation p-value < 0.05 denoted by a *. Selected correlation plots are shown and indicated in red (b), purple (c), green (d), and blue (e). The Pearson’s correlation analysis performed in (b), (c), (d), and (e) are quantified using a two-tailed Pearson’s correlation test with p-values denoted in the brackets. Figure data included in Source Data.
CD127+ % and S12, which provide a measure of the average size of T cells, both showed significant differences and strongly correlated for CAR T cells manufactured in Breez and G-Rex. On the other hand, no difference in CAR expression was seen in both bioreactors, whereas correlations with CTM markers (such as C14), a combination of cell deformability and size features, remain strong. A complete correlation heatmap showing 152 flow cytometry gating and 88 CTM markers can be seen in Supplementary Fig. 8.
CTM biophysical features correlate with CAR T potency and cytokine profiles
To functionally measure the CD19 targeting and killing potential of the CAR T cell product, the CAR T cells were challenged with target NALM6 cells to quantify the CAR T killing and lysis (Fig. 6a). Multiple CTM features were identified to correlate strongly with killing potency (Fig. 6b and Supplementary Table 4). Three selected CTM features (S5, D21, and C6) were plotted against the 1:5 specific cell lysis %, showing strong linear correlations Fig. 6c–e. CTM features such as S5 show strong correlations for both Breez and G-Rex CAR T cells ( | ρall | > 0.95) while D21 and C6 show bioreactor-specific correlations with different ρBreez and ρGrex, suggesting functional biophysical correlations that are specific to the CAR T manufacturing platform (Fig. 6d, e). Our CTM biophysical features show that the manufacturing protocols, in addition to donor-specificity, can impact the biophysical properties, cellular phenotypes, and corresponding functionality of CAR T cells6,7.
a shows the raw plots of donors 1, 2, and 3 in the CAR T potency assay for CAR T cells manufactured using Breez and G-Rex. All points in the plot represent the mean ± SD. b displays the correlation heatmap of CTM features versus effector:target (E:T) ratios. 1:5 E:T ratios best describe the kinetics of the potency assay with close to 50% killing efficiency, and the correlative plots are selectively displayed for S5, D21, and C6 versus 1:5 E:T specific lysis % in (c), (d), and (e), respectively. All data presented are mean ± SD. ρall represents the two-tailed Pearson’s correlation of Breez and G-Rex measurements combined, while ρBreez and ρGrex represent the Pearson’s correlation coefficient of Breez and G-Rex separately. f shows the correlation heatmap of the CTM features and selected cytokine profiles identified by the colored dot with Pearson’s correlation profiles of ρcytokine. Selected data points are shown in (g), (h), (i), and (j) for S26, S17, D6, and S4 compared to IFN-γ, GM-CSF, IL-5, and GZB, respectively. All data presented here are mean ± SD. All correlation heatmap profiles are based on Pearson’s correlation analysis, and strongly correlated features with a two-tailed p-value < 0.05 are marked with * in the heatmap. For individual plots, Pearson’s correlative values are shown in ρ while the significances of the correlation are labeled as *, **, ****, and ns, denoting two-tailed p-value < 0.05, 0.01, 0.0001, and not significant, respectively. Figure data included in Source Data.
Following co-culture with target NALM6 cells, cytokine secretion of n = 3 donors was also profiled for Breez (B1, 2, and 3) and G-Rex (G1, 2, and 3) as seen in the Supplementary Table 5. Pearson’s two-tailed correlation analysis with CTM assay was performed (Fig. 6f and Supplementary Fig. 9). Negative correlations were seen for IL-3, TNF, LTα, and GM-CSF, as these are known co-factors that regulate immune cell responses32.
Positive correlations with CTM profiles were observed for IL-5 and IL-13. Both IL-5 and IL-13 have cellular pathways that are somewhat interrelated, as they both ultimately contribute to Th2 immune responses and have been shown to relate to certain pathophysiology, such as asthma33,34. Selected correlation plots are shown in Fig. 6g–j. Minimal correlations across the CTM features were observed for IL-6, IFN-y, and GZB (Granzyme b). The strongest correlation of GZB is observed in Fig. 6j with a correlation of 0.95 (p-value = 0.004). Collectively, CTM features show a strong correlation with CAR T cell function in potency and cytokine secretion profiling.
Aggregated CAR T cell linear biophysical indexing
To enable comparison across all culture conditions and experiments, a biophysical index based on linear principal component analysis (PCA) was performed to reduce the 88 features into two principal components in PCA (Fig. 7a and Supplementary Fig. 10). The principal component 1 captures 44% of the aggregated CTM features variance and was used as the linear biophysical index (Supplementary Table 6 and Supplementary Table 7).
The schematics for workflow on biophysical indexing of profiled T cell biophysical features, which are dimensionally reduced using linear PCA with the dominant principal component (PC1) used as an indexing measure, given its representation of the CTM biophysical features of maximal variance (a). The biophysical indexing was performed on n = 3 healthy donors with triplicates each (b), a single donor with n = 6 replicates (c), and n = 4 healthy versus patient CAR T comparison on Breez (d). The box plots show the bounded 1st and 3rd quartiles with median as center line; whiskers show the minimum and maximum values (see Supplementary Table 7). In addition to the box plots, the mean ± SD value is denoted beside the boxplot. The two-tailed Student’s t-test is used to perform the statistical analysis for (a), (b), and (c) with **, ****, and ns denoting p-value < 0.01, 0.0001, and not significant, respectively. Figure data included in Source Data.
The CTM biophysical index showed a significant difference between mean ± SD for Breez (−0.083 ± 0.74) and G-Rex (−1.90 ± 0.46) manufactured CAR T cells, which was profiled on the final day of manufacture Fig. 7b. This data is in agreement with earlier analysis where individual CTM features showed statistically significant differences for CAR T cells in the n = 3 donors each in a triplicate study shown in Fig. 4b, d, e. To investigate the effects of donor CAR T heterogeneity during manufacturing, a single donor sample was used in n = 6 independent batches of CAR T manufacturing runs for Breez and G-Rex each (Fig. 7c). Each batch of the CAR T products was frozen in liquid nitrogen (LN2). Together, the samples were thawed, and the CTM biophysical profiling, flow cytometry, and potency assay were performed for the donor replicates on the same day. The single donor replicates in Fig. 7c show that the variation between the same donor can be significantly large after 12 days of CAR T manufacturing for Breez (1.70 ± 1.10), while variations of G-Rex (−0.62 ± 0.42) remain comparable to Fig. 7b CAR T cells from 3 donor triplicates (−1.90 ± 0.46). There is still a significant biophysical difference between cryopreserved CAR T products from Breez and G-Rex (Fig. 7c).
CAR T cells from 4 cancer patients were manufactured in Breez and compared to 4 healthy donors (Fig. 7d). Each CAR T batch was cryopreserved, and the CAR T cell characterizations were performed after resting for two days after thawing. The aggregated biophysical index showed that there is no significant difference between the patient (1.70 ± 0.66) and healthy donor (1.16 ± 0.72) CAR T cells manufactured in Breez. Interestingly, the patient CAR T cells have the same biophysical index as the same donor replicate Breez CAR T cells in Fig. 7c. The patient-derived CAR T potency and CTM correlations are shown in Supplementary Fig. 11.
In this experiment, all independent Breez (n = 23) and G-Rex (n = 15) CAR T samples showed significant biophysical profile differences of 0.91 ± 1.17 versus −1.39 ± 0.77, respectively. (Supplementary Fig. 12). Principal component indexing performed on aggregated flow cytometry phenotype and potency assay shows very similar indexing trends with the CTM biophysical index (Supplementary Fig. 13, Supplementary Table 6). With multiple CTM features showing correlation with clinical CAR T potency test, it is plausible that CTM index strongly correlates with CAR T potency and flow cytometry phenotype with two-tailed Pearson’s correlations coefficient of −0.82 (p-value = 3.6 × 10−10) and 0.68 (p-value = 2.9 × 10−6), respectively (Supplementary Fig. 14).
Collectively, the results show CTM biophysical phenotype distinctly differentiating manipulated T cells in 24-well plate cultures and manufactured CAR T cells from different bioreactors. These biophysical properties correlate with both surface markers phenotyping and functional profiles of the T cells.
Discussion
While immune cell morphological features, such as size, deformability, and shape, have been shown to be strong indicators of cell activation, metabolic activity, cell cycle, and disease pathology35,36,37,38, these parameters are not yet adapted for direct assessment of autologous CAR T cells in manufacturing. This study demonstrates deformability characteristics of CAR T cells as correlative and potentially surrogate markers for critical quality attributes, including killing potency. Of note, our CTM assay detected distinct biophysical signatures between CAR T cells manufactured from two different culture platforms, Breez and G-Rex, despite using the same donors with the same transfection and activation modality. Using an aggregated biophysical index, comparison with single donor replicates and patient samples shows consistent biophysical profile characteristics for each bioreactor. Similarly, Song et al. evaluated CAR T cells manufactured from different bioreactors and found significant phenotypic differences between CAR T cells manufactured using different bioreactors31, albeit using a potency functional assay and flow cytometry phenotyping. The potential for such a system to rapidly measure functional phenotypes of CAR T cells when deployed closer to the bioreactors is invaluable for the growing field of CAR T bioprocessing, especially in the adaptive manufacturing of next-generation CAR T products18,39.
Hale et al.’s recent publication on morphological phenotypes of CD3 cells driving unique functional characteristics is new evidence of the potential biophysical correlation to cell states previously unknown40. While discussions on cell deformability are lacking, it can be hypothesized, based on evidence from other groups on cell deformability41,42, that the immune cell architecture will influence the biophysical properties of size and deformability, which determine function. In the case of CAR T cells for immunotherapy, biophysical changes due to variations in activation protocols may impact how T cells interact with the endothelium in different tissues, resulting in differences in in vivo cell distribution and trapping26,43,44,45. Waugh et al. showed that selected subpopulation activation showed different biophysical changes in naïve T cells versus CD8+ T cells44. Natelie et al. investigated that a wide range of inflammatory conditions influence the mechanical properties of immune cells, which can significantly impact their function independent of T cell surface maturation markers26. Thus, the potency of CAR T cells to kill effectively depends on their ability to detect tumor chemokine signatures and on cell migration or extravasation into the tumor cells. The CTM assay specifically probes cell size and deformation of T cells, which may be indirectly related to the cell extravasation potential46. Thus, a correlation between the CTM features and CAR T potency is surprising yet probable.
The mechanobiology of T cells is a nascent but growing field, where conventional techniques to probe the mechanical response of cells focus on static, single-cell responses in micro pipetting technique43,44 and atomic force microscopy47,48. These techniques are manual and have low throughput; therefore, they are not adequate in cell manufacturing control, where automation, speed, and consistency are required. Microfluidic technologies such as cell deformability cytometry38,49 and optical force cytometry50 offer high-content single-cell biophysical measurements albeit with the need for high magnification laser optical setup or costly highspeed cameras; constriction pillar microfluidics are often easier to operate albeit with increased likelihood of clogging using gap sizes smaller than the target profiled cell51; The use of viscoelastic fluids and impedance to probe cell deformability shows distinct in vitro activated states of neutrophils52. While the CTM requires a sequential change of flow rates to induce cell deformation, it can handle a large range of cell concentrations in native culture media and uses a 4× objective magnification optical setup with a standard machine vision camera.
Herein, we also investigated the correlation between surface marker phenotyping of T cells and found that certain biophysical features strongly correlated with CAR T cell phenotypes. T cell geometrical deformations directly induce nuclear morphological changes and alter gene expression through the nuclear factor of activated T cells and other signaling cascades53. Thus, the biophysical and biochemical domains of T cells are complicated but clearly linked54,55. In the current manufacturing protocols of autologous CAR T cells, monitoring of cell phenotypes (CD4, CD8, Anti-CD19 CAR) is not performed daily, mainly because of the limited cell number produced, and the time and risk associated with the manual handling56. Given the correlative potential, our label-free assay only requires 20 µL of sample and completes in 10 min, enabling potential use for perfusion and in-process monitoring of quality attributes of CAR T cells for the next generation of adaptive manufacturing18,57.
Flow cytometry-based phenotyping between G-Rex and Breez-produced CAR T cells showed significant, if subtle, differences, which we believe represent subtle potential functional and phenotypic differences. The CTM profiles are in agreement with this difference, and the observed biophysical correlation and distinction of CAR T cells in both bioreactors would be a valuable area of future study. Insights from various ongoing research in biophysical deformability profiling measures the collective functionality and intracellular dynamics of cells, can go beyond what flow cytometry surface markers can measure on their own58,59.
Continuous testing during autologous CAR T manufacturing and wide testing at release is hampered by limited cell availability (e.g., Novartis T Charge with a total dose of >2.5 million CAR T cells60,61). This necessitates the use of simple markers or surrogate assays for potency62,63. Unfortunately, simple markers such as CD19 CAR expression levels and dose11 do not correlate well with clinical efficacy. There are no precise immunophenotypic profiles that are direct predictors of CAR T cell function64. In vitro killing or surrogate assays are arguably the closest to validating in vivo CAR T potency64,65. Existing killing assays typically require >10 h of manual cell processing and handling, excluding sample preparation, making it not feasible for pre-infusion testing of autologous CAR T cell products66. Alternatively, the biophysical label-free CTM assay, demonstrating potential correlation with function and in vitro potency, emerges as a strong surrogate for in-process monitoring, requiring only a small volume (20 μL) with at least 1000 T cells for histogram distribution. Its rapid turnaround time is particularly advantageous for autologous products with limited shelf life before patient administration, ensuring timely functional and potency testing. While there is a strong correlation between our assay and in vitro killing assays, this does not suggest a correlation with in vivo efficacy, and downstream validations are in the works.
The CTM assay is a continuous flow and non-destructive cell profiling method, and can be potentially integrated into the bioreactors for in-line Process Analytics Technology (PAT) or even as a closed-loop cell culture monitoring and control system for adaptive manufacturing of next-generation CAR T products18,39. With a compact detection modality using commercially available machine vision cameras, low magnification bright field objectives, and standard laptop processors, the CTM assay is adaptable and deployable when compared to current start-of-the-art cell deformability measurement tools utilizing ultra-high-speed cameras and server specification computers for fast frame rate captures and image processing35,67. However, there are several limitations to the current CTM assay. First, it remains unclear whether the selected transformational features are comprehensive and optimal for biophysical measurements. Furthermore, how immune cells deform and interact with different microfluidic structures as a function of flow rate is not well understood, making it challenging to determine the ideal conditions for modulating these structures. As a result, the micropillar configurations explored in this study are limited to those that are already well-characterized. Additionally, while the CTM assay is effective in profiling shifts in immune cell distribution, sorting cells within individual CTM bins remains highly challenging as it entails more than 30 outlet sorting channels. However, this approach may eventually provide valuable single-cell mechanical insights.
This study introduces a microfluidic assay that offers a rapid and non-destructive means to profile the biophysical attributes of T cells, particularly CAR T cells, and to correlate these attributes with functional phenotypes. This assay addresses a critical need in CAR T cell therapy by enabling real-time, label-free monitoring of critical cellular characteristics during and after the manufacturing process. Ultimately, this approach supports adaptive manufacturing strategies by offering insights into CAR T cell production, which may, in turn, influence their in vivo efficacy.
Methods
Microfluidic chip fabrication
The CTM microfluidic assay, as shown in Fig. 1a, consists of a single microfluidic chip integrating two microfluidic devices with designs adapted from Zeming et al.22,68 with the L and inverse-L micropillar structures. The improvements to chip designs include an expanded dynamic range to measure an approximated output size range from 6 to 12 µm with a single sample inlet and outlet (Supplementary Fig. 1). The microfluidic device specifications are provided in Supplementary Fig. 15 and Supplementary Table 8. The CTM device consists of polydimethylsiloxane (PDMS) material (Sylgard 184, Dow Corning) bonded to a clear soda lime glass slide. The device is fabricated based on established soft photolithography means. Briefly, starting with computer-aided drawing, the designs are transferred to a glass chromed photomask (JD photodata), followed by using UV lithography means. An SU-8 coating of 20 microns is coated on the surface of a 4-inch wafer, baked, and exposed to UV light using the chrome photomask. The final developed features on the wafer are coated with hydrophobic trichloro(1,1,2,2-perfluoroocytl)silane (Sigma, Singapore) using chemical vapor deposition. Finally, PDMS is mixed using a 1:10 curing ratio, degassed, and poured onto the wafer to be baked at 80 °C for 90 min. The cured PDMS is peeled off the wafer and processed by cutting it into the correct device dimensions with reservoirs and tubing insert holes punched. The final device is obtained by bonding the fabricated PDMS to a clear glass slide using a plasma generator.
CTM assay
The CTM assay shown in Fig. 1a depicts a 4-condition process to modulate the trajectory of cell flow measured at the locating bins (Supplementary Fig. 2). At least 20 µL of T cell samples are directly loaded onto the sample loading reservoirs on the chip. The CTM assay can process undiluted cell concentrations at a minimum of 1 × 106 cells mL−1. Cell concentrations processed here range from 1 × 106 cells mL−1 to 80 × 106 cells mL−1. The CTM assay is driven by negative pressure at the outlet with flow driving pressure for 1× at −50 mBar while 10× flow is driven at −500 mBar (see Supplementary Fig. 1). The custom CTM optical setup shown in Supplementary Fig. 1 uses a 4× Olympus infinite objective lens with a 2 mm field of view and is focused using a 60 mm lens to project the image to a machine vision camera (BFS-U3-16S2M, FLIR Blackfly2). The cell outlet trajectory positions are recorded on the camera to identify single cells (see Supplementary Fig. 2) and capture the images at 15 FPS for 1× flow and 150 FPS for 10× flow. The time taken to record the locating bins at each outlet in Fig. 1a for the 4 conditions is 4, 4, 1, and 1 min to establish the modulated biophysical signatures in H1, H2, H3, and H4, respectively.
CTM data processing
To enable analysis to output the H1–4 biophysical signatures, video frames recorded from the camera are processed in buffered real-time data using Python 3.10.11 scripts for OpenCV 4.6.0.66 background subtraction, blob detection, and histogram bin tabulation for sequential conditions. The processed data is saved and post-processed to the CTM features using a secondary script to import the H1-4 and process the transform functions, resulting in 88 features. Machine classification of the features and phenotype data was performed using existing modules in Python, using Numpy 1.26.2, Pandas 1.5.3, Scikit-learn 1.3.2, and SciPy 1.11.4, for all data normalization, statistical analysis, correlation analysis, PCA data, and hierarchical clustering. All statistical analyses for sample mean testing were performed using a two-tailed Student’s t-test, while correlation analysis was evaluated with Pearson’s correlation. The data visualization and plots were performed using Matplotlib 3.8.2 and Seaborn 0.11.2 Python modules. Data from all Figures for CTM, flow cytometry, and potency are tabulated in Source Data, cytokine data is shown in Supplementary Table 5. All processed data were normalized on a logarithmic scale. The data of PCA models for CTM, flow cytometry, and potency are tabulated in Supplementary Table 6.
Quantitative phase imaging
Quantitative phase images (QPI) were taken on a Leica microscope with incubator attachments for culture well plates to allow imaging of T cells in the culture wells for the longitudinal studies in Fig. 3. The QPI method69 and setup are based on a previous publication by Lee et al.70. The T cell QPI images were acquired on Days 0, 2, 4, 6, 8,10, 12, and 14 to characterize the morphology of the cells. The raw QPI data are shown in Source Data with strong correlations with CTM biophysical features (see Supplementary Fig. 4).
T cell isolation from PBMCs
CD3+ T cells were isolated using the EasySep Human T Cell Isolation Kit (STEMCELL Technologies, Cat. No. 17951). Activation using CD3/CD28 Dynabeads (Thermo Fisher Scientific, Cat. No. 11141D) was performed for 0–24 h in 24-well plate cultures (Thermo Fisher Scientific) for naïve control, CD3/CD28 activation, and 10% (v/v) DMSO experiments. CD3/CD28 beads were removed prior to the CTM assay. T cell experiments in Fig. 3 were activated using ImmunoCult Human CD3/CD28 T Cell Activator (STEMCELL Technologies, Cat. No. 10971). Cells were transferred into a 12-well glass-bottom plate for QPI imaging.
Cell lines
Lenti-X 293T Cell Line was purchased from Takara Bio Inc. and maintained in HyClone Dulbecco’s Modified Eagle Medium (DMEM) with high glucose (Thermo Fisher Scientific, Cat. No. 11965-084) supplemented with 10% Fetal Bovine Serum (Thermo Fisher Scientific, Cat. No. 16140-071) and 1% Penicillin-Streptomycin (Thermo Fisher Scientific, Cat. No. 15140-122). Jurkat, Clone E6-1 (TIB-152) and NALM6, clone G5 (CRL-3273) cell lines were purchased from ATCC and maintained in HyClone RPMI 1640 medium (Cytiva, Cat. No. SH30255.01) supplemented with 10% Fetal Bovine Serum (Thermo Fisher Scientific, Cat. No. 16140-071) and 1% penicillin-streptomycin (Thermo Fisher Scientific, Cat. No. 15140-122).
Plasmids
pMDLg/pRRE, pRSV-Rev, and pMD2.G packaging plasmids were gifts from Didier Trono (Addgene plasmid # 12251, 12253, and 12259)71. A second-generation anti-CD19 CAR-IRES-EGFP transfer plasmid, encoding a codon-optimized anti-CD19 CAR comprised of Myc-epitope-tagged FMC63 single-chain variable fragments, IgG4 hinge, CD28 transmembrane domain, and human 4-1BB and CD3zeta intracellular signaling domains72 was PCR amplified from geneblocks (IDT) and cloned into the third-generation lentiviral vector pHIV-EGFP (Addgene plasmid # 21373) using Gibson Assembly. Research-grade packaging and transfer plasmids were then prepared by GenScript Biotech.
Lentiviral vector production and titration
Lentiviral vectors were generated by first transfecting the Lenti-X 293T Cell Line with the packaging plasmids and transfer plasmid complexed with Lipofectamine 3000 Transfection Reagent (Thermo Fisher Scientific, Cat. No. L3000-075). Medium was changed every 4–6 h after transfection, and lentiviral supernatants were collected 48–96 h after transfection. The supernatant was then centrifuged for 5 min at 300 g, 4 °C, and filtered through a 0.45-μm low-protein-binding filter. The clarified supernatant was then ultracentrifuged for 1.5 h at 75,000 × g, 4 °C. The lentiviral pellet was then resuspended in Opti-MEM Reduced Serum Medium (Thermo Fisher Scientific, Cat. No. 31985-070) at 4 °C and stored in aliquots at −80 °C. The reagents used are shown in Supplementary Table 9. The concentrated lentiviral vectors were titrated on Jurkat cells. The percentages of GFP+ Jurkat cells were measured by flow cytometry two days post-transduction, and lentiviral titers were calculated and expressed in TU/mL.
CAR manufacturing in bioreactors
Frozen human peripheral blood mononuclear cells (PBMC) from 3 healthy donors were purchased from STEMCELL Technologies. For patient samples, PBMCs were donated from four anonymous lymphoma patient samples (NCT05648019) and extracted from discarded leukapheresis tubing sets (Spectra Optia Apheresis System, Terumo BCT). All uses of human clinical material have been approved by the SingHealth Centralized Institutional Review Board (CIRB) approval: #2018/2677 and #2022/2322. All recruited volunteers provided written informed consent. All recruited volunteers provided written informed consent. The PBMCs were isolated using Ficoll-Paque density gradient centrifugation (Cytiva, Cat. No. 17144003) at 400 × g for 30 min, cryopreserved in CryoStor CS10 (STEMCELL Technologies, Cat. No. 07930), and later thawed for CAR T cell production. T cells were isolated using EasySep Human T Cell Isolation Kit (STEMCELL Technologies, Cat. No. 17951) and resuspended in AIM V Medium (Thermo Fisher Scientific, Cat. No. 12055-083) supplemented with 2% human male AB serum (Merck Sigma–Aldrich, Cat. No. H4522) and 100 IU mL−1 recombinant human IL-2 (Miltenyi Biotec, Cat. No. 130-097-748). The detailed protocols, preparation, and operation of G-Rex and Breeze for CAR T manufacturing were detailed in Sin et al.30. Briefly, for activation, purified T cells were mixed with DynaBeads Human T-Expander CD3/CD28 (Thermo Fisher Scientific, Cat. No. 11141D) at a 1:1 cell:bead ratio and seeded into each gas-permeable well of a G-Rex 24-well plate (Wilson Wolf) and each microbioreactor cassette (Breez Biosystems, MilliporeSigma). The list of reagents is shown in Supplementary Table 10. One day after activation, lentiviral vectors were added to the wells or cassettes at a multiplicity of infection (MOI) of 5. T cells were expanded over 10 days for a total process duration of 12 days.
Potency assay
Cryopreserved end-of-production T cells were thawed, rested overnight (for experiments in Figs. 4–5) or for 2 days, and co-cultured overnight with NALM6-Luciferase cells at various effector:target (E:T) ratios (CAR+ T cell: NALM6 cell) in RPMI 1640 medium (Cytiva, Cat. No. SH30255.01) supplemented with 10% Fetal Bovine Serum (Thermo Fisher Scientific, Cat. No. 16140-071) and 1% Penicillin-Streptomycin (Thermo Fisher Scientific, Cat. No. 15140-122). For the luciferase luminescence-based cytotoxicity assay with NALM6-Luciferase cells, co-cultured cells were transferred to a white 96-well opaque-bottom plate. Bright-Glo Luciferase Assay reagent (Promega, Cat. No. E2610) was added to the wells, and luminescence signals were measured using an Infinite M200 Pro plate reader (Tecan).
Cytokine profiling
Freshly produced T cells were co-cultured overnight with NALM6 cells at a 1:1 effector-to-target (E:T) ratio (CAR+ T cell:NALM6 cell) in AIM V Medium (Thermo Fisher Scientific), supplemented with 2% human male AB serum (Merck Sigma–Aldrich) and 100 IU mL−1 recombinant human IL-2 (Miltenyi Biotec). Cell culture supernatants were collected and stored at −80 °C. Cytokine and chemokine secretion profiles were assessed using the Human XL Cytokine Luminex Performance Assay 46-plex Fixed Panel (R&D Systems, Cat. No. LKTM014B) and analyzed on a Luminex FLEXMAP 3D system (Thermo Fisher Scientific). Due to high levels of Granzyme B and IFN-γ exceeding the assay’s standard range, these cytokines were re-evaluated using the Human Granzyme B and IFN-γ ProcartaPlex Kit (Thermo Fisher Scientific, Cat. No. EPX01A-12027-901, EPX01A-10228-901) on the same Luminex FLEXMAP 3D system.
Flow cytometry panels
For the experiments in Fig. 4 and Supplementary Figs. 5 and 6, flow cytometry was performed using the gating strategy shown in Supplementary Fig. 16. with catalog and dilution factor shown in Supplementary Table 11. These are the following antibodies: LIVE/DEAD Fixable Violet (Thermo Fisher Scientific, Cat. No. L34955), BV510 anti-CD4 (clone OKT4, 317444), BV605 anti-CD45RA (clone HI100, 304134), BV650 anti-CD8a (clone RPA-T8, 301042), PerCP/Cy5.5 anti-LAG-3 (clone 11C3C65, 369312), PE anti-CCR7 (clone G043H7, 353204), PE/Dazzle 594 anti-CD127 (clone A019D5, 351336), PE/Cy7 anti-CD57 (clone HNK-1, 359624), APC anti-CD62L (clone DREG-56, 304810), AF700 anti-PD-1 (clone EH12.2H7, 329952), and APC/Fire 750 anti-TIM-3 (clone F38-2E2, 345044) (Biolegend). Cells were washed with PBS (Thermo Fisher Scientific, Cat. No. 10010-031) and stained by incubating with LIVE/DEAD Fixable Violet Stain for 20 min at room temperature. Following this, cells were washed with FACS Buffer [PBS containing 2% Fetal Bovine Serum (Thermo Fisher Scientific) and 0.1% sodium azide (Merck Sigma–Aldrich)], then incubated with antibodies for 20 min at room temperature for staining. The cells were washed again with FACS Buffer before being analyzed on a CytoFLEX S flow cytometer (Beckman Coulter).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are included in the Supplementary Information or available from the authors, as are unique reagents used in this Article. The raw numbers for charts and graphs are available in the Source Data file whenever possible. Source data are provided with this paper.
References
Cliff, E. R. S. et al. High cost of chimeric antigen receptor T-cells: challenges and solutions. Am. Soc. Clin. Oncol. Educ. Book, 43 e397912, (2023).
Bersenev, A. & Kili, S. Management of ‘out of specification’ commercial autologous CAR-T cell products. Cell Gene Ther. Insights 4, 1051–1058 (2018).
Cohet, G. et al. Failure and out of specification manufacturing of autologous CAR-T cells could be associated with a high concentration of total nucleated cells, CD3 + cells and neutrophils in the apheresis product. Blood 142, 3520 (2023).
Novartis Pharmaceuticals Corporation. BLA 125646, tisagenlecleucel. In U.S. Department of Health and Human Services, Food and Drug Administration, US Oncologic Drugs Advisory Committee Meeting (Food and Drug Administration, 2017).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 26, 1878–1887 (2020).
Levine, B. L., Miskin, J., Wonnacott, K. & Keir, C. Global manufacturing of CAR T cell therapy. Mol. Ther. - Methods Clin. Dev. 4, 92–101 (2017).
Tyagarajan, S., Spencer, T. & Smith, J. Optimizing CAR-T cell manufacturing processes during pivotal clinical trials. Mol. Ther. Methods Clin. Dev. 16, 136–144 (2020).
Fraietta, J. A. et al. Determinants of response and resistance to CD19 chimeric antigen receptor (CAR) T cell therapy of chronic lymphocytic leukemia. Nat. Med. 24, 563–571 (2018).
Capelli, C. et al. Potency assays and biomarkers for cell-based advanced therapy medicinal products. Front. Immunol. 14, 1186224 (2023).
Abramson, J. S. et al. Lisocabtagene maraleucel for patients with relapsed or refractory large B-cell lymphomas (TRANSCEND NHL 001): a multicentre seamless design study. Lancet 396, 839–852 (2020).
Awasthi, R. et al. Tisagenlecleucel cellular kinetics, dose, and immunogenicity in relation to clinical factors in relapsed/refractory DLBCL. Blood Adv. 4, 560–572 (2020).
Summary Basis for Regulatory Action for BLA 125643 (YESCARTA™) (U.S. Food and Drug Administration, Review Committee, 2017).
Holthaus, L. et al. CD4(+) T cell activation, function, and metabolism are inhibited by low concentrations of DMSO. J. Immunol. Methods 463, 54–60 (2018).
de Abreu Costa, L. et al. Dimethyl sulfoxide (DMSO) decreases cell proliferation and TNF-α, IFN-γ, and IL-2 cytokines production in cultures of peripheral blood lymphocytes. Molecules https://doi.org/10.3390/molecules22111789 (2017).
Kondo, A. T. et al. Thawing and infusion of CAR-T cell products. J. Bone Marrow Transpl. Cell. Ther. 3, 165 (2022).
Fesnak, A. D. The challenge of variability in chimeric antigen receptor T cell manufacturing. Regen. Eng. Transl. Med. 6, 322–329 (2020).
Abou-el-Enein, M. et al. Scalable manufacturing of CAR T cells for cancer immunotherapy. Blood Cancer Discov. 2, 408–422 (2021).
Blache, U., Popp, G., Dünkel, A., Koehl, U. & Fricke, S. Potential solutions for manufacture of CAR T cells in cancer immunotherapy. Nat. Commun. 13, 5225 (2022).
Huang, L. R., Cox, E. C., Austin, R. H. & Sturm, J. C. Continuous particle separation through deterministic lateral displacement. Science 304, 987–990 (2004).
Hochstetter, A. et al. Deterministic lateral displacement: challenges and perspectives. ACS Nano 14, 10784–10795 (2020).
Holmes, D. et al. Separation of blood cells with differing deformability using deterministic lateral displacement(dagger). Interface focus 4, 20140011 (2014).
Zeming, K. K. et al. Label-free biophysical markers from whole blood microfluidic immune profiling reveal severe immune response Signatures. Small 17, 2006123 (2021).
Gironi, B. et al. Effect of DMSO on the mechanical and structural properties of model and biological membranes. Biophys. J. 119, 274–286 (2020).
Golan, M. et al. AFM monitoring the influence of selected cryoprotectants on regeneration of cryopreserved cells mechanical properties. Front. Physiol. https://doi.org/10.3389/fphys.2018.00804 (2018).
Charpentier, J. C. et al. Macropinocytosis drives T cell growth by sustaining the activation of mTORC1. Nat. Commun. 11, 180 (2020).
Bufi, N. et al. Human primary immune cells exhibit distinct mechanical properties that are modified by inflammation. Biophys. J. 108, 2181–2190 (2015).
Kirouac, D. C. & Zandstra, P. W. The systematic production of cells for cell therapies. Cell Stem Cell 3, 369–381 (2008).
Fraessle, S. P. et al. Activation-inducible CAR expression enables precise control over engineered CAR T cell function. Commun. Biol. 6, 604 (2023).
Watanabe, N., Mo, F. & McKenna, M. K. Impact of manufacturing procedures on CAR T cell functionality. Front. Immunol. https://doi.org/10.3389/fimmu.2022.876339 (2022).
Sin, W.-X. et al. A high-density microfluidic bioreactor for the automated manufacturing of CAR T cells. Nat. Biomed. Eng. 8, 1571–1591 (2024).
Song, H. et al. 145Comparison of CAR-T cell manufacturing platforms reveals distinct phenotypic and transcriptional profiles. J. Immunother. Cancer 9, A153–A153 (2021).
Borriello, F. et al. GM-CSF and IL-3 modulate human monocyte TNF-α production and renewal in in vitro models of trained immunity. Front. Immunol. https://doi.org/10.3389/fimmu.2016.00680 (2017).
Tomasiak-Łozowska, M. M., Bodzenta-Łukaszyk, A., Tomasiak, M., Skiepko, R. & Zietkowski, Z. [The role of interleukin 13 and interleukin 5 in asthma]. Postepy Hig. Med. Dosw. 64, 146–155 (2010).
Agache, I. et al. Serum IL-5 and IL-13 consistently serve as the best predictors for the blood eosinophilia phenotype in adult asthmatics. Allergy 71, 1192–1202 (2016).
Herbig, et al. In Flow Cytometry Protocols (eds Teresa S. Hawley & Robert G. Hawley) 347–369 (Springer New York, 2018).
Bashant, K. R., Vassallo, A., Herold, C. & Berner, R. Real-time deformability cytometry reveals sequential contraction and expansion during neutrophil priming. J. Leukoc. Biol. 105, 1143–1153 (2019).
Lin, W. et al. Morphological change of CD4+ T cell during contact with DC modulates T-cell activation by accumulation of F-actin in the immunology synapse. BMC Immunol. 16, 49 (2015).
Rosendahl, P., Plak, K., Jacobi, A. & Kraeter, M. Real-time fluorescence and deformability cytometry. Nat. Methods 15, 355–358 (2018).
Ran, T., Eichmüller, S. B., Schmidt, P. & Schlander, M. Cost of decentralized CAR T-cell production in an academic nonprofit setting. Int. J. Cancer 147, 3438–3445 (2020).
Hale, B. D. et al. Cellular architecture shapes the naïve T cell response. Science 384, eadh8697 (2024).
Crawford, K. et al. Rapid biophysical analysis of host immune cell variations associated with sepsis. Am. J. Respir. Crit. Care Med. 198, 280–282 (2018).
Ekpenyong, A. E., Toepfner, N., Chilvers, E. R. & Guck, J. Mechanotransduction in neutrophil activation and deactivation. Biochim. Biophys. Acta 1853, 3105–3116 (2015).
Guillou, L. et al. T-lymphocyte passive deformation is controlled by unfolding of membrane surface reservoirs. Mol. Biol. Cell 27, 3574–3582 (2016).
Waugh, R. E., Lomakina, E., Amitrano, A. & Kim, M. Activation effects on the physical characteristics of T lymphocytes. Front. Bioeng. Biotechnol. 11, 1175570 (2023).
Wienke, J. et al. T cell interaction with activated endothelial cells primes for tissue-residency. Front. Immunol. 13, 827786 (2022).
Jung, S., Wulftange, W. J., Fadaei, P., Man, Y. & Gurkan, U. A. Microfluidic assessment of T cell deformability and capillary network occlusion. Blood 142, 3651–3651 (2023).
Hu, K. H. & Butte, M. J. T cell activation requires force generation. J. Cell Biol. 213, 535–542 (2016).
Harrison, D. L., Fang, Y. & Huang, J. T-Cell mechanobiology: force sensation, potentiation, and translation. Front. Phys. https://doi.org/10.3389/fphy.2019.00045 (2019).
Nawaz, A. A. et al. Intelligent image-based deformation-assisted cell sorting with molecular specificity. Nat. Methods 17, 595–599 (2020).
Hebert, C. G., Terray, A. & Hart, S. J. Toward label-free optical fractionation of blood—optical force measurements of blood cells. Anal. Chem. 83, 5666–5672 (2011).
An, L., Ji, F., Zhao, E., Liu, Y. & Liu, Y. Measuring cell deformation by microfluidics. Front. Bioeng. Biotechnol. 11, 1214544 (2023).
Petchakup, C. et al. Microfluidic impedance-deformability cytometry for label-free single neutrophil mechanophenotyping. Small 18, e2104822 (2022).
Dutta, D. et al. Recruitment of calcineurin to the TCR positively regulates T cell activation. Nat. Immunol. 18, 196–204 (2017).
Mukhopadhyay, M. Immunomechanics. Nat. Methods 20, 35–35 (2023).
Li, R. & Ma, C. The CAR T-cell mechanoimmunology at a glance. Adv. Sci. 7, 2002628 (2020).
Wang, X. & Rivière, I. Clinical manufacturing of CAR T cells: foundation of a promising therapy. Mol. Ther. Oncolytics https://doi.org/10.1038/mto.2016.15 (2016).
Hort, S. et al. Toward rapid, widely available autologous CAR-T cell therapy – artificial intelligence and automation enabling the smart manufacturing hospital. Front. Med. https://doi.org/10.3389/fmed.2022.913287 (2022).
Hur, S. C., Henderson-MacLennan, N. K., McCabe, E. R. B. & Di Carlo, D. Deformability-based cell classification and enrichment using inertial microfluidics. Lab Chip 11, 912–920 (2011).
Makhija, E., Jokhun, D. S. & Shivashankar, G. V. Nuclear deformability and telomere dynamics are regulated by cell geometric constraints. Proc. Natl Acad. Sci. USA 113, E32–E40 (2016).
Sperling, A. S. et al. P1446: Phase I study data update of Phe885, a fully human BCMA-directed CAR-T cell therapy manufactured using the T-Chargetm platform for patients with relapsed/refractory (R/R) multiple myeloma (Mm). HemaSphere 6, 1329–1330 (2022).
Dickinson, M. J. et al. A novel autologous CAR-T therapy, YTB323, with preserved T-cell stemness shows enhanced CAR T-cell efficacy in preclinical and early clinical development. Cancer Discov. 13, 1982–1997 (2023).
European Medicines Agency. Guideline on the Quality, Non-Clinical and Clinical Aspects of Gene Therapy Medicinal Products (European Medicines Agency, 2018).
Salmikangas, P., Carlsson, B., Klumb, C., Reimer, T. & Thirstrup, S. Potency testing of cell and gene therapy products. Front. Med. 10, 1190016 (2023).
Dias, J., Cadiñanos-Garai, A. & Roddie, C. In Potency Assays for Advanced Stem Cell Therapy Medicinal Products (ed. Burns, J. S.) 117–137 (Springer International Publishing, 2023).
Lu, R. et al. iSECRETE: Integrating Microfluidics and DNA Proximity Amplification for Synchronous Single-Cell Activation and IFN-γ Secretion Profiling. Adv. Sci. 11, 2309920 (2024).
Kiesgen, S., Messinger, J. C., Chintala, N. K., Tano, Z. & Adusumilli, P. S. Comparative analysis of assays to measure CAR T-cell-mediated cytotoxicity. Nat. Protoc. 16, 1331–1342 (2021).
Di Carlo, D. A mechanical biomarker of cell state in medicine. J. Lab. Autom. 17, 32–42 (2012).
Zeming, K. K., Salafi, T., Chen, C.-H. & Zhang, Y. Asymmetrical deterministic lateral displacement gaps for dual functions of enhanced separation and throughput of red blood cells. Sci. Rep. 6, 22934 (2016).
Chen, M., Phillips, Z. F. & Waller, L. Quantitative differential phase contrast (DPC) microscopy with computational aberration correction. Opt. Express 26, 32888–32899 (2018).
Lee, E. et al. Multimodal Microscopy for Label-free Monitoring of Mesenchymal Stromal Cells. In Frontiers in Optics / Laser Science (eds Mazzali, B. Lee, C. Mazzali, K. Corwin, and R. Jason Jones) OSA Technical Digest FW7E.8 (Optica Publishing Group 2020).
Dull, T. et al. A third-generation lentivirus vector with a conditional packaging system. J. Virol. 72, 8463–8471 (1998).
Gordon, K. S. et al. Screening for CD19-specific chimaeric antigen receptors with enhanced signalling via a barcoded library of intracellular domains. Nat. Biomed. Eng. 6, 855–866 (2022).
Acknowledgements
This research is supported by Agency for Science, Technology And Research (A*STAR) under the Industry Alignment Fund - Pre-positioning Programme (IAF-PP) grant: “Assembling Screening, Productions, High-throughput Analytics, and Lentiviral Targeting for T cells (ASPHALT) (H24J4a0031)” and National Research Foundation, Prime Minister’s Office, Singapore, under its Campus for Research Excellence and Technological Enterprise (CREATE) program through the Singapore-MIT Alliance for Research and Technology (SMART): Critical Analytics for Manufacturing Personalized-Medicine (CAMP) Inter-Disciplinary Research Group.
Author information
Authors and Affiliations
Contributions
K.K.Z., J.H. designed the CTM assay chip. K.Y.Q. and K.K.Z. planned and performed the CTM assay experiments. K.K.Z. and C.R.G. assembled the optics and mechanical parts for the CTM assay. E.L. developed and performed the phase contrast imaging. K.K.Z. develops the data processing pipeline and GUI to operate and process the CTM assay. W.X.S., K.C., D.B.L.T., F.K. and K.Y.Q. performed the T cell isolation, transduction, CAR T manufacturing, potency assay, and flow cytometry data. F.L.W.I.L., M.S.-F.S., and S.Y.S. provided patient samples. K.K.Z., W.X.S., K.C., M.B., and J.H. wrote and edited the manuscript. All authors reviewed and approved the manuscript prior to submission.
Corresponding authors
Ethics declarations
Competing interests
M.E.B. is a founder, consultant, and equity holder of Kelonia Therapeutics and Abata Therapeutics. J.H. and K.K.Z. have filed a patent (pending) WO/2021/141539 based on the designs of the biophysical microfluidic assay. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maté Biro, Weiqiang Chen, and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeming, K.K., Quek, K.Y., Sin, WX. et al. Cell trajectory modulation: rapid microfluidic biophysical profiling of CAR T cell functional phenotypes. Nat Commun 16, 4775 (2025). https://doi.org/10.1038/s41467-025-59789-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41467-025-59789-w